Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Enhancing the stability of silver nanoparticles in biomimetic environments through biomembrane hybridization
Tatsuhiro
Yokoyama
*ab,
Hiroyuki
Harada
a,
Shin-ichi
Sawada
d,
Kazunari
Akiyoshi
c,
Hikaru
Takaya
e,
Ryosuke
Mizuta
b and
Yoshihiro
Sasaki
 b
b
aDepartment of Oral and Maxillofacial Surgical Oncology, Graduate School of Medical and Dental Sciences, Institute of Science Tokyo, 1-5-45 Yushima, Bunkyo-ku, Tokyo, 113-8510, Japan
bDepartment of Polymer Chemistry, Graduate School of Engineering, Kyoto University, Kyotodaigaku-katura, Nishikyo-ku, Kyoto 615-8510, Japan
cGraduate School of Medicine, Kyoto University, Yoshida-Konoe-cho, Sakyo-ku, Kyoto, 606-8501, Japan
dcSIMVa, Chiba University, Chuo-ku Inohana, Chiba, 260-8670, Japan
eFaculty of Life and Environmental Sciences, Teikyo University of Science, Adachi-ku, Tokyo, 120-0045, Japan
First published on 5th January 2026
Abstract
Silver nanoparticles (AgNPs) have unique properties that make them valuable for biomedical applications. However, their instability in vivo has limited their use. Hybridizing AgNPs with organic materials is an effective way to address this issue. Biomembranes, as natural organic materials, offer stability in vivo and have various functions that are beneficial for biomedical applications. Despite these advantages, biomembrane hybridization remains challenging, making it one of the factors that hinder the biomedical application of biomembrane hybrid nanoparticles. In addition, the hybridization of biomembranes with AgNPs remains an underexplored area of research. In this study, we apply our previously reported floating cell layer penetration method to AgNPs, enabling the preparation of biomembrane hybrid AgNPs (hybrid) without membrane isolation or complex processing. Characterization of the resulting hybrid revealed changes in particle size, zeta potential, UV-vis spectra, and transmission electron microscopy (TEM) images. In addition, quantification of phospholipids confirmed that AgNPs were hybridized with sufficient phospholipids for coating, and lipidomics analysis showed that those phospholipids were reported in donor cells. These results indicate successful hybridization of AgNPs with biomembrane derived from source cells. In addition, biomembrane hybridization was suggested to improve the colloidal stability of AgNPs in in vivo-mimetic environments containing various ions or biomolecules. In summary, we constructed biomembrane hybrid AgNPs using a simple and reproducible approach, highlighting their potential for biomedical applications.
Introduction
Silver nanoparticles (AgNPs) have unique chemical and physical properties, making them valuable for various applications. Consequently, AgNPs have been extensively studied in multiple fields, including biomedical research. For instance, Khatoon et al. reported the antibacterial and antifungal activities of AgNPs.1 AgNPs have also been shown to exhibit antitumor activity against various types of cancer, and antiviral activity against certain viruses.2,3 Owing to these properties, AgNPs have been investigated both in vitro and in vivo for biotechnology applications. For example, AgNPs have been incorporated into scaffolding materials for regenerative medicine, with the antimicrobial activity of the materials being an important contributor to good tissue regeneration. Honda et al. reported that combining AgNPs with hydroxyapatite resulted in antimicrobial and biocompatible scaffolding materials for bone tissue regeneration.4 In addition, Takallu et al. developed a collagen hydrogel containing AgNPs for periodontal tissue regeneration.5 AgNPs are also useful in diagnosis. For example, AgNPs can function as a radiocontrast agent for the diagnosis of various diseases. Zou et al. developed AgNP-based radiocontrast agents that demonstrated significant contrast enhancement in micro-CT scanning.6 Therefore, AgNPs are useful for both diagnosis and treatment, and have attracted attention in numerous areas of medical research, including cancer treatment.7Despite their potential, several challenges should be addressed if AgNPs are to be used for biomedical applications. For example, AgNPs are cytotoxic to various types of cells.8 Additionally, AgNPs are cleared by immune cells and can induce immunotoxicity.9 Another significant challenge is their low stability in in vivo environments owing to interactions with various ions and biomolecules. For instance, chloride ions induce the formation of AgCl on AgNPs, leading to aggregation.10 Additionally, AgNPs release silver ions as a result of the ion content or pH of the surrounding environment, which could be a factor contributing to cytotoxicity.8,11 Proteins—as one of the major biomolecule groups—also interact with AgNPs forming a layer known as a “protein corona”. For a short time, the protein corona is reversible. However, over time, it becomes more stable and irreversible, ultimately leading to nanoparticle clearance by the immune system.12
To address these challenges and enhance the functionality of AgNPs, hybridization of AgNPs with organic materials has been explored. For example, Abdellatif et al. reported that synthesizing and coating AgNPs with cellulose-based polymer improved the stability of AgNPs in aqueous solutions without affecting the antimicrobial activity.13 In addition, Cheng et al. developed pH-triggered polymer hybrid silver nanoparticles that showed remarkable efficacy against various microbial diseases.14 Hybridizing AgNPs with lipids or peptides could also be effective methods for improving the cytotoxicity of AgNPs.15,16
Biomembrane hybridization has been reported to enhance the stability of nanoparticles in in vivo conditions. For example, several studies have shown that cell membrane hybridization prevents protein corona formation around the nanoparticles.17–19 As a result, nanoparticle clearance by immune cells could be prevented. In addition, the functions of membrane proteins could be conferred to nanoparticles, which would extend the in vivo persistence of the particles.20 For instance, nanoparticles coated with red blood cell membrane, with self-recognized proteins, exhibited a prolonged circulation time in vivo without immune cell uptake.21 Therefore, biomembranes and membrane proteins are beneficial for enhancing the in vivo stability of nanoparticles. In addition, their other functions, for example targeting ability—one of the notable functions of membrane proteins—are attractive for more accurate disease diagnosis and enhancement of therapeutic efficacy.22–26 Despite the advantages, direct hybridization of nanoparticles with the biomembrane remains difficult.27 In addition to this challenge, there are limited reports of AgNPs hybridized with biomembrane, and the appropriate method for hybridizing biomembrane with AgNPs has not yet been established.28 Therefore, devising an easy and reproducible method to fabricate biomembrane hybrid AgNPs, and precisely evaluating the effect of the hybridization, are crucial for expanding their biomedical potential.
We previously reported a floating cell layer penetration method, which enables direct coating of nanoparticles with biomembranes without the need for complex membrane isolation steps.29 This method utilizes density gradient centrifugation to form cell layers at a liquid interface. Nanoparticles are then passed through the layers via centrifugation, allowing them to become hybridized with the biomembranes in a single step. The key advantage of this approach is the significant simplification of the process compared with conventional methods that require prior membrane extraction. This study applies this established method to AgNPs for the first time. Our primary objectives were to construct biomembrane-coated AgNPs and to investigate whether the coating improves their colloidal stability in environments mimicking in vivo conditions. To evaluate the properties of the resulting biomembrane hybrid AgNPs (hybrid), we conducted particle size measurement, zeta potential analysis, UV-vis spectroscopy, phospholipid quantification on AgNPs, and lipidomic analysis of the biomembrane. These analyses confirmed the successful hybridization of AgNPs with biomembrane using a straightforward method. Furthermore, lipidomic profiling suggested that the hybrid nanoparticles incorporated biomembrane components derived from donor cells. In addition, biomembrane hybridization was suggested to improve the colloidal stability of AgNPs in biomimetic environments containing electrolytes and biomolecules. Although biomembrane hybridization is expected to enhance nanoparticle stability in vivo, its application to AgNPs has not been widely reported, leaving this approach largely unexplored. This study extends the promising strategy to AgNPs.
Experimental
Materials
Silver nanoparticles were purchased from DOWA Electronics (Tokyo, Japan). Trisodium citrate dihydrate was purchased from FUJIFILM Wako Pure Chemical Corporation (Osaka, Japan). Poly vinylidene di-fluoride (PVDF) filters (0.45 μm) were purchased from Merck Millipore Ltd (Belize, USA). EM Stainer, a negative stain for electron microscopy, was purchased from Nisshin EM (Tokyo, Japan). Tripan Blue Stain 0.4% was purchased from Aligned Genetics (Gyeonggi-do, South Korea). Optiprep™ (60% iodixanol solution) was purchased from Cosmo Bio Co., Ltd. (Tokyo, Japan). CellBrite™ Green was purchased from Cosmo Bio Co., Ltd. (Tokyo, Japan) and Methanol was purchased from FUJIFILM Wako Pure Chemical Corporation (Osaka, Japan). The mouse macrophage-like cell line Raw264.7 was purchased from Japanese Collection of Research Bioresources Cell Bank (JCRB Cell Bank).Preparation of AgNP dispersion liquid
A mixture of pure AgNPs (1 mg mL−1) and 2 mM citric acid in MilliQ (pH = 7.3) was sonicated in a sonication homogenizer (Central scientific commerce, Tokyo, Japan) for 30 min. After sonication, the AgNPs solution was filtrated using a 0.45 μm PVDF filter.Cell culture
RAW264.7 cells were cultured in Dulbecco's modified Eagle medium (DMEM) (Thermo Scientific, MA, USA) containing 10% fetal bovine serum (FBS) and 1% Gibco antibiotic–antimycotic (AA) (Thermo Scientific, MA, USA). Cell number and availability were measured using tripan-blue staining with a Luna™ fl dual fluorescence cell counter (Aligned Genetics).Preparation of floating cell layer
RAW264.7 cells were suspended in 500 μL of 30% iodixanol solution. Ultracentrifuge tubes were filled with 1.5 mL of 50 mM HEPES/KOH buffer (pH = 7.4) and the cell suspension was stacked underneath. Centrifugation at 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g was then performed for 5 min at 4 °C.
000×g was then performed for 5 min at 4 °C.
Fluorescent staining of RAW264.7
RAW264.7 cells (1 × 107) were suspended in PBS, and CellBrite™ Green was added according to the product protocol. Samples were then incubated for 30 min. To remove the free CellBrite, washing was carried out three times.Nanoparticle penetration and purification
AgNPs (0.2 mg) were added to the top of the prepared floating cell layer and centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g for 10 min. The supernatant was removed and the particles were dispersed in HEPES/KOH buffer using a bath sonicator at 28 KHz for 3 min. After redispersion, the tubes were centrifuged at 10
000×g for 10 min. The supernatant was removed and the particles were dispersed in HEPES/KOH buffer using a bath sonicator at 28 KHz for 3 min. After redispersion, the tubes were centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g for 5 min. The supernatant was removed and the same volume of HEPES/KOH buffer was added three times, and then the particles were redispersed. Finally, the tube was sonicated at 28 KHz for 3 min to disperse the hybrid nanoparticles.
000×g for 5 min. The supernatant was removed and the same volume of HEPES/KOH buffer was added three times, and then the particles were redispersed. Finally, the tube was sonicated at 28 KHz for 3 min to disperse the hybrid nanoparticles.
Evaluation of biomembrane hybrid nanoparticles
The Dhy, polydispersity index, and ζ potential of the prepared particles were measured using a Zetasizer Nano ZS (Malvern Instruments, Worcestershire, UK). The UV-vis spectra were obtained using a Spectrophotometer U-3900H (HITACHI, Japan). A 10 μL dispersion of biomembrane hybrid nanoparticles was cast of an ion sputter plasma-treated elastic carbon grid and allowed to stand for 10 min. The particles were then negatively stained with EM Stainer, diluted five times in 50 mM HEPES/KOH buffer for 30 min, and the morphology was observed using an HT7700 transmission electron microscope (HITACHI, Japan).Quantification of the biomembrane on hybrid nanoparticles
The amount of total phospholipid derived from the biomembranes was determined using a Phospholipids C Test Wako Kit (FUJIFILM Wako Pure Chemical Corporation, Osaka, Japan). AgNPs and hybrid were prepared at a concentration of 0.05 mg mL−1 based on UV-vis spectra. The sample (100 μL) was then centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g for 5 min at 4 °C. After removing 90 μL of the supernatant, the same volume of 25% methanol diluted with the buffer from the kit was added. The sample was then sonicated at 28 KHz for 1 min and centrifuged at 10
000×g for 5 min at 4 °C. After removing 90 μL of the supernatant, the same volume of 25% methanol diluted with the buffer from the kit was added. The sample was then sonicated at 28 KHz for 1 min and centrifuged at 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g for 5 min at 4 °C. The supernatant was used for measurement. The color development solution was prepared using 10 mg mL−1 of the color reagent included in the kit. Sample (80 μL) and color development solution (40 μL) were incubated at 37 °C in the dark. Then, the sample was measured using UV-vis spectrometry. The phospholipid standard solution included in the kit was used to create a calibration curve based on absorbance at 590 nm.
000×g for 5 min at 4 °C. The supernatant was used for measurement. The color development solution was prepared using 10 mg mL−1 of the color reagent included in the kit. Sample (80 μL) and color development solution (40 μL) were incubated at 37 °C in the dark. Then, the sample was measured using UV-vis spectrometry. The phospholipid standard solution included in the kit was used to create a calibration curve based on absorbance at 590 nm.
Lipid extraction for lipidomics analysis
The lipid samples (100 μL each) were extracted using a chloroform-free Lipid Extraction Kit (Cell Biolabs, California, USA, Catalog #STA-612) according to the protocol (https://www.cellbiolabs.com/sites/default/files/STA-612-lipid-extraction-kit.pdf). Briefly, each 100 μL particle suspension was mixed with 500 μL of lipid extraction reagent A and vortexed vigorously for 10 min. Sequentially, 250 μL of lipid extraction reagent B was added twice (for a total of 500 μL B), followed by 5 min of vortex mixing with each addition. Next, 500 μL of lipid extraction reagent C was added to the mixture, which was again vortexed for 5 min. The sample was then centrifuged at 1000×g for 5 min to separate the phases, and the upper organic layer (containing the lipids) was carefully transferred to a fresh tube. The remaining aqueous phase was re-extracted twice by adding fresh volumes of the organic phase reagent: first, 530 μL of reagent B (vortex 5 min, centrifuge 1000×g for 5 min), pooling the resulting organic layer with the first extract; then, 420 μL of reagent B (vortex 5 min, centrifuge again), pooling this final organic layer with the previous extracts. The combined organic extract was left uncapped in a vacuum concentrator (or in a 37 °C dry incubator) until completely dry. The resulting lipid residue was finally diluted to 2.0 mL with the initial mobile phase solvents for LC–MS/MS analysis and gently vortexed to prepare a homogenous solution ready for the subsequent LC–MS/MS lipidomics analysis.Lipidomic analysis of biomembrane hybrid AgNPs
In this study, lipid species are abbreviated by their class (e.g., PC for phosphatidylcholine) and annotated as (total number of carbon atoms![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) number of double bonds) in the fatty acyl chains, e.g., PC (36
number of double bonds) in the fatty acyl chains, e.g., PC (36![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2). Lipidomic analyses were performed on a Shimadzu triple quadrupole LC–MS/MS system consisting of a Nexera UHPLC coupled to an LCMS-8045 mass spectrometer (https://www.shimadzu.com/an/sites/shimadzu.com.an/files/pim/pim_document_file/applications/application_note/13968/an_c237-en.pdf). The dried lipid extracts prepared using the methods described above were dissolved in 2.0 mL of acetonitrile/2-propanol (80
2). Lipidomic analyses were performed on a Shimadzu triple quadrupole LC–MS/MS system consisting of a Nexera UHPLC coupled to an LCMS-8045 mass spectrometer (https://www.shimadzu.com/an/sites/shimadzu.com.an/files/pim/pim_document_file/applications/application_note/13968/an_c237-en.pdf). The dried lipid extracts prepared using the methods described above were dissolved in 2.0 mL of acetonitrile/2-propanol (80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 (v/v)) and spiked with 10 μL of PC (17
20 (v/v)) and spiked with 10 μL of PC (17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 0/20
0/20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4) (1 μg mL−1 in acetonitrile) for phospholipid analysis as an internal standard prior to injection. Chromatographic separation was carried out using a C8 reversed-phase column (Phenomenex Kinetex C8, 2.1 mm i.d. × 150 mm length, 2.6 μm) maintained at 45 °C for phospholipid analysis. The mobile phases comprised 20 mM HCO2NH4 in H2O (A) and acetonitrile/2-propanol (50
4) (1 μg mL−1 in acetonitrile) for phospholipid analysis as an internal standard prior to injection. Chromatographic separation was carried out using a C8 reversed-phase column (Phenomenex Kinetex C8, 2.1 mm i.d. × 150 mm length, 2.6 μm) maintained at 45 °C for phospholipid analysis. The mobile phases comprised 20 mM HCO2NH4 in H2O (A) and acetonitrile/2-propanol (50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 (v/v)) (B) for phospholipid analysis. The HPLC conditions, such as gradient program and flow rate, were applied as specified in the Shimadzu MRM library method manual. Mass spectrometric detection was performed using electrospray ionization in both positive and negative modes, with multiple reaction monitoring (MRM) acquisition for targeted lipid species. The Shimadzu MRM library method for phospholipid profiling was used, containing predefined MRM transitions for the major lipid classes. The phospholipid classes monitored included phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylglycerol (PG), phosphatidylinositol (PI), phosphatidylserine (PS), and sphingomyelin (SM), together with their lysophospholipid counterparts (LPC, LPE, LPG, LPI, and LPS). Both diacyl and monoacyl (lyso-) species were quantified using class-specific MRM transitions, with detectable acyl-chain lengths spanning C14–C22 and unsaturation levels from 0 to 6 double bonds as defined by the Shimadzu phospholipid MRM library (https://www.shimadzu.com/an/products/liquid-chromatograph-mass-spectrometry/lc-ms-software/lcmsms-mrm-library-for-phospholipid-profiling/index.html). The MRM transitions were chosen to capture class-specific fragment ions (e.g. headgroup ions for phospholipid classes) as well as fatty acid fragment ions, enabling identification of the class and acyl composition of each lipid. Data acquisition and analysis were conducted with Shimadzu LabSolutions LCMS software. Complete results of lipidomic analysis are provided in SI.
50 (v/v)) (B) for phospholipid analysis. The HPLC conditions, such as gradient program and flow rate, were applied as specified in the Shimadzu MRM library method manual. Mass spectrometric detection was performed using electrospray ionization in both positive and negative modes, with multiple reaction monitoring (MRM) acquisition for targeted lipid species. The Shimadzu MRM library method for phospholipid profiling was used, containing predefined MRM transitions for the major lipid classes. The phospholipid classes monitored included phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylglycerol (PG), phosphatidylinositol (PI), phosphatidylserine (PS), and sphingomyelin (SM), together with their lysophospholipid counterparts (LPC, LPE, LPG, LPI, and LPS). Both diacyl and monoacyl (lyso-) species were quantified using class-specific MRM transitions, with detectable acyl-chain lengths spanning C14–C22 and unsaturation levels from 0 to 6 double bonds as defined by the Shimadzu phospholipid MRM library (https://www.shimadzu.com/an/products/liquid-chromatograph-mass-spectrometry/lc-ms-software/lcmsms-mrm-library-for-phospholipid-profiling/index.html). The MRM transitions were chosen to capture class-specific fragment ions (e.g. headgroup ions for phospholipid classes) as well as fatty acid fragment ions, enabling identification of the class and acyl composition of each lipid. Data acquisition and analysis were conducted with Shimadzu LabSolutions LCMS software. Complete results of lipidomic analysis are provided in SI.
Evaluation of biomembrane hybrid nanoparticle colloidal stability in in vivo-mimetic conditions
AgNPs before and after biomembrane hybridization (100 μL, 0.03 mg mL−1 (buffer: 50 mM HEPES/KOH)) was mixed with 900 μL PBS (Gibco Thermo fisher scientific) or DMEM containing 10% FBS and 1% AA at 25 °C in the dark. After mixing, we measured the particle size with DLS and observed the changes over a period of time.Results and discussion
Preparation of silver nanoparticle (AgNP) dispersion
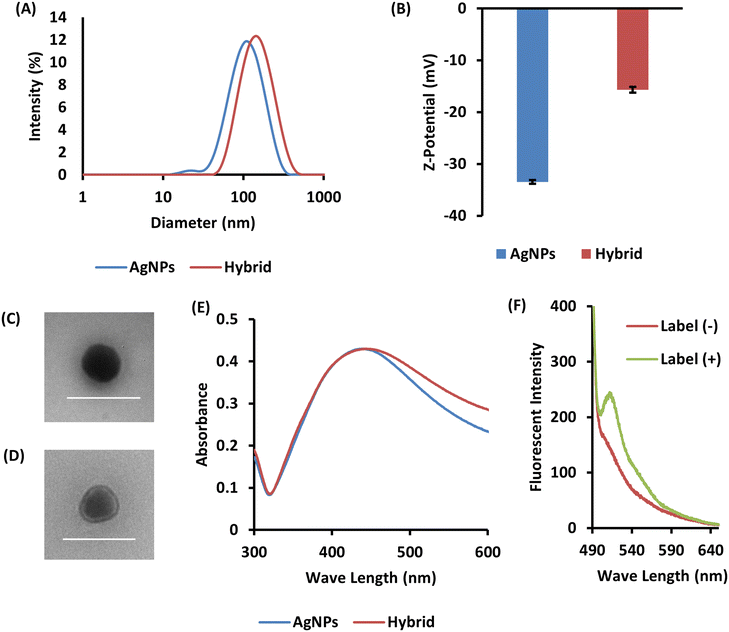
In this study, we used non-coated solid AgNPs. We used sodium citrate as a dispersant, which provides a citrate ion coating.30 We first mixed AgNPs with a 2 mM sodium citrate solution and sonicated the mixture for 30 min to disperse the AgNPs in the solution. Then, the solution was filtered with a 0.45 μm pore size filter. We characterized the AgNPs dispersed in the solution by dynamic light scattering (DLS) measurements and UV-vis spectroscopy. The DLS measurements showed that the average hydrodynamic diameter (Dhy) of AgNPs was 93 ± 0.3 nm with a slightly broad polydispersity index (PdI) of 0.20 ± 0.0 (Fig. 2A). The UV-vis spectra showed a maximum absorbance wavelength (λmax) of 432 nm, which is the characteristic absorption wavelength of AgNPs < 100 nm (Fig. 2E).30 Thus, the UV-vis spectra were consistent with the DLS results, and we confirmed the preparation of an AgNP dispersion.Hybridization of nanoparticles and biomembrane by floating cell layer penetration
We hybridized AgNPs with biomembrane using a previously described procedure (Fig. 1).29 This approach enables direct utilization of live cells as a membrane source, minimizing the need for membrane isolation. In this study, approximately 80% of cells remained viable after the process, indicating that the method had limited impact on cell viability while enabling AgNPs–biomembrane hybridization. Furthermore, the processes were performed easily with centrifugation and ultrasonication.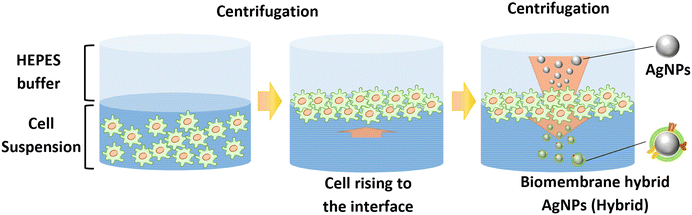 | ||
| Fig. 1 Schematic illustration of preparation of AgNPs hybridized with biomembrane. The sizes of the cell and AgNPs are shown schematically for clarity and are not to scale. | ||
Characterization of the biomembrane hybrid AgNPs
The particles were then redispersed in 50 mM HEPES/KOH buffer and their Dhy was measured using DLS (Fig. 2A). The Dhy of purified AgNPs was greater than that of the particles before penetration through the floating cell layers, indicating that the Dhy increased upon complexation with the biomembrane. To evaluate the biomembrane hybridization, we analyzed the surface properties of the nanoparticles. We determined the zeta (ζ) potential of hybrid and AgNPs with the same preparation process without floating cell layers (Fig. 2B). The zeta potential of AgNPs, which were strongly negative charged, increased significantly when AgNPs were passed through the floating cell layers. Changes in zeta potential are widely used as evidence of the presence of biomembrane on nanoparticles.31 Therefore, the zeta potential increase indicated that the biomembrane had hybridized with the AgNPs. The particles were observed using transmission electron microscopy (TEM) to determine their shape. Negative staining was used for biomembrane observation. After the hybridization process, a membrane-like structure was observed around the AgNPs (Fig. 2C and D). These images indicate that the biomembrane layer was on the surface of the AgNPs after the hybridization process. Furthermore, UV-vis spectra showed that the absorbance increased toward longer wavelengths compared with AgNPs prepared using the same process without cell layers (Fig. 2E). This observation can be explained by localized surface plasmon resonance (LSPR). Plasmonic metal nanoparticles, for example, AgNPs or gold nanoparticles, have an intrinsic resonance wavelength range, and LSPR is responsive to alterations to the surface coating on the particles, including biomembrane hybridization.15,32 Therefore, the UV-vis spectra indicated that the cellular components including biomembrane were hybridized with the AgNPs. In addition, hybridization was performed using cells with fluorescently labeled membranes, and fluorescence was observed on the AgNPs (Fig. 2F). This result indicated that biomembranes including cell membrane were hybridized on the AgNPs.Quantification of biomembrane hybridized with AgNPs
We estimated the amount of biomembrane coating by quantifying the phospholipids using an enzymatic colorimetric method. Approximately 170 ± 21 μg of phospholipids were associated with 1 mg of AgNPs. To evaluate this coating amount, 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC), a representative phospholipid, was used as a reference. DOPC forms bilayers similar to biomembranes and is widely used for such comparisons.31 The thickness of a lipid bilayer is typically 5 nm.33 On the basis of geometric and molecular volume calculations, 4.4 × 10−8 mol of DOPC would be required to fully cover 1 mg of AgNPs (93 nm in diameter) with a monolayer. In our samples, 2.2 ± 0.3 × 10−7 mol of phospholipid was detected per mg of AgNPs, able effective coating.Lipidomic analysis of the biomembrane on the AgNPs
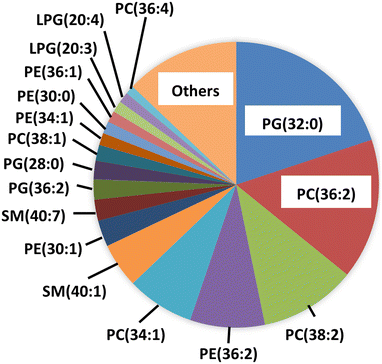
To characterize the biomembrane on hybrid, mass spectrometry of the constituent lipids was performed. The analysis of the lipids extracted from hybrid was conducted using targeted multiple reaction monitoring (MRM) on a triple quadrupole LC–MS/MS system. Then, lipidomic analysis quantified and identified the 422 lipid classes, of which specific species were detected (Fig. 3). Approximately 90% of the detected lipids corresponded to previously reported species in RAW264.7 cells.34–36 This result indicates that AgNPs were hybridized with donor cell-derived lipids, which were found in various cellular components, including biomembranes.In our previous study, we performed the biomembrane hybridization onto silica nanoparticles.29 Since oxide surfaces such as silica typically present hydrophilic functionalities and hydration layers, interfacial behaviors for lipid assembly can differ from those on metallic surfaces. In particular, spontaneous vesicle rupture/fusion to form supported lipid bilayers is strongly surface-condition dependent and can be difficult on certain metal substrates.37
Importantly, our method does not rely on spontaneous vesicle rupture/fusion on the particle surface; rather, cell-derived membrane fragments are physically recruited and transferred onto nanoparticles during the penetration process. In this study, we employed citrate capping to enhance the colloidal stability of AgNPs and to provide a hydrophilic, negatively charged interface. Citrate-based surface modification has been reported not to hinder lipid–particle interactions, and the construction of lipid layers on silver surfaces has also been reported.38,39 Under these conditions, we achieved biomembrane-coated citrate-capped AgNPs with coating characteristics supported by DLS, ζ-potential and TEM results (Fig. 2A–D).
Colloidal stability of the biomembrane hybrid AgNPs in an in vivo-mimetic environment
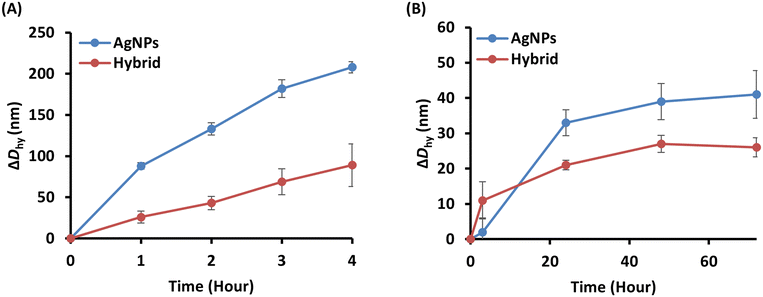
The colloidal stability of AgNPs in vivo is important for biomedical applications. Electrolytes and biomolecules are factors that are known to contribute to colloidal instability of AgNPs. We monitored the increase in particle size of hybrid, which is common metric of colloidal stability, under two different conditions, and observed a smaller size growth compared with unmodified AgNPs, suggesting the potential improvement in colloidal stability. First, we investigated the effect of electrolytes on AgNPs and hybrid. The nanoparticles were mixed with phosphate-buffered saline (PBS) and the particle size was monitored over time (Fig. 4A). The aggregation rate of hybrid was slower than that of AgNPs, suggesting that biomembrane hybridization may contribute to improved colloidal stability of AgNPs in an environment rich in electrolytes. These results are attributed to the coating layer around the AgNPs, which prevented interaction between the nanoparticle surface and the reagents and ions present.40 The layer thickness and density are the main factors affecting the reactions. In this study, the biomembrane layer, a thick layer containing high molecular weight compounds such as proteins, was deposited on the surface of AgNPs. As a result, biomembrane hybridization may protect AgNPs from ions, leading to decrease the reaction rate. Next, we evaluated the influence of biomolecules on AgNPs. Both AgNPs and hybrid were mixed with Dulbecco's modified Eagle medium (DMEM) containing 10% fetal bovine serum (FBS). The Dhy was measured over time. The particle size of hybrid was increased, but finally, ΔDhy of hybrid was smaller than that of unmodified AgNPs (Fig. 4B). Hybrid exhibited a similar degree of size change compared with unmodified AgNPs, suggesting that the biomembrane coating did not markedly compromise colloidal stability in biomolecule-rich environments. In summary, biomembrane hybridization implied enhanced colloidal stability of AgNPs under electrolyte-rich conditions, while preserving colloidal stability in biomolecule-rich media. These advantages are useful to deliver AgNPs to target cells, suggesting the potential applicability of this approach in biomedical research, for example, using AgNPs as drug delivery carrier.Conclusions
In this study, we demonstrated that our established method for direct biomembrane hybridization can be applied to AgNPs, suggesting the potential enhancement of their colloidal stability in vivo mimicking conditions. Colloidal stability is one of the most important factors for biomedical applications of AgNPs. Therefore, our method provides a straightforward strategy to prepare biomembrane-coated AgNPs for biomedical research and highlights their potential applicability in the clinic.Author contributions
The manuscript was written through the contributions of all authors. All authors have approved the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
All data supporting the findings of this study are available within the article and its supporting information (SI) file.Supplementary information is available. See DOI: https://doi.org/10.1039/d5lf00329f.
Acknowledgements
This research was supported by grants from the Japan Society for the Promotion of Science (JSPS) Grants-in-Aid for Scientific Research (Grant number JP16H06313, JP22H00585 (KA), JP21K18324, JP22H02199, JP21H04954, JP18H02089, JP22H00585, JP23H04934, JP23K23466 and JP24K21340 (YS)). We thank Sarah Dodds, PhD, from Edanz (https://jp.edanz.com/ac) for editing a draft of this manuscript.References
- U. T. Khatoon, G. V. S. N. Rao, K. M. Mohan, A. Ramanaviciene and A. Ramanavicius, Vacuum, 2017, 146, 259–265 Search PubMed.
- R. R. Miranda, I. Sampaio and V. Zucolotto, Colloids Surf., B, 2022, 210, 112254 CrossRef CAS PubMed.
- K. Naumenko, S. Zahorodnia, C. V. Pop and N. Rizun, J. Virus Erad., 2023, 9, 100330 CrossRef CAS PubMed.
- M. Honda, Y. Kawanobe, K. Ishii, T. Konishi, M. Mizumoto, N. Kanzawa, M. Matsumoto and M. Aizawa, Mater. Sci. Eng., C, 2013, 33, 5008–5018 Search PubMed.
- S. Takallu, F. Kakian, A. Bazargani, H. Khorshidi and E. Mirzaei, Sci. Rep., 2024, 14, 7262 CrossRef CAS PubMed.
- J. Zou, M. Hannula, S. Misra, H. Feng, R. H. Labrador, A. S. Aula, J. Hyttinen and I. Pyykkö, NANO, 2015, 13, 5 Search PubMed.
- B. K. Kashyap, V. V. Singh, M. K. Solanki, A. Kumar, J. Ruokolainen and K. K. Kesari, ACS Omega, 2023, 8, 14290–14320 CrossRef CAS PubMed.
- M. Ahameda, M. S. Alsalhi and M. K. J. Siddiqui, Clin. Chim. Acta, 2010, 411, 1841–1848 CrossRef PubMed.
- J. Bi, C. Mo, S. Li, M. Huang, Y. Lin, P. Yuan, Z. Liu, B. Jia and S. Xu, Biomater. Sci., 2023, 11, 4151–4183 RSC.
- A. M. E. Badawy, T. P. Luxton, R. G. Silva, K. G. Scheckel, T. M. Tolaymat and M. T. Suidan, Environ. Sci. Technol., 2010, 44, 1260–1266 CrossRef PubMed.
- W. Zhang, Y. Yao, N. Sullivan and Y. Chen, Environ. Sci. Technol., 2011, 45, 4422–4428 CrossRef CAS PubMed.
- E. Casals, T. Pfaller, A. Duschl, G. J. Oostingh and V. Puntes, ACS Nano, 2010, 4(7), 3623–3632 CrossRef CAS PubMed.
- A. A. H. Abdellatif, H. N. H. Alturki and H. M. Tawfeek, Sci. Rep., 2021, 11, 84 CrossRef CAS PubMed.
- X. Cheng, X. Pei, W. Xie, J. Chen, Y. Li, J. Wang, H. Gao and Q. Wan, Small, 2022, 18, 2200915 CrossRef CAS PubMed.
- T. J. Miesen, A. M. Engstrom, D. C. Frost, R. Ajjarapu, R. Ajjarapu, C. N. Lira and M. R. Mackiewicz, RSC Adv., 2020, 10, 15677–15693 RSC.
- L. Zhou, X. Zhao, M. Li, Y. Lu, C. Ai, C. Jiang, Y. Liu, Z. Pan and J. Shi, Appl. Microbiol. Biotechnol., 2021, 105, 3759–3770 Search PubMed.
- L. Rao, Q. Meng, L. Bu, B. Cai, Q. Huang, Z. Sun, W. Zhang, A. Li, S. Guo, W. Liu, T. Wang and X. Zhao, ACS Appl. Mater. Interfaces, 2017, 9, 2159–2168 CrossRef CAS PubMed.
- D. Zou, Z. Wuc, X. Yic, Y. Huid, G. Yang, Y. Liu, Tengjisi, H. Wangb, A. Brooks, H. Wanga, X. Liu, Z. P. Xu, M. S. Roberts, H. Gaog and C. Zhao, Proc. Natl. Acad. Sci. U. S. A., 2023, 120(1), e2214757120 CrossRef CAS PubMed.
- I. Ferreira-Faria, S. Yousefiasl, A. Macário-Soares, M. Pereira-Silva, D. Peixoto, H. Zafar, F. Raza, H. Faneca, F. Veiga, M. R. Hamblin, F. R. Tay, J. Gao, E. Sharifi, P. Makvandi and A. C. Paiva-Santos, J. Controlled Release, 2022, 351, 174–197 CrossRef CAS PubMed.
- W. Liu, M. Zou, S. Qin, Y. Cheng, Y. Ma, Y. Sun and X. Zhang, Adv. Funct. Mater., 2020, 30, 2003559 CrossRef CAS.
- Q. Pei, X. Hu, X. Zheng, S. Liu, Y. Li, X. Jing and Z. Xie, ACS Nano, 2018, 12, 1630 CrossRef CAS PubMed.
- H. Sun, J. Su, Q. Meng, Q. Yin, L. Chen, W. Gu, Z. Zhang, H. Yu, P. Zhang, S. Wang and Y. Li, Adv. Funct. Mater., 2017, 27, 1604300 Search PubMed.
- A. E. Boniakowski, A. S. Kimball, A. Joshi, M. Schaller, F. M. Davis, A. denDekker, A. T. Obi, B. B. Moore, S. L. Kunkel and K. A. Gallagher, Eur. J. Immunol., 2018, 48, 1445–1455 CrossRef CAS PubMed.
- S. Gordon, Cell, 2002, 111, 927–930 CrossRef CAS PubMed.
- L. Zou, Y. Zhang, N. Cheraga, O. D. Abodunrin, K. Qu, L. Qiao, Y. Ma, Y. Hang, N. Huang and L. Chen, Small, 2024, 20, 2304110 CrossRef CAS PubMed.
- J. An, X. Jiang, Q. Wu, Z. Zou, Y. Li, J. Sun, X. Liu, Y. Xiong, C. Wu, X. Mei and H. Tian, ACS Appl. Nano Mater., 2023, 6, 19073–19087 CrossRef CAS.
- L. Liu, X. Bai, M. Martikainen, A. Kårlund, M. Roponen, W. Xu, G. Hu, E. Tasciotti and V. Lehto, Nat. Commun., 2021, 12, 5726 Search PubMed.
- Q. Zhao, X. Y. Sun, B. Wu, Y. Shang, X. Huang, H. Dong, H. Liu, W. Chen, R. Gui and J. Li, Mater. Sci. Eng., C, 2021, 119, 111648 CrossRef CAS PubMed.
- R. Mizuta, E. Kanao, K. Ukyo, S. Kuwada, S. Sawada, Y. Ishihama, K. Akiyoshi and Y. Sasaki, Nano Lett., 2024, 24, 12907–12914 CAS.
- N. G. Bastús, F. Merkoçi, J. Piella and V. Puntes, Chem. Mater., 2014, 26, 2836–2846 CrossRef.
- L. Liu, D. Pan, S. Chen, M. Martikainen, A. Kårlund, J. Ke, H. Pulkkinen, H. Ruhanen, M. Roponen, R. Käkelä, W. Xu, J. Wang and V. Lehto, Nat. Commun., 2022, 13, 6181 Search PubMed.
- I. Srivastava, R. Xue, J. Jones, H. Rhee, K. Flatt, V. Gruev and S. Nie, ACS Nano, 2022, 16, 8051–8063 CrossRef CAS PubMed.
- W. Stillwell, An Introduction to Biological Membranes: Composition, Structure and Function, Elsevier, 2nd edn, 2016 Search PubMed.
- C. A. Rouzer, P. T. Ivanova, M. O. Byrne, S. B. Milne, L. J. Marnett and H. A. Brown, Biochemistry, 2006, 45, 14795–14808 Search PubMed.
- A. Y. Andreyev, E. Fahy, Z. Guan, S. Kelly, X. Li, J. G. McDonald, S. Milne, D. Myers, H. Park, A. Ryan, B. M. Thompson, E. Wang, Y. Zhao, H. A. Brown, A. H. Merrill, C. R. H. Raetz, D. W. Russell, S. Subramaniam and E. A. Dennis, J. Lipid Res., 2010, 51, 2785–2797 Search PubMed.
- M. Santos, T. Maurício, R. Domingues and P. Domingues, Arch. Biochem. Biophys., 2025, 768, 110384 CrossRef CAS PubMed.
- R. P. Richter, R. Bérat and A. R. Brisson, Langmuir, 2006, 22, 3497–3505 Search PubMed.
- F. Wang, D. E. Curry and J. Liu, Langmuir, 2015, 31, 13271–13274 Search PubMed.
- H. Zhu, J. Zhang, X. Dai, V. St, D. Mesias, H. Chi, C. Wang, C. S. Yeung, Q. Chen, W. Liu and J. Huang, Anal. Bioanal. Chem., 2023, 415, 3243–3253 CrossRef CAS PubMed.
- X. Li, J. J. Lenhart and H. W. Walker, Langmuir, 2012, 28, 1095–1104 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |