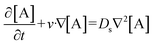Open Access Article
Open Access ArticleAn automated microfluidic system based on V-groove chip for rapid immunohistochemistry
Lu
Zhong
a,
Hang
Chen
b,
Hong-Lei
Chen
b,
Jun
Peng
c and
Zhi-Ling
Zhang
 *a
*a
aCollege of Chemistry and Molecular Sciences, Wuhan University, Wuhan 430072, P. R. China. E-mail: zlzhang@whu.edu.cn
bDepartment of Pathology, School of Basic Medical Sciences, Wuhan University, Wuhan 430071, P. R. China
cWuhan Pathologic Biotechnology Limited Company, Wuhan 430074, P. R. China
First published on 10th March 2026
Abstract
The demand for efficient detection of tumor biomarkers in clinical settings is growing. Traditional immunohistochemistry (IHC) is time-consuming and labor-intensive, making rapid IHC technologies essential for improving efficiency in pathological diagnosis. Microfluidic technology, characterized by miniaturization, high throughput, and automation, has emerged as one of the most promising approaches for advancing rapid tissue diagnostics. This study designed a fluid distribution channel with excellent uniformity and developed a microfluidic chip featuring an upper V-groove structure, based on a passive mixing strategy in microfluidic systems. This chip effectively enhances the mixing efficiency of antigens and antibodies without relying on external field-driven mechanisms. Furthermore, a novel, automated, and integrated microfluidic platform was constructed to achieve rapid, reliable, and automated immunohistochemical staining. Experimental results demonstrated that the immunohistochemical staining performance obtained with this chip is comparable to that of conventional methods, exhibiting excellent uniformity and reproducibility. The staining time for markers such as CK and Ki-67 in tissue samples can be reduced to 11 minutes, representing a 90% time saving compared to traditional methods.
Introduction
Tumor biomarkers are critical tools for achieving early screening, early diagnosis, and dynamic monitoring of tumors. Among these, IHC technology enables highly efficient in situ detection of these markers in tissue through specific antigen–antibody binding. As one of the gold standards in clinical pathology, IHC not only aids in distinguishing benign from malignant tumors, and tracing the origin of metastatic tumors, but also provides key molecular characteristics for accurate staging and personalized therapy. It plays an indispensable role in pathological diagnosis, treatment decision-making, and prognostic evaluation.1,2 It involves fixing thin sections of tumor tissue onto glass slides, followed by the application of various specific detection reagents such as antibodies targeting biomarkers of interest. These slides are incubated in a humidity-controlled chamber at constant temperature for a designated duration. Visualization is achieved through fluorescence or chromogenic bright-field microscopy techniques.3 IHC typically requires several hours to even days to complete, making it difficult to meet the clinical demand for timely diagnosis and treatment. In contrast, rapid IHC can accurately identify tumor molecular subtypes and therapeutic targets in a significantly shorter time-frame. This capability markedly reduces diagnostic waiting periods and accelerates therapeutic decision-making, thereby establishing rapid IHC as an indispensable technology for improving diagnostic efficiency and advancing precision medicine.4–6Traditional IHC technology faces inherent limitations. Under static incubation conditions, once the antibody solution is applied to the tissue sample surface, specific antigen–antibody binding occurs immediately at the solid–liquid interface, forming antigen–antibody complexes. The binding rate is governed by reaction kinetics and is positively correlated with the binding rate constant. As the reaction proceeds, the local antibody concentration at the interface decreases rapidly due to binding consumption, resulting in a localized depletion zone. When the interfacial antibody concentration becomes significantly lower than the bulk concentration, a concentration gradient is established, triggering a diffusion-dominated mass transfer process.7,8
According to diffusion kinetics theory, the thickness of the depletion zone δ varies with time as  , where D is the diffusion coefficient of the antibody and t is the incubation time. Under these conditions, the reaction rate is determined by the diffusive flux:
, where D is the diffusion coefficient of the antibody and t is the incubation time. Under these conditions, the reaction rate is determined by the diffusive flux: 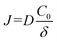 , where C0 is the bulk concentration of the antibody. The reaction rate is subsequently determined by the diffusion flux, which depends on the bulk antibody concentration C0 and δ.9 As the incubation time increases, the continuous growth of the diffusion layer thickness δ leads to a gradual decline in the reaction rate. In conventional immunoassays, the detection of a single biomarker typically requires 5 to 24 hours. This prolonged duration is primarily due to two limiting factors: first, the affinity constant of the antigen–antibody binding reactions (reaction-controlled phase); and second, the mass transfer efficiency of antibody molecules to the reaction interface (diffusion-controlled phase).10 These temporal limitations severely restrict the fulfillment of clinical rapid diagnostic needs, urgently requiring the optimization of reaction conditions or improvement of detection methods to enhance detection efficiency.
, where C0 is the bulk concentration of the antibody. The reaction rate is subsequently determined by the diffusion flux, which depends on the bulk antibody concentration C0 and δ.9 As the incubation time increases, the continuous growth of the diffusion layer thickness δ leads to a gradual decline in the reaction rate. In conventional immunoassays, the detection of a single biomarker typically requires 5 to 24 hours. This prolonged duration is primarily due to two limiting factors: first, the affinity constant of the antigen–antibody binding reactions (reaction-controlled phase); and second, the mass transfer efficiency of antibody molecules to the reaction interface (diffusion-controlled phase).10 These temporal limitations severely restrict the fulfillment of clinical rapid diagnostic needs, urgently requiring the optimization of reaction conditions or improvement of detection methods to enhance detection efficiency.
In response to the aforementioned challenges, current research primarily focuses on enhancing molecular recognition efficiency through antibody engineering (e.g., nanobodies, affinity optimization) to develop high-affinity antibody reagents,11 while studies dedicated to improving mass transfer efficiency to reduce the time required for antibodies to reach the reaction interface remain relatively scarce. Conventional techniques for accelerating mass transfer, such as mild agitation, are often limited by inconsistent performance and a lack of precise control.
Microfluidic chips, also known as lab-on-a-chip, possess a high surface area to volume ratio and enable the manipulation of fluids at the microscale.12,13 Microfluidic chips leverage high flow rates and micron-scale channel heights to physically constrain the growth of diffusion boundary layers. Active fluid driving enables proactive antibody transport and convective mass transfer, where the dimensionless Péclet number (Pe = U0L0/D), defined by characteristic flow velocity U0, characteristic length L0, and diffusion coefficient D, is far greater than 1. This indicates that the convective transport rate far exceeds the diffusion rate and dominates the overall transport process due to the direction of fluid motion.14 To enhance mass transfer efficiency, related studies have introduced microfluidic technology into IHC.15–19 Ciftlik et al.16 developed a rapid microfluidic tissue processor, that enables primary antibody incubation to be completed within 2 minutes, while also distinguishing ambiguous HER-2 expression in breast cancer. Similarly, Saška Brajkovic et al.15 established an ALK immunofluorescence technique based on a microfluidic tissue processor. When detecting formalin-fixed and paraffin-embedded (FFPE) samples of lung adenocarcinoma, primary antibody incubation reached signal saturation in just four minutes, with total staining requiring only 18 minutes. However, the aforementioned microfluidic immunostaining methods based on conventional straight microchannels still have inherent limitations in mass-transfer efficiency. Under low Reynolds number conditions, fluid flow within a microchannel is typically laminar, and mass exchange between fluid streams primarily relies on molecular diffusion in the direction perpendicular to the flow.20 To overcome laminar flow constraints, mechanisms that promote radial mixing must be introduced. There are two main approaches to enhance radial mixing in microfluidic chips: active mixing and passive mixing techniques.21 Active mixing technology utilizes external energy fields, such as acoustic22 or magnetic23 fields, to induce forced fluid mixing. For instance, Muaz S. Draz et al.18 attached a water-soluble adhesive to bond a piezoelectric actuator to the outer surface of a microfluidic staining system, thereby generating vortex-like acoustic streaming through actuation. This approach reduced the primary antibody incubation time to 1 minute for HER-2 and CK staining in breast cancer BT-474 cells. Passive mixing, on the other hand, employs specially designed microstructures within the channels (such as herringbone24 or chevron25 structures) to alter streamlines and induce secondary flows and vortices. This effectively disrupts the conventional diffusion boundary layer and substantially reduces the depletion layer thickness δ (δ ∼ D/u), where u represents the characteristic flow velocity,26 thereby significantly accelerating the antigen–antibody reaction rate at the solid–liquid interface.27 Designing chaotic advection structures within microchannels eliminates the reliance on external energy fields, offering notable advantages over active mixing technologies in terms of energy consumption, cost, and fabrication convenience.
This study developed an automated immunohistochemical staining system based on an integrated V-groove upper-structure microfluidic chip for rapid IHC staining. This platform incorporates a control system, fluidic delivery module, temperature regulation unit, and an on-chip reaction chamber, enabling fully automated execution of critical procedural steps, including antibody incubation, washing, and chromogenic development. Two types of fluid distribution channel architecture were designed and systematically evaluated to assess their impact on staining homogeneity. Concurrently, the relationship between flow rate and staining intensity was investigated by optimizing flow velocity to enhance reaction kinetics. The staining performance of our system was benchmarked against conventional manual IHC protocols by comparing signal intensity, staining uniformity, and experimental reproducibility.
Materials and methods
Microfluidic chip fabrication
The PDMS microfluidic device with a V-groove upper structure was fabricated using a two-step soft lithography process. A bare silicon wafer was spin-coated with SU-8 2050 photoresist to form a 50 μm-thick layer. After soft baking, the resist was exposed to ultraviolet (UV) light, and then post-exposure baked, to define the first-layer microchannel structure. A second spin-coating step, followed by alignment exposure, development, and hard baking, was then performed to create the master mold with the upper V-shaped microstructure. The mold was treated with trimethylchlorosilane (TMC) for 30 minutes to facilitate demolding. A PDMS prepolymer mixture (base-to-curing agent, 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 mass ratio) was poured onto the mold surface. After degassing, it was cured in an oven at 75 °C, and the solidified PDMS device was peeled from the master mold. SU-8 2050 was purchased from AZ Electronic Materials, polydimethylsiloxane (PDMS) from Momentive (USA), trimethylchlorosilane from Shanghai Lingfeng Chemical Reagent Co., Ltd., and SU-8 developer from Kayaku Advanced Materials (USA).
1 mass ratio) was poured onto the mold surface. After degassing, it was cured in an oven at 75 °C, and the solidified PDMS device was peeled from the master mold. SU-8 2050 was purchased from AZ Electronic Materials, polydimethylsiloxane (PDMS) from Momentive (USA), trimethylchlorosilane from Shanghai Lingfeng Chemical Reagent Co., Ltd., and SU-8 developer from Kayaku Advanced Materials (USA).
Tissue samples processing and immunostaining
The FFPE tissues were sectioned at 4–5 μm using a microtome (Leica, Germany). The sections were baked in a 65 °C oven for 60 minutes and then subjected to three rounds of dewaxing with a dewaxing agent, each lasting 8 minutes. Following dewaxing, the sections were hydrated through a graded alcohol series (95%, 85%, and 70%) for 5 minutes each. Antigen retrieval was performed using a pH 9.0 EDTA buffer under microwave irradiation: first heated at 100% power for 3 minutes, then at 30% power for 12 minutes. After cooling to room temperature, the sections were incubated with 3% hydrogen peroxide solution at room temperature for 10 minutes to block endogenous peroxidase activity. The tissue sections were then placed in a staining apparatus for sequential antibody incubation and chromogenic development. The paraffin-embedded tissues were obtained from Zhongnan Hospital of Wuhan University (this study was approved by the Institutional Review Board of Zhongnan Hospital). The dewaxing agent, absolute ethanol, primary antibodies, PBS buffer, and pH 9.0 EDTA antigen retrieval buffer were purchased from Guangzhou Anbiping Co., Ltd. The secondary antibodies and DAB (3,3′-diaminobenzidine) chromogenic reagent were obtained from Dako, Denmark.Image analysis
Tissue sections were imaged using a microscope (BX53, Olympus, Japan) equipped with a CCD camera (DP72, Olympus, Japan). Image analysis was performed using the open-source software ImageJ, released by the National Institutes of Health (NIH). The DAB staining channel was isolated using the Color Deconvolution plugin, and after background correction, the optical density (OD) of positive areas was calculated using the formula OD = log10(255/I), where I represents pixel intensity. The average optical density (AOD) was used to quantify the staining intensity. Additionally, H-score was calculated using inForm 2.6 software (Akoya Biosciences, USA). This scoring system converts the distribution and proportion of positively stained cells into a semi-quantitative value ranging from 0 to 300, providing a comprehensive assessment of protein expression levels. The H-score was calculated using the formula: H = ∑Pii, where Pi represents the percentage of positive cells exhibiting staining intensity i, and i denotes the staining intensity grade, typically classified into four levels: 0 (negative), 1 (weak), 2 (moderate), and 3 (strong).Numerical simulations
The numerical simulation was performed using the COMSOL Multiphysics simulation platform, establishing a coupled model of analyte transport and surface-immobilized target reaction kinetics within a microfluidic system. The binding of antigen and antibody is a reversible, specific bimolecular reaction, the essence of which is the highly complementary binding of immune recognition molecules in three-dimensional space. This process relies on the spatial conformational matching between the antigenic epitope and the complementarity-determining regions (CDRs) within the variable region of the antibody. The association rate constant (k_ads) reflects the efficiency of molecular collision and successful binding, while the dissociation rate constant (k_des) embodies the stability of the complex.28 When the concentration of the immobilized ligand is significantly lower than that of the analyte in solution, the binding process can be approximated and treated using pseudo-first-order reaction kinetics.| [A] + [B] ⇌ [AB] |
The simulation assumes the reaction follows a first-order Langmuir adsorption model. The equations describe the temporal evolution of the target density (B0), the concentration of the analyte at the reactive surface ([A]surface), and the concentration of the resulting analyte-target complex (AB), denoted as Cs. R represents the reaction rate.29
The initial surface density of targets, B0, was set to 1 × 10−7 mol m−2.30 The dissociation rate constant, k_des, was set to 3 × 10−4 s−1, and the association rate constant, k_ads, was 2.5 × 102 m3 mol−1 s−1.31 The initial bulk concentration of analyte A, C0, was 1 × 10−4 mol m−3. The transport of the reagent within the microfluidic chip is governed by Fick's second law of diffusion:
Results and discussion
An automated microfluidic system was developed for rapid IHC staining. The system consists of four main modules: a control panel, an injection pump, a temperature control unit, and a microfluidic chip (Fig. 1a). A photograph of the experimental setup is provided in Fig. S1a. Reaction parameters, including incubation time, reagent volume, and flow rate, can be preset via the control panel (Fig. S1c). The injection pump, which operates according to programmed parameters, is equipped with eight inlet ports connected to reagent reservoirs and one outlet port connected to the microfluidic chip inlet. The microfluidic chip was reversibly bonded within a metal fixture (Fig. S1b) and placed on a heating plate maintained at 37 °C. Upon initiation, the system automatically executes the entire staining process, including antibody incubation, washing, and chromogenic development.Design for flow rate uniformity within microfluidic chips
In IHC, staining intensity, specificity, and uniformity are critical metrics for evaluating assay success. Uniformity refers to the consistent distribution of staining across the entire tissue section without local over-staining, under-staining, or edge effects. In conventional IHC, achieving uniform staining requires ensuring that each reagent fully covers the tissue section and extends slightly beyond its edges to prevent inhomogeneity due to incomplete coverage. Similarly, in microfluidic-based immunostaining, it is essential to ensure that reagents uniformly and indiscriminately cover every corner of the chip's reaction area. By designing the microchannel microstructure and the reaction zone geometry, a stable, uniform flow profile can be established. This enables reagents to flow through the tissue region at a consistent velocity and concentration, thereby ensuring highly uniform fluid shear stress, reagent concentration, and reaction time regardless of whether the sample is located at the center or the periphery of the chip.The microfluidic chip comprised a fluid distribution network and a reaction chamber. If the antibody reagent flows unidirectionally within the microchip and is discharged immediately after passing through the microchannel once, the unbound antibodies are directly discarded without being utilized, resulting in significant reagent consumption. To enhance mass transfer efficiency by employing a higher flow rate while conserving the antibody reagent, a bidirectional recirculating pumping strategy was adopted. This approach enables the reagent to flow back and forth within the closed microchip, allowing unbound antibodies to regain opportunities for binding with antigens during the reverse flow (Fig. 1b). Fig. 1c shows a side view of reagent flow within the microfluidic chip. Consequently, the distribution channels employ a fully symmetric geometric design to ensure a consistent flow-field distribution under both forward and reverse flow directions. In this work, two distinct symmetric distribution architectures were proposed: a tree topology structure (Fig. 2a) and a pillar-array-based flow-splitting structure (Fig. 2c). Computational fluid dynamics (CFD) simulations were performed to analyze the hydrodynamic behavior of both structures at a flow rate of 10 μL s−1. The simulations indicate that at this flow rate, both designs can rapidly achieve complete reagent exchange within the reaction chamber. Analysis of the velocity distribution, however, reveals notable differences: the tree topology chip produces a nearly uniform, “front-aligned” flow profile under pressure-driven conditions, with highly consistent velocities throughout the chamber (Fig. 2b). In contrast, the pillar array flow-splitting chip exhibits a markedly non-uniform velocity distribution, characterized by higher velocities in the central region and lower velocities near the sidewalls, which may lead to spatial variations in mass-transfer rates across the reaction zone (Fig. 2d).
The tissue sections stained with the two types of chips were divided into five regions—upper, middle, lower, left, and right (Fig. 2e). From each region, ten images were randomly selected and processed using ImageJ software. First, color deconvolution was applied to separate the DAB-positive regions from the hematoxylin-stained (blue, nuclear) areas. The threshold was then adjusted to identify the strongly stained regions, and the integrated optical density (IOD)—defined as the sum of the optical densities of all pixels within these regions—was calculated. Finally, the AOD values were obtained using the formula AOD = IOD/A, where A denotes the total area of the DAB-positive region. A higher AOD value indicates stronger staining intensity, and a more concentrated AOD distribution reflects greater staining uniformity across the tissue section. The AOD values obtained with the tree topology chip (ranging from 0.312 to 0.354) were notably more clustered than those obtained with the pillar array flow-splitting chip (ranging from 0.288 to 0.350).
The interquartile range (IQR) is a statistical measure that quantifies the dispersion of a dataset. It is defined as the difference between the third quartile (Q3) and the first quartile (Q1), expressed as IQR = Q3 – Q1. The IQR captures the spread of the middle 50% of the data; a smaller IQR indicates that the central portion of the data is more concentrated and uniformly distributed, whereas a larger IQR suggests a higher degree of dispersion. Under the tree topology structure (Fig. 2f), the IQR values were 0.00335 in the x-direction and 0.00145 in the y-direction, demonstrating excellent overall uniformity with low dispersion in both directions. Notably, the y-direction exhibited even lower dispersion and superior uniformity compared to the x-direction. In contrast, under the pillar array flow-splitting structure (Fig. 2g), the IQR value in the x-direction increased significantly to 0.01375, while the IQR in the y-direction also rose noticeably to 0.008, indicating a marked decline in overall uniformity. The increase in dispersion was particularly pronounced in the x-direction, where the IQR reached 411% of that observed in the tree topology structure. Similarly, dispersion in the y-direction increased substantially, with an IQR value 552% higher than that of the tree topology structure. This clear shift from the tree topology structure to the pillar array flow-splitting structure resulted in a transition from a highly uniform, low-dispersion state to one characterized by significantly increased dispersion and reduced uniformity. The issue of dispersion was especially prominent in the x-direction, while uniformity in the y-direction also deteriorated. Consequently, the symmetric tree topology fluid distribution channel was ultimately selected, as it effectively ensures spatial uniformity during reciprocating reagent flow within the chip, thereby providing a fluidic foundation for highly consistent immunostaining in subsequent steps.
Enhanced staining intensity and rapid immunohistochemistry achieved by a V-groove upper-structure microfluidic chip
To achieve efficient and uniform IHC staining, a microfluidic chip featuring a V-groove upper-layer structure was designed and fabricated. The chip reaction area measures 15 mm × 15 mm, which was sufficiently large to cover the tissue section. Its height was set at 50 μm, approximately ten times greater than the thickness of a typical tissue section (4–5 μm), thereby preventing mechanical compression of the specimen. Key geometric parameters of the V-groove structure—including the apex angle, width, and depth—were optimized using computational fluid dynamics (CFD) simulations, with the concentration of target antigen–antibody complexes generated at 100 s serving as the metric for reaction rate. The simulation results indicated that the mass transfer efficiency was maximized when the V-groove depth was 10 μm, the apex angle was 90°, and the width was 0.5 mm (Fig. 3a–d).IHC staining was performed on consecutive breast cancer tissue sections using the microfluidic chips and compared with conventional manual staining. Staining within the chip was conducted at 37 °C: first, 150 μL of reagent was injected at a flow rate of 20 μL s−1 to achieve initial filling, followed by bidirectional reciprocating flow at 10 μL s−1, creating an “aspiration-ejection” pumping pattern. The washing step was carried out by flushing at 20 μL s−1 for 20 s to ensure thorough removal of unbound antibodies. Under identical incubation time and temperature conditions, immunostaining was performed separately using the two chip types (Fig. 4b and c), with conventional staining results serving as the control (Fig. 4a). Fig. 4d–f show magnified views of the red-boxed areas, respectively. The staining intensity obtained on the chips was normalized, and the staining outcomes were evaluated based on the microscopic image signal intensity. The experimental results showed that the staining intensity of the V-groove chip was 102% of that achieved by the conventional method, while the planar microfluidic chip (P-chip) exhibited only about 88% of the conventional staining intensity, corresponding to an approximately 15% enhancement in staining signal strength for the V-groove chip compared to the planar chip (Fig. 4g). This result confirms that the V-groove upper-layer structure enhances the antigen–antibody binding rate, providing a reliable hydrodynamic enhancement mechanism for achieving rapid and efficient IHC staining.
Further optimization of the key reaction steps was performed. Without compromising staining quality, the DAB chromogenic time was reduced to 1 min (Fig. S2a), the secondary antibody incubation time to 2 min (Fig. S2b), and the primary antibody incubation time to 4 min (Fig. S2c). Fig. S2d–f show the average optical density values under the corresponding conditions. The entire immunostaining process could thus be completed within 11 min. However, the traditional immunohistochemical staining process takes 122 minutes, and the microfluidic system reduces the time by 90% (Table S1). When comparing V-chip staining with off-chip static staining under the same total reaction time (11 min), the latter exhibited a significant reduction in staining intensity. To quantitatively evaluate the staining performance, cytokeratin (CK) markers in four different tissue samples—breast cancer, lung cancer, cervical cancer, and gastric cancer—were stained using the V chip (Fig. 5a–d) and the off-chip method (Fig. 5e–h), followed by semi-quantitative analysis using the H-score (Fig. 5i). Independent sample t-tests indicated a highly statistically significant difference in staining intensity between the two groups (P < 0.001). These results demonstrated that the V-groove-based microfluidic platform substantially shortens IHC processing time, highlighting its strong potential for rapid pathological diagnosis.
Numerical simulation was employed to analyze the surface reaction dynamics within the microfluidic chip and elucidate the mechanism by which the V-groove structure enhances antigen–antibody binding. The simulation results demonstrated that under identical flow conditions, the microfluidic chip with a V-groove upper-layer structure exhibits a higher production rate of target antigen–antibody reaction products compared to the planar-channel chip, with an approximately 22% increase in the initial reaction rate (Fig. 6a–c).
Flow-field analysis revealed that in the planar structure, fluid flow follows a typical laminar pattern, with radial reagent transport predominantly driven by molecular diffusion, resulting in a diffusion-limited reaction process. In contrast, within the V-groove structure, flow disturbances induced by the groove geometry generate a pronounced vertical velocity component (Fig. 6d), which enhances the transport of reagents perpendicular to the primary flow direction. This convection-augmented effect significantly diminishes the boundary-layer influence, increases the collision frequency between antigen and antibody molecules, and thereby accelerates immune-complex formation.
The effect of flow rate within microfluidic chips on immunohistochemical staining outcomes
In microfluidic chip-based IHC, the flow rate directly influences the final immunostaining outcome by regulating the binding efficiency between antibodies and antigens. At low flow rates, mass transport of antibodies to the reaction interface is dominated by diffusion, which significantly limits the transport rate and impedes rapid replenishment of antibodies consumed at the surface, potentially leading to an antibody-depleted zone at the interface. This markedly slows the kinetics of antigen–antibody binding, prolongs the incubation time required for sufficient staining, and thereby reduces detection efficiency. Conversely, excessively high flow rates may impose substantial shear stress on tissue samples, posing a potential risk to structural integrity. Hence, precise modulation of flow rate is critical for successful microfluidic IHC assays.To identify the optimal flow rate, a systematic flow rate optimization experiment was conducted using consecutive tissue sections (Fig. 7a–f). The results demonstrated that within the range of 1–60 μL min−1, the AOD values corresponding to staining signals increased gradually with rising flow rates. Beyond this range, the AOD values plateaued, indicating that antigen–antibody binding approached saturation (Fig. 7g). Increasing the flow rate beyond 60 μL min−1 resulted in no significant effect on staining intensity. Therefore, the incubation flow rate can be appropriately selected within the range of 60 μL min−1 to 900 μL min−1. In this experiment, a relatively high flow rate of 600 μL min−1 was chosen to ensure staining quality. Regarding sample safety, a flow rate of 600 μL min−1 does not generate excessive shear stress; it neither causes tissue sample detachment nor structural damage, thereby preserving the integrity of clinical specimens.
Validation of reproducibility and clinical sample staining in microfluidic immunohistochemistry
To validate the consistency and reproducibility of microfluidic IHC in both qualitative interpretation and semi-quantitative evaluation of staining intensity, dozens of FFPE tissues, including breast cancer, tonsil, lung cancer, and gastric cancer specimens, were analyzed. Immunohistochemical staining was performed for five biomarkers: CK (14 cases), Ki-67 (11 cases), HER-2 (5 cases), CK5/6 (5 cases), and CD3 (5 cases). All samples were independently assessed by pathologists, with conventional IHC results used as the reference standard (Table S2). Staining in the experimental group was performed using a microfluidic system, followed by microscopic imaging to acquire signals.The staining results (positive/negative or graded scores) for all markers showed complete concordance between the two methods. No false-positive or false-negative results were observed among the 40 cases, indicating that the microfluidic chip approach accurately reproduces the qualitative interpretation of conventional IHC. Staining intensity was evaluated using AOD values. Pearson correlation analysis of AOD values obtained from both methods across the 40 cases revealed a very strong positive correlation between conventional IHC and the microfluidic chip method (r = 0.9552, P < 0.0001) (Fig. 8a).
These data confirm that the microfluidic chip method not only aligns with conventional IHC in qualitative assessment but also demonstrates excellent agreement in semi-quantitative staining intensity evaluation.
For reproducibility assessment, five consecutive sections from the same FFPE sample were subjected to microfluidic IHC staining for CK, and AOD values were calculated. The standard deviation was 0.0235, with a coefficient of variation (CV) of 3.00% (Fig. 8b). These results indicate robust stability in reagent delivery and incubation conditions within the microfluidic system, demonstrating that microfluidic IHC offers excellent reproducibility for FFPE tissue analysis. The method proved equivalent to conventional IHC while providing substantially faster processing and higher automation.
Conclusions
In this study, an automated immunohistochemistry staining system based on a microfluidic chip with a V-groove upper structure was developed to address limitations of conventional IHC methods, including labor-intensive procedures and long processing times. Utilizing a reagent recirculation scheme within the chip and a V-shaped microchannel designed to enhance passive mixing, the system effectively promotes vertical convective mass transfer between antigens and antibodies without external actuation, thereby significantly shortening the immunoreaction duration. Experiments demonstrated that staining for CK in breast cancer tissue sections was completed within 11 minutes, representing a 90% time reduction compared to standard protocols, while achieving comparable staining intensity and uniformity. Results for multiple biomarkers across various clinical samples showed excellent concordance with traditional methods, with a correlation coefficient of 0.9552 and good reproducibility. Furthermore, computational fluid dynamics simulations and structural optimizations elucidated the underlying fluidic mechanism: the V-groove upper structure enhances the reaction rate by inducing secondary flow to disrupt the diffusion boundary layer. The design of a tree topology distributor was also validated to ensure uniform staining. This work not only provides a reliable platform for rapid, automated IHC but also offers substantial experimental and theoretical support for implementing microfluidics in pathological diagnostics.Author contributions
Lu Zhong: writing – original draft, writing – review and editing, formal analysis, methodology, investigation. Hang Chen: investigation, formal analysis. Hong-Lei Chen: writing – review and editing, supervision, resources, methodology. Peng Jun: resources, methodology. Zhi-Ling Zhang: writing – review and editing, supervision, resources, methodology, funding acquisition.Conflicts of interest
The authors do not have any competing or financial interests.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: the SI contains: Fig. S1, a schematic diagram of the microfluidic immunohistochemical staining system setup; Fig. S2, optimization of staining time conditions (DAB chromogenic time, HRP incubation time, and cytokeratin incubation time); Table S1, comparison of staining time between conventional and microfluidic immunohistochemistry methods; and Table S2, comparison of staining results for five biomarkers between the two methods. See DOI: https://doi.org/10.1039/d6lc00089d.Acknowledgements
This work was supported by the National Natural Science Foundation of China (22274118 and 22574124).Notes and references
- P. Brandtzaeg, J. Immunol. Methods, 1998, 216, 49–67 CrossRef CAS PubMed.
- D. Y. Mebratie and G. G. Dagnaw, Semin. Diagn. Pathol., 2024, 41, 154–160 CrossRef PubMed.
- J. A. Ramos-Vara and M. A. Miller, Vet. Pathol., 2014, 51, 42–87 CrossRef CAS PubMed.
- R. D. Lovchik, D. Taylor and G. Kaigala, Microsyst. Nanoeng., 2020, 6, 94 CrossRef CAS PubMed.
- K. Imai, H. Nanjo, W. Shigeeda, T. Sugai, T. Ito, Y. Maniwa, S. Takashima, H. Saito, N. Yanagawa, Y. Tanaka, T. Doi, Y. Hiroshima, K. Nomura, M. Tanino, S. Tanaka, Y. Minamiya and R. I. S. Grp, Cancer Sci., 2023, 114, 702–711 CrossRef CAS PubMed.
- K. Terata, H. Saito, H. Nanjo, Y. Hiroshima, S. Ito, K. Narita, Y. Akagami, R. Nakamura, H. Konno, A. Ito, S. Motoyama and Y. Minamiya, Sci. Rep., 2017, 7, 2810 CrossRef.
- M. S. Johnson, J. N. Mueller, C. Daniels, H. N. Najm and J. Zádor, J. Phys. Chem. A, 2024, 128, 3685–3702 CrossRef CAS.
- H. Nygren, M. Werthen and M. Stenberg, J. Immunol. Methods, 1987, 101, 63–71 CrossRef CAS PubMed.
- H. Nygren and M. Stenberg, J. Colloid Interface Sci., 1985, 107, 560–566 CrossRef CAS.
- M. Stenberg, L. Stiblert and H. Nygren, J. Theor. Biol., 1986, 120, 129–140 CrossRef CAS PubMed.
- H. X. Li, Q. Zhou, N. Cao, C. H. Hu, J. C. Wang, Y. He, S. Jiang, Q. Li, M. Chen, L. Gong, M. Luo, X. Z. Deng and Z. G. Luo, Cell Commun. Signaling, 2025, 23, 271 CrossRef.
- A. Bange, H. B. Halsall and W. R. Heineman, Biosens. Bioelectron., 2005, 20, 2488–2503 CrossRef CAS.
- K. N. Han, C. A. Li and G. H. Seong, Annu. Rev. Anal. Chem., 2013, 6, 119–141 CrossRef CAS.
- T. Gervais and K. F. Jensen, Chem. Eng. Sci., 2006, 61, 1102–1121 CrossRef CAS.
- S. Brajkovic, B. Pelz, M. G. Procopio, A. L. Leblond, G. Repond, A. Schaub-Clerigué, D. G. Dupouy and A. Soltermann, Diagn. Pathol., 2018, 13, 79 CrossRef CAS.
- A. T. Ciftlik, H. A. Lehr and M. A. M. Gijs, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 5363–5368 CrossRef CAS.
- S. Kwon, C. H. Cho, Y. Kwon, E. S. Lee and J. K. Park, Sci. Rep., 2017, 7, 45968 CrossRef CAS.
- M. S. Draz, D. Dupouy and M. A. M. Gijs, Lab Chip, 2023, 23, 3258–3271 RSC.
- G. Cappi, D. G. Dupouy, M. A. Comino and A. T. Ciftlik, Virchows Arch., 2019, 9, 4489 Search PubMed.
- T. M. Squires and S. R. Quake, Rev. Mod. Phys., 2005, 77, 977–1026 CrossRef CAS.
- Z. H. Li, B. L. Zhang, D. Dang, X. L. Yang, W. G. Yang and W. F. Liang, Sens. Actuators, A, 2022, 344, 113757 CrossRef CAS.
- Z. Z. Chen, L. Shen, X. Zhao, H. Q. Chen, Y. X. Xiao, Y. H. Zhang, X. P. Yang, J. H. Zhang, J. J. Wei and N. J. Hao, Appl. Mater. Today, 2022, 26, 101356 CrossRef.
- M. Wojnicki, X. G. Yang, P. Zabinski and G. Mutschke, Micromachines, 2025, 16, 422 CrossRef.
- S. L. Stott, C. H. Hsu, D. I. Tsukrov, M. Yu, D. T. Miyamoto, B. A. Waltman, S. M. Rothenberg, A. M. Shah, M. E. Smas, G. K. Korir, F. P. Floyd, A. J. Gilman, J. B. Lord, D. Winokur, S. Springer, D. Irimia, S. Nagrath, L. V. Sequist, R. J. Lee, K. J. Isselbacher, S. Maheswaran, D. A. Haber and M. Toner, Proc. Natl. Acad. Sci. U. S. A., 2010, 107, 18392–18397 CrossRef CAS PubMed.
- Y. Zhang, X. D. Tong, L. Yang, R. L. Yin, Y. Li, D. Zeng, X. Y. Wang and K. Deng, Sens. Actuators, B, 2021, 332, 129511 CrossRef CAS.
- A. D. Stroock, S. K. W. Dertinger, A. Ajdari, I. Mezic, H. A. Stone and G. M. Whitesides, Science, 2002, 295, 647–651 CrossRef CAS.
- J. P. Golden, T. M. Floyd-Smith, D. R. Mott and F. S. Ligler, Biosens. Bioelectron., 2007, 22, 2763–2767 CrossRef CAS PubMed.
- D. R. Davies, E. A. Padlan and S. Sheriff, Annu. Rev. Biochem., 1990, 59, 439–473 CrossRef CAS.
- D. B. Hibbert, J. J. Gooding and P. Erokhin, Langmuir, 2002, 18, 1770–1776 CrossRef CAS.
- H. Hajji, L. Kolsi, W. Hassen, A. Al-Rashed, M. N. Borjini and M. A. Aichouni, Phys. E, 2018, 104, 177–186 CrossRef CAS.
- C. K. Yang, J. S. Chang, S. D. Chao and K. C. Wu, J. Appl. Phys., 2008, 103, 084702 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |