Open Access Article
Open Access ArticleFast and precise magnetophoresis of superparamagnetic nanoparticles on a micro-magnetic substrate in a static liquid environment
Elise
Bou
 *ab,
Claudia
de la Fuente
ab,
Etienne
Orsini
*ab,
Claudia
de la Fuente
ab,
Etienne
Orsini
 a,
Sarah
Delshadi
a,
Sarah
Delshadi
 c,
Orphée
Cugat
c,
Orphée
Cugat
 *b and
Franz
Bruckert
*a
*b and
Franz
Bruckert
*a
aUniversité Grenoble Alpes (UGA), CNRS, Grenoble INP, LMGP, 38000 Grenoble, France. E-mail: elise.bou@grenoble-inp.fr; franz.bruckert@grenoble-inp.fr
bUniversité Grenoble Alpes, CNRS, Grenoble INP, G2Elab, 38000 Grenoble, France. E-mail: orphee.cugat@grenoble-inp.fr
cMagIA Diagnostics, 15 Rue Maréchal Leclerc, 38130 Échirolles, France
First published on 10th February 2026
Abstract
Micro-magnet arrays coupled with external rotating magnetic fields have been reported for the manipulation of individual superparamagnetic particles in stationary microfluidics, requiring no fluidic actuation. In addition to their magnetic properties preventing particle aggregation, superparamagnetic particle size reduction advantageously increases their interaction with surrounding liquid environment. However, precise magnetophoresis of submicrometric particles is challenging, owing to their low magnetic content. We present a micro-magnetofluidic chip for fast and collective transport of superparamagnetic nanoparticles (SNPs) over centimetric distances. In our innovative approach, SNPs are captured on a line pattern on top of a micro-magnet array covered with a spacer, which allows for SNP precise positioning along the surface using a rotating external magnetic field. The impact of SNP size on magnetophoresis was explored by characterizing the velocility ranges of 100 nm and 200 nm SNPs. For 100 nm size, a critical speed of 500 μm s−1 was reached beyond which the particles get resuspended, whereas for the 200 nm size, our system maximum rotation speed conveys the particles at up to 1.4 mm s−1 without reaching a critical speed. The mobility of SNPs was investigated in different buffers commonly used for biological analysis, and shown to be enhanced in low ionic strength conditions. As an example of future application, we demonstrate nucleic acid detection by SNP accumulation enabling to concentrate the fluorescence signal. This system provides the opportunity to be easily integrated in lab-on-a-chip (LOC) devices and could help relieve current development limitations related to complex and costly fluidic actuation instrumentation.
Introduction
Micro-magnetofluidics exploit magnetic fields and gradients for the contactless manipulation of magnetic particles within microfluidic chips by magnetophoresis. A wide range of functions can be implemented, such as mixing or pumping liquids, and for trapping or sorting magnetic particles.1–3 Particle surface functionalization allows for specific interactions with biomolecules and cells, enabling biological analysis integration in microfluidics.4 Hence, manipulation of magnetic particles in microfluidics has been widely explored for the development of lab-on-a-chip (LOC) devices.5,6 Superparamagnetic particles, composed of nano-sized magnetic inclusions within polymer or silica matrices, exhibit particular magnetic properties resulting in zero-net particle magnetization in the absence of an external magnetic field, which prevents their magnetostatic aggregation and is thus highly beneficial for practical manipulation in microfluidics.Stationary microfluidics, for which no fluidic actuation is required, represent an opportunity for future industrialization of LOC devices by reducing the manufacturing cost and the associated instrumentation complexity. Magnetophoresis can be implemented in stationary microfluidics to move particles, using variable magnetic field gradients. Strong and long-range magnetophoretic forces are able to move particles without the need for a microfluidic flow.
Micro-magnet arrays in microfluidic chips have been reported as efficient tools to manipulate magnetic particles in static liquid environments. A panorama of geometries, materials and applications of micro-magnet arrays for individual superparamagnetic particle manipulation have been reviewed in detail.7 These micro-magnet arrays usually consist in an arrangement of discrete magnetic structures embedded within a substrate. They have been produced in various soft and hard magnetic materials, usually iron-nickel alloy and cobalt, or in ferrite garnet thin films exhibiting spontaneous domains with alternating magnetization directions after deposition.7 A variety of structure geometries have been explored including disks,8 square and triangle shapes,9 or strips.10,11 Such structures can generate or concentrate magnetic fields and produce strong magnetic field gradients, up to 105 T m−1, in localized volumes. Magnetized particles are attracted by the gradients towards the substrate regions with magnetic field maxima. Magnetic particle actuation on the substrate is implemented using a rotating external magnetic field to modulate the local fields by shifting the locations of field maxima. A field rotation in the plane of the substrate is used to actuate the particles along the borders of the micro-magnetic structures.12 A magnetic field rotation out of the plane of the substrate is used to induce step-wise motion from one trapping position to the next. This longitudinal transport configuration was implemented to discriminate different particle sizes or particles bound to specific biomolecules.8–10,13 Disposition of a layer on top of a 1D micro-magnet array has been studied to allow continuous displacement of 50–200 μm particle aggregates.14 However, these micro-magnet arrays are fabricated using clean room processes, which precludes their large-scale use. The cost of producing micro-magnet array is therefore a first challenge.
Reducing particle size to submicrometric dimensions increases their diffusivity, thus reducing their sedimentation, while increasing their surface-to-volume ratio. These combined effects enhance interactions between functionalized submicrometric superparamagnetic particles and targeted elements within a liquid environment.15 The second challenge nevertheless lies in applying sufficient magnetophoretic forces to manipulate these particles presenting reduced magnetic content, especially in stationary liquids.
In this context, the start-up company MagIA diagnostics has developed a technology that addresses these two challenges. The system consists in a micro-magnetofluidic chip with an embedded plastoferrite micro-magnetic substrate, and employs 200 nm superparamagnetic particles to target specific biomolecules in the sample to analyze. Magnetophoretic-induced convective attraction of the particles suspended in the sample towards the substrate enables the rapid capture of the whole nanoparticle population.16 The innovative no-wash analysis protocol relies on their capture on a pre-determined line pattern, enabling the differential detection from a single image, of localized fluorescence from the biomolecules bound to the particles.17 An immunoassay portable device was developed for the rapid and simultaneous detection of a panel of infectious diseases (HIV, HBV and HBC).
The objective of our work is to implement lateral magnetophoresis for continuous, collective superparamagnetic nanoparticle (SNP) transport and for their accumulation in a stationary microfluidic environment. Using micro-magnet arrays similar to those developed for the MagIA diagnostics systems, we investigate the role of a spacer on top of the micro-magnet array for effective magnetophoresis and precise positioning of SNPs along the surface. We explore the transport of 100 nm and 200 nm SNPs over centimetric distances, at velocities reaching 1.4 mm s−1. We also study SNP mobility in various buffer solutions relevant for biological applications. Finally, we improve the detection of fluorescent DNA thanks to SNP magnetophoresis and accumulation along a line.
Working principle
When a SNP is subjected to an external magnetic field H, it exhibits a magnetic moment m, which is dependent on its magnetic material magnetization M and volume V:| m = MV | (1) |
The magnetization M follows a linear relationship with the external magnetic field H at low magnetic fields (below 10 mT):
| M = χH | (2) |
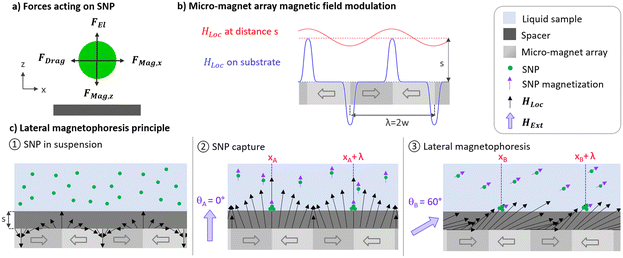
Magnetophoresis is related to the motion of magnetized particles subjected to a magnetophoretic force when positioned in a magnetic field gradient. Forces involved in particle magnetophoresis within a liquid environment have been extensively described in the literature.4,18–22 SNP magnetophoresis is implemented at the surface of a flat magnetic substrate magnetized with a periodic micro-magnetic pattern. Fig. 1a illustrates the main forces acting on a single SNP moving near a substrate. The magnetophoretic force FMag applied to one single SNP can be expressed as:
| FMag = μ0(m·∇)H | (3) |
| FDrag = 6πηrvfD | (4) |
Due to the reduced size and magnetic content of SNPs, strong magnetic gradients are required to generate a magnetophoretic force sufficient to displace them in liquid. To this end, we use a micro-magnet array, since reducing the size of magnets increases the magnetic field gradients generated in their vicinity.24 The micro-magnetic array embedded in our substrate consists in juxtaposed magnetic domains of width w, presenting alternating magnetization directions along x. This micro-magnetic array results in a strip pattern of defined periodicity λ = 2w, generating a local periodical magnetic field HLoc (Fig. 1b). As shown earlier for micro-magnet arrays, magnetic field and gradient profiles along x evolve from a sharp peak-to-valley profile to a weaker sinusoidal-shape profile as the distance from the substrate increases along z.25 We exploit this smoothing of magnetic field and gradient profiles as a function of the distance z from the substrate by disposing a spacing layer of appropriate thickness s (Fig. 1c). This spacer separates the sample from the micro-magnetic substrate, at which distance s the magnetic field generated by the micro-magnet array becomes insufficient to magnetize and attract suspended SNPs towards the substrate. This allows the suspended SNPs to diffuse and interact with the elements of interest present in the sample during an incubation time. It is possible to attract the suspended SNPs towards the substrate by magnetizing them through an external homogeneous magnetic field HExt applied along any direction θ in the (xz) plane (Fig. 1c, step 2). Junctions between adjacent magnetic patterns separated by the distance λ present higher magnetic fields and gradients, corresponding to minimum magnetic potential energy locations where SNPs are captured. The patterns and their junctions are linear, resulting in SNP capture on a line pattern in the (xy) plane with periodicity λ along x. Once the SNPs are captured on the line pattern at the surface of the spacer, they can undergo lateral magnetophoresis along the substrate. Fig. 1c depicts the principle of SNP lines lateral magnetophoresis in our specific configuration. Rotating the external magnetic field HExt by θ degrees in the (xz) plane results in displacing the capture sites along x by a distance d expressed as follows:
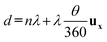 | (5) |
The driving speed of SNP lines vDrive along the substrate is directly proportional to the external magnetic field rotation speed ω and the micro-magnet period λ:
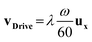 | (6) |
Materials and methods
SNPs and reagents
Nano-screen mag ARA/G SNPs of diameter 100 nm and 200 nm (4415-5, Chemicell) are composed of a magnetite core, a layer of green fluorescent lipophilic dye and a polysaccharide matrix shell. Supplier massic concentration is 25 mg mL−1. SNPs were resuspended at a massic concentration of 29 μg mL−1 in Tris HCl 10 mM buffer (pH 8.3), corresponding to 5.2 × 1010 and 6.4 × 109 particles per mL for 100 nm particles and 200 nm particles, respectively. Alternative resuspension buffers were prepared from a Denhardt 50× solution (D2532 Sigma Aldrich), an hybridization 2× solution (H7140, Sigma-Aldrich) and a DNA binding buffer solution (Buffer PB, Quiagen) containing guanidinium chloride and isopropanol.Estapor carboxylated SNPs of diameter 200 nm (M1-020/50, Merck) were used for nucleic acid transport and detection experiments. These particles are non-fluorescent and composed of multiple magnetite inclusions in a polystyrene matrix. Supplier massic concentration is 100 mg mL−1. SNPs were functionalized with streptavidin as described previously.26 For nucleic acid transport and detection experiments, SNPs were diluted at a massic concentration of 15 μg mL−1 corresponding to 2.3 × 109 particles per mL, in a solution containing 0.5× hybridization solution, 1× Denhardt solution, 10 mM Tris HCl buffer and 4 μg mL−1 of amplified biotinylated fluorescent DNA. This amplified 461 bp DNA was obtained by PCR, using biotinylated and A488-labeled primers, as described previously.26
SNP hydrodynamic diameter characterization
SNP hydrodynamic diameter were characterized by dynamic light scattering (DLS) technique (Zetasizer Ultra, Malvern Panalytical). All samples were diluted in filtered deionised water to a concentration of 12 μg mL−1 to avoid multiple scattering effects and were equilibrated at 25 °C prior to analysis. In the case of Estapor 200 nm SNPs, analysis was performed in three conditions including bare non-functionalized SNPs, streptavidin-functionalized SNPs and streptavidin-functionalized SNPs incubated with biotinylated DNA. Biotinylated DNA was incubated for 5 minutes with streptavidin-functionalized SNPs, and the bound complexes were washed to remove unbound nucleic acids before resuspension in deionized water. Measurements were acquired in triplicate for each sample. Intensity-weighted hydrodynamic diameter was calculated using the instrument software analysis. Reported data in Table 1 are the mean hydrodynamic diameters and standard deviations, and size distributions are presented in SI, Fig. S1.| Nanoscreen mag 100 | Nanoscreen mag 200 | Estapor 200 | |||
|---|---|---|---|---|---|
| Bare | Streptavidin | DNA | |||
| Mean (nm) | 213 | 293 | 186 | 275 | 602 |
| Standard deviation (nm) | 1 | 21 | 1 | 8 | 23 |
Measured hydrodynamic diameters obtained for Nanoscreen mag 100 nm and 200 nm SNP are 213 ± 1 nm and 293 ± 21 nm respectively and exceed supplier nominal values. This may be due to an instability of the polysaccharide matrix structure. However, the measured 80 nm difference in diameter between the two SNP references is consistent. Hydrodynamic diameter results of 186 ± 1 nm, 275 ± 8 nm, and 602 ± 23 nm were obtained respectively for the bare, streptavidin-functionalized and DNA bound streptavidin-functionalized Estapor 200 nm SNPs. Following covalent functionalization with streptavidin, a clear increase in hydrodynamic diameter is observed which confirms the formation of a protein layer at the SPN surface by successful grafting. The measured difference in diameter between bare and streptavidin-functionalized SNPs is around 90 nm. This value exceeds expectations based on reported streptavidin size of 5–11 nm.27,28 This large increase in diameter may be due to the potential formation of aggregates of several streptavidin layers,29 or to a change of refractive index induced by streptavidin grafting and affecting DLS measurement.30 A substantial increase of 235 nm in diameter between streptavidin-functionalized SNPs with and without biotinylated DNA incubation suggests specific DNA adsorption on SPNs. This measured diameter difference is consistent with the estimated 156 nm size of DNA double-strands synthetized for this study, which has been previously described.26
SNP magnetic characterization
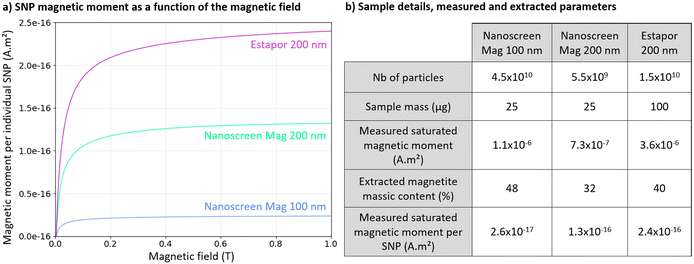
Magnetic characterization of the different SNP types was performed using a VSM SQUID magnetometer (Quantum Design). Magnetic moment as a function of magnetic field was measured in the range 0–1 T at room temperature for Nanoscreen mag ARA/G 100 nm and 200 nm (4415-5, Chemicell) and Estapor carboxylated 200 nm (M1-020/50, Merck) SNPs. Measurement curves are presented in Fig. 2a. Samples were prepared by pipetting 1 μL of supplier SNP solution for each reference on a 4 × 4 mm2 silicon substrate. Samples dried at room temperature within a few minutes resulting in 4.5 × 1010 and 5.5 × 109 particles corresponding to 25 μg sample mass for Nanoscreen mag 100 nm and 200 nm respectively, and 1.5 × 1010 particles corresponding to 100 μg sample mass for Estapor 200 nm. Measured saturated magnetic moments at 1 T were 1.1 × 10−6 A m2, 7.3 × 10−7 A m2 and 3.6 × 10−6 for Nanoscreen mag 100 nm, Nanoscreen mag 200 nm and Estapor 200 nm SNP samples respectively. Magnetite massic content contained in the various SNP references was determined using a magnetization per unit mass of 91 A m2 kg−1 and a density of 5.18 × 103 kg m−3.31 Resulting extracted magnetite massic contents are 48%, 32% and 40%, for Nanoscreen mag 100 nm, Nanoscreen mag 200 nm and Estapor 200 nm particles respectively. Results and sample details are summarized in Fig. 2.Micro-magnetofluidic chip
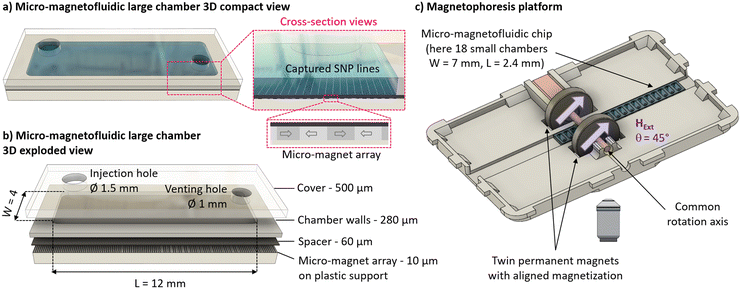
We developped a magnetophoresis platform for SNP lateral transport within a micro-magnetofluidic chip which does not require fluidic actuation. The micro-magnetofluidic chip concept used in this work has been developed by MagIA diagnostics company for magnetically localized immunoassays (MLFIA).32 The chip cross-section is presented in Fig. 3a. Materials for chip fabrication are provided by MagIA diagnostics, and have been described previously.17,32 A microfluidic chip is fabricated upon a flat micro-magnet array substrate composed of a 10 μm-thick layer of plastoferrite, a polymer composite loaded with magnetite powder, itself laminated on a millimeter-thick plastic surface for mechanical stability. The micro-magnet array is embodied by a pattern of magnetized strips presenting opposite in-plane magnetization directions with a period λ = 2w = 110 μm. This strip pattern generates local magnetic fields BLoc and gradients up to 50 mT and 5000 T m−1 respectively, at the surface of the substrate. A spacer consisting of a polypropylene film and acrylic adhesive is disposed on top of the magnetic substrate. The thickness s of the spacer is either 60 μm for a single layer or 120 μm for a double. Next, a 280 μm-thick polymer layer, consisting in polyester film sandwiched between two acrylic adhesive layers, is laser cut to form the outline of the fluidic chamber, and disposed on top of the spacer. Finally, a 500 μm thick PMMA cover is pressed on top of the double-sided adhesive layer to seal the fluidic chamber. Injection and venting holes, of diameters 1.5 mm and 1.1 mm respectively, are laser cut into the PMMA cover. A single chip has a footprint of 90 × 11 mm2 and can hold several independent rectangular chambers of assorted sizes (Fig. 3b). In the frame of this work, two chip types are used: one containing 18 small fluidic chambers (2.4 × 7 mm2, 6 μL each), and another containing 5 larger chambers (12 × 4 mm2, 15 μL each).UV-O3 surface treatment
To increase surface hydrophilicity, UV-O3 surface treatment was performed on the spacer covering the micromagnetic substrate. Surfaces were exposed during 30 min to an UV-ozone cleaner device (Novascan PSD). The change in water surface contact angles before (93.8° ± 2.6°, n = 4) and after (70.0° ± 3.2°, n = 4) treatment indicates the increased hydrophilicity of the microfluidic chamber induced by this procedure.Lateral magnetophoresis platform
For lateral SNP transport on the chamber substrate, the micro-magnetofluidic chip is inserted in the magnetophoresis platform (Fig. 3c). The platform is 3D printed by fused filament technique with PLA-Ultra polymer. The platform (110 mm wide and 150 mm long) is designed to be geometrically compatible with an inverted fluorescence microscope stage (IX71, Olympus), and presents a 11 mm circular window underside to allow observation through an inverted objective. The fluidic chip sits in a central groove, along which it can be manually positioned to observe the various chambers through the window. Two ring-shaped permanent magnets (neodymium N42, Neomagtec GmbH) are fixed with a 26 mm inner gap on a non-magnetic rotation axis, which is mounted in the plane of the platform. This way, the two ring magnets are placed either side of the fluidic chip and observation window. Magnetizations are along the magnets diameter, and aligned in the same direction. Magnets dimension are 30 mm outer diameter, 10 mm inner diameter and 6 mm thickness and their permanent magnetization is 1.25 T. Size, magnetization and gap between the magnets result in a uniform magnetic field BExt (65 mT at their center), as characterized by simulations and measurements, over an area large enough to cover the largest fluidic chambers. The magnets common rotation axis is connected to a miniature DC geared motor, driven by a standard variable power supply. The external magnetic field intensity and its rotation speed range were characterized using a Hall probe placed in the center of the platform, in the position of the observed chambers.Fluorescence microscopy
Fluorescence microscopy of fluorescent SNPs was performed using an Olympus IX70 microscope equipped with a 5× or 10× objective, a white LED lamp, a GFP fluorescence cube and a DP30BW 12 bit camera (1360 × 1024 pixels2, gain 8). Exposure parameters for imaging were 100% lamp power and 15 ms exposure time for GFP fluorescence and 1% lamp power and 0.5 ms exposure time for bight field. Time-lapse movies were recorded at a φ = 1.6 frames per second acquisition rate. In the case of nucleic acid transport and detection experiments, the lamp power was reduced to 60% and exposure time increased to 30 ms, to minimize photobleaching.Image processing
Vertically integrated fluorescence intensity profiles were extracted from fluorescence images using Image Pro Plus (MediaCybernetics). Processing of fluorescence intensity profiles, consisting in background signal elimination with baseline subtraction and immobile SNP fraction analysis, was performed using Python. Initial peaks were measured when the external magnetic field is at angle θ = 0°. A linear baseline was defined by the minima between peaks separated by λ, and the fluorescence profile was integrated above the baseline from one minimum to the next. The immobile fractions were measured after rotating the external magnetic field by an angle θ < 360°. After rotation, two peaks are visible: one at the same position as the initial peak corresponding to the SNP immobile fraction, and another one in-between successive initial peaks positions corresponding to the mobile fraction. The fluorescence under immobile and initial peaks was integrated in order to define the immobile fraction. The line velocity was determined on image sequences using ImageJ as: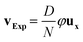 | (7) |
Results and discussion
Precise SNP position control
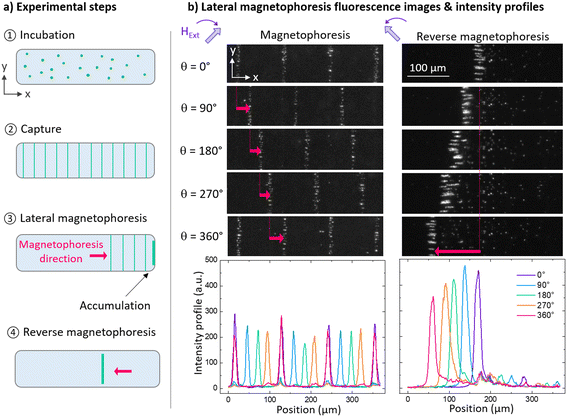
A typical lateral magnetophoresis experiment is depicted in Fig. 4a. A sample containing SNPs is injected in one chamber of the micro-magnetofluidic chip. The micro-magnetofluidic chip is then placed in the magnetophoresis platform, where the external magnetic field magnetizes the SNPs enough to get them captured by the micro-magnetic substrate. The time required to capture the suspended SNPs contained in the fluidic chamber volume on parallel lines (Fig. 4a, step 2) depends on SNP type and concentration.16 Lateral magnetophoresis is then activated by rotating the external magnetic field to displace the SNP lines along the chamber. SNP displacement is controlled by the angle θ of the rotating field, as shown on Fig. 4b, left (Movie S1). After multiple rotations, the SNPs have been driven against a side wall of the chamber, where they accumulate into a single wider line. This accumulation results in increased peak height and amplitude in the fluorescence intensity profile. Reversing the rotation direction drives this single line back towards the center of the chamber. In some cases, the wider accumulated line splits into a few thinner lines, as in the case of the experiment presented for DNA detection. By controling the number of revolutions and adjusting the θ angle, it is possible to precisely position this single line anywhere along the substrate (Fig. 4b, right, Movie S2). The images presented in Fig. 4b were obtained using fluorescent Nanoscreen mag 200 nm SNPs, at a concentration of 29 μg mL−1 in Tris HCl 10 mM buffer in a 2.4 × 7 mm2 chamber.In the literature, micrometric33 and even sub-micrometric position control has been achieved for single superparamagnetic micro-particle manipulation,34 using domain wall structures in thin magnetic films and external rotating magnetic fields. To our knowledge, other types of micro-magnet arrays used to transport SNPs have not reached the translational continuity position control and precision achieved in our work, as these systems rely on discrete fixed capture sites over bare micro-patterned magnet arrays previously reviewed by Rampini et al.7 Our configuration, allowing spatially continuous linear positioning of particle capture sites along the whole micro-magnet array thanks to the spacer, enables an increased translational position control. The position precision is linked to the angle θ precision enabled by the actuation system, and the micro-magnet periodicity λ. In our existing system, the precision on θ is ∼10° corresponding to ∼3 μm SNP positioning precision on the micro-magnetic substrate of periodicity λ = 110 μm; position precision, speed and controllability are currently being improved by implementing a stepper motor and associated digital driver. Shorter periodicity λ of the micro-magnet array is also a potential way to improve position precision.
SNP size, magnetic field rotation speed and spacer thickness effects on lateral magnetophoresis
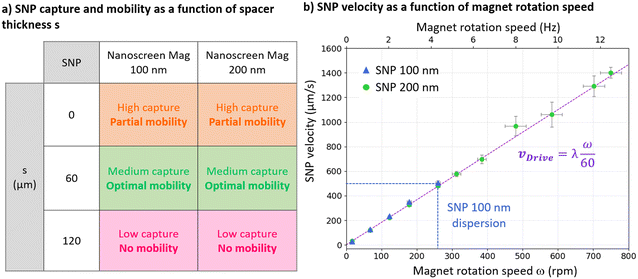
The following experiments were conducted with two SNP sizes, using Nanoscreen mag of diameter 100 nm and 200 nm, at a massic concentration of 29 μg mL−1 in Tris HCl 10 mM buffer, corresponding to 5.2 × 1010 and 6.4 × 109 particles per mL respectively. The fluidic chamber used for these experiments is 2.4 × 7 mm2 in size.SNP lateral magnetophoresis was first experimentally investigated for different spacer thicknesses: s = {0 μm, 60 μm, 120 μm}, corresponding to no polymer adhesive layer on the micro-magnetic substrate, one layer, and a double layer. SNPs were captured on the substrate for 5 min before applying magnetophoresis at rotation speed ω = 17 rpm. Fig. 5a presents a table summarizing experimental observations. Due to the intense magnetic field gradients generated at the surface of the substrate, SNP capture is high without spacer (s = 0 μm). However, the local magnetic field HLoc exhibiting a sharp peak-to-valley profile, modulation with the external field HExt is not appropriate for optimal displacement of the lines and we observe partial SNP mobility in this configuration. Gradients at the surface of a double spacer (s = 120 μm) are low, which results in low SNP capture and in particle resuspension when activating lateral magnetophoresis. At the surface of a single layer (s = 60 μm), gradients are moderate which allows for medium capture and appropriate lateral magnetophoresis. Moreover, the near-sinusoidal profile of HLoc at this distance s = 60 μm from the substrate is well suited for smooth and continuous displacement of SNP lines along the surface. A single 60 μm spacer was therefore selected for subsequent experiments.
The effect of a spacer on SNP velocity,35 and on continuous displacement of SNP aggregates has been previously investigated.14 In our case, the presence of the spacer does not affect SNP velocity, which follows vDrive (eqn (6)) over the characterized magnet rotation speed range.
Lateral magnetophoresis was implemented at various external magnetic field rotation speeds ω comprised in the range 17–750 rpm to experimentally determine the velocity range achievable for SNP transport (Fig. 5b). SNPs were first captured on the substrate forming single lines during 5 minutes, before activating magnetophoresis at 17 rpm during 2 minutes to accumulate the SNP lines towards one side of the chamber. Reverse magnetophoresis was then implemented to drive back a single 1D accumulated SNP line, and repeated over the magnetic field rotation range to characterize SNP experimental velocity vExp. Time-lapse images were recorded and vExp was computed by tracking the line position as a function of time (eqn (7)). Fig. 5b presents SNP velocities from n = 3 independent experiments for each external magnetic field rotation speed ω for 100 nm and 200 nm SNP sizes. The driving speed vDrive, proportional to rotation speed (eqn (6)), is represented as a dashed purple line. We observe that SNP velocity closely follows the linear capture sites displacement velocity driven by the rotating field. A critical speed of 506 ± 16 μm s−1 is achieved for 100 nm particles at ω = 260 ± 6 rpm, above which the particles disperse and are released in suspension. Hydrodynamic drag force FDrag becomes comparable to the magnetophoretic driving force FMag at this critical SNP speed. A drag force FDrag of 1.5 pN has been calculated using eqn (4) for 100 nm individual SNP at the measured critical speed of 500 μm s−1, using a drag coefficient fD ≈ 3 in the presence of a solid wall at the vicinity of the particle.4 We therefore estimate the order of magnitude of the magnetophoresis force FMag to be around ∼1.5 pN for individual 100 nm SNPs. Experimental 200 nm SNP velocities as high as 1403 ± 43 μm s−1 are reached at ω = 750 ± 30 rpm. This maximal magnet rotation speed at which SNP velocities were experimentally characterized is a current limitation of our motor performance. We expect a critical speed above 1400 μm s−1 at which 200 nm SNPs would disperse, as observed for 100 nm particles.
Yellen et al. developed an analytical model for single particle step-wise translational magnetophoresis over a micro-magnet array using a rotating external magnetic field. Particle velocity follows the tendency of an overdamped non-linear oscillator presenting a proportional evolution with magnetic field driving frequency below a critical frequency, above which the velocity is drastically reduced due to the damping induced by Stoke's drag.8 Although their described micro-magnet array and superparamagnetic particle type are different from our work, we can benefit from this study to establish an analytical understanding of our experimental results, taking note that magnetic interactions between particles are not negligible in our configuration due to their aggregation occurring on the linear capture sites. In reported studies investigating magnetophoresis on micro-magnet arrays, particle velocity is reduced or particles get immobilized on the substrate above a critical rotation speed.8,9,11,13 The behavior is different in our configuration where we observe dispersion and resuspension.
Velocities reaching up to 1 mm s−1 have been previously reported for superparamagnetic particle circular motion around the perimeter of a micro-magnet.36 To our knowledge, velocities exceeding 1 mm s−1 have not been reported in the literature for SNP translational displacement over a micro-magnet array in static liquid environment. This result opens possibilities for a range of applications involving the manipulation of biochemical molecules in microfluidics.
Immobile SNP fraction and electrostatic effects
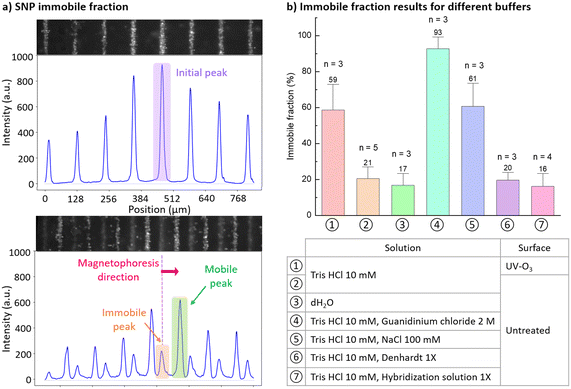
In magnetophoretic experiments, we observed that some SNPs were strongly attached to the spacer and not displaced by the rotating magnetic field. In order to better understand and optimize the displacement, the immobile SNP fraction was quantitatively studied in various buffers and solutions. SNPs used in these experiments were Nanoscreen mag 200 nm at a concentration of 29 μg mL−1 in a 2.4 × 7 mm2 chamber. Displacement was measured at low rotation speed ω = 17 rpm. SNP capture time was set at 5 min. Image acquisition at ×5 magnification allows to characterize 8 lines per frame. Fluorescence is integrated vertically along the x direction resulting in fluorescence peaks which correspond to captured SNP lines. Areas of initial peaks before starting magnetophoresis were extracted as described in Fig. 6a. After starting magnetophoresis, the initial peaks split into two sets of peaks, one corresponding to moving particles, the other to immobile ones. This immobile fraction is calculated as the average ratio between the areas of immobile peaks highlighted in orange on Fig. 6a(bottom) and of initial peaks highlighted in purple (top).The role of surface charges is evidenced by the effect of UV-O3 treatment of the spacer, which strongly increases the immobile fraction to 59 ± 14% (condition 1) as compared to the measured 21 ± 6% of the non-treated spacer (condition 2). This treatment is likely to generate functional groups such as carbonyl and hydroxyl groups,37,38 and therefore to increase the presence of negative charges. Both the SNPs and spacer therefore present repulsive negative charges. This experimental situation is described by the DLVO theory, which states that two energy minima exist, one at close distance where van der Waals interactions dominate, and one at larger distance where electrostatic repulsive interactions dominate, separated by an activation barrier. The vertical magnetophoretic forces exerted on the SNPs are possibly able to overcome this activation barrier and therefore to force the particles to come in close contact with the plastic surface where they become immobilized. In this theory, increasing the ionic strength decreases the energy barrier, favoring SNP adsorption on the surface.
Experimentally, we show that increasing the ionic strength by adding 100 mM NaCl notably increases the immobile fraction to 61 ± 13% (condition 5), which is consistent with DLVO theory and a previous experimental study by Wirix-Speetjens et al.39 Decreasing the ionic strength by suspending SNPs in dH2O reduces particle immobilization to 17 ± 7% (condition 3), which does not significantly differ from the results obtained in the condition with Tris HCl 10 mM buffer (condition 2).
Polymers are known to increase colloidal stability by preventing close interaction between particles and surfaces. In our experimental results, adding 1× Denhardt solution does not significantly reduce the immobile fraction (20 ± 4%, condition 6) in comparison with Tris HCl buffer (21 ± 6%, condition 2), suggesting that the presence of surface bound polymers is also insufficient to prevent particle immobilization. Adding 1× hybridization solution results in 16 ± 7% immobile fraction (condition 7) which again does not significantly differ from results with Tris HCl buffer (condition 2). However, hybridization 1× contains 750 mM NaCl, which represents a greater ionic strength than 100 mM NaCl (61 ± 13%, condition 5). Denaturated salmon DNA fragments and other polymers present in the hybridization solution have a significant effect on the decrease of the immobile fraction in a solution with high ionic strength.
Interestingly, addition of 2 M guanidinium chloride (condition 4) results in 93 ± 7% immobile fraction. This could be explained by the formation of salt-bridges between the positively charged guanidinium ions and carboxylic acids, as reported by Huerta-Viga et al.40 Future investigations on surface treatment and buffer composition are necessary to further reduce the SNP immobile fraction and optimize the transport of cargo molecules.
As we observe SNP aggregation on the linear capture sites, where magnetic interactions between SNPs are not negligible, the measured fluorescence is not proportional to SNP numbers because some fluorescence emission is hidden when particles stack vertically. Thus, the immobile fraction results should be interpreted as semi-quantitative values. Nevertheless, this analysis demonstrates clear electrostatics effects depending on the buffer nature, and the presented tendency can be used as a basis for further investigations.
DNA detection by SNP accumulation
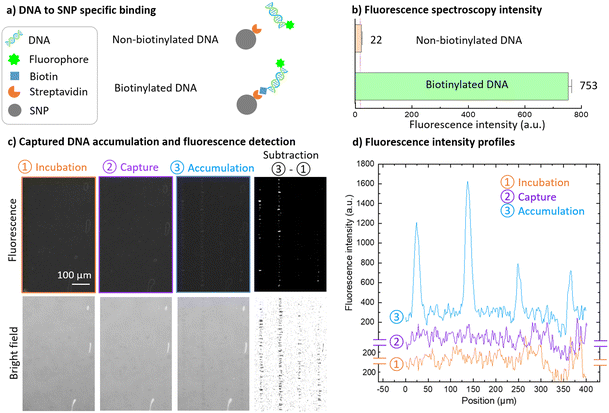
In this part, we take advantage of the possibility to accumulate SNPs on a few lines, as presented on Fig. 3, to detect amplified biotinylated and fluorescent DNA produced by PCR as previously detailed.26 In this experiment a large 12 × 4 mm2 chamber was used. A solution containing 4 μg mL−1 of fluorescent biotinylated DNA and 15 μg mL−1 of Estapor non-fluorescent carboxylated 200 nm SNPs (2.3 × 109 particles per mL) functionalized with streptavidin was introduced in the chamber. The specific biotinylated DNA binding to SNP via streptavidin–biotin interaction (Fig. 7a) was characterized by fluorescence spectroscopy using non-biotinylated DNA as a control (Fig. 7b). The solution also contains 10 mM Tris HCl, 1× Denhardt solution and 0.5× hybridization solution. Experimental details are presented in SI Fig. S2. The presence of non-labeled denatured salmon DNA fragments from the hybridization solution reduces non-biotinylated DNA non-specific binding and increases biotinylated DNA specific binding (SI, Fig. S3). Preliminary experiments using washed SNPs coupled to fluorescent DNA at a concentration of 29 μg mL−1 in Tris HCl 10 mM and Denhardt 1× solution showed that the immobile fraction was 12 ± 2%. Movie S3 shows the lateral magnetophoresis of fluorescent DNA bound to SNP. To ensure maximal DNA capture on the particles, functionalized SNPs were incubated with DNA for 10 min before exposition to the external magnetic field. SNPs were then captured by the micro-magnet array during 5 min. Lateral magnetophoresis was applied at 67 rpm for 3 min to concentrate SNPs lines onto the right wall of the 12 mm long chamber. Magnetophoresis was then reversed at 17 rpm for 3 min to position the concentrated SNP lines back in the center of the chamber. At each step (incubation, capture, accumulation), overlapping fluorescent and bright field images were acquired at the same position in the center of the chamber (Fig. 7c). Exposure parameters were kept constant for the acquisition of all images at each step, enabling direct comparison of associated fluorescence intensity profiles presented in Fig. 7d. Bright field images are shown below the corresponding fluorescence images. Additional images were computed by subtracting the images obtained at the incubation step from those obtained at the accumulation step.Fluorescence images and corresponding fluorescence intensity profiles do not exhibit any significant signal at the incubation and capture steps, while a significant signal is observed after the accumulation step. Biotinylated DNA has been previously detected by SNP capture in identical micro-magnet array and spacer conditions in a previous study by Orsini et al.26 In order to show the potential of lateral magnetophoresis and SNP accumulation for improving detection sensitivity, we worked at a reduced SNP concentration in comparison with this previous study. Therefore, we observe no significant fluorescence signal at the capture step in our experimental conditions, as observed in fluorescence image (Fig. 7c) and fluorescence intensity profile (Fig. 7d, plotted in purple), for capture step 2. After SNP accumulation, SNP lines are visible on the fluorescence image (Fig. 7c, accumulation step 3), and enhanced on the subtraction image, and a very significant signal is observed on the vertical intensity profile (Fig. 7d, accumulation step 3, plotted in blue). Four peaks can be distinguished separated by 110 μm, which correspond to 4 concentrated lines over a distance of 330 μm, allowing for biotinylated DNA detection. Fluorescence intensity profiles obtained during the incubation step (Fig. 7d, plotted in orange) where SNPs are in suspension before capture correspond to the background noise. To simply quantify biotinylated DNA detection improvement between SNP capture and accumulation steps, a peak-based signal-to-noise ratio (SNR) was estimated as follows:
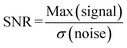 | (8) |
Conclusions
In this work, we demonstrated rapid and efficient SNP transport and accumulation within a micro-magnetofluidic chip, requiring no external fluidic actuation and offering precise position control. The technology developed consists in planar magnetic substrates, which are magnetically micro-patterned into a strip array in order to generate periodic areas exhibiting intense magnetic gradients. Applying an external homogeneous magnetic field magnetizes the nanoparticles, which subsequently become attracted towards the substrate, forming a line-array on the capture sites at the junctions between magnetic domains. Rotating the external magnetic field translates the magnetic energy landscape, inducing directional motion of the SNP lines along the substrate.This principle of potential energy landscape translation over a micro-magnet array by means of an external rotating magnetic field has been previously reported.7–9,41,42 However, in these studies, SNP stable positions on the micro-magnet arrays are discrete. One innovative contribution of our work consists in disposing a spacer on top of the micro-magnet array to smooth the potential energy landscape to which SNPs are subjected. The presence of this spacer enables a smooth and continuous displacement of the SNP lines over the substrate, which can be precisely controlled with the external magnetic field rotation angle. This characteristic represents an advantage for future applications where particle delivery at a specific location is needed. The spacer plays another important role in biochemical applications: in the absence of an external magnetic field, it allows SNPs to remain in suspension and be free to interact with molecules in the surrounding medium, during the time required for adequate incubation.
We demonstrate SNP size dependence on magnetophoresis by characterizing 100 nm and 200 nm SNP velocity as a function of the field rotation speed. A critical speed of around 500 μm s−1 was achieved for 100 nm SNP above which particles disperse and get resuspended in liquid. No critical speed was reached for 200 nm SNP in the limit of the maximal 750 rpm rotation speed delivered by our current device, at which particles reached 1.4 mm s−1 velocity without stalling nor resuspending.
The mobility of SNP was investigated in different buffers commonly used for biological analysis. Semi-quantitative SNP immobile fraction results vary from 16% to 93% depending on the buffer conditions, showing that SNP mobility is favored in low ionic strength conditions and with the presence of denatured DNA and polymers in hybridization solution.
We show SNP accumulation against a solid wall of the microfluidic chamber to concentrate fluorescence on a few SNP lines, enabling to overcome noise and detect specific fluorescent DNA signal that was not detectable at the capture step where SNPs are distributed over multiple lines.
Transport of a high number of submicrometric SNPs (∼108 particles) was implemented over centimetric distances by lateral magnetophoresis in static liquid environment. Our results constitute a proof of concept and a basis for further investigations and improvements required for practical application of this technology. Today, a gap still exists between LOC proofs of concept and the development of operational devices, owing to a lack of standardization and sophisticated microfluidic systems, resulting in manufacturing process incompatibilities.43,44 We believe our lateral magnetophoresis approach for collective and precise SNP transport represents a versatile and low-cost tool, and is promising for integrating sensitive biochemical assays into miniaturized setups, contributing towards accessible microfluidics systems for the development of lab-on-a-chip devices.
Author contributions
Conceptualization, EB, EO, SD, OC and FB; methodology, EB, EO, OC and FB; investigation, EB, CdF, OC and FB; writing – original draft preparation, EB; writing – review and editing, EB, SD, OC and FB; supervision, SD, OC and FB; project administration, SD, OC, and FB; funding acquisition, FB.Conflicts of interest
The authors disclosed the following conflicts of interest: SD is employed by MagIA diagnostics, the company developing MLFIA technology. The authors declare that this study received funding from “Pack Ambition Recherche 2021—Projet MagMol” of the Auvergne-Rhône-Alpes Region. The funders were not involved in the study design; collection, analysis, or interpretation of data; the writing of this article; or the decision to submit it for publication. SD holds position on MagIA advisory board. FB and OC are co-authors of a patent on capture kit for a molecule in sample and detection method owned by CNRS and licensed to MagIA diagnostics. OC is the co-author of a patent on a kit and method for the capture of a molecule using magnetic means owned by CNRS and licensed to MagIA diagnostics.Data availability
The raw data associated with this manuscript, which consists in images and image sequences, are available from the authors upon request.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5lc01072a
Acknowledgements
The authors thank A. Beleza-Azevedo and T. Devillers from Institut Néel (CNRS) for the SNP magnetic characterization using a VSM-SQUID magnetometer. This work was supported by a grant “Pack Ambition Recherche 2021—Projet MagMol” from the Auvergne-Rhône-Alpes Region. This work was also partially supported by the Micro and Nanotechnologies Federation (FMNT) in Grenoble.References
- N.-T. Nguyen, Microfluid. Nanofluid., 2012, 12, 1–16 CrossRef.
- R.-J. Yang, H.-H. Hou, Y.-N. Wang and L.-M. Fu, Sens. Actuators, B, 2016, 224, 1–15 CrossRef.
- N. Pamme, Lab Chip, 2006, 6, 24–38 RSC.
- M. A. M. Gijs, F. Lacharme and U. Lehmann, Chem. Rev., 2010, 110, 1518–1563 CrossRef PubMed.
- C. Ruffert, Micromachines, 2016, 7, 21 CrossRef PubMed.
- N. Pamme, Curr. Opin. Chem. Biol., 2012, 16, 436–443 CrossRef PubMed.
- S. Rampini, P. Li and G. U. Lee, Lab Chip, 2016, 16, 3645–3663 RSC.
- B. B. Yellen, R. M. Erb, H. S. Son, R. Hewlin, Jr., H. Shang and G. U. Lee, Lab Chip, 2007, 7, 1681 RSC.
- S. Rampini, P. Li, D. Gandhi, M. Mutas, Y. F. Ran, M. Carr and G. U. Lee, Sci. Rep., 2021, 11, 5302 CrossRef PubMed.
- M. Donolato, B. T. Dalslet and M. F. Hansen, Biomicrofluidics, 2012, 6, 024110 CrossRef PubMed.
- P. Tierno, F. Sagués, T. H. Johansen and T. M. Fischer, Phys. Chem. Chem. Phys., 2009, 11, 9615 RSC.
- X. Hu, S. R. Goudu, S. R. Torati, B. Lim, K. Kim and C. Kim, Lab Chip, 2016, 16, 3485–3492 RSC.
- M. Ouk and G. S. D. Beach, J. Magn. Magn. Mater., 2017, 444, 218–226 CrossRef.
- S. Van Pelt, A. Frijns and J. Den Toonder, Lab Chip, 2017, 17, 3826–3840 RSC.
- H. A. Cho, J. Hyun Min, J. Hua Wu, J. Woo Jang, C.-S. Lim and Y. Keun Kim, J. Appl. Phys., 2014, 115, 17B512 CrossRef.
- M. Fratzl, S. Delshadi, T. Devillers, F. Bruckert, O. Cugat, N. M. Dempsey and G. Blaire, Soft Matter, 2018, 14, 2671–2681 RSC.
- M. Fratzl, P. Bigotte, R. Gorbenkov, G. Goubet, P. Halfon, P. Kauffmann, D. Kirk, V. Masse, X. Payet-Burin, O. Ramel and S. Delshadi, SLAS Technol., 2024, 29, 100119 CrossRef CAS PubMed.
- A. Munaz, M. J. A. Shiddiky and N.-T. Nguyen, Biomicrofluidics, 2018, 12, 031501 CrossRef PubMed.
- A. O. Ayansiji, A. V. Dighe, A. A. Linninger and M. R. Singh, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 30208–30214 CrossRef CAS PubMed.
- G. Cai, Z. Yang, Y.-C. Chen, Y. Huang, L. Liang, S. Feng and J. Zhao, Cyborg Bionic Syst., 2023, 4, 0023 CrossRef CAS PubMed.
- K. Nandy, S. Chaudhuri, R. Ganguly and I. K. Puri, J. Magn. Magn. Mater., 2008, 320, 1398–1405 CrossRef CAS.
- J. Lim, C. Lanni, E. R. Evarts, F. Lanni, R. D. Tilton and S. A. Majetich, ACS Nano, 2011, 5, 217–226 Search PubMed.
- V. Agmo Hernández, ChemTexts, 2023, 9, 10 Search PubMed.
- O. Cugat, J. Delamare and G. Reyne, IEEE Trans. Magn., 2003, 39, 3607–3612 CrossRef.
- M. Kustov, P. Laczkowski, D. Hykel, K. Hasselbach, F. Dumas-Bouchiat, D. O’Brien, P. Kauffmann, R. Grechishkin, D. Givord, G. Reyne, O. Cugat and N. M. Dempsey, J. Appl. Phys., 2010, 108, 063914 CrossRef.
- E. Orsini, F. Bruckert, M. Weidenhaupt, O. Cugat, P. Kauffmann and S. Delshadi, Biosensors, 2025, 15, 195 CrossRef CAS.
- C. S. Neish, I. L. Martin, R. M. Henderson and J. M. Edwardson, Br. J. Pharmacol., 2002, 135, 1943–1950 CrossRef CAS PubMed.
- L. Häussling, B. Michel, H. Ringsdorf and H. Rohrer, Angew. Chem., Int. Ed. Engl., 1991, 30, 569–572 CrossRef.
- A. Moyano, E. Serrano-Pertierra, M. Salvador, J. Martínez-García, Y. Piñeiro, S. Yañez-Vilar, M. Gónzalez-Gómez, J. Rivas, M. Rivas and M. Blanco-López, Biosensors, 2020, 10, 80 CrossRef CAS PubMed.
- R. D'Agata, P. Palladino and G. Spoto, Beilstein J. Nanotechnol., 2017, 8, 1–11 CrossRef PubMed.
- C. P. Hunt, B. M. Moskowitz and S. K. Banerjee, in Rock Physics & Phase Relations, ed. T. J. Ahrens, 1995, pp. 189–204 Search PubMed.
- S. Delshadi, M. Fratzl, O. Ramel, P. Bigotte, P. Kauffmann, D. Kirk, V. Masse, M. P. Brenier-Pinchart, H. Fricker-Hidalgo, H. Pelloux, F. Bruckert, C. Charrat, O. Cugat, N. M. Dempsey, T. Devillers, P. Halfon, A. Leroy, M. Weidenhaupt and P. N. Marche, Lab Chip, 2023, 23, 645–658 RSC.
- L. E. Helseth, T. M. Fischer and T. H. Johansen, Phys. Rev. Lett., 2003, 91, 208302 Search PubMed.
- M. Donolato, P. Vavassori, M. Gobbi, M. Deryabina, M. F. Hansen, V. Metlushko, B. Ilic, M. Cantoni, D. Petti, S. Brivio and R. Bertacco, Adv. Mater., 2010, 22, 2706–2710 Search PubMed.
- S. R. Goudu, X. Hu, B. Lim, K. Kim, K. Kim, H. Kim, J. Yoon and C. Kim, IEEE Trans. Magn., 2018, 54, 1–4 Search PubMed.
- E. Rapoport and G. S. D. Beach, Appl. Phys. Lett., 2012, 100, 082401 CrossRef.
- H.-Y. Nie, M. J. Walzak, B. Berno and N. S. McIntyre, Appl. Surf. Sci., 1999, 144–145, 627–632 Search PubMed.
- M. J. Walzak, S. Flynn, R. Foerch, J. M. Hill, E. Karbashewski, A. Lin and M. Strobel, J. Adhes. Sci. Technol., 1995, 9, 1229–1248 Search PubMed.
- R. Wirix-Speetjens, W. Fyen, Xu Kaidong, Jo De Boeck and G. Borghs, IEEE Trans. Magn., 2005, 41, 4128–4133 Search PubMed.
- A. Huerta-Viga, S. R. Domingos, S. Amirjalayer and S. Woutersen, Phys. Chem. Chem. Phys., 2014, 16, 15784–15786 Search PubMed.
- R. S. Conroy, G. Zabow, J. Moreland and A. P. Koretsky, Appl. Phys. Lett., 2008, 93, 203901 CrossRef.
- L. Gao, M. A. Tahir, L. N. Virgin and B. B. Yellen, Lab Chip, 2011, 11, 4214 RSC.
- G. Gharib, İ. Bütün, Z. Muganlı, G. Kozalak, İ. Namlı, S. S. Sarraf, V. E. Ahmadi, E. Toyran, A. J. Van Wijnen and A. Koşar, Biosensors, 2022, 12, 1023 CrossRef CAS PubMed.
- M. I. Mohammed, S. Haswell and I. Gibson, Procedia Technol., 2015, 20, 54–59 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |