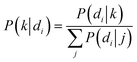Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Size-based sorting of dynamic bacterial clusters
Elham
Akbari
 a,
Jason P.
Beech
a,
Jason P.
Beech
 a,
Johannes
Kumra Ahnlide
a,
Johannes
Kumra Ahnlide
 b,
Sebastian
Wrighton
b,
Sebastian
Wrighton
 bc,
Pontus
Nordenfelt
bc,
Pontus
Nordenfelt
 b and
Jonas O.
Tegenfeldt
b and
Jonas O.
Tegenfeldt
 *a
*a
aDepartment of Physics, Division of Solid State Physics, Lund University, Lund, Sweden. E-mail: jonas.tegenfeldt@ftf.lth.se; Tel: +46 222 8063
bDepartment of Clinical Sciences, Division of Infection Medicine, Faculty of Medicine, Lund University, Lund, Sweden
cCenter for Pathophysiology, Infectiology, and Immunology, Institute of Immunology, Medical University of Vienna, 1090 Vienna, Austria
First published on 15th January 2026
Abstract
Group A Streptococcus (GAS) forms highly deformable aggregates with broad variations in size and morphology, complicating controlled separation and biological analysis. Reliable methods to isolate fractions of GAS clusters with defined properties are essential for studying host–pathogen interactions that depend on cluster size. Here, we present a simple deterministic lateral displacement (DLD) microfluidic device to separate complex suspensions of bacterial aggregates into two size-enriched fractions. We use a DLD with a small displacement angle to accommodate the large range of particle sizes above the critical size. We introduce an intermediate outlet, in addition to the conventional zigzag and displacement outlets, to collect the aggregates which exhibit a large dispersion due to their broad variety in shape and sizes close to the device critical diameter. In this way, we can demonstrate fractionation of GAS clusters with >90% purity based on effective size while causing minimal fragmentation or additional aggregation, as demonstrated by image analysis and dual-colour experiments. Finally, we show biological relevance through a live immune-cell assay, where human immune cells migrate more rapidly in the presence of larger GAS clusters than in smaller clusters or single bacteria. These results demonstrate that DLD-based separation provides biologically meaningful fractions of bacterial aggregates and enables new analyses of how cluster size influences immune responses.
1. Introduction
Streptococcus pyogenes, which is commonly referred to as group A streptococcus (GAS), is a Gram-positive human pathogenic bacterium. This human pathogen causes diseases with an estimated 700 million infected people each year, with a mortality rate of over 25% in severe cases such as necrotizing fasciitis and sepsis. A rapid recent increase in clindamycin resistance,1 the antibiotics routinely used for treating GAS infection, combined with the lack of vaccines or alternative antibiotics, makes understanding GAS pathogenicity important.GAS, is a round-shaped bacterium (coccus) that has varying tendencies to grow in long chains which form clusters when grown in vitro in a liquid medium.2,3 Based on a study by Frick et al.,4 this property of growing in clusters is crucial for adherence, the resistance to phagocytosis, and the virulence of GAS. Other studies on different bacteria such as Streptococcus pneumoniae,5 and on yeast such as Saccharomyces, have shown that bio-particles of different sizes and shapes are thought to contribute differently to virulence and colonization.6 Therefore, sorting GAS bacteria and clusters based on their physical properties can be an important first step in the study of how these properties affect pathogenicity.
Generating size-based fractions from suspensions of cultured GAS is inherently challenging because GAS form clusters when cultured.7 With cluster sizes of hundreds of bacteria,4 size distributions are large, and any sorting scheme needs to accommodate this large range. With smaller groups of bacteria forming chains,8 aspect ratios can be large. Large clusters, while more spherical than shorter chains, can have intricate three-dimensional shapes where rotational effects add to the complexity by making size a poorly defined property.9 Moreover, surface molecules which help GAS adhere to one another and to various surfaces8 allow clusters to stick together and to surfaces, leading to size distributions that vary over time.
Microfluidic systems are popular alternatives to traditional sorters and separators such as membrane filters and centrifugal elutriators and have been used for the separation of biological particles, including bacterial cells.10 These systems can separate a wide range of biological particles and cells, from bacteria,11 mammalian,12,13 and cancer cells14 to viruses, extracellular vesicles,15 and DNA.16 Label-free, microfluidics techniques functioning without the intervention of any external forces, other than those that drive flows, are among the most studied methods for the sorting of biological particles.17–20 Filter-based sorting methods have been employed for isolating biological particles, but they often face challenges such as the clogging of pores by larger cells or debris.18,21 In various studies, inertial focusing,22,23 pinched flow fractionation,24 and viscoelastic sorting25,26 have been used to isolate biological particles, including bacteria, from human physiological samples for diagnostic purposes. These methods, sort particles of different sizes and shapes using hydrodynamic forces, with viscoelastic sorting27 also utilizing elastic forces.28 In addition to size, these methods have been shown to be sensitive to other physical properties of the sample, including shape29 and deformability,21–23,25,30,31 However, there is a general lack of the application of these methods to the generation of fractions with well-defined properties from samples with highly heterogeneous sizes and morphologies such as cultures of GAS bacteria.
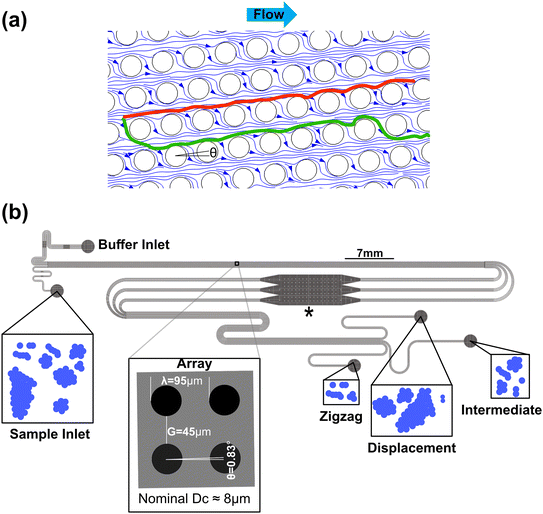
DLD is a label-free, passive, and continuous microfluidic-based sorting method with sensitivity to size,32 morphology,33,34 and deformability34 of biological particles. This sensitivity comes from interactions between particles and pillars which force particles into specific trajectories. Arrays can be designed to be more or less sensitive to different physical properties for example with optimised pillar shapes,35,36 but size is most often the dominant parameter. There are two dominant trajectories through a DLD array which are divided by a threshold diameter, known as the critical diameter, Dc, see Fig. 1a. In the ideal operation, particles smaller than Dc follow the direction of the fluid flow in a trajectory that is referred to as the zigzag trajectory (and are collected in the zigzag outlet) and those larger than Dc follow the displacement trajectory (and are collected in the displacement outlet).
For a quadratic pillar array, Dc depends on the gap size, G, between pillars and the angle, θ, at which the array is tilted with respect to the flow direction, or equivalently the periodicity, N = 1/tan![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) θ of the array along the channel. The critical diameter37,38 can be approximated by a simple expression.
θ of the array along the channel. The critical diameter37,38 can be approximated by a simple expression.
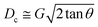 | (1) |
Instead of resorting to a complicated device with multiple separation sections on one chip, where each section removes particles of a different size range sequentially,39,40 we present a simplified solution with a deterministic lateral displacement (DLD) device based on a single separation array for separating cultured GAS bacteria into two well-defined, size-based fractions. Eqn (1) shows that we can accommodate the wide range of sizes in the displaced fraction by minimizing the array angle. However, the small array angle makes the separation sensitive to flow perturbations, and together with a sample that exhibits a wide variety of shapes, we expect particles with sizes close to the Dc to disperse, thereby contaminating the zigzag and displacement outlets. We addressed that problem by adding an intermediate outlet that collects most of the dispersed particles. This way we ensured that the collected fractions were as pure as possible with respect to size, allowing us to isolate two populations in the zigzag and displacement outlets with sufficient numbers of clusters for downstream biological assays.
We demonstrate the ability of the device to generate size-based fractions from the complex GAS cluster population. Careful comparison of GAS cluster size and morphology before and after sorting was used to detect any changes in shape or size as the sample is passed through the device. By running a mixture of two GAS cluster populations pre-stained in two different colours and quantifying dual colour clusters in the outlets, we could identify and estimate the extent of exchange between clusters in the device. Finally, a preliminary proof of concept study of the effect of GAS cluster size was performed with two distinct, size-based fractions. In this assay, immune cells (neutrophils) were imaged via time-lapse microscopy as they interacted with the two fractions containing different cluster sizes.
2. Experimental
2.1. DLD device design
The aim for the device was to generate one fraction containing singlets, doublets and small clusters of GAS up to approximately 10 μm in diameter for collection in the zigzag outlet and a second fraction containing clusters above 10 μm in diameter in the displacement outlet. Our design aims at achieving high throughput, low level of sample dilution, high purity and low level of sample break-up.It is well known that in practice, the flow angle of particles moving in a DLD as a function of particle size is not described by a sharp step-function between the direction of the average liquid flow and the direction given by the obstacle array. This can be traced back to factors related to the applied flow rates, the characteristics of the particles and the device design. For low Péclet numbers, i.e. small particles and slow flow rates, the transition between the two flow directions is gradual, due to diffusion.32 For very high concentrations, particle–particle interactions become important, causing a shift of the critical diameter to lower values as well as a slight broadening of the particle trajectories.16,41 For array angles such that the periodicity N is not an integer, additional particle trajectories can appear,42 which in practice may also blur the transition. For arrays that deviate from a square lattice such as rhombic arrays and arrays with different lateral and longitudinal distances between the pillars, the flow permeability may be anisotropic, which in turn may change the fluid flow direction locally.43–46 For high Reynolds numbers, flow anisotropy in the array may cause deviations in the flow trajectories as a function of flow rate.47 Since the Dc depends on the flow direction, these factors may introduce dispersion in the flow trajectories of the particles as well as shifts in the critical diameter. Moreover, deformable particles with non-spherical shape may change their shape and orientation dynamically as they move from pillar to pillar in the array,48 changing their effective size, which, in turn, may have a large impact on their trajectories.
The sizes of the GAS clusters range from single bacteria (1 μm) to hundreds of bacteria (many tens of micrometers). In order to create two fractions, one with single cells, doublets and small clusters and another with larger clusters, our device is required to simultaneously have a rather small critical diameter, while have large enough gaps G to allow all clusters to pass without clogging the array. From eqn (1), we can see that this can be achieved by minimizing the array angle, corresponding to using the largest possible periodicity of the array, N. Moreover, large N has been shown to improve separation for soft particles especially at high flow rates16 which makes it even more important for our bacterial clusters. Large N lead to smaller deformations at each event since for a given lateral displacement the number of post interactions scales as 1/tan(θ). Devices with small θ (large N) therefore spread the lateral driving force over a larger number of particle–obstacle interactions.
To balance the need for large N with the practical requirement to fit the device on a manageable 55 × 75 mm glass slide (considering that the large particles are displaced by 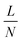 , where L is the length of the array), we chose to make a device with
, where L is the length of the array), we chose to make a device with 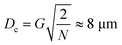 , N = 69, which corresponds to a gap of G = 45 μm. The length of the DLD array is 65
, N = 69, which corresponds to a gap of G = 45 μm. The length of the DLD array is 65![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 590 μm and it is 100 μm deep. The width is selected to be 950 μm which corresponds to all large particles that enter in the inlet channel ill also hit the wall once they reach the end of the array. To minimize the problems described above with rhombic arrays, we utilized a square array,46 rotated at an angle θ = 0.83° with respect to the flow direction to achieve N = 69. Although higher periodicities have been reported (N = 100 (ref. 49)) in the literature, we choose an intermediate value to keep the sensitivity to any perturbations to the particles trajectories within a range that we can handle. An important concern with DLD arrays is the effects of non-integer periodicities of the array, resulting in anomalous trajectories in addition to the standard zigzag and displacement trajectories. It is however mainly a problem for small N. In our case, it can be shown that the angles for these anomalous trajectories amount to less than 1.5% of the angle of the displacement trajectory. This in turn translates into 14 μm lateral displacement at the end of the device, which is significantly less than the gap between to pillars in the DLD array. We therefore conclude that the effect can be neglected for our device.
590 μm and it is 100 μm deep. The width is selected to be 950 μm which corresponds to all large particles that enter in the inlet channel ill also hit the wall once they reach the end of the array. To minimize the problems described above with rhombic arrays, we utilized a square array,46 rotated at an angle θ = 0.83° with respect to the flow direction to achieve N = 69. Although higher periodicities have been reported (N = 100 (ref. 49)) in the literature, we choose an intermediate value to keep the sensitivity to any perturbations to the particles trajectories within a range that we can handle. An important concern with DLD arrays is the effects of non-integer periodicities of the array, resulting in anomalous trajectories in addition to the standard zigzag and displacement trajectories. It is however mainly a problem for small N. In our case, it can be shown that the angles for these anomalous trajectories amount to less than 1.5% of the angle of the displacement trajectory. This in turn translates into 14 μm lateral displacement at the end of the device, which is significantly less than the gap between to pillars in the DLD array. We therefore conclude that the effect can be neglected for our device.
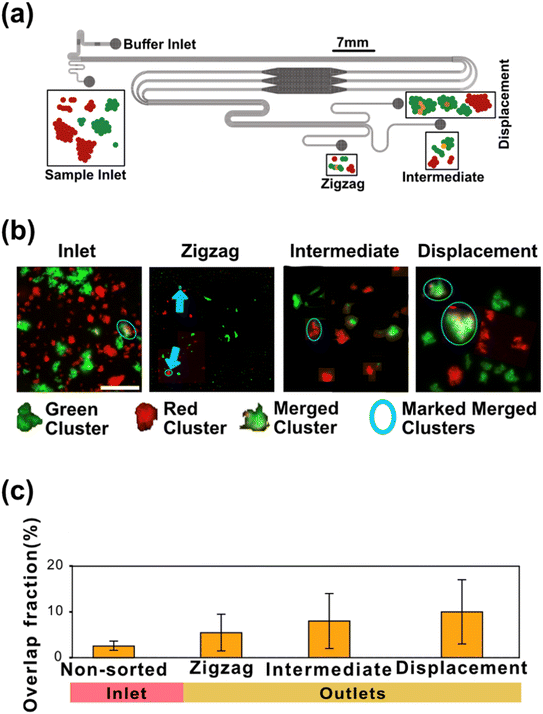
Finally, we want to minimize the volume of liquids handled by the device and minimize dilution of the sample by keeping it as narrow as possible with a width corresponding to one displacement array, the length of the DLD array divided by N. Due to the small array angle, the device becomes sensitive to any perturbations of the flow due to overall misalignment of the flow. With a narrow device, the performance is affected by local changes in flow patterns close to the wall.50 While the wall effect can be compensated for by careful and high-resolution modifications of the pillars at the boundary of the array,51,52 other effects, e.g. from complex particles are not. We therefore resort to a simple solution. To ensure the purity of our two collected fractions, we introduce a third outlet to our device, in addition to the zigzag and displacement outlets, which we call intermediate outlet, that collects all sample that does not move consistently in either of the zigzag or displacement trajectories (see Fig. 1b for detailed description of the device). We expect to gain in terms of purity of the sample and simplicity of the device design at a cost of losing some of the sample.
Fig. 1b shows a schematic of the device with sample inputs and outputs and illustrations depicting a typical GAS sample. The inlet and outlet channels are designed to have equal fluidic resistances per unit width respectively although the outlet channels are longer with higher resistances, which we have observed to lead to stable, laminar flow throughout the separation array. Finite element simulations used in the design process can be found in S1.
2.2. DLD device fabrication
Full details of device fabrication protocols and device characterization can be found in the S2. In short, mask-based UV-lithography was used to create a master mould in a 100 μm thick dry film resist (SUEX®, K100, DJ Microlaminates, Sudbury, Massachusetts, USA) laminated onto a 4″ silicon wafer and this was then used to create replicas in PDMS that were plasma bonded to glass slides.2.3. Sample preparation: bacteria culture
GAS strains of the M1 serotype, SF370 were cultured statically in Todd-Hewitt Yeast media (THY) at 37 °C, 5% CO2. Three weeks before culture, the strain was maintained on THY agar plates. Bacteria were grown to the logarithmic phase by diluting an overnight culture 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 in fresh THY. After dilution, bacteria were grown for around 2–2.5 h and then washed 2 times with PBS. After washing with phosphate buffer saline (PBS, Merck Life Science AB, Stockholm, Sweden), the bacteria were heat-killed. This was achieved using 15 minutes on ice, followed by a 5-minute heat shock at 80 °C and a final 15 minutes on ice. The heat-killed bacteria were then stained according to the type of experiment performed. For the size-based separation characterisations and bioassays GAS were stained with Oregon Green (OG) 488-X succinimidyl ester (ThermoFisher, Massachusetts, United States) at a concentration of 20 mg mL−1 (a 2 μL dye solution was added to 1 mL of bacteria at 37 °C under gentle rotation whilst protected from light for 30 min). The two-colour study of cluster dynamics was done using both OG and Wheat Germ Agglutinin (WGA) conjugated fluorescent dye 640 nm emission (CR®640, Biotium, Inc., Fremont, CA, United States), (100 μL of dye solution prepared in 0.15 M (molar) NaCl buffer solution with a concentration of 1 mg mL−1 was added to twice buffer-washed bacteria and incubated at room temperature under gentle rotation, protected from light, for 20 min). For all experiments, bacterial samples were diluted 40 times with bovine serum albumin (BSA, Merck Life Science AB, Stockholm, Sweden) 1 w/v% in water and a total of 1 mL of sample prepared this way was used as inputs for sorting. Heat killing of bacteria is a common practice and is not expected to change the short-term interaction of bacteria with immune cells.53 Note that for work with pathogenic bacteria, adherence to standard safety routines and precautions including wearing personal protective equipment is a requirement. This includes using protective gloves, goggles and face shields as appropriate as well as working in a ventilated dedicated workspace. Routines must be in place to ensure that users are properly trained accordingly.
20 in fresh THY. After dilution, bacteria were grown for around 2–2.5 h and then washed 2 times with PBS. After washing with phosphate buffer saline (PBS, Merck Life Science AB, Stockholm, Sweden), the bacteria were heat-killed. This was achieved using 15 minutes on ice, followed by a 5-minute heat shock at 80 °C and a final 15 minutes on ice. The heat-killed bacteria were then stained according to the type of experiment performed. For the size-based separation characterisations and bioassays GAS were stained with Oregon Green (OG) 488-X succinimidyl ester (ThermoFisher, Massachusetts, United States) at a concentration of 20 mg mL−1 (a 2 μL dye solution was added to 1 mL of bacteria at 37 °C under gentle rotation whilst protected from light for 30 min). The two-colour study of cluster dynamics was done using both OG and Wheat Germ Agglutinin (WGA) conjugated fluorescent dye 640 nm emission (CR®640, Biotium, Inc., Fremont, CA, United States), (100 μL of dye solution prepared in 0.15 M (molar) NaCl buffer solution with a concentration of 1 mg mL−1 was added to twice buffer-washed bacteria and incubated at room temperature under gentle rotation, protected from light, for 20 min). For all experiments, bacterial samples were diluted 40 times with bovine serum albumin (BSA, Merck Life Science AB, Stockholm, Sweden) 1 w/v% in water and a total of 1 mL of sample prepared this way was used as inputs for sorting. Heat killing of bacteria is a common practice and is not expected to change the short-term interaction of bacteria with immune cells.53 Note that for work with pathogenic bacteria, adherence to standard safety routines and precautions including wearing personal protective equipment is a requirement. This includes using protective gloves, goggles and face shields as appropriate as well as working in a ventilated dedicated workspace. Routines must be in place to ensure that users are properly trained accordingly.
2.4. Neutrophil preparation and phagocytosis assay captured by time-lapse microscopy
Neutrophils were isolated from a single healthy donor for each experiment. All experiments with blood cells were performed in accordance with the Guidelines of the Declaration of Helsinki, and Experiments were approved by the Swedish Ethical Review Authority at Lund, permit number 2015/801. Informed consent was obtained from all human participants in this study. Isolation was done using a Polymorphoprep density gradient (Serumwerk Bernburg AG, Bernburg, Germany) according to the manufacturer's instructions. If there was any contamination with erythrocytes then the cells were pelleted and the supernatant removed. The cells were then resuspended in 1 mL of Milli-Q (Merck Millipore, Darmstadt, Germany), immediately followed by 14 mL of 1× PBS to reinstate an isotonic state. Cells were pelleted again and lysis was repeated until the cell pellet appeared clear of erythrocytes. Cells were then resuspended in Na Media (5.6 mM glucose, 127 mM NaCl, 10.8 mM KCl, 2.4 mM KH2PO4, 1.6 mM MgSO4, 10 mM HEPES, 1.8 mM CaCl2; pH adjusted to 7.3 with NaOH). Cells were counted using a Bürker chamber and the concentration was adjusted to 3 × 105 mL. The cell-permeant nucleic acid stain, Hoechst 33342 (ThermoFisher Scientific, Waltham, MA USA) was added at a final concentration of 2 μM and the cells were then kept on ice before further use. Imaging was done using 35 mm dishes with 20 mm inner chambers that were fitted with untreated No 1.5 coverslips (Cellvis, Mountain View, CA, USA). The coverslip was coated with 20 μg mL−1 of fibronectin after which it was blocked with 1% BSA in PBS. For the phagocytosis experiment, the bacteria were first heat-killed, stained as described above and added directly to the coated coverslips after sorting using the DLD device. Clusters were allowed to settle and adhere to the coated coverslip. The supernatant was then gently aspirated and 1 mL of the prepared neutrophil suspension was added before transferring the dish to the microscope with a stage incubator for time-lapse live cell imaging (37 °C, 5% CO2). The dish was left undisturbed for 30 minutes to allow the neutrophils to adhere before time-lapse microscopy was initiated at a rate of one frame every second minute.2.5. Fluidics and microscopy
Flow was generated in our device by keeping the outlets at ambient pressure and applying nitrogen at overpressures that were carefully adjusted to obtain a stable and even flow straight through the pillar array. The pressures were applied to two 15 mL tubes using an MFCS-4C pressure controller (Fluigent, Paris, France), specifically 140 mbar for the sample and 150 mbar for the buffer. The resulting flow rate through the device is 30 ± 2 μL min−1 corresponding to an average shear rate of 210 s−1 and, assuming the viscosity of water, a shear stress of 0.2 Pa. This is within typical ranges for biofilm studies, under physiological conditions in the airways and in e.g. microfluidic perfusion systems.54The fluids flow via 768 μm inner diameter capillaries from these tubes into the device. Devices were prepared by rinsing with the running buffer (BSA 1% w/v) for 10 min prior to running the samples. Because GAS have a higher density than the running buffer, they tend to sediment in the sample collection outlets where the concentration can become high enough that clusters interact and can stick together. To avoid this, separated fractions were moved off-chip directly after sorting by allowing them to flow downwards out of the device, through short capillary tubes. The large (displacement) and middle-sized (intermediate) fractions flowed directly into two separate wells of an Ibidi, microscope slide-based well plate (Ibidi μ-slides, 8-well high, Ibidi GmbH, Gräfelfing, Germany) coated with fibronectin. The small fraction (zigzag), which was at too high a concentration for both image analysis and the phagocytosis assay, flowed first into a gently shaken 1.5 mL centrifuge tube half-filled with running buffer to first decrease the concentration and was then transferred to a third well of the fibronectin-covered Ibidi slide. Individual GAS cells and clusters were allowed to settle and adhere to the fibronectin-covered surface of the slide. Orienting the device and collecting the sample in this way minimized the effects of sedimentation inside the capillaries which tended to lead to aggregation post separation. After collection in the petri dishes the separated fractions were left undisturbed for 30 minutes to assure the sedimentation of bacteria and clusters onto the bottom of the glass slides where they could be imaged using fluorescent microscopy. All subsequent image analyses were performed on samples prepared in this way. Phagocytosis assays were subsequently performed on samples prepared the same way but refrigerated for 2 hours and only including the zigzag and displacement fractions.
All fluidics experiments were performed under observation using an inverted epifluorescence microscope (Nikon Eclipse TE2000-U, Nikon Corporation, Tokyo, Japan) in transmission mode together with a scientific CMOS camera (Andor NEO sCMOS, Andor Technology, Belfast, Northern Ireland) at various magnifications (4×, 10×, 20× and 50×). Size distributions of GAS clusters in pre and post sorted samples together with two colour cluster dynamics measurements were done on a similar microscope (Nikon Eclipse Ti, Nikon Corporation, Tokyo, Japan) with cMOS camera (ORCA-flash 4.0, Hamamatsu Photonics, Japan) or an EMCCD camera (Andor iXon, Andor Technology, Belfast, Northern Ireland), with FITC and Cy5 filter cubes and an LED light source (Sola light engine, Lumencore, Beaverton, Oregon, USA).
For the immuno assay the microscope (Nikon Ti2, Nikon Corporation, Tokyo, Japan) featured a cage incubator (Okolab, Ottaviano, Italy) to allow for live cell imaging (37 °C, 5% CO2). Images were acquired with a 20×/0.75 N.A. Plan Apo λ objective (Nikon) with an additional 1.5× zoom (Nikon OptoVar, Nikon Corporation, Tokyo, Japan). All images were captured using a DS-Qi2 camera (Nikon Corporation, Tokyo, Japan), giving a size of the field of view of 1200 μm × 797 μm.
Further details can be found in S3 and S4.
2.6. Image and statistical analysis
Automated scans with multiple fields of view were captured of non-sorted and sorted samples in Ibidi μ-slide wells. A full description of the analysis pipeline can be found in the S4. Briefly, images were classified into two main types using signal to background noise analysis. Those containing many single bacteria and small clusters (non-sorted and small sorted fractions) were analysed using Ilastik, a deep learning computer vision software55 which is time and compute expensive but gives much better results for the semantic segmentation of small, high density objects with low signal to noise. Images with fewer, larger objects, and better resulting signal to noise, were segmented into background and foreground using intensity thresholding with Otsu's algorithm. Images were also manually inspected in Fiji56 at a range of contrast levels to ensure that no small objects were missed in the segmentation process. Automated image analyses returning unrealistic size values (e.g. outside the expected range of between 1.8 and 500 pixels at 20× magnification) where rerun in Ilastik under increased user supervision. Two-colour cluster analysis involved performing all of the above in two separate colour channels and characterising and counting clusters of both single and mixed colours. For the cell motility assay, neutrophils were segmented using the pretrained StarDist 2D_versatile_fluo model57 on Gaussian-filtered nuclear channel images and tracked across frames by solving a linear assignment problem based on Euclidean distances between cell centroids. The tracking code is available at at https://github.com/ahnlabb/sbsdbc.3. Results and discussion
3.1. Sorting of GAS clusters by size
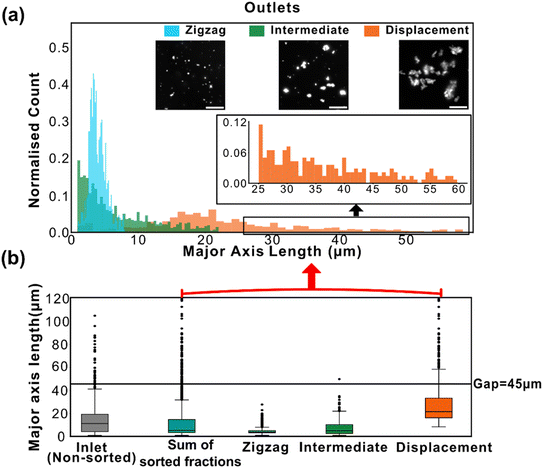
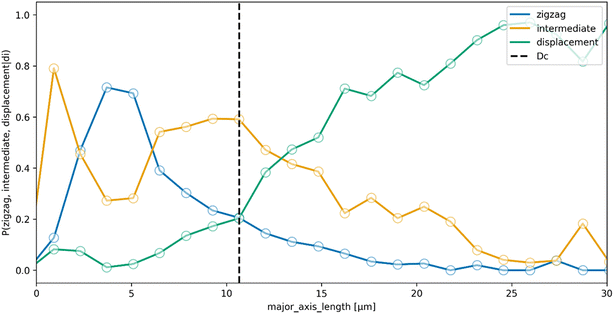
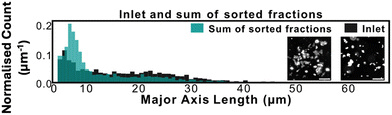
GAS clusters were separated into distinct size fractions. Fig. 2 shows example microscopy images of GAS samples after separation with obvious differences in the sizes and numbers of clusters. As can be seen in Fig. 2a and b, there is a significant difference in major axis length between the collected zigzag and displacement fractions, while the intermediate fraction major axis lengths span over a broader range, overlapping with the two other fractions. This clearly shows how the additional intermediate outlet successfully captures particles that move with trajectories outside the zigzag and displacement modes which would, greatly decrease the purity of the collected fractions in a conventional two-outlet device. Fig. 2b also shows that there are clusters in the separated fractions that, with a major axis up to ≈120 μm, are larger than the gaps in the array (45 μm wide and 100 μm deep), indicating that either they are narrow enough to fit through the array and are rotating in order to do so, or unlike microspheres, they are able to deform and squeeze through the gaps. A third possibility is that the clusters dynamically break up and reform in the device before we perform the image analysis as indicated by the two-colour mixing experiment.To estimate the probabilities of a particle of a given size to end up in each one of the three outlets we use an Bayesian approach. We used the measured size distributions of the bio particles (mainly bacterial clusters) in the three outlets to derive the routing probabilities, see Fig. 3. The probability of a particle of size di ending up in outlet k is given by:
We find that the purity for the small fraction is 91% and for the large fraction is 90%, with yields 74% for the small and 73% for the large. Expressions for the purities and yields are defined in S4.
3.2. GAS cluster dynamics during separations
To assess the cluster breakup and reaggregation in our experiments, we analysed the problem from two perspectives. To start with, we compared the size distributions of the unsorted (inlet) and the pooled outlet subpopulations, and found an increase among the small clusters. This raises the question whether the change in size distribution is due to selective losses or due to breakup of the clusters. We therefore followed up by mixing two identical bacterial cluster populations that differ only in colour. The mixture was studied in bulk as well as in our separation device.We plotted the sum of all measured clusters in the three outlets and compared the resulting distribution to that measured for the sample that we introduced in the inlet, see Fig. 4. We observed a moderate decrease in relative abundance of large clusters. This can either be due some large clusters not entering the device or being stuck in the device. Alternatively, this could be due to large clusters breaking into smaller ones. Since we could not see any major clogging in our device, and since we observed a noticeable increase in the relative abundance of small clusters (with major axis length <10 μm), we can speculate that some clusters break into smaller pieces. Note that the data is normalised as a probability density distribution so that any increase in small particles by definition leads to a smaller fraction of larger particles. The apparent decrease in larger particles is therefore expected to be somewhat exaggerated.
To test for any breakup and reaggregation of the clusters in more detail during DLD separation, we performed a two-colour mixing experiment. In these experiments two identical suspensions of GAS clusters are stained each with a different colour and subsequently mixed immediately before sorting. The final concentration of the mixture is the same as that of the bacterial solution used for sorting GAS (50 μL of each sample, resulting in a final volume of 100 μL in 4 mL of 1% BSA buffer). Any clusters containing both colours must have formed based on an exchange between the clusters. Fig. 5 shows how this experiment was done. The separation process took 30 minutes, with the inlet bacterial sample continuously shaken in 15 mL Falcon tubes at 20 rpm. After collection at the outlets, the samples were left to rest for 30 minutes to allow sedimentation at the bottom of Petri dishes. Fig. 5b shows examples of clusters with those containing both colours marked with a blue ring. Due to the finite time between mixing the two suspensions and taking the images before loading them into the device, during which clusters can interact and merge, we detect a small number of two-colour clusters in the inlet population. For the collected outlet fractions, there is a small but significant increase in the numbers of two-colour clusters when compared to the inlet. We can also observe that the number of mixed clusters is greater for the intermediate and the displacement outlets. This can be explained by two mechanisms. First, it is reasonable to assume that clusters consisting of a combination of two clusters are larger than other clusters and therefore are found in the displacement outlet. Second, since the contact between the particles and the pillars in the DLD array is significantly more frequent for displacement mode than for zigzag mode, roughly by a factor of N, the shear experienced by the large clusters is greater, which may cause them to shed more smaller clusters, which in turn may bind randomly to the surrounding clusters. Note that care needs to be taken when interpreting the absolute numbers of clusters because the high density of clusters in the images may lead to cluster overlap, even if they have not physically joined. Taken together the above results show a relative clustering of about 10% for the outlets. See Fig. 5c. We can thus conclude that we have a good size-based separation of GAS clusters with only minor cluster breakup and merging taking place in the device with sufficient GAS clusters in each fraction for bioassays. Control experiments of cluster dynamics in bulk are described in S5.
3.3. Effect of GAS cluster size on immune cell behaviour
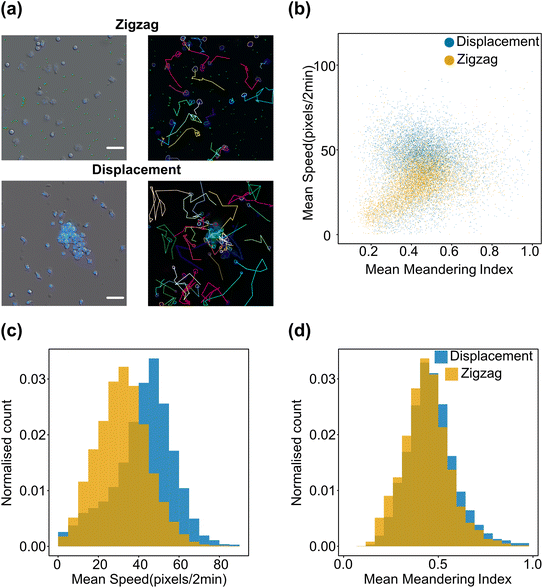
To show that our device is generating two subpopulations with sufficient sample volume and purity for bioassays, we performed a cell motility study where we exposed neutrophils to each of the two sorted subpopulations of the bacterial clusters (singlets/doublets and clusters up to ∼10 μm in one fraction, and clusters larger than ∼10 μm in the other). The assay demonstrates that neutrophils are more mobile and exhibit faster migration speed when exposed to large bacterial clusters. However, the meandering index, which quantifies directionality, does not show a significant difference suggesting that the size of clusters of GAS bacteria may not contribute to chemotactic signalling between immune cells and bacteria. This observation is also evident in the scatter plot, where, although distinct regions correspond to each sample, the difference is primarily reflected in the speed of neutrophil cells rather than their meandering index. While very preliminary, this proof-of-concept analysis shows that DLD-based separation could be helpful to start teasing apart the complex biology underlying host–bacteria interactions. See Fig. 6.4. Conclusions
We have demonstrated that we can overcome the challenges of separating GAS bacteria suspensions that have very broad distributions in size and morphology using a simple DLD device with a single Dc. The large N allows us to handle a broad range of particle sizes without clogging and it minimizes deleterious effects of shear on the clusters. The device is narrow which minimizes the dilution of the sample. We demonstrate the usefulness of the intermediate outlet to remove particles with sizes close to Dc, thereby improving the purity of the zigzag and displacement output fractions. There is a minimal change of the cluster size distribution and the device generates two populations of high purity. We were able collect sufficient sample for a proof of principle of downstream bio assays by testing the activity of neutrophils on collected fractions. To fully address the biological question, more extensive studies are needed. These may be based on pure size-based sorting or they may be complemented by including the effect of shape, deformability and other physical characteristics on the particle sorting using DLD, which, in turn, will pave the way for a refined separation of, not only GAS populations, but also a plethora of other types of complex and dynamic samples.Conflicts of interest
There are no conflicts to declare.Data availability
Data for this article, including spread sheets, videos and images are available at Harvard Dataverse at https://dataverse.harvard.edu/dataverse/GAS_clusters_DLD_tegenfeldt_lab. Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5lc01111f.Acknowledgements
We acknowledge financial support by the Swedish Research Council (grants number 2019-02355 and 2019-04102), and from NanoLund (grants number p20-2019, s01-2024, and staff01-2020). All device fabrication was performed in the cleanroom of Lund Nano Lab at Lund University. The authors are indebted to Christelle Prinz for extensive and thoughtful feedback that greatly strengthened the manuscript's focus and readability.References
- S. Lapthorne, R. McWade, N. Scanlon, S. Ní Bhaoill, A. Page, C. O’Donnell, G. Dornikova, M. Hannan, B. Lynch, M. Lynch and D. Brady, Access Microbiol., 2024, 6, 000772.v4 Search PubMed.
- J. R. Carapetis, A. C. Steer, E. K. Mulholland and M. Weber, Lancet Infect. Dis., 2005, 5, 685–694 Search PubMed.
- T. de Neergaard, A. Bläckberg, H. Ivarsson, S. Thomasson, V. K. Ahnlide, S. Chowdhury, H. Khakzad, W. Bahnan, J. Malmström, M. Rasmussen and P. Nordenfelt, Microbiol. Spectrum, 2022, 10, e02486-22 CrossRef PubMed.
- I. M. Frick, M. Morgelin and L. Bjorck, Mol. Microbiol., 2000, 37, 1232–1247 CrossRef CAS PubMed.
- J. L. Rodriguez, A. B. Dalia and J. N. Weiser, Infect. Immun., 2012, 80, 3454–3459 CrossRef CAS PubMed.
- A. B. Dalia and J. N. Weiser, Cell Host Microbe, 2011, 10, 486–496 CrossRef CAS PubMed.
- P. P. Cleary and A. Larkin, J. Bacteriol., 1979, 140, 1090–1097 CrossRef CAS PubMed.
- A. Matysik, F. K. Ho, A. Q. Ler Tan, A. Vajjala and K. A. Kline, Biofilm, 2020, 2, 100013 Search PubMed.
- S. H. Au, J. Edd, A. E. Stoddard, K. H. K. Wong, F. Fachin, S. Maheswaran, D. A. Haber, S. L. Stott, R. Kapur and M. Toner, Sci. Rep., 2017, 7, 2433 CrossRef PubMed.
- T. Laurell, F. Petersson and A. Nilsson, Chem. Soc. Rev., 2007, 36, 492–506 RSC.
- J. P. Beech, B. D. Ho, G. Garriss, V. Oliveira, B. Henriques-Normark and J. O. Tegenfeldt, Anal. Chim. Acta, 2018, 1000, 223–231 CrossRef CAS PubMed.
- M. M. Wang, E. Tu, D. E. Raymond, J. M. Yang, H. Zhang, N. Hagen, B. Dees, E. M. Mercer, A. H. Forster, I. Kariv, P. J. Marchand and W. F. Butler, Nat. Biotechnol., 2005, 23, 83–87 CrossRef CAS PubMed.
- S. C. Hur, A. J. Mach and D. Di Carlo, Biomicrofluidics, 2011, 5, 22206 CrossRef PubMed.
- A. Mishra, T. D. Dubash, J. F. Edd, M. K. Jewett, S. G. Garre, N. M. Karabacak, D. C. Rabe, B. R. Mutlu, J. R. Walsh, R. Kapur, S. L. Stott, S. Maheswaran, D. A. Haber and M. Toner, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 16839–16847 Search PubMed.
- M. Bayareh, Chem. Eng. Process., 2020, 153, 107984 Search PubMed.
- O. E. Strom, J. P. Beech and J. O. Tegenfeldt, Micromachines, 2022, 13, 1754 Search PubMed.
- T. Zhang, D. Di Carlo, C. T. Lim, T. Zhou, G. Tian, T. Tang, A. Q. Shen, W. Li, M. Li, Y. Yang, K. Goda, R. Yan, C. Lei, Y. Hosokawa and Y. Yalikun, Biotechnol. Adv., 2024, 71, 108317 Search PubMed.
- C. W. Shields Iv, C. D. Reyes and G. P. Lpez, Lab Chip, 2015, 15, 1230–1249 Search PubMed.
- G. M. Whitesides, Nature, 2006, 442, 368–373 CrossRef CAS PubMed.
- E. K. Sackmann, A. L. Fulton and D. J. Beebe, Nature, 2014, 507, 181–189 Search PubMed.
- A. Farahinia, W. J. Zhang and I. Badea, J. Sci.:Adv. Mater. Devices, 2021, 6, 303–320 CAS.
- J. M. Martel and M. Toner, Annu. Rev. Biomed. Eng., 2014, 16, 371–396 Search PubMed.
- M. Masaeli, E. Sollier, H. Amini, W. B. Mao, K. Camacho, N. Doshi, S. Mitragotri, A. Alexeev and D. Di Carlo, Phys. Rev. X, 2012, 2, 031017 Search PubMed.
- G. de Timary, C. J. Rousseau, L. Van Melderen and B. Scheid, Lab Chip, 2023, 23, 659–670 RSC.
- P. Liu, H. Liu, D. Yuan, D. Jang, S. Yan and M. Li, Anal. Chem., 2021, 93, 1586–1595 Search PubMed.
- T. Zhang, A. K. Cain, L. Semenec, L. Liu, Y. Hosokawa, D. W. Inglis, Y. Yalikun and M. Li, Anal. Chem., 2023, 95, 2561–2569 Search PubMed.
- P. Liu, H. Liu, L. Semenec, D. Yuan, S. Yan, A. K. Cain and M. Li, Microsyst. Nanoeng., 2022, 8, 7 Search PubMed.
- Y. Cho, M. H. Lee, S. Lee and Y. Cho, Microchim. Acta, 2024, 191, 738 Search PubMed.
- L. Wang, G. Qian, K. Wang, Z. Wu, H. Yan, L. Shi and T. Zhou, J. Chromatogr. A, 2024, 1730, 465126 Search PubMed.
- J. Zhou and I. Papautsky, Microsyst. Nanoeng., 2020, 6, 113 Search PubMed.
- D. Yuan, S. H. Tan, Q. B. Zhao, S. Yan, R. Sluyter, N. T. Nguyen, J. Zhang and W. H. Li, RSC Adv., 2017, 7, 3461–3469 RSC.
- L. R. Huang, E. C. Cox, R. H. Austin and J. C. Sturm, Science, 2004, 304, 987–990 CrossRef CAS PubMed.
- S. H. Holm, J. P. Beech, M. P. Barrett and J. O. Tegenfeldt, Lab Chip, 2011, 11, 1326–1332 RSC.
- J. P. Beech, S. H. Holm, K. Adolfsson and J. O. Tegenfeldt, Lab Chip, 2012, 12, 1048–1051 Search PubMed.
- S. Ranjan, K. K. Zeming, R. Jureen, D. Fisher and Y. Zhang, Lab Chip, 2014, 14, 4250–4262 RSC.
- K. K. Zeming, S. Ranjan and Y. Zhang, Nat. Commun., 2013, 4, 1625 CrossRef PubMed.
- J. A. Davis, Microfluidic separation of blood components through deterministic lateral displacement, PhD, Princeton University, 2008 Search PubMed.
- D. W. Inglis, J. A. Davis, R. H. Austin and J. C. Sturm, Lab Chip, 2006, 6, 655–658 Search PubMed.
- S. H. Holm, J. P. Beech, M. P. Barrett and J. O. Tegenfeldt, Anal. Methods, 2016, 8, 3291–3300 RSC.
- J. A. Davis, D. W. Inglis, K. J. Morton, D. A. Lawrence, L. R. Huang, S. Y. Chou, J. C. Sturm and R. H. Austin, Proc. Natl. Acad. Sci. U. S. A., 2006, 103, 14779–14784 Search PubMed.
- S. H. Holm, Z. M. Zhang, J. P. Beech, G. Gompper, D. A. Fedosov and J. O. Tegenfeldt, Phys. Rev. Appl., 2019, 12, 014051 Search PubMed.
- B. R. Long, M. Heller, J. P. Beech, H. Linke, H. Bruus and J. O. Tegenfeldt, Phys. Rev. E:Stat., Nonlinear, Soft Matter Phys., 2008, 78, 046304 Search PubMed.
- S.-C. Kim, B. H. Wunsch, H. Hu, J. T. Smith, R. H. Austin and G. Stolovitzky, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, E5034–E5041 Search PubMed.
- R. Quek, D. V. Le and K. H. Chiam, Phys. Rev. E:Stat., Nonlinear, Soft Matter Phys., 2011, 83, 056301 CrossRef PubMed.
- T. Kulrattanarak, R. G. van der Sman, Y. S. Lubbersen, C. G. Schroen, H. T. Pham, P. M. Sarro and R. M. Boom, J. Colloid Interface Sci., 2011, 354, 7–14 Search PubMed.
- R. Vernekar, T. Kruger, K. Loutherback, K. Morton and D. W. Inglis, Lab Chip, 2017, 17, 3318–3330 Search PubMed.
- C. Mallorie, R. Vernekar, B. Owen, D. W. Inglis and T. Krüger, Phys. Rev. Fluids, 2024, 9, 024203 CrossRef.
- D. W. Inglis, R. E. Nordon, J. P. Beech and G. Rosengarten, Micromachines, 2021, 12(7), 783 Search PubMed.
- K. Loutherback, K. S. Chou, J. Newman, J. Puchalla, R. H. Austin and J. C. Sturm, Microfluid. Nanofluid., 2010, 9, 1143–1149 CrossRef.
- E. Pariset, C. Pudda, F. Boizot, N. Verplanck, J. Berthier, A. Thuaire and V. Agache, Small, 2017, 13, 1701901 Search PubMed.
- D. W. Inglis, Appl. Phys. Lett., 2009, 94, 013510 CrossRef.
- D. Inglis, R. Vernekar, T. Krüger and S. Feng, Microfluid. Nanofluid., 2020, 24, 18 CrossRef.
- P. Nordenfelt, S. Waldemarson, A. Linder, M. Mörgelin, C. Karlsson, J. Malmström and L. Björck, J. Exp. Med., 2012, 209, 2367–2381 CrossRef CAS PubMed.
- L. Kim, Y.-C. Toh, J. Voldman and H. Yu, Lab Chip, 2007, 7, 681–694 RSC.
- S. Berg, D. Kutra, T. Kroeger, C. N. Straehle, B. X. Kausler, C. Haubold, M. Schiegg, J. Ales, T. Beier, M. Rudy, K. Eren, J. I. Cervantes, B. Xu, F. Beuttenmueller, A. Wolny, C. Zhang, U. Koethe, F. A. Hamprecht and A. Kreshuk, Nat. Methods, 2019, 16, 1226–1232 CrossRef CAS PubMed.
- J. Schindelin, I. Arganda-Carreras, E. Frise, V. Kaynig, M. Longair, T. Pietzsch, S. Preibisch, C. Rueden, S. Saalfeld, B. Schmid, J. Y. Tinevez, D. J. White, V. Hartenstein, K. Eliceiri, P. Tomancak and A. Cardona, Nat. Methods, 2012, 9, 676–682 Search PubMed.
- U. Schmidt, M. Weigert, C. Broaddus and G. Myers, Cell Detection with Star-Convex Polygons, International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), 2018, pp. 265–273 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |