Open Access Article
Open Access ArticleEV-Blade: an automated centrifugal-pneumatic cartridge for size- and affinity-based exosome isolation from whole blood
Lucas
Poncelet
 ab,
Keith J.
Morton
a,
Matthew
Shiu
a,
Gaétan
Veilleux
a,
Chantal
Richer
b,
Liviu
Clime
a,
Daniel
Sinnett
bc and
Teodor
Veres
*a
ab,
Keith J.
Morton
a,
Matthew
Shiu
a,
Gaétan
Veilleux
a,
Chantal
Richer
b,
Liviu
Clime
a,
Daniel
Sinnett
bc and
Teodor
Veres
*a
aLife Sciences Division, National Research Council of Canada, Boucherville, Quebec, Canada. E-mail: teodor.veres@cnrc-nrc.gc.ca
bDivision of Hemato-Oncology, Sainte-Justine Hospital Pediatric Research Center, Montreal, Quebec, Canada
cPediatrics, University of Montreal Faculty of Medicine, Montreal, Quebec, Canada
First published on 20th January 2026
Abstract
Extracellular vesicles (EVs), especially the exosome-sized subset are increasingly exploited as minimally invasive cancer biomarkers. These small vesicles are abundant in biofluids and play crucial roles in intercellular communication and disease progression by transporting bioactive molecules. Exosomes offer distinct diagnostic and prognostic advantages over traditional cancer biomarkers, but purifying and extracting exosomes from blood remains challenging. There is a need to simply and cost-effectively isolate exosomes from milliliter quantities of whole blood for transcriptional and other omics-based research. Addressing this gap, we propose a microfluidic cartridge, the EV-Blade, for size and affinity-based purification of exosomes on a centrifugal microfluidic platform. We demonstrate a method to automate exosome purification from whole blood samples on a single microfluidic cartridge. The EV-Blade system combines blood centrifugation, plasma filtration for EV size selection and immunomagnetic capture using functionalized superparamagnetic nanoparticles targeting CD9, CD63, and CD81 exosomal surface proteins. We report on the device performance, purity of exosome recovery and the quality of RNA collected following on-chip EV lysis. We use this automated method to detect relevant long coding and non-coding RNA transcripts in circulating blood exosomes, showcasing the EV-Blade for use in cancer patient risk stratification. The system presented herein represents a significant advancement in exosome purification, offering a robust and automated platform for liquid biopsy-based cancer research and clinical applications. This innovation holds promise for cancer diagnosis, prognosis, and monitoring through non-invasive biomarkers.
Introduction
Extracellular vesicles (EVs), including exosomes, have emerged as promising vehicles for biomarker discovery in cancer research.1 Here we define exosomes as a subset of small EVs (sEVs)2 with an outer diameter under 200 nm and bearing various combinations of the CD9, CD63 and CD81 transmembrane proteins.3 They are secreted by various cells in the body and can be found abundantly in biofluids such as blood, urine, and saliva.4 They play critical roles in intercellular communication, disease progression, and therapeutic resistance by shuttling various bioactive molecules, including proteins, metabolites, and RNA.5–9 Consequently, exosomes have attracted significant attention as potential sources for non-invasive diagnostic and prognostic information in cancer patients.10–12In the context of cancer, exosomes are implicated in multiple processes, including modulation of the tumor microenvironment, drug resistance, and the establishment of pre-metastatic niches.5,7–9,13,14 Furthermore, exosomes offer several advantages over traditional cancer biomarkers, such as tumor-associated antigens and circulating tumor cells. Exosomes are more abundant in peripheral blood and can be detected at earlier disease states,15 making them highly promising for liquid biopsy-based strategies in cancer diagnosis, prognosis, and monitoring. However, the purification of exosomes from peripheral blood remains a challenging and time-consuming process due to their small size and complex physical properties. While differential ultracentrifugation has long been considered the gold standard,16 it is often accompanied by further purification steps based on density gradient or affinity capture techniques. Other purification methods, such as size exclusion chromatography,17,18 precipitation with volume-excluding polymers19 and filtration,20 have also been explored. Notably, size-based pre-selection combined with immuno-affinity capture have shown promising results for biomarker discovery using total RNA sequencing (RNA-seq).21,22 Multiple microfluidic devices have been developed to purify EVs from various types of biofluids. Whole blood is a complex matrix, so most devices require initial sample pre-treatment by centrifugation and then use plasma or serum as inputs in EV filtration,23–25 deterministic lateral displacement26 or immuno-affinity27–31 protocols. Other devices have implemented sized-based only32–34 or affinity-based only35 EV purification starting from whole blood. Only two devices currently combine both selection methods36,37 from whole undiluted blood, with starting volumes in the 150 to 300 μL range. EVs derived from blood contain a low amount of RNA, usually within the nanograms per milliliter (ng mL−1) range.38 Blood volumes between 1 and 3 mL may be required to obtain quality RNA-seq data for biomarker discovery and disease monitoring.18,39 Despite the progress made in exosome purification techniques, there is still a need for a simple, cost-effective workflow capable of efficiently isolating specific subsets of sEVs from milliliter volumes of whole blood. The complexity of current, labour-intensive EV separation and purification methods hinders the widespread adoption of exosome-based liquid biopsies in a clinical setting.
Building on our previous work in active pneumatic, centrifugal microfluidics40–42 and microfluidic-based immuno-magnetics,43,44 we present here the EV-Blade, implementing an automated method to purify exosomes from milliliter quantities of whole blood performed wholly on a low-cost microfluidic cartridge compatible with large-scale manufacturing. During continuous centrifugal rotation, the EV-Blade cartridge combines size-based EV pre-selection, using both whole blood centrifugation and plasma cross-flow filtration, together with EV immunomagnetic capture with functionalized superparamagnetic nanoparticles (MNPs) targeting well-established exosomal surface proteins CD9, CD63, and CD81.45,46 The automated device enabled repeated recovery of pure exosomes lysates, with yields equivalent to manual reference protocols for combined size and affinity-based selection. Following on-chip lysis of exosomes purified from whole blood samples, we recovered high quality RNA. Using RT-qPCR we detected relevant long coding and non-coding exosomal RNA transcripts representative of B-cell acute lymphoblastic leukemia (B-ALL),22 highlighting the potential for translation of our method in the field of exosome-based liquid biopsies for non-invasive cancer biomarkers monitoring.
Experimental
Active pneumatic control in centrifugal microfluidics
In standard centrifugal microfluidics, the centrifugal force generated by rotation drives on-chip fluid displacement. Valves are used for temporization of protocol steps and spatial control of both samples and reagents. Commonly, capillary valves,47 hydrophobic valves48 or passive siphons49 enable or restrict fluid transfers at specific rotation speeds. Other dynamic approaches to centrifugal valving include magnetic50 or mechanical actuation51 and dissolvable materials52 which significantly increase cartridge complexity. Moreover, these valves typically operate irreversibly and once open the entire fluid content dispenses from one reservoir to the next. Mixing actions can be achieved using complex microstructures,53,54 active agitation with magnetic particles or shaking cycles.55 Given the inherent limitations of passive and mechanically complex approaches to valving, mixing, and fluid handling, alternative solutions such as active pneumatic pumping have been proposed. Previous work by Clime et al.40 describe the implementation of the PowerBlade system, enabling active pneumatic pumping within a centrifugal microfluidics platform while in rotation. Briefly, a computer-controlled pressure manifold is integrated on the platform's rotor, capable of delivering positive and negative pressure to the device cartridge through individually addressable ports.On the PowerBlade system fluid displacement can now be controlled by balancing centrifugal forces and pneumatic pressure gradients. This capability enables movement of liquids against the centrifugal force (reverse pumping), from the device periphery back to the center of rotation, and by extension, back and forth movements through fluidic structures which are particularly useful for operations involving incubation or washing. Pneumatic actuated siphon valves can be reused repeatedly, enabling precise dispensing of fluid aliquots from the same reservoir. Injecting air into liquid filled device reservoirs through the pneumatic manifold generates bubbles that rise in the liquid against the centrifugal force field enabling fast and efficient mixing. Finally, routing flow from a single starting reservoir towards two or more different paths (flow switching) is easily achieved. Overall, active pneumatic pumping on the PowerBlade platform offers great flexibility for fluid handling inside centrifugal microfluidic devices.
Microfluidic cartridge fabrication
The main body of the EV-Blade fluidic cartridge was designed with Solidworks (Dassault Systèmes, FR) and fabricated with a CNC milling machine (Q350, Menig Automation, USA) from custom bio-compatible thermoplastic blanks in-house injected, cyclo olefin polymer (COP) Zeonor 1060R (Zeon Chemicals LP, USA) for low volume prototyping, followed by a small volume injection molding run (1000 units) of the microfluidic cartridges in Zeonor 1060R (Protolabs, USA).The magnetic capture sub-unit uses a previously developed magnetic gradient concentrator called the M-Chip.43 It consists of an array of micron-sized, high aspect ratio cylindrical pillars (20 μm diameter, 50 μm pitch) hot embossed in a polymer substrate. A master mold with multiple insert dies was defined on a silicon wafer using photolithography and deep reactive ion-etching (80 μm depth). The micropattern was first replicated by hot embossing in a 300 μm thick fluorinated ethylene polymer (FEP) film to create a re-usable working stamp for serial replication of the microstructured pillar array in 1 mm thick, 150 mm diameter Zeonor 1060R blanks. After embossing, individual arrays were die-cut and coated with 5.5 ± 0.5 μm of nickel followed by 200 nm of gold via electroless deposition to protect and passivate the nickel surface. The fluidic layer was fabricated by hot embossing in-house, extruded, thermoplastic elastomer (TPE, Mediprene OF 400M) sheets using an epoxy mold replicated from a PDMS mold, itself replicated from a 50 μm thick negative photoresist (SU8 1075, Gersteltec, CH) patterned on a silicon wafer. After punching inlet and outlet through hole vias in the TPE fluidic layer, it was bonded to the microstructured pillar array by contact at room temperature (RT). Finally, the through hole vias were aligned with the top of the EV-Blade cartridge and bonded by contact. The complete assembly was then annealed at 60 °C for 12 h in an oven to increase adhesion and release residual stress in the injection molded parts.
The cross-flow filtration sub-unit uses a PVDF 0.22 μm filtration membrane (Sterlitech, USA) bonded to a seat structure inside the filtration chamber and fixed using a laser-cut ring of pressure sensitive adhesive (PSA, Adhesive Research, USA).
The EV-Blade cartridge microfluidic channels and reservoirs were sealed by laminating a PSA (Adhesive Research, USA) together with a thin COP (Zeon Chemicals LP, USA) backing layer.
Cell culture
REH (ETV6-RUNX1+ human B cell precursor leukemia; ATCC CRL-1567) and 697 (TCF3-PBX1+ human B cell precursor leukemia; DSMZ ACC-42) cells were first cultured in RPMI 1640 medium (Wisent, CA) supplemented with 10% heat-inactivated fetal bovine serum (FBS) (Wisent, CA) and 1% penicillin/streptomycin (Wisent, CA). They were then transferred gradually to 100% X-VIVO 15 serum-free hematopoietic cell medium (Lonza, CH) over the course of 4 passages. All cell lines were cultured in a 37 °C incubator with 5% CO2. Conditioned cell culture medium (CCM) was collected 48 h after passage and apoptosis was kept under 5%.Blood collection
Blood samples were collected in K2 EDTA coated tubes (BD, USA) by venipuncture from healthy donors. The institutional review board of the CHU Sainte-Justine Research Center approved the research protocol and written informed consent was obtained from all participants. The study was performed in accordance with the Declaration of Helsinki.Manual EVs isolation from CCM for blood spiking
For all the cellular models, 45 mL of CCM was collected at 70% cell confluency and centrifuged at 300×g for 10 min at RT to remove suspended cells followed by a second round of centrifugation at 2500×g for 10 min at RT to remove large cell debris. The supernatant was collected and concentrated down to 900 μL by ultrafiltration using Amicon Ultra-15 centrifugal filter units with 100 kDa molecular weight cut-off (Millipore Sigma, USA). Concentrated CCM was used as input for EVs isolation by size-exclusion chromatography SEC using qEV original 70 nm columns (Izon Science, NZ). For each sample, 4 × 500 μL fractions were collected, filtered through a 0.22 μm centrifugal filter unit at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000g for 4 min (Ultrafree-MC, Millipore Sigma, USA) to remove large EVs (>200 nm in diameter) and concentrated down to 0.5 mL using Amicon Ultra-0.5 centrifugal filter units with 30 kDa molecular weight cut-off spun at 14
000g for 4 min (Ultrafree-MC, Millipore Sigma, USA) to remove large EVs (>200 nm in diameter) and concentrated down to 0.5 mL using Amicon Ultra-0.5 centrifugal filter units with 30 kDa molecular weight cut-off spun at 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g. The volume of concentrated sEVs added to the blood samples was adjusted to align with Cq values of B-ALL biomarker candidates we previously measured in patient samples via RT-qPCR.22
000×g. The volume of concentrated sEVs added to the blood samples was adjusted to align with Cq values of B-ALL biomarker candidates we previously measured in patient samples via RT-qPCR.22
Manual exosomes and sEVs purification from blood
Freshly collected blood was pooled and divided into 1 mL aliquots. Aliquots used for modeling exosome based liquid biopsies were spiked with concentrated sEVs to align with Cq values of B-ALL biomarker candidates we previously measured in patient samples via RT-qPCR.22For all manual methods, blood samples underwent centrifugation at 1100×g for 10 min for plasma separation. For size-exclusion chromatography (SEC), the collected plasma was filtered using a syringe and a 0.8 μm cellulose acetate filter membrane (GVS, Italy) following recommendations of Gaspar et al.18 Then, 500 μL of filtered plasma was used as input for EVs isolation via SEC using qEV original 70 nm columns (Izon Sience, New-Zealand). Four 500 μL fractions per sample were collected and centrifuged at 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g to remove larger microvesicles.
000×g to remove larger microvesicles.
For manual immunomagnetic exosomes purification, the plasma was filtered through 0.22 μm centrifugal filter units at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g for 4 min (Ultrafree-MC, Millipore Sigma, USA) to remove large EVs (>200 nm in diameter). Each filtered plasma sample was then processed with the EasySep Human Pan-Extracellular Vesicle Positive Selection Kit (Stemcell, CA), purifying EVs bearing CD9, CD63 and CD81 surface proteins following the manufacturer's protocol. After the final wash step, bead bound EVs were either lysed in RL buffer (Norgen, CA) for RNA extraction or in RIPA buffer (Sigma) for protein analysis.
000×g for 4 min (Ultrafree-MC, Millipore Sigma, USA) to remove large EVs (>200 nm in diameter). Each filtered plasma sample was then processed with the EasySep Human Pan-Extracellular Vesicle Positive Selection Kit (Stemcell, CA), purifying EVs bearing CD9, CD63 and CD81 surface proteins following the manufacturer's protocol. After the final wash step, bead bound EVs were either lysed in RL buffer (Norgen, CA) for RNA extraction or in RIPA buffer (Sigma) for protein analysis.
Tunable resistive pulse sensing (TRPS)
TRPS measurements were conducted using a qNano instrument (Izon Science, NZ). Various polyurethane nanopores appropriately stretched (NP100, NP150, NP200, NP400, NP800, Izon Science, NZ) and the matching diluted calibration polystyrene beads (CPC100, CPC200, CPC400, CPC800, Izon Science, NZ) were used to obtain the particle size distribution (PSD) of the EV samples. All samples were diluted in phosphate buffered saline (PBS) with 25 mM trehalose and 0.05% Tween20 to minimize EV aggregation. Each measurement was performed at two different pressures (5 and 10 mbar). Data were processed and analyzed using the Izon Control Suite software v3.3.2.2001.Optimization of magnetic capture element
The capture efficiency of magnetic nanoparticles (MNPs) using the M-Chip device was assessed by optical absorbance measurements as described previously.43,56 Briefly, stand-alone M-Chip devices with various gap sizes between the nickel/gold coated pillars were assembled onto in-house, injection molded luer-lock cover plates and connected to a syringe pump. See Poncelet et al.44 for more details on the test device assembly, noting that a single, larger N52 magnet was used for the current study to model the configuration of the EV-Blade cartridge. A custom setup, consisting of a pair of tube holders each containing a photodiode and an 850 nm wavelength led coupled to a signal amplifier and associated computer software was use to record the measurements in real-time. Optical absorbance in the outlet tubing was compared to a reference tube filled with the MNPs dilution buffer. For each measurement, 100 μL unlabelled EasySep MNPs (Stemcell, CA) were diluted in 1 mL PBS. The M-Chip array was first filled with the dilution buffer to record a baseline then switched to diluted MNPs. Once maximal absorbance at the outlet was reached MNPs capture was initiated in the M-Chip device by placing it either between a pair of permanent NdFeB magnets (K&J Magnetics, USA) or along the side of a single magnet. After reaching an absorbance plateau corresponding to the maximal capture capacity for a set flowrate and magnet configuration, dilution buffer was flowed through the M-Chip to simulate washing steps and the recording was ended.MNPs preparation for on-chip processing
The manufacturer's original protocol starts with incubation of the pre-processed plasma sample with 50 μL mL−1 of antibodies cocktail for 10 min at RT, followed by incubation with 100 μL mL−1 of MNPs for 10 min at RT before the washing and EVs lysis steps. Due to limited availability of pressure ports on the PowerBlade instrument, constraining the total number of steps possible to automate on a single device, we introduced the following variation. The MNPs and antibodies cocktail were incubated together off-chip for 10 min at RT before loading in the EV-Blade device.Assessment of CD9+ CD63+ CD81+EV recovery and purity with the EV-Blade
Concentration of known EV protein markers CD9, CD63, CD81 and syntenin-1 (Synt-1) as well as cytochrome-C (Cyt-C) as a marker of cellular organelles contamination and potential contamination by various apolipoproteins (Apo-A1, Apo-A2, Apo-B, Apo-C2, Apo-E) were measured using bead-based immunoassay panels (exosome characterization 6-plex human and human apolipoprotein ProcartaPlex panels, Invitrogen, USA) on a CS 1000 Autoplex Analyzer (PerkinElmer, USA) after lysis of purified EVs in radioimmunoprecipitation assay (RIPA) buffer. Starting from aliquots of the same pooled blood sample, we compared protein content in sEVs purified by SEC, manually purified with the EasySep Human Pan-Extracellular Vesicle Positive Selection Kit (Stemcell, CA) and EVs purified from whole blood with the EV-Blade cartridge on the PowerBlade. All samples were lysed using 100 μL of RIPA buffer and topped-up to 200 μL with PBS.RNA extraction and quantification
For the samples purified with the EV-Blade microfluidic cartridges, bead-bound exosomes trapped in the magnetic pillar array were lysed on-chip with RL buffer (Norgen, CA). The lysate was collected from the cartridge at the outlet of the processed sample chamber for off-chip RNA purification. Total RNA was extracted using the Total RNA Purification Micro Kit (Norgen, CA) following the manufacturer's instructions with on-column DNAse digestion (Norgen, CA) following the RNA binding step. For conventionally purified samples, the magnetic beads were removed from the lysate by centrifugation before further processing through the purification columns. For all samples, RNA yield and quality was assessed by electrophoresis on a 2100 bioanalyzer with the RNA 6000 nano or pico kits (Agilent Technologies, USA).Results and discussion
EV-Blade microfluidic cartridge
The EV-Blade cartridge (Fig. 1) consists of multiple fluidic sub-units designed to automate sequential steps of the full EV purification protocol. These sub-units are connected by linking microfluidic channels. Handling and flow of sample and reagent fluids is controlled by active pneumatic pumping using eight pressure ports on the microfluidic cartridge. These cartridge ports interface directly with eight individually controlled pressure manifold ports on the PowerBlade instrument, as described previously.40 The blood fractionation chamber is designed to accommodate up to 1 mL of whole blood. After white and red blood cells are sedimented by on-chip centrifugation, the plasma is transferred to the subsequent cross-flow filtration step through an outlet channel to collect just the upper plasma fraction. The filtration unit uses a two-chamber design. One chamber houses a 13 mm diameter membrane filter with 0.22 μm pore size, supported on a seat structure consisting of an array of 1 mm diameter posts and fixed in place using a laser-cut ring of pressure sensitive adhesive. The second chamber serves as a plasma overflow reservoir during the transfer process. The wash (700 μL) and lysis (100 μL) buffer reservoirs store reagents on-board until needed at specific time points in the protocol. During the protocol, waste products are directed to a collection reservoir (up to 3 mL). At the end of the protocol, lysate from purified EVs (100 μL) is sent to a dedicated processed sample reservoir. All the reservoirs have a dedicated port to allow manual reagent loading by pipetting before sealing with 4 mm diameter discs of PCR compatible adhesive foil prior to loading the microfluidic EV-Blade cartridge on the PowerBlade instrument. Finally, the magnetic capture unit has three design elements: (i) a source chamber, (ii) a magnetic capture insert and (iii) a sink chamber. During sample processing, each reagent or mixture is first transferred from its holding reservoir of origin into the source chamber. Next the sample is transferred through the magnetic capture insert channels and magnetic pillar array and into the sink chamber. After filling of the sink chamber, the liquid can be transferred either to the waste collection chamber or to the EV lysate collection chamber. By appropriate actuation of the pressure ports, liquids can be transferred from the sink chamber back through the capture insert to the source chamber, allowing multiple passes through the magnetic capture array. This unique back and forth fluid handling capability of the EV-Blade platform not only maximizes magnetic capture efficiency, but maximizes washing and lysis efficacy as well. By harnessing the dual fluidic handling modes of centrifugal microfluidics and active pneumatic pumping together with the additional filtration and immuno-magnetic capture functional units, the EV-Blade enables the implementation of complex and clinically relevant EV preparation protocols on low-cost, industry scalable microfluidic devices.Plasma filtering sub-unit
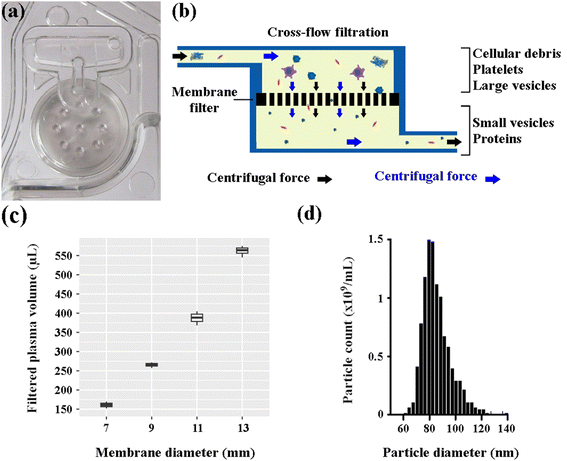
Fractionation of whole blood by centrifugation removes most of the cells, platelets and larger cell-debris from the sample. The filtering sub-unit shown in Fig. 2a is designed to remove the remaining platelets, cell debris and larger EVs from the plasma sample via cross-flow filtration (Fig. 2b) prior to immuno-capture. A membrane cut-off size of 0.22 μm typically permits most sEVs to pass through. We assessed plasma recovery post-filtration for four different membrane diameters, ranging from 7 mm to 13 mm (Fig. 2c), determining that 13 mm membranes are suitable for processing up to 1 mL of whole blood on the EV-Blade cartridge. Additionally, we observed that filtration capacity is directly proportional to the membrane area, providing design criteria for cross-flow filtration of plasma on centrifugal microfluidics platforms. Fig. 2d shows a total particle count in the solution after filtration on the cartridge. The resulting PSD is consistent with typical profiles observed for sEV preparations.17,18,57 At this stage, the sample is also expected to contain lipoproteins of comparable size.Magnetic capture sub-unit
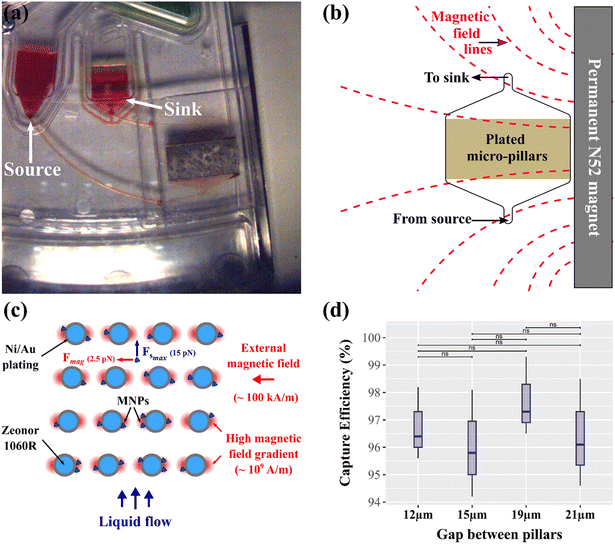
Fig. 3a shows a close-up photograph of the magnetic capture sub-unit with the source and sink chamber that regulate flow through the M-Chip. We used 400 nm diameter MNPs for exosome immunocapture within the M-Chip, a Ni/Au-coated micropillar magnetic array. Compared to the same mass of 2.8 μm Dynabeads, these MNPs provide approximately nine times more capture surface area and 350 times more particles in suspension, significantly accelerating binding kinetics due to increased target–particle interactions. While the reduced magnetic content of the MNPs poses a challenge for separation, the EV-Blade design overcomes this by positioning a magnetic gradient concentrator (the M-Chip) adjacent to a compact N52 permanent magnet (Fig. 3b). The pillar array generates local magnetic field gradients several orders of magnitude higher than the external magnet alone (Fig. 3c), enabling rapid MNP capture using magnets that can reasonably be mounted on a centrifuge rotor. All calculations regarding the MNP capture dynamics are detailed in the SI. The EV-Blade also incorporates a novel centrifugal-pneumatic filling strategy (Fig. 3c) that drives liquid into the M-Chip against the centrifugal force, efficiently displacing air and preventing bubble entrapment, a common limitation in previous syringe-pump implementations,43,44 thus enabling stable operation above 200 μL min−1 without flow disruption. The absence of trapped bubbles ensures consistent and uniform liquid exchange, enabling faster washing steps and supporting a higher overall sample processing capacity within shorter timeframes. Further exploration of this deterministic filling phenomenon is presented in Fig. S1. Importantly, capture efficiency remains consistent despite minor manufacturing variations in pillar geometry (Fig. 3d and SI), underlining the robustness of the system for high-throughput, scalable immunocapture from complex biofluids at high flow rates.Automated exosomes purification protocol
The sequential exosomes purification protocol including pressure values, rotation speeds, durations and open ports on the pressure manifold is summarized in Table 1. Fig. 4 shows selected stroboscopic images taken during an experiment to illustrate the key steps of the automated protocol. A video of the entire experiment is available in the SI (Movie Fig. S4). The protocol starts with fractionation of the whole blood sample using just the centrifugal force function (step 1, Fig. 4a). The plasma fraction is then transferred to the filtration chamber by applying positive pressure through port #1. Plasma flow through the filtration membrane is driven both by the centrifugal force and a gentle negative pressure applied to ports #2 to #8 (step 2, Fig. 4b). The filtered plasma then flows directly into the incubation chamber containing a pre-loaded solution of MNPs functionalized with a cocktail of EV specific antibodies. By applying negative pressure to port #2, air is entrained through the plasma in a pressure differential with port #3 (at atmosphere), inducing bubble formation at the base of the incubation chamber. Buoyed to the chamber top by the centrifugal force, the air bubbles create a convection current within the chamber to efficiently mix the functionalized MNPs with the filtered plasma (step 3, Fig. 4c). This bubble mixing step is repeated periodically during the incubation step (10 min duration) to maintain MNP dispersion in the solution. Separately, at 5 min before the end of the incubation step, a small aliquot of pre-loaded wash buffer is transferred into the source chamber by applying positive pressure to port #5. Complete, bubble-free filling of the magnetic capture element is controlled by balancing the centrifugal force with active pneumatic pumping; this controlled wetting and filling primes the M-Chip micro-pillar array insert without trapping air, which would otherwise disrupt the flow through pillars and impact capture efficiency (step 4, Fig. 4d). The filtered plasma/MNPs mixture is transferred to the source chamber by applying positive pressure to port #2. As the suspended MNPs transit the primed M-Chip, the entire population is trapped on the Ni/Au coated pillars, leaving the plasma in the sink chamber depleted of target EVs. The depleted plasma is then itself discarded into the waste collection chamber by applying negative pressure to port #8 (step 5, Fig. 4e). After capture, multiple, sequential aliquots of wash buffer are transferred to the source chamber by applying positive pressure to port #5 and then subsequently transferred through the M-Chip to remove most non-vesicular contaminants (step 6, Fig. 4f). By oscillating ports #6 to #8 between positive and negative pressure, buffer is shuttled back and forth through the M-Chip to increase overall washing efficiency. Each wash buffer aliquot entering the sink chamber is then also discarded into the waste chamber. Next, the lysis buffer is transferred to the source chamber by applying positive pressure to port #4. Over the course of 5 min, the lysis buffer is cycled back and forth through the M-Chip to lyse the captured EVs (step 7, Fig. 4g) without needing to release the MNP from the microstructure pillars. Finally, the lysate collected in the sink chamber is transferred to the lysate collection chamber by applying negative pressure to port #6 (step 8, Fig. 4h). The EVs lysate can then be manually extracted from the device for downstream analyses. The EV-Blade cartridge enables automated purification of EVs sub-populations and lysate recovery in 60 min.| Step | Operation | Active ports | Applied pressure (psi) | Rotation speed (rpm) | Duration |
|---|---|---|---|---|---|
| a Suspension containing 50 μL EasySep MNPs incubated with 25 μL pan-EVs positive selection cocktail (Stemcell). b PBS. c Buffer RL (Total RNA Purification Micro Kit, Norgen). | |||||
| Blood fractionation | |||||
| 1 | Centrifugation | — | — | 800 | 30 min |
| Plasma filtration | |||||
| 2a | Transfer plasma to filtration chamber | 1 | 4.5 | 400 | 30 s |
| 2b | 0.22 μm plasma filtration | 1, 2 | 1 | 500 | 60 s |
| Affinity-capture | |||||
| 3a | Re-suspend MNPsa by bubble mixing | 3 | 2 | 600 | 5 × 1 s |
| 3b | Plasma incubation with MNPs | — | — | 100 | 10 min |
| Bubble mixing during incubation | 3 | 2 | 600 | 10 × 2 s | |
| Mchip priming | |||||
| 4a | Send wash bufferb aliquot to sink | 5 | 1 | 500 | 1 s |
| 4b | Flow wash buffer in M-Chip | 6, 7, 8 | −0.5 | 200 | 60 s |
| 4c | Empty drain to waste | 8 | −2 | 500 | 3 s |
| Magnetic capture | |||||
| 5a | Send plasma + MNPs to sink | 4 | 1 | 500 | 5 s |
| 5b | Flow beads in M-Chip | 6, 7, 8 | −0.3 | 200 | 120 s |
| 5c | Empty drain to waste | 8 | −2 | 500 | 3 s |
| Wash (repeat 3 times) | |||||
| 6a | Send wash aliquot to sink | 5 | 1 | 600 | 2 s |
| 6b | Flow wash buffer in M-Chip (back and forth) | 6,7,8 | ±0.5 | 200 | 3 × 45 s |
| 6c | Empty drain to waste | 8 | −2 | 500 | 3 s |
| EVs lysis | |||||
| 7a | Send lysis bufferc to sink | 4 | 1.5 | 500 | 3 s |
| 7b | Flow lysis buffer in M-Chip (back and forth) | 6, 7, 8 | ±0.5 | 200 | 6 × 30 s |
| Lysate collection | |||||
| 8 | Send EVs lysate to processed sample chamber | 6 | −2 | 500 | 10 s |
 | ||
| Fig. 4 Selected stroboscopic images of the EV-Blade cartridge at different stages of the EVs purification protocol. Operational parameters for each step are detailed in Table 1. (a) Fractionation of the blood sample by centrifugation. (b) The plasma fraction is transferred to the cross-flow filtration chamber. The liquid flow through the membrane is driven by negative pressure and the centrifugal force, both generated by the PowerBlade instrument. (c) Air bubbles are injected via a channel at the bottom of the incubation chamber to induce chaotic mixing of the filtered plasma with the MNPs. (d) An aliquot of wash buffer is used to fully wet the surface of the micropillar array. (e) The plasma/MNPs mixture is flowed through the pillars for instantaneous flow-through magnetic capture. The MNP-free plasma is then transferred to the waste chamber (f) multiple aliquots of wash buffer are flowed back and forth through the M-Chip and discarded into the waste chamber. (g) The lysis buffer is transferred to the source chamber and flowed back and forth through the M-Chip for incubation with the EVs bound to the captured MNPs. (h) The EVs lysate is transferred to the processed sample chamber for manual collection and off-chip downstream analyses. | ||
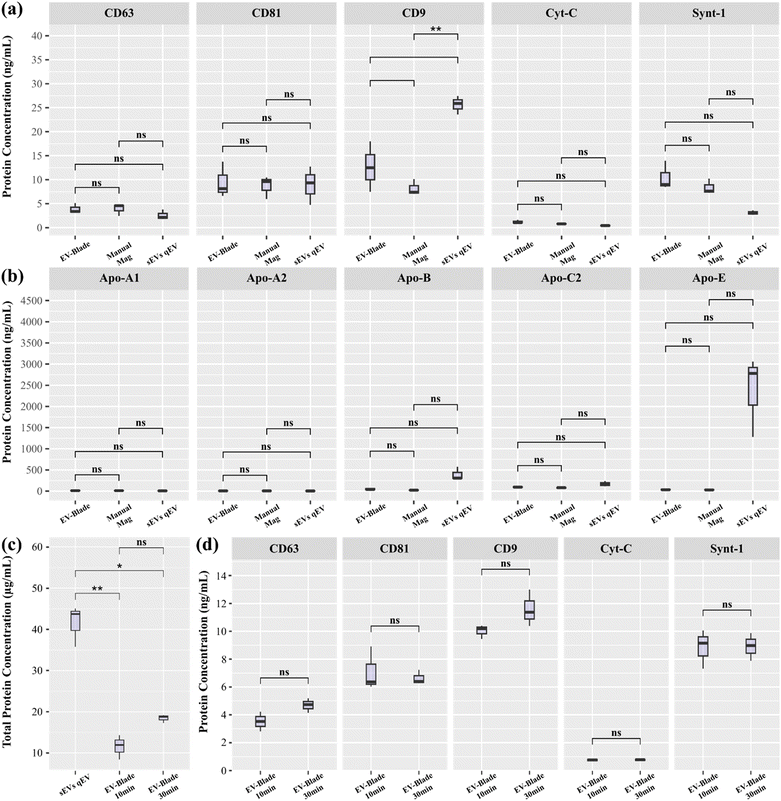
Exosome purification efficiency
We compared the exosome purification performance of the EV-Blade against SEC and a manual size- and affinity-based exosomes purification protocol that combines centrifugation-based blood fractionation, 0.22 μm filtration using a centrifugal filtering column and immunomagnetic capture in a tube. Freshly collected whole blood samples were pooled and divided into 1 mL aliquots. We processed 3 blood samples with each method (SEC, manual size- and affinity-based purification and EV-Blade), and measured the protein concentration of known exosomes-enriched transmembrane proteins (CD63, CD81, CD9 and Synt-1) in the recovered lysates. Both methods yielded almost identically pure exosomes populations, with negligible cellular contamination, as indicated by the absence of Cyt-C (Fig. 5a). Additionally, preparations from both size- and affinity-based methods (Manual Mag and EV-Blade) were free from 5 lipoproteins (Apo-A1, Apo-A2, Apo-B, Apo-C2 and Apo-E) whereas SEC preparations (sEVs qEV) contained traces of Apo-B and a significant amount of Apo-E, indicating contamination by high-density lipoprotein (HDL) particles in SEC purified EVs (Fig. 5b). The total protein yield was higher for SEC purified EVs compared to EV-Blade purified exosomes (Fig. 5c), which can be attributed to HDL particle contamination in SEC samples and the higher specificity of the EV-Blade capture protocol targeting a subset of blood derived sEVs enriched in CD63, CD81 and CD9.Increasing the immuno-capture incubation time in the EV-Blade method resulted in more total protein (Fig. 5c) but did not significantly alter exosomes-specific protein levels (Fig. 5d), suggesting 10 min incubation is sufficient. Extending incubation does not further increase exosome yield and may reduce purity through non specific binding. Reference values measured for exosomal transmembrane proteins and lipoproteins in filtered plasma samples are shown in Fig. S2. Combined with the PSD shown in Fig. 2c, these results confirm that the EV-Blade cartridge effectively purifies exosomes from whole blood. Processed samples are free from size-similar intra-cellular organelle contaminants and yield results comparable to existing manual protocols, with expected differences in purity when comparing size selection only (sEVs qEV) versus size- and affinity-based purification methods (Manual Mag and EV-Blade).
Analysis of exosomes derived circulating RNA
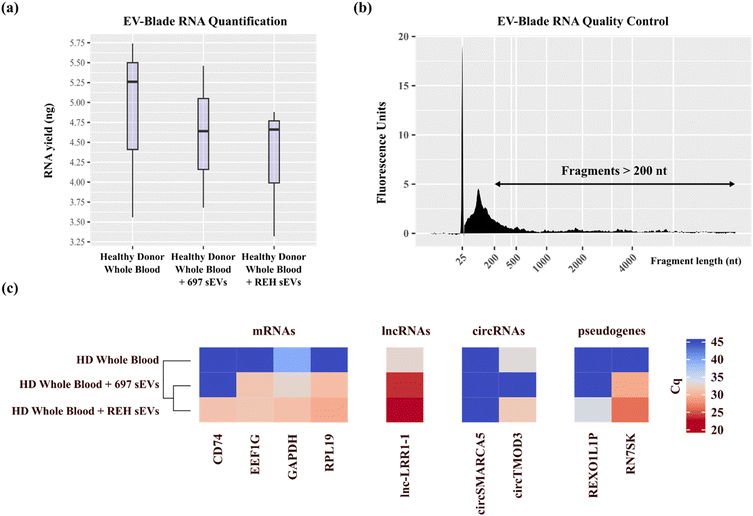
We evaluated the EV-Blade cartridge as a companion tool for exosomal RNA-based cancer liquid biopsy. In previous work, we analyzed long circulating RNAs found in extracellular vesicles purified from the peripheral blood of childhood B-ALL patients using total RNA sequencing.22 Given the very limited availability of plasma from childhood B-ALL patients at diagnosis (900 μL per patient), most extracted RNA was used for sequencing and RT-qPCR validation. To simulate similar conditions for the purpose of validating our technology, we started with a pooled sample of whole blood collected from healthy donors and purified exosomes from three 1 mL aliquots under the three following conditions: (i) healthy donor whole blood, (ii) healthy donor whole blood spiked with sEVs manually purified from REH cell CCM and (iii) healthy donor whole blood spiked with sEVs manually purified from 697 cell CCM. The concentration of spiked sEVs was adjusted to align with Cq values of selected potential biomarker candidates we previously measured in patient samples via RT-qPCR.22 These conditions represent healthy individuals, one subtype of low-risk B-ALL (ETV6::RUNX1 positive) patients and one subtype of high-risk B-ALL (TCF3::PBX1 positive) patients forming a model for liquid biopsy application in cancer patient risk stratification. Fig. 6a shows RNA yields from EV-Blade purified samples, ranging from 3.32 to 5.74 ng. RNA concentrations in spiked plasma samples were similar or slightly lower than those in whole blood samples. The collected lysates exhibited a broad distribution of RNA fragment sizes, with the majority measuring <200 nucleotides and larger fragments present at lower abundance, as shown in Fig. 6b. On average, 30% of the total fragments exceeded 200 nucleotides (DV200), with RNA integrity number (RIN) values ranging from 1.7 to 2.4. Previous work by us and others22,58–61 have shown the utility of long coding and non-coding RNA transcripts including mRNAs, lncRNAs, circRNAs and pseudogenes biotypes for risk stratification of B-ALL patients. We previously identified potential RNA-based biomarkers in patients' blood-derived exosomes and cellular models that correlate with B-ALL molecular subtypes associated with high- and low-risk conditions.22 We selected a panel of genes in these 4 biotypes and assessed their expression by RT-qPCR (Fig. 6c), revealing significant expression differences for 8 out of 9 genes across the three model conditions, indicating that our exosome-derived RNA profiles can distinguish between different B-ALL subtypes. The EV-Blade exosome purification results show that starting from a clinically relevant volume of whole peripheral blood, we were able to recover sufficient RNA to successfully screen a panel of potential B-ALL biomarkers correlating with high- and low-risk conditions in model samples. Further validation with primary patient samples will be critical to confirm these findings and assess the device's sensitivity, specificity, and robustness for routine clinical diagnostics.Conclusion
This study presents the EV-Blade, the first integrated centrifugal–pneumatic microfluidic cartridge to automate size- and affinity-based exosome isolation using cyclic flow-through magnetic capture. This innovative integration enables a complex purification protocol to be fully automated at high flow rates, crucial for processing larger sample volumes (in the milliliter range) compared to previously reported methods that were limited to smaller blood volumes. Automating this process is crucial for clinical adoption as it simplifies and reduces the costs associated with the otherwise complex purification workflow. Handling larger volumes is essential for detecting lowly expressed biomarkers, thereby enhancing disease detection capabilities during the early stages from minimally invasive biofluid collections, such as peripheral blood draws, rather than invasive tissue biopsies.The EV-Blade efficiently isolates specific subsets of sEVs, particularly exosomes, from whole blood in 60 min. Its robust design supports scalable fabrication and consistent capture performance despite manufacturing variations, making it a cost-effective and clinically viable solution for exosome purification. The device removes blood cells, larger EVs, and cellular debris while retaining sEVs within the 60 to 120 nm size range in high quantities. On-chip immunomagnetic capture uses functionalized magnetic nanoparticles assembled in porous magnetic clouds for cyclic flow-through incubation. This process further purifies exosomes by targeting well-established exosomal surface proteins, effectively eliminating size-similar intracellular organelle contaminants and lipoproteins from recovered samples. We showcase the EV-Blade's capabilities in a clinically relevant workflow by processing model samples for liquid biopsies aimed at risk stratification in B-ALL patients. This demonstration highlights the EV-Blade's ability to efficiently recover both coding and non-coding RNA transcripts from blood exosomes, facilitating the screening of potential B-ALL biomarkers which can be used for the differentiation of B-ALL subtypes. These results underscore the practical applications of the EV-Blade in cancer research and diagnostics, offering a powerful tool for non-invasive biomarker discovery and disease monitoring.
In summary, the EV-Blade offers an automated and scalable method to isolate exosomes from whole peripheral blood, efficiently retaining sEVs in plasma through fractionation and cross-flow filtration and providing robust immunomagnetic capture of sEV subsets (e.g., exosomes) for practical omics-based applications in cancer research. Given the simplicity of the device and its ability to fully automate the complex immunomagnetic separation protocol, which can be easily customized for targeting different circulating targets by simply changing the magnetic bead functionalization, this method holds great promise for advancing liquid biopsy-based approaches. It can be easily translated towards multiple applications in cancer research and ultimately in clinical practice, improving the diagnosis and monitoring of various diseases.
Conflicts of interest
The authors declare no conflicts of interest.Data availability
The data supporting this article have been included in the main body of the article and as part of the supplementary information (SI).Supplementary information is available. See DOI: https://doi.org/10.1039/d5lc00977d.
Acknowledgements
This work was funded by the NRC Medical Devices Research Center, the NRC Pandemic Response Challenge Program and the Francois-Karl Viau (FKV) chair in pediatric oncogenomics. LP was supported by a doctoral fellowship from the Cole Foundation and a scholarship from the National Sciences and Engineering Research Council. DS holds the FKV chair in pediatric oncogenomics.References
- V. C. Kok and C.-C. Yu, Int. J. Nanomed., 2020, 15, 8019–8036 Search PubMed.
- J. Kowal, M. Tkach and C. Théry, Curr. Opin. Cell Biol., 2014, 29, 116–125 CrossRef CAS PubMed.
- K. W. Witwer and C. Théry, J. Extracell. Vesicles, 2019, 8, 1648167 Search PubMed.
- A.-K. Ludwig and B. Giebel, Int. J. Biochem. Cell Biol., 2012, 44, 11–15 Search PubMed.
- X. Zang, J. Gu, J. Zhang, H. Shi, S. Hou, X. Xu, Y. Chen, Y. Zhang, F. Mao, H. Qian, T. Zhu, W. Xu and X. Zhang, Cell Death Dis., 2020, 11, 215 Search PubMed.
- Q. Xiao, C. Lin, M. Peng, J. Ren, Y. Jing, L. Lei, Y. Tao, J. Huang, J. Yang, M. Sun, J. Wu, Z. Yang, Z. Yang and L. Zhang, Front. Oncol., 2022, 12, 1033143 Search PubMed.
- J. Nehrbas, J. T. Butler, D. W. Chen and P. Kurre, Front. Oncol., 2020, 10, 90 Search PubMed.
- M. Layoun, J. Huan, A. M. Skinner and P. Kurre, Blood, 2011, 118, 3751 Search PubMed.
- M. Ge, Z. Qiao, Y. Kong, H. Lu and H. Liu, Cancer Sci., 2020, 111, 1279–1290 Search PubMed.
- W. Yu, J. Hurley, D. Roberts, S. K. Chakrabortty, D. Enderle, M. Noerholm, X. O. Breakefield and J. K. Skog, Ann. Oncol., 2021, 32, 466–477 Search PubMed.
- Q. Li, Y. Shao, X. Zhang, T. Zheng, M. Miao, L. Qin, B. Wang, G. Ye, B. Xiao and J. Guo, Tumor Biol., 2015, 36, 2007–2012 Search PubMed.
- J. Wang, Y. Zhou, J. Lu, Y. Sun, H. Xiao, M. Liu and L. Tian, Med. Oncol., 2014, 31, 1–8 Search PubMed.
- D. Wang, X. Ming, J. Xu and Y. Xiao, Hematol. Oncol., 2021, 39, 390–400 Search PubMed.
- A. Samii and F. Razmkhah, Stem Cell Rev. Rep., 2020, 16, 1081–1091 Search PubMed.
- M. He and Y. Zeng, J. Lab. Autom., 2016, 21, 599–608 Search PubMed.
- C. Théry, S. Amigorena, G. Raposo and A. Clayton, Curr. Protoc. Cell Biol., 2006, 30, 3.22.21–23.22.29 Search PubMed.
- T. Baranyai, K. Herczeg, Z. Onódi, I. Voszka, K. Módos, N. Marton, G. Nagy, I. Mäger, M. J. Wood, S. El Andaloussi, Z. Pálinkás, V. Kumar, P. Nagy, Á. Kittel, E. I. Buzás, P. Ferdinandy and Z. Giricz, PLoS One, 2015, 10, e0145686 CrossRef PubMed.
- L. S. Gaspar, M. M. Santana, C. Henriques, M. M. Pinto, T. M. Ribeiro-Rodrigues, H. Girão, R. J. Nobre and L. Pereira de Almeida, Mol. Ther.--Methods Clin. Dev., 2020, 18, 723–737 CrossRef CAS PubMed.
- R. J. Lobb, M. Becker, S. W. Wen, C. S. F. Wong, A. P. Wiegmans, A. Leimgruber and A. Möller, J. Extracell. Vesicles, 2015, 4, 27031 Search PubMed.
- R. Grant, E. Ansa-Addo, D. Stratton, S. Antwi-Baffour, S. Jorfi, S. Kholia, L. Krige, S. Lange and J. Inal, J. Immunol. Methods, 2011, 371, 143–151 Search PubMed.
- N. McNamee, R. Daly, J. Crown and L. O'Driscoll, Transl. Oncol., 2022, 15, 101274 Search PubMed.
- L. Poncelet, C. Richer, A. Gutierrez-Camino, T. Veres and D. Sinnett, Int. J. Mol. Sci., 2025, 26, 3956 Search PubMed.
- Z. Han, C. Peng, J. Yi, D. Zhang, X. Xiang, X. Peng, B. Su, B. Liu, Y. Shen and L. Qiao, Sens. Actuators, B, 2021, 333, 129563 Search PubMed.
- F. Liu, O. Vermesh, V. Mani, T. J. Ge, S. J. Madsen, A. Sabour, E.-C. Hsu, G. Gowrishankar, M. Kanada, J. V. Jokerst, R. G. Sierra, E. Chang, K. Lau, K. Sridhar, A. Bermudez, S. J. Pitteri, T. Stoyanova, R. Sinclair, V. S. Nair, S. S. Gambhir and U. Demirci, ACS Nano, 2017, 11, 10712–10723 Search PubMed.
- Z. Wang, H. J. Wu, D. Fine, J. Schmulen, Y. Hu, B. Godin, J. X. Zhang and X. Liu, Lab Chip, 2013, 13, 2879–2882 Search PubMed.
- J. T. Smith, B. H. Wunsch, N. Dogra, M. E. Ahsen, K. Lee, K. K. Yadav, R. Weil, M. A. Pereira, J. V. Patel, E. A. Duch, J. M. Papalia, M. F. Lofaro, M. Gupta, A. K. Tewari, C. Cordon-Cardo, G. Stolovitzky and S. M. Gifford, Lab Chip, 2018, 18, 3913–3925 Search PubMed.
- H. Fang, M. Liu and W. Jiang, Appl. Biochem. Biotechnol., 2023, 195, 3109–3121 Search PubMed.
- H. Shao, J. Chung, K. Lee, L. Balaj, C. Min, B. S. Carter, F. H. Hochberg, X. O. Breakefield, H. Lee and R. Weissleder, Nat. Commun., 2015, 6, 6999 CrossRef CAS PubMed.
- C. Chen, J. Skog, C. H. Hsu, R. T. Lessard, L. Balaj, T. Wurdinger, B. S. Carter, X. O. Breakefield, M. Toner and D. Irimia, Lab Chip, 2010, 10, 505–511 Search PubMed.
- Z. Zhao, Y. Yang, Y. Zeng and M. He, Lab Chip, 2016, 16, 489–496 RSC.
- M. He, J. Crow, M. Roth, Y. Zeng and A. K. Godwin, Lab Chip, 2014, 14, 3773 Search PubMed.
- R. T. Davies, J. Kim, S. C. Jang, E.-J. Choi, Y. S. Gho and J. Park, Lab Chip, 2012, 12, 5202–5210 RSC.
- V. Sunkara, C. J. Kim, J. Park, H. K. Woo, D. Kim, H. K. Ha, M. H. Kim, Y. Son, J. R. Kim and Y. K. Cho, Theranostics, 2019, 9, 1851–1863 CrossRef CAS PubMed.
- M. Wu, Y. Ouyang, Z. Wang, R. Zhang, P.-H. Huang, C. Chen, H. Li, P. Li, D. Quinn, M. Dao, S. Suresh, Y. Sadovsky and T. J. Huang, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 10584–10589 CrossRef CAS PubMed.
- M. Sancho-Albero, V. Sebastián, J. Sesé, R. Pazo-Cid, G. Mendoza, M. Arruebo, P. Martín-Duque and J. Santamaría, J. Nanobiotechnol., 2020, 18, 150 CrossRef CAS PubMed.
- Y.-S. Chen, Y.-D. Ma, C. Chen, S.-C. Shiesh and G.-B. Lee, Lab Chip, 2019, 19, 3305–3315 RSC.
- J. Rho, J. Chung, H. Im, M. Liong, H. Shao, C. M. Castro, R. Weissleder and H. Lee, ACS Nano, 2013, 7, 11227–11233 CrossRef CAS PubMed.
- S. R. Kumar, E. T. Kimchi, Y. Manjunath, S. Gajagowni, A. J. Stuckel and J. T. Kaifi, Sci. Rep., 2020, 10, 2800 CrossRef CAS PubMed.
- R. S. Rodosthenous, E. Hutchins, R. Reiman, A. S. Yeri, S. Srinivasan, T. G. Whitsett, I. Ghiran, M. G. Silverman, L. C. Laurent, K. Van Keuren-Jensen and S. Das, iScience, 2020, 23, 101182 CrossRef CAS PubMed.
- L. Clime, D. Brassard, M. Geissler and T. Veres, Lab Chip, 2015, 15, 2400–2411 Search PubMed.
- D. Brassard, L. Clime, M. Mounier and T. Veres, Proceedings of 20th International Conference on Miniaturized Systems for Chemistry and Life Sciences (MicroTAS 2016), 2016, 2016, pp. 31–32 Search PubMed.
- D. Brassard, M. Geissler, M. Descarreaux, D. Tremblay, J. Daoud, L. Clime, M. Mounier, D. Charlebois and T. Veres, Lab Chip, 2019, 19, 1941–1952 Search PubMed.
- L. Malic, X. Zhang, D. Brassard, L. Clime, J. Daoud, C. Luebbert, V. Barrere, A. Boutin, S. Bidawid, J. Farber, N. Corneau and T. Veres, Lab Chip, 2015, 15, 3994–4007 Search PubMed.
- L. Poncelet, L. Malic, L. Clime, M. Geissler, K. J. Morton, C. Nassif, D. Da Fonte, G. Veilleux and T. Veres, Analyst, 2021, 146, 7491–7502 RSC.
- W. Stoorvogel, M. J. Kleijmeer, H. J. Geuze and G. Raposo, Traffic, 2002, 3, 321–330 Search PubMed.
- C. Thery, L. Zitvogel and S. Amigorena, Nat. Rev. Immunol., 2002, 2, 569–579 CrossRef CAS PubMed.
- J. M. Chen, P.-C. Huang and M.-G. Lin, Microfluid. Nanofluid., 2007, 4, 427–437 Search PubMed.
- Y. Feng, Z. Zhou, X. Ye and J. Xiong, Sens. Actuators, A, 2003, 108, 138–143 CrossRef CAS.
- J. Siegrist, R. Gorkin, L. Clime, E. Roy, R. Peytavi, H. Kido, M. Bergeron, T. Veres and M. Madou, Microfluid. Nanofluid., 2009, 9, 55–63 CrossRef.
- Z. Cai, J. Xiang, B. Zhang and W. Wang, Sens. Actuators, B, 2015, 206, 22–29 Search PubMed.
- Z. Cai, J. Xiang and W. Wang, Sens. Actuators, B, 2015, 221, 257–264 Search PubMed.
- B. S. Lee, J.-N. Lee, J.-M. Park, J.-G. Lee, S. Kim, Y.-K. Cho and C. Ko, Lab Chip, 2009, 9, 1548–1555 RSC.
- A. Shamloo, P. Vatankhah and A. Akbari, Chem. Eng. Process.: Process Intesif., 2017, 116, 9–16 CrossRef CAS.
- C. W. Hsu, P. T. Shih and J. M. Chen, Micromachines, 2020, 11, 1110 CrossRef PubMed.
- M. Grumann, A. Geipel, L. Riegger, R. Zengerle and J. Ducrée, Lab Chip, 2005, 5, 560–565 Search PubMed.
- L. Clime, X. D. Hoa, N. Corneau, K. J. Morton, C. Luebbert, M. Mounier, D. Brassard, M. Geissler, S. Bidawid, J. Farber and T. Veres, Biomed. Microdevices, 2015, 17, 17 CrossRef PubMed.
- C. Thery, L. Zitvogel and S. Amigorena, Nat. Rev. Immunol., 2002, 2, 569–579 CrossRef CAS PubMed.
- M. Lajoie, S. Drouin, M. Caron, P. St-Onge, M. Ouimet, R. Gioia, M. H. Lafond, R. Vidal, C. Richer, K. Oualkacha, A. Droit and D. Sinnett, PLoS One, 2017, 12, e0174124 CrossRef PubMed.
- T. H. Tran, S. Langlois, C. Meloche, M. Caron, P. Saint-Onge, A. Rouette, A. R. Bataille, C. Jimenez-Cortes, T. Sontag, H. Bittencourt, C. Laverdière, V.-P. Lavallée, J.-M. Leclerc, P. D. Cole, L. M. Gennarini, J. M. Kahn, K. M. Kelly, B. Michon, R. Santiago, K. E. Stevenson, J. J. G. Welch, K. M. Schroeder, V. Koch, S. Cellot, L. B. Silverman and D. Sinnett, Blood Adv., 2022, 6, 1329–1341 CrossRef CAS PubMed.
- T. R. Fernando, N. I. Rodriguez-Malave, E. V. Waters, W. Yan, D. Casero, G. Basso, M. Pigazzi and D. S. Rao, Mol. Cancer Res., 2015, 13, 839–851 CrossRef CAS PubMed.
- A. Buratin, M. Paganin, E. Gaffo, A. Dal Molin, J. Roels, G. Germano, M. T. Siddi, V. Serafin, M. De Decker, S. Gachet, K. Durinck, F. Speleman, T. Taghon, G. Te Kronnie, P. Van Vlierberghe and S. Bortoluzzi, Blood Adv., 2020, 4, 5902–5914 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |