Open Access Article
Open Access ArticleDevelopment of a 3D-printed microfluidic chip for retinal organoid–endothelial co-culture
Rodi
Kado Abdalkader
 *a,
Shigeru
Kawakami
b,
Yuuki
Takashima
c and
Takuya
Fujita
d
*a,
Shigeru
Kawakami
b,
Yuuki
Takashima
c and
Takuya
Fujita
d
aRitsumeikan Global Innovation Research Organization (R-GIRO), Ritsumeikan University, Japan. E-mail: rodi@fc.ritsumei.ac.jp
bGraduate School of Biomedical Sciences, Nagasaki University, Japan
cFaculty of Pharmacy, Juntendo University, Japan
dCollege of Pharmaceutical Sciences, Ritsumeikan University, Japan
First published on 2nd February 2026
Abstract
Pathological angiogenesis, such as that observed in wet age-related macular degeneration (AMD), is difficult to reproduce in vitro using human-relevant models. Although organ-on-chip (OoC) systems incorporating retinal pigment epithelium (RPE) and endothelial barriers have been reported, models integrating human retinal organoids with vascular networks remain limited. Here, we present a fully 3D-printed microfluidic platform for co-culture of human induced pluripotent stem cell (hiPSC)-derived retinal organoids containing intrinsic RPE regions with endothelial cells. The device, fabricated from flexible thermoplastic polyurethane (TPU) on a transparent polyvinyl chloride (PVC) substrate, supports three-dimensional co-culture within a fibrin–Matrigel matrix. In this system, endothelial cells formed organized vascular networks that localized around RPE-associated regions of retinal organoids without direct tissue invasion. Organoid–endothelial co-culture resulted in increased VEGF secretion, while exogenous VEGF further enhanced endothelial localization near RPE regions without affecting organoid growth. Functional assays using fluorescent dextran and rhodamine-labeled liposomal nanoparticles demonstrated spatially restricted and time-dependent transport along vascularized regions adjacent to the organoid interface. This retinal organoid-on-chip provides a simple and robust in vitro platform for studying retinal–vascular interactions and vascular-mediated transport processes.
Introduction
Among all eye diseases, retinal diseases can cause severe and irreversible vision loss.1 Age-related macular degeneration (AMD), diabetic retinopathy, and retinitis pigmentosa are some of the very common illnesses with widespread loss of vision and even blindness.2,3 The most severe form of AMD, also known as wet AMD, includes abnormal growth of choroidal blood vessels into the subretinal space, causing blurring of vision. The intricate architecture of retinal tissue presents severe challenges to unravelling the pathophysiology of AMD disease aetiology, which in turn hinders effective therapy design.To advance our knowledge of AMD disease mechanisms, the development of advanced more human-like in vitro models of the retinal tissue is critical. Traditional in vivo models, while valuable, tend to fall short of simulating human retinal pathology due to serious anatomical and genetic differences.4 Standard in vitro models, on the other hand, can offer controlled environments for the investigation of disease onset, cell interactions, and drug efficacy. However, current in vitro models tend to be less complex and functional compared to the human retinal tissue and thus necessitate more innovative methods for better modeling of retinal disease.5,6 Another limitation lies in the availability of reliable biological sources of human retinal cells, as donor-derived tissue is scarce, expensive, and loses phenotypic stability after a few passages.7
To address these challenges, we focused on two complementary platforms. First, human induced pluripotent stem cell (hiPSC)-derived retinal organoids have shown great promise, as they can differentiate into multiple retinal cell types and self-organize into layered structures that closely resemble the native retina.8–11 However, these organoids typically lack endothelial cells, which are essential for vascularization, nutrient and waste exchange, and for modeling angiogenic pathologies relevant to diseases such as AMD.9 Second, organ-on-chip (OoC) technology represents a major advance in retinal modeling and has rapidly emerged as a powerful approach for constructing physiologically relevant in vitro systems.12,13 OoC platforms have been successfully applied to model a wide range of human tissues, including the liver,14 intestine,15 brain,16 and eye.17–19 By using microfluidic devices, these systems recreate tissue microarchitecture and physiological conditions by positioning living cells within three-dimensional (3D) microenvironments under precisely controlled flow.12,20,21
In the context of retinal disease research, OoC approaches offer major advantages: the ability to simulate blood-retina barriers, support ECM remodeling, and observe real-time cellular behavior under dynamic flow.19,22
These systems can closely mimic the in vivo environment and therefore provide a more physiologically relevant platform for studying retinal diseases and screening new therapeutic modalities. Early retinal OoC models were constructed in relatively simple configurations, highlighting the importance of the retinal pigment epithelium (RPE) and endothelial cells within microfluidic devices to model the outer blood-retinal barrier (oBRB).23–25 However, while valuable, these models typically employed cell lines and relied on layered structures that lacked key retinal cell types such as photoreceptors.26 More advanced approaches introduced the co-culture of hiPSC-derived retinal organoids placed on a porous membrane overlaid with an RPE monolayer within microfluidic device. This model enabled direct interaction between the organoids and RPE cells, offering a more robust model that incorporates both photoreceptors and RPE cells.22,27,28 Despite these advances, direct co-culture of hiPSC-derived retinal organoids with endothelial networks—necessary for reconstructing choroidal-like vasculature—remains limited.
We previously reported a simple and low-cost microfluidic chip fabricated by fused deposition modeling (FDM)-based 3D printing of thermoplastic polyurethane (TPU) on a transparent polyvinyl chloride (PVC) substrate, and demonstrated its biocompatibility with hiPSC-derived optic vesicle organoids and endothelial cells.29 Compared with conventional polydimethylsiloxane (PDMS)-based platforms, this fabrication approach offers advantages including mechanical flexibility, optical transparency, reduced cost, and rapid prototyping. Building on this strategy, we developed a retinal organoid-on-chip platform that enables direct co-culture of hiPSC-derived retinal organoids containing intrinsic RPE regions with endothelial cells within a fibrin–Matrigel matrix. The fully 3D-printed device eliminates the need for semipermeable membranes and supports organoid–endothelial interactions in a three-dimensional configuration. Using this platform, we show that endothelial cells self-organize into vascular networks that spatially associate with retinal organoids under basal conditions driven by endogenous VEGF secretion from the organoids, while enhanced endothelial localization near RPE regions is observed upon exogenous VEGF stimulation. In addition, we demonstrate vascular network-mediated transport of fluorescent liposomes and their localization at the organoid interface. Together, these results establish a simple and robust retinal organoid-on-chip platform that complements existing retinal-on-chip systems and extends their application to studies of retinal–vascular interactions, angiogenic processes, and vascular-mediated transport.
Experimental
1. Device fabrication and characterization
The microfluidic device was fabricated using our previously established FDM 3D-printing protocol (Anycubic Technology Co., Ltd., Shenzhen, China) with 1.75 mm translucent clear TPU filaments (SainSmart, Lenexa, KS, USA).29,30 Device designs were generated in Tinkercad (https://www.tinkercad.com) and consisted of six cell culture chambers, each connected to inlets and outlets. STL files were sliced and processed using Ultimaker Cura software with the following parameters: layer height, 0.01–0.02 mm; shell thickness, 2–10 layers. TPU filaments were directly deposited onto 1 mm-thick clear PVC sheets, after which the printed constructs were trimmed and stored at room temperature. The fidelity of the printing was assessed by measuring the diameter of printing organoid chamber using imageJ software according to the equation:| PF = A1/A0 |
2. Cell culturing and organoid generation
hiPSCs, 585A1,32 was procured from Riken Cell Bank and handled in accordance with the guidelines stipulated by the ethics committees of Ritsumeikan University (Approved number 2021-004-3). Culture dishes were pre-coated with imatrix-511 (Nippi Inc., Tokyo, Japan) overnight at 37 °C. 1 × 104 viable cells in mTeSR™ plus medium (STEMCELL Technologies Inc., Vancouver, Canada) supplemented with 10 μM Y27632 (FUJIFILM Wako, Osaka, Japan) were seeded in 6-well plate. The next day cell culture medium was changed to mTeSR™ plus only. The culture medium was refreshed with fresh mTeSR™ plus medium every two days. Subsequently, embryoid bodies (EBs) were produced by adding 1 × 105 viable cells in mTeSR™ plus medium supplemented with 10 μM Y27632 in 10 cm ultra-low attachment dishes (PrimeSurface®; Sumitomo Co., Ltd., Tokyo, Japan). After three days, the mTeSR™ plus medium was replaced with the chemical induction medium, consisting of fresh E6 medium (Thermo Fisher Scientific Inc., Waltham, MA, USA) supplemented with 10 μM TGF-β kinase/activin receptor-like kinase inhibitor (SB-505124; Santa Cruz Biotechnology Inc., Dallas, TX, USA), 10 μM Y27632 and BMP4 (20 ng mL−1) (PeproTech, Rocky Hill, NJ, USA). These conditions were maintained for 3 days. EBs were then transferred into 6-well plates coated with Matrigel (Geltrex; Thermo Fisher Scientific Inc., Waltham, MA, USA) (6–8 EBs per well). Subsequently, the medium was switched to E6 medium (Thermo Fisher Scientific Inc., Waltham, MA, USA) supplemented with 1% N2 supplement. On day 20, optical vesicle (OV) aggregates were picked with 27G syringe needle (Terumo Corp., Tokyo, Japan) and transferred in the differentiation medium (DM) consisted of DMEM/F12 (3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) medium supplemented with 5% FBS, 2% B27 (Thermo Fisher Scientific Inc., Waltham, MA, USA) and 1% MEM none-essential amino acids (FUJIFILM Wako, Osaka, Japan) to 10 cm ultra-low attachment dishes (PrimeSurface®; Sumitomo Co., Ltd., Tokyo, Japan). The culture medium was refreshed with DM medium supplemented 100 μM taurine (Sigma-Aldrich, St. Louis, MO, USA) every two days. From day 50, 500 nM of retinoic acid (FUJIFILM Wako, Osaka, Japan) was added to the DM for inducing maturation.
1) medium supplemented with 5% FBS, 2% B27 (Thermo Fisher Scientific Inc., Waltham, MA, USA) and 1% MEM none-essential amino acids (FUJIFILM Wako, Osaka, Japan) to 10 cm ultra-low attachment dishes (PrimeSurface®; Sumitomo Co., Ltd., Tokyo, Japan). The culture medium was refreshed with DM medium supplemented 100 μM taurine (Sigma-Aldrich, St. Louis, MO, USA) every two days. From day 50, 500 nM of retinoic acid (FUJIFILM Wako, Osaka, Japan) was added to the DM for inducing maturation.
For culturing human umbilical vein endothelial cells (HUVECs) (Lonza), cells were passaged on 0.1% gelatin pre-coated dishes in DMEM medium supplemented with 10% v/v FBS, 10 ng mL−1 bFGF (FUJIFILM Wako, Osaka, Japan) and 1 ng mL−1 EGF (FUJIFILM Wako, Osaka, Japan) to be known as HUVEC basal medium. The medium was replaced, and the culture medium was refreshed daily. Cell culturing was limited to 9 passages.
3. Endothelial organization in retinal organoid co-culture and VEGF modulation
HUVECs were suspended in an embedding gel consisting of 100 μL bovine fibrinogen (5 mg mL−1; FUJIFILM Wako, Osaka, Japan), 320 μL Matrigel (Geltrex; Thermo Fisher Scientific, Waltham, MA, USA), and 4 μL aprotinin (FUJIFILM Wako, Osaka, Japan). Separately, 1 × 107 cells were resuspended in 425 μL DMEM and then mixed with the embedding gel to yield the final cell–gel mixture, which was kept on ice. Microfluidic devices were sterilized by UV irradiation for 60 min before use. Retinal organoids were transferred to the central chamber of each device using a wide-bore pipette tip, overlaid with 40 μL of the cell–gel mixture, and immediately treated with 10 μL thrombin (5 U mL−1; FUJIFILM Wako, Osaka, Japan). After incubation at 37 °C with 5% CO2 for 10 min to allow gel polymerization, the medium was replaced with HUVEC basal medium supplemented with 1 μg mL−1 aprotinin. For VEGF supplementation experiments, VEGF was added to the HUVEC basal medium at a final concentration of 300 ng mL−1 (FUJIFILM Wako, Osaka, Japan) and applied daily to the device reservoirs. FITC–dextran (0.1 mg mL−1; Sigma-Aldrich, St. Louis, MO, USA) was applied as a fluorescent tracer to visualize vascular network formation. For the lipid nanoparticle delivery assay, rhodamine-labeled liposomes were prepared according to the previous method33 and characterized as described in Method S1 (Table S1). Liposomes were diluted in Hank's balanced salt solution supplemented with calcium and magnesium (HBSS+; Thermo Fisher Scientific) to a final concentration of 0.1 mg mL−1 of lipids and then 10 μL was introduced into the inlet of the microfluidic device. Liposome distribution was assessed using time-lapse fluorescence microscopy, acquiring images at 1 min intervals over a 30 min period (KEYENCE, Tokyo, Japan). After 60 min of exposure, co-cultures were fixed with 4% paraformaldehyde at room temperature for 30 min. Samples were then washed with PBS and imaged by confocal laser scanning microscopy (FV3000; Olympus Corp., Tokyo, Japan).4. Microscopic imaging, histology, and immunofluorescence analysis
For confocal imaging and whole-mount immunofluorescence staining, samples were washed twice with PBS and fixed with 4% paraformaldehyde (PFA) in PBS for 60 min at 25 °C. Fixed samples were permeabilized with 0.1% Triton X-100 overnight at 4 °C, followed by blocking with 5% bovine serum albumin and 0.1% Tween-20 for 90 min at room temperature. Samples were then incubated overnight at 4 °C with primary antibodies diluted in blocking buffer (Table S2). After washing, samples were incubated with secondary antibodies (Alexa Fluor 555 goat anti-mouse IgG or Alexa Fluor 488 goat anti-rabbit IgG; 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500; Thermo Fisher Scientific) for 60 min at 25 °C. Alternatively, samples were stained with ActinGreen 488 (Thermo Fisher Scientific, Waltham, MA, USA) overnight at 4 °C. After two PBS washes, nuclei were counterstained with Hoechst 33342 (10 μg mL−1; Dojindo Molecular Technologies, Kumamoto, Japan) for 60 min at 25 °C. Confocal imaging was performed using an FV3000 scanning microscope (Z-stack thickness: 200 μm (5 μm per slice)) (Olympus Corp., Tokyo, Japan), and images were analysed using FV31S-SW software. Cell morphology and fluorescence intensity were further evaluated using ImageJ software. For histology and section-based immunofluorescence, retinal organoids were fixed with 4% PFA overnight at 4 °C and processed to generate paraffin blocks. Sections (5 μm thickness) were collected and stored at 4 °C. For H&E staining, slides were deparaffinized and stained using standard procedures. For immunofluorescence, sections underwent antigen retrieval in Tris-EDTA buffer (pH 9.0) by heating to boiling for 15 min, followed by permeabilization with 0.5% Triton X-100 in PBS overnight at 4 °C. After blocking with 5% bovine serum albumin and 0.1% Tween-20 for 90 min at room temperature, sections were incubated overnight at 4 °C with primary antibodies (Table S1). After washing, secondary antibodies (Alexa Fluor 555 goat anti-mouse IgG or Alexa Fluor 488 goat anti-rabbit IgG; 1
500; Thermo Fisher Scientific) for 60 min at 25 °C. Alternatively, samples were stained with ActinGreen 488 (Thermo Fisher Scientific, Waltham, MA, USA) overnight at 4 °C. After two PBS washes, nuclei were counterstained with Hoechst 33342 (10 μg mL−1; Dojindo Molecular Technologies, Kumamoto, Japan) for 60 min at 25 °C. Confocal imaging was performed using an FV3000 scanning microscope (Z-stack thickness: 200 μm (5 μm per slice)) (Olympus Corp., Tokyo, Japan), and images were analysed using FV31S-SW software. Cell morphology and fluorescence intensity were further evaluated using ImageJ software. For histology and section-based immunofluorescence, retinal organoids were fixed with 4% PFA overnight at 4 °C and processed to generate paraffin blocks. Sections (5 μm thickness) were collected and stored at 4 °C. For H&E staining, slides were deparaffinized and stained using standard procedures. For immunofluorescence, sections underwent antigen retrieval in Tris-EDTA buffer (pH 9.0) by heating to boiling for 15 min, followed by permeabilization with 0.5% Triton X-100 in PBS overnight at 4 °C. After blocking with 5% bovine serum albumin and 0.1% Tween-20 for 90 min at room temperature, sections were incubated overnight at 4 °C with primary antibodies (Table S1). After washing, secondary antibodies (Alexa Fluor 555 goat anti-mouse IgG or Alexa Fluor 488 goat anti-rabbit IgG; 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 500) were applied for 60 min at 25 °C. Finally, slides were mounted using antifade medium containing DAPI (Fluoro-KEEPER; Nacalai Tesque, Kyoto, Japan) and imaged using a fluorescence microscope (KEYENCE, Tokyo, Japan). For the investigation of cell morphology, area, and quantification of mean fluorescence intensity (MFI), images were analyzed using ImageJ software. CD31-positive endothelial area and RPE dark-region area were independently segmented and quantified, and the ratio of CD31-positive area localized within RPE-associated regions was calculated to determine the fraction of RPE-localized endothelial signal.
500) were applied for 60 min at 25 °C. Finally, slides were mounted using antifade medium containing DAPI (Fluoro-KEEPER; Nacalai Tesque, Kyoto, Japan) and imaged using a fluorescence microscope (KEYENCE, Tokyo, Japan). For the investigation of cell morphology, area, and quantification of mean fluorescence intensity (MFI), images were analyzed using ImageJ software. CD31-positive endothelial area and RPE dark-region area were independently segmented and quantified, and the ratio of CD31-positive area localized within RPE-associated regions was calculated to determine the fraction of RPE-localized endothelial signal.
5. VEGF quantification by ELISA
VEGF secretion was quantified from culture medium collected from the outlet reservoirs of the microfluidic devices. Samples were collected and stored at −30 °C until analysis. Human VEGF-A concentrations were measured using a commercially available ELISA kit (human VEGF-A ELISA kit; Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer's instructions. Absorbance was measured at 450 nm using a microplate reader. VEGF concentrations were calculated from a standard curve generated using recombinant human VEGF-A and normalized to the well volume (0.2 mL). Samples below the detection limit were reported as not detected (N.D.).6. Data visualization and statistics
All quantitative data are presented as mean ± standard error of the mean (SEM) of at least three biological replicates. Graphs generated using GraphPad Prism 9 (GraphPad Software, San Diego, CA, USA).Results
Design and fabrication of a 3D-printed microfluidic device for retinal organoid co-culture with endothelial cells
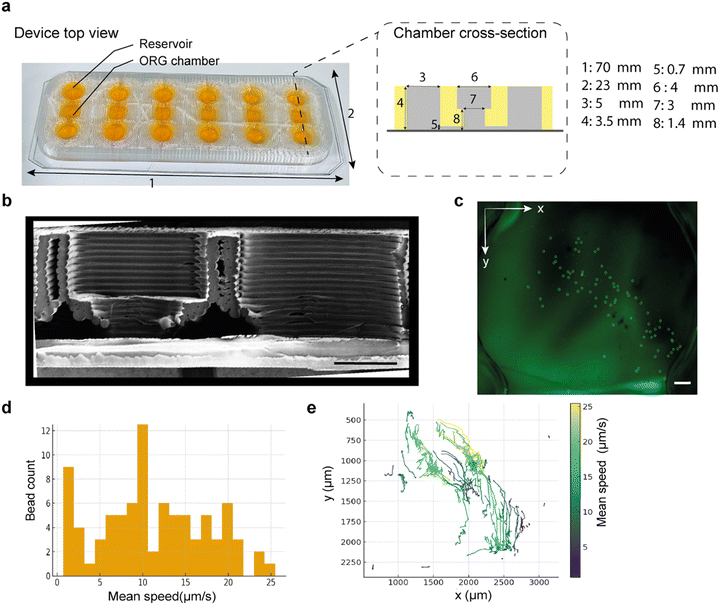
To develop a physiologically relevant co-culture platform for retinal organoids and endothelial cells without relying on a physical barrier such as a porous membrane, we fabricated a multiwell microfluidic device using flexible TPU polymer via FDM 3D printing, bonded onto a PVC substrate (Fig. 1). The rationale for this open co-culture design was to allow direct organoid–endothelial interactions, allowing tissue–tissue communication and enabling paracrine and matrix-mediated signaling across a continuous gel interface without artificial separation. Each device comprises six independent chambers, each with a central cylindrical compartment (3.5 mm diameter; 1.5 mm height) for organoid culture and two lateral side reservoirs for media exchange (Fig. S1). A schematic view of the chamber geometry and flow path is shown in Fig. 2a. The chamber architecture was selected to support gel injection, stable organoid placement, and unidirectional perfusion from reservoir to chamber outlet. To assess fabrication accuracy, we calculated the printing fidelity (PF) ratio as the ratio of printed to designed dimensions, which was 0.89 (10–11% deviation) (Fig. S2). This level of fidelity is consistent with the expected dimensional variation for FDM-printed microfluidic devices, particularly those using flexible filaments such as TPU, as reported in prior studies.34 SEM imaging revealed well-defined filament deposition, consistent interlayer adhesion, and preserved chamber walls (Fig. 2b). The characteristic layered morphology of FDM printing was observed but did not compromise chamber integrity or bonding to the PVC substrate. Flow behaviour within the device was examined by introducing fluorescent 2 μm latex microbeads into one of the side reservoirs and monitoring their passive movement through the reservoir–chamber–outlet pathway (Fig. 2c) (Video S1). The resulting bead velocities ranged primarily from 5–15 μm s−1 (Fig. 2d), and trajectory mapping demonstrated reproducible bead displacement across the chamber (Fig. 2e). No stagnant regions or discontinuities in movement were observed. The observed velocity variations reflect expected flow patterns in open-well geometries and FDM-printed microstructures, where local chamber shape and surface texture produce minor differences in streamline behaviour. Importantly, beads moved continuously throughout the chamber, indicating unobstructed connectivity suitable for subsequent co-culture experiments. Overall, these results confirm that the TPU–PVC device exhibits consistent structural fidelity, stable bonding, and reliable passive fluid transport. These characteristics support its application as a practical microfluidic platform for combining retinal organoids with endothelial networks in a shared hydrogel environment.Characterization of hiPSC-derived retinal organoids for integration into the microfluidic platform
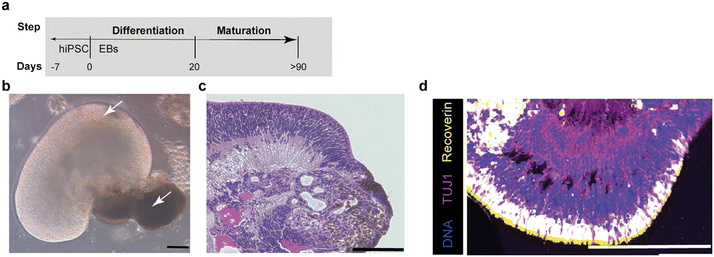
To establish a physiologically relevant retinal component for our Ooc system, we first generated retinal organoids from hiPSCs following our established differentiation and maturation workflow (Fig. 3a). EBs were formed from undifferentiated hiPSCs and transitioned through early retinal differentiation. Organoids exhibited key morphological features of developing retina—including a laminated neuroepithelium and distinct RPE-like pigmented regions (Fig. 3b, arrows)—consistent with observations reported in previous studies.35–37 Histological examination using hematoxylin and eosin staining confirmed the presence of organized retinal layers, showing dense neuroepithelial structures, emerging outer retinal regions, and clearly identifiable pigmented RPE areas indicative of early retinal patterning (Fig. 3c). Immunofluorescence staining further validated this architecture, revealing strong TUJ1 expression in neuronal populations and Recoverin expression in photoreceptors, arranged in spatially distinct layers typical of developing human retina (Fig. 3d). These findings demonstrate that organoids already contain neuronal and photoreceptor elements and recapitulate key features of retinal tissue organization. Collectively, these results confirm that our hiPSC-derived retinal organoids possess the structural complexity and cellular diversity necessary for physiologically relevant retinal modeling. The presence of photoreceptor, neuronal, and RPE-like compartments provides a robust biological foundation for their subsequent integration into the microfluidic platform.Establishment of vascular networks around retinal organoids
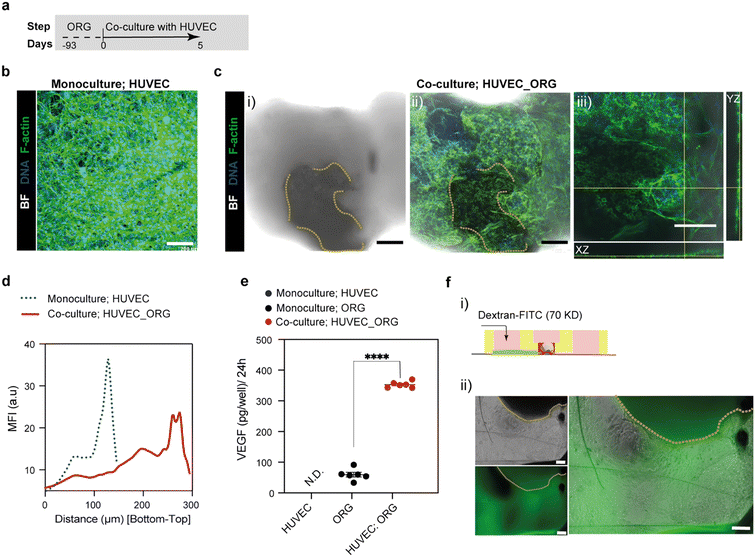
hiPSC-derived retinal organoids containing RPE regions were generated over 93 days and subsequently co-cultured with HUVEC embedded in a fibrin-based matrix within the microfluidic device for 5 days (Fig. 4a). In HUVEC monoculture, endothelial cells formed a dense but largely planar vascular network with limited vertical organization (Fig. 4b). In contrast, co-culture with retinal organoids induced pronounced endothelial remodeling, characterized by spatially organized and interconnected vascular structures preferentially distributed around the organoid RPE dark regions, where a polygonal morphology was observed by F-actin staining, without direct invasion into the organoid tissue (Fig. 4c). Fluorescence imaging and orthogonal XZ and YZ views revealed three-dimensional endothelial architectures extending along the RPE–gel interface, forming a thin layer with an approximate thickness of 2–4 μm, reminiscent of the Bruch's membrane–RPE interface reported in vivo.38,39 This spatial organization indicates active endothelial reorganization driven by organoid-derived cues rather than passive surface association. Quantitative fluorescence intensity profiling along the vertical axis further demonstrated a marked redistribution of endothelial signal in co-culture, with a broader and shifted mean fluorescence intensity profile toward the lower regions corresponding to the RPE-localized area, compared with monoculture (Fig. 4d), consistent with organoid-guided endothelial positioning within the 3D microenvironment. To investigate RPE-derived paracrine signaling, VEGF secretion was quantified. Retinal organoids cultured alone secreted VEGF at levels of approximately 60 pg per well, consistent with previous reports of VEGF secretion from hiPSC-derived RPE cells in the basolateral direction of microfluidic devices.22 In contrast, VEGF levels were below the detection limit in HUVEC monoculture. Notably, VEGF secretion was significantly increased in the organoid–endothelial co-culture condition, reaching approximately 350 pg per well (Fig. 4e), suggesting synergistic bidirectional signaling between retinal tissue and endothelial cells.Similar endothelial-mediated enhancement of VEGF levels in RPE has been reported in choroidal tissues from murine models,40 highlighting a close interplay between RPE-derived VEGF signaling and the adjacent choroidal endothelium. Building on this observation, we next assessed macromolecular transport across the organoid–endothelial interface. Functional assessment using 70 kDa FITC–dextran revealed localized and spatially restricted transport (Fig. 4f), indicating that the remodeled endothelial network influences macromolecular diffusion rather than permitting unrestricted permeation.
VEGF-dependent enhancement of RPE-associated vascular organization
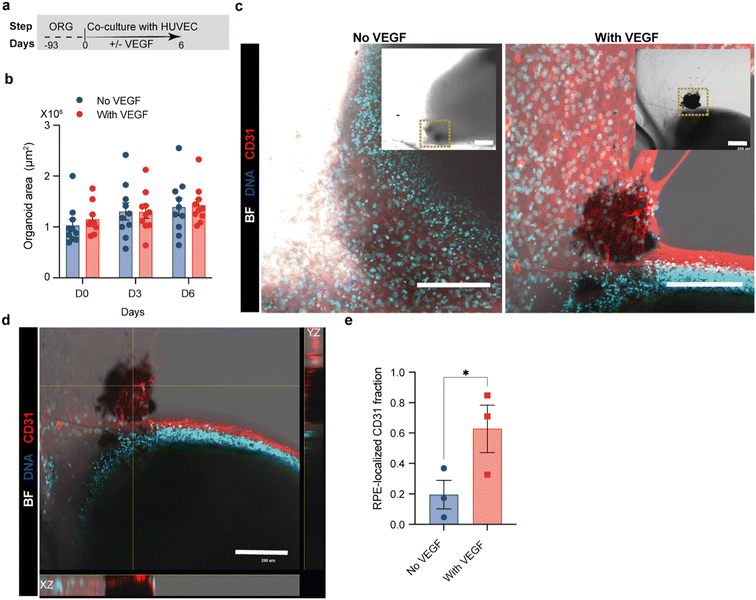
To further evaluate the role of VEGF in regulating endothelial organization in retinal organoids, hiPSC-derived retinal organoids were co-cultured with HUVECs in the presence or absence of exogenously supplemented VEGF for up to 6 days (Fig. 5a). Quantification of organoid area over time revealed no significant differences between VEGF-treated and untreated conditions at days 0, 3, or 6 (Fig. 5b), indicating that VEGF supplementation did not measurably affect organoid growth or gross morphology over the experimental period. In the absence of VEGF, endothelial cells formed vascular networks that remained largely distributed within the surrounding gel, exhibiting limited association with the organoid surface (Fig. 5c, left). In contrast, VEGF supplementation markedly enhanced endothelial accumulation and organization at the organoid interface, with dense CD31-positive structures preferentially localized to RPE-associated regions of the organoid (Fig. 5c, right). Orthogonal XZ and YZ views further confirmed that VEGF promoted the formation of three-dimensional endothelial structures extending along the RPE–gel interface, rather than inducing uniform vascular expansion throughout the extracellular matrix (Fig. 5d). Quantitative analysis demonstrated a significant increase in the fraction of CD31-positive signal localized to RPE regions in the VEGF-treated condition compared with the no-VEGF control (Fig. 5e), indicating that VEGF acts to potentiate organoid-guided endothelial positioning rather than inducing nonspecific angiogenic growth. This spatially biased endothelial response is consistent with previous report24 showing that elevated VEGF levels are sufficient to induce choroidal neovascularization (CNV)-like properties, a pathological process characterized by aberrant vascular growth toward and through the RPE–Bruch's membrane interface. Together, these findings suggest that while retinal organoids provide intrinsic spatial cues for endothelial organization, excessive VEGF functions as a key modulatory factor that amplifies RPE-associated vascular patterning. Accordingly, the present organoid-on-chip platform captures VEGF-responsive interface remodeling relevant to CNV biology and may therefore serve as a useful system for evaluating anti- or pro-angiogenic strategies targeting CNV-associated pathways.Dynamic transport of lipid nanoparticles at the organoid-vascular interface
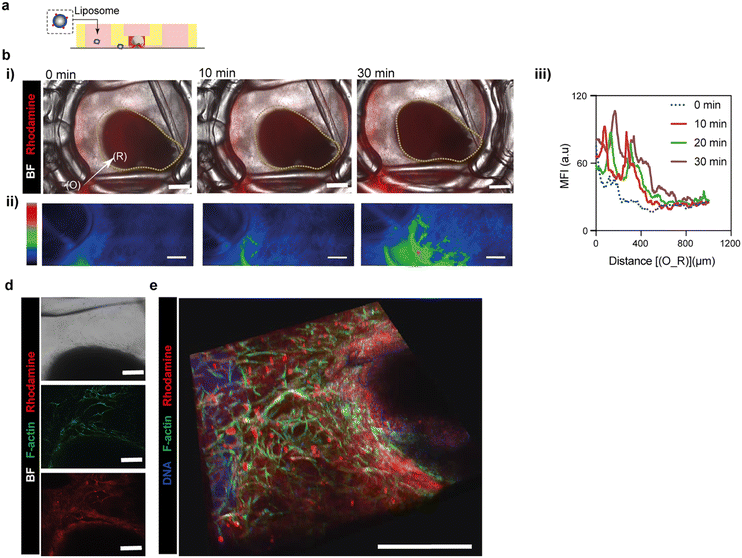
To assess the functional transport properties of the vascularized retinal organoid-on-chip system, rhodamine-labeled liposomes were introduced into the perfusion channel, and their spatiotemporal distribution was monitored in real time (Fig. 6a). Time-lapse imaging revealed progressive accumulation of liposomal signal in regions proximal to the retinal organoid over 30 min (Fig. 6b-i) (Video S2), with fluorescence intensity increasing preferentially along the organoid-facing side of the vascularized gel. Pseudocolor fluorescence mapping further illustrated the directional spread of liposomes toward the organoid interface, indicating non-uniform transport rather than homogeneous diffusion through the matrix (Fig. 6b-ii). In contrast, organoid-only controls embedded in gel without vascular networks exhibited minimal liposomal signal and limited spatial distribution (Fig. S3), highlighting the contribution of the vascular architecture to nanoparticle transport. Quantitative analysis confirmed a time-dependent increase in mean fluorescence intensity along the distance from the perfusion region toward the organoid boundary, with a clear shift in signal distribution over time (Fig. 6b-iii), demonstrating controlled and gradual liposomal nanoparticle transport within the device. High-magnification imaging revealed colocalization of rhodamine-labeled liposomes with endothelial structures in the vascular network adjacent to the organoid (Fig. 6d), suggesting that endothelial architecture influences liposomal nanoparticle localization and retention. Three-dimensional reconstruction further confirmed that liposomes distributed along vascularized regions surrounding the organoid rather than directly penetrating the organoid tissue (Fig. 6e), indicating that transport is regulated at the organoid–vascular interface. These results indicate that endothelial networks formed in the device can mediate nanoparticle distribution toward organoid tissue, providing a basis for future drug testing in a controlled retinal model.Limitations and future directions
While the presented co-culture system successfully recapitulates key features of choroid-like endothelial organization and VEGF-modulated vascular remodeling, several limitations should be acknowledged. First, the fibrin-based extracellular matrix (ECM), although well suited for three-dimensional endothelial network formation and spatial control, introduces partial physical separation between endothelial cells and RPE regions. Consequently, endothelial cells may also interact with exposed retinal domains, including photoreceptor-containing regions. Future modifications to the device architecture and ECM configuration may help promote more defined and sustained RPE–endothelial coupling. Second, although HUVECs represent a robust and widely used endothelial model, they may not fully reflect the molecular identity and functional properties of choroidal endothelial cells. Incorporation of hiPSC-derived or primary choroidal endothelial cells would further improve tissue specificity and physiological relevance. Finally, future studies could leverage patient-derived retinal organoids, particularly from individuals with AMD, to examine disease-dependent differences in endogenous VEGF secretion, endothelial organization, and barrier function. Systematic comparison of vascular network formation in patient-derived organoids with and without VEGF supplementation may enable evaluation of the platform's ability to reproduce choroidal vascular abnormalities associated with AMD. Together, advances in device/ECM designs that strengthen RPE–endothelial communication, combined with deeper molecular profiling (e.g., transcriptomics and proteomics), will facilitate the development of more fully integrated vascularized retinal tissue-on-chip platforms and expand their utility for patient-specific disease modeling and predictive therapeutic assessment.Conclusions
In this study, we established a fully 3D-printed retinal organoid-on-chip platform that enables direct co-culture of hiPSC-derived retinal organoids containing intrinsic RPE regions with endothelial cells. By eliminating semipermeable barriers and embedding both cell types within a fibrin–Matrigel matrix, the system supports physiologically relevant organoid–endothelial interactions. We demonstrated that endothelial cells self-organize into vascular networks that spatially associate with retinal organoids. Under co-culture conditions, endogenous VEGF secretion was significantly increased, while exogenous VEGF supplementation further promoted vascular network expansion in proximity to RPE regions without affecting organoid growth. In addition, the platform supported transport of liposomal nanoparticles through the engineered vascular networks, demonstrating its applicability for drug delivery studies. Overall, this platform provides a useful in vitro tool for investigating retinal angiogenesis and evaluating therapeutic strategies.Author contributions
Rodi Kado Abdalkader: conceptualized and managed the project, designed, and performed experiments, analyzed, and interpreted data, visualized the data, and wrote the manuscript. Shigeru Kawakami, Yuuki Takashima, and Takuya Fujita: provided resources and contributed to manuscript refinement. All authors critically reviewed the manuscript and agreed with the publication.Conflicts of interest
All authors declare no competing financial interest.Data availability
Data supporting this article, have been included as part of the supplementary information (SI).Supplementary information is available. See DOI: https://doi.org/10.1039/d5lc00939a.
Acknowledgements
We acknowledge the Ritsumeikan Global Innovation Research Organization (R-GIRO). We are also gratefully acknowledging the Biomedical Engineering Center (BMEC) at Ritsumeikan University for providing access to the confocal scanning microscope. This work was generously supported by the Japan Society for the Promotion of Science 24K15712 and 24K22396.Notes and references
- S. Resnikoff, D. Pascolini, D. Etya'ale, I. Kocur, R. Pararajasegaram, G. P. Pokharel and S. P. Mariotti, Bull. W. H. O., 2004, 82, 844–851 Search PubMed.
- Y. Deng, L. Qiao, M. Du, C. Qu, L. Wan, J. Li and L. Huang, Genes Dis., 2022, 9, 62–79 CrossRef CAS PubMed.
- P. Mitchell, G. Liew, B. Gopinath and T. Y. Wong, Lancet, 2018, 392, 1147–1159 CrossRef PubMed.
- R. W. N. Slijkerman, F. Song, G. D. N. Astuti, M. A. Huynen, E. van Wijk, K. Stieger and R. W. J. Collin, Prog. Retinal Eye Res., 2015, 48, 137–159 CrossRef PubMed.
- K. C. Eldred and T. A. Reh, Dev. Biol., 2021, 480, 114–122 CrossRef CAS PubMed.
- K. Achberger, J. C. Haderspeck, A. Kleger and S. Liebau, Adv. Drug Delivery Rev., 2019, 140, 33–50 CrossRef CAS PubMed.
- Y. Zhu, B. Cao, A. Tolone, J. Yan, G. Christensen, B. Arango-Gonzalez, M. Ueffing and F. Paquet-Durand, Front. Neurosci., 2022, 16, 938089 CrossRef PubMed.
- M. Völkner, M. Zschätzsch, M. Rostovskaya, R. W. Overall, V. Busskamp, K. Anastassiadis and M. O. Karl, Stem Cell Rep., 2016, 6, 525–538 CrossRef PubMed.
- X. Zhong, C. Gutierrez, T. Xue, C. Hampton, M. N. Vergara, L.-H. Cao, A. Peters, T. S. Park, E. T. Zambidis, J. S. Meyer, D. M. Gamm, K.-W. Yau and M. V. Canto-Soler, Nat. Commun., 2014, 5, 4047 CrossRef CAS PubMed.
- A. Sridhar, A. Hoshino, C. R. Finkbeiner, A. Chitsazan, L. Dai, A. K. Haugan, K. M. Eschenbacher, D. L. Jackson, C. Trapnell, O. Bermingham-McDonogh, I. Glass and T. A. Reh, Cell Rep., 2020, 30, 1644–1659.e4 CrossRef CAS PubMed.
- C. S. Cowan, M. Renner, M. De Gennaro, B. Gross-Scherf, D. Goldblum, Y. Hou, M. Munz, T. M. Rodrigues, J. Krol, T. Szikra, R. Cuttat, A. Waldt, P. Papasaikas, R. Diggelmann, C. P. Patino-Alvarez, P. Galliker, S. E. Spirig, D. Pavlinic, N. Gerber-Hollbach, S. Schuierer, A. Srdanovic, M. Balogh, R. Panero, A. Kusnyerik, A. Szabo, M. B. Stadler, S. Orgül, S. Picelli, P. W. Hasler, A. Hierlemann, H. P. N. Scholl, G. Roma, F. Nigsch and B. Roska, Cell, 2020, 182, 1623–1640.e34 CrossRef CAS PubMed.
- M. W. van der Helm, A. D. van der Meer, J. C. T. Eijkel, A. van den Berg and L. I. Segerink, Tissue Barriers, 2016, 4, e1142493 Search PubMed.
- D. E. Ingber, Nat. Rev. Genet., 2022, 23, 467–491 CrossRef CAS PubMed.
- E. Moradi, S. Jalili-Firoozinezhad and M. Solati-Hashjin, Acta Biomater., 2020, 116, 67–83 CrossRef CAS PubMed.
- S. Jalili-Firoozinezhad, F. S. Gazzaniga, E. L. Calamari, D. M. Camacho, C. W. Fadel, A. Bein, B. Swenor, B. Nestor, M. J. Cronce, A. Tovaglieri, O. Levy, K. E. Gregory, D. T. Breault, J. M. S. Cabral, D. L. Kasper, R. Novak and D. E. Ingber, Nat. Biomed. Eng., 2019, 3, 520–531 Search PubMed.
- L. Amirifar, A. Shamloo, R. Nasiri, N. R. de Barros, Z. Z. Wang, B. D. Unluturk, A. Libanori, O. Ievglevskyi, S. E. Diltemiz, S. Sances, I. Balasingham, S. K. Seidlits and N. Ashammakhi, Biomaterials, 2022, 285, 121531 CrossRef CAS PubMed.
- R. Abdalkader and K. Kamei, Lab Chip, 2020, 20, 1410–1417 RSC.
- D. Bennet, Z. Estlack, T. Reid and J. Kim, Lab Chip, 2018, 18, 1539–1551 RSC.
- T. Gensheimer, D. Veerman, E. M. van Oosten, L. Segerink, A. Garanto and A. D. van der Meer, Lab Chip, 2025, 25, 996–1014 RSC.
- S. N. Bhatia and D. E. Ingber, Nat. Biotechnol., 2014, 32, 760–772 Search PubMed.
- D. E. Ingber, Adv. Sci., 2020, 7, 2002030 Search PubMed.
- K. Achberger, C. Probst, J. Haderspeck, S. Bolz, J. Rogal, J. Chuchuy, M. Nikolova, V. Cora, L. Antkowiak, W. Haq, N. Shen, K. Schenke-Layland, M. Ueffing, S. Liebau and P. Loskill, eLife, 2019, 8, e46188 CrossRef PubMed.
- S. Lee, S. Kim and J. S. Jeon, Lab Chip, 2022, 22, 4359–4368 Search PubMed.
- M. Chung, S. Lee, B. J. Lee, K. Son, N. L. Jeon and J. H. Kim, Adv. Healthcare Mater., 2018, 7, 1700028 Search PubMed.
- M. Cipriano, K. Schlünder, C. Probst, K. Linke, M. Weiss, M. J. Fischer, L. Mesch, K. Achberger, S. Liebau, M. Mesquida, V. Nicolini, A. Schneider, A. M. Giusti, S. Kustermann and P. Loskill, Commun. Biol., 2022, 5, 52 Search PubMed.
- U. Nam, S. Lee and J. S. Jeon, ACS Biomater. Sci. Eng., 2023, 9, 4929–4939 Search PubMed.
- K. Achberger, M. Cipriano, M. J. Düchs, C. Schön, S. Michelfelder, B. Stierstorfer, T. Lamla, S. G. Kauschke, J. Chuchuy, J. Roosz, L. Mesch, V. Cora, S. Pars, N. Pashkovskaia, S. Corti, S.-M. Hartmann, A. Kleger, S. Kreuz, U. Maier, S. Liebau and P. Loskill, Stem Cell Rep., 2021, 16, 2242–2256 Search PubMed.
- S. Corti, K. H. Kim, T. Chen, A. Botezatu, V. Cora, K. Ma, N. Pashkovskaia, A. Bernal Vergara, D. Sperlich, K. Dave, A. Tolone, R. M. Reddinger, C. B. Tully, M. Higgins, A. Kleger, M. Breunig, P. Lopatta, S. Wingerter, M. Cipriano, S. Bolz, M. Ueffing, N. Buss, P. Loskill, S. Liebau and K. Achberger, Cell Rep. Med., 2025, 6, 102244 Search PubMed.
- R. Kado Abdalkader, S. Konishi and T. Fujita, Sci. Rep., 2025, 15, 6125 CrossRef CAS PubMed.
- R. Abdalkader, S. Konishi and T. Fujita, Biomimetics, 2021, 7, 2 Search PubMed.
- J. Schindelin, I. Arganda-Carreras, E. Frise, V. Kaynig, M. Longair, T. Pietzsch, S. Preibisch, C. Rueden, S. Saalfeld, B. Schmid, J.-Y. Tinevez, D. J. White, V. Hartenstein, K. Eliceiri, P. Tomancak and A. Cardona, Nat. Methods, 2012, 9, 676–682 Search PubMed.
- K. Okita, T. Yamakawa, Y. Matsumura, Y. Sato, N. Amano, A. Watanabe, N. Goshima and S. Yamanaka, Stem Cells, 2013, 31, 458–466 CrossRef CAS PubMed.
- R. Abdalkader, S. Kawakami, J. Unga, Y. Higuchi, R. Suzuki, K. Maruyama, F. Yamashita and M. Hashida, Drug Delivery, 2017, 24, 320–327 CrossRef CAS PubMed.
- S. Waheed, J. M. Cabot, N. P. Macdonald, T. Lewis, R. M. Guijt, B. Paull and M. C. Breadmore, Lab Chip, 2016, 16, 1993–2013 RSC.
- A. Hoshino, R. Ratnapriya, M. J. Brooks, V. Chaitankar, M. S. Wilken, C. Zhang, M. R. Starostik, L. Gieser, A. La Torre, M. Nishio, O. Bates, A. Walton, O. Bermingham-McDonogh, I. A. Glass, R. O. L. Wong, A. Swaroop and T. A. Reh, Dev. Cell, 2017, 43, 763–779.e4 CrossRef CAS PubMed.
- W.-L. Deng, M.-L. Gao, X.-L. Lei, J.-N. Lv, H. Zhao, K.-W. He, X.-X. Xia, L.-Y. Li, Y.-C. Chen, Y.-P. Li, D. Pan, T. Xue and Z.-B. Jin, Stem Cell Rep., 2018, 10, 1267–1281 CrossRef CAS PubMed.
- K. C. Eldred, S. E. Hadyniak, K. A. Hussey, B. Brenerman, P.-W. Zhang, X. Chamling, V. M. Sluch, D. S. Welsbie, S. Hattar, J. Taylor, K. Wahlin, D. J. Zack and R. J. Johnston, Science, 2018, 362, eaau6348 CrossRef PubMed.
- G. S. Hageman, P. J. Luthert, N. H. V. Chong, L. V. Johnson, D. H. Anderson and R. F. Mullins, Prog. Retinal Eye Res., 2001, 20, 705–732 CrossRef CAS PubMed.
- R. S. Ramrattan, T. L. Van der Schaft, C. M. Mooy, W. C. De Bruijn, P. G. H. Mulder and P. T. V. M. De Jong, Invest. Ophthalmol. Visual Sci., 1994, 35, 2857–2864 Search PubMed.
- H. Wang, P. Geisen, E. S. Wittchen, B. King, K. Burridge, P. A. D'Amore and M. E. Hartnett, Invest. Ophthalmol. Visual Sci., 2011, 52, 570–578 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |