 Open Access Article
Open Access ArticleMicrofluidic capillary transit velocity as a functional measure for sickle cell disease and in vitro-derived red blood cells
Solomon
Oshabaheebwa
 a,
Utku
Goreke
a,
Utku
Goreke
 b,
Yuxuan
Du
c,
Christopher L.
Wirth
b,
Yuxuan
Du
c,
Christopher L.
Wirth
 d,
Zoe
Sekyonda
a,
Bryan L.
Benson
d,
Zoe
Sekyonda
a,
Bryan L.
Benson
 ei,
Payam
Fadaei
c,
Yusang B.
Ley
a,
Nathan M.
Perez
b,
Petros
Giannikopoulos
b,
David N.
Nguyen
ei,
Payam
Fadaei
c,
Yusang B.
Ley
a,
Nathan M.
Perez
b,
Petros
Giannikopoulos
b,
David N.
Nguyen
 bfg,
Michael A.
Suster
h,
Pedram
Mohseni
bfg,
Michael A.
Suster
h,
Pedram
Mohseni
 *ah and
Umut A.
Gurkan
*ah and
Umut A.
Gurkan
 *ac
*ac
aDepartment of Biomedical Engineering, Case Western Reserve University, Cleveland, OH, USA
bInnovative Genomics Institute, University of California, Berkeley, CA, USA
cDepartment of Mechanical and Aerospace Engineering, Case Western Reserve University, Cleveland, OH 44106, USA. E-mail: umut@case.edu; Tel: +1 (216) 368 6447
dDepartment of Chemical and Biomolecular Engineering, Case Western Reserve University, Cleveland, OH, USA
eDepartment of Anesthesiology, Case Western Reserve University School of Medicine, Cleveland, OH, USA
fDepartment of Medicine, University of California, San Francisco, CA, USA
gDepartment of Molecular and Cell Biology, University of California, Berkeley, CA, USA
hDepartment of Electrical, Computer, and Systems Engineering, Case Western Reserve University, Cleveland, OH 44106, USA. E-mail: pxm89@case.edu; Tel: +1 (216) 368 5263
iDepartment of Anesthesiology, University Hospitals Cleveland Medical Center, Cleveland, OH, USA
First published on 21st January 2026
Abstract
Emerging therapies in sickle cell disease (SCD) aim to restore healthy red blood cell (RBC) function, but they often yield heterogeneous cellular responses. There are no proven techniques to evaluate restored rheological functionality and heterogeneity in these RBCs. We present a biomimetic microcapillary network, high-speed imaging, and computational algorithms to analyze RBC capillary velocity profiles of the entire sample population at single-cell resolution. Using peripheral RBCs from SCD patients and healthy donors, we showed that RBC capillary transit velocity correlated with cell shape, hydrodynamic adaptability, and elongation index. Healthy RBCs exhibited a velocity distribution skewed toward higher values, whereas RBCs from individuals with SCD showed a shift toward lower velocities. SCD samples had a greater fraction of slow RBCs than healthy controls (42.1% ± 12.0% vs. 19.0% ± 4.9%, p < 0.0001). We tested mixtures of healthy and SCD RBCs to simulate heterogeneous therapeutic effects and demonstrated that the assay was sensitive to small fractions of abnormal RBCs. The slow RBC fraction emerged as a potential biomarker associated with SCD disease severity. This fraction significantly increased under hypoxia showing sensitivity to hypoxia-induced sickling. Finally, we assessed in vitro-derived RBCs and observed distinct velocity profiles for nucleated and enucleated cells. Processing methods to enrich enucleated RBCs improved the velocity profile, producing a distribution that was more comparable to that of peripheral RBCs. This platform's ability to assess individual RBCs and generate a velocity profile from a small number of cells makes it well suited for evaluating the rheological properties of in vitro-derived RBCs.
Introduction
Sickle cell disease (SCD) is the most common genetic blood disorder affecting more than 7 million people globally.1 It is caused by a single point mutation in the hemoglobin molecule that results in the production of sickle hemoglobin (HbS).2 HbS molecules bind together when deoxygenated to form stiff fibers which distort RBCs into spiked or crescent shaped cells with reduced deformability and increased stickiness.2,3 These stiff and sticky sickle RBCs may be trapped in narrow capillaries and venules and contribute to acute vaso-occlusive events (VOEs) which are the leading cause of hospitalization in people living with SCD.4,5 Circulation of abnormal RBCs throughout the body results in a wide range of clinical complications, including stroke, acute chest syndrome (ACS), pulmonary hypertension, kidney disease, splenic complications, and other musculoskeletal effects.6,7 The manifestation of these complications is clinically variable across patients making disease prognosis highly unpredictable.6Next-generation therapies for SCD target hematopoietic stem cells (HSCs) to correct or modify the underlying genetic mutation. This approach enables production of RBCs with non-polymerizing hemoglobin variants. However, these therapies show heterocellular effects, with individual RBCs showing different levels of the therapeutic hemoglobin.8,9In vitro-derived RBCs are increasingly used to evaluate SCD gene therapies before advancing to animal and human studies.10,11 In this approach, HSCs are edited to correct hemoglobin mutations or sustain HbF production, then differentiated in vitro to assess the therapeutic effect in the resulting RBCs.12–14 While most studies measure intended and unintended indels, or HbF/hemoglobin correction, functional assays remain limited. Some groups have applied static sickling assays,12,15 but no validated methods exist for evaluating rheological properties of in vitro-derived RBCs, an important indication of therapeutic efficacy. Assessment of in vitro-derived RBCs is further complicated by small sample volumes and high variability in cell morphology.16 These challenges underscore the need for technologies with sensitivity to cell-to-cell variability rather than bulk rheology assays. Cell-to-cell variability in therapeutic response has also been reported in other pharmacological drugs including Hydroxyurea which increases production of fetal hemoglobin (HbF) with heterocellular variability.17 A recent study also found that voxelotor, a precursor to GBT02601, binds heterogeneously to hemoglobin at the molecular level.18 These cellular variations have profound ramifications on the rheological properties of blood, particularly in the microcirculation where the size of the individual RBC approaches and, in the case of capillaries, exceeds the vessel diameter. The heterogeneity in rheological properties of RBCs could explain the clinical variability in disease expression.
Conventional methods of assessing cellular deformability at the single cell level include atomic force microscopy (AFM),19 optical tweezers,20 and micropipette aspiration.21 These have a low throughput which prevents a thorough characterization of the entire RBC population. Additionally, they require specialized equipment and skilled personnel which makes them inaccessible to most research and clinical settings. To overcome these limitations, bulk approaches such as ektacytometry and rheometry have been employed.22,23 However, these provide an average measurement of the entire cell population, masking single-cell-level heterogeneity. These techniques are also designed for large-volume peripheral samples and cannot accurately characterize in vitro-derived RBCs given their small volumes and intrinsic heterogeneity.
Recently, microfluidic devices have been utilized to measure the deformability of individual RBCs in capillary-like environments leveraging the ability to mimic capillary geometry.24–27 RBC deformability can be quantified by measuring capillary transit velocity or transit time where deformable RBCs readily pass through narrow channels, while rigid RBCs experience increased resistance and longer transit times. Alternatively, RBC imaging techniques have been utilized to measure the elongation index by comparing the major and minor radii of the cell subjected to shear stress. While these microfluidic techniques measure RBC mechanical properties with single-cell resolution, they have limitations in throughput,26 or utilize channels wider than the RBC diameter that may not fully replicate the confined environment and cell–wall interactions present in capillaries.27 Many of these techniques implement single channels that might not fully replicate the complex and varied flow conditions found in the in vivo capillary network.25 Recently, Williams et al., developed a microfluidic platform that simultaneously measured single RBC deformability and oxygen saturation using a high-speed camera at 2000 fps.24 While this allowed for detailed biophysical measurements, the requirement for such sophisticated imaging equipment limits its accessibility in most research or clinical facilities. Additionally, there are several commercially-available microfluidic systems that assess RBC properties including shape and velocity with single-cell resolution including Erysense,28 AcCellerator/Shape-In,29 and LaVision Micro-Particle Image Velocimetry (μPIV) systems.30 These systems are however characterized by highly sophisticated hardware and software packages that remain prohibitive in most research settings. Importantly, these techniques have not been validated for evaluating the rheological properties of in vitro-derived RBCs, a critical gap in the advancement of next-generation SCD gene therapies.
Here we present methods of quantifying RBC rheological properties including capillary transit velocity, RBC shape, elongation index and hydrodynamic adaptability in a microfluidic capillary network at single-cell resolution. We developed a high-throughput assay that generated capillary transit velocity profiles for the entire red blood cell population and quantified the slow RBC fraction to identify cells with impaired capillary flow dynamics. To validate the assay's clinical relevance, we investigated associations between the slow RBC fraction in SCD and conventional SCD severity markers including lactate dehydrogenase (LDH), absolute reticulocyte count (ARC), C-reactive protein (CRP) and the frequency of hospital admissions. We also evaluated the sensitivity of the assay to hypoxia-induced RBC sickling by evaluating the velocity profiles under hypoxia with and without treatment with GBT021601, an inhibitor of HbS polymerization. Finally, we utilized the assay to analyze the velocity profiles of in vitro-derived RBCs.
Methods
Design and fabrication of the microfluidic devices
Microfluidic devices were fabricated with polydimethylsiloxane (PDMS) using soft lithography techniques as previously described.31,32 Each microfluidic device comprised six discrete arrays of micropillars. The spacing between micropillars was uniform within each array but decreased progressively in subsequent arrays from 12 μm in the first array (nearest to the inlet) to 3 μm in the sixth array (Fig. 1A). The PDMS substrate imprinted with the capillary network was bonded to a plain glass slide using oxygen plasma, yielding a fully assembled microfluidic device. Prior to running the experiments, the microfluidic devices were washed with pure ethanol, rinsed with 1X PBS, adsorbed with 3% (w/v) bovine serum albumin, and incubated overnight at 4 °C to prevent non-specific binding as previously described.31Development of a gas chamber for modulating oxygen tension in the microfluidic device
To modulate the oxygen environment in the capillary network, a gas chamber was integrated on top of the PDMS block containing the capillary network. The gas chamber was fabricated using poly(methyl methacrylate) (PMMA) sheets that were cut with a VLS2.30 laser cutting machine (Universal Laser Systems, AZ, USA) to make the walls and top cover of the gas chamber. The PMMA layers were bonded with double sided adhesive tape (Fig. 5A). The gas chamber was supplied with either compressed air (21% O2) to achieve normoxia conditions, or with compressed nitrogen gas (95% N2 and 5% CO2) to establish a hypoxia environment. Before sample perfusion, the gas chamber was perfused with the controlled gas for 20 minutes to facilitate oxygen depletion of the capillary network across the gas-permeable PDMS layer.Sample preparation
Healthy blood samples (HbAA) and sickle cell blood samples (HbSS) were collected in EDTA tubes from the Hematopoietic Biorepository and Cellular Therapy Core at Case Western Reserve University and the University Hospitals Cleveland Medical Center (UHCMC), respectively under institutional review board (IRB)-approved protocols. All samples were stored at 4 °C and tested within 24 hours of blood collection. The blood samples were centrifuged at 300g for 5 minutes. The buffy coat and plasma were extracted and discarded leaving only the RBC pellet which was then resuspended in phosphate buffered saline (PBS) to achieve a 0.5% hematocrit. In experiments involving GBT021601 treatment, the RBC pellet was resuspended in a PBS buffer containing either 60 mM of GBT021601 (MedChemExpress, HY-148788) or buffer plus the carrier dimethyl sulfoxide (DMSO, for untreated control samples) and incubated for 2 hours at 37 °C. Clinical data including LDH and ARC in HbSS samples were obtained from hospital records. RBC hemoglobin profiles were determined using HPLC (VARIANT™ II, Bio-Rad Laboratories, Inc., Hercules, CA).Erythroid differentiation and preparation of in vitro-derived RBC samples
Erythroid differentiation was performed in three phases from commercially sourced hematopoietic stem and progenitor cells (HSPCs) as previously described.10 Phase 1 media (days 1–7), consisted of Iscove's modified Dulbecco's medium (IMDM) supplemented with 1% L-glutamine, 2% penicillin–streptomycin (Gibco #15-140-122; stock 100%), 3% heat-inactivated human plasma AB+ (Stemcell Technologies #70039.6), 2% human AB serum (Fisher BioReagents #BP2525100; stock 100%), 330 μg mL−1 holo-human transferrin (Sigma #T0665; stock 25 mg mL−1), 10 μg mL−1 recombinant human insulin (Sigma #I9278; stock 10 mg mL−1), 2 IU mL−1 heparin (SIAL-H3149; stock 50 mg mL−1), 3 IU mL−1 recombinant erythropoietin (Stemcell #78007; stock 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 U mL−1), hydrocortisone at 10−6 M (Stemcell #7926; stock 50 μg mL−1), SCF at 100 ng mL−1 (PeproTech #30007100UG; stock 50 μg mL−1) and IL-3 at 5 ng mL−1 (PeproTech #200-03; stock 50 μg mL−1). In phase 2 (days 7–11), cells were cultured in phase 1 media without IL-3 and hydrocortisone. In phase 3 (days 11–21), cells were cultured in phase 2 media without SCF. Cells were maintained daily at a density of 5 × 104 to 2 × 105 cells per ml (phase 1 & 2) and 1 × 106 per ml (phase 3). On day 21, the first sample was collected for analysis which comprised of a heterogeneous mixture of nucleated and enucleated cells. An aliquot of this sample was filtered through an Acrodisc WBC filter (Pall Corporation, Port Washington, NY) to deplete nucleated cells, yielding a second sample enriched for enucleated RBCs.
000 U mL−1), hydrocortisone at 10−6 M (Stemcell #7926; stock 50 μg mL−1), SCF at 100 ng mL−1 (PeproTech #30007100UG; stock 50 μg mL−1) and IL-3 at 5 ng mL−1 (PeproTech #200-03; stock 50 μg mL−1). In phase 2 (days 7–11), cells were cultured in phase 1 media without IL-3 and hydrocortisone. In phase 3 (days 11–21), cells were cultured in phase 2 media without SCF. Cells were maintained daily at a density of 5 × 104 to 2 × 105 cells per ml (phase 1 & 2) and 1 × 106 per ml (phase 3). On day 21, the first sample was collected for analysis which comprised of a heterogeneous mixture of nucleated and enucleated cells. An aliquot of this sample was filtered through an Acrodisc WBC filter (Pall Corporation, Port Washington, NY) to deplete nucleated cells, yielding a second sample enriched for enucleated RBCs.
Each sample consisted of 2 million cells. The in vitro-derived cells were centrifuged at 500g for 5 minutes and resuspended in 300 μl of PBS. Control samples were prepared by isolation of 2 million peripheral RBCs from a healthy donor via centrifugation and resuspending them in 300 μl of PBS.
Experimental setup
The microfluidic devices were mounted on an Olympus IX83 inverted motorized microscope (Evident, Tokyo, Japan) with a 40× objective and a Prime BSI Express sCMOS camera (Teledyne, ON, Canada). The samples were perfused through the microfluidic devices at precisely controlled driving pressure generated by a Fluigent Flow-EZ™ system (Fluigent, MA, USA). The driving pressure was set to 200 mBar until the RBC flow profile showed even distribution across the capillary network. The microscope stage was then moved to position the field of view (FOV) at the centre of the capillary in network in the desired micropillar array. The pressure was then reduced to 50 mBar and images were captured with the camera at 94 frames per second (fps). 3000 frames were analyzed per experiment. The 4 μm-wide array was used in all experiments as it showed the optimal separation between HbAA and HbSS RBCs (Fig. S1). Experiments presented in Fig. 2 followed the same procedure except that the videos were recorded with an iPhone 13 mini (Apple, CA, USA) mounted on a Leica microscope (Leica Microsystems GmbH, Germany) with a 63× oil-immersion objective and 10x eye piece.Measurement of RBC velocity in the capillary network
RBCs were tracked using an approach in which the image stack was initially processed with Fiji,33 and then subsequently analyzed with MATLAB version 9.13.0 (R2022b, Natick, MA) to identify and track individual RBCs. First, a custom script was written in Fiji to subtract the micropillars from view, generating a frame-by-frame tiff file with isolated RBCs on a clear white background. Briefly, this was achieved by converting each frame to an 8-bit image with binary pixel intensities by thresholding. The first frame was then subtracted from the other frames to eliminate micropillars which were stationary in all frames. The output of the Fiji script was a sequence of frames showing RBCs moving across the FOV with a uniform white background. This sequence of frames was then analyzed with a custom MATLAB code based on existing particle tracking methods commonly used for other problems in colloid science.34–36 The code works by first identifying all RBCs in the frame(s) and then creates tracks based on the relative changes in position in subsequent frames. Velocities were then calculated from tracks of individual RBCs as the ratio of displacement to elapsed time.Quantification of the slow RBC fraction
A kernel density estimation was employed to generate a smooth representation of the RBC velocity distribution. The kernel density estimate was normalized to an area under the curve (AUC) of 1 to determine RBC fractions in various velocity ranges. The absolute RBC velocities from the HbAA and HbSS samples are shown in Fig. S2. To minimize the effect of microfluidic imperfections across devices, the RBC velocities were normalized relative to the maximum RBC velocity observed in the same sample (i.e., normalized velocity = V/Vmax). Vmax was defined from the kernel density distribution as the velocity at which the probability density in the tail approached zero, to avoid bias from outlier events. This normalization method was based on the assumption that each sample contained a population of healthy RBCs that flowed through the capillaries mostly unencumbered by wall collisions or friction. Any decrease in normalized RBC velocity was therefore attributed to alterations in the RBC mechanical properties that increased resistance when traversing narrow capillaries. RBC velocity distribution profiles from 20 healthy donors were analyzed to establish a threshold for identifying the slow RBC fraction. This threshold was defined as one standard deviation below the modal velocity observed in the healthy samples. The slow RBC fraction was then quantified by calculating the AUC below this threshold.Computational modeling of microfluidic hemodynamics
The hemodynamics environment in the microfluidic capillary network was modeled using finite element analysis in COMSOL Multiphysics 6.0 to visualize bulk flow. The fluid was perfused at a fixed inlet pressure with the outlet set to atmospheric pressure. All walls were treated as rigid structures with a no-slip condition. The velocity profile and shear rate were obtained using Navier–Stokes modeling equations for incompressible creeping flow. The Carreau-Yasuda model was implemented which accounts for non-Newtonian behavior by expressing fluid viscosity as a function of shear rate, as shown in eqn (1). Where, η is viscosity, γ is shear rate, μ∞ is viscosity at infinite shear rate, μ0 is viscosity at zero shear rate, λ, a and b are material coefficients. The parameter values utilized were μ∞ = 0.0035 Pa s, μ0 = 0.16 Pa s, λ = 8.2 s, a = 1.23, and b = 0.64.37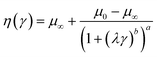 | (1) |
Statistical analysis
Statistical analyses were performed in GraphPad Prism Version 9.4.1 (GraphPad Software, Boston, MA). All results were reported as mean ± standard deviation (SD). The appropriate statistical tests were used, including the paired t test, unpaired t test, and Pearson's correlation analysis. The specific tests employed for each analysis were reported in the figure captions. A P-value of 0.05 was considered as the threshold for statistical significance.Results
Characterization of bulk velocity profile in the microfluidic capillary network
The microfluidic architecture employed a strategically staggered micropillar configuration to create complex flow patterns with spatially heterogeneous velocity magnitudes (Fig. 1B). The staggered micropillar configuration generated bifurcating and converging velocity streamlines analogous to flow patterns observed in in vivo capillary networks. This phenomenon was also observed in the cross-sectional 2D-plots of velocity magnitude where the velocity profile across the 4 μm-wide channel shows parabolic flow with maximum velocity at the channel centerline (Fig. 1C). However, in the downstream wide channel, the velocity profile had multiple velocity peaks centered at the inlets of the subsequent 4 μm-wide channels (Fig. 1D). Importantly, this microfluidic micropillar configuration enabled visualization of multiple RBCs flowing in parallel channels for high-throughput analysis of capillary transit velocity ultimately aiding in differentiation of fast and slow RBCs (Fig. 1E). Notably, the micropillar configuration included “open spaces” between capillaries to prevent channel blockage and flow perturbations by any occluding RBCs. This design may however result in flow phenomena different from those observed in vivo.Characterization of RBC shape, hydrodynamic adaptability, elongation index and capillary velocity
To investigate the dynamics of RBC flow and factors that affect capillary velocity, we recorded high resolution images of RBCs flowing through the capillary network at approximately 240 fps with an iPhone 13 mini using the slow-motion video settings. The iPhone was mounted on a microscope with a 63× oil-immersion objective and 10× eye piece. Example images from this setup are shown in Fig. 1E, 2A and G. Individual RBCs were manually tracked using Kinovea (version 0.9.5), an open-source video annotation tool commonly used for frame-by-frame video analysis and motion tracking.38–40 This tracking method enabled us to record the cell shape, assess hydrodynamic adaptability, calculate elongation index and determine the velocity of each individual RBC.The RBC shapes were categorized manually as croissant, slipper, oval, discocyte, spherocyte or other shapes as shown in the example images in Fig. 2A. RBCs frequently transitioned between croissant and slipper shapes as others have previously noted,29,41 prompting consolidation of these two categories in subsequent analysis. In HbAA samples, 80 ± 8% of RBCs were either croissant or slipper-shaped versus 22 ± 1% in HbSS samples. The HbSS samples also contained Oval, discocyte, and spherocyte RBC shapes that were not seen in the HbAA samples. These shapes were attributed to RBC membrane alterations due to HbS polymerization and cell dehydration typical in SCD. Both HbAA and HbSS samples contained a fraction of RBCs with other shapes that were too unique and variable to be classified into any one category. These “other” shapes were also more numerous in HbSS compared to HbAA (54 ± 2% versus 20 ± 8%, Fig. 2B). Similar shaped RBCs had comparable normalized velocities with no significant differences in both HbAA and HbSS samples. Croissant or slipper-shaped RBCs were however significantly faster than all other shapes while oval, discocyte and spherocyte-shaped RBCs, that were exclusive to SCD, also had the slowest velocities (Fig. 2C). The presence of RBCs with abnormal shapes therefore plays a major role in the impairment of microvascular rheology in SCD.
Hydrodynamic adaptability was defined as the cell's capacity to navigate through constrictions and maintain alignment with the primary flow direction while avoiding obstacle collisions. We hypothesized that RBCs would follow the streamlines in Fig. 1B and navigate through the micropillar constrictions without collisions. While this was an idealistic hypothesis, as healthy RBCs have been shown to collide with the endothelial walls at bifurcations exhibiting “lingering effects”,42 it allowed us to study how this phenomenon varied in healthy versus SCD RBCs. Our results showed that some RBCs deviated from the preferred trajectory and exhibited collision events (off-axis wall collisions) with the upstream faces of the micropillars perpendicular to the primary flow direction (Fig. 2D). This phenomenon is relevant to understanding RBC behavior in vivo, where cells encounter similar geometric constraints and flow disturbances at bifurcations in the microvasculature. We quantified RBC hydrodynamic adaptability by counting off-axis wall collision events as the RBCs traversed six consecutive micropillar constrictions along the flow direction. A collision event was defined as cell contact with the perpendicular face of a micropillar at a distance >2 μm from the edge of the intended flow path (Fig. 2D). The frequency of off-axis wall collisions in this region was significantly higher in HbSS versus HbAA (2.0 ± 1.4 versus 0.8 ± 0.9, Fig. 2E). A higher frequency of off-axis wall collisions was also significantly associated with reduced normalized velocity (P < 0.0001, Fig. 2F) as the colliding RBCs lost momentum and often dragged along the wall in a region with lower velocity magnitude compared to the midline.
Elongation index of the RBCs was measured in the wide channels before entering the narrow constriction. RBCs in this region were elongated due to a shear stress exerted by the bulk fluid and regulated by the pressure source at the inlet. The elongation index was calculated using the axis of the RBC in the direction of flow (a) and the axis perpendicular to the first axis (b) as shown in eqn (2) (Fig. 2G). This equation was previously shown as a robust measure of RBC deformability.43
 | (2) |
For each RBC the elongation index was measured at five distinct locations along its path and the measurements were averaged to reduce error. HbSS RBCs had a lower elongation index than HbAA RBCs (0.19 ± 0.09 vs. 0.28 ± 0.09, P < 0.0001) likely due to decreased deformability typical of HbSS RBCs (Fig. 2H). The relationship between RBC elongation index and normalized velocity was non-linear revealing a critical threshold effect where additive decreases in RBC deformability triggers severe microvascular rheology impairment. The effect of elongation on velocity progressively diminished at high elongation indices until reaching saturation at a maximum velocity. As shown in Fig. 2I, the steep portion of the curve was predominantly occupied by HbSS RBCs, reflecting their inability to achieve the critical deformation thresholds required for efficient capillary transit. This non-linear relationship between cellular rigidity and microvascular perfusion provides critical insight into how a small fraction of poorly deformable RBCs can precipitate the severe micro rheology defects typically observed in SCD patients.
Capillary velocity profiles in SCD versus heathy peripheral RBCs
We analyzed RBC velocities from 34 HbSS samples and 20 HbAA samples, automatically analyzing 1073 ± 534 RBCs per experiment. For each sample, velocity distributions were generated using probability density function analysis. RBCs in HbAA samples were more densely distributed at higher velocities while the velocity profile for HbSS samples was shifted left with a substantial proportion of RBCs flowing at low velocities (Fig. 3A). The combined distribution of all HbAA samples exhibited a modal normalized velocity of 0.83 and a standard deviation (SD) of 0.12. To establish a threshold for identifying slow RBCs, we subtracted one SD from this modal normalized velocity, yielding a cutoff value of 0.71. The slow RBC fraction was quantified from the AUC to represent the percentage RBC population below this threshold. HbSS individuals showed a higher fraction of slow RBCs and reduced mean normalized RBC velocity. They had a mean slow RBC fraction of 42.1 ± 12.0% significantly higher than 19.0 ± 4.9% for HbAA samples (Fig. 3B). Their mean normalized RBC velocity was also significantly lower than that of HbAA samples (0.70 ± 0.02 vs. 0.78 ± 0.03, p < 0.0001, Fig. 3C). Additionally, we evaluated assay reproducibility by measuring the slow RBC fraction of one HbAA and one HbSS sample across three different devices each. The HbAA and HbSS samples had coefficients of variation of 4.7% and 8.1% respectively (Fig. S3).To validate the effect of abnormal RBCs fractions on the velocity profile, HbSS RBCs were gradually added to HbAA RBCs to make 0%, 25%, 50%, 75% and 100% HbSS RBCs in the sample mixture. As the proportion of HbSS RBCs increased, there was a leftward shift in the velocity distribution graphs indicating a decrease in the population of RBCs flowing in the high velocity ranges and a corresponding increase in the population of RBCs flowing in the low velocity ranges (Fig. 3D). In five HbSS samples, each mixed with HbAA RBCs from a different healthy donor, we observed a significant increase in the slow RBC fraction (Fig. 3E) and a corresponding decrease in mean normalized RBC velocity (Fig. 3F) as the proportion of HbSS RBCs increased. We expect that the native HbSS samples contained a heterogenous mix of stiff and deformable RBCs as previously reported.44 However, these results indicate that the abnormal RBC fractions in SCD were responsible for the increased slow RBC fraction and the leftward shift in the velocity profile. Our assay was therefore sensitive to increasing proportions of abnormal HbSS RBCs within a mixed RBC population.
Clinical significance of the slow RBC fraction in SCD
To demonstrate the clinical and pathological significance of the slow RBC fraction in SCD, we analyzed its distribution across all samples including HbAA and HbSS to identify a cutoff threshold for elevated slow RBC fraction (SRF) within the participants of the study. We identified a bimodal distribution of the kernel density of the SRF and determined a classification threshold at the local minima of 32.2% (Fig. 4A). The threshold was utilized to stratify samples in the low SRF group or high SRF group with mean slow RBC fractions of 20.0 ± 5.3% and 46.3 ± 8.4% respectively (Fig. 4B). SCD subjects in the high SRF group exhibited significantly elevated hemolytic and inflammatory markers, including LDH, ARC, and CRP (Fig. 4C–E), indicating more severe disease activity.Further, we investigated whether the slow RBC fraction generated useful clinical predictions particularly examining the number of hospitalizations in the past 12 years, which is clinically meaningful and impacts quality of life of patients with SCD. Participants in the high SRF group experienced significantly more annual hospital admissions than those in the low SRF group (Fig. 4F). We then used regression modelling to identify the variables that best predict the frequency of hospitalization. In this dataset, only CRP and SRF had any association with the number of hospitalizations, with CRP being the most predictive (R squared = 0.168, versus 0.094 for SRF alone). However, the addition of SRF to the model contributed predictive value beyond CRP alone based on Akaike information criterion (AIC), with resultant R squared 0.227. This had the most impact on predicting hospitalizations for patients with low values of CRP, who may nevertheless have a higher number of hospitalizations with a high SRF (Fig. 4). From a biological context, this could potentially imply that the slow RBC fraction, reflecting poorer capillary transit dynamics, contributes a distinct risk than C-reactive protein, reflecting a systemic inflammatory state.
Effects of GBT021601 on RBC capillary velocity under hypoxia
To evaluate the sensitivity of the assay to hypoxia-induced RBC sickling, a gas chamber was setup as shown in Fig. 5A, to control the oxygen levels in the capillary network. Ten HbSS samples were either incubated with GBT021601 or buffer plus the carrier DMSO (untreated) and perfused through the device under normoxia or hypoxia. GBT021601 is a novel pharmacological agent that prevents RBC sickling by covalently binding to HbS hemoglobin, thereby increasing the hemoglobin's affinity for oxygen. GT021601 was shown to prevent HbS polymerization and significantly decrease RBC sickling in human and murine samples even under extreme hypoxia.45In untreated samples, the velocity distribution graph shifted leftward under hypoxia compared to normoxia due to an increase in stiff RBCs due to HbS polymerization. Conversely, the velocity distribution graph of GBT021601-treated RBCs under hypoxia was comparable to the normoxia graph (Fig. 5B). The slow RBC fraction was significantly higher under hypoxia than normoxia in the untreated samples (50.4 ± 12.5% vs. 28.8 ± 9.1%, p < 0.0001, Fig. 5C), while in GBT021601-treated samples, it remained low under hypoxia conditions (33.4% ± 11.4%, p = 0.25 vs. normoxia, Fig. 5C). Similarly, the mean normalized velocity of untreated RBCs decreased under hypoxia compared to normoxia but remained steady under both conditions in GBT021601-treated samples (Fig. 5D). The increase in slow RBC fraction under hypoxia (slow RBC fraction under hypoxia minus normoxia), was associated with the % HbS levels in individual samples (Fig. 5E). The SCD patients in this study were on transfusion therapy thus had a mixture of HbSS and HbAA RBCs from healthy donors. Patients who were recently transfused had a lower % HbS and a modest increase in slow RBC fraction under hypoxia. Two patients were on supportive therapy and were not recently transfused thus had % HbS levels greater than 70% and consequently the largest increase in slow RBC fraction under hypoxia (Fig. 5E). The fraction of occluding RBCs in untreated samples under hypoxia was 2.4% ± 4.0% (Fig. S4). Importantly, even non-occluding RBCs exhibited reduced capillary velocity, a major finding, often overlooked in prior studies that primarily focused on fully occluding cells while excluding those that passed through constrictions.32 Our assay captures these subtle yet physiologically relevant changes, reflecting the effect of transient hypoxia that circulating RBCs typically encounter while passing through capillaries. These results indicate that the assay was sensitive to the compromised capillary velocity caused by sickled RBCs. GBT021601 mitigated the impairment of capillary velocity under hypoxia suggesting that the assay could be used to assess the effects of novel SCD therapies on RBC rheology under varying oxygen tension.
Capillary velocity profiles of in vitro-derived RBCs
We assessed the velocity profiles of two in vitro-derived samples prepared with different methods. One sample comprised a heterogeneous mix of nucleated and enucleated cells, and the second sample was processed post erythroid differentiation to enrich enucleated RBCs. The samples were characterized using fluorescence microscopy and flow cytometry to visualize and distinguish the erythroid maturation stages. Cells were stained with fluorophore-conjugated antibodies against CD71, which is highly expressed on nucleated erythroid precursors but lost upon maturation, and absent on resting leucocytes, and CD235a, which is expressed on both nucleated and enucleated RBCs but absent on leucocytes.46 Both Imaging and flow cytometry confirmed the heterogeneity of the unprocessed sample, revealing leucocytes, nucleated RBCs, reticulocytes (CD71+ CD235a+), and terminally differentiated enucleated RBCs (CD712− CD235a+; Fig. 6A and S5). The microfluidic assay showed that heterogeneous sample exhibited a bimodal velocity distribution, with peaks at normalized velocities of 0.78 and 0.24, where the lower velocity component corresponded to the more rigid, nucleated cell fraction. In contrast, the processed sample with enucleated cells demonstrated a velocity profile concordant with a peripheral RBC sample (Fig. 6B). The slow RBC fraction in the processed sample was significantly higher than the peripheral RBC sample (32.9 ± 0.96% vs. 29.8 ± 0.70%, p = 0.01). The heterogeneous sample had the highest slow RBC fraction of 67.2 ± 6.4% (p = 0.001 vs. the processed sample, Fig. 6C). Given the anticipated differences in the biomechanical properties of in vitro-derived versus peripheral RBCs, we analyzed the absolute velocities and observed a pronounced leftward shift in the velocity distribution of enucleated in vitro-derived RBCs compared to peripheral RBCs (Fig. 6D and E), with mean absolute velocities of 0.79 ± 0.05 mm s−1vs. 0.93 ± 0.006 mm s−1 (p = 0.01). The heterogeneous sample exhibited an even lower mean velocity of 0.45 ± 0.05 mm s−1. These differences in absolute velocity reflect the distinct rheological characteristics of in vitro-derived versus peripheral RBCs. Our results demonstrate that processing methods to enrich enucleated RBCs improved the sample's velocity profile, generating RBCs with relative rheology characteristics more comparable to peripheral RBCs. This assay could thus be utilized to analyze in vitro-derived RBCs generated from various erythroid differentiation methods, from HSCs of various sources or carrying specific genetic modifications.Discussion
We analyzed several RBC rheological properties including capillary transit velocity, RBC shape, hydrodynamic adaptability and elongation index. Our results reveal that capillary transit velocity correlated with the RBC shape, hydrodynamic adaptability, and elongation index. While healthy RBCs have a biconcave (discocyte) shape at rest, they adapt their shape to flow conditions taking on the croissant or slipper shapes or a plethora of other shapes to minimize hydrodynamic resistance.41,47 We found that RBCs in SCD had unique shapes including oval, discocytes and spherocytes as well as a higher frequency of “other” shapes compared to healthy RBC samples. The presence of unique shapes exclusive to HbSS samples corroborates previous studies that have shown that HbS polymerization has permanent effects of RBC morphology.48,49 The abnormally shaped RBCs had markedly reduced capillary velocity suggesting that specific RBC shapes under flow may be linked to impaired vascular rheology in SCD. HbSS RBCs also had lower elongation index and a higher frequency of off-axis wall collisions compared to healthy RBCs. This suggests that RBCs in SCD have impaired hydrodynamic adaptability thus cannot efficiently navigate the microcapillary network. Unlike healthy RBCs that can deform and elongate to navigate through confined spaces, stiff cells in SCD cannot change their shape to accommodate the vessel geometry and effectively follow the streamlines. Other studies have also found increased margination in SCD as the sickled RBCs tend to concentrate near vessel walls,50 which may play a role in increasing off-axis wall collisions. These aberrant RBCs make direct contact with the capillary walls, exerting abnormal mechanical stress on the endothelial walls during passage.51 This mechanical stress not only impedes blood flow but may also contribute to endothelial damage and inflammation which further exacerbates SCD pathology. The slow RBC fraction therefore represents the proportion of RBCs with increased hydrodynamic resistance and increased interaction with the endothelium lining the capillary walls which may pose a higher risk of contributing to vascular pathology via endothelial activation.52Importantly, we developed a method for rapid, high throughput assessment of capillary transit velocity in which we generate a velocity profile to represent the entire RBC population from approximately 1000 cells analyzed with single cell resolution, from a 30 seconds video. The RBC capillary transit velocity represents the functional ability of RBCs to navigate narrow capillaries at healthy velocities. We also measured the slow RBC fraction to quantify the proportion of the cell population with impaired capillary transit dynamics. Capillary transit time is particularly relevant in SCD pathology as HbS polymerization has a characteristic delay period before the polymers start to form.53 HbS-containing RBCs can transit through hypoxic capillaries and return to the lungs for reoxygenation faster than the delay time, thereby avoiding the formation of the HbS polymers. RBCs with longer transit times (the slow RBC fraction) therefore have prolonged exposure to the hypoxic environment in the capillaries resulting in a higher probability of sickling and contributing to VOEs. Notably, this study utilized samples at low levels of hematocrit (0.5%) thus had minimal cell-to-cell interactions. At physiological hematocrit (∼25% in SCD54), RBCs with impaired velocity could cause traffic congestion in the capillaries, curtailing flux to increase the capillary transit time of relatively healthy, unaffected cells. By curtailing total flux in microcirculation, a small fraction of slow RBCs could initiate a viscous cycle that accelerates polymer formation in a larger fraction of HbS-containing RBCs resulting in severe VOEs. It is therefore critical that new therapies minimize the slow RBC fraction. Our assay may be utilized to detect the remaining small populations of RBCs that may have significant contributions to microcirculation defects including VOEs.
The clinical relevance of this assay is underscored by the significant association between the slow RBC fraction and conventional biomarkers of SCD severity. HbS polymer-containing RBCs are dramatically stiffer, and a greater proportion of these stiffened cells lead to increased hemolysis and elevated inflammatory response reflected by elevated LDH, ARC and CRP levels. Furthermore, a higher fraction of stiff RBCs in SCD are more likely to impede blood flow in the microcapillaries, leading to VOEs, the primary cause of hospitalization in SCD patients.5 Indeed, our finding showed that the high slow RBC fraction was associated with a higher frequency of annual hospital admissions. Importantly, the slow RBC fraction improved the prediction value of a model for identifying patients with a historically high frequency of hospitalizations. Future work may assess the utility of the slow RBC fraction alongside other biochemical markers in the prediction of SCD clinical outcomes including the potency of novel treatments, VOCs, and trends in overall disease morbidity.
We utilized mixtures of healthy and HbSS RBCs to simulate heterogeneous treatment effects and validate the assay's sensitivity to small factions of abnormal RBCs in a mixed sample. This study revealed a shift in the velocity profile toward low velocities and an increase in the slow RBC fraction with increasing proportion of diseased HbSS RBCs in a healthy sample. This demonstrates that the assay is sensitive to abnormal subpopulations that may be present following therapeutic intervention. Previous studies exploring the effects of hypoxia on RBC rheology induced high levels of capillary occlusion by using methods that differ from in vivo conditions, such as pre-exposing samples to hypoxia before they enter the microchannels or perfusing them at low velocities of approximately 0.1 mm s−1.32,55,56 While these approaches increase sensitivity to HbS-containing cells, they may obscure rheological characteristics that occur in vivo. In contrast, our assay more closely replicates transient hypoxia conditions in the physiological environment. We exposed cells to hypoxia while they flowed through the microchannels at absolute velocities of 0.75 ± 0.06 mm s−1 (for HbSS samples, Fig. S2), which falls within the typical capillary velocity range of 0.5–1.5 mm s−1.57 In this design, the cells traversed the entire chip in under 30 seconds. Our results demonstrate that RBC capillary velocity is significantly compromised even under these transient hypoxic conditions. Importantly, the assay was sensitive to the effects of GBT02601 under hypoxia, demonstrating potential for assessing the effectiveness of similar drugs in preventing hypoxia-induced RBC sickling. Future studies may compare the effectiveness of various therapies in reducing the slow RBC fraction under various levels of oxygen tension, for both ex vivo and in vivo drug administration.
This assay detected heterogeneity in in vitro-derived RBCs showing distinct velocity profiles for nucleated and enucleated cells. These results suggest that assay may be used for quality control in biomanufacturing applications to assess variability in rheological properties of the RBC products from various erythroid differentiation techniques. In recent studies, several groups have differentiated RBCs in vitro from genetically modified HSCs to study the potency of various SCD gene therapies in expressing therapeutic hemoglobin variants and improving RBC deformability.10,58 Although in vitro-derived RBCs may differ morphologically from peripheral RBCs, recent advances in erythroid differentiation are narrowing the gap between these cell types. Several groups are advancing differentiation protocols to increase the degree of maturation to fully enucleated, biconcave erythrocytes.59,60 For example, Claessen et al. showed that supplementing differentiation cultures with 5.0 mg dL−1 of free cholesterol produced in vitro-derived RBCs that were more deformable than a reference reticulocyte sample.61 These advances support the use of in vitro-derived RBCs to investigate the therapeutic effect of SCD gene therapies on multiple RBC functions, including rheology, with comparison to peripheral RBCs. Our assay complements this work by providing a method assessing the effect of next generation SCD therapies on the rheological properties of in vitro-derived RBCs. Future work may validate this assay's sensitivity to various degrees of gene editing efficiency and the applicability of the assay to screen multiple gene therapy modalities to predict the most potent candidates with the highest probability of success in animal and human studies.
In conclusion, we present a platform for assessment of capillary transit velocity in SCD and in vitro-derived RBCs. The assay measures the capillary transit velocity of individual cells flowing through a microfluidic capillary network and generates a profile to characterize all RBCs in the sample. We validated the assay using peripheral RBCs from SCD and healthy donors and showed that the capillary transit velocity correlated with the cell shape, hydrodynamic adaptability and elongation index. We established the slow RBC fraction as a potential biomarker that correlates with key biomarkers for SCD severity. Future studies will utilize this assay to evaluate the rheological properties of in vitro-derived RBCs generated from HSCs treated with gene therapies designed to modify RBC mechanics.
Ethics statement
All experiments were performed in accordance with the Guidelines of the United States Food and Drug Administration (US FDA) and Declaration of Helsinki and approved by the ethics committee at the University Hospitals Cleveland Medical Center (UHCMC IRB# 05-14-07C). Informed consents were obtained from human participants of this study.Author contributions
SO: conceptualization, methodology, investigation, data analysis, visualization, software development, writing – original draft. UG: methodology, validation, data analysis, resources, visualization, investigation, writing – review & editing. YD: conceptualization, validation, data analysis, visualization, investigation, writing – review & editing. CLW: conceptualization, methodology, software development, writing – review & editing. ZS: methodology, data analysis, investigation, writing – review & editing. BB: methodology, data analysis, writing – review & editing. PF: investigation, validation, data analysis, writing – review & editing. YBL: investigation, validation, writing – review & editing. NMP: methodology, investigation. PG: methodology, funding acquisition, resources, supervision. DNN: methodology, funding acquisition, resources, supervision. MAS: conceptualization, methodology, funding acquisition, supervision, project administration, writing – review & editing. PM: conceptualization, methodology, funding acquisition, supervision, project administration, writing – review & editing. UAG: conceptualization, methodology, resources, funding acquisition, supervision, project administration, writing – review & editing.Conflicts of interest
UAG and Case Western Reserve University have financial interests in Hemex Health Inc. UAG and Case Western Reserve University have financial interests in BioChip Labs Inc. UAG, PM, MAS and Case Western Reserve University have financial interests in Xatek Inc. UAG has financial interests in DxNow Inc. PM has financial interests in Haima Therapeutics LLC. Financial interests include licensed intellectual property, stock ownership, research funding, employment, and consulting. Hemex Health Inc. offers point-of-care diagnostics for hemoglobin disorders, anemia, and malaria. BioChip Labs Inc. offers commercial clinical microfluidic biomarker assays for inherited or acquired blood disorders. Xatek Inc. offers Point-of-care global assays to evaluate the hemostatic process. DxNow Inc. offers microfluidic and bio-imaging technologies for in vitro fertilization, forensics, and diagnostics. The competing interests of Case Western Reserve University employees are overseen and managed by the Conflict of Interests Committee according to a Conflict-of-Interest Management Plan.Data availability
The corresponding author of this publication will fulfill all reasonable requests for materials and data.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5lc00769k.
Acknowledgements
The authors acknowledge the following funding sources: National Heart, Lung, and Blood Institute under Awards: R56HL165946, R42HL162214, R41HL172662, and R44HL140739; National Institutes of Health Office of the Director under Award No. U01AI176469; National Institute of Diabetes and Digestive and Kidney Diseases Small Business Innovation Research Program grant R42DK119048; NIH – Fogarty International Center grant No. D43TW012260. U. A. G. acknowledges support by the Clinical and Translational Science Collaborative of Northern Ohio which is funded by the National Center for Advancing Translational Sciences (NCATS) of the National Institutes of Health, UM1TR004528. Z. S. acknowledges the American Society of Hematology Graduate Hematology Award. This article's contents are solely the authors' responsibility and do not necessarily represent the official views of the National Institutes of Health.References
- A. M. Thomson, T. A. McHugh, A. P. Oron, C. Teply, N. Lonberg, V. V. Tella, L. B. Wilner, K. Fuller, H. Hagins and R. G. Aboagye, Lancet Haematol., 2023, 10, 585–599 CrossRef PubMed
.
- G. J. Kato, F. B. Piel, C. D. Reid, M. H. Gaston, K. Ohene-Frempong, L. Krishnamurti, W. R. Smith, J. A. Panepinto, D. J. Weatherall and F. F. Costa, Nat. Rev. Dis. Primers, 2018, 4, 1–22 CrossRef PubMed
.
- P. Sundd, M. T. Gladwin and E. M. Novelli, Annu. Rev. Pathol., 2019, 14, 263–292 CrossRef CAS PubMed
.
- T. Jang, M. Poplawska, E. Cimpeanu, G. Mo, D. Dutta and S. H. Lim, J. Transl. Med., 2021, 19, 397 Search PubMed
.
- N. Shah, M. Bhor, L. Xie, J. Paulose and H. Yuce, PLoS One, 2019, 14, e0214355 CrossRef CAS PubMed
.
- S. K. Ballas, Clin. Hemorheol. Microcirc., 2018, 68, 105–128 Search PubMed
.
- E. Vichinsky, Hematology Am. Soc. Hematol. Educ. Program, 2017, 2017, 435–439 CrossRef PubMed
.
- A. A. Abraham and J. F. Tisdale, Blood, 2021, 138, 932–941 CrossRef CAS PubMed
.
- D. C. De Souza, N. Hebert, E. B. Esrick, M. F. Ciuculescu, N. M. Archer, M. Armant, É. Audureau, C. Brendel, G. Di Caprio, F. Galactéros, D. Liu, A. McCabe, E. Morris, E. Schonbrun, D. Williams, D. K. Wood, D. A. Williams, P. Bartolucci and J. M. Higgins, Nat. Commun., 2023, 14, 5850 CrossRef CAS PubMed
.
- G. A. Newby, J. S. Yen, K. J. Woodard, T. Mayuranathan, C. R. Lazzarotto, Y. Li, H. Sheppard-Tillman, S. N. Porter, Y. Yao, K. Mayberry, K. A. Everette, Y. Jang, C. J. Podracky, E. Thaman, C. Lechauve, A. Sharma, J. M. Henderson, M. F. Richter, K. T. Zhao, S. M. Miller, T. Wang, L. W. Koblan, A. P. McCaffrey, J. F. Tisdale, T. A. Kalfa, S. M. Pruett-Miller, S. Q. Tsai, M. J. Weiss and D. R. Liu, Nature, 2021, 595, 295–302 CrossRef CAS PubMed
.
- S. Pellegrin, C. E. Severn and A. M. Toye, Haematologica, 2021, 106, 2304 CrossRef CAS PubMed
.
- T. Mayuranathan, G. A. Newby, R. Feng, Y. Yao, K. D. Mayberry, C. R. Lazzarotto, Y. Li, R. M. Levine, N. Nimmagadda, E. Dempsey, G. Kang, S. N. Porter, P. A. Doerfler, J. Zhang, Y. Jang, J. Chen, H. W. Bell, M. Crossley, S. V. Bhoopalan, A. Sharma, J. F. Tisdale, S. M. Pruett-Miller, Y. Cheng, S. Q. Tsai, D. R. Liu, M. J. Weiss and J. S. Yen, Nat. Genet., 2023, 55, 1210–1220 CrossRef CAS PubMed
.
- J.-Y. Métais, P. A. Doerfler, T. Mayuranathan, D. E. Bauer, S. C. Fowler, M. M. Hsieh, V. Katta, S. Keriwala, C. R. Lazzarotto and K. Luk, Blood Adv., 2019, 3, 3379–3392 CrossRef PubMed
.
- S. H. Park, C. M. Lee, D. P. Dever, T. H. Davis, J. Camarena, W. Srifa, Y. Zhang, A. Paikari, A. K. Chang, M. H. Porteus, V. A. Sheehan and G. Bao, Nucleic Acids Res., 2019, 47, 7955–7972 Search PubMed
.
- C. Antoniani, V. Meneghini, A. Lattanzi, T. Felix, O. Romano, E. Magrin, L. Weber, G. Pavani, S. El Hoss and R. Kurita, Blood, 2018, 131, 1960–1973 CrossRef CAS PubMed
.
- S. Shah, X. Huang and L. Cheng, Stem Cells Transl. Med., 2014, 3, 346–355 CrossRef CAS PubMed
.
- N. Hebert, M. G. Rakotoson, G. Bodivit, E. Audureau, L. Bencheikh, L. Kiger, N. Oubaya, S. Pakdaman, M. Sakka and G. Di Liberto, Am. J. Hematol., 2020, 95, 1235–1245 CrossRef CAS PubMed
.
- M. de la Cueva Tamanaha, E. F. Cabrera, J. Sargeant, P. D. Gershon, P. P. S. Russell and M. J. Cocco, bioRxiv, 2025, preprint, DOI:10.1101/2025.05.12.653546.
- V. Sergunova, S. Leesment, A. Kozlov, V. Inozemtsev, P. Platitsina, S. Lyapunova, A. Onufrievich, V. Polyakov and E. Sherstyukova, Sensors, 2022, 22, 2055 CrossRef CAS PubMed
.
- M. Dao, T. L. Chwee and S. Subra, J. Mech. Phys. Solids, 2003, 51, 2259–2280 CrossRef
.
- M. J. Oh, F. Kuhr, F. Byfield and I. Levitan, J. Visualized Exp., 2012, 1, 3886 Search PubMed
.
- A. Sadaf, K. G. Seu, E. Thaman, R. Fessler, D. G. Konstantinidis, H. A. Bonar, J. Korpik, R. E. Ware, P. T. McGann, C. T. Quinn and T. A. Kalfa, Front. Physiol., 2021, 12, 636609 CrossRef PubMed
.
- M. Gutierrez, M. Shamoun, K. G. Seu, T. Tanski, T. A. Kalfa and O. Eniola-Adefeso, Sci. Rep., 2021, 11, 7909 CrossRef CAS PubMed
.
- D. C. Williams and D. K. Wood, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2313755120 CrossRef CAS PubMed
.
- P. Guruprasad, R. G. Mannino, C. Caruso, H. Zhang, C. D. Josephson, J. D. Roback and W. A. Lam, Am. J. Hematol., 2019, 94, 189–199 CrossRef CAS PubMed
.
- D. W. Lee, I. Doh, F. A. Kuypers and Y. H. Cho, Biomed. Microdevices, 2015, 17, 102 CrossRef PubMed
.
- M. S. Bazzi, J. M. Valdez, V. H. Barocas and D. K. Wood, Biophys. J., 2020, 119, 2307–2315 CrossRef CAS PubMed
.
- S. M. Recktenwald, M. G. Lopes, S. Peter, S. Hof, G. Simionato, K. Peikert, A. Hermann, A. Danek, K. van Bentum and H. Eichler, Front. Physiol., 2022, 13, 884690 CrossRef PubMed
.
- F. Reichel, J. Mauer, A. A. Nawaz, G. Gompper, J. Guck and D. A. Fedosov, Biophys. J., 2019, 117, 14–24 CrossRef CAS PubMed
.
- K. L. Pitts and M. Fenech, J. Visualized Exp., 2013, 74, e50314 Search PubMed
.
- S. Oshabaheebwa, C. A. Delianides, A. A. Patwardhan, E. N. Evans, Z. Sekyonda, A. Bode, F. M. Apio, C. K. Mutuluuza, V. A. Sheehan and M. A. Suster, Biosens. Bioelectron., 2024, 258, 116352 CrossRef CAS PubMed
.
- Y. Man, E. Kucukal, R. An, Q. D. Watson, J. Bosch, P. A. Zimmerman, J. A. Little and U. A. Gurkan, Lab Chip, 2020, 20, 2086–2099 RSC
.
- J. Schindelin, I. Arganda-Carreras, E. Frise, V. Kaynig, M. Longair, T. Pietzsch, S. Preibisch, C. Rueden, S. Saalfeld, B. Schmid, J. Y. Tinevez, D. J. White, V. Hartenstein, K. Eliceiri, P. Tomancak and A. Cardona, Nat. Methods, 2012, 9, 676–682 CrossRef CAS PubMed
.
- E. M. Furst, Particle tracking with Matlab, University of Delaware, Newark, 2015 Search PubMed.
- M. W. Issa, H. Yu, M. C. Roffin, S. V. Barancyk, R. M. Rock, J. F. Gilchrist and C. L. Wirth, Langmuir, 2022, 38, 11581–11589 CrossRef CAS PubMed
.
- J. C. Crocker and D. G. Grier, J. Colloid Interface Sci., 1996, 179, 298–310 CrossRef CAS
.
- F. Abraham, M. Behr and M. Heinkenschloss, Comput. Methods Biomech. Biomed. Eng., 2005, 8, 127–137 CrossRef PubMed
.
- A. Puig-Diví, C. Escalona-Marfil, J. M. Padullés-Riu, A. Busquets, X. Padullés-Chando and D. Marcos-Ruiz, PLoS One, 2019, 14, e0216448 CrossRef PubMed
.
- N. Shishov, K. Elabd, V. Komisar, H. Chong and S. N. Robinovitch, PLoS One, 2021, 16, e0258923 CrossRef CAS PubMed
.
- B. Pueo, A. Penichet-Tomas and J. M. Jimenez-Olmedo, Physiol. Behav., 2020, 227, 113144 CrossRef CAS PubMed
.
- S. M. Recktenwald, K. Graessel, F. M. Maurer, T. John, S. Gekle and C. Wagner, Biophys. J., 2022, 121, 23–36 CrossRef CAS PubMed
.
- Y. Rashidi, G. Simionato, Q. Zhou, T. John, A. Kihm, M. Bendaoud, T. Krüger, M. O. Bernabeu, L. Kaestner, M. W. Laschke, M. D. Menger, C. Wagner and A. Darras, Biophys. J., 2023, 122, 1526–1537 CrossRef CAS PubMed
.
- A. Saadat, D. A. Huyke, D. I. Oyarzun, P. V. Escobar, I. H. Øvreeide, E. S. Shaqfeh and J. G. Santiago, Lab Chip, 2020, 20, 2927–2936 RSC
.
- A. Mozzarelli, J. Hofrichter and W. A. Eaton, Science, 1987, 237, 500–506 CrossRef CAS PubMed
.
- K. Dufu, C. Alt, S. Strutt, T. Tang, H. Liao-Zou, Y. Yuan, B. E. Cathers and D. Oksenberg, Blood, 2020, 136, 7–8 CrossRef
.
- K. E. McGrath, Front. Hematol., 2024, 3, 1496916 CrossRef
.
- M. Nouaman, A. Darras, C. Wagner and S. M. Recktenwald, Biomicrofluidics, 2024, 18, 024104 CrossRef CAS PubMed
.
- F. A. Kuypers, Hematol./Oncol. Clin. North Am., 2014, 28, 155–179 CrossRef PubMed
.
- C. D'Costa, O. Sharma, R. Manna, M. Singh, S. Singh, S. Singh, A. Mahto, P. Govil, S. Satti, N. Mehendale, Y. Italia and D. Paul, Bioeng. Transl. Med., 2024, 9, e10643 CrossRef PubMed
.
- X. Cheng, C. Caruso, W. A. Lam and M. D. Graham, Sci. Adv., 2023, 9, eadj6423 CrossRef PubMed
.
- K. Katoh, Int. J. Mol. Sci., 2023, 24, 16518 CrossRef PubMed
.
- C. Caruso, X. Cheng, M. E. Michaud, H. M. Szafraniec, B. E. Thomas, M. E. Fay, R. G. Mannino, X. Zhang, Y. Sakurai and W. Li, Blood, 2024, 144, 2050–2062 CrossRef CAS PubMed
.
- J. Hofrichter, P. D. Ross and W. A. Eaton, Proc. Natl. Acad. Sci. U. S. A., 1974, 71, 4864–4868 CrossRef CAS PubMed
.
- S. G. Ahmed, U. A. Ibrahim and A. W. Hassan, Ann. Saudi Med., 2006, 26, 439–443 CrossRef PubMed
.
- E. Du, M. Diez-Silva, G. J. Kato, M. Dao and S. Suresh, Proc. Natl. Acad. Sci. U. S. A., 2015, 112, 1422–1427 CrossRef CAS PubMed
.
- Y. Qiang, A. Sissoko, Z. L. Liu, T. Dong, F. Zheng, F. Kong, J. M. Higgins, G. E. Karniadakis, P. A. Buffet and S. Suresh, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2217607120 CrossRef CAS PubMed
.
- A. G. Hudetz, Microcirculation, 1997, 4, 233–252 CrossRef CAS PubMed
.
- G. L. S. Martins, C. K. V. Nonaka, E. A. Rossi, A. V. R. de Lima, C. S. A. Adanho, M. S. Oliveira, S. C. M. A. Yahouedehou, C. L. e. M. de Souza, M. D. S. Gonçalves and B. D. Paredes, Cells, 2023, 12, 1121 CrossRef CAS PubMed
.
- S. Heshusius, E. Heideveld, P. Burger, M. Thiel-Valkhof, E. Sellink, E. Varga, E. Ovchynnikova, A. Visser, J. H. Martens and M. von Lindern, Blood Adv., 2019, 3, 3337–3350 CrossRef PubMed
.
- Y. K. Cho, H.-K. Kim, S. S. Kwon, S.-H. Jeon, J.-W. Cheong, K. T. Nam, H.-S. Kim, S. Kim and H. O. Kim, Stem Cell Res. Ther., 2023, 14, 106 CrossRef CAS PubMed
.
- M. J. A. G. Claessen, N. Yagci, K. Fu, E. Brandsma, M. J. Kersten, M. von Lindern and E. van den Akker, Sci. Rep., 2024, 14, 15592 CrossRef CAS PubMed
.
| This journal is © The Royal Society of Chemistry 2026 |






