Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Real-time high-throughput characterisation of the surface elasticity of suspended cells
Ziyu
Guo
,
Yi
Sui
 * and
Wen
Wang
* and
Wen
Wang
School of Engineering and Materials Science, Queen Mary University of London, London, E1 4NS, UK. E-mail: y.sui@qmul.ac.uk
First published on 20th January 2026
Abstract
The intrinsic elasticity of the cell membrane–cortex complex, i.e., the cell surface, is a promising biomarker for cell status and disease and has widespread biological and biomedical applications. However, measuring cell surface elasticity in real time with high throughput has not been achieved so far. Here we develop a system and demonstrate that it can characterise the intrinsic surface elasticity of up to 411 cells per second, with a low latency of less than 1 millisecond per cell from an image to predicted elasticity. Our key innovation is to integrate a multi-layer perception (MLP) based machine learning algorithm, which infers the surface elasticity of cells from their camera-recorded steady-deformation profiles in a microchannel, with a high-fidelity mechanistic model, which resolves the cell surface, cytoplasm and nucleus and can accurately predict the flow-induced cell deformation. Applied to human prostate cancer PC-3 and leukaemia K-562 cell lines, the system enables measuring tens of thousands of cells within minutes to explore the cell mechano-heterogeneity, the relation between surface elasticity and cell size, and the possibility of using surface elasticity and cell size for cell classification. We show that the measured cell surface elasticity is little affected by flow conditions when doubling the flow speed or suspension fluid viscosity. The system is also sensitive enough to detect a reduction of cell surface elasticity as a result of the cytochalasin D-induced actin disassembly. By enabling real-time high-throughput characterisation of the surface elasticity of cells, the present method may inspire new applications.
1 Introduction
The cell membrane–cortex complex (MCC), i.e., the cell surface, is composed of the cell plasma membrane and the underlying cortical cytoskeleton. It is one of the most important structures of cells which provides the cell's mechanical strength, regulates the cross-membrane mass transport, and interacts with the extracellular environment.1–3 The mechanical properties of the cell surface, such as elasticity and viscosity, influence a wide range of biological processes of cells, including differentiation, growth, migration, endocytosis and exocytosis.2,4–6 They are also promising biomarkers for the diagnosis and prognosis of diseases, such as sickle cell anaemia, malaria, and cancers.7–11Measuring the mechanical properties of the cell surface is extremely time-consuming and labour-intensive due to the cells' small size, fragility, and often the huge quantity (e.g. thousands to millions) required for blood-related applications. Classical characterisation methods, such as atomic force microscopy,12,13 micropipette aspiration,14–16 or magnetic twisting cytometry,17–19 often measure cell deformation under a well-defined force and then fit the deformation–force relation to a theoretical model to inversely infer the cell mechanical properties. Due to the lack of automation, those methods often suffer from low throughput rates that are limited to 10–1000 cells per hour.20
To address the need for faster measurement, in the recent decade, microfluidic-based deformability cytometry (DC) has emerged as a transformative technology for high-throughput cell mechanical phenotyping.21–35 DC technologies can flow thousands of suspended cells through a microfluidic channel in a second and deform the cells with precisely controlled fluid forces. The state-of-the-art systems can measure, in real time, the deformation parameters of the cells by integrating high-speed imaging, data streaming and online image processing.25,34 However, the intrinsic mechanical properties of the cells still need to be obtained through post-processing of the experimental data,36 as the inverse-fitting process to infer mechanical properties from cell deformation is very time-consuming. It should be noted that as a mechanical biomarker, intrinsic mechanical properties of cells have significant advantages over cell deformation parameters such as the length-to-width ratio and non-circularity. Cell deformation parameters strongly depend on the DC flow geometry and operational flow conditions such as the suspension viscosity and flow speed, leading to difficulties in comparing cell deformability measured by different DC systems.
In addition, current DC systems consider cells as a homogeneous object and measure the deformability of the whole cell. Characterising the mechanical properties of the cell surface has not been achieved so far. A major problem is the lack of a high-fidelity cell mechanical model, which takes into account the subcellular structures including the cell surface and can accurately predict the flow-induced deformation of suspended cells in microchannels.
In recent years, machine learning, in particular artificial neural networks (ANNs), has rapidly emerged as a powerful tool in a wide range of engineering applications, including data-driven reconstruction of flow fields,37,38 inverse design of material structures for targeted mechanical performance,39,40 and prediction of cell or tissue mechanics from images.41–43 In DC systems, ANNs have been employed to analyse cell images or deformation features for cell classification and sorting.44–46 However, real-time inference of intrinsic cell mechanical properties has yet to be achieved for the challenges mentioned above.
To address those challenges, recently we developed a three-dimensional (3D) computational model to simulate the transient flow-induced deformation of suspended cancer cells in microchannels of different geometries.47–49 The model considers cells as a three-layer compound structure, including a viscoelastic surface representing the MCC, a viscous cytoplasm, and a nucleus modelled as a smaller stiffer microcapsule. The computational approach can accurately recover the transient deformation of human leukaemia (e.g., HL-60, K-562) and prostate cancer PC-3 cells47,49 in shear and extensional flows, providing evidence that it could serve as a general modelling framework for cancer and more broadly eukaryotic cells in suspensions. Although simpler mechanical models for suspended cells exist, for example, those considering cells as liquid droplets50,51 or deformable microcapsules,52–54 the models often cannot quantitatively predict the flow-induced cell deformation profiles or consider the effect of the cell nucleus. For inverse inference of cell mechanical properties from cell deformation, it is crucial to fit the experimentally measured cell profile to a quantitative cell mechanical model.
We also developed a number of machine learning approaches based on a deep convolutional neural network (DCNN),55,56 a DCNN coupled with a long short-term memory network (LSTM),57 and a multi-layer perceptron (MLP)54 to inversely infer, in real time, the intrinsic surface elasticity of bioartificial microcapsules through their flow-induced deformation in microchannels. Those microcapsules could serve as a qualitative mechanical model for suspended cells.
In this study, we build a system integrating our recent developments in high-fidelity cell mechanical modelling,47,49 machine learning-based real-time mechanical characterisation,54 with an established microchannel platform by Guck's group25 to generate high-throughput flow-induced cell deformation. We demonstrate that the system can characterise the intrinsic surface elasticity of cells at a throughput rate of up to 411 cells per second, with an image-to-surface-elasticity latency of less than 1 millisecond on a personal computer without using parallel computing. Measuring tens of thousands of leukaemia K-562 and prostate cancer PC-3 cells, our results indicate a high level of mechanical heterogeneity of both cell lines, where the surface elasticity spans more than two orders of magnitude, considerably greater than previous studies that measured tens to hundreds of cells. The surface elasticity of K-562 cells increases with cell size, while the opposite trend is observed for PC-3 cells. We also find that the measured results are little affected by the flow condition when doubling the flow speed or the suspension fluid viscosity. In addition, the present system can detect a reduction of the cell surface elasticity, following disassembly of cell actin filaments using cytochalasin D (CytoD).
2 Results and discussion
2.1 Design of the system
Our system consists of four major modules, as illustrated in Fig. 1. The first module is a microfluidic platform to flow through and deform hundreds of suspended cells per second under precisely controlled flow conditions. The microchannel geometry follows the classical design of Otto et al.,25 is illustrated in Fig. 8, and is detailed in section 4.1 of Materials and methods. Human prostate cancer PC-3 and leukaemia K-562 cells are suspended in phosphate-buffered saline (PBS, Sigma-Aldrich) added with 2–4% (w/w) methylcellulose (15 cP, Sigma-Aldrich) to a concentration of around 107 cells per ml. At room temperature, the cell suspension is driven through a straight microchannel that has a square cross section with a width of 20 μm, at an average speed of 7–14 cm s−1, by syringe pumps (PHD ULTRA, Harvard Apparatus). Hydrodynamic focusing using sheath flow is employed to centre the cells horizontally, and wall lift forces acting on the deformable cells help to position them vertically to the channel centreline. Shortly after entering the straight channel, cells deform into steady bullet shapes under the effect of fluid shear. The cell steady profile is captured by a high-speed streaming camera (exo174MU3, SVS-Vistek), which is the second module of the system. The camera feeds the images in real time to a personal computer for online prediction of the intrinsic cell surface elasticity. This challenging prediction task is conducted by an MLP-based machine learning algorithm, the third module and the core of the present system.Details of the MLP-based prediction algorithm are covered in section 4.4 in Materials and methods, and here we only provide a brief introduction. The MLP is one of the most widely used neural networks in supervised learning.54,58 It is a fully connected feed-forward network, which means a neuron of a layer is connected to all neurons of the next layer, and the information only flows from one layer to the next. Due to the MLP's simplicity, the universal approximation property for nonlinear relations between data and their corresponding measurements, as well as the feature that its training is conducted offline and its prediction process only involves a limited number of algebraic calculations, the MLP is a promising tool for fast inverse analysis of cell mechanical properties.
The fourth module of the system is a high-fidelity mechanistic model which can simulate the flow-induced deformation of cells in a microchannel. The model generates labelled data to train the MLP algorithm. We consider the deformation of biological cells with a wide range of size and surface elasticity in the straight square channel in Fig. 8. The cell is modelled as a three-layer viscoelastic structure consisting of the membrane–cortex complex, cytoplasm and cell nucleus (detailed in section 4.5). The cell deformation and its interaction with the surrounding fluid is solved with a well-tested immersed-boundary lattice Boltzmann method (see section 4.6 for details). The steady cell deformation profiles, together with the corresponding cell parameters, are used to train the MLP-based prediction algorithm.
2.2 Performance of the cell mechanical model
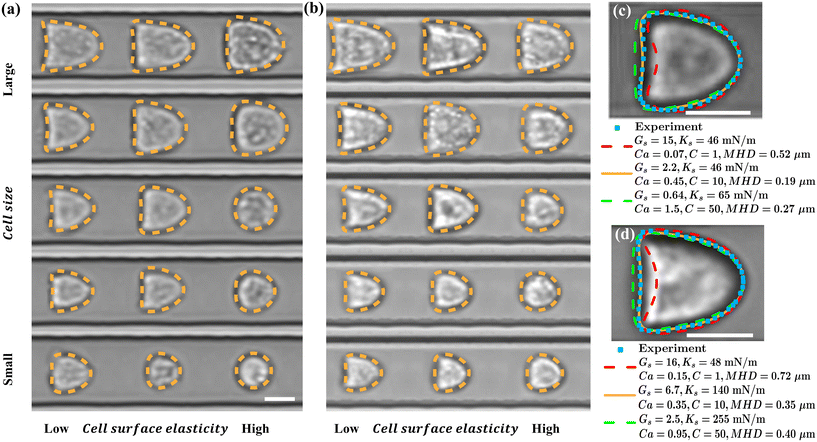
The present system uses an MLP-based algorithm trained with simulation data generated by a cell mechanical model to infer the surface elasticity of cells from their steady deformation in a microchannel. Therefore, the capability of the cell mechanical model to accurately predict the flow-induced cell deformation is essential to the accuracy of the system. We conduct extensive comparisons of the cell steady profiles obtained from the present numerical simulations and experiments for both K-562 and PC-3 cells and have two main observations.Firstly, we find that the cell mechanical model can accurately recover the steady deformation of both cell lines for cells in a wide range of sizes and surface elasticity. Some examples for the K-562 and PC-3 cells are shown in Fig. 2a and b, respectively. Our simulation setup, as detailed in section 4.5, matches the experimental conditions that are described in section 4.1. In brief, the straight square part of the microchannel has a cross-sectional dimension of W = H = 20 μm. No-slip boundary conditions are applied at the channel walls. The suspension medium in the simulations follow the same rheological properties of the PBS with methylcellulose used in the experiments (see section 4.1), and the average flow speed is also identical to that of the experiments (i.e., U = 7 cm s−1). As discussed in section 4.1, the steady cell deformation is mainly determined by the capillary number Ca = μU/Gs, which measures the relative importance of the fluid shear and cell elastic forces as well as the confinement ratio β = 2a/W, which compares the cell diameter with the channel dimension.
For each cell shown in Fig. 2a and b, the cell size, and thus the confinement ratio, in the simulation is extracted directly from the corresponding experimental image by matching the footprint area. For the cell surface elasticity, we have set the surface hardness parameter at C = 10 (so that the cell surface area-dilatational elasticity Ks can be calculated from the shear elasticity Gs by Ks = (2C + 1)Gs, justification is provided later). Under fixed flow conditions matching experiments, we vary the cell surface elasticity Gs in a wide range, covering 0.01 ≤ Ca ≤ 3, with increments of δCa = 0.01 for Ca ≤ 0.1 and δCa = 0.05 for larger Ca. We obtain the steady-deformed cell profiles from numerical simulations and quantify their differences with the profile recorded in experiment using the mean Hausdorff distance (MHD).59 To explain the MHD, consider two sets of points, R = r1, r2, r3, …, rm and S = s1, s2, s3, …, sn, representing the steady cell profiles obtained from numerical simulation and experiment, respectively. Assuming that the two sets of points have the same centre of mass, the MHD ![[h with combining macron]](https://www.rsc.org/images/entities/i_char_0068_0304.gif) (R, S) of the two profiles is defined as:
(R, S) of the two profiles is defined as:
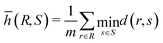 | (1) |
![[h with combining macron]](https://www.rsc.org/images/entities/i_char_0068_0304.gif) (R, S) is taken as the best fit. We present the best-fit parameters for all the cases of Fig. 2a and b in Table S1 of the SI. Note that in the present cell mechanical model the cell surface shear and area-dilatation elastic moduli Gs and Ks in the SK law are two-dimensional surface mechanical properties with a unit of N m−1. They quantify the resistance to in-plane shear and area-dilatation deformation of the thin cell surface, respectively. The three-dimensional bulk shear elastic modulus of the cell MCC, which has a unit of Pa, can be estimated by G = Gs/h, where h represents the thickness of the cell MCC.60
(R, S) is taken as the best fit. We present the best-fit parameters for all the cases of Fig. 2a and b in Table S1 of the SI. Note that in the present cell mechanical model the cell surface shear and area-dilatation elastic moduli Gs and Ks in the SK law are two-dimensional surface mechanical properties with a unit of N m−1. They quantify the resistance to in-plane shear and area-dilatation deformation of the thin cell surface, respectively. The three-dimensional bulk shear elastic modulus of the cell MCC, which has a unit of Pa, can be estimated by G = Gs/h, where h represents the thickness of the cell MCC.60
In general, for both cell lines, larger cells with lower surface elasticity tend to deform into bullet shapes, with a larger curvature at the front region, in order to balance the pressure drop outside the cell along the flow direction. With the cell size decreasing or surface elasticity increasing, the cells become more rounded.
Secondly, we find that by fitting the steady cell deformation profile recorded in an experiment to those predicted by numerical simulations using the present model, one can uniquely determine the cell surface shear and dilatational elastic moduli Gs and Ks. Examples for a K-562 and a PC-3 cell are shown in Fig. 2c and d, respectively, where the experimental results are compared with simulations with three combinations of Gs and Ks. Note that the three combinations have all resulted in an almost identical deformation index (DI), 0.06 and 0.05, respectively, for the K-562 and PC-3 cells. The cell deformation index is calculated as 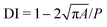 , where A and P are the area and perimeter of the cell's projection on the mid-depth plane of the channel. However, only one combination of parameters leads to excellent agreement with the experiment in terms of the cell shape, indicated by the minimum MHD. In our fitting, the parametric space of the numerical simulations covers 1 ≤ C ≤ 50 and 0.01 ≤ Ca ≤ 3. The MHD between the cell steady profiles obtained from simulations and experiments is shown in Fig. S1 of the SI. For both cells of Fig. 2c and d, at the best fit, we find that the membrane hardness parameter C in the SK law is close to 10. This is consistent with our earlier studies of the transient deformation of HL-60 cells flowing through a constricted channel,47 and of PC-3 and K-562 cells flowing through constricted and cross-slot microchannels,49 where we found that using the SK law with C = 10 is necessary for good agreements between simulations and experiments. Based on those observations, in Fig. 2a and b and the rest of this paper, we have set C = 10 to save computational effort. We discuss the limitations of this simplification in section 3.
, where A and P are the area and perimeter of the cell's projection on the mid-depth plane of the channel. However, only one combination of parameters leads to excellent agreement with the experiment in terms of the cell shape, indicated by the minimum MHD. In our fitting, the parametric space of the numerical simulations covers 1 ≤ C ≤ 50 and 0.01 ≤ Ca ≤ 3. The MHD between the cell steady profiles obtained from simulations and experiments is shown in Fig. S1 of the SI. For both cells of Fig. 2c and d, at the best fit, we find that the membrane hardness parameter C in the SK law is close to 10. This is consistent with our earlier studies of the transient deformation of HL-60 cells flowing through a constricted channel,47 and of PC-3 and K-562 cells flowing through constricted and cross-slot microchannels,49 where we found that using the SK law with C = 10 is necessary for good agreements between simulations and experiments. Based on those observations, in Fig. 2a and b and the rest of this paper, we have set C = 10 to save computational effort. We discuss the limitations of this simplification in section 3.
2.3 Performance of the MLP algorithm
We test the prediction accuracy and latency of the MLP-based algorithm using data generated from our numerical simulations, where the ground truth information, such as the cell surface elasticity and size, is exactly known. To train the MLP, we have used ∼600 sets of steady deformation profiles of cells with a wide range of dimensionless size, i.e., 0.4 ≤ 2a/W ≤ 0.9, and capillary number, i.e., 0.001 ≤ Ca ≤ 3.0. Here a and W respectively represent the cell radius and the channel width. The capillary number Ca is a key dimensionless parameter governing the cell deformation and is defined in section 4.5. The cell surface elasticity Gs is related to the capillary number by Gs = μU/Ca, where μ is the suspension fluid viscosity and U is the average flow speed in the channel. The parameter space considered has largely covered the range of cell deformation observed in the present experiments. Our testing samples, consisting of 120 cases in the same parameter space, have not been used during the training of the MLP. We compare the MLP predicted Ca of the testing samples with the corresponding ground truth in Fig. 3a. The insets show the extents of cell deformation in the low and high Ca regimes. In general, very good agreement has been observed. The mean absolute percentage error (MAPE) of the predicted Ca from the corresponding ground truth calculated from the entire testing samples is 5.8%, which suggests excellent prediction accuracy. The MAPE of a parameter y is defined as: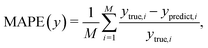 | (2) |
In wet experiments, cells are often not perfectly aligned to the channel centreline, even after using a sheath flow to focus the cells. We test the influence of a small initial offset, doc, defined as the distance between the cell mass centre and the centreline of the square straight channel, on the prediction accuracy of the present MLP. Note that the MLP has been trained by samples of perfectly aligned cells. Fig. 3b–d show the comparisons of the MLP predicted Ca with the corresponding ground truth, for cells with 2a/W = 0.7 and different offset distances along the z-direction (i.e., dzoc), y-direction (i.e., dyoc) and both directions. The coordinate system is defined in Fig. 10b, with the flow along the x-axis, the channel depth along the y-axis, and the channel width along the z-axis. Interestingly, from Fig. 3b–d we find that the MLP can still reach reasonably high accuracy, even for slightly off-centred cells. The MAPEs for the cells of Fig. 3b–d are shown in Fig. 3e. With doc increasing from 0 to 0.05W, the MAPE only grows moderately, from about 6% to 8%. In the present experiments, which we will show results later, we mainly focus on cells with doc ≤ 0.05W (i.e., 1 μm).
We also consider the effect of the number of cell membrane nodes, which is, to some extent, equivalent to the resolution of the input of the MLP, on the prediction accuracy and time (or latency) of the MLP. The same training and testing samples of Fig. 3a have been used here. As can be seen from Fig. 4, the prediction accuracy, indicated by the MAPE of the predicted Ca, generally increases with the number of membrane nodes and reaches a plateau when there are 60 membrane nodes. It is encouraging to find that even with 10 membrane nodes, the MLP can achieve a MAPE of 10.2%. With the number of membrane nodes increasing from 20 to 80, the prediction time only moderately grows from 0.2 to 0.24 ms.
 | ||
| Fig. 4 Effect of the number of cell membrane nodes representing a cell's steady profile on prediction accuracy (square) and latency (circle) of the present MLP. | ||
Due to the lack of quantitative experimental data of the surface elasticity of suspended cells, in particular for K-562 and PC-3 cells considered in the present study, it is not possible to rigorously test the accuracy of the present prediction algorithm. However, in a relevant recent study on mechanical characterisation of bioartificial microcapsules, we demonstrated that a similar MLP-based algorithm can accurately predict the capsules' membrane dilational modulus from its steady deformation in tube flow.54 Specifically, with 60 membrane nodes representing the capsule profile, extracted from an experimental image that has 42 pixels covering the capsule's diameter, the MLP reached a MAPE of 13% from the experimental measurements. In the present study, we have used comparable resolutions for the cell image and the membrane profile. Therefore we expect a similar level of prediction accuracy for the cell surface elasticity.
We test the prediction latency of the MLP-based algorithm for processing experimental cell images. The prediction latency is defined as the time it takes for the algorithm to predict the cell surface elasticity from a single camera-recorded image, without using any parallel computing. It consists of two parts: the image processing time and the network prediction time. The results are presented in Table 1. With 60 membrane nodes to represent the cell steady deformation profile, the present MLP-based algorithm can achieve a total latency of 0.64 ms, consisting of 0.42 ms in image processing and 0.22 ms in MLP prediction, using a laptop with an Intel i7-10850H, 2.7 GHz CPU. If we tolerate a higher MAPE of 7.4% by reducing the cell membrane resolution to 20 nodes, the total latency can be down to 0.48 ms.
| Image processing time (ms) | Prediction time (ms) | |
|---|---|---|
| 20 membrane nodes | 0.28 | 0.2 |
| 60 membrane nodes | 0.42 | 0.22 |
Regarding the throughput rate of the present system for characterisation of the surface elasticity of cells, based on a total latency of 0.64 ms per cell, the system can reach a theoretical throughput rate of 1562 cells per second. However, in our experiments, due to the limitation that the cell suspension cannot be too dense for avoiding the formation of cell clusters, we have recorded a peak throughput rate of 411 cells per second.
2.4 Surface mechanical heterogeneity of K-562 and PC-3 cells
We use the system to characterise the surface elasticity of human leukaemia K-562 and prostate cancer PC-3 cells. The scatter plots of the cell surface shear elastic modulus Gsversus cell size, i.e., the diameter of a cell assuming an initially spherical shape, measured from 71![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 414 K-562 and 96
414 K-562 and 96![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 998 PC-3 cells, are shown in Fig. 5a and b, respectively. The experimental conditions are identical to those of Fig. 2a and b for K-562 and PC-3 cells, respectively. Interestingly, for both cell lines, we find that the cell surface Gs spans more than two orders of magnitude. Such high levels of cell mechanical heterogeneity have not been reported previously for K-562 or PC-3 cells. In previous experiments of mechanical characterisation of the two cell lines, often the apparent Young's modulus of tens of cells was measured, and the variations are within one order of magnitude.61,62 The data in Fig. 5a and b are captured within 10 minutes of the flow experiments. The results collected at the beginning and end of the experiments, presented in Fig. S2 in the SI, show little difference, suggesting that the observed cell mechanical heterogeneity is not due to cytoskeletal remodelling of the cells during the experiments.
998 PC-3 cells, are shown in Fig. 5a and b, respectively. The experimental conditions are identical to those of Fig. 2a and b for K-562 and PC-3 cells, respectively. Interestingly, for both cell lines, we find that the cell surface Gs spans more than two orders of magnitude. Such high levels of cell mechanical heterogeneity have not been reported previously for K-562 or PC-3 cells. In previous experiments of mechanical characterisation of the two cell lines, often the apparent Young's modulus of tens of cells was measured, and the variations are within one order of magnitude.61,62 The data in Fig. 5a and b are captured within 10 minutes of the flow experiments. The results collected at the beginning and end of the experiments, presented in Fig. S2 in the SI, show little difference, suggesting that the observed cell mechanical heterogeneity is not due to cytoskeletal remodelling of the cells during the experiments.
The surface Gs and size of the K-562 and PC-3 cells in Fig. 5a and b are compared in Fig. 5c. In general, PC-3 cells are larger and have a stiffer membrane compared with K-562 cells. The average diameters of the K-562 and PC-3 cells in suspensions are 12.7 ± 1.9 μm (average ± standard deviation) and 16.2 ± 2.0 μm, respectively. The average surface Gs of the two cell lines are respectively 2.4 ± 9.6 mN m−1 and 8.6 ± 19.2 mN m−1. The high standard deviation of Gs is due to the huge mechanical heterogeneity of the cells as well as the log-normal distribution of the cell surface Gs, which we discuss later. Assuming a cell MCC thickness of h = 200 nm,3 we can estimate the bulk shear modulus G of the MCC through G = Gs/h.60 For K-562 and PC-3 cells, the average bulk shear modulus of the MCC is 12 ± 45 kPa and 43 ± 95 kPa, respectively. These are considerably higher than the apparent Young's modulus of the same cell lines characterised using AFM, which were reported to be 0.42 ± 0.38 kPa measured from 25 K-562 cells,61 and 2.9 ± 1.2 kPa measured from 40 PC-3 cells.62 The discrepancy is not surprising. The present system considers the cell surface as a thin material layer and measures its in-plane shear elasticity, while the AFM experiments had both used the Hertz contact model,61,62 simplified cells as semi-infinite homogeneous elastic solids, and probed the out-of-plane indentation response of the whole cell. With an advanced mechanical model accounting for the thin cell cortex layer, Cartagena-Rivera et al. reported AFM measurement of the cortical elasticity of primary human foreskin fibroblast cells of the order of 10 kPa,63 which is comparable to the present estimation.
Interestingly, from Fig. 5a one can find that larger K-562 cells tend to have a higher surface Gs. The two variables are positively correlated, with Spearman's rank correlation coefficient ρ = 0.128. In contrast, as can be seen from Fig. 5b, larger PC-3 cells tend to have a lower Gs, with a correlation coefficient of ρ = −0.552, indicating a negative correlation between the cell size and the surface Gs. The relations between cell size and surface Gs for the two cell lines are first reported in the present study. Although the underlying biophysical mechanisms are beyond the scope of the present study, several factors including the cytoskeletal organization and membrane–cortex coupling could contribute to the observation. A study of adherent cells has reported that larger cells can exhibit a thinner and/or less cross-linked actin cortex,64 which would be expected to reduce the cell surface elasticity. Further studies combining cell surface Gs measurement with direct assays of the cell cortical organisation for cells of different sizes and types will be an interesting direction of future work.
Fig. 5d and e present the histograms of the size and surface Gs of K-562 and PC-3 cells, respectively. It can be seen that for both cell lines, the cell size follows a normal distribution, and the cell surface Gs distribution seems to be consistent with a log-normal curve. Due to the heavy-tailed nature of the log-normal distribution, a small fraction of cells with very high surface Gs has substantially increased the average and standard deviation of Gs of both K-562 and PC-3 cells. When restricting the analyses to cells within the 10% density contour of Gsvs. size plots (i.e., data enclosed by the red curves in Fig. 5a and b), the variations of Gs become much smaller, with Gs = 1.4 ± 1.2 mN m−1 for K-562 and 5.3 ± 4.5 mN m−1 for PC-3 cells. The heavy-tailed log-normal distribution of the cell surface Gs also explains the fact that the average Gs is much higher than the median, which is 1.0 mN m−1 for K-562 cells and 3.9 mN m−1 for PC-3 cells. The peak of the Gs distribution is located closer to the median than the average.
We also test the possibility of classifying the two cell lines entirely based on their surface Gs and size, using the data of Fig. 5a and b. With a simple decision tree (DT) model,65 as detailed in section 4.7, we can reach an overall classification accuracy of 90.4%. Assuming PC-3 is the positive group, the sensitivity and specificity of the present DT are 0.89 and 0.93, respectively. The decision boundary in the cell size–surface Gs domain is shown in Fig. 5f. Despite the overlap of the cell size and the surface Gs of the two cell lines, the decision boundary of the DT is clear and no fragments of classes appear, indicating a good probabilistic basis for classification of the two types of cells based on the two physical parameters.
2.5 Effects of flow parameters on measured cell surface elasticity
We examine the effects of flow conditions, such as the flow speed and suspension fluid viscosity, on the measured surface Gs. The results for PC-3 cells of the same batch of Fig. 5b are presented in Fig. 6. We first increase the flow speed in experiments from 0.07 to 0.14 m s−1 while using the same suspension medium. This leads to stronger fluid force applied to the cells and noticeable larger cell deformation. The scatter plot of cell surface shear elastic modulus Gsversus cell size, characterised from 88![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 281 PC-3 cells at 0.14 m s−1, is shown in Fig. 6a. Similar to the result of the baseline experiment (Fig. 5b), the cell size and surface Gs are negatively correlated, with Spearman's rank correlation coefficient of ρ = −0.581. Fig. 6c compares the surface Gs and size of the PC-3 cells measured at the two different flow speeds. At the higher speed, the average cell surface Gs is 8.9 ± 16.4 mN m−1 with a median of 4 mN m−1. These are very close to the baseline results.
281 PC-3 cells at 0.14 m s−1, is shown in Fig. 6a. Similar to the result of the baseline experiment (Fig. 5b), the cell size and surface Gs are negatively correlated, with Spearman's rank correlation coefficient of ρ = −0.581. Fig. 6c compares the surface Gs and size of the PC-3 cells measured at the two different flow speeds. At the higher speed, the average cell surface Gs is 8.9 ± 16.4 mN m−1 with a median of 4 mN m−1. These are very close to the baseline results.
 | ||
Fig. 6 Effects of the flow speed and suspension fluid viscosity on the inferred surface Gs of PC-3 cells. Scatter plots of Gsversus size of (a) 88![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 281 PC-3 cells suspended in PBS added with 3% methylcellulose flowing at a speed of 0.14 m s−1 and (b) 60 281 PC-3 cells suspended in PBS added with 3% methylcellulose flowing at a speed of 0.14 m s−1 and (b) 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 863 PC-3 cells suspended in PBS added with 4% methylcellulose flowing at 0.07 m s−1. The result of the baseline experiment using the same batch of PC-3 cells is shown in Fig. 5b. (c) Box plots of the surface Gs and size of PC-3 cells measured under three different flow conditions. 863 PC-3 cells suspended in PBS added with 4% methylcellulose flowing at 0.07 m s−1. The result of the baseline experiment using the same batch of PC-3 cells is shown in Fig. 5b. (c) Box plots of the surface Gs and size of PC-3 cells measured under three different flow conditions. | ||
Next, we test the effect of the suspension fluid viscosity by flowing cells in a more viscous medium consisting of PBS added with 4% (w/w) methylcellulose at the same flow speed as the baseline case, i.e., 0.07 m s−1. The average fluid viscosity for the fully developed flow in the narrow straight channel is 58 mPa s. Compared with the baseline results, a higher fluid medium viscosity results in an increase in the cell steady deformation. The scatter plot of the cell surface Gsversus cell size, characterised from 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 863 PC-3 cells, is shown in Fig. 6b, and the box plots are presented in Fig. 6c. The average cell surface Gs is 8.6 ± 14.1 mN m−1, with a median of 3.9 mN m−1. Both are comparable to those of the baseline case, where cells are measured in a less viscous suspension medium.
863 PC-3 cells, is shown in Fig. 6b, and the box plots are presented in Fig. 6c. The average cell surface Gs is 8.6 ± 14.1 mN m−1, with a median of 3.9 mN m−1. Both are comparable to those of the baseline case, where cells are measured in a less viscous suspension medium.
Similar tests have also been conducted on K-562 cells, and the results are presented in Fig. S3 in the SI. For both cell lines, we find that the measured cell surface Gs is little affected by doubling the flow speed or suspension medium viscosity.
It is worth mentioning that biological cells have been found to exhibit strain-hardening behaviour, where the apparent stiffness of a cell increases with the cell deformation.66,67 In the present cell mechanical model, the SK law has already taken into account the strain-hardening effect of the cell surface, therefore it is not surprising that the inferred cell surface Gs has changed little with flow conditions.
2.6 Effect of cytochalasin D on cell surface Gs
To test the sensitivity of the present system in detecting changes of cell surface elasticity, we characterise PC-3 cells treated with a cytoskeletal drug, cytochalasin D (CytoD). CytoD disassembles the actin network of cells, including the actin cortex beneath the cell plasma membrane, and therefore reduces the cell stiffness.68,31 We treat PC-3 cells with CytoD at a concentration of 5 μM for 2 hours and measure their surface Gs using the present platform under a flow condition identical to that of Fig. 5b. Since the CytoD is dissolved in a DMSO solvent before being added to the cell culture medium, we also characterise PC-3 cells treated with the same concentration of DMSO as an extra control group to consider its possible effects on the cell surface Gs.The scatter plots of the cell surface Gsversus cell size of 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 141 CytoD-treated and 26
141 CytoD-treated and 26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 355 DMSO-treated PC-3 cells are shown in Fig. 7a and b, respectively. Both experiments show a negative correlation between cell size and surface Gs, similar to that in Fig. 5b that serves as a baseline. As shown in Fig. 7c, compared with the baseline result, DMSO added into the cell culture medium at 0.05% (v/v) does not affect the cells' surface Gs. However, after the CytoD treatment, the cell surface Gs drops significantly to an average of 5.1 ± 18.8 mN m−1, and the median decreases almost by half to 2.1 mN m−1. In Fig. 7a there is a small peak in the size distribution around 2a ∼8 μm. Likely, these are large cell-derived particles, including vesicles and apoptotic bodies, following the CytoD treatment that has strongly disrupted a small fraction of cells.69–71
355 DMSO-treated PC-3 cells are shown in Fig. 7a and b, respectively. Both experiments show a negative correlation between cell size and surface Gs, similar to that in Fig. 5b that serves as a baseline. As shown in Fig. 7c, compared with the baseline result, DMSO added into the cell culture medium at 0.05% (v/v) does not affect the cells' surface Gs. However, after the CytoD treatment, the cell surface Gs drops significantly to an average of 5.1 ± 18.8 mN m−1, and the median decreases almost by half to 2.1 mN m−1. In Fig. 7a there is a small peak in the size distribution around 2a ∼8 μm. Likely, these are large cell-derived particles, including vesicles and apoptotic bodies, following the CytoD treatment that has strongly disrupted a small fraction of cells.69–71
 | ||
Fig. 7 Effect of CytoD on the surface Gs of PC-3 cells. Scatter plots of Gsversus size of (a) 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 141 PC-3 cells treated with a 5 μM CytoD solution for 2 hours and (b) 26 141 PC-3 cells treated with a 5 μM CytoD solution for 2 hours and (b) 26![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 355 PC-3 cells treated with a 0.05% (v/v) DMSO solution for 2 hours. The result of the baseline experiment using PC-3 cells of the same batch without any chemical treatment is shown in Fig. 5b. (c) Box plots of the surface Gs and size of PC-3 cells treated with CytoD and DMSO and not treated with any chemical. 355 PC-3 cells treated with a 0.05% (v/v) DMSO solution for 2 hours. The result of the baseline experiment using PC-3 cells of the same batch without any chemical treatment is shown in Fig. 5b. (c) Box plots of the surface Gs and size of PC-3 cells treated with CytoD and DMSO and not treated with any chemical. | ||
3 Conclusions
The present study aims to develop a system for real-time high-throughput mechanical characterisation of the intrinsic surface elasticity of suspended cells. To achieve this goal, we build an automatic processing pipeline, which seamlessly integrates a microfluidic device for controlled flow-induced cell deformation, a high-speed streaming camera which records the steady-deformation profiles of the cells in the microfluidic and passes the data to a personal computer, an MLP-based machine learning algorithm on the computer predicting the intrinsic cell surface elasticity from the steady cell deformation image, and finally a high-fidelity mechanistic model which simulates cell deformation in the microfluidic in wide parametric space and provides training data to the MLP algorithm.A major merit of the present system is that it can measure the surface elasticity of large-population suspended cells. This is enabled by the integration of the MLP, as a method for fast inverse analysis, and the cell mechanistic model, which resolves the cell surface, cytoplasm and nucleus. Compared with cell deformation parameters, which are primary outcomes of previous high-throughput DC systems, intrinsic cell surface elasticity is less affected by the flow geometry49 and therefore may facilitate comparisons of cell mechanical properties measured by different systems.
The present system also features an unprecedented sub-millisecond per cell image-to-property latency, and can measure the surface elasticity of up to 411 cells per second. This is several orders of magnitude faster than the current state-of-the-art systems that can measure intrinsic mechanical properties of cells, such as the AFM or micropipette. It will enable researchers to thoroughly investigate the mechanical heterogeneity of the cell surface, a fundamental property that might correlate to molecular differences in the cell membrane–cortex complex.72 By measuring tens of thousands of K-562 and PC-3 cells, we have found high levels of heterogeneity of the cell surface elasticity for both cell lines, which are one order of magnitude greater than previously reported cell apparent elasticity. We also find that the surface elasticity of K-562 cells increases with the cell size, while the PC-3 cells show the opposite trend.
It has been well established that the mechanics of the cell membrane–cortex complex plays a crucial role in controlling cell division, growth and migration.5 The present system may therefore benefit research along those lines by enabling screening the surface elasticity of large-population heterogeneous cells in a short time. Some cancer drugs specifically target the cell surface. For example, 4-hydroxyacetophenone could reduce cancer metastasis by increasing the cell cortex stiffness.73 The present system may be useful in monitoring the efficacy of such drugs. Indeed, we have demonstrated that the system is sensitive enough to detect the reduction of the cell surface elasticity as a result of the CytoD-induced actin disassembly.
In the context of label-free cell classification and sorting, our results suggest that the cell surface elasticity, together with the cell size, could provide a good probabilistic basis for classification of cell types. Using them to build a decision tree, we show that one can classify K-562 and PC-3 cells with an accuracy of 90.4%. Together with the sub-millisecond prediction latency of the present MLP algorithm, the system could be supplemented by a sorting module to achieve high-throughput cell sorting based on cell surface elasticity and size.
One limitation of the present study is that we have fixed the relation between the cell surface shear and the area-dilatation moduli in the SK law in our computational model, following our previous and the present tests on several cancer cells lines including the HL-60, PC-3 and K-562 cells in different flow setups.47,49 While this simplifies the inverse problem, biological cells exhibit significant heterogeneity. This assumption should be released in future work. The training sample will need to be significantly expanded. Previous studies including ours56,74 have shown that it is possible to simultaneously infer the surface shear and area-dilatational elastic moduli of microcapsules by fitting the steady deformation profiles obtained from experiments and model predictions.
Besides, although we have separately tested the accuracy of the cell mechanical model and the MLP algorithm, the two key components of the present system, rigorous cross-validation of the present measurements against a different well-established system has not been achieved. The main reason is the lack of quantitative data for the surface elasticity of suspended cells. Encouragingly, the present results of the surface elasticity of both K-562 and PC-3 cells are largely consistent with our recent measurements.49 There we flowed the same cell lines through both constricted and cross-slot microchannel geometries and fit the cell surface elasticity by minimising the mean Hausdorff distance between model-predicted and experimentally measured cell deformation profiles. From tens of cells of the two cell lines, we found that the surface Gs of K-562 and PC-3 cells are respectively in the ranges of 3.6–5.2 and 5.8–20.7 mN m−1.49 These fall into the parametric space measured in the present study on much larger samples.
In principle, the present system could be validated against measurements of micropipette aspiration of the cell surface. However, in the latter approach, cells are often considerably larger than the micropipette, leading to physical contact and strong mechanical interaction between the cell and the glass capillary wall that affect cell deformation. These are currently not included in our mechanical model. Another possibility is to measure well-characterised synthetic bioartificial microparticles such as hydrogel microcapsules. We had demonstrated in a recent study that an MLP-based method, similar to the present algorithm, can accurately characterise the surface area-dilatational modulus of bio-artificial capsules with a human serum albumin–alginate membrane from the capsule's steady deformation in tube flow.54 The MAPE of the MLP-predictions from the experimental measurements using a parallel plate compression method reached 13%. However, those microcapsules often follow mechanical constitutive laws that are different from the present cell model.75 They are also much larger than biological cells, with a diameter of above 100 μm.
4 Materials and methods
4.1 Experimental setup
The geometry of the microchannel which we use to flow through and deform human prostate cancer PC-3 and leukaemia K-562 cells is shown in Fig. 8. It mainly consists of a hydrodynamic sheath flow module with a 45 constriction connected by a long straight square channel. Filter posts are placed before the channel inlets to remove any cell clusters or debris. The cell suspension and sheath flow are driven by two syringe pumps (PHD ULTRA, Harvard Apparatus) with a flow rate ratio of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2, and the average flow speed in the straight channel is 0.07–0.14 m s−1. As illustrated in the photo inset of the lower panel of Fig. 8, when approaching the narrow straight channel, a cell is elongated and reaches the maximum extension at the entrance. Inside the narrow straight channel, the cell gradually deforms into a steady bullet shape under the effect of fluid shear. The cell steady profile in the region of interest (RoI) is illuminated by an LED lamp (LA-HDF-7010, HAYASHI) and recorded with a high-speed streaming camera (EXO174MU3, SVS-Vistek). The camera is equipped with a Mitutoyo 20× long working distance objective and works at a frame rate of 1400 frames per second with a shutter speed of 19 μs. The channel width is covered by 68 pixels. As shown in Fig. 8, we have observed that cell boundaries in our images are clear and there are no halos, indicating that when reaching the RoI cells are largely flowing along the channel centreline, which is in the focal plane of imaging.
2, and the average flow speed in the straight channel is 0.07–0.14 m s−1. As illustrated in the photo inset of the lower panel of Fig. 8, when approaching the narrow straight channel, a cell is elongated and reaches the maximum extension at the entrance. Inside the narrow straight channel, the cell gradually deforms into a steady bullet shape under the effect of fluid shear. The cell steady profile in the region of interest (RoI) is illuminated by an LED lamp (LA-HDF-7010, HAYASHI) and recorded with a high-speed streaming camera (EXO174MU3, SVS-Vistek). The camera is equipped with a Mitutoyo 20× long working distance objective and works at a frame rate of 1400 frames per second with a shutter speed of 19 μs. The channel width is covered by 68 pixels. As shown in Fig. 8, we have observed that cell boundaries in our images are clear and there are no halos, indicating that when reaching the RoI cells are largely flowing along the channel centreline, which is in the focal plane of imaging.
The long straight square channel of the present system has a cross-sectional dimension of W = H = 20 μm. This is due to the average diameter of the K-562 and PC-3 cells considered in the present study, which is 2a ∼13 and 13 μm, respectively. We find that an ideal confinement ratio 2a/W should be in the range of 0.5–0.9. A channel that is too wide results in weaker confinement, leading to smaller cell deformation under the same average flow speed. It has been well established that accurately inferring the mechanical properties of microcapsules and cells undergoing small deformation is very challenging.56,76 It is also more difficult to hydrodynamically focus cells to the centreline of a wider channel. On the other hand, if the channel is too narrow, there is a higher chance that the large cells in a suspension would contact the channel wall, requiring additional wall–cell interaction to be accounted for in the mechanical model, and some large cells would even block the channel. A narrow channel also needs a high pressure drop to maintain a specific flow speed, which is essential to the processing throughput rate.
The cell suspension buffer is made of PBS added with 2–4% (w/w) methylcellulose (15 cP, Sigma-Aldrich) to control its viscosity. We measure the viscosity using a cone-plate rheometre (AR2000, TA Instruments) at room temperature and find the buffer is weakly shear-thinning, following a power law:
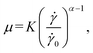 | (3) |
![[small gamma, Greek, dot above]](https://www.rsc.org/images/entities/i_char_e0a2.gif) is the local shear rate and
is the local shear rate and ![[small gamma, Greek, dot above]](https://www.rsc.org/images/entities/i_char_e0a2.gif) 0 = 1 s−1. At methylcellulose concentrations of 2%, 3%, and 4%, the values of K are 0.016, 0.059 and 0.442 Pa s, respectively, and α = 0.99, 0.94 and 0.78.
0 = 1 s−1. At methylcellulose concentrations of 2%, 3%, and 4%, the values of K are 0.016, 0.059 and 0.442 Pa s, respectively, and α = 0.99, 0.94 and 0.78.
4.2 Device fabrication
The microchannels are fabricated using a standard soft lithography method with polymethylsiloxane (PDMS).77 To produce a master mould, a negative photoresist (SU8-2025, Kayaku) is spread at 500 rpm for 5 s and spin-coated on a 4 inch silicon wafer (Test Grade, PI-KEM, UK) at 4000 rpm for 30 s using a spin coater (SPIN150i, SPS-POLOS) to generate a 20 μm photoresist film, which is the height of the microchannel. After spinning, the wafer is soft-baked at 95 C for 5 minutes to evaporate the photoresist solvent. The chip design, depicted on a film mask, is transferred on the coated substrate via UV exposure at 160 mJ cm−2 (UV-KUB 9, Kloe), followed by post-exposure baking at 95 C for another 5 minutes and development in SU8 developer (Kayaku) for 4 minutes. The master that carries the microchannel design is inspected using a profilometer (Profilm 3D, Filmmetrics). After master fabrication, PDMS (base-to-curing agent ratio of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 w/w, Slygard 184, Dow) is degassed, poured over the master, and cured at 60 C for 2 hours. Following the cutting of the chips and the punching of 0.7 mm inlet and outlet holes, the PDMS replica is cleaned with isopropanol and treated using oxygen plasma (Zepto, Diener) and then bonded together. The bonded device is incubated at 60 C for 12 h to strengthen the bonding, and finally, PTFE tubing is connected to the inlets and outlet for flow experiments.
1 w/w, Slygard 184, Dow) is degassed, poured over the master, and cured at 60 C for 2 hours. Following the cutting of the chips and the punching of 0.7 mm inlet and outlet holes, the PDMS replica is cleaned with isopropanol and treated using oxygen plasma (Zepto, Diener) and then bonded together. The bonded device is incubated at 60 C for 12 h to strengthen the bonding, and finally, PTFE tubing is connected to the inlets and outlet for flow experiments.
4.3 Cell culture
PC-3 and K-562 cells are provided by the Barts Cancer Institute (London, UK) and cultured at 37 C with 5% CO2 according to standard mammalian tissue culture protocols.78 PC-3 cells are cultured in RPMI-1640 medium (Sigma-Aldrich, Cat. no. R8758) with 10% (v/v) fetal bovine serum (FBS, Sigma-Aldrich, Cat. no. F7524) and 1% (v/v) penicillin–streptomycin (Sigma-Aldrich, Cat. no. P4333). The cells are passaged every third day and harvested for experiments at a confluency of around 80% when cells are growing at the log phase. During harvesting, PC-3 cells are washed once with PBS, detached using trypsin–EDTA (Sigma-Aldrich, Cat. no. T4049), centrifuged at 300g for 3 minutes, and finally resuspended into the PBS with methylcellulose at a concentration of around 107 cells per ml for flow experiments.K-562 cells are cultured in the same medium as the PC-3 cells with extra 1% (v/v) L-glutamine (Sigma-Aldrich, Cat. no. G7513). The cells are split every third day and resuspended into the same new medium at a concentration of around 0.2 million per ml. When the cells are growing at the log phase to a concentration of around 0.7 million per ml, they are centrifuged at 150g for 5 minutes and then resuspended into the PBS solution with 2% (w/w) methylcellulose at a concentration of around 107 cells per ml for experiments.
To test the sensitivity of the present system, we consider PC-3 cells treated with cytochalasin D (CytoD, 5 mg ml−1 in DMSO, Sigma-Aldrich, Cat. no. C2618). Before harvesting, PC-3 cells are cultured with CytoD stock solution added to the cell culture medium at 0.05% (v/v), equivalent to 5 μM, for 2 hours. To test the effect of the DMSO solvent on cell surface elasticity, we also measure PC-3 cells from the same batch that are cultured with DMSO added to the cell culture medium at 0.05% (v/v) for 2 hours.
4.4 MLP-based prediction algorithm
The MLP-based prediction algorithm follows the method we developed recently,54 where more details can be found. The network consists of one input, two hidden and one output layers, with each hidden layer having 256 perceptrons, which are the basic network elements. The perceptrons conduct nonlinear input–output mapping defined by the regression problem. Each perceptron receives weighted inputs from all perceptrons of the previous layer, and its output is calculated by passing the sum of weighted input signals through a nonlinear activation function. The output of the jth perceptron in the nth layer can be obtained from: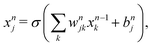 | (4) |
| σ(xnj) = max(0, xnj). | (5) |
The samples for training of the MLP have been obtained from numerical simulations resolving the mechanical interaction between the cell and the suspension fluid using an immersed-boundary lattice Boltzmann method as described in sections 4.5 and 4.6. A sample consists of a steady deformation profile of a cell in the channel flow and its labels in the forms of the corresponding surface elasticity and cell size. We discretize the cell profile into membrane nodes that have equal arc-length distances and assemble their coordinates into a 1D vector. During training, internal parameters of the MLP, such as the weight and bias, are adjusted to minimize a mean square root (MSE) loss function, which describes how close the prediction is to the corresponding ground truth. The MSE function is defined as:
| L = |ylabel − y2|, | (6) |
To optimize the trainable parameters in the MLP, we obtain the gradients of the loss functions with respect to the trainable parameters and then update the values of the parameters with an optimizer based on a stochastic gradient descent algorithm called ADAM80 with an initial learning rate of 0.001. Mini-batch mode training with a batch size of 128 is applied, and exposing all training samples to the MLP once is called an epoch. At the end of each epoch, the MLP is validated using a small portion (10% in the present study) of training samples that have not been actually used in training. With the process iterating, the losses decrease and converge to small values, indicating network predictions are approaching their corresponding ground truth. To avoid overfitting, batch-normalization81 and dropout regularization82 at a dropout rate of 0.3 have been applied. The training is terminated when the values of loss functions no longer decrease over several iterations even after reducing the learning rate.
After training, the MLP can predict the cell surface elasticity and size from the steady profile of a cell in a channel flow. We build an image processing pipeline, as illustrated in Fig. 9, to convert a cell image into a 1D coordinates vector that can be processed by the MLP. Firstly, the camera-recorded image is subtracted from its background to remove the channel and static noise to strengthen the cell features. The image is then binarized, and holes on the binarized image within the cell are filled. A cell is detected from the binarized image by finding any connected region above a threshold area, that is set to 50 μm2, considering the sizes of cells characterised in the present study. Next, the boundary of the detected cell is extracted from the binarized image with a border following algorithm,83 and the boundary nodes are sorted counterclockwise. As can be seen in Fig. 9, the cell boundary directly extracted from the binary image is not smooth, which can significantly reduce the prediction accuracy of the present method. To solve the problem, we use a piecewise second-order polynomial function to approximate the cell boundary smoothly and boundary nodes with equal arc-length distance from the fitted curve. Finally, the coordinates of those boundary nodes are built into a 1D vector, in the same format as the training data, that can be processed by the MLP.
4.5 Cell mechanical model
We simulate the flow-induced deformation of an initially spherical cell with radius a in a square channel with width W, as illustrated in Fig. 10. A three-dimensional Cartesian coordinate system is defined with the x-axis along the flow direction, the y- and z-axes along the depth and width of the channel, and the origin at the centre of the channel inlet plane. Following the model of Wang et al.,47 we consider a cell as a compound three-layer structure including a viscoelastic MCC, a viscous cytoplasm, and a nucleus that is smaller and stiffer than the cell. | ||
| Fig. 10 Illustrations of (a) the cell mechanical model, which accounts for the cell surface, cytoplasm, and nucleus, and (b) an initially spherical cell flowing through a straight square channel. | ||
The elasticity of the MCC follows the strain-hardening Skalak's (SK) law,66 with a strain energy function:
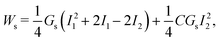 | (7) |
Since the MCC is assumed to be infinitely thin in the model, we take into account a small bending resistance using Helfrich's bending energy formulation,84
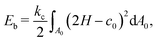 | (8) |
The cell cytoplasm is modelled as a Newtonian liquid. The cell nucleus is represented by a small capsule with radius an, which consists of a viscous fluid core enclosed by a hyperelastic nucleus membrane that follows the SK law. Its membrane elastic moduli are assumed to be twice those of the cell membrane to represent the fact that a cell nucleus is generally stiffer than the whole cell.
We are mainly interested in the steady deformation of cells in the channel flow. At the steady state, the cell profile is not changing and the cytoplasm is largely in solid translation, therefore, viscous effects of the cell subcellular components have little effect on the cell steady profile.
In the present study, inertial effect is negligible; the steady deformation of the cells is mainly determined by the following two dimensionless parameters:
• The capillary number Ca = μU/Gs, which compares the viscous fluid force acting on the cell and the membrane elastic force. The terms μ and U, respectively, are the average viscosity and flow speed of the fluid in the straight channel. At a higher Ca, a cell will undergo larger deformation.
• The confinement ratio β = 2a/W, which compares the size of the cell to the width of the channel.
In the range of small to moderate cell deformation, which has been the focus of the present study, we find that a cell nucleus with nucleus-to-cell size ratio an/a ≤0.7 will not significantly affect the cell steady profile. One example is shown in Fig. 11, for a cell with β = 0.75, 0.5 ≤ an/a ≤ 0.7 at Ca = 3.0, where the cell is undergoing moderate deformation. This is again due to the solid translational motion of the cell interior. In the experiments we measure the an/a of both the K-562 and the PC-3 cell lines. The cell nuclei are stained with DAPI and visualised using a fluorescence microscope (Axioscope, ZEISS), while the cells are visualised with bright field imaging. Measuring from ∼100 cells of each of the cell lines, we find that the average an/a for K-562 and PC-3 cells are 0.62 and 0.59, respectively, with 76% K-562 and 75% PC-3 cells having an/a ≤0.7. To save computational efforts, in the present simulations, we have assumed an/a = 0.6.
 | ||
| Fig. 11 Cell nucleus with an/a ≤0.7 has little effect on the steady cell deformation profile in the moderate deformation regime at Ca = 3.0, β = 0.75. | ||
4.6 Computational method
The present computational framework is based on a well-tested immersed boundary lattice Boltzmann method.47–49,86–91 Briefly, the fluid flow is governed by the three-dimensional Navier–Stokes equations, which are solved using a 3D nineteen-velocity lattice Boltzmann model. For the viscosity of the channel fluid, a truncated power-law model is used.92 At the walls of the square channel, the non-slip boundary condition is applied using a second-order bounce back scheme.93 A second-order non-equilibrium extrapolation method94 was employed to impose the pressure boundary conditions at the inlet and outlet, and the pressure drop is adjusted to match the flow conditions in the experiments. The fluid–cell interaction is solved using an immersed boundary method.95,96 The membrane of the cell is discretised into flat triangular elements following Ramanujan.97 A finite element method is used to calculate the membrane elastic forces from the membrane deformation and its strain energy function. We conduct a mesh convergence test; the fluid grid size that is finally chosen is Δx = 0.01W, and the MCC of the cell was discretised into 32![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 768 flat triangular elements connecting 16
768 flat triangular elements connecting 16![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 386 nodes.
386 nodes.
4.7 Decision tree for cell classification
The decision tree (DT) is a supervised machine learning algorithm that is widely used for classification tasks. Compared with other machine learning methods, DTs are not black-box models, can be easily expressed by rules, and therefore have been used in a vast range of applications such as drug discovery,98 emotion recognition99 and disease treatment.100A DT consists of root, internal and leaf nodes connected in a hierarchical tree structure. Each decision node carries a rule that helps split the data set reaching the node into several subsets. In this study, the data set consists of the cell size, surface shear elastic modulus Gs, and the associated cell type, which are shown in Fig. 5a and b. The rule on a decision node is a threshold of either the cell size or the surface Gs that can be used to split the cells into K-562 and PC-3 groups. The quality of the data split is measured by the Gini index, defined as:
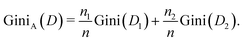 | (9) |
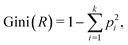 | (10) |
Author contributions
Y. S. designed the research. Z. G. and Y. S. developed and built the system. Z. G. conducted the experiments. All authors analyzed the results and wrote and reviewed the manuscript.Conflicts of interest
The authors declare no conflicts of interests.Data availability
The authors confirm that the data supporting the findings of this study are available within the main article. Raw data can be obtained from the corresponding author upon request. Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5lc00909j.Acknowledgements
Y. S. and W. W. acknowledge research grants from the UK Royal Society (IES/R2/212075, NIF/R1/191510) and the Engineering and Physical Sciences Research Council (EPSRC, EP/Z002818/1). Z. G. acknowledges the PhD studentships provided by the Queen Mary University of London (QMUL) and the Chinese Scholarship Council. The simulations were performed using the high-performance computer clusters of QMUL (funded by the UK EPSRC grants EP/K000128/1, EP/P020194/1 and EP/T022213/1). All authors thank Prof Yong-Jie Lu of the Barts Cancer Institute for providing the cell lines. The authors also thank the two referees whose comments have improved the quality of the paper.Notes and references
- A. Yeung and E. Evans, Biophys. J., 1989, 56, 139–149 CrossRef CAS PubMed.
- A. Diz-Muñoz, O. D. Weiner and D. A. Fletcher, Nat. Phys., 2018, 14, 648–652 Search PubMed.
- L. Lamparter and M. Galic, Front. Cell Dev. Biol., 2020, 8, 684 Search PubMed.
- M. P. Sheetz, Nat. Rev. Mol. Cell Biol., 2001, 2, 392–396 CrossRef CAS PubMed.
- G. Salbreux, G. Charras and E. Paluch, Trends Cell Biol., 2012, 22, 536–545 Search PubMed.
- A. Diz-Muñoz, D. A. Fletcher and O. D. Weiner, Trends Cell Biol., 2013, 23, 47–53 CrossRef PubMed.
- J. Sleep, D. Wilson, R. Simmons and W. Gratzer, Biophys. J., 1999, 77, 3085–3095 CrossRef CAS PubMed.
- P. Katira, M. H. Zaman and R. T. Bonnecaze, Phys. Rev. Lett., 2012, 108, 028103 Search PubMed.
- A. Callan-Jones, O. E. A. Arriagada, G. Massiera, V. Lorman and M. Abkarian, Biophys. J., 2012, 103, 2475–2483 CrossRef CAS PubMed.
- M. F. Coughlin, D. R. Bielenberg, G. Lenormand, M. Marinkovic, C. G. Waghorne, B. R. Zetter and J. J. Fredberg, Clin. Exp. Metastasis, 2013, 30, 237–250 CrossRef CAS PubMed.
- C. Händel, B. S. Schmidt, J. Schiller, U. Dietrich, T. Möhn, T. R. Kießling, S. Pawlizak, A. W. Fritsch, L.-C. Horn and S. Briest, et al. , New J. Phys., 2015, 17, 083008 CrossRef.
- M. J. Rosenbluth, W. A. Lam and D. A. Fletcher, Biophys. J., 2006, 90, 2994–3003 Search PubMed.
- T. G. Kuznetsova, M. N. Starodubtseva, N. I. Yegorenkov, S. A. Chizhik and R. I. Zhdanov, Micron, 2007, 38, 824–833 Search PubMed.
- E. Evans and A. Yeung, Biophys. J., 1989, 56, 151–160 Search PubMed.
- R. M. Hochmuth, J. Biomech., 2000, 33, 15–22 CrossRef CAS PubMed.
- H. Wang, F. Zhou, Y. Guo and L. A. Ju, Eur. Biophys. J., 2022, 51, 119–133 Search PubMed.
- B. Fabry, G. N. Maksym, J. P. Butler, M. Glogauer, D. Navajas and J. J. Fredberg, Phys. Rev. Lett., 2001, 87, 148102 CrossRef CAS PubMed.
- V. M. Laurent, E. Planus, R. Fodil and D. Isabey, Biorheology, 2003, 40, 235–240 Search PubMed.
- M. Irmscher, A. M. de Jong, H. Kress and M. W. Prins, Biophys. J., 2012, 102, 698–708 Search PubMed.
- P.-H. Wu, D. R.-B. Aroush, A. Asnacios, W.-C. Chen, M. E. Dokukin, B. L. Doss, P. Durand-Smet, A. Ekpenyong, J. Guck and N. V. Guz, et al. , Nat. Methods, 2018, 15, 491–498 CrossRef CAS PubMed.
- G. Tomaiuolo, M. Barra, V. Preziosi, A. Cassinese, B. Rotoli and S. Guido, Lab Chip, 2011, 11, 449–454 RSC.
- D. R. Gossett, H. T. Tse, S. A. Lee, Y. Ying, A. G. Lindgren, O. O. Yang, J. Rao, A. T. Clark and D. Di Carlo, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 7630–7635 Search PubMed.
- S. Byun, S. Son, D. Amodei, N. Cermak, J. Shaw, J. H. Kang, V. C. Hecht, M. M. Winslow, T. Jacks and P. Mallick, et al. , Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 7580–7585 Search PubMed.
- E. M. Darling and D. Di Carlo, Annu. Rev. Biomed. Eng., 2015, 17, 35–62 CrossRef CAS PubMed.
- O. Otto, P. Rosendahl, A. Mietke, S. Golfier, C. Herold, D. Klaue, S. Girardo, S. Pagliara, A. Ekpenyong and A. Jacobi, et al. , Nat. Methods, 2015, 12, 199–202 Search PubMed.
- J. Shaw Bagnall, S. Byun, S. Begum, D. T. Miyamoto, V. C. Hecht, S. Maheswaran, S. L. Stott, M. Toner, R. O. Hynes and S. R. Manalis, Sci. Rep., 2015, 5, 18542 CrossRef CAS PubMed.
- K. D. Nyberg, K. H. Hu, S. H. Kleinman, D. B. Khismatullin, M. J. Butte and A. C. Rowat, Biophys. J., 2017, 113, 1574–1584 Search PubMed.
- N. Toepfner, C. Herold, O. Otto, P. Rosendahl, A. Jacobi, M. Kräter, J. Stächele, L. Menschner, M. Herbig and L. Ciuffreda, et al. , eLife, 2018, 7, e29213 Search PubMed.
- F. J. Armistead, J. G. De Pablo, H. Gadêlha, S. A. Peyman and S. D. Evans, Biophys. J., 2019, 116, 1127–1135 CrossRef CAS PubMed.
- B. Fregin, F. Czerwinski, D. Biedenweg, S. Girardo, S. Gross, K. Aurich and O. Otto, Nat. Commun., 2019, 10, 415 Search PubMed.
- M. Urbanska, H. E. Muñoz, J. Shaw Bagnall, O. Otto, S. R. Manalis, D. Di Carlo and J. Guck, Nat. Methods, 2020, 17, 587–593 Search PubMed.
- R. Gerum, E. Mirzahossein, M. Eroles, J. Elsterer, A. Mainka, A. Bauer, S. Sonntag, A. Winterl, J. Bartl and L. Fischer, et al. , eLife, 2022, 11, e78823 Search PubMed.
- M. Urbanska and J. Guck, Annu. Rev. Biophys., 2024, 53, 367–395 CrossRef CAS PubMed.
- F. Reichel, R. Goswami, S. Girardo and J. Guck, Lab Chip, 2024, 24, 2440–2453 RSC.
- B. Hartmann, F. Reichel, C. Möckel and J. Guck, bioRxiv, 2025, preprint, 2025–04, DOI:10.1101/2025.04.17.649258.
- A. Mietke, O. Otto, S. Girardo, P. Rosendahl, A. Taubenberger, S. Golfier, E. Ulbricht, S. Aland, J. Guck and E. Fischer-Friedrich, Biophys. J., 2015, 109, 2023–2036 Search PubMed.
- M. Raissi, A. Yazdani and G. E. Karniadakis, Science, 2020, 367, 1026–1030 Search PubMed.
- G. E. Karniadakis, I. G. Kevrekidis, L. Lu, P. Perdikaris, S. Wang and L. Yang, Nat. Rev. Phys., 2021, 3, 422–440 CrossRef.
- V. Sekar, M. Zhang, C. Shu and B. C. Khoo, AIAA J., 2019, 57, 993–1003 Search PubMed.
- P. Jiao, C. Zhang, W. Meng, J. Wang, D. Jang, Z. Wu, N. Agarwal and A. H. Alavi, ACS Appl. Mater. Interfaces, 2025, 17, 2991–3001 CrossRef CAS PubMed.
- C. Stashko, M.-K. Hayward, J. J. Northey, N. Pearson, A. J. Ironside, J. N. Lakins, R. Oria, M.-A. Goyette, L. Mayo and H. G. Russnes, et al. , Nat. Commun., 2023, 14, 3561 CrossRef CAS PubMed.
- M. S. Schmitt, J. Colen, S. Sala, J. Devany, S. Seetharaman, A. Caillier, M. L. Gardel, P. W. Oakes and V. Vitelli, Cell, 2024, 187, 481–494 CrossRef CAS PubMed.
- Z. Wu, Y. Feng, R. Bi, Z. Liu, Y. Niu, Y. Jin, W. Li, H. Chen, Y. Shi and Y. Du, Cell Regener., 2025, 14, 21 Search PubMed.
- A. A. Nawaz, M. Urbanska, M. Herbig, M. Nötzel, M. Kräter, P. Rosendahl, C. Herold, N. Toepfner, M. Kubánková and R. Goswami, et al. , Nat. Methods, 2020, 17, 595–599 CrossRef CAS PubMed.
- C. Combs, D. D. Seith, M. J. Bovyn, S. P. Gross, X. Xie and Z. S. Siwy, Biomicrofluidics, 2022, 16, 014104 Search PubMed.
- M. Herbig, A. Jacobi, M. Wobus, H. Weidner, A. Mies, M. Kräter, O. Otto, C. Thiede, M.-T. Weickert and K. S. Götze, et al. , Sci. Rep., 2022, 12, 870 Search PubMed.
- Z. Wang, R. Lu, W. Wang, F. Tian, J. Feng and Y. Sui, Biomech. Model. Mechanobiol., 2023, 22, 1129–1143 Search PubMed.
- R. Lu, P. Yu and Y. Sui, Soft Matter, 2024, 20, 4057–4071 RSC.
- R. Lu, J. Li, Z. Guo, Z. Wang, J. Feng and Y. Sui, Biomech. Model. Mechanobiol., 2025, 24, 489–506 CrossRef CAS PubMed.
- F. Y. Leong, Q. Li, C. T. Lim and K. H. Chiam, Biomech. Model. Mechanobiol., 2011, 10, 755–766 CrossRef PubMed.
- X. Zhang, X. Chen and H. Tan, Microfluid. Nanofluid., 2017, 21, 146 CrossRef.
- N. Takeishi, Y. Imai, T. Yamaguchi and T. Ishikawa, Phys. Rev. E, 2015, 92, 063011 Search PubMed.
- J. Cui, Y. Liu, L. Xiao, S. Chen and B. M. Fu, Biomech. Model. Mechanobiol., 2021, 20, 243–254 Search PubMed.
- Z. Guo, T. Lin, D. Jing, W. Wang and Y. Sui, Biomech. Model. Mechanobiol., 2023, 22, 1209–1220 CrossRef PubMed.
- T. Lin, Z. Wang, R. Lu, W. Wang and Y. Sui, Phys. Fluids, 2021, 33, 1–12 Search PubMed.
- T. Lin, Z. Wang, R. Lu, W. Wang and Y. Sui, Adv. Appl. Math. Mech., 2022, 14, 79–100 Search PubMed.
- T. Lin, Z. Wang, W. Wang and Y. Sui, Soft Matter, 2021, 17, 4027–4039 RSC.
- F. Murtagh, Neurocomputing, 1991, 2, 183–197 CrossRef.
- M.-P. Dubuisson and A. K. Jain, Proceedings of 12th international conference on pattern recognition, 1994, pp. 566–568 Search PubMed.
- D. Barthes-Biesel, Annu. Rev. Fluid Mech., 2016, 48, 25–52 CrossRef.
- M. Islam, R. Mezencev, B. McFarland, H. Brink, B. Campbell, B. Tasadduq, E. K. Waller, W. Lam, A. Alexeev and T. Sulchek, Cell Death Dis., 2018, 9, 239 CrossRef PubMed.
- K. Pogoda, E. PiĘta, M. Roman, N. Piergies, D. Liberda, T. P. Wrobel, P. A. Janmey, C. Paluszkiewicz and W. M. Kwiatek, Arch. Biochem. Biophys., 2021, 697, 108718 CrossRef CAS PubMed.
- A. X. Cartagena-Rivera, J. S. Logue, C. M. Waterman and R. S. Chadwick, Biophys. J., 2016, 110, 2528–2539 Search PubMed.
- R. Kumar, S. Saha and B. Sinha, Biochim. Biophys. Acta, Mol. Cell Res., 2019, 1866, 118516 CrossRef CAS PubMed.
- S. B. Kotsiantis, Artif. Intell. Rev., 2013, 39, 261–283 CrossRef.
- R. Skalak, A. Tozeren, R. Zarda and S. Chien, Biophys. J., 1973, 13, 245–264 CrossRef CAS PubMed.
- J. Xu, Y. Tseng and D. Wirtz, J. Biol. Chem., 2000, 275, 35886–35892 CrossRef CAS PubMed.
- C. Rotsch and M. Radmacher, Biophys. J., 2000, 78, 520–535 CrossRef CAS PubMed.
- G. C. Godman, A. F. Miranda, A. D. Deitch and S. W. Tanenbaum, J. Cell Biol., 1975, 64, 644–667 CrossRef CAS PubMed.
- S. R. White, P. Williams, K. R. Wojcik, S. Sun, P. S. Hiemstra, K. F. Rabe and D. R. Dorscheid, Am. J. Respir. Cell Mol. Biol., 2001, 24, 282–294 CrossRef CAS PubMed.
- D. Di Vizio, M. Morello, A. C. Dudley, P. W. Schow, R. M. Adam, S. Morley, D. Mulholland, M. Rotinen, M. H. Hager and L. Insabato, et al. , Am. J. Pathol., 2012, 181, 1573–1584 Search PubMed.
- S. J. Altschuler and L. F. Wu, Cell, 2010, 141, 559–563 CrossRef CAS PubMed.
- D. S. Bryan, M. Stack, K. Krysztofiak, U. Cichoń, D. G. Thomas, A. Surcel, E. S. Schiffhauer, M. A. Beckett, N. N. Khodarev and L. Xue, et al. , Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 22423–22429 Search PubMed.
- A. Koolivand and P. Dimitrakopoulos, Microfluid. Nanofluid., 2017, 21, 89 CrossRef.
- C. de Loubens, J. Deschamps, G. Boedec and M. Leonetti, J. Fluid Mech., 2015, 767, R3 CrossRef.
- X.-Y. Wang, A. Merlo, C. Dupont, A.-V. Salsac and D. Barthès-Biesel, Flow, 2021, 1, E8 CrossRef.
- D. Qin, Y. Xia and G. M. Whitesides, Nat. Protoc., 2010, 5, 491 CrossRef CAS PubMed.
- K. Phelan and K. M. May, Curr. Protoc. Cell Biol., 2015, 66, 1 Search PubMed.
- I. Goodfellow, Y. Bengio and A. Courville, Deep learning, MIT press, Cambridge, Massachusetts, 2016, pp. 185–191 Search PubMed.
- D. P. Kingma and J. Ba, Adam: A Method for Stochastic Optimization, 2017 Search PubMed.
- S. Ioffe and C. Szegedy, Proceedings of the 32nd international conference on machine learning, Lille, France, 2015, pp. 448–456 Search PubMed.
- N. Srivastava, G. Hinton, A. Krizhevsky, I. Sutskever and R. Salakhutdinov, J. Mach. Learn. Res., 2014, 15, 1929–1958 Search PubMed.
- S. Suzuki, et al. , Comput. Vis. Graphics Image Process., 1985, 30, 32–46 Search PubMed.
- O. Y. Zhong-Can and W. Helfrich, Phys. Rev. A: At., Mol., Opt. Phys., 1989, 39, 5280–5288 CrossRef PubMed.
- G. T. Charras, M. Coughlin, T. J. Mitchison and L. Mahadevan, Biophys. J., 2008, 94, 1836–1853 Search PubMed.
- Y. Sui, Y. T. Chew, P. Roy and H. T. Low, J. Comput. Phys., 2008, 227, 6351–6371 Search PubMed.
- Y. Sui, Y. T. Chew, P. Roy, Y. P. Cheng and H. T. Low, Phys. Fluids, 2008, 20, 112106 Search PubMed.
- Z. Wang, Y. Sui, A. V. Salsac, D. Barthès-Biesel and W. Wang, J. Fluid Mech., 2016, 806, 603–626 Search PubMed.
- Z. Wang, Y. Sui, A. V. Salsac, D. Barthès-Biesel and W. Wang, J. Fluid Mech., 2018, 849, 136–162 CrossRef CAS.
- R. X. Lu, Z. Wang, A. V. Salsac, D. Barthès-Biese, W. Wang and Y. Sui, J. Fluid Mech., 2021, 923, A11 Search PubMed.
- R. Lu, Z. Guo, P. Yu and Y. Sui, J. Fluid Mech., 2023, 962, A26 Search PubMed.
- S. Gabbanelli, G. Drazer and J. Koplik, Phys. Rev. E, 2005, 72, 046312 CrossRef PubMed.
- M. Bouzidi, M. Firdaouss and P. Lallemand, Phys. Fluids, 2001, 13, 3452–3459 CrossRef CAS.
- Z.-L. Guo, C.-G. Zheng and B.-C. Shi, Chin. Phys., 2002, 11, 366–374 Search PubMed.
- C. S. Peskin, Acta Numer., 2002, 11, 479–517 CrossRef.
- F. A. Amiri, G. Le, Q. Chen and J. Zhang, J. Comput. Phys., 2020, 423, 109800 Search PubMed.
- S. Ramanujan and C. Pozrikidis, J. Fluid Mech., 1998, 361, 117–143 Search PubMed.
- Y. You, X. Lai, Y. Pan, H. Zheng, J. Vera, S. Liu, S. Deng and L. Zhang, Signal Transduction Targeted Ther., 2022, 7, 156 Search PubMed.
- E. H. Houssein, A. Hammad and A. A. Ali, Neural Comput. Appl., 2022, 34, 12527–12557 Search PubMed.
- C. W. Olanow, R. L. Watts and W. C. Koller, Neurology, 2001, 56, S1–S88 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |