Open Access Article
Open Access ArticleAutonomous microfluidic labs: progress and prospects
Suyash
Damir
 a,
Fernando
Delgado-Licona
a,
Fernando
Delgado-Licona
 b,
Andrew
deMello
b,
Andrew
deMello
 *a and
Milad
Abolhasani
*a and
Milad
Abolhasani
 *b
*b
aDepartment of Chemistry and Applied Biosciences, Institute of Chemical and Bioengineering, ETH Zurich, Vladimir-Prelog-Weg 1, Zürich 8093, Switzerland. E-mail: andrew.demello@chem.ethz.ch
bDepartment of Chemical & Biomolecular Engineering, North Carolina State University, Raleigh, NC, USA. E-mail: abolhasani@ncsu.edu
First published on 17th February 2026
Abstract
Global challenges such as climate change, escalating energy demands, and health equity require new scientific innovations able to deliver timely solutions. Self-driving laboratories (SDLs) combine robotics and lab automation with artificial intelligence to efficiently explore complex experimental spaces, reduce human effort, and speed up discovery through intelligent experimentation. Central to this transformation is responsible research acceleration (RRA). This ensures that advances are reproducible, transparent, and resource-efficient, and lays the foundation for sustainable innovation. Microfluidics, with its precise control of heat and mass transfer rates, minimal reagent use, and seamless integration with real-time sensing and automation, represents an ideal platform to embody RRA principles within SDLs. This perspective explores the synergy between microfluidics and autonomous experimentation, highlights key challenges, and proposes strategies for fully autonomous microfluidic workflows. We argue that flow-based platforms are essential to expedite discovery and that stronger academia–industry collaboration is critical in shortening the path from scientific insight to real-world implementation and impact.
1. Introduction
Scientific and technological innovation is increasingly constrained not by possibility, but by time. Urgent global challenges, ranging from climate change and energy transition to healthcare, food security, and sustainable manufacturing, demand solutions on timescales that conventional research methods struggle to meet. Whilst the scientific community continues to produce groundbreaking concepts and insights, the time it takes to translate these ideas into tangible impacts remains a bottleneck. As the gap widens between the pace of global crises and the speed of conventional discovery pipelines, there is growing recognition that accelerating the research cycle is not just desirable, but necessary.1In this context, a new research paradigm is emerging, and centered on the integration of automation, artificial intelligence (AI), and closed-loop experimentation. At the core of this shift is the self-driving laboratory (SDL), which is a physical–digital platform that automates the iterative design–make–test–analyze (DMTA) cycle (Fig. 1). SDLs replace linear workflows with adaptive ones, where machine learning (ML) models guide the selection of experiments, interpret outcomes, and rapidly refine hypotheses. This approach enables researchers to explore vast design spaces systematically and efficiently, reducing the number of physical experiments needed and increasing the likelihood of identifying high-performing solutions.2
Importantly, SDLs do not replace human insight but rather extend it. Researchers define goals, constraints, and hypotheses, with the SDL executing experiments, analyzing results, and proposing new directions. This hybrid model of human–AI–robot collaboration is especially well-suited for use in domains where experimentation is expensive, slow, or multidimensional, such as materials discovery, catalysis, and pharmaceutical development. That said, as SDLs begin to reshape these fields, questions of trust, reproducibility, and equitable access become more pressing. Accelerating discovery must not come at the cost of rigor. To ensure this balance, the concept of responsible research acceleration (RRA) offers a guiding framework. Building on the principles of responsible AI ecosystems,3,4 RRA emphasizes transparency, reproducibility, sustainability, and equitable access across all parts of the DMTA cycle. In practice, this means documenting experimental decisions, reporting uncertainty, minimizing waste and energy use, and ensuring that the benefits of accelerated discovery are broadly distributed. Platforms that inherently support these values, by generating reliable data at scale with minimal resource consumption, are particularly valuable in the SDL landscape.5
Microfluidics represents one such enabling platform. By miniaturizing and compartmentalizing chemical and biological processes, microfluidic systems reduce reagent use, accelerate reaction times, and offer exquisite control over experimental conditions. Their intrinsic scalability and compatibility with automation make them a natural fit for closed-loop experimentation, minimizing variability and enhancing reliability. Moreover, the ability to multiplex experiments, integrate in-line sensing, and achieve high spatial and temporal resolution enables rich, multimodal data collection, which is an essential (but often ignored) ingredient for effective predictive ML modeling and optimization. Microfluidic platforms can thus serve as efficient engines for SDLs, supporting both rapid exploration and reproducibility.
This perspective article explores the intersection of microfluidics and autonomous experimentation, focusing on how recent advances are enabling the realization of fully autonomous microfluidic labs. We examine the current state of the field, identify unmet challenges, and highlight promising directions for research and deployment. We argue that microfluidics is not merely compatible with the SDL paradigm, but rather constitutes a central technology for realizing rapid, reproducible, and resource-efficient autonomous experimentation.
2. Microfluidics evolution
Microfluidics involves the manipulation of fluids at the micrometer scale, where fluid behavior differs fundamentally from that observed at larger length scales. At these dimensions, the governing physical forces shift: surface-to-volume ratios rise dramatically, inertial forces diminish, viscous and interfacial forces dominate, and flows become strictly laminar.6,7 Small-scale fluid manipulation eliminates turbulence and renders transport processes (especially mixing) diffusion-limited and deterministic. Such precise control over mass and heat transfer is crucial for conducting reproducible, high-fidelity chemical and biological experiments. Combined with minimal reagent consumption and the capacity for parallelization, microfluidics offers a compelling platform for combinatorial exploration, materials discovery, and process optimization under resource-constrained conditions.8The field of microfluidics emerged in the early 1980s from the need to enhance the sensitivity, reproducibility, and longevity of chemical sensors.9 As researchers recognized that miniaturization could reduce reagent usage and shorten analysis times, microfluidic systems evolved rapidly, giving rise to integrated lab-on-a-chip platforms (or micro total analysis systems, μTAS) in late 1990s.10,11 These platforms enabled the consolidation of laboratory functions such as sample preparation, reaction, detection, and analysis onto a single device, transforming workflows across disciplines.
Research in microfluidics has demonstrated substantial impact across chemistry,12–14 biology,15–19 and materials science.20–23 Perhaps the most transformative contribution has been in single-cell and single-molecule analysis,15,17,24–30 and nucleic acid amplification,31–36 where microfluidic compartmentalization underpins technologies such as digital polymerase chain reaction (PCR)37 and single-cell genomics,38 enabling measurements that are fundamentally inaccessible to bulk assays. Further, in chemical synthesis and materials discovery, microfluidic reactors have provided precise control over heat and mass transfer rates, enabling reproducible synthesis, access to transient reaction regimes, and high-throughput exploration of parameter spaces with drastically reduced material consumption.39–41 These capabilities establish microfluidics as a mature and distinct experimental modality, enabling the investigation of complex chemical and biological systems with a level of precision, throughput, and data richness that is difficult to achieve by conventional means.
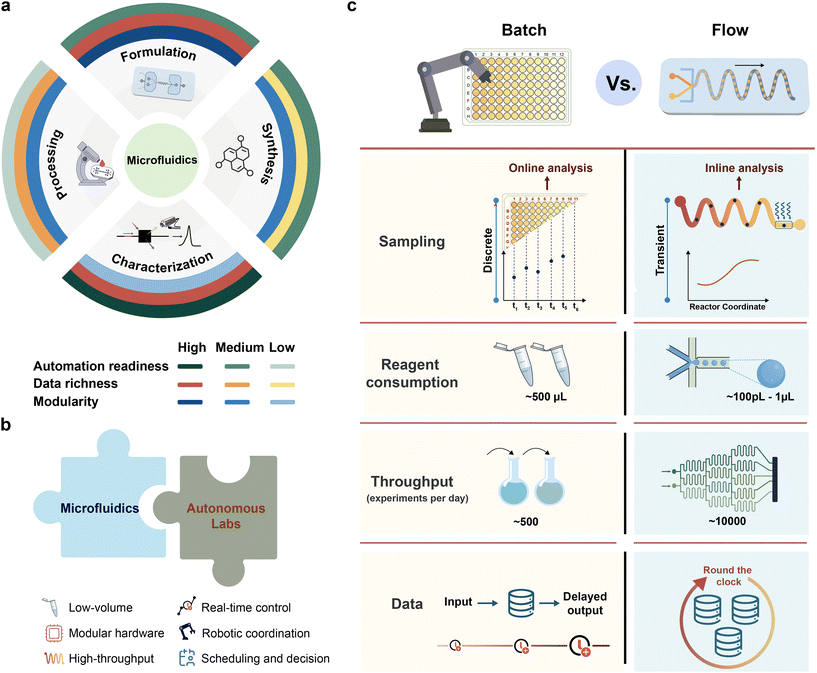
Despite the success of miniaturized workflows, monolithic μTAS platforms are often limited in terms of scalability. Their rigid architectures restrict configurational flexibility, making it difficult to iterate on designs or troubleshoot failures without extensive refabrication. In contrast, modular microfluidic systems, composed of discrete, task-specific units, offer enhanced flexibility, enabling rapid reconfiguration and easier integration with external automation (Table 1).42,43 Each component in such modular workflows typically serves one of four primary functions: (a) formulation and mixing with precise compositional control, (b) continuous or droplet-based synthesis, (c) in-line or on-line detection using integrated sensors, and (d) downstream processing such as separation, purification, sorting, and archiving (Fig. 2a).
| Features | μTAS | Modular microfluidic systems |
|---|---|---|
| Scalability | Low (exponentially high after design is fixed) | High (number of modules decide the limit) |
| Reconfigurability | Medium (rigid architecture) | High (discretized units) |
| Reproducibility | Excellent (with established microfabrication practices) | Good (for standardized modules), can vary with assembly |
| Automation | High (but less flexible) | High (dynamic automation via module integration) |
| Failure recovery | Poor (single point failure may require full replacement) | Excellent (swap individual modules) |
| Cost | Low (for large-scale commercial applications) | High upfront costs (decreasing with methods like 3D printing) |
This modularity proves particularly advantageous when investigating dynamic and multidimensional experimental landscapes. For example, enzymatic assays often require iterative optimization of substrate ratios, pH, temperature, and reaction time44—parameters that demand flexible experimental workflows capable of accommodating diverse assay formats and detection methods. While integrated μTAS platforms excel at executing defined protocols with high precision, their architectures are optimized for specific experimental workflows, making it challenging to adapt to fundamentally different assay requirements or incorporate new analytical techniques without substantial redesign effort.45 Similarly, in colloidal nanomaterial synthesis, where product quality is highly sensitive to flow profiles and mixing dynamics, modular platforms allow individual tuning of each stage while enabling real-time adjustments based on sensor feedback.46 In both cases, modular microfluidic systems reduce development time, enhance reproducibility, and allow selective upgrades or repairs without disrupting the entire workflow.
The convergence of microfluidics and SDLs is thus a natural progression. Modular microfluidic platforms satisfy key requirements for autonomous experimentation: precise control, scalability, and compatibility with real-time sensing and actuation.114 Reactions can be executed with high temporal and spatial resolution, and outcomes continuously measured, producing comprehensive, structured datasets that are ideally suited for ML. When combined with robotic liquid handling, programmable pumps, and digital control systems (Fig. 2b), these platforms move beyond static, miniaturized versions of traditional labs toward truly adaptive systems that learn from data and respond dynamically to new information.47,48
In this context, miniaturization is not simply shrinking workflows but a foundational technology that supports rapid iteration, efficient modular reconfigurability, and data-rich experimentation, features crucial when performing complex, multistep assays in fully autonomous SDLs.
3. Data-rich SDLs: moving towards flow
The choice between batch and flow processing fundamentally shapes the responsiveness, adaptability, and data richness of an SDL's experimental platform. (Table 2) Batch reactions, typically conducted in well plates, vials, or flasks, remain the mainstay of chemical and biological research due to their simplicity, ergonomics, familiarity, and compatibility with a wide range of assays. They benefit from decades of methodological development and can be automated using robotic liquid handlers. As a result, batch formats have become deeply embedded in early-stage screening and assay development workflows.49 However, these systems are inherently discrete and asynchronous; reactions are initiated and terminated in bulk, and measurements are often limited to endpoint analysis (Fig. 2c). This experimentation format restricts the ability to perform time-resolved experiments or the dynamic adjustment of conditions in response to real-time feedback, capabilities essential for closed-loop discovery.| Features | Modular microfluidic systems | Modular batch systems |
|---|---|---|
| a Dependent on system design and level of standardization. | ||
| Operating mode | Continuous (state depends on flow conditions) | Discrete (state progresses by scheduled steps) |
| Search-space | Ideal for continuous variables; can handle categorical changes with reagent switching modules | Ideal for discrete variables and broad formulation libraries |
| Experimental coordinate | Residence time/position | Experiment index (well/vial ID) and time |
| Data richness per unit material | Higha (continuous signals yield rich information) | Moderatea (can increase with repeated sampling) |
| Data latency | Low (time-resolved traces along reactor coordinate) | Moderatea (often endpoint or sparse time points) |
| Reconfigurability | High (discretized units, facile module integration) | Medium (discretized units, steep workflow integration) |
| Ease of protocol transfer | Higha (aided by standardized platform ecosystems/well-specified systems), custom builds are prevalent | Higha (aided by standardized platform ecosystems) |
| Cost | Medium (lower reagent use; maintenance on sub-modules) | High (higher consumables, i.e., tips and plates, CapEx depends on robots and modules) |
| Typical failure modes | Clogging, pump drift, sensor fouling/calibration; mitigated by inline monitoring and control logic | Mechanical drift, evaporation, liquid-handling variance, sensor calibration |
| Safety | Small internal volumes reduce hazards | Larger discrete volumes and open handling can increase exposure |
| Workflow diversity & complexity | Higha (if modularization is prioritized; multi-step trains possible) | High (excellent for discrete libraries and multi-step handling) |
| Cross-contamination | Effects can arise from fouling; managed by flushing, surface and secondary phase choices | Disposable consumables simplify isolation; managed by protocols |
| Residence time control | Direct and precise via flow rate and reactor volume | Indirect; set by protocol timing and mixing/incubation history |
| Material transport control | Transport engineered via channel geometry/flow regime; mixing is often predictable, reproducible, and model-informed | Handles slurries/solids; mixing governed by geometry/agitation; potential edge effects |
| Energy transport control | High surface-to-volume ratios enable enhanced thermal control; reduced characteristic lengths enable uniform energy flux | Thermal gradients are possible across the reactors/plates; slower heating/cooling rates |
Continuous flow systems, by contrast, enable a fundamentally different experimental architecture. In microfluidic reactors, operated either as segmented (droplet-based) or continuous flows, reaction conditions are controlled in both space and time. The spatial progression of fluid through the reactor maps directly onto reaction time, enabling kinetic resolution and fine temporal control without process interruption.50–52
A key consequence of using continuous systems is the seamless integration of flow reactors with real-time, in-line characterization techniques.53 Unlike batch workflows that rely on discrete sampling and aliquoting, in-line sensing allows continuous, non-invasive monitoring of reaction evolution, supporting the rapid acquisition of large, time-resolved datasets and access to transient phenomena that are impossible to capture off-line. Optical spectroscopy (including UV-vis and near infrared) is the most widely adopted technique in microfluidics, as it can be integrated readily via bespoke flow cells or fiber-optic probes and provide low-latency readouts of reaction outcome (concentration, conversion, and optical properties of compounds), making it particularly suitable for closed-loop optimization, where rapid feedback is essential.54–58
Complementary modalities extend this analytical space in microfluidic reactors. Specifically, infrared (IR) spectroscopy offers chemically distinct fingerprints of functional group transformations and is commonly implemented using in-line flow cells to mitigate substrate transparency and path-length constraints.59–62 Emission-based techniques such as fluorescence and photoluminescence provide high sensitivity at low concentrations and are particularly effective for nanomaterial synthesis and biochemical assays,63–65 though their use in autonomous workflows requires accounting for photobleaching, inner-filter effects, and temperature-dependent signal drift.66,67 Beyond optical methods, on-line mass spectrometry enables molecular-level insight into intermediates and reaction pathways, albeit with added latency from flow splitting and the need to synchronize residence time with data acquisition.68,69 Together with specialized approaches such as Raman,70 nuclear magnetic resonance (NMR),71,72 electron paramagnetic resonance (EPR),73 and X-ray spectroscopy,74,75 these techniques provide a versatile and dynamic characterization toolbox for multimodal sensing in autonomous microfluidic platforms, capabilities that remain difficult to replicate in batch-based autonomous workflows.
While these analytical tools enable data-rich experimentation, their effective deployment in autonomous microfluidic systems depends on more than just sensor availability. Reliable flow characterization often requires stable calibration over long experimental campaigns, resistance to fouling and bubble formation, and seamless integration with control software. As such, in-flow characterization in SDLs is best viewed as a co-designed component of the experimental workflow, rather than a modular add-on, with sensing, data handling, and decision-making tightly coupled. These features collectively support more adaptive, efficient, and informative experimentation, which are critical to SDLs.
Recent demonstrations highlight the performance and efficiency gains of microfluidic flow systems in autonomous research settings. In the AlphaFlow platform, for example, over 9000 distinct reaction conditions were explored using less than 30 μL of each reagent, achieving rapid optimization of quantum dot optical properties within 1000 hours, an outcome inaccessible to standard mini-batch workflows.39 Beyond throughput, the small reaction volumes (typically nanoliters to microliters) lead to dramatic reductions in material consumption and waste. In the case of atropine synthesis, transitioning from batch to flow reduced waste by nearly two orders of magnitude while improving product purity and throughput.76 A shift to continuous formats also reduces the dependence on single-use consumables and improves energy efficiency, offering clear advantages for sustainable and toxicity-sensitive applications.77
While continuous- and segmented-flow platforms are ideally suited for data-rich autonomous experimentation, the majority of existing SDLs rely on batch-based, macroscale formats. This reflects practical considerations in implementing microfluidic SDLs rather than fundamental limitations of continuous flow approaches.48 In particular, issues related to robustness, system integration, and product volumes obtained for post-processing become increasingly important when translating microfluidic concepts into autonomous workflows. Initial setup costs, including microreactors, pumps, and analytical modules, can also exceed those of traditional batch infrastructure.78 Additionally, as platforms scale to include multiple parallelized flow paths or downstream processes, engineering complexity increases, demanding reliable integration strategies and modular design principles.79,80
Nonetheless, batch and microfluidic-based SDLs address different practical needs within modular experimentation workflows. Robotic, batch-based platforms are particularly effective when the DMTA cycle relies on decoupled, multi-step procedures involving solid and heterogeneous mixtures, where high-dimensional variables are explored as discrete, programmable and audited protocols. Commercial robotic/batch ecosystems facilitate protocol transfer due to their commercial maturity and widespread adoption. However, the quantity and reliability of the resulting datasets can be limited in part by more constrained engineered controls over material and energy transport, and by the complexity of integrating in-process characterization, which often results in endpoint readouts with limited temporal sampling. These limitations are frequently compounded by a higher cost per datapoint, driven by material consumption and reliance on single-use consumables. If the process is amenable to flow (i.e., no significant solid precursors) the choice for microfluidic modules allows finely engineered control over transport phenomena, enabling reproducible, time-resolved measurements and seamless integration of in-line analytics under tightly defined reaction conditions. Such capabilities directly support closed-loop, adaptive discovery by supporting reproducible, high-fidelity measurements, albeit with increased demands on system robustness, integration, and modular standardization.113 Moving forward, integrating microfluidic modules into SDL architectures offers a practical route to scaling data quality, control, and autonomy while complementing batch-based scientific workflows.
4. Challenges and perspectives
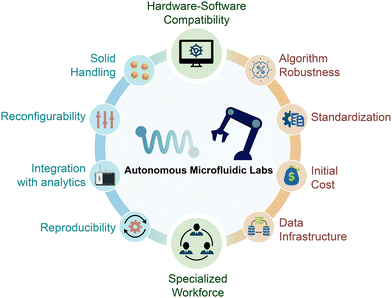
While microfluidic technologies offer clear advantages for autonomous experimentation, several barriers must be addressed to realize their full potential in SDLs (Fig. 3). Realizing autonomous microfluidic laboratories require the integration of sensing, automation, and intelligent control, as well as a careful consideration of the underlying reaction modality. In modular microfluidic workflows, a persistent challenge is the handling of solid particulates, either introduced as reagents or formed in situ during synthesis. This is particularly relevant in reactions involving heterogeneous catalysis, crystallization, or nanoparticle formation, where solid phases are intrinsic to the chemistry.20 At the microscale, even minor aggregation or precipitation events can obstruct narrow channels, disrupt flow profiles, and compromise sensor accuracy. These fouling events degrade performance and may also generate misleading data that propagate through the closed-loop system, undermining optimization and reproducibility.Studies from industrial settings underscore the scale of this issue. For instance, a seminal assessment by Lonza indicated that more than 60% of reactions amenable to continuous processing were rendered impractical on the microscale due to solid-related complications.40 Perovskite nanocrystal synthesis serves as a representative example, where shifts in precursor concentration or temperature can induce premature nucleation, leading to salt precipitation and channel blockage. Without reliable fouling detection and compensation strategies, such events introduce errors into the data stream that can misguide decision-making algorithms and compromise the integrity of subsequent iterations. In practice, fouling detection can be realized through multiple modalities. Optical microscopy remains the most widely used approach, with bright-field and fluorescence imaging enabling real-time visualization of aggregation, deposition, and adsorption in microchannels.81–84 Complementary techniques discussed for in-line characterization, such as Raman (including surface-enhanced Raman) spectroscopy,85 IR spectroscopy,86 NMR imaging,87,88 and X-ray methods89 have also been employed to probe early-stage fouling and particle composition without labeling, albeit with higher integration complexity. Beyond direct imaging, indirect indicators such as real-time knowledge analysis of pressure-drops across reactor length or deviations in volumetric flow rate under constant actuation provide simple, robust signatures of fouling or blockage that are well suited for autonomous operation.
Addressing challenges related to solid aggregation requires a combination of physical, chemical, and algorithmic solutions. Passive strategies include surface modifications to reduce particle adhesion90 and hydrodynamic control schemes that keep solids away from channel walls. Active techniques such as acoustic streaming,91 magnetic field manipulation, and electrophoretic separation92 enable dynamic control of solids within microchannels. However, these methods remain underutilized in autonomous workflows and often lack generalizability across chemistries. Incorporating redundancy into system architecture, such as parallelized flow paths or self-cleaning loops, can also increase robustness by enabling fault isolation and recovery. Furthermore, integrating in-line imaging or pressure-based sensing could allow SDLs to autonomously detect clogging and adapt protocols in real time, thereby preserving data quality and experimental continuity.
A second major challenge is the substantial upfront costs and specialized expertise required to establish the SDL infrastructure. Off-the-shelf automation systems can exceed 0.1–0.5 M USD, creating steep entry barriers that often concentrate operational knowledge among a few trained users.93 To save time and cost, labs resort to bespoke solutions that function locally but may lack the reproducibility and transferability needed for industrial adoption.
This is compounded by another barrier: integration of modular microreactor components with automated hardware and data infrastructure. While individual instruments (e.g., pumps, reactors, and sensors) have matured considerably, they are rarely designed for seamless machine-to-machine communication. Most commercial tools are optimized for manual operation, often requiring proprietary software interfaces, limiting interoperability. For SDLs to function autonomously and scalably, experimental components must operate as part of an orchestrated ecosystem capable of sharing data, triggering actions, and responding to sensor feedback in real time. Currently, this orchestration is largely implemented through ad hoc workarounds. For example, synchronizing a commercial syringe pump with an in-line spectrometer often demands microcontroller-based intermediaries, reverse-engineered communication protocols, or custom Python scripts emulating manual inputs.48 These solutions, while functional, are fragile and difficult to maintain, hindering reproducibility and slowing deployment and widespread adoption.
Moving forward, two complementary strategies have the potential to drastically accelerate progress. First, democratizing access through open-source hardware (e.g., FINDUS, OTTO) and standardized software stacks, supported by community-driven training, can reduce both financial and knowledge barriers.48,94,95 Second, the research community would benefit from the development and adoption of hardware-agnostic communication standards tailored to automated laboratories, analogous to the OPC-UA protocol in industrial automation or SCPI in instrumentation control. Initiatives like SiLA 2, combined with open data standards like AniML represent important steps toward this goal.96 Broader adoption of such frameworks, coupled with efforts to design SDL-compatible instrumentation with standardized application programming interfaces (APIs) are likely to significantly reduce integration overheads, particularly in academic and early-stage applications where resources are most constrained.
Beyond technical integration, establishing unified data structures for capturing experimental metadata, instrument settings, and sensor outputs is critical. Such a unified data structure ensures interoperability across devices and also supports long-term goals in ML, benchmarking, and collaboration. Emerging efforts toward “FAIR” (findable, accessible, interoperable, reusable) data standards in autonomous experimentation provide a promising direction and should be extended to microfluidic contexts, where the richness of spatial–temporal data is especially valuable.97
While the majority of the SDL development has focused on chemical synthesis and materials science, significant opportunities exist in expanding these capabilities to biological systems. Applications such as single-cell genomics, where microfluidics enables the isolation and analysis of individual cells,18,24,29 or genotype–phenotype linkage studies that track cellular responses under controlled microenvironments,98,99 represent particularly promising directions for autonomous experimentation. Notably, several widely adopted commercial platforms illustrate the practical maturity of microfluidics in this domain. Commercially available digital PCR systems, which partition samples into millions of isolated volumes for absolute nucleic acid quantification, have become a diagnostic gold standard in the last decade.100,101 Similarly, droplet-based microfluidic workflows revolutionized single-cell analysis, fundamentally advancing our understanding of cellular heterogeneity in cancer and immunology.102–104 These successes underscore how microfluidics can support scalable, standardized, and data-intensive biological workflows. Translating such capabilities into autonomous microfluidic SDLs will require advances in integrated biological sensing, sterile and robust fluid handling, and biocompatible system design, offering a compelling pathway toward accelerating discovery in synthetic biology and personalized medicine.105
Beyond microscale optimization, an important consideration for autonomous microfluidic laboratories is the translation of knowledge to larger-scale processes. While scaling microreactors through numbering-up or parallelization has often been proposed,106 it remains a nontrivial challenge in practice. Instead, microfluidic SDLs are best viewed as discovery engines that generate transferable kinetic and mechanistic insights under well-defined conditions.107
The precise control over transport phenomena in microreactors enables the systematic mapping of reaction performance through dimensionless variables, such as the Damköhler number for reaction-transport coupling, the Péclet number for mixing efficiency, and the Biot number for heat transfer dynamics.108 Importantly, the degree to which these parameters must be preserved during scale-up depends on the controlling regime. For chemistries that are kinetically-limited (Da ≪ 1), translation to larger scales primarily requires maintaining practical homogeneity and thermal control rather than matching mixing-related dimensionless variables exactly. In contrast, when reactions outpace mixing (Da > 1) observed rates and selectivity can become functions of mass transfer; operating in this regime can disrupt kinetics, making transport similarity (and the associated dimensionless variables) a primary scale-up target for preserving performance.109 Additionally, microfluidic platforms rapidly identify process boundaries, such as precipitation thresholds, thermal runaway conditions, or selectivity changes, that might take months to map using traditional batch screening.110 These fundamental insights, captured as dimensionless correlations and operating windows, guide the design of production-scale reactors where transport limitations differ significantly from microscale conditions.
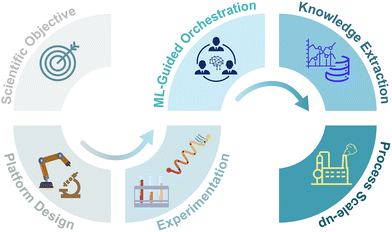
Looking forward, the ultimate vision for microfluidic SDLs extends beyond individual experimental optimization to encompass end-to-end workflow automation, from application selection and device design to implementation and scale-up (Fig. 4). Such automation would leverage ML algorithms not only to optimize experimental conditions but also to guide the selection of appropriate microfluidic architectures, predict optimal reactor materials, and even suggest novel reaction pathways based on accumulated experimental data.111,112 This systems-level approach would democratize advanced chemical and biological research by making elaborate experimental capabilities accessible to researchers without specialized microfluidics expertise.
In sum, advancing autonomous microfluidic labs requires the confrontation of both chemical and infrastructural challenges. Solutions will likely emerge at the intersection of hardware innovation, software interoperability, and systems-level thinking. The development of robust solid-handling strategies, standardized communication protocols, and open-source control frameworks will be central to this effort. As these components mature, microfluidic SDLs are poised to become more capable and resilient while improving accesibility to the broader scientific community, ushering in a new era of scalable, data-rich, and intelligent experimentation.
Conclusions
The convergence of microfluidics and SDLs signals a shift toward research infrastructure designed for faster, more reproducible, and more sustainable practices. While these systems promise to deliver high-quality data at reduced material and energy costs, their adoption will depend on addressing persistent barriers such as solid handling, development of modular components, and standardized workflow automation. Progress will require community-wide efforts to establish interoperable standards, reducing the barriers through open-source and modular designs. In parallel, advancing the digitalization of microfluidic platforms, including their design, simulation, and integration with virtual twins, represents a critical next step to enable predictive, scalable, and fully autonomous experimentation. By embedding principles of responsible research acceleration into every stage of SDLs development, autonomous microfluidic labs can evolve into reliable platforms that accelerate the discovery timescales while ensuring that the benefits of innovation are distributed equitably and sustainably.Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting the findings of this study is available within the main text.Acknowledgements
F. D. L. and M. A. gratefully acknowledge the financial support from the National Science Foundation (award no. 1940959, 2315996 and 2420490) and the University of North Carolina Research Opportunities Initiative (UNC-ROI) program. S. D. and A. dM. gratefully acknowledge the Swiss National Science Foundation for financial support through NCCR Catalysis (grant number 225147), a National Centre of Competence in Research.References
- M. Abolhasani and E. Kumacheva, Nat. Synth., 2023, 2, 483–492 CrossRef.
- J. Schrier, A. J. Norquist, T. Buonassisi and J. Brgoch, J. Am. Chem. Soc., 2023, 145, 21699–21716 CrossRef PubMed.
- B. C. Stahl, Sci. Rep., 2023, 13, 7586 CrossRef PubMed.
- E. Papagiannidis, P. Mikalef and K. Conboy, J. Strateg. Inf. Syst., 2025, 34, 101885 CrossRef.
- F. Delgado-Licona, D. Addington, A. Alsaiari and M. Abolhasani, Nat. Chem. Eng., 2025, 2, 277–280 CrossRef.
- T. M. Squires and S. R. Quake, Rev. Mod. Phys., 2005, 77, 977–1026 CrossRef.
- A. J. deMello, Nature, 2006, 442, 394–402 CrossRef PubMed.
- S. S. Damir, J. Probst, A. deMello and S. Stavrakis, Chem.:Methods, 2025, 202500069 Search PubMed.
- D. R. Reyes, D. Iossifidis, P.-A. Auroux and A. Manz, Anal. Chem., 2002, 74, 2623–2636 CrossRef PubMed.
- A. Manz, N. Graber and H. Widmer, Sens. Actuators, B, 1990, 1, 244–248 CrossRef.
- A. R. Wheeler, W. R. Throndset, R. J. Whelan, A. M. Leach, R. N. Zare, Y. H. Liao, K. Farrell, I. D. Manger and A. Daridon, Anal. Chem., 2003, 75, 3581–3586 CrossRef PubMed.
- A. Abou-Hassan, O. Sandre and V. Cabuil, Angew. Chem., Int. Ed., 2010, 49, 6268–6286 Search PubMed.
- L. Bezinge, R. M. Maceiczyk, I. Lignos, M. V. Kovalenko and A. J. deMello, ACS Appl. Mater. Interfaces, 2018, 10, 18869–18878 Search PubMed.
- J. M. Granda, L. Donina, V. Dragone, D.-L. Long and L. Cronin, Nature, 2018, 559, 377–381 CrossRef PubMed.
- C. Zhang and D. Xing, Chem. Rev., 2010, 110, 4910–4947 Search PubMed.
- Z. Yu, S. Lu and Y. Huang, Anal. Chem., 2014, 86, 9386–9390 Search PubMed.
- D. A. Cusanovich, R. Daza, A. Adey, H. A. Pliner, L. Christiansen, K. L. Gunderson, F. J. Steemers, C. Trapnell and J. Shendure, Science, 2015, 348, 910–914 Search PubMed.
- A. Isozaki, H. Mikami, H. Tezuka, H. Matsumura, K. Huang, M. Akamine, K. Hiramatsu, T. Iino, T. Ito, H. Karakawa, Y. Kasai, Y. Li, Y. Nakagawa, S. Ohnuki, T. Ota, Y. Qian, S. Sakuma, T. Sekiya, Y. Shirasaki, N. Suzuki, E. Tayyabi, T. Wakamiya, M. Xu, M. Yamagishi, H. Yan, Q. Yu, S. Yan, D. Yuan, W. Zhang, Y. Zhao, F. Arai, R. E. Campbell, C. Danelon, D. Di Carlo, K. Hiraki, Y. Hoshino, Y. Hosokawa, M. Inaba, A. Nakagawa, Y. Ohya, M. Oikawa, S. Uemura, Y. Ozeki, T. Sugimura, N. Nitta and K. Goda, Lab Chip, 2020, 20, 2263–2273 Search PubMed.
- E. Biselli, E. Agliari, A. Barra, F. R. Bertani, A. Gerardino, A. De Ninno, A. Mencattini, D. Di Giuseppe, F. Mattei, G. Schiavoni, V. Lucarini, E. Vacchelli, G. Kroemer, C. Di Natale, E. Martinelli and L. Businaro, Sci. Rep., 2017, 7, 12737 Search PubMed.
- S. Krishnadasan, R. J. C. Brown, A. J. deMello and J. C. deMello, Lab Chip, 2007, 7, 1434 Search PubMed.
- R. W. Epps, M. S. Bowen, A. A. Volk, K. Abdel-Latif, S. Han, K. G. Reyes, A. Amassian and M. Abolhasani, Adv. Mater., 2020, 32, 2001626 Search PubMed.
- K. Abdel-Latif, R. W. Epps, F. Bateni, S. Han, K. G. Reyes and M. Abolhasani, Adv. Intell. Syst., 2021, 3, 2000245 Search PubMed.
- J.-M. Lu, H.-F. Wang, Q.-H. Guo, J.-W. Wang, T.-T. Li, K.-X. Chen, M.-T. Zhang, J.-B. Chen, Q.-N. Shi, Y. Huang, S.-W. Shi, G.-Y. Chen, J.-Z. Pan, Z. Lu and Q. Fang, Nat. Commun., 2024, 15, 8826 Search PubMed.
- A. Huebner, M. Srisa-Art, D. Holt, C. Abell, F. Hollfelder, A. J. deMello and J. B. Edel, Chem. Commun., 2007, 1218–1220 Search PubMed.
- F. S. O. Fritzsch, K. Rosenthal, A. Kampert, S. Howitz, C. Dusny, L. M. Blank and A. Schmid, Lab Chip, 2013, 13, 397–408 Search PubMed.
- K.-W. Huang, Y.-C. Wu, J.-A. Lee and P.-Y. Chiou, Lab Chip, 2013, 13, 3721–3727 Search PubMed.
- A. Liu, W. Liu, Y. Wang, J.-C. Wang, Q. Tu, R. Liu, J. Xu, S. Shen and J. Wang, Microfluid. Nanofluid., 2013, 14, 515–526 Search PubMed.
- M. Zhao, P. G. Schiro, J. S. Kuo, K. M. Koehler, D. E. Sabath, V. Popov, Q. Feng and D. T. Chiu, Anal. Chem., 2013, 85, 2465–2471 Search PubMed.
- S. Köster, F. E. Angilè, H. Duan, J. J. Agresti, A. Wintner, C. Schmitz, A. C. Rowat, C. A. Merten, D. Pisignano, A. D. Griffiths and D. A. Weitz, Lab Chip, 2008, 8, 1110 Search PubMed.
- R. Mohammadi, I. Sakic, A. Jain, P. Mathur, T. Weiss, A. J. deMello, S. Stavrakis and M. Asghari, Small, 2025, 21, e07981 Search PubMed.
- A. T. Woolley, D. Hadley, P. Landre, A. J. deMello, R. A. Mathies and M. A. Northrup, Anal. Chem., 1996, 68, 4081–4086 Search PubMed.
- A. J. deMello, Nature, 2003, 422, 28–29 Search PubMed.
- J. Reboud, Y. Bourquin, R. Wilson, G. S. Pall, M. Jiwaji, A. R. Pitt, A. Graham, A. P. Waters and J. M. Cooper, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 15162–15167 Search PubMed.
- J. A. Lounsbury, A. Karlsson, D. C. Miranian, S. M. Cronk, D. A. Nelson, J. Li, D. M. Haverstick, P. Kinnon, D. J. Saul and J. P. Landers, Lab Chip, 2013, 13, 1384–1393 Search PubMed.
- Q. Zhu, Y. Gao, B. Yu, H. Ren, L. Qiu, S. Han, W. Jin, Q. Jin and Y. Mu, Lab Chip, 2012, 12, 4755–4763 Search PubMed.
- S. Petralia, R. Verardo, E. Klaric, S. Cavallaro, E. Alessi and C. Schneider, Sens. Actuators, B, 2013, 187, 99–105 Search PubMed.
- H. Wu, X. Cao, Y. Meng, D. Richards, J. Wu, Z. Ye and A. J. deMello, Biosens. Bioelectron., 2022, 211, 114377 Search PubMed.
- N. Shembekar, C. Chaipan, R. Utharala and C. A. Merten, Lab Chip, 2016, 16, 1314–1331 Search PubMed.
- A. A. Volk, R. W. Epps, D. T. Yonemoto, B. S. Masters, F. N. Castellano, K. G. Reyes and M. Abolhasani, Nat. Commun., 2023, 14, 1403 Search PubMed.
- D. M. Roberge, L. Ducry, N. Bieler, P. Cretton and B. Zimmermann, Chem. Eng. Technol., 2005, 28, 318–323 Search PubMed.
- J. P. McMullen and K. F. Jensen, Annu. Rev. Anal. Chem., 2010, 3, 19–42 Search PubMed.
- J. Wu, H. Fang, J. Zhang and S. Yan, J. Nanobiotechnol., 2023, 21, 85 Search PubMed.
- Y.-Q. Fan, H.-L. Wang, K.-X. Gao, J.-J. Liu, D.-P. Chai and Y. J. Zhang, Chin. J. Anal. Chem., 2018, 46, 1863–1871 Search PubMed.
- J. Wang, Electrophoresis, 2002, 23, 713–718 Search PubMed.
- Á. Á. Ríos and M. Zougagh, TrAC, Trends Anal. Chem., 2015, 69, 105–113 Search PubMed.
- A. A. Volk, R. W. Epps and M. Abolhasani, Adv. Mater., 2021, 33, 2004495 Search PubMed.
- E. A. Galan, H. Zhao, X. Wang, Q. Dai, W. T. S. Huck and S. Ma, Matter, 2020, 3, 1893–1922 Search PubMed.
- G. Tom, S. P. Schmid, S. G. Baird, Y. Cao, K. Darvish, H. Hao, S. Lo, S. Pablo-García, E. M. Rajaonson, M. Skreta, N. Yoshikawa, S. Corapi, G. D. Akkoc, F. Strieth-Kalthoff, M. Seifrid and A. Aspuru-Guzik, Chem. Rev., 2024, 124, 9633–9732 Search PubMed.
- C. Holtze and R. Boehling, Curr. Opin. Chem. Eng., 2022, 36, 100798 Search PubMed.
- J. Wegner, S. Ceylan and A. Kirschning, Adv. Synth. Catal., 2012, 354, 17–57 Search PubMed.
- D. Hess, A. Rane, A. J. deMello and S. Stavrakis, Anal. Chem., 2015, 87, 4965–4972 Search PubMed.
- D. Hess, V. Dockalova, P. Kokkonen, D. Bednar, J. Damborsky, A. deMello, Z. Prokop and S. Stavrakis, Chem, 2021, 7, 1066–1079 Search PubMed.
- N. Mukhin, P. Jha and M. Abolhasani, Matter, 2025, 8, 102205 Search PubMed.
- J. Probst, P. Howes, P. Arosio, S. Stavrakis and A. deMello, Anal. Chem., 2021, 93, 7673–7681 Search PubMed.
- I. Lignos, V. Morad, Y. Shynkarenko, C. Bernasconi, R. M. Maceiczyk, L. Protesescu, F. Bertolotti, S. Kumar, S. T. Ochsenbein, N. Masciocchi, A. Guagliardi, C.-J. Shih, M. I. Bodnarchuk, A. J. deMello and M. V. Kovalenko, ACS Nano, 2018, 12, 5504–5517 Search PubMed.
- W. Ferstl, T. Klahn, W. Schweikert, G. Billeb, M. Schwarzer and S. Loebbecke, Chem. Eng. Technol., 2007, 30, 370–378 Search PubMed.
- J. P. McMullen and K. F. Jensen, Org. Process Res. Dev., 2011, 15, 398–407 Search PubMed.
- R. W. Epps, A. A. Volk, K. Abdel-Latif and M. Abolhasani, React. Chem. Eng., 2020, 5, 1212–1217 Search PubMed.
- P. Hinsmann, M. Haberkorn, J. Frank, P. Svasek, M. Harasek and B. Lendl, Appl. Spectrosc., 2001, 55, 241–251 Search PubMed.
- K. L. A. Chan, S. Gulati, J. B. Edel, A. J. deMello and S. G. Kazarian, Lab Chip, 2009, 9, 2909–2913 Search PubMed.
- C. F. Carter, H. Lange, S. V. Ley, I. R. Baxendale, B. Wittkamp, J. G. Goode and N. L. Gaunt, Org. Process Res. Dev., 2010, 14, 393–404 Search PubMed.
- J. S. Moore and K. F. Jensen, Org. Process Res. Dev., 2012, 16, 1409–1415 Search PubMed.
- I. Lignos, S. Stavrakis, G. Nedelcu, L. Protesescu, A. J. deMello and M. V. Kovalenko, Nano Lett., 2016, 16, 1869–1877 Search PubMed.
- I. Lignos, R. M. Maceiczyk, M. V. Kovalenko and S. Stavrakis, Chem. Mater., 2020, 32, 27–37 Search PubMed.
- H. Nagai, N. Miwa, M. Segawa, S.-I. Wakida and K. Chayama, J. Appl. Phys., 2009, 105, 102015 Search PubMed.
- J. Zhang, R. Zhou, D. Tang, X. Hou and P. Wu, TrAC, Trends Anal. Chem., 2019, 110, 183–190 Search PubMed.
- T. Wang, L.-H. Zeng and D.-L. Li, Appl. Spectrosc. Rev., 2017, 52, 883–908 Search PubMed.
- M. C. Mitchell, V. Spikmans, A. Manz and A. J. deMello, J. Chem. Soc., Perkin Trans. 1, 2001, 514–518 Search PubMed.
- L. Fidalgo, G. Whyte, B. Ruotolo, J. Benesch, F. Stengel, C. Abell, C. Robinson and W. Huck, Angew. Chem., Int. Ed., 2009, 48, 3665–3668 Search PubMed.
- S. Schwolow, F. Braun, M. Rädle, N. Kockmann and T. Röder, Org. Process Res. Dev., 2015, 19, 1286–1292 Search PubMed.
- V. Sans, L. Porwol, V. Dragone and L. Cronin, Chem. Sci., 2015, 6, 1258–1264 Search PubMed.
- Y. Takahashi, M. Nakakoshi, S. Sakurai, Y. Akiyama, H. Suematsu, H. Utsumi and T. Kitamori, Anal. Sci., 2007, 23, 395–400 Search PubMed.
- T. Moragues, M. Agrachev, S. Mitchell, G. Jeschke, J. Pérez-Ramírez and A. J. deMello, Small Methods, 2025, 9, 2401771 Search PubMed.
- J. Probst, C. N. Borca, M. A. Newton, J. Van Bokhoven, T. Huthwelker, S. Stavrakis and A. deMello, ACS Meas. Sci. Au, 2021, 1, 27–34 Search PubMed.
- T. Moragues, G. Giannakakis, A. Ruiz-Ferrando, C. N. Borca, T. Huthwelker, A. Bugaev, A. J. deMello, J. Pérez-Ramírez and S. Mitchell, Angew. Chem., Int. Ed., 2024, 63, e202401056 Search PubMed.
- A.-C. Bédard, A. R. Longstreet, J. Britton, Y. Wang, H. Moriguchi, R. W. Hicklin, W. H. Green and T. F. Jamison, Bioorg. Med. Chem., 2017, 25, 6233–6241 Search PubMed.
- D. Dallinger and C. O. Kappe, Curr. Opin. Green Sustainable Chem., 2017, 7, 6–12 Search PubMed.
- R. L. Hartman, Curr. Opin. Chem. Eng., 2020, 29, 42–50 Search PubMed.
- B. Mosadegh, T. Bersano-Begey, J. Y. Park, M. A. Burns and S. Takayama, Lab Chip, 2011, 11, 2813–2818 Search PubMed.
- F. Su, K. Chakrabarty and R. Fair, 'IEEE Trans. Comput.-Aided Des. Integr. Circuits Syst., 2006, 25, 211–223 Search PubMed.
- A. Perazzo, L. Sicignano, G. Tomaiuolo, R. Marotta, R. Andreozzi and S. Guido, Chem. Eng. Sci., 2019, 207, 581–587 Search PubMed.
- L. Sicignano, G. Tomaiuolo, A. Perazzo, S. P. Nolan, P. L. Maffettone and S. Guido, Chem. Eng. J., 2018, 341, 639–647 Search PubMed.
- K. B. Neeves and S. L. Diamond, Lab Chip, 2008, 8, 701–709 Search PubMed.
- C. Güell, M. Ferrando and F. López, Monitoring and Visualizing Membrane-Based Processes, John Wiley & Sons, 2009 Search PubMed.
- L. Cui, M. Yao, B. Ren and K.-S. Zhang, Anal. Chem., 2011, 83, 1709–1716 Search PubMed.
- S. G. Kazarian, Anal. Bioanal. Chem., 2007, 388, 529–532 Search PubMed.
- F. Arndt, S. Schuhmann, G. Guthausen, S. Schütz and H. Nirschl, J. Membr. Sci., 2017, 524, 691–699 Search PubMed.
- S. Buetehorn, L. Utiu, M. Küppers, B. Blümich, T. Wintgens, M. Wessling and T. Melin, J. Membr. Sci., 2011, 371, 52–64 Search PubMed.
- K. Park, J. H. Kim, B. J. Kim, S. J. Cho, J. Hong and G. Lim, Langmuir, 2020, 36, 10548–10554 Search PubMed.
- M. Ashokkumar and T. J. Mason, Kirk-Othmer Encyclopedia of Chemical Technology, John Wiley & Sons, Ltd, 2007 Search PubMed.
- F. Wang, C. H. Chon and D. Li, J. Colloid Interface Sci., 2010, 352, 580–584 Search PubMed.
- B. S. Flowers and R. L. Hartman, Challenges, 2012, 3, 194–211 Search PubMed.
- H. Tegally, J. E. San, J. Giandhari and T. de Oliveira, BMC Genomics, 2020, 21, 729 Search PubMed.
- F. Barthels, U. Barthels, M. Schwickert and T. Schirmeister, SLAS Technol., 2020, 25, 190–199 Search PubMed.
- D. C. Florian, M. Odziomek, C. L. Ock, H. Chen and S. A. Guelcher, Sci. Rep., 2020, 10, 13663 Search PubMed.
- N. Hillson, M. Caddick, Y. Cai, J. A. Carrasco, M. W. Chang, N. C. Curach, D. J. Bell, R. Le Feuvre, D. C. Friedman, X. Fu, N. D. Gold, M. J. Herrgård, M. B. Holowko, J. R. Johnson, R. A. Johnson, J. D. Keasling, R. I. Kitney, A. Kondo, C. Liu, V. J. J. Martin, F. Menolascina, C. Ogino, N. J. Patron, M. Pavan, C. L. Poh, I. S. Pretorius, S. J. Rosser, N. S. Scrutton, M. Storch, H. Tekotte, E. Travnik, C. E. Vickers, W. S. Yew, Y. Yuan, H. Zhao and P. S. Freemont, Nat. Commun., 2019, 10, 2040 Search PubMed.
- R. B. Canty, J. A. Bennett, K. A. Brown, T. Buonassisi, S. V. Kalinin, J. R. Kitchin, B. Maruyama, R. G. Moore, J. Schrier, M. Seifrid, S. Sun, T. Vegge and M. Abolhasani, Nat. Commun., 2025, 16, 3856 Search PubMed.
- A. Mencattini, D. Di Giuseppe, M. C. Comes, P. Casti, F. Corsi, F. R. Bertani, L. Ghibelli, L. Businaro, C. Di Natale, M. C. Parrini and E. Martinelli, Sci. Rep., 2020, 10, 7653 Search PubMed.
- B. Poschkamp and S. Bekeschus, Cytometry, Part A, 2024, 105, 356–367 Search PubMed.
- K. A. Heyries, C. Tropini, M. VanInsberghe, C. Doolin, O. I. Petriv, A. Singhal, K. Leung, C. B. Hughesman and C. L. Hansen, Nat. Methods, 2011, 8, 649–651 Search PubMed.
- B. M. Paegel, R. G. Blazej and R. A. Mathies, Curr. Opin. Biotechnol., 2003, 14, 42–50 Search PubMed.
- P. See, J. Lum, J. Chen and F. Ginhoux, Front. Immunol., 2018, 9, year Search PubMed.
- X. Wang, Y. He, Q. Zhang, X. Ren and Z. Zhang, Genomics, Proteomics Bioinf., 2021, 19, 253–266 Search PubMed.
- A. J. Black, E. Wiseman, P. Wang, D. Wilson, F. Torres and J. Modarelli, Cancer Res., 2023, 83, 241–241 Search PubMed.
- A. Stephenson, L. Lastra, B. Nguyen, Y.-J. Chen, J. Nivala, L. Ceze and K. Strauss, ACS Synth. Biol., 2023, 12, 3156–3169 Search PubMed.
- A. M. Nightingale, J. H. Bannock, S. H. Krishnadasan, F. T. F. O'Mahony, S. A. Haque, J. Sloan, C. Drury, R. McIntyre and J. C. deMello, J. Mater. Chem. A, 2013, 1, 4067 Search PubMed.
- S. Battat, D. A. Weitz and G. M. Whitesides, Lab Chip, 2022, 22, 530–536 Search PubMed.
- M. C. Ruzicka, Chem. Eng. Res. Des., 2008, 86, 835–868 Search PubMed.
- J. H. A. Schuurmans, S. D. A. Zondag, A. Chaudhuri, J. Van Der Schaaf and T. Noël, ACS Cent. Sci., 2026, acscentsci.5c01825 Search PubMed.
- J. Zhang, K. Wang, A. R. Teixeira, K. F. Jensen and G. Luo, Annu. Rev. Chem. Biomol. Eng., 2017, 8, 285–305 Search PubMed.
- E. E. Tsur, Annu. Rev. Biomed. Eng., 2020, 22, 285–307 Search PubMed.
- D. McIntyre, A. Lashkaripour, P. Fordyce and D. Densmore, Lab Chip, 2022, 22, 2925–2937 Search PubMed.
- R. B. Canty and M. Abolhasani, Nat. Synth., 2024, 3, 1327–1339 Search PubMed.
- A. V. Volk, Z. S. Campbell, M. Y. S. Ibrahim, J. A. Bennett and M. Abolhasani, Annu. Rev. Chem. Biomol. Eng., 2022, 13, 45–72 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |