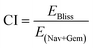Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
CombiCTx: screening diffusion gradients of anti-cancer drug combinations
Christina
Stelzl
 ab,
Ada
Lerma-Clavero
a,
Selina
Camenisch
abc,
Benoit
Simon
ab,
Olle
Eriksson
ab,
Oliver
Degerstedt
ab,
Ada
Lerma-Clavero
a,
Selina
Camenisch
abc,
Benoit
Simon
ab,
Olle
Eriksson
ab,
Oliver
Degerstedt
 d,
Hans
Lennernäs
d,
Femke
Heindryckx
a,
Johan
Kreuger
ab and
Paul
O'Callaghan
d,
Hans
Lennernäs
d,
Femke
Heindryckx
a,
Johan
Kreuger
ab and
Paul
O'Callaghan
 *ab
*ab
aDepartment of Medical Cell Biology, Uppsala University, Uppsala, Sweden. E-mail: paul.ocallaghan@mcb.uu.se
bThe Science for Life Laboratory, Uppsala University, Uppsala, Sweden
cDepartment of Health Science and Technology, ETH Zurich, Zurich, Switzerland
dDepartment of Pharmaceutical Biosciences, Uppsala University, Uppsala, Sweden
First published on 1st January 2026
Abstract
The reduced effectiveness of chemotherapy in many patients undergoing treatment highlights the need for novel drug combinations that target drug resistance mechanisms contributing to tumor survival. Dynamic conditions within the tumor microenvironment influence the response to anti-cancer drugs. Accordingly, identifying effective drug concentrations and interactions (additive, synergistic, or antagonistic) in relevant tumor tissue models will inform new treatment combinations. To address this need for combinatorial chemotherapeutic (CTx) screening assays, we have developed a new assay called CombiCTx, which uses a device with three reservoirs containing gels loaded with anti-cancer drugs. The drug-loaded device is inverted and placed in a standard culture dish above cancer cells, and both are then enclosed in gel. Drugs diffuse from the reservoirs and expose cancer cells to overlapping dynamic drug gradients. We imaged diffusion of the anti-cancer drug doxorubicin in the assay using time-lapse microscopy, and established an imaging protocol for quantifying MDA-MB-231 breast cancer cell survival responses along drug gradients. Finally, evaluating combination effects of navitoclax and gemcitabine with CombiCTx revealed localized effects of navitoclax, attributed to limited diffusion, while gemcitabine seemed to diffuse readily throughout the assay and revealed a mild synergy in navitoclax affected regions. These data demonstrate the capacity of CombiCTx to evaluate the cytotoxic effects of anti-cancer drug combinations while accounting for drug diffusion differences, which is relevant in the context of the 3D tumor environment and may thereby help inform clinical treatment strategies.
Introduction
The last decades have seen a surge in the development of anti-cancer drugs targeting specific molecules and pathways required for cancer cell survival and cell division.1,2 However, the potential of such targeted therapies may be compromised by various cellular mechanisms that confer drug resistance, and factors within the tumor microenvironment that reduce drug efficacy.3,4 It is therefore necessary to identify novel combination therapies, where two or more drugs that act by different mechanisms or modes of action are combined in an optimal dose ratio to reduce the probability that resistance will develop.5 There are several examples from clinical trials in which the advantages and disadvantages of combinatorial treatments on patient outcomes have been assessed.6–9 Confirming the patient benefits of such drug combinations requires clinical data; however, the initial analysis of drug combinations relies heavily on validated in vitro assays or well-established animal models.10,11 The latter are valuable model systems, but given the vast number of possible drug combinations that could be evaluated it is often not a logistically, economically or ethically reasonable approach.Pre-clinical in vitro testing is valuable to establish whether specific drug combinations produce additive, synergistic or antagonistic effects.12 For many clinically approved drug combinations, efficacy benefits are predominantly attributed to additive rather than synergistic effects.13 Nonetheless, assay systems that facilitate the detection of synergistic or unforeseen antagonistic effects of drug combinations are useful, though the translation of such synergistic combinations to patients is challenging.12–14 A recent in vitro study evaluating the potency and efficacy of 2025 two-drug combinations of clinical interest on 125 breast, colorectal and pancreatic cancer cell lines, demonstrated the benefits of combining different therapies, but synergies were only detected in 5.2% of the tested cancer cell lines with drug combination pairs.15 Such studies yield a wealth of mechanistic insights and are invaluable datasets. However, in studies with such high throughput analyses the assay conditions are typically simplified to conventional 2D well-plate systems, and while highly efficient, these systems come at the expense of typically not being able to account for the complexity of the tumor microenvironment. This is an important consideration as cellular responses to drug treatments are also context-dependent and modulated by a host of factors such as the composition and mechanical properties of the surrounding extracellular matrix (ECM), which may impact the diffusion of drugs through the tissue microenvironment.16–18
There are ongoing efforts to develop scalable 3D assay systems that better recapitulate aspects of the tumor microenvironment with the aim of more efficiently identifying promising combination therapies.18–24 Many of these approaches rely on relatively complex microfluidic and organ-on-chip systems, and while these technologies offer various advantages, their operation typically requires experienced users and specialized equipment. In the current study we have developed CombiCTx, an anti-cancer drug screening assay. The assay is relatively simple to perform and is largely compatible with standard cell culture handling. A 2D culture of cancer cells is encased in a 3D volume of hydrogel into which the CombiCTx device is embedded. The device consists of three reservoirs preloaded with anti-cancer drugs prepared in the same hydrogel, from which the drugs diffuse and create dynamic gradient landscapes over the entire cell population. We describe a detailed protocol on how to conduct the CombiCTx assay, and demonstrate an application to quantify diffusion of the chemotherapeutic doxorubicin (DOX). We establish a time-lapse fluorescence imaging protocol to record apoptosis and cell death of MDA-MB-231 breast cancer cells, using a caspase 3/7 reporter and propidium iodide, and quantify responses to the apoptotic agent staurosporine in specific regions of the CombiCTx drug diffusion landscape. Finally, we apply this protocol and use CombiCTx to conduct a drug combination study using the anti-cancer agents navitoclax and gemcitabine.
Materials and methods
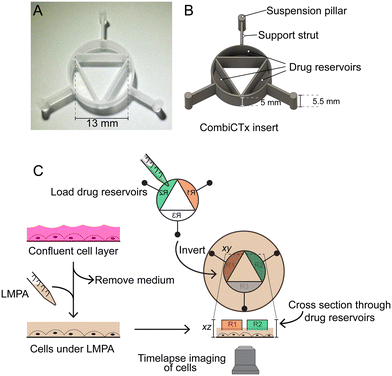
Design and fabrication of the CombiCTx insert
The CombiCTx insert was designed using Autodesk Fusion 360 (Autodesk, San Francisco, USA) computer-aided design (CAD) software. Prototype CombiCTx inserts were 3D-printed using a Prusa i3 MK3S+ printer (Prusa Research, Prague, Czech Republic) with white X-PLA 1.75 mm filament (Add North 3D AB, Ölsremma, Sweden), which facilitated the testing of various design iterations. Prior to use the inserts were sterilized by extensive spraying with 70% ethanol, followed by 30 min of UV irradiation in a cell culture hood. These printed inserts were designed to fit into a 35 mm diameter cell culture dish and each reservoir holds a volume of 100 μL. The finalized CombiCTx insert design was milled from 8 mm thick sheets of polycarbonate using a CNC machine (Datron neo, Datron, CA, USA). This approach permits the production of a large number of inserts, and requires no post-processing. Prior to use these inserts were sterilized by autoclaving, and while relatively similar in dimensions to the 3D-printed inserts, had a larger reservoir volume of 124 μL.Cell culture
The MDA-MB 231 breast cancer cell line was obtained from the American Type Culture Collection (ATCC, LGC Standards GmbH, Wesel, Germany). The genetic identity was validated25 by Idexx Bioanalytics (Ludwigsburg, Germany). Cells were grown in Dulbecco's modified Eagle's medium (DMEM) Glutamax (Thermo Fisher Scientific, Uppsala, Sweden), supplemented with 10% fetal bovine serum (FBS) (Thermo Fisher Scientific), and routinely grown and passaged before reaching confluency, at 37 °C and 5% CO2 in a humidified incubator. For the various drug assays, cells were seeded to a density of 55![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per cm2 in three wells of a tissue culture treated plastic 6-well plate one day before the start of the assay to ensure a near confluent cell layer.
000 cells per cm2 in three wells of a tissue culture treated plastic 6-well plate one day before the start of the assay to ensure a near confluent cell layer.
The CombiCTx assay setup
The CombiCTx insert consists of three reservoirs, which can be loaded with drugs of interest that are diluted in a low melting point agarose (LMPA; Thermo Fisher Scientific, Uppsala, Sweden) solution prepared with phenol red-free OptiMEM reduced serum medium (Thermo Fisher Scientific, Uppsala, Sweden). The insert is ready to be used once the LMPA has polymerized in the reservoirs. To activate a CombiCTx assay, cell media is removed from a culture dish (35 mm in diameter) in which a 2D layer of cancer cells of interest have been seeded in advance. The cells are covered with LMPA (700 μL; 37 °C), and thereafter the inverted CombiCTx is immediately inserted into the culture dish to activate the assay. As the cell-covering LMPA polymerizes it embeds the drug-containing LMPA reservoirs of the CombiCTx insert forming a continuous gel through which diffusion can occur (Fig. 1). The inverted insert is lowered into the culture dish at an angle such that two of the support pillars first touch the plate, followed by gently lowering the remaining pillar of the insert into place. This ensures minimal disturbance of the cell layer and avoids the trapping of air beneath the insert. To ensure the inserts remain stable during the polymerization process, a weight (stainless steel disc) is placed on top of the insert during polymerization, which typically occurred within 10–15 min. Media (1 ml) is then added to the culture dish to ensure all exposed surfaces are submerged, whereafter the assay is moved to the microscope for imaging, which is typically initiated within 10 min of activating the assay. We identified the LMPA-embedding of the loaded CombiCTx device in the cell culture dish to be a critical step. The objective of this process is to establish a continuum between the pre-polymerized LMPA in the reservoirs with the LMPA in the culture dish. Therefore, it is important not to disturb the setup during the polymerization process, and to subsequently handle the assay cautiously so as not to disturb the device in the gel, which could interrupt the interface between the hydrogels or introduce tears. Such imperfections may introduce channeling effects that will impact the diffusion from the reservoirs and we attributed a number of assay failures to such issues. Depending on the specific conditions of the diffusion assays, the LMPA gel solution used to cover the cell layer contained combinations of the following cell stains: 2 μM propidium iodide (PI; Invitrogen Thermo Fisher Scientific), 2 μM Cell event caspase-3/7 green detection reagent (Thermo Fisher Scientific) and NucBlue live cell stain (Invitrogen Thermo Fisher Scientific), added in accordance with the manufacturer's instructions.Fluorophore preparation for CombiCTx assays using FITC and TRITC
Fluorescein 5/6-isothiocyanate (FITC) or tetramethylrhodamine-5-isothiocyanate (TRITC) (Thermo Fisher Scientific, Uppsala, Sweden) were diluted to a concentration of 25 μM in 0.3% LMPA and added to separate adjacent reservoirs of a 3D-printed CombiCTx insert, and the remaining well was filled with 0.3% LMPA only.Drug preparation for CombiCTx assays using doxorubicin, staurosporine, navitoclax and gemcitabine
For the drug diffusion assays, doxorubicin (DOX; 2 mg ml−1 (3.68 mM) in DMSO; Accord Healthcare, Solna, Sweden; (Vnr: 189790)) was diluted to 100 μM, or staurosporine (10 mM stock in DMSO; Abcam, Cambridge, United Kingdom) was diluted to 50 μM in 0.6% LMPA and used to fill the desired number of reservoirs. For the combinatorial drug testing assays, navitoclax (10 mM stock in DMSO, CAS number 923564-51-6, Selleck Chemicals, Fisher Scientific) and gemcitabine (100 mM stock in DMSO, CAS number 122111-03-9, Selleck Chemicals, Fisher Scientific), were each diluted to a concentration of 300 μM in 0.6% LMPA. To ensure consistent LMPA concentrations, and to control for any potential effects of the DMSO solvent, DMSO was added to drug-free gel reservoirs as follows. For DOX experiments, drug-free reservoirs were loaded with LMPA containing 2.7% (v/v) DMSO. For the experiments with navitoclax and gemcitabine drug-free reservoirs were loaded with LMPA containing 3% (v/v) DMSO, to control for the highest DMSO concentration added in the navitoclax reservoirs.Physicochemical properties of the compounds used in the study
To facilitate discussions about potential differences in the diffusion of the studied molecules, the molecular mass; pKα; partition coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P; a measure of a compound's lipophilicity or hydrophobicity); and the distribution coefficient (log
P; a measure of a compound's lipophilicity or hydrophobicity); and the distribution coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D; which additionally accounts for the compound's ionization state) are summarized in Table 1. The predicted log
D; which additionally accounts for the compound's ionization state) are summarized in Table 1. The predicted log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P and log
P and log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D (at pH 7.4) values were collected from the Chemspider database (https://www.chemspider.com/) and were calculated using the PhysChem module of the ACD/Labs Percepta Platform software. Notably, predicted physicochemical values differ between databases depending on the calculation software used, and we provided the Chemspider records in Table 1 as it was the only source with log
D (at pH 7.4) values were collected from the Chemspider database (https://www.chemspider.com/) and were calculated using the PhysChem module of the ACD/Labs Percepta Platform software. Notably, predicted physicochemical values differ between databases depending on the calculation software used, and we provided the Chemspider records in Table 1 as it was the only source with log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P and log
P and log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D (at pH 7.4) values for all the molecules studied here, with the exception of TRITC. While we provide specific values in Table 1 we discuss differences in the lipophilicity between molecules and the potential effect on diffusion behavior in relative terms.
D (at pH 7.4) values for all the molecules studied here, with the exception of TRITC. While we provide specific values in Table 1 we discuss differences in the lipophilicity between molecules and the potential effect on diffusion behavior in relative terms.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P and log
P and log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D (at pH 7.4) values were collected from the Chemspider database (https://www.chemspider.com/), which were calculated with the PhysChem module of the ACD/Labs Percepta Platform software. All data was collected February 2025 (n/a; not available)
D (at pH 7.4) values were collected from the Chemspider database (https://www.chemspider.com/), which were calculated with the PhysChem module of the ACD/Labs Percepta Platform software. All data was collected February 2025 (n/a; not available)
| Compound | Abbreviation | Molecular mass (g mol−1) | pKα | log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P P |
log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D (at pH 7.4) D (at pH 7.4) |
|---|---|---|---|---|---|
| Tetramethylrhodamine-5-isothiocyanate | TRITC | 443.5 | n/a | n/a | n/a |
| Fluorescein 5/6-isothiocyanate | FITC | 389.4 | n/a | 4.00 | 4.64 |
| Doxorubicin | DOX | 543.5 | 7.34 (acidic, 1); 8.46 (2); 9.46 (basic, 3) | 2.82 | −0.79 |
| Staurosporine | STA | 466.5 | 13.46 (acidic), 9.55 (basic) | 3.29 | 3.30 |
| Gemcitabine | Gem | 263.2 | 11.52 (acidic), 3.65 (basic) | −0.47 | −1.40 |
| Navitoclax | Nav | 974.6 | 4.26 (acidic), 8.30 (basic) | 12.14 | 8.31 |
Confocal microscopy, image acquisition, processing and analysis
All imaging was performed on an LSM700 confocal laser scanning microscope (Zeiss, Jena, Germany) using a 5×/0.16 objective. Time-lapse images of fluorescently labelled cells, or fluorescent drugs were acquired using Zen imaging software (Zeiss, Jena, Germany). The tile scan function was applied to acquire 16-bit images from 11 × 11 fields of view using a digital zoom setting of 0.6, which yielded images with total x, y dimensions of 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 350 μm × 21
350 μm × 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 350 μm (2559 × 2559 pixels). The durations of the recordings are specified in the respective figures. Image analysis was performed using the Fiji version of ImageJ.26 For the doxorubicin (DOX) and staurosporine data, the time-lapse series were acquired in one continuous imaging session (using a temperature-controlled enclosure), whereas for the combinatorial drug experiments cells were returned to the incubator between the three timepoints (0, 24 and 48 hours), which were combined for each well. Images were then aligned in ImageJ using the rigid body transformation of the Hyperstack Registration plugin.27 To reduce edge-effects associated with the tile-scanning mode of acquisition (discussed below), the tile-scanned images of FITC and TRITC fluorescence were processed with the ImageJ Gaussian blur filter using a filter sigma (radius) of 75 pixels. The same Gaussian filter was applied to visualize DOX fluorescence over time in a subregion of the CombiCTx device (Fig. 3C), and DOX intensity was pseudocolored using the ‘Thermal’ color scheme in ImageJ. Quantitative analysis of DOX fluorescence was performed on the unfiltered original images. As outlined in the region of interest (ROI) maps, four key ROIs associated with each of the three reservoirs were analyzed. These were labelled as centre (c), edge (e), middle (m) and interaction zone (i). This generated 10 independent ROIs (i.e. R1(c), R1(e), R1(i), R2(c), R2(e), R2(i), R3(c), R3(e), R3(i), and (m), which was in the middle of the device between all reservoirs). Subsequently, the mean fluorescence intensity of each ROI in all three fluorescence channels was recorded for each well and at each specified timepoint. For instances where the registration procedure failed to align the images satisfactorily, the ROIs were positioned separately, and the analysis was performed as for all other wells. Quantification of relative caspase 3/7 activity and PI signals were performed as follows: The average fluorescence from each ROI was determined for each time point and for each of the three signals i.e. caspase 3/7, PI and NucBlue. To account for potential differences in cell number between ROIs the caspase 3/7 or PI signal for each timepoint was expressed relative to the NucBlue signal acquired at the first timepoint. In the experiments with navitoclax and gemcitabine we additionally normalized the quantification between the four biological replicates. Each technical replicate consisted of one CombiCTx device loaded with navitoclax alone, one device with gemcitabine alone, and one device with navitoclax and gemcitabine. The caspase 3/7/NucBlue or PI/NucBlue values for each set of technical replicates were normalized relative to the navitoclax only experiments, using the value acquired in the ROI marked R3 at the 0 h timepoint. The differences in signals collected at the 0 h and 48 h timepoints were used to assess statistical differences between the ROIs and assay conditions.
350 μm (2559 × 2559 pixels). The durations of the recordings are specified in the respective figures. Image analysis was performed using the Fiji version of ImageJ.26 For the doxorubicin (DOX) and staurosporine data, the time-lapse series were acquired in one continuous imaging session (using a temperature-controlled enclosure), whereas for the combinatorial drug experiments cells were returned to the incubator between the three timepoints (0, 24 and 48 hours), which were combined for each well. Images were then aligned in ImageJ using the rigid body transformation of the Hyperstack Registration plugin.27 To reduce edge-effects associated with the tile-scanning mode of acquisition (discussed below), the tile-scanned images of FITC and TRITC fluorescence were processed with the ImageJ Gaussian blur filter using a filter sigma (radius) of 75 pixels. The same Gaussian filter was applied to visualize DOX fluorescence over time in a subregion of the CombiCTx device (Fig. 3C), and DOX intensity was pseudocolored using the ‘Thermal’ color scheme in ImageJ. Quantitative analysis of DOX fluorescence was performed on the unfiltered original images. As outlined in the region of interest (ROI) maps, four key ROIs associated with each of the three reservoirs were analyzed. These were labelled as centre (c), edge (e), middle (m) and interaction zone (i). This generated 10 independent ROIs (i.e. R1(c), R1(e), R1(i), R2(c), R2(e), R2(i), R3(c), R3(e), R3(i), and (m), which was in the middle of the device between all reservoirs). Subsequently, the mean fluorescence intensity of each ROI in all three fluorescence channels was recorded for each well and at each specified timepoint. For instances where the registration procedure failed to align the images satisfactorily, the ROIs were positioned separately, and the analysis was performed as for all other wells. Quantification of relative caspase 3/7 activity and PI signals were performed as follows: The average fluorescence from each ROI was determined for each time point and for each of the three signals i.e. caspase 3/7, PI and NucBlue. To account for potential differences in cell number between ROIs the caspase 3/7 or PI signal for each timepoint was expressed relative to the NucBlue signal acquired at the first timepoint. In the experiments with navitoclax and gemcitabine we additionally normalized the quantification between the four biological replicates. Each technical replicate consisted of one CombiCTx device loaded with navitoclax alone, one device with gemcitabine alone, and one device with navitoclax and gemcitabine. The caspase 3/7/NucBlue or PI/NucBlue values for each set of technical replicates were normalized relative to the navitoclax only experiments, using the value acquired in the ROI marked R3 at the 0 h timepoint. The differences in signals collected at the 0 h and 48 h timepoints were used to assess statistical differences between the ROIs and assay conditions.
Bliss independence calculations for drug combination effects
We applied the Bliss independence model to calculate the expected combination effect (EBliss) of navitoclax and gemcitabine based on the cell death results from the same ROIs obtained in assays using navitoclax only or gemcitabine only, and compared them to the results obtained in assays in which navitoclax and gemcitabine were combined (Fig. 5C–E). We applied the same approach to compare distinct ROIs within assays in which cells were treated with the combination of navitoclax and gemcitabine (Fig. 5F). We selected the Bliss independence model as it assumes that the drugs in question have distinct modes of action or targets,28 which is the case for navitoclax and gemcitabine, and was calculated using the formula:| EBliss = ENav only + EGem only − (ENav only·EGem only) |
CI values that were below, equal to, or above 1 were respectively indicative of drug synergies, additive effects, or antagonisms.29
Statistical analyses
Statistical analyses were performed using Graphpad Prism 10 software. The type of statistical test used to analyze specific datasets and the number of replicates included in the analyses is described in the associated figure legends. Briefly, a Shapiro–Wilk test was used to test for normality of data distribution. Paired Student's t-test was applied to compare differences between two groups when data was collected from within the same CombiCTx device. Significant main effect differences between more than two groups were assessed using two-way ANOVA, and the differences between specific groups was identified using the Tukey's multiple comparisons post hoc test.Results and discussion
The CombiCTx assay is based on the previously developed CombiANT assay, used for antibiotic interaction testing,30 and redesigned here for the purpose of studying the combined interactions and effects of anti-cancer drugs on adherent cancer cells. Central to the assay is the CombiCTx insert (Fig. 1A) that comprises three reservoirs, which can be filled with drugs of interest that are dissolved in low melting point agarose (LMPA). The CombiCTx insert features three support struts, each radiating outward from the midpoint of the outer wall of each reservoir, with a suspension pillar at their distal end that fixes a spatial distance of 0.5 mm from the bottom of the cell culture dish to the drug-loaded reservoir (Fig. 1B). The assay is activated in a two-step procedure: firstly, the CombiCTx reservoirs are loaded with the LMPA containing the drugs of interest and allowed to polymerize, next a 2D culture of cells is covered in a layer of LMPA, and the drug-loaded CombiCTx is inverted and gently lowered into the LMPA covering the cell layer prior to its polymerization (Fig. 1C). As the drug reservoirs face downwards towards the cells, once the LMPA covering the cells polymerizes it forms a continuous connection with the drug-containing LMPA in the reservoirs. This ensures that drugs can diffuse from each of the three drug reservoirs in a continuous gel towards the cancer cells, and thereby form dynamic gradient landscapes that expose cells to a wide range of drug concentrations.Imaging patterns of combined fluorophore diffusion in the CombiCTx assay
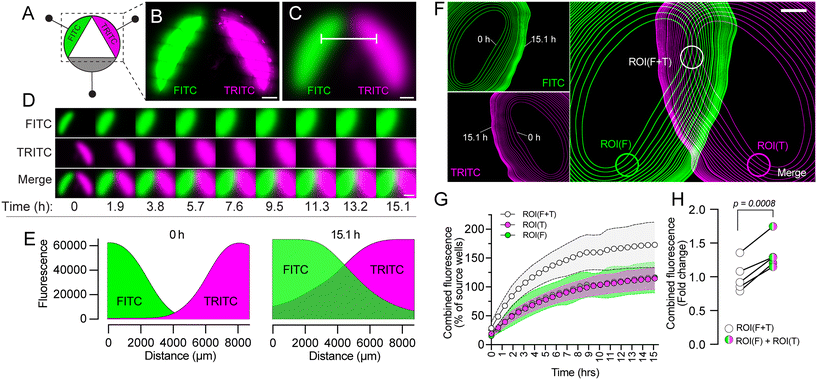
Diffusion in hydrogels will be influenced by the molecular mass as well as other physiochemical properties of the compounds of interest; potential interactions between the compounds and the diffusion matrix; as well as characteristics such as the pore size and molecular occupancy of the hydrogel.31–34 To visually assess and characterize the formation of dynamic gradients in the CombiCTx assay we imaged the diffusion of two fluorophores FITC and TRITC from two different reservoirs of the CombiCTx insert, outlined in Fig. 2A. FITC and TRITC have respective molecular mass of 389.4 g mol−1 and 443.5 g mol−1, which are similar to commonly used anti-cancer drugs such as DOX (543.5 g mol−1), and the reported pore sizes for agarose gels of a similar percentage to those used here are in the low micrometer range.35 Using Molview (https://molview.org/) we estimated the molecular diameters of FITC and TRITC to be respectively 1.25 nm and 1.46 nm, and so the relatively larger hydrogel mesh size is unlikely to have size-exclusion effects on fluorophore diffusion.31 The isothiocyanate group in both FITC and TRITC reacts with amine groups at pH 7–9, but as agarose gels are comprised of linear galactose-based polysaccharides with chemical formula (C12H18O9)n,36 no such interactions are expected.37 Further, given that the average surface charge on agarose is predicted to be neutral,38 the diffusion of fluorophores would not be expected to be extensively electrostatically impeded. Together, this suggests that FITC and TRITC would diffuse freely in a similar manner in the LMPA gels. To visualize this a field of view incorporating the FITC- and TRITC-filled reservoirs was acquired by tile-scanning, which stitches together multiple acquisition frames (Fig. 2B). Signal reduction was observed at the edges between adjacent frames within the tile-scan, which is a recognized imaging artefact sometimes associated with this acquisition mode.39 In an effort to smooth the transitions between frames we processed the tile-scanned images in ImageJ with a Gaussian blur (sigma radius of 75 pixels) filter (Fig. 2C). A montage of the time-lapse imaging for the fluorophores illustrates their diffusion patterns over a 15.1 h duration, during which they entered the central triangular area of the device (Fig. 2D). As the area directly below the apex of the triangle is closest to the FITC and TRITC reservoirs, it would be the first region in which diffusing FITC and TRITC will combine. To visualize this combination the fluorescence profile between the FITC and TRITC reservoirs was quantified using a line plot (white line in Fig. 2C) in the 0 h and 15.1 h images, which revealed the formation of overlapping and opposing FITC and TRITC gradients (Fig. 2E). To better visualize the temporal and spatial changes and overlap in FITC and TRITC diffusion, a threshold was set for the FITC and TRITC fluorescence from the time-lapse in Fig. 2D and the detectable moving front of these signals were outlined for each timepoint, and projected into a single FITC, TRITC and Merge image (Fig. 2F). From the earliest timepoint the combination of FITC and TRITC diffusion gradients between the adjacent FITC and TRITC reservoirs was apparent as an overlap of FITC and TRITC fronts in the upper region of the device. At later timepoints this overlap spread to the centre of the device. In regions where the FITC and TRITC reservoirs were adjacent to the lower empty reservoir no FITC and TRITC signal overlap was apparent (Fig. 2F). To quantitatively assess these differences, the total FITC and TRITC fluorescence from three regions of interest (ROIs) were quantified from five independent experiments. As expected, the combination of FITC and TRITC gradients in ROI(F + T), resulted in a higher degree of total fluorescence than in ROI(F) or ROI(T), in which only FITC or TRITC gradients had been observed (Fig. 2G). Notably, at the endpoint of the assay the sum of the fluorescence recorded from ROI(F) + ROI(T) was 1.352 ± 0.056 (mean ± SEM) fold higher than the total fluorescence observed in ROI(F + T) (Fig. 2H). This indicates that fewer FITC molecules entered ROI(F + T) than ROI(F), and fewer TRITC molecules entered ROI(F + T) than ROI(T). This is likely due to differences in the concentration gradient of total fluorophores that forms over time between ROI(F + T) and the FITC and TRITC reservoirs, which would be expected to be less steep than the gradients between ROI(F) and the FITC reservoir, and ROI(T) and the TRITC reservoir. This likely reduces the diffusion of fluorophores into ROI(F + T) and may explain why the fluorescence in ROI(F + T) is less than, rather than equal to, the sum of fluorescence in ROI(F) + ROI(T). Together this data demonstrates the diffusion landscapes that can be formed between different compounds in the CombiCTx device, and highlights how distinct regions within the device can be used to compare combinatorial interactions and effects with those of individual compounds.Imaging doxorubicin (DOX) diffusion gradients in the CombiCTx assay
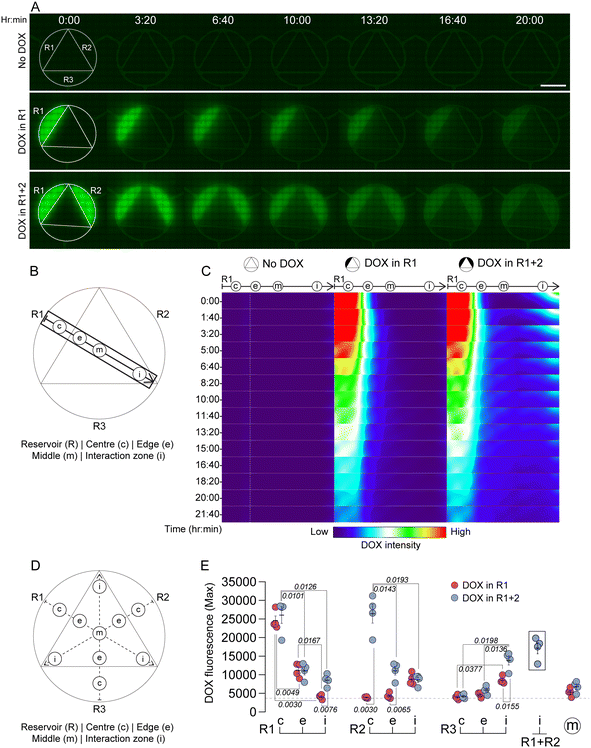
To characterize the formation of dynamic gradients of a small molecule anti-cancer drug within the CombiCTx assay we imaged the diffusion of DOX, a widely studied and clinically used anthracycline chemotherapeutic.40 DOX has a molecular mass of 543.5 g mol−1, and its distinct photophysical properties permit quantification of drug concentrations by measuring absorbance or fluorescence,34,41 which enabled us to visualize drug diffusion by tracking DOX fluorescence in the CombiCTx assay. The assay was conducted with LMPA-covered MDA-MB-231 cells, which were labelled with the NucBlue nuclear stain. The CombiCTx device was loaded without DOX (No DOX), with DOX loaded in reservoir 1, or DOX loaded in reservoirs 1 and 2. Time-lapse confocal microscopy images were collected over a total of 21 h 40 min, and the montage in Fig. 3A represent images from seven timepoints during 20 hours of imaging. To facilitate visualization of DOX diffusion from one reservoir over time, a rectangular subregion was isolated from the tile-scanned images, and included the following regions of interest (ROI): the centre (c) of reservoir 1 (R1); the area at the outside edge (e) of the R1 wall; the middle (m) of the entire device; and the interaction zone (i) that is furthest from R1, but closest to both R2 and R3 (Fig. 3B). A montage of images from this subregion was prepared for the entire timeseries (14 images over 21 h 40 min) for each condition, and as in Fig. 2, these were processed using the ImageJ Gaussian blur filter (75 pixels). The DOX fluorescence intensity was pseudocolored with a gradient color scheme, such that areas of high DOX intensity were red and low DOX intensity were dark blue (Fig. 3C). The DOX-free condition displayed a consistent level of background fluorescence across all time points. In the conditions in which DOX was loaded in R1, or in R1 + R2 simultaneously, the fluorescence from within the centre (c) of the reservoir decreased over time as DOX diffused outwardly into the surrounding gel. In comparison, the DOX intensity appeared to increase in areas towards the middle of the device, and was most pronounced in the conditions with DOX in R1 + R2 (Fig. 3C). To quantitatively assess differences in DOX fluorescence intensities throughout the CombiCTx device, the average fluorescence associated with the (c), (e), (m) and (i) ROIs associated with R1, R2 and R3 were measured and plotted over time for conditions with DOX in R1, and DOX in R1 + R2 (Fig. S1A and B).To compare the highest levels of DOX exposure between each of the conditions, we analyzed maximum DOX fluorescence values from each ROI (Fig. 3D and E). When DOX was loaded only in R1, significant differences were detected between the selected ROIs that were positioned along the concentration gradient. Specifically, relative to the average maximum DOX values in the centre (c) of the reservoir, DOX values were 54.3% lower at the edge (e) ROI, 78.2% lower in the middle (m) of the device, and 83.4% lower in the ROI that we term the interaction zone (i) between R2 and R3. This is the ROI that is furthest from R1, but still contained within the perimeter of the CombiCTx device (Fig. 3E). When DOX was loaded in R1 and R2 near symmetrical gradients of DOX were observed from each side of the device, which could be confirmed by comparing the maximum DOX fluorescence between spatially equivalent paired positions. Such that, R1(e) was 54.3% lower than R1(c), and R2(e) was 54.7% lower than R2(c); R1(i) was 70.1% lower than R1(c), and R2(i) was 68.3% lower than R2(c). For DOX in R1 + R2 we also compared DOX exposure in the “interaction zones” between reservoirs i.e. R1(i), R2(i) and R3(i) (Fig. 3D and E). The maximum DOX fluorescence in R3(i) (between the DOX-containing reservoirs R1 and R2) was 1.63 ± 0.17 (mean ± SD) fold higher than that in R1(i) and 1.60 ± 0.09 (mean ± SD) fold higher than that in R2(i). This was expected due to the fact that both R1(i) and R2(i) are adjacent to the DOX-free reservoir R3. However, as observed for FITC and TRITC (Fig. 2G), the DOX fluorescence in R3(i) was still 0.81 ± 0.06 (mean ± SD) fold lower than what would have been predicted from the sum of the values recorded in R1(i) and R2(i) (Fig. 3E, values enclosed by black rectangle).
To assess differences in DOX fluorescence within the perimeters of the reservoirs we identified two additional regions of interest in the peripheries of each reservoir (distal 1 (d1), and distal 2 (d2)) and compared the maximum DOX fluorescence between these and the reservoir's centre (c) (Fig. S1C). The maximum DOX fluorescence in d1 and d2 of R1 were on average lower than that recorded in the centre (c) of R1, and this was similar between conditions (18.5% reduction for DOX in R1, and 18.2% for DOX in R1 + R2). A similar reduction (20.7%) was also detected for the d1 and d2 positions in R2, compared to its centre (c); however, possibly due to variability in the individual values, none of these reductions were statistically significant (Fig. S1D). Nonetheless, by using DOX fluorescence as a proxy for drug concentration, these data demonstrate that there may be location-dependent differences within the perimeter of the reservoirs. This may also reflect differences in local concentration gradients, as DOX molecules in regions close to the perimeter (e.g. the distal ROIs); encounter a steeper gradient and therefore diffuse more readily into the surrounding DOX-free gel than DOX molecules originating from the centre (c) of the reservoir, which are initially surrounded by gel containing similarly high DOX concentrations.
To assess DOX interactions with the MDA-MB-231 cells seeded under the CombiCTx devices in Fig. 3 we had imaged NucBlue fluorescence in parallel with DOX fluorescence. At the 0 h timepoint the NucBlue signal from the middle of the device was similar between the No DOX, DOX in R1, and DOX in R1 + R2 conditions, and revealed homogenous distributions of cells (Fig. S2A and B). However, in the area contained within the perimeter of R1 and R2, the presence of DOX was associated with a substantial reduction in the NucBlue signal. As DOX diffused outwardly into the device the NucBlue signal similarly decreased (Fig. S2C). This decrease was most evident during the initial 6 h 40 min of the assay, but persisted until the end, and a greater overall decrease in NucBlue signal was observed for the condition with DOX in R1 + R2 (Fig. S2C). DNA intercalation of DOX following cellular uptake has previously been established to compete with binding of NucBlue (which is a formulation of Hoechst 33242), and we have previously demonstrated a similar time-dependent, inverse relationship between NucBlue and DOX fluorescence for HepG2 tumor cells exposed to a gradient of DOX.42,43 Therefore, the decrease in the NucBlue signal associated with the MDA-MB-231 cells in Fig. S2C provides indirect evidence of cellular uptake of DOX and provides an additional proxy measure of DOX diffusion in the CombiCTx device.
Together these data further support the observations in Fig. 2, which highlight the utility of analyzing regions of interest within the same CombiCTx device for assessing dynamic concentration gradients of drugs. The establishment of such gradients bears similarities with how drugs diffuse in tissues, and such an in vitro assay that accounts for these diffusion effects could provide valuable translational knowledge for locoregional anti-cancer treatments, such as transarterial chemoembolization (TACE) and local prostate injections.44–46
Quantitative image analysis of apoptosis and cell death in the CombiCTx assay
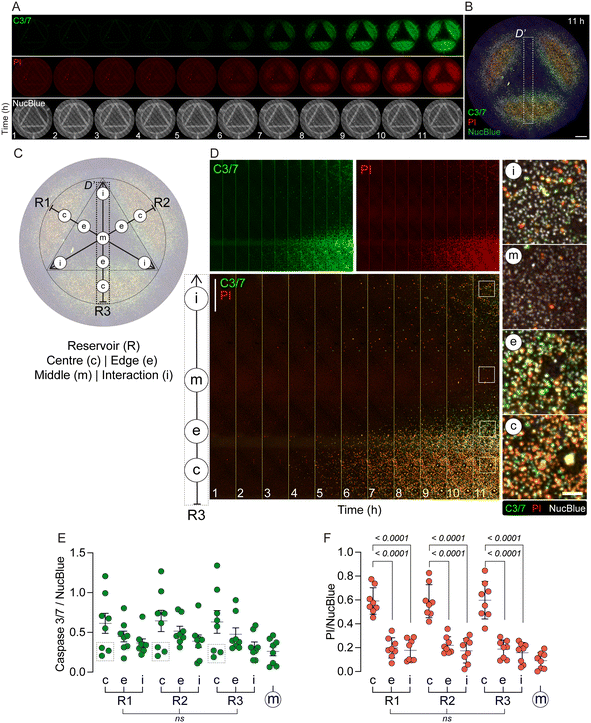
To facilitate quantification of the cell response to anti-cancer agents using the CombiCTx assay, we next adapted an imaging protocol using a caspase 3/7 reporter to detect apoptotic cells and the nuclear stain propidium iodide (PI, which is impermeable to living cells) to detect dead cells. MDA-MB-231 breast cancer cells were pre-seeded on the bottom of a cell culture dish and cultured under standard conditions. Prior to initiating the CombiCTx assay, all three CombiCTx reservoirs were loaded with 50 μM staurosporine, which has a molecular mass of 466.5 g mol−1, which is similar though somewhat smaller than DOX; however, other physiochemical properties such as lipophilicity, differs between the two, and may alter their respective diffusion (Table 1). DOX was replaced by staurosporine in these experiments due to the fact that DOX's fluorescence emission profile overlaps with that of the caspase 3/7 reporter. Staurosporine is a potent protein kinase C inhibitor and an established inducer of apoptosis,47 capable of acting through both caspase-dependent and caspase-independent mechanisms.48 We have previously demonstrated that staurosporine induces detectable caspase 3/7 activation in MDA-MB-231 cells after approximately 5 h of exposure.49 Prior to assay initiation, the cancer cells were covered with an LMPA solution in which the caspase 3/7 reporter, PI, and NucBlue stains were diluted. Time-lapse confocal microscopy imaging of labelled MDA-MB-231 cells under and around the entire CombiCTx insert was carried out to visualize the induction of apoptosis and occurrence of cell death over an 11 h time-lapse experiment (Fig. 4A and B). Gradual caspase 3/7 activation and increased cell death (i.e. PI-positive cells) were primarily observed in the regions directly under or close to the drug reservoirs (Fig. 4A–C). The subregion D′ (Fig. 4B and C), which included the centre (c) of R3; the R3-associated edge (e); the middle (m) of the device; and the R3 interaction zone (i) between R1 and R2, was extracted from the tile-scanned time-lapse images and montages of the caspase 3/7, PI and merged images are presented in Fig. 4D. While a number of caspase 3/7 and PI-positive cells are visible during the initial 4 h of imaging, in and around R3(c), both signals most clearly started to increase from the 5 h timepoint through to the final 11 h timepoint. This revealed a gradient of apoptotic and necrotic (PI-positive) cells that decreases towards the middle (m) of the CombiCTx insert and increases towards the interaction zone (i) between R1 and R2. These profiles are in agreement with expected staurosporine concentration gradients. Higher magnification images of the (c), (e), (m) and (i) ROIs illustrate the differences in caspase 3/7 and PI staining relative to the NucBlue-positive cells in each (Fig. 4D).To quantitatively assess differences in caspase 3/7 and PI fluorescence within the CombiCTx device, the average fluorescence within the (c), (e), (m) and (i) ROIs associated with R1, R2 and R3 were measured and plotted over time for a total of eight independent experiments in which staurosporine was loaded into R1, R2 and R3 (Fig. S3A and B). Differences between ROIs were evaluated for the final 11 h timepoint (Fig. 4E and F) where the caspase 3/7 and PI signals were normalized to the average NucBlue signal recorded in the associated areas, to account for potential differences in cell numbers. For caspase 3/7 and PI there was no significant differences between R1, R2, or R3 for their respective ROI values, which suggests that in applications of the CombiCTx assay where the same drug is loaded in all reservoirs then spatially matched ROIs can function as technical replicates of each other. There was a high degree of variability for the caspase 3/7 signals, particularly within the centre (c) ROIs for each reservoir. As expected, the mean signal was highest there, and was on average 24% lower in the edge (e) ROIs; 43% lower in the interaction zones (i) ROIs; and 57% lower in the middle (m) of the device. However, these differences between ROIs were determined not to be significant based on Tukey's multiple comparisons (Fig. 4E). In contrast, compared to the PI signals in the centre (c) of each reservoir, the mean signals in the edge (e) ROIs were on average 66% lower, and 71% lower in the interaction zone (i) ROIs, which represented significant signal reductions. The PI signal was further reduced by 85% in the middle (m) of the device. While the differences between the (e) and (i) ROIs were not significantly different they were on average lower in (i) than (e). Nonetheless, this suggests that the concentration of staurosporine was greater at the edge (e) of the reservoirs than in the interaction zone (i). This contrasts somewhat with what might have been expected based on the DOX experiments, where DOX fluorescence was greater in the interaction zone R3(i) between R1 and R2, than in R1(e) or R2(e) (Fig. 3E). These differences may reflect how the specific physiochemical properties of DOX and staurosporine impact their diffusion. We also tested versions of the assay in which staurosporine (50 μM) was loaded in R1 only or R1 + R2, and at the 11 h timepoint the caspase 3/7 and PI signals were in line with expected diffusion patterns for the drug i.e. the greatest areas of cell death were detected in the respective reservoirs and gradually decreased outwardly into the surrounding gel areas (Fig. S3C).
As mentioned, caspase 3/7 values recorded in ROI(c) were variable for each of the reservoirs. For three experiments in particular (values enclosed by a dashed square in Fig. 4E) the caspase 3/7 values in R1–R3(c), were lower than the average signal from the caspase 3/7 values in the adjacent R1–R3(e) ROIs. However, all of the eight PI values from R1–R3(c), which were acquired in the same experiments, were significantly higher than the R1–R3(e) ROIs. Therefore, MDA-MB-231 cells in the (c) ROIs would have lost membrane integrity, becoming PI-positive, without initiating caspase 3/7 activity. Given that equal staurosporine concentrations were used throughout, and induced reproducible levels of PI-positive cells, this variability in caspase 3/7 activation is difficult to reconcile; although, as mentioned above staurosporine has previously been demonstrated to induce cell death through caspase-independent mechanisms.48 These data illustrate the benefits of being able to assess cell viability with more than one reporter in the CombiCTx assay, which could be particularly useful when assessing cell responses to drugs where the mode of cell death is not established.
In summary, this imaging strategy permits the quantitative assessment of anti-cancer drug effects on cell viability, and can be spatially and temporally resolved in the CombiCTx assay to study combinations of dynamic drug diffusion gradients.
Assessing the cytotoxic effects of combinations of navitoclax and gemcitabine using the CombiCTx assay
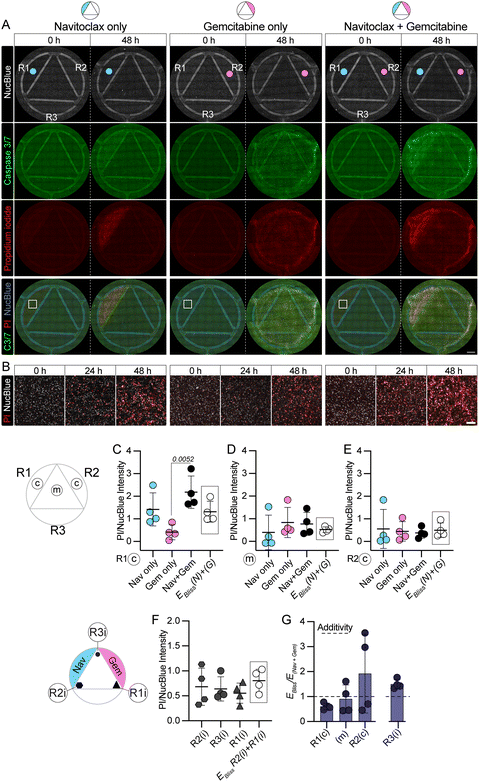
Finally, we applied the CombiCTx assay to assess the cytotoxicity of combinations of two chemotherapeutic drugs with different physicochemical properties and mechanistic targets; namely, navitoclax and gemcitabine. Navitoclax is a Bcl2-inhibitor that has been investigated in phase I and II clinical trials.50 Gemcitabine, an approved chemotherapeutic treatment of the anti-metabolite class, inhibits DNA synthesis through integrating into the new DNA strand and inhibiting the nucleoside salvage pathway, thus hindering DNA replication.51 Jaaks et al. reported robust synergy between a navitoclax and gemcitabine combination treatment of basal-like breast cancer cell lines (including MDA-MB-231 cells) in a 2D assay, with synergies detected in approximately 69% of the 23 breast cancer cell lines included in the validated subset.15 Navitoclax and gemcitabine have also been tested in a combinatorial dosing regimen in a phase I clinical trial in patients with solid tumors and aimed to determine safety, optimal dosing, and clinical activity. The regimen was well tolerated, but provided a best response of stable disease i.e. the extent and severity of the cancer neither increased or decreased.52The MDA-MB 231 cells were exposed to navitoclax-only, gemcitabine-only, or navitoclax and gemcitabine. These three conditions were tested in separate CombiCTx devices such that navitoclax was loaded in R1; gemcitabine was loaded in R2; or navitoclax was loaded in R1 and gemcitabine was loaded in R2 (Fig. 5A). Reservoirs not loaded with drugs were loaded with LMPA containing DMSO, as a solvent control. Imaging was performed directly after activation of the CombiCTx assay and again after 24 h and 48 h, between which the assay was returned to the incubator. The apoptosis and cell death imaging protocol described in Fig. 4 was applied to determine the extent of cytotoxicity (Fig. 5A). Examples of the collected tile scan images from each of the three experiment conditions at the 0 h and 48 h timepoints are presented in Fig. 5A. In Fig. 5B, an enlarged view of the NucBlue and PI signal within the centre (c) of R1 is presented for all three timepoints and illustrates how the PI signal increased over time in each of the conditions.
Quantification of relative caspase 3/7 activity and PI signals per NucBlue intensity was performed as detailed in the Materials and Methods, and the values acquired in each ROI from each time-point are presented in Fig. S4A–C. Caspase 3/7 activity increased over time, as expected, with the highest activity recorded at 48 h in the navitoclax + gemcitabine assay in the region directly under the navitoclax drug reservoir (Fig. S4B). Similarly, the maximum PI signal was recorded after 48 h under the navitoclax-filled drug reservoir in the navitoclax + gemcitabine assay (Fig. S4C). Two-way ANOVA revealed a significant main effect difference for time in both the caspase 3/7 and PI data, but overall, the variability in the relative caspase 3/7 intensity was high between replicates, similar to that observed in Fig. 4 for the experiments with staurosporine. Therefore, we focused our analysis on the PI dataset. Notably, at the 0 h timepoints we detected elevated PI fluorescence under wells containing navitoclax (Fig. S4C). As this timepoint is acquired directly after activation of the CombiCTx assay, the PI signal is not readily attributed to cell death, and to our knowledge navitoclax has no intrinsic fluorescence properties. Nonetheless, to ensure that subsequent quantitative comparisons would only account for the degree to which the relative PI signal changed during the 48 h time period, we subtracted the 0 h timepoint intensities from the 48 h data for the data presented in Fig. 5C–F.
As the establishment of diffusion gradients requires time, locations within the CombiCTx assay that are closer to one drug source than the other can also be used to assess the effects of sequential treatments. We compared the relative PI signal intensity in the centre (c) of R1, in which navitoclax was loaded in the navitoclax-only, and navitoclax + gemcitabine conditions, and which was drug-free in the gemcitabine only conditions. In the navitoclax + gemcitabine assay, cells in this location would be first exposed to a high concentration of navitoclax, followed later by a relatively lower concentration of gemcitabine. After 48 h, the relative PI signal in R1(c) was significantly higher in the navitoclax + gemcitabine assay compared to the gemcitabine-only assay (9.00 ± 5.91 fold; mean ± SEM), and while not significant it was also higher (1.66 ± 0.22 fold; mean ± SEM) than that observed for navitoclax-only (Fig. 5C). This PI signal from the combined treatment assays were also 1.69 ± 0.17 (mean ± SEM) fold higher than the EBliss (Nav only) + (Gem only) values for R1(c), which calculates the expected effect of combining these two drugs based on the effects recorded in the single drug assays (Fig. 5C, datapoints within the rectangle). Furthermore, the combination indices calculated from these EBliss R1(c) values and the corresponding data for the effects (E) recorded in the navitoclax + gemcitabine assays (ENav+Gem) were all below 1 (Fig. 5G), which was indicative of mildly synergistic drug effects. In contrast, comparisons between the centrally located ROI (m) for the different assays revealed no significant differences (Fig. 5D). If both drugs diffused at an equal rate from their respective reservoirs the combined drug concentration in ROI (m) would be predicted to be higher in the navitoclax + gemcitabine than in the navitoclax-only or gemcitabine-only assays. However, for the navitoclax-only conditions the relative PI signal was close to 0 for the majority of replicates (0.40 ± 0.37; mean ± SEM), while elevated signals were detected for the gemcitabine-only condition (0.84 ± 0.33; mean ± SEM) and the navitoclax + gemcitabine condition (0.77 ± 0.25; mean ± SEM) (Fig. 5D). A similar pattern was observed in the centre (c) of R2, which was loaded with gemcitabine in the gemcitabine only and navitoclax + gemcitabine assay, and gemcitabine-free in the navitoclax-only assay (Fig. 5E). This suggests that the degree of navitoclax diffusion within the assay is much lower than that of gemcitabine. The EBliss/ENav+Gem calculated for the (m) ROIs and R2(c) were also highly variable, which may in part be due to the relatively low navitoclax concentrations reaching these locations (Fig. 5G). A possible explanation for these diffusion differences is the respective differences in the molecular mass and lipophilicity of these two drugs. Gemcitabine is the smallest molecule in our study (263.2 g mol−1),53 and is substantially less lipophilic than navitoclax (see Table 1) and has negligible protein binding affinities.54 In contrast, navitoclax was the largest drug tested in the assay with a molecular mass of 974.6 g mol−1 and high lipophilicity (see Table 1), with strong protein binding properties.55,56 Therefore, it is likely that gemcitabine readily diffuses throughout the assay space impacting viability at a greater range, while navitoclax diffuses more slowly causing more localized effects, and may be further hindered by interactions with the cells seeded directly under the navitoclax reservoir. To compare the combined effect of navitoclax and gemcitabine exposure with the effect from predominantly navitoclax or gemcitabine alone within the same CombiCTx assay, we compared the relative PI signal recorded in the interaction zones associated with each reservoir i.e. R1(i), R2(i) and R3(i). A similar degree of cell death was observed in all three locations, and the predicted combination effect (EBliss R2(i) + R1(i)) calculated by combining the cell death observed in R2(i) (nearest navitoclax) with that in R1(i) (nearest gemcitabine) was not observed in R3(i) (which is equally near the navitoclax and gemcitabine reservoirs; Fig. 5F). As a result, the EBliss/ENav+Gem indices in R3(i) were above 1, indicating a mildly antagonistic effect of the combined drugs in this location (Fig. 5G). As noted, Jaaks et al. demonstrate that a navitoclax and gemcitabine combination has synergistic effects in 2D cultures of breast cancer cells.15 We similarly observed evidence of synergy using a standard 2D cell viability assay, where combinations of gemcitabine (fixed at 20 μM) with increasing concentrations of navitoclax were added in solution to cultures of MDA-MB-231 cells (Fig. S5A–C). A significant decrease in viability was observed after 48 h when cells treated with increasing concentrations of navitoclax were simultaneously treated with gemcitabine (20 μM) (Fig. S5B). The 20 μM concentration of gemcitabine alone reduced viability on average to approximately 75%. Viability differences between navitoclax conditions with or without gemcitabine proved negligible when navitoclax concentrations exceeded 1 μM, which reduced cell viability to approximately 30% or lower. As in Fig. 5 we calculated the EBliss/ENav+Gem indices and determined that synergistic effects (i.e. <1) on cell death were detected when 20 μM of gemcitabine was combined with navitoclax concentrations at or below 1 μM (Fig. S5C). This demonstrates that low doses of navitoclax can potentiate the effects of gemcitabine when cells are exposed to solutions of the drugs in a conventional plate assay format. In the CombiCTx experiments we identified a potential synergy between navitoclax and gemcitabine, compared to navitoclax alone in the R1(c) position (Fig. 5C and G), but no effect was observed in the R2(c) position (Fig. 5D and G) i.e. where high gemcitabine concentrations would have been expected to combine with low navitoclax concentrations if it had readily diffused. Together these observations highlight the importance of complimenting standard 2D drug tests with assays such as CombiCTx that can account for the effects of the extracellular environment on drug diffusion and potentially facilitate better predictions of drug efficacies.
Limitations of the current study
While the CombiCTx assay represents some important similarities with how drugs are delivered to tumor cells in the body, there are a number of limitations to be considered. Firstly, the tumor cells are grown in standard tissue culture dishes as 2D monocultures of MDA-MB-231 cells. Therefore, while the extracellular matrix that covers the cells to mimic the tumor microenvironment is 3D in nature, the cell culture is not. An advantage of culturing cells in 2D is that they are easily imaged with standard protocols, and the assay could be further adapted for other high content imaging protocols such as Cell Painting to phenotypically profile cell responses within gradients of drugs or other effector combinations.59–61In vivo the presence of multiple different cell types in the tumor stroma may impact the effect of tumor cell drug responses, and heterogeneity between tumor cells will result in different susceptibilities to treatments. This complexity is not recapitulated in the applications described here, which is limited to analyses of the MDA-MB-231 triple negative breast cancer cell line. However, it would be possible to label and co-culture different tumor cell types with and without established drug resistance, and/or stromal cells of interest, to compare cell-type specific responses within the CombiCTx device. The assay could also be adapted for 3D cell cultures; for example, tumor spheroids could be separately cultured and then distributed throughout the gel landscape, which could be pre-populated with other cells of interest. Another simplification in the current assay format is the use of low melting point agarose (LMPA) to model the 3D extracellular environment. To increase complexity, it would be interesting to apply the CombiCTx assay to evaluate how gradients and cell responses to drugs are affected by different types of tumor-like matrices.34,57–62 Determining what drug concentrations to load in the CombiCTx assay also requires consideration. For example, for navitoclax we loaded 300 μM in the device reservoirs. A Phase I trial with navitoclax detected peak plasma concentrations of approximately 3.2 μM, following administration of a 150 mg oral dose.6 From the reported area under the curve (AUC) data, after 24 h the plasma concentration remain above approximately 1 μM, and in each 21-day treatment regimen patients were administered a daily navitoclax dose for three or five consecutive days. Therefore, while these are substantially lower drug concentrations than those loaded here, there is a sustained exposure to the drug throughout the three or five days of treatment. It is difficult to translate such clinical data into a recommended dose to use in the CombiCTx device, but for readily diffusible drugs that will establish dynamic concentration gradients throughout the assay over time, it is likely necessary to consider loading higher drug concentrations than those found to be effective in conventional 2D assays, to ensure that the full spectrum of drug effects is captured. Relatedly, due to the design of the device, we assume that cell responses in locations that are suitably far from the reservoirs can serve as drug-free controls. However, we cannot rule out the possibility that drug-induced paracrine signals may diffuse into these control regions and elicit unforeseen responses. Such effects could potentially be accounted for by initially characterizing control cell responses in drug-free devices. Further developments of the CombiCTx device could incorporate modular dosing reservoirs, to accommodate repeat dosing regimens. Additional complexity could be added through the integration of fluidic channels, which would permit the evaluation of perfusion effects on drug diffusion, cell permeability, and potentially even drug clearance. Further, enclosing the assay in such a microfluidic system would also facilitate regulation of other parameters such as pH, oxygen availability and nutrient access, which will also alter sensitivity to drugs, and a number of relevant systems are already under development by others.24,63–65In conclusion, this study demonstrates the capacity of CombiCTx to evaluate the cytotoxic effect of combinations of anti-cancer drugs in an assay format that captures differences in drug diffusion characteristics. As the tumor microenvironment may similarly differentially impact drug combinations in vivo, the assay provides a useful testing ground to evaluate such effects. Such data will better inform the selection of drug combinations, and in future research the platform may be applied to assess chemotherapy combinations for TACE-like treatment strategies, which involve the locoregional delivery of chemotherapies to the tumor.
Author contributions
Conceptualization: C. S., O. E., J. K., P. O'C. Methodology: C. S., A. L. C., S. C., B. S., O. E., O. D., H. L., F. H., J. K., P. O'C. Validation: C. S., A. L. C., S. C., B. S., O. E., O. D. Investigation: C. S., A. L. C., S. C., B. S., O. E., O. D., P. O'C. Formal analysis: C. S., A. L. C., P. O'C. Resources: H. L., F. H., J. K. Data curation: C. S., A. L. C., P. O'C. Visualization: C. S., A. L. C., J. K., P. O'C. Writing original draft: C. S., J. K., P. O'C. Writing – review and editing: C. S., A. L. C., S. C., B. S., O. E., O. D., H. L., F. H., J. K., P. O'C. Supervision: H. L., F. H., J. K., P. O'C. Funding acquisition: H. L., F. H., J. K.Conflicts of interest
Johan Kreuger is a co-founder of Rx Dynamics that develop CombiANT technology and a co-applicant on a related patent application. The CombiANT assay is closely related to the CombiCTx assay described here.Data availability
The quantitative image analysis data and the supplementary materials and methods, and Fig. S1–5, have been provided as Supplementary Information (SI) files. See DOI: https://doi.org/10.1039/d5lc00686d. The CAD files for the CombiCTx device are available on request.Acknowledgements
This study was conducted as part of the Additive Manufacturing for the Life Sciences (AM4Life) consortium, which is funded by Sweden's Innovation Agency Vinnova (grant number 2019-00029), and by additional grants to J. K. from the Swedish Cancer Society (Cancerfonden; grant numbers 20 1285 PjF and 23 2692 Pj 01 H), and the Göran Gustafsson's Foundation (Göran Gustafssons Stiftelser). F. H. is supported by the Swedish Cancer Society (Cancerfonden; grant number 23 2776 Pj), the Swedish Society for Medical Research (grant number S17-0092), The Swedish Research Council (Vetenskapsrådet; grant number 2021-01628) and Göran Gustafsson's Foundation (Göran Gustafssons Stiftelser). H. L. is supported by the Swedish Cancer Society (Cancerfonden; grant number CAN 24 3519 Pj 01 H) and the Swedish Research Council (Vetenskapsrådet; grant numbers 2020-02367 and 2024-03166). 3D printing was performed at U-PRINT: Uppsala University's 3D-printing facility at the Disciplinary Domain of Medicine and Pharmacy and SciLifeLab Uppsala. We would like to thank Johan Öhman from the Dept. of Cell and Molecular Biology, Uppsala University for his assistance with the CNC milling of the CombiCTx inserts.References
- D. Hanahan, Cancer Discovery, 2022, 12, 31–46 CrossRef CAS PubMed
.
- D. Hanahan and R. A. Weinberg, Cell, 2011, 144, 646–674 CrossRef CAS PubMed
.
- S. Negrini, V. G. Gorgoulis and T. D. Halazonetis, Nat. Rev. Mol. Cell Biol., 2010, 11, 220–228 CrossRef CAS PubMed
.
- S. Yuan, J. Almagro and E. Fuchs, Nat. Rev. Cancer, 2024, 24, 274–286 CrossRef CAS PubMed
.
- J. S. Lopez and U. Banerji, Nat. Rev. Clin. Oncol., 2017, 14, 57–66 CrossRef CAS PubMed
.
- M. Puglisi, L. R. Molife, M. J. de Jonge, K. H. Khan, L. V. Doorn, M. D. Forster, M. Blanco, M. Gutierrez, C. Franklin, T. Busman, J. Yang and F. A. Eskens, Future Oncol., 2021, 17, 2747–2758 CrossRef CAS PubMed
.
- C. Vernieri, M. Prisciandaro, M. Milano, M. S. Cona, C. Maggi, M. Brambilla, A. Mennitto, C. Fabbroni, E. Farè, S. Cresta, L. Celio, G. Mariani, G. Bianchi, G. Capri and F. de Braud, Clin. Breast Cancer, 2019, 19, e306–e318 CrossRef CAS PubMed
.
- L. M. Dredze, M. Friger, S. Ariad, M. Koretz, B. Delgado, R. Shaco-Levy, M. Tokar, M. Bayme, R. Agassi, M. Rosenthal, V. Dyomin, O. Belochitski, S. Libson, T. Mizrahi and D. B. Geffen, Breast Cancer Res. Treat., 2022, 193, 597–612 CrossRef CAS PubMed
.
- P. Martin-Romano, I. Baraibar, J. Espinós, J. Legaspi, J. M. López-Picazo, J. M. Aramendía, O. A. Fernández and M. Santisteban, Breast J., 2018, 24, 473–479 CrossRef CAS PubMed
.
- T. C. Chou, Leuk. Lymphoma, 2008, 49, 2059–2080 CrossRef PubMed
.
- M. R. Carvalho, D. Barata, L. M. Teixeira, S. Giselbrecht, R. L. Reis, J. M. Oliveira, R. Truckenmüller and P. Habibovic, Sci. Adv., 2019, 5, eaaw1317 CrossRef CAS PubMed
.
- R. S. Narayan, P. Molenaar, J. Teng, F. M. G. Cornelissen, I. Roelofs, R. Menezes, R. Dik, T. Lagerweij, Y. Broersma, N. Petersen, J. A. Marin Soto, E. Brands, P. van Kuiken, M. C. Lecca, K. J. Lenos, S. In 't Veld, W. van Wieringen, F. F. Lang, E. Sulman, R. Verhaak, B. G. Baumert, L. J. A. Stalpers, L. Vermeulen, C. Watts, D. Bailey, B. J. Slotman, R. Versteeg, D. Noske, P. Sminia, B. A. Tannous, T. Wurdinger, J. Koster and B. A. Westerman, Nat. Commun., 2020, 11, 2935 CrossRef CAS PubMed
.
- H. Hwangbo, S. C. Patterson, A. Dai, D. Plana and A. C. Palmer, Nat. Cancer, 2023, 4, 1693–1704 CrossRef CAS PubMed
.
- K. Peters, A. Lerma Clavero, F. Kullenberg, M. Kopsida, D. Dahlgren, F. Heindryckx, H. Lennernäs and M. Sjöblom, PLoS One, 2024, 19, e0307414 CrossRef CAS PubMed
.
- P. Jaaks, E. A. Coker, D. J. Vis, O. Edwards, E. F. Carpenter, S. M. Leto, L. Dwane, F. Sassi, H. Lightfoot, S. Barthorpe, D. van der Meer, W. Yang, A. Beck, T. Mironenko, C. Hall, J. Hall, I. Mali, L. Richardson, C. Tolley, J. Morris, F. Thomas, E. Lleshi, N. Aben, C. H. Benes, A. Bertotti, L. Trusolino, L. Wessels and M. J. Garnett, Nature, 2022, 603, 166–173 CrossRef CAS PubMed
.
- M. Binnewies, E. W. Roberts, K. Kersten, V. Chan, D. F. Fearon, M. Merad, L. M. Coussens, D. I. Gabrilovich, S. Ostrand-Rosenberg, C. C. Hedrick, R. H. Vonderheide, M. J. Pittet, R. K. Jain, W. Zou, T. K. Howcroft, E. C. Woodhouse, R. A. Weinberg and M. F. Krummel, Nat. Med., 2018, 24, 541–550 CrossRef CAS PubMed
.
- J. Kreuger and M. Phillipson, Nat. Rev. Drug Discovery, 2016, 15, 125–142 CrossRef CAS PubMed
.
- H. Hwangbo, S. Chae, W. Kim, S. Jo and G. H. Kim, Theranostics, 2024, 14, 33–55 CrossRef CAS PubMed
.
- J. Rodrigues, M. A. Heinrich, L. M. Teixeira and J. Prakash, Trends Cancer, 2021, 7, 249–264 CrossRef CAS PubMed
.
- P. Datta, M. Dey, Z. Ataie, D. Unutmaz and I. T. Ozbolat, npj Precis. Oncol., 2020, 4, 18 CrossRef PubMed
.
- Y. Wang and H. Jeon, Trends Pharmacol. Sci., 2022, 43, 569–581 CrossRef CAS PubMed
.
- N. Choo, S. Ramm, J. Luu, J. M. Winter, L. A. Selth, A. R. Dwyer, M. Frydenberg, J. Grummet, S. Sandhu, T. E. Hickey, W. D. Tilley, R. A. Taylor, G. P. Risbridger, M. G. Lawrence and K. J. Simpson, SLAS Discovery, 2021, 26, 1107–1124 CrossRef CAS PubMed
.
- V. Brancato, J. M. Oliveira, V. M. Correlo, R. L. Reis and S. C. Kundu, Biomaterials, 2020, 232, 119744 CrossRef CAS PubMed
.
- R. Fevre, G. Mary, N. Vertti-Quintero, A. Durand, R. F. Tomasi, E. Del Nery and C. N. Baroud, iScience, 2023, 26, 106651 CrossRef CAS PubMed
.
- M. Yu, S. K. Selvaraj, M. M. Liang-Chu, S. Aghajani, M. Busse, J. Yuan, G. Lee, F. Peale, C. Klijn, R. Bourgon, J. S. Kaminker and R. M. Neve, Nature, 2015, 520, 307–311 CrossRef CAS PubMed
.
- J. Schindelin, I. Arganda-Carreras, E. Frise, V. Kaynig, M. Longair, T. Pietzsch, S. Preibisch, C. Rueden, S. Saalfeld, B. Schmid, J.-Y. Tinevez, D. J. White, V. Hartenstein, K. Eliceiri, P. Tomancak and A. Cardona, Nat. Methods, 2012, 9, 676–682 CrossRef CAS PubMed
.
- V. Sharma, Zenodo, https://zenodo.org/record/2252521#.YzagunZBz-g, 2018.
- D. Duarte and N. Vale, Curr. Res. Pharmacol. Drug Discov., 2022, 3, 100110 CrossRef PubMed
.
- J. Foucquier and M. Guedj, Pharmacol. Res. Perspect., 2015, 3, e00149 CrossRef PubMed
.
- N. Fatsis-Kavalopoulos, R. Roemhild, P.-C. Tang, J. Kreuger and D. I. Andersson, PLoS Biol., 2020, 18, e3000856 CrossRef CAS PubMed
.
- O. Lieleg and K. Ribbeck, Trends Cell Biol., 2011, 21, 543–551 CrossRef CAS PubMed
.
- O. Lieleg, R. M. Baumgärtel and A. R. Bausch, Biophys. J., 2009, 97, 1569–1577 CrossRef CAS PubMed
.
- A. Pluen, P. A. Netti, R. K. Jain and D. A. Berk, Biophys. J., 1999, 77, 542–552 CrossRef CAS PubMed
.
- O. Degerstedt, P. O'Callaghan, A. L. Clavero, J. Gråsjö, O. Eriksson, E. Sjögren, P. Hansson, F. Heindryckx, J. Kreuger and H. Lennernäs, Drug Delivery Transl. Res., 2024, 14, 970–983 CrossRef CAS PubMed
.
- M. J. Moore, F. Bodera, C. Hernandez, N. Shirazi, E. Abenojar, A. A. Exner and M. C. Kolios, Nanoscale, 2020, 12, 21420–21428 RSC
.
- H. Nosrati, M. Khodaei, Z. Alizadeh and M. Banitalebi-Dehkordi, Int. J. Biol. Macromol., 2021, 192, 298–322 CrossRef CAS PubMed
.
-
G. T. Hermanson, in Bioconjugate Techniques, ed. G. T. Hermanson, Academic Press, Boston, 3rd edn, 2013, pp. 395–463, DOI:10.1016/B978-0-12-382239-0.00010-8
.
- M. Kadlec, M. Pekař and J. Smilek, Colloids Surf., A, 2024, 700, 134791 CrossRef CAS
.
- F. B. Legesse, O. Chernavskaia, S. Heuke, T. Bocklitz, T. Meyer, J. Popp and R. Heintzmann, J. Microsc., 2015, 258, 223–232 CrossRef CAS PubMed
.
- A. Bisht, D. Avinash, K. K. Sahu, P. Patel, G. Das Gupta and B. D. Kurmi, Drug Delivery Transl. Res., 2025, 15, 102–133 CrossRef PubMed
.
- O. Degerstedt, J. Gråsjö, A. Norberg, E. Sjögren, P. Hansson and H. Lennernäs, Eur. J. Pharm. Sci., 2022, 172, 106150 CrossRef CAS PubMed
.
- O. Hovorka, V. Šubr, D. Větvička, L. Kovář, J. Strohalm, M. Strohalm, A. Benda, M. Hof, K. Ulbrich and B. Říhová, Eur. J. Pharm. Biopharm., 2010, 76, 514–524 CrossRef CAS PubMed
.
- O. Degerstedt, P. O'Callaghan, A. L. Clavero, J. Gråsjö, O. Eriksson, E. Sjögren, P. Hansson, F. Heindryckx, J. Kreuger and H. Lennernäs, Drug Delivery Transl. Res., 2024, 14, 970–983 CrossRef CAS PubMed
.
- E. Lilienberg, I. R. Dubbelboer, A. Karalli, R. Axelsson, T. B. Brismar, C. Ebeling Barbier, A. Norén, F. Duraj, M. Hedeland, U. Bondesson, E. Sjögren, P. Stål, R. Nyman and H. Lennernäs, Mol. Pharmaceutics, 2017, 14, 448–458 CrossRef CAS PubMed
.
- E. Sjögren, T. L. Tammela, B. Lennernäs, K. Taari, T. Isotalo, L.-Å. Malmsten, N. Axén and H. Lennernäs, Mol. Pharmaceutics, 2014, 11, 3097–3111 CrossRef PubMed
.
- E. Lilienberg, I. R. Dubbelboer, A. Karalli, R. Axelsson, T. B. Brismar, C. Ebeling Barbier, A. Norén, F. Duraj, M. Hedeland, U. Bondesson, E. Sjögren, P. Stål, R. Nyman and H. Lennernäs, Mol. Pharmaceutics, 2017, 14, 448–458 CrossRef CAS PubMed
.
- R. Bertrand, E. Solary, P. O'Connor, K. W. Kohn and Y. Pommier, Exp. Cell Res., 1994, 211, 314–321 CrossRef CAS PubMed
.
- C. A. Belmokhtar, J. Hillion and E. Ségal-Bendirdjian, Oncogene, 2001, 20, 3354–3362 CrossRef CAS PubMed
.
- P. O’Callaghan, A. Engberg, O. Eriksson, N. Fatsis-Kavalopoulos, C. Stelzl, G. Sanchez, O. Idevall-Hagren and J. Kreuger, J. Cell Sci., 2022, 135(7), jcs258809 CrossRef PubMed
.
- N. N. Mohamad Anuar, N. S. Nor Hisam, S. L. Liew and A. Ugusman, Front. Pharmacol., 2020, 11, 564108 CrossRef PubMed
.
- M. L. Alvarellos, J. Lamba, K. Sangkuhl, C. F. Thorn, L. Wang, D. J. Klein, R. B. Altman and T. E. Klein, Pharmacogenet. Genomics, 2014, 24, 564–574 CrossRef CAS PubMed
.
- J. M. Cleary, C. M. S. R. Lima, H. I. Hurwitz, A. J. Montero, C. Franklin, J. Yang, A. Graham, T. Busman, M. Mabry, K. Holen, G. I. Shapiro and H. Uronis, Invest. New Drugs, 2014, 32, 937–945 CrossRef CAS PubMed
.
- https://www.ebi.ac.uk/chembl/compound_report_card/CHEMBL888/ .
- D. R. Liston and M. Davis, Clin. Cancer Res., 2017, 23, 3489–3498 CrossRef CAS PubMed
.
- E. F. Choo, J. Boggs, C. Zhu, J. W. Lubach, N. D. Catron, G. Jenkins, A. J. Souers and R. Voorman, Drug Metab. Dispos., 2014, 42, 207–212 CrossRef CAS PubMed
.
-
M. D. Wendt, in Protein-Protein Interactions, ed. M. D. Wendt, Springer Berlin Heidelberg, Berlin, Heidelberg, 2012, pp. 231–258, DOI:10.1007/978-3-642-28965-1_7
.
- M. Asmani, S. Velumani, Y. Li, N. Wawrzyniak, I. Hsia, Z. Chen, B. Hinz and R. Zhao, Nat. Commun., 2018, 9, 2066 CrossRef PubMed
.
- A. Fernández-Colino, L. Iop, M. S. Ventura Ferreira and P. Mela, Adv. Drug Delivery Rev., 2019, 146, 17–36 CrossRef PubMed
.
- C. Calitz, J. Rosenquist, O. Degerstedt, J. Khaled, M. Kopsida, M. Fryknäs, H. Lennernäs, A. Samanta and F. Heindryckx, Sci. Rep., 2023, 13, 748 CrossRef CAS PubMed
.
- G. S. Hussey, J. L. Dziki and S. F. Badylak, Nat. Rev. Mater., 2018, 3, 159–173 CrossRef CAS
.
- M. W. Pickup, J. K. Mouw and V. M. Weaver, EMBO Rep., 2014, 15, 1243–1253 CrossRef CAS PubMed
.
- S. Bakhshandeh, U. Heras, H. M. Taïeb, A. R. Varadarajan, S. M. Lissek, S. M. Hücker, X. Lu, D. S. Garske, S. A. E. Young, A. Abaurrea, M. M. Caffarel, A. Riestra, P. Bragado, J. Contzen, M. Gossen, S. Kirsch, J. Warfsmann, K. Honarnejad, C. A. Klein and A. Cipitria, Sci. Adv., 2024, 10, eadr3997 CrossRef CAS PubMed
.
- J. Komen, E. Y. Westerbeek, R. W. Kolkman, J. Roesthuis, C. Lievens, A. van den Berg and A. D. van der Meer, Lab Chip, 2020, 20, 3167–3178 RSC
.
- S. Chakrabarty, W. F. Quiros-Solano, M. M. P. Kuijten, B. Haspels, S. Mallya, C. S. Y. Lo, A. Othman, C. Silvestri, A. van de Stolpe, N. Gaio, H. Odijk, M. van de Ven, C. M. A. de Ridder, W. M. van Weerden, J. Jonkers, R. Dekker, N. Taneja, R. Kanaar and D. C. van Gent, Cancer Res., 2022, 82, 510–520 CrossRef CAS PubMed
.
- E. Steinberg, R. Friedman, Y. Goldstein, N. Friedman, O. Beharier, J. A. Demma, G. Zamir, A. Hubert and O. Benny, Commun. Biol., 2023, 6, 1157 CrossRef PubMed
.
| This journal is © The Royal Society of Chemistry 2026 |