Open Access Article
Open Access ArticleLab-on-a-chip for enzyme activity monitoring in industrial solid-state fermentation processes compatible with R2R fabrication
Verónica
Mora-Sanz
 *a,
Alvaro
Conde
b,
Elisabeth
Hengge
*a,
Alvaro
Conde
b,
Elisabeth
Hengge
 c,
Conor
O'Sullivan
d,
Andoni
Rodriguez
e,
Caroline
Hennigs
f,
Maciej
Skolimowski
b,
Nastasia
Okulova
d,
Jan
Kafka
d,
Bernd
Nidetzky
c,
Conor
O'Sullivan
d,
Andoni
Rodriguez
e,
Caroline
Hennigs
f,
Maciej
Skolimowski
b,
Nastasia
Okulova
d,
Jan
Kafka
d,
Bernd
Nidetzky
 c,
Ana
Ayerdi
c,
Ana
Ayerdi
 a,
Matija
Strbac
a,
Martin
Smolka
a,
Matija
Strbac
a,
Martin
Smolka
 g,
Goran
Bijelic
a and
Nerea
Briz
*a
g,
Goran
Bijelic
a and
Nerea
Briz
*a
aTECNALIA, Basque Research and Technology Alliance (BRTA), Mikeletegi Pasealekua 2, 20009 Donostia-San Sebastian, Spain. E-mail: veronica.mora@tecnalia.com; nerea.briz@tecnalia.com
bMicronit BV, Colosseum 15 7521 PV, Enschede, The Netherlands
cInstitute of Biotechnology and Biochemical Engineering, Graz University of Technology, Petersgasse 12, Graz 8010, Austria
dInmold A/S, Teglbuen 10 2990, Nivå, Denmark
eBionic surface technologies GmbH, Liebenauer Hauptstraße 2-6, 8041 Graz, Austria
fNaturstoff-Technik GmbH, Marie-Curie-Straße 11, 27711 Osterholz-Scharmbeck, Germany
gJOANNEUM RESEARCH Forschungsgesellschaft mbH, Materials-Institute for Surface Technologies and Photonics, A-8160 Weiz, Austria
First published on 30th January 2026
Abstract
We present a disposable lab-on-a-chip (LoC) for colorimetric enzyme activity monitoring in solid-state fermentation (SSF) processes. The microfluidic chip structures are fabricated via roll-to-roll (R2R) extrusion coating, which reduces costs and enhances efficiency. The LoC operates on capillary-driven flow microfluidics in which a droplet added at the inlet self-fills the chip by capillary action, reaching the reaction chamber. A capillary pump then removes excess liquid, isolating the detection area where the enzymatic reaction takes place. The selection of the target enzymes (α-amylase and cellulase) was made based on their relevance to the industrial biodetergent production processes. For LoC compatibility, enzymatic assays must deliver a strong signal and must be user-friendly. One-step colorimetric assays meet these criteria by releasing a dye from a substrate through enzymatic action. To make the chip easier to handle, the enzymatic substrates were integrated into its reaction chamber in dryed form. For this purpose, two strategies for integration were tested: drop-casting followed by freeze-drying, and piezoelectric deposition with air-drying. Additionally, storage conditions were optimized to enhance shelf-life and reagent stability. To measure enzymatic activity, a pocket-sized colorimetric reader was developed and adapted to the LoC geometry while an Android app was created to enable smartphone-based control of the reader. Furthermore, validation with commercial enzymes established the limit of detection (LoD), and subsequent tests with SSF samples from an industrial plant confirmed the functionality of the system. The enzymatic activity measurements are completed in under 10 minutes, revealing increasing enzymatic activity as fermentation progresses. In conclusion, the LoC provides a quick and cost-effective solution for detecting α-amylase and cellulase in samples derived from SSF processes.
1. Introduction
Industrial enzymes are biocatalysts that accelerate chemical reactions to obtain valuable products.1 They are employed across various industries, including food,2 beverage,3 pharmaceutical,4 biofuels,5 personal care,6 biological laundry detergents7 and other commercial sectors.8 Biocatalysts are replacing traditional chemical catalysts due to efficient kinetics, economic benefits and lower toxicity and by-product generation. Additionally, the sector has experienced rapid growth in recent years due to the ability of enzymes to operate under mild conditions, with high specificity and productivity.9The techniques employed for large-scale enzyme production are submerged fermentation (SmF) and solid-state fermentation (SSF). In SmF, microorganisms are cultivated in a stirred and aerated liquid medium under tightly controlled physicochemical conditions, whereas SSF involves microbial growth on moist solid substrates with little or no free water.10 These distinct environments lead to different process performances and operational requirements.
In recent years, the SSF technique has gained popularity due to its simplicity, cost-effectiveness, and eco-friendly nature.11 The advantages include high enzyme yields, the use of nutrient-rich agricultural waste as substrate, minimal need for process parameter regulation, lower effluent waste and foam production, and straightforward purification of final products.12 These features make SSF particularly attractive for large-scale enzyme production, but they also underline the importance of appropriate monitoring strategies to ensure consistent process performance.
Monitoring enzymatic activity at different stages is highly desirable to control process efficiency and product quality.
However, in current practice, monitoring is largely limited to environmental parameters13 such as temperature, pH, water content and carbon cycle-related variables (e.g., biomass production, CO2 concentration). The ability to monitor enzymatic activity in real time during SSF would enable rapid adjustment of process conditions, optimization of production and improved batch-to-batch consistency.
Currently no practical tools exist for rapid, on-site enzyme activity monitoring in SSF. Main challenges in measuring enzymatic activity in SSF include the need for specialized equipment and the use of methods that are often labor-intensive, time-consuming and costly. These methods typically require trained personnel and, due to their complexity, are often unsuitable for rapid on-site measurements, limiting the ability for real-time monitoring and immediate process adjustments. Several studies on lignocellulolytic enzymes and cellulase production in SSF have emphasized that accurate activity measurements rely on expensive, time-intensive protocols, which are difficult to implement at industrial scale.14,15 In addition, quality control workflows are hindered by delays associated with sending samples to centralized laboratories and waiting for results, which postpones batch release and increases storage and logistics costs.
In light of these challenges, lab-on-a-chip (LoC) technologies emerge as a promising solution, revolutionizing chemical and biological analysis through miniaturized, portable devices that enable fast and affordable measurements.16 These devices facilitate the automatization of standard and routine laboratory procedures and the implementation of chemical and biochemical analysis in a miniaturized system. This approach has been made possible by the technology of microfluidics through manipulation of small fluid volumes in channels with micrometre-scale dimensions.17 The potential of LoC technology for monitoring fermentation processes has been demonstrated by the measurement of biomolecule concentrations, like glucose, glutamate and glutamine.18,19
Here, we present a user-friendly, microfluidic disposable LoC for colorimetric enzyme activity monitoring in industrial SSF processes. To the best of our knowledge, this is the first study of microfluidic sensor system specifically designed for enzyme activity measurements in biofermentation processes, with a particular focus on SSF-based detergent production.
This study focuses on laundry enzymes detection in real samples from SSF processes. Nowadays, main application of enzymes is in the laundry industry, accounting for 30–40% of the total enzyme market.20 The major classes of detergent enzymes are proteases,21 lipases,22 α-amylase,23 cellulase,24 mannanases.25 Among these, α-amylase and cellulase are of particular interest in SSF-based biodetergent manufacturing due to their key roles in starch removal and fabric care and therefore were selected as target enzymes in this work. Enzyme-based detergents have evolved from early formulations with animal-derived proteases to modern products containing stable microbial proteases, α-amylases, cellulases and lipases.26–28 For implementation in a LoC devices, enzymatic assays should provide a robust signal output and easy handling for the final user. These requirements are fulfilled by one-step colorimetric assays, where a dye is released from a substrate by enzymatic action.
The LoC device is fabricated as a disposable microfluidic chip fabricated on cyclic olefin copolymer (COC) foil using a scalable roll-to-roll process, which is compatible with mass production and optical detection. The resulting microfluidic chips on polymer foil substrates are also referred as lab-on-a-foil devices.29 The reagents required for the colorimetric assays are pre-stored in dried form within the chip to simplify handling for the final user and enable point-of-need measurements. This study includes an assessment of the stability of the integrated reagents. A customized portable colorimetric reader, specifically developed for this work, operated via a smartphone application was used to acquire and process the assay signals, making the system suitable for on-site measurements.
This work presents the design, laboratory environment validation, and demonstration of the developed LoC system for enzymatic measurement in samples derived from relevant SSF processes. First, we characterize the analytical performance of the platform using commercial enzymes, including the limit of detection for α-amylase and cellulase. Finally, the whole system has been validated in a relevant environment with SSF samples obtained from the manufacturing process of a biodetergent producer, demonstrating its applicability for in situ monitoring of changes in amylase and cellulase enzymatic activity throughout the chosen fermentation process.
2. Materials
α-Amylase from Aspergillus oryzae (10065), α-glucosidase from Saccharomyces cerevisiae (G5003), Cellulase from Aspergillus niger (22178), β-hydroxy-4-morpholinepropanesulfonic acid (MOPSO) (M8389), D-(+)-trehalose dihydrate (T0167-100G), 4-nitrophenyl phosphate disodium salt hexahydrate (pNP) (71768), Triton X-100 (1086431000), dimethyl sulfoxide (DMSO) (D2438), COC 96-well plate (M3812), Amicon Ultra 10 K filters (UFC501024) and 3 mm glass beads (1040150500) were purchased from Merck. Resorufin-β-D-cellobioside (ab274307) and resorufin (ab244308) were supplied by Abcam. Ethylidene-4-nitrophenyl-α−D-maltoheptaoside (pNP-G7) (PG7-01C) was obtained from Sorachim. Tween-20 (TW00200250) was purchased from Scharlab and sodium acetate (141632) from PanReac. The hydrophilic coating S100 was supplied by Joninn. The pressure-sensitive adhesive (94090) was obtained from Adhesive Research.3. Methods
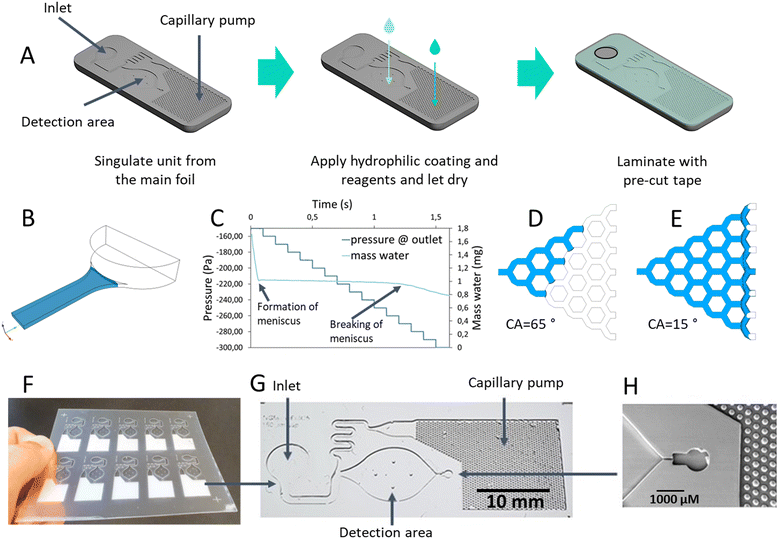
The development of the LoC system comprised a sequential and integrated workflow that fulfil the whole value chain (Fig. 1). First, the enzymatic assays were established and optimized in microplate format to define the conditions required for the implementation on the LoC. Second, reagent stability was evaluated under different storage conditions. In parallel, the microfluidic concept was designed and optimized through computational fluid dynamics (CFD) simulations, followed by roll-to-roll (R2R) extrusion-coating production of the chips. Subsequently, biofunctionalization strategies were developed for integrating the enzymatic reagents onto the foil-based device. Later the backend processes include the application of a hydrophilic coating in the capillary pump and closing the chip with a laser precut pressure sensitive tape. A portable colorimetric reader was then designed to enable real-time kinetic measurements including the calculation of enzymatic activity. The analytical workflow concludes with, first the functional validation of the system using commercial enzymes and finally with real SSF samples.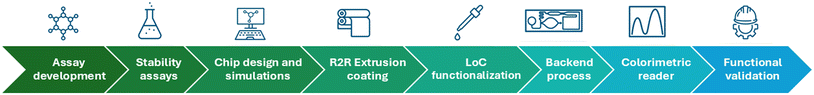 | ||
| Fig. 1 Sequential workflow for the design, fabrication, and validation of the microfluidic lab-on-a-foil chip. | ||
3.1. Assay development
The colorimetric enzymatic reactions selected should be compatible with a LoC produced by R2R extrusion coating process. Due to the inherent volumetric limits of foil-based devices steps like filtration or precipitation are difficult to integrate. In view of this, one step assays were chosen to simplify the process and to take advantages of synergies in chip design. In both enzymatic reactions, a specific substrate is added and then by the action of the enzyme an optical measurable compound is released. UV-visible measurements in 96-well microplate geometry were performed on a Synergy H1 microplate reader (BioTek) controlled by BioTek Gen5 Software during assay development prior to the LoC implementation.3.2. Stability assays experiments
The stability of a microfluidic LoC reagent refers to its ability to maintain expected and consistent performance over time without degradation. It relates to performance of a product within a specified time. To investigate the stability of the enzymatic reagents (pNP-G7 and α-glucosidase for the α-amylase assay, and resorufin-β-D-cellobioside for the cellulase assay), tests were conducted at different temperatures and storage durations (equivalent to 1, 3, and 6 months). Additionally, the effects of trehalose as a preservative, storage under inert atmosphere, and the use of silica gel to prevent humidity were examined. To simulate storage in an inert atmosphere with silica gel, a heat-sealable airtight bag was used. The bag was filled with argon and included a silica gel packet before being sealed. For evaluating the storage temperatures of 4 °C and 23 °C, accelerated stability tests were performed at 37 °C, with the equivalence to real-time storage calculated according to the standard guide for accelerated aging of sterile barrier systems for medical devices (ASTM). A detailed protocol can be found in the SI for the calculation of the accelerated aging time. The −20 °C condition was not simulated but analysed in real-time.For this, the enzymatic substrates were deposited in the surface of a COC 96-well plate by drop casting followed by freeze-drying. For the α-amylase reaction, 100 μL of a solution containing 10 mM pNP-G7 and 2 mg mL−1 α-glucosidase in 0.05 mM MOPSO buffer at pH 7 was used. For the cellulase reaction, 100 μL of a 0.5 mM resorufin-β-D-cellobioside solution in 0.5 mM acetate buffer at pH 6 was lyophilized. In the conditions where trehalose was added, it was incorporated to a final concentration of 2.5% (w/v). The reagents were applied via drop casting into the wells of the microplate, immediately frozen at −40 °C, and freeze-dried overnight (ON) at 0 °C and 12 mbar.
After the storage period, fresh enzyme was added, and the enzymatic activity was measured. In both cases, commercial enzymes were used: for the α-amylase reaction, 100 μL of 1 mg mL−1 enzyme solution in 0.05 mM MOPSO buffer at pH 7 was added, and for the cellulase reaction, 100 μL of 4 mg mL−1 enzyme solution in 0.5 mM acetate buffer at pH 6 was used. The enzymatic activity was then compared to that obtained with fresh reagents.
3.3. Lab-on-a-chip concept and simulations
The LoC works on the principle of capillary-driven microfluidics. The assembly process is shown in Fig. 3A. The chip is made of two flexible transparent layers: the bottom layer contains microstructures, while the top layer is made from pressure-sensitive adhesive tape, which is structured by laser cutting. The chip design consists of an inlet, a detection area, and a capillary pump (Fig. 3A). This pump was chosen to enable the autonomous flow of samples trough the LoC. Incorporating capillary pumps into LoC systems offers several advantages: enabling passive fluid transport by harnessing surface tension, eliminating the need for external power sources and reducing system complexity. This passive mechanism not only simplifies the device design but also enhances portability, making LoC devices more suitable for point-of-need applications.33The chip operates as follow: first, the user adds a drop of the solution containing the enzyme into the inlet, and the chip self-fills by capillary action. The sample reaches the reaction chamber, where the reaction substrates are deposited. The flow continues until it reaches the capillary pump, which is coated with a hydrophilic material that removes the excess liquid and isolates the detection area where the enzymatic reaction takes place. A schematic of the chip filling process is shown Fig. S2 in the SI.
As fluid is placed at the inlet, it is induced into the channel solely due to capillary effects. The so-called meniscus is formed inside the channel due to the surface tension and the contact angle of the surface, hence creating an under pressure that drives the fluid into the chip. At some point, as most of the fluid at the inlet has been already absorbed, a new meniscus forms exactly at the transition between the inlet chamber and the channel due to the sharp edge of the geometry. That new meniscus will stabilize, opposing that which drives the fluid in. Overcoming that meniscus is key for the filling to proceed, and so is the knowing of the suction pressure required to do so. The capillary pressure needed to overcome the inlet meniscus, what is referred to as “breaking” it, was obtained using CFD simulations, namely a simplified digital model of the inlet. A transient simulation was set in a way that an increasing suction pressure was applied, which allowed the emptying of the water from the simulation domain, while monitoring the inlet meniscus. CFD simulations were also used to optimize the design of the capillary pump, which is responsible for breaking the meniscus in the first place. The process involved testing various geometries to achieve the best balance between performance and the requirements of R2R fabrication.
3.4. R2R extrusion coating production of the chips
The chip developed for this work was produced using R2R imprinting, an approach for high throughput fabrication of microfluidic chip components which got recent attention in various studies.34–37 In this case, R2R extrusion coating was applied,37 which is a continuous process for manufacturing flexible thermoplastic materials in rolls. A modified R2R extrusion coating process was used to fabricate the microfluidic chips. A negative of the desired microfluidic pattern was mounted onto a cooling roller. Molten thermoplastic material was continuously extruded into a gap (nip) between the cooling roller and a second roller, referred to as the pressure roller. The pressure roller squeezes the molten thermoplastic film against the mold, ensuring sufficient filling and transfer of the microfluidic design. As the thermoplastic travels along the circumference of the cooling roller it cools and hardens. Removal of the solidified thermoplastic from the cooling roller occurs and the imprinted foil is wound onto a collection roller, freeing the mold before it again reaches the nip. The process continues in a continuous manner in which the supplied molten thermoplastic is constantly imprinted, cooled, removed and stored.38This method offers several advantages: high speed and scalability for enhanced efficiency and productivity, cost reductions through optimized material use and energy efficiency, flexibility for diverse applications and integrated processes, consistent quality with improved process control and fewer defects, support for rapid prototyping and advanced technologies, and reduced environmental impact through lower waste and energy consumption39,40 Cyclic olefin copolymer (COC) was chosen for the chip fabrication due to its flexibility required in R2R manufacturing, its thermoplastic nature needed for the extrusion coating and its transparency necessary for colorimetric detection.41
3.5. Biofunctionalization strategies
The immobilization of the various reagents on the chip reduces the need for user interaction with the LoC, making the device easier to handle. Incorporating reagents into LoC devices can be achieved through different methods such as drop casting followed by freeze-drying or piezoelectric deposition with subsequent air drying.The integration of reagents for the α-amylase was carried out by the former method. Drop casting involves depositing a solution containing the reactive component onto the chip surface, followed by freeze-drying to preserve the stability of the components and activity. This technique effectively preserves the reagents, allowing for their storage and immediate use upon rehydration. It has been employed to integrate biomolecules into microfluidic devices, enhancing their shelf life and functionality.42 In this work 5 μL of 100 mM pNP-G7 and 5 μL of 20 mg mL−1 α-glucosidase directly into the reaction chamber of the chips. This was followed by immediate freezing at −40 °C and ON freeze-drying at 0 °C and 12 mbar.
In contrast, the reagents for the cellulase reaction were introduced by spotting and air drying. Piezoelectric deposition involves the precise spotting of reagent droplets onto specific areas of the chip using piezoelectric dispensers. Following deposition, the reagents are air-dried, which simplifies the integration process and maintains reagent functionality. This method allows for controlled placement and uniform distribution of components, facilitating consistent biochemical reactions within the microfluidic channels. In this work spotting was performed using a sciFLEXARRAYER S12 (Scienion) spotter, equipped with a “PDC90 (uncoated)” nozzle. The spotting conditions were a 10 × 10 microarray, 600 nL per chip, and a 2 mM print solution of resorufin-β-D-cellobioside.
3.6. Portable colorimetric reader and operating procedure
A custom device (Photogent) was developed to track the activity of enzyme extracts obtained through fermentation methods. The device was designed to measure absorbance at two specific wavelengths: 405 nm (for α-amylase) and 570 nm (for cellulase) over time. The measurements are taken in kinetic mode, allowing for continuous monitoring of enzyme activity as the reaction progresses.The system consists of a custom-designed hardware and an Android application that manages measurements. The hardware is composed of five main components: LED light sources, a light detector, a microcontroller unit (MCU) with a Bluetooth module, and a battery supply. The LED light sources include purple (405 nm) and green (570 nm) LEDs, each with a low spectral bandwidth. The light-emitting mode can be selected via the Android app, based on the wavelength required for the specific enzyme being measured.
The light detector functions as a transimpedance amplifier, where the variations in light intensity, due to enzymatic reactions, induce changes in the current through the amplifier. The absorbance values are monitored in real-time via the Android App.
The device is powered by a lithium-ion battery, which can be recharged via a USB-C connection and can support up to 200 hours of operation. Detailed device specifications are provided in the Table S1 of the SI.
The device communicates with the Android app. via a low-energy Bluetooth connection. Before the measurement, several parameters can be set through the app, including the light-emitting mode, streaming period, and total measurement duration.
In practical use, the device operates as showed in Fig. 2. Before each measurement, the microfluidic chip containing the dried reagents is inserted into the detection chamber of the Photogent reader, ensuring correct alignment between the reaction chamber and the optical path. Once the chip is placed, the user opens the Android application and selects the appropriate wavelength mode (405 nm for α-amylase or 570 nm for cellulase). The kinetic acquisition parameters are set through the interface. The sample is then added to the inlet of the microfluidic chip, initiating the reaction. The user closes the lid and starts the measurement via the app, after which the chosen LED illuminates the reaction chamber continuously while the photodetector records changes in transmitted light intensity as the reaction progresses. These signals are processed and transferred to the smartphone in real time. Once the acquisition ends, the app displays the absorbance-time dataset, which can be exported as a .txt file for external analysis or directly processed using the enzyme activity calculation tool. The device does not require additional calibration between measurements and is ready for subsequent assays immediately after data export.
 | ||
| Fig. 2 Operation of the photogent reader. A user inserts the microfluidic chip into the reader while the smartphone app shows the real-time absorbance signal recorded during the enzymatic assay. | ||
3.7. Enzymatic activity calculations
The calculation of enzyme activity for α-amylase and cellulase is based on measuring changes in absorbance at specific wavelengths in kinetic mode, with readings taken at 1 minute intervals. For α-amylase, absorbance is measured at 405 nm, while for cellulase, it is measured at 570 nm. The difference in absorbance (ΔAbs) between two time points (T1 and T2) reflects the amount of product generated by the respective enzyme during the reaction. This difference is interpolated on a calibration curve (pNP for α-amylase, resorufin-β-D-cellobioside for cellulase) to determine the amount of product generated. Finally, enzyme activity is calculated by considering the reaction chamber volume (12 μL), reaction time, and the mass of the biodetergent sample. A detailed protocol can be found in the SI for the calculation of enzyme activity. These calculations for α-amylase and cellulase activity can be made in a digital calculation sheet after downloading the raw data from the smartphone. For calculations in the software app., T1 and T2 were fixed at 0 minutes and 10 minutes, respectively.3.8. Functional validation with commercial enzymes
The functional validation of the α-amylase and cellulase LoC was carried out using commercial enzymes. The enzymes were dissolved in the optimised reaction buffer at different concentrations, and 50 μL of each solution was introduced through the inlet of the microfluidic chip. A calibration curve was constructed, and the LoD was determined. The LoD was calculated as three times the standard deviation of the blank absorbance signal, divided by the slope of the regression equation, yielding the result in mg mL−1. To convert the LoD into α-amylase activity units (μg mL−1), the LoD value in mg mL−1 was interpolated using the calibration curve.3.9. Functional validation with real samples
Real samples obtained at various stages of the fermentation process, each with different enzymatic concentrations, were shipped at room temperature and stored under the same conditions until their measurement. A simple method was developed for the extraction of enzymes from solid biodetergent samples, designed for implementation in an industrial fermentation plant without laboratory facilities and requiring minimal instrumentation. Five grams of the enzyme-containing sample were weighed using a digital spoon and transferred to a 50 mL tube. Next, four grams of 3 mm glass microbeads were added, followed by 15 mL of reaction buffer. The mixture was vortexed for 2 minutes to ensure thorough mixing. Afterward, 1000 μL of the supernatant was collected with a syringe and filtered through a 0.8 μm filter to remove solid residues. Finally, 50 μL of the sample extracted from the biodetergent is introduced into the microfluidic chip inlet.4. Results and discussion
4.1. Microfluidic chip design
The chip design was optimized via CFD simulations, testing different capillary pump geometries until an optimal balance between design and R2R fabrication constraints was achieved. The main task of the capillary pump is to overcome the forces opposing the fluid flow, one of which, very critical, is the creation of a meniscus at the inlet region of the channel. The sharp edge at the inlet chamber-to-channel transition facilitates the creation of a meniscus at the point where enough liquid has already been removed from the inlet chamber. A visual representation of this meniscus, displayed as the liquid-air interface at the inlet, is shown in Fig. 3B. In the simulations, a continuously increasing suction pressure was applied at the created channel outlet. Fig. 3C shows the variation in pressure at the outlet together with the remaining water mass over time. The light blue line represents the changing mass of water, showing a drastic decrease until the meniscus is stabilized at about −220 Pa, strongly impeding the discharge. With the increase of the suction pressure, a value −270 Pa was shown to be needed to overcome the meniscus. At that pressure, the capillary pressure is overcome, allowing the water to flow through the inlet, as indicated by the steeper decrease of the water mass. The events of formation and breaking of the meniscus are also indicated in the chart. This simulation helps to obtain the required pressure to break the meniscus and initiate fluid movement, for which the microfluidic pump is responsible. The designed pump geometry, consisting of 100 μm micropillars, was tested with a hydrophilic coating to enhance liquid flow. COC has a hydrophobic nature with a contact angle of approximately 80° (Fig. 3D). The addition of this coating reduces the contact angle to 15° (Fig. 3E), enabling the breakage of the meniscus and the subsequent removal of excess liquid from the fluidic network, effectively compartmentalizing the detection area chamber. In addition to this geometry, an alternative design featuring irregular hexagons was evaluated using CFD simulations. However, this design did not meet the required performance criteria. The detailed results of this alternative design are provided in the Fig. S5. Similar approaches have been reported in the literature, demonstrating the impact of microstructure geometry on capillary-driven flow. Capillary pumps incorporating micropillars and other patterned microstructures have been shown to influence flow rates and pressure generation in passive fluidic systems.43 Additionally, studies have demonstrated that structured capillary pumps with varying microstructure sizes and arrangements can effectively control fluid movement by modifying capillary pressure.44 Moreover, highly ordered micropillar arrays fabricated in silicon have been used to enhance capillary flow and improve fluidic resistance control.45 Our findings align with these studies, highlighting the importance of pump geometry in ensuring efficient liquid transport and compartmentalization in LoC systems.After optimizing the chip design, the next step was to scale up its production using the R2R extrusion coating process. This method allows for high-throughput manufacturing of the microfluidic chips, ensuring consistent quality and reproducibility across large volumes. A foil, with a size of 80 mm × 85 mm, containing multiple chip units, imprinted in COC via the R2R extrusion coating process is shown in Fig. 3F. The final design of a single chip unit, including the inlet, detection area, and capillary pump, is depicted in Fig. 3G The overall dimensions of the LoC are 40 mm × 15 mm. Additionally, Fig. 3H provides a detailed view (SEM micrograph) of the capillary stop valve, where the 100 μm micropillars are clearly visible.
4.2. Stability assays of the enzymatic substrates
The enzymatic substrates were deposited on the surface of a COC 96-well plate as described in the Methods section. After the storage period, fresh enzyme was added, and the enzymatic activity was measured and compared with the activity obtained using fresh reagents.In the case of α-amylase detection, both the chromogenic substrate pNP-G7 and the auxiliary enzyme α-glucosidase are required in addition to α-amylase. To simplify handling for the end user, both reagents were stored together in the reaction chamber of the chip. Although better stability was achieved when pNP-G7 was stored alone (Fig. S6), the combined formulation was evaluated to enable user-friendly operation. The enzymatic reaction of cellulase, however, only requires the use of a chromogenic substrate (resorufin-β-D-cellobioside).
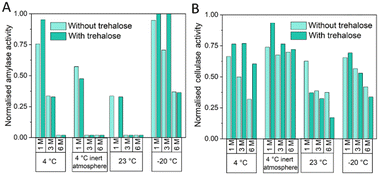
Fig. 4A shows normalized α-amylase activity under various temperature and storage conditions: 4 °C, 4 °C under inert atmosphere, and 23 °C under accelerated conditions, as well as −20 °C in real-time storage, evaluated over 1, 3, and 6 months. Storage at 4 °C under inert atmosphere and with silica gel and at 23 °C are not suitable for preserving the reagents, as α-amylase activity drops below 60% within the first month in all tested conditions. This result could be attributed to over-drying of reagents caused by silica gel application. Since enzyme stability can rely on small amounts of moisture for stability, this can lead to their destabilization. Enzymes like α-glucosidase may lose activity or denature when exposed to excessive drying conditions, such as those caused by silica gel desiccants. This phenomenon has been reported in studies highlighting the negative impact of over-drying on protein stability, where antibodies undergo denaturation and loss of biological activity when exposed to low moisture levels.46 At 4 °C, α-amylase activity declines significantly over time, with trehalose providing slight improvement and maintaining activity above 75% during the first month of storage. At −20 °C, α-amylase activity remains nearly 100% with trehalose throughout the first 3 months of storage. In summary, trehalose enhances stability but long-term preservation is only achieved at −20 °C.
As it is shown in Fig. 4B 23 °C and −20 °C are not adequate temperatures for the storage of freeze dried resorufin-β-D-cellobioside. However, the stability of resorufin-β-D-cellobioside at 4 °C is improved when the substrate is stored under inert atmosphere and with silica gel. Under these conditions resorufin-β-D-cellobioside can be stored for 6 months at 4 °C without decreasing its functionality. The addition of trehalose improves the stability of the substrate.
4.3. Chips self-filling optimization and integration and solubilization of the enzymatic substrates in the reaction chamber of the chip
The integration of the reagents into the reaction chamber of the chip reduces user interaction and simplifies handling. As a result, all reagents were incorporated in dry format directly into the reaction chamber of the chip. Two different biofunctionalization strategies were evaluated to introduce enzymatic substrates: one method involved manual drop casting followed by freeze drying, while the other used reagent deposition through spotting followed by air drying.The integration of the reagents required for the α-amylase reaction (pNP-G7 and α-glucosidase) was performed employing the former method, as described in the Methods section. The self-filling of the chip was optimized using the reaction buffer (MOPSO buffer) with different concentrations of surfactants (0.1%, 0.2%, and 0.3% (v/v) of TWEEN-20 and TRITON X-100. These surfactants were tested to reduce surface tension, improve wettability, and ensure consistent filling of the microfluidic channels. By lowering the surface tension of the liquid, the detergents promoted smoother and faster capillary action, allowing the fluid to flow more efficiently through the microchannels, minimizing air bubbles and enhancing overall performance.47
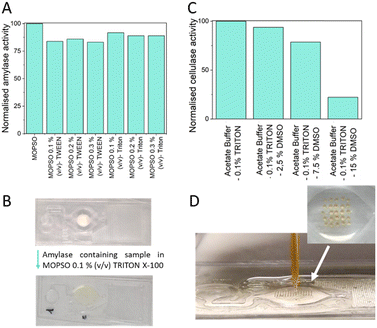
First, the α-amylase reaction was performed using the MOPSO buffer with the different detergent concentrations to ensure that the addition of detergents did not affect the α-amylase activity. The results, shown in Fig. 5A, indicate that none of the tested conditions reduced α-amylase activity by more than 10%. Thus, all conditions are suitable for performing the α-amylase assay.
Next, the self-filling of the chip was tested using the different buffer/surfactant compositions. The reaction chamber filling speed and an image of the chip 10 minutes after filling are shown in Table S2 in the SI. If the surfactant percentage is too low, the flow inside the chip stops, but if it is too high, the liquid overflows and spills out of the reaction chamber. The optimal condition was achieved with 0.1% (v/v) TRITON X-100, as the filling time was appropriate (10 seconds) and the liquid remained within the reaction chamber after 10 minutes.
Finally, the optimized filling buffer (MOPSO with 0.1% (v/v) TRITON X-100) was tested with the freeze-dried reagents. Fig. 5B shows the chip with the reagents lyophilized in the reaction chamber, along with an image of the chip 10 minutes after the introduction of a sample containing commercial α-amylase in MOPSO with 0.1% (v/v) TRITON X-100. The liquid in the reaction chamber turns yellow due to the release of pNP from the action of α-amylase.
Since resorufin-β-D-cellobioside is only soluble in organic-based solutions and not in water-based solutions, introducing the reagents in dry state via dispensing and freeze-drying was not possible, as re-dissolving of the uniformly covered surface was unfeasible. The results of the attempts to solubilize resorufin-β-D-cellobioside in the LoC after drop casting followed by freeze drying are shown in Fig. S7 and S8 of the SI. Therefore, piezoelectric deposition was chosen as the integration method. This approach allows the deposition of small arrays of low volumes, making it easier to dissolve the substrate during the reaction when the enzyme and buffer are added. Resorufin-β-D-cellobioside is soluble in DMSO but has a low solubility in the aqueous buffer in which the cellulase reaction takes place. To ensure complete substrate dissolution, the enzyme had to be added to the chip in a buffer containing DMSO. Thus, it was necessary to determine the maximum percentage of DMSO (2.5%, 7.5%, and 15% (v/v)) that could be included in the reaction buffer (0.5 M acetate buffer, pH 6, 0.1% (v/v) TRITON X-100) without significantly inhibiting cellulase activity (Fig. 5C). The results showed that DMSO inhibits enzymatic activity. The highest DMSO concentration allowing measurable enzymatic activity, with less than a 30% reduction in signal, was 7.5%, which was used in further experiments to balance substrate dissolution and enzymatic activity inhibition by DMSO. Under these conditions, the spotted substrate was successfully re-dissolved. Fig. 5D shows an image of the chip during substrate spotting in the reaction chamber, along with a detailed view of the resorufin-β-D-cellobioside array.
4.4. Laboratory functional validation and chip shelf life
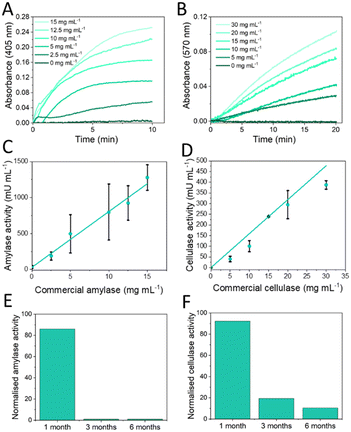
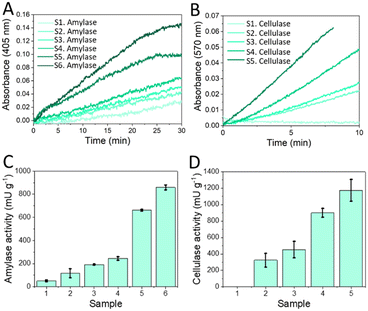
The custom-designed colorimetric reader was employed for the functional laboratory validation of α-amylase and cellulase chips by detecting absorbance changes within the reaction chamber of the chip. Fig. 6A offers an external view of the portable colorimetric reader, highlighting its compact design and portability. Fig. 6B displays the measurement chamber specifically adapted to the geometry of the microfluidic chip, ensuring precise absorbance readings. Lastly, Fig. 6C illustrates the working device with purple emitting mode. In addition, the performance of the portable reader was compared with a commercial microplate reader pNP standard solutions in PBS. For this comparison, the same LoC was measured in both systems by employing a 3D-printed holder that positioned the reaction chamber precisely over a single well of the microplate reader. The portable reader, whose optical design is fully matched to the geometry of the chip, exhibited a slightly higher sensitivity, as reflected in the steeper slope of its calibration curve. The calibration plots are provided in the Fig. S9 of the SI. The functional validation of both the α-amylase and cellulase chips, was performed with commercial enzymes. Six different concentrations of enzymes with three replicates were tested to create the calibration curve. The raw data, directly extracted from the smartphone connected to the chip reader, is shown in Fig. 7A and B for amylase and cellulase respectively. For α-amylase, absorbance at 405 nm was measured over time, showing an increase in the reaction rate as the concentration of α-amylase increased. Similarly, for cellulase, absorbance was recorded at 570 nm, and a similar trend of increasing reaction rate with increasing cellulase concentration was observed. From this data, the enzymatic activity was calculated, and the calibration curves for both enzymes were established, with the calibration curve for α-amylase shown in Fig. 7C and that for cellulase in Fig. 7D. The limit of detection (LOD) was also determined, with α-amylase having an LOD of 136.7 mU mL−1 and cellulase an LOD of 9.2 mU mL−1. Both the LOD and activity ranges for the two enzymes are within the expected values for real samples from the fermentation plant.Additionally, the stability of the reagents was re-evaluated to calculate the shelf life of the chip. Accelerated storage studies were conducted at 37 °C, replicating previously established optimal conditions in which α-amylase reagents were stored for one month at 4 °C, while cellulase reagents were stored at 4 °C in an inert atmosphere and with silica gel for six months.
In this study, stability testing was conducted with the reagents already placed in the reaction chamber of the chip, using drop casting and freeze-drying for α-amylase reagents, and spotting for cellulase enzymatic substrate.
For α-amylase, similar results to those in Fig. 4A were obtained, as the reagents maintained over 80% of their functionality after one month of storage at 4 °C, and activity fell below 35% after three months (Fig. 7E). However, for cellulase, reagent stability decreased significantly when stored on the chip following spotting, with the reagents preserving over 95% of their activity for only one month, then dropping below 35% by the third month (Fig. 7F). This discrepancy with previous results from microplate-based assays (Fig. 4B) is likely due to differences in deposition methods. In prior studies, resorufin-β-D-cellobioside was applied by drop casting and freeze-drying, while in the current study, spotting was used. Since spotting was performed with DMSO, the drops were not completely dried before storage, which may have led to residual solvent remaining on the reagents. This incomplete removal of DMSO could create a microenvironment that destabilizes the cellulase reagents over time, potentially accelerating degradation processes and leading to a more rapid decay in their functional activity than was seen with freeze-dried samples. Moreover, because the droplets are not fully dried, the amount of DMSO added is no longer determined only by the volume introduced with the cellulase buffer. There is an additional contribution of DMSO originated from the incompletely dried droplets, which cannot be quantified. As shown in Fig. 5C, the DMSO concentration influences the enzymatic activity and may therefore also contribute to the observed decrease in activity. For the piezoelectric deposition of resorufin-β-D-cellobioside a DMSO-specific piezo dispense capillary was required (‘PDC90 (uncoated)’ nozzle). Although Fig. 4B demonstrates that the addition of trehalose increases the shelf life of the reagent, this strategy could not be applied here because trehalose caused clogging of the capillary and prevented reproducible dispensing. This represents a known limitation and improving the long-term stability of the formulation will be essential for any future commercialization of the LoC.
4.5. Functional validation with real samples
The enzyme extraction protocol described in the Methods section was used to assess the concentration of α-amylase and cellulase in the samples from the fermentation plant. To evaluate the reproducibility of the extraction step, an initial assessment was performed in which four operators each carried out five consecutive enzyme extractions from a cellulase containing sample. The enzymatic activity of all extracts was measured in a 96-well microplate and the resulting data were analysed using one-way ANOVA test. This test is used to compare the means of three or more independent groups and determine whether there are significant differences among them.No significant differences were detected among operators (ANOVA, p = 0.46), indicating that the sample extraction method does not introduce substantial operational errors under the tested conditions. Further experiments involving a greater number of operators and conditions would be needed to confirm the applicability of the extraction protocol in industrial settings. However, this extended study is out of the scope of this work. The corresponding plot illustrating this analysis is provided in the Fig. S10 of the SI.
Samples were taken at different time points during the microorganism growth phase, where increasing enzymatic activity was expected. The raw kinetic data for both enzymes were downloaded directly from the smartphone connected to the chip reader. For α-amylase, Fig. 8A shows the change in absorbance at 405 nm over time, while for cellulase, Fig. 8B illustrates the absorbance at 570 nm. As fermentation advances, an increase in reaction rate is observed for both enzymes, reflecting higher concentrations of α-amylase and cellulase in the samples. This is consistent with the sample sequence from 1 to 6 for amylase and 1 to 5 for cellulase, where higher sample numbers correspond to more advanced stages of the fermentation process. The data were processed according to the calculations outlined in the Methods section to convert the kinetic measurements into enzymatic activity values. Fig. 8C shows the enzymatic activity for α-amylase and Fig. 8D for cellulase. In the case of sample 1 containing cellulase the enzymatic activity could not be detected because a change in absorbance at 570 nm was not observed over time. The results show that enzymatic activity significantly increases in samples taken at more advanced stages of the fermentation process, indicating higher enzyme production as fermentation progresses. These results indicates that the developed microfluidic chip technology works for quick analysis of enzyme activity in enzyme containing ferments and formulations thereof.
To assess potential cross-reactivity, several control experiments were performed. Extracts rich in cellulase (S3. Cellulase) were incubated with pNP-G7 and monitored at 405 nm, both in the absence (control 1) and presence (control 2) of added glucosidase, while extracts rich in amylase (S3. Amylase) were incubated with pNP-G7 without glucosidase (control 3) and with resorufin-β-D-cellobioside (control 4) at 570 nm. In all cases, no significant increase in absorbance was observed, indicating that there was no cross-reactivity between the enzymes and substrates. A detailed description of these controls and the corresponding results are provided in Fig. S11 in the SI.
Conclusions
In this study, a microfluidic lab-on-a-foil was successfully fabricated using R2R extrusion coating. The device operates via self-filling microfluidics, enabled by a hydrophilic pump consisting of 100 μm pillar-shaped microstructures. To enhance liquid flow, a hydrophilic coating was applied to reduce the contact angle of the COC chip material, facilitating efficient meniscus breakage at the chip inlet. The necessary reagents for measuring α-amylase and cellulase activity were successfully integrated using two distinct strategies based on the nature of the substrates: drop-casting followed by freeze-drying and spotting followed by air-drying. The shelf life of both chip configurations was determined to be one month when stored at 4 °C with the integrated reagents. Additionally, the incorporation of Triton X-100 into the reaction buffer improved chip filling. To detect the enzymatic reaction signal in situ, an absorbance reader was developed, and the LoC system was validated in two stages. First, enzymatic assays with commercial enzymes confirmed the proper functionality of both the chip and the reader. Subsequently, samples from a SSF plant were analysed at different time points during the exponential phase. The system successfully detected increasing enzymatic activities of α-amylase and cellulase over time, demonstrating its applicability for enzyme monitoring in SSF processes. The developed LoC and reader are readily adaptable for other SSF-relevant laundry enzymes that produce a soluble, chromogenic product. For example, lipase activity could be measured using pNP–palmitate, and β-1,4-mannanase activity using pNP–mannotetraoside. In addition to their use in the laundry industry, these enzymes have broad applications across food and beverage, textile, paper, biofuel, pharmaceutical, and industrial sectors, highlighting the versatility and potential of the LoC platform.Author contributions
VMS, AC, AR, CH, CH, MS, NO, JK, BN, MSt, MSm, GB and NB were involved in development of conceptualization and methodology steps of this study. VMS, AC, EH, CO, AR and CH were involved in investigation sections. VMS, AC, AR done validation and visualization. VMS contributed to writing – original draft. VMS, AC, CO, AR, JK, AY, MSm and NB done writing – review and editing.Conflicts of interest
There are no conflicts to declare.Data availability
All relevant data are available from the authors on reasonable request and/or are included within the article and the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5lc00528k.Acknowledgements
This work was supported by the European Union's 2020 research and innovation programme under grant agreement no. 862092 (NextGen Microfluidics project).Notes and references
- S. Wu, R. Snajdrova, J. C. Moore, K. Baldenius and U. T. Bornscheuer, Angew. Chem., Int. Ed., 2021, 60, 88–119 CrossRef CAS PubMed.
- A. Kumar, S. Dhiman, B. Krishan, M. Samtiya, A. Kumari, N. Pathak, A. Kumari, R. E. Aluko and T. Dhewa, Food Prod., Process. Nutr., 2024, 6, 85 CrossRef CAS.
- F. Cosme, A. Inês and A. Vilela, Fermentation, 2023, 9(4), 385 CrossRef CAS.
- G. K. Meghwanshi, N. Kaur, S. Verma, N. K. Dabi, A. Vashishtha, P. D. Charan, P. Purohit, H. S. Bhandari, N. Bhojak and R. Kumar, Biotechnol. Appl. Biochem., 2020, 67, 586–601 CrossRef CAS PubMed.
- P. Kalita, B. Basumatary, P. Saikia, B. Das and S. Basumatary, Energy Nexus, 2022, 6, 100087 CrossRef CAS.
- R. Ciriminna, A. Scurria and M. Pagliaro, Biofuels, Bioprod. Biorefin., 2021, 15, 1604–1610 CrossRef CAS.
- M. A. Farooq, S. Ali, A. Hassan, H. M. Tahir, S. Mumtaz and S. Mumtaz, Arch. Microbiol., 2021, 203, 1281–1292 CrossRef CAS PubMed.
- J. Chapman, A. E. Ismail and C. Z. Dinu, Catalysts, 2018, 8(6), 238 CrossRef.
- F. N. Niyonzima, S. M. Veena and S. S. More, Industrial Production and Optimization of Microbial Enzymes, 2020, vol. 11 Search PubMed.
- A. Fasim, V. S. More and S. S. More, Curr. Opin. Biotechnol., 2021, 69, 68–76 CrossRef CAS PubMed.
- R. R. Singhania, A. K. Patel, C. R. Soccol and A. Pandey, Biochem. Eng. J., 2009, 44, 13–18 CrossRef CAS.
- A. M. Musaalbakri and C. Webb, J. Appl. Biotechnol. Bioeng., 2017, 4, 511–532 Search PubMed.
- V. Bellon-Maurel, O. Orliac and P. Christen, Process Biochem., 2003, 38, 881–896 CrossRef CAS.
- A. Boondaeng, J. Keabpimai, C. Trakunjae, P. Vaithanomsat, P. Srichola and N. Niyomvong, Heliyon, 2024, 10, e26601 Search PubMed.
- V. H. Uchida, J. Waste Resour. Reprocess., 2020, 1, 1–10 Search PubMed.
- S. Gupta, K. Ramesh, S. Ahmed and V. Kakkar, Int. J. Bio-Science Bio-Technology, 2016, 8, 311–322 CrossRef.
- G. M. Whitesides, Nature, 2006, 442, 368–373 CrossRef CAS PubMed.
- M. Bäcker, D. Rakowski, A. Poghossian, M. Biselli, P. Wagner and M. J. Schöning, J. Biotechnol., 2013, 163, 371–376 Search PubMed.
- K. Pontius, D. Semenova, Y. E. Silina, K. V. Gernaey and H. Junicke, Front. Bioeng. Biotechnol., 2020, 8, 1–15 Search PubMed.
- S. Ito, Extremophiles Handbook, 2011 Search PubMed.
- L. Vojcic, C. Pitzler, G. Körfer, F. Jakob, R. Martinez, K. H. Maurer and U. Schwaneberg, New Biotechnol., 2015, 32, 629–634 CrossRef CAS PubMed.
- F. Hasan, A. A. Shah, S. Javed and A. Hameed, Afr. J. Biotechnol., 2010, 9, 4836–4844 CAS.
- R. A. Arfah, S. Sarlan, A. Karim, A. Anita, A. Ahmad, P. Taba, H. Karim, S. H. Larekeng, D. A. Rampisela and R. B. Ladju, Karbala Int. J. Mod. Sci., 2024, 10, 431–441 Search PubMed.
- S. A. Ladeira, E. Cruz, A. B. Delatorre, J. B. Barbosa and M. L. L. Martins, Electron. J. Biotechnol., 2015, 18, 110–115 Search PubMed.
- A. Dawood and K. Ma, Front. Bioeng. Biotechnol., 2020, 8, 1–17 Search PubMed.
- V. N. Jisha, R. B. Smitha, S. Pradeep, S. Sreedevi, K. N. Unni, S. Sajith, P. Priji, M. Sarath Josh and S. Benjamin, Adv. Enzyme Res., 2013, 1, 39–51 Search PubMed.
- P. W. Tardioli, R. Sousa, R. C. Giordano and R. L. C. Giordano, Enzyme Microb. Technol., 2005, 36, 555–564 Search PubMed.
- S. Gurkok, Int. J. Sci. Eng. Res., 2019, 10, 75–81 Search PubMed.
- M. Focke, D. Kosse, C. Müller, H. Reinecke, R. Zengerle and F. Von Stetten, Lab Chip, 2010, 10, 1365–1386 RSC.
- Š. Janeček, B. Svensson and E. A. MacGregor, Cell. Mol. Life Sci., 2014, 71, 1149–1170 CrossRef PubMed.
- R. Visvanathan, M. Qader, C. Jayathilake, B. C. Jayawardana, R. Liyanage and R. Sivakanesan, J. Sci. Food Agric., 2020, 100, 2836–2847 CrossRef CAS PubMed.
- D. J. Coleman, M. J. Studler and J. J. Naleway, Anal. Biochem., 2007, 371, 146–153 CrossRef CAS PubMed.
- E. Khanjani, A. Fergola, J. A. López Martínez, S. Nazarnezhad, J. Casals Terre, S. L. Marasso and B. Aghajanloo, Front. Lab Chip Technol., 2025, 4, 1502127 Search PubMed.
- J. Hiltunen, C. Liedert, M. Hiltunen, O. H. Huttunen, J. Hiitola-Keinänen, S. Aikio, M. Harjanne, M. Kurkinen, L. Hakalahti and L. P. Lee, Lab Chip, 2018, 18, 1552–1559 Search PubMed.
- P. Toren, M. Smolka, A. Haase, U. Palfinger, D. Nees, S. Ruttloff, L. Kuna, C. Schaude, S. Jauk, M. Rumpler, B. Hierschlager, I. Katzmayr, M. Sonnleitner, M. W. Thesen, M. Lohse, M. Horn, W. Weigel, M. Strbac, G. Bijelic, S. Hemanth, N. Okulova, J. Kafka, S. Kostler, B. Stadlober and J. Hesse, Lab Chip, 2020, 20, 4106–4117 RSC.
- C. Liedert, L. Rannaste, A. Kokkonen, O. H. Huttunen, R. Liedert, J. Hiltunen and L. Hakalahti, ACS Sens., 2020, 5, 2010–2017 CrossRef CAS PubMed.
- P. Khumwan, S. Ruttloff, J. Götz, D. Nees, C. O'Sullivan, A. Conde, M. Lohse, C. Wolf, N. Okulova, J. Brommert, R. Benauer, I. Katzmayr, N. Ladenhauf, W. Weigel, M. Skolimowski, M. Sonnleitner, M. Smolka, A. Haase, B. Stadlober and J. Hesse, Biosensors, 2025, 15, 4 Search PubMed.
- J. Park, K. Shin and C. Lee, Int. J. Precis. Eng. Manuf., 2016, 17, 537–550 CrossRef.
- S. Murthy, H. Pranov, H. C. Pedersen and R. Taboryski, J. Vac. Sci. Technol., B, 2026, 34, 06KM02 Search PubMed.
- S. Murthy, M. Matschuk, Q. Huang, N. K. Mandsberg, N. A. Feidenhans'l, P. Johansen, L. Christensen, H. Pranov, G. Kofod and H. C. Pedersen, Adv. Eng. Mater., 2015, 18, 484–489 CrossRef.
- H. Cong and N. Zhang, Biomicrofluidics, 2022, 16(2), 021301 CrossRef CAS PubMed.
- I. Rautsola, M. Haapala, L. Huttunen, O. Korhonen and T. Sikanen, Lab Chip, 2024, 24, 4211 Search PubMed.
- N. Vasilakis, K. I. Papadimitriou, H. Morgan and T. Prodromakis, Microfluid. Nanofluid., 2017, 21, 103 Search PubMed.
- M. Zimmermann, H. Schmid, P. Hunziker and E. Delamarche, Lab Chip, 2007, 7, 119–125 Search PubMed.
- Y. Ye, Y. Zhao, J. Cheng, M. Li and C. Huang, Micro Nano Lett., 2018, 13, 1682–1687 CrossRef CAS.
- A. Duralliu, P. Matejtschuk, P. Stickings, L. Hassall, R. Tierney and D. R. Williams, Pharmaceutics, 2020, 12(4), 303 Search PubMed.
- H. Madadi and J. Casals-Terré, Microsyst. Technol., 2013, 19, 143–150 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |