 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Green reduction of graphene oxide mediated by Sporosarcina pasteurii under harsh conditions, changing the paradigm of rGO production with a non-pathogenic, nanomaterial-resistant bacterium
Massimiliano
Papi†
 abc,
Francesco
Amato†
abc,
Francesco
Amato†
 d,
Andrea Giacomo
Marrani
d,
Andrea Giacomo
Marrani
 d,
Leonardo
Giaccari
d,
Francesca
Sciandra
e,
Marco
De Spirito
ab and
Valentina
Palmieri
d,
Leonardo
Giaccari
d,
Francesca
Sciandra
e,
Marco
De Spirito
ab and
Valentina
Palmieri
 *abc
*abc
aDipartimento di Neuroscienze, Università Cattolica del Sacro Cuore, Largo Francesco Vito 1, 00168 Rome, Italy. E-mail: valentina.palmieri@cnr.it
bFondazione Policlinico Universitario A. Gemelli IRCCS, 00168 Rome, Italy
cIstituto dei Sistemi Complessi, CNR, Via dei Taurini 19, 00185 Rome, Italy
dDipartimento di Chimica, Sapienza Università di Roma, P.le Aldo Moro 5, 00185 Rome, Italy
eIstituto di Scienze e Tecnologie Chimiche, SCITEC-CNR, c/o Università Cattolica del Sacro Cuore, Largo Francesco Vito 1, 00168 Roma, Italy
First published on 6th March 2026
Abstract
The reduction of graphene oxide (GO) to reduced GO (rGO) is pivotal for producing graphene-based materials with numerous applications in the fields of electronics, energy storage, sensing, and biomedicine. Traditional reduction methods often involve toxic reagents and high temperatures, which pose significant environmental and safety concerns. Therefore, developing green reduction techniques has become essential. In this work, we demonstrate that the non-pathogenic bacterium Sporosarcina pasteurii is capable of reducing GO under nutrient-deprived aqueous conditions at both low (4 °C) and moderate (30 °C) temperatures. Spectroscopic analyses (UV-vis, Raman, FTIR, and XPS) confirmed the depletion of oxygen-containing functional groups and the partial restoration of the π-conjugated carbon network, indicating the successful reduction of GO. In particular, XPS investigations revealed that epoxide groups are preferentially removed during the bacterial reduction process. Morphological characterization revealed direct association between GO sheets and bacterial cells without compromising cell integrity or long-term viability. Fractionation experiments showed that reduction is mediated by both soluble extracellular components and membrane-associated factors, suggesting a mechanism involving extracellular electron transfer rather than active metabolism. NADH consumption in the presence of GO supports its role as an extracellular electron acceptor, contributing to the cellular redox balance under nutrient-limited conditions. These findings introduce S. pasteurii as a robust biocatalyst for the green, low-cost, and scalable production of rGO under mild conditions.
Green foundation1. Our work advances green chemistry by introducing a sustainable, nutrient-free and non-toxic method to reduce graphene oxide using Sporosarcina pasteurii. Unlike conventional chemical approaches that require hazardous reagents or high temperatures, our method operates in water at mild or even low temperatures (4–30 °C), showing efficiency without compromising bacterial viability. This represents a qualitative leap towards the safer and scalable production of reduced graphene oxide.2. The key green chemistry achievement is the replacement of toxic reductants with a non-pathogenic bacterium, demonstrating comparable reduction efficiency, as confirmed by spectroscopy, while avoiding the generation of chemical wastes and enabling low-energy processes. 3. Future research could make the process greener by optimizing large-scale bioreactors, recycling bacterial by-products as additional reducing agents, and integrating renewable energy sources for cultivation and processing, further lowering the environmental footprint. |
Introduction
Graphene oxide (GO) and its derivative, reduced graphene oxide (rGO), have gained enormous importance for their applications in many fields of science such as energy storage, environmental remediation, biosensing, drug delivery, tissue engineering and 3D printing.1–4 GO, characterized by a high density of oxygen-containing functional groups, exhibits excellent dispersibility in aqueous media and enables easy chemical modification. This makes GO particularly advantageous in biomedical applications such as targeted drug delivery systems,5 antibacterial coatings,6,7 and as a scaffold material in tissue engineering.8,9 Moreover, the ability of GO to adsorb heavy metals, dyes and organic pollutants has been widely explored for water purification and environmental clean-up.10,11 On the other hand, rGO, which results from the partial restoration of the π-conjugated network of GO via chemical, thermal, or green reduction, exhibits significantly improved electrical conductivity and mechanical integrity. These properties make rGO more suitable for applications such as supercapacitors, lithium-ion batteries, flexible electronics, and sensors.12 The electrical conductivity of rGO is beneficial for interfacing with electrically active cells, such as neurons and cardiomyocytes, promoting cellular responses through electrical stimulations.13,14 Moreover, the high surface area and residual functional groups of rGO allow it to adsorb and interact with a wide range of ions and biomolecules. This property has been shown to enhance specific cell differentiation pathways, such as osteogenesis by mesenchymal stem cells.8 Among the various strategies for obtaining rGO, chemical reduction remains the most commonly used approach. Typical chemical methods involve the use of potent reducing agents such as hydrazine hydrate, sodium borohydride, or hydroquinone, which are capable of removing oxygen-containing functional groups and partially restoring the sp2-hybridized carbon lattice.15 However, these chemical processes raise significant environmental and health concerns due to the generation of hazardous wastes. In addition to toxicity concerns, currently employed GO reduction strategies differ significantly in terms of the process complexity, cost, and scalability. Chemical reduction methods using hydrazine hydrate or sodium borohydride are effective but rely on highly reactive and potentially hazardous reagents, raising safety and environmental concerns that limit their applicability in biomedical or environmentally sensitive applications.16,17 Alternatively, thermal annealing represents a chemically cleaner approach; however, it typically requires high-temperature furnaces (>1000 °C), resulting in substantial energy consumption and associated greenhouse gas emissions, which negatively impact process sustainability at scale.18 Green chemical reductants such as ascorbic acid are less-toxic alternatives.19 Nevertheless, the reduction of compounds using such reductants during industrial production generally involves microbial fermentation, followed by chemical purification steps, which may increase process complexity and labor requirements when implemented at a larger scale.16,17In response to these limitations, there has been growing interest in the development of “green” reduction methods that utilize environmentally benign and biocompatible alternatives.20–22 These include the use of biological agents such as plant extracts, bacterial cultures, fungi, and amino acids. Green reduction methods not only mitigate environmental and health risks but also improve the biocompatibility of the resulting rGO, which is particularly advantageous in biomedical contexts. Moreover, in the case of non-pathogenic microbial-mediated GO reduction, these approaches may offer a scalable and safe platform for large-scale production.23,24 A series of microbial species have indeed been used to obtain a low cost, green rGO: Shewanella spp.24–29Geobacter spp.30–32 which use mainly c-type cytochromes as well as soluble electron shuttles in the process,31E. coli and P. aeruginosa,32–34 halofiles,35Glucorobacter spp.36Desulfuromonas spp.31 are the species for which the effect was reported. Since most microbial reductions of GO take place at room temperature, this method is considerably more environmentally friendly than other reduction techniques. In addition, it offers a more cost-effective option for large-scale graphene production compared to hydrothermal methods. However, microbial reduction typically requires a longer reaction time—usually between 24 and 72 hours—than chemical or thermal approaches, and the presence of GO itself can negatively affect bacterial viability.23 Furthermore, microbial reduction processes are dependent on bacterial growth conditions, such as precise anaerobic atmosphere, controlled temperature and specific nutrients in the medium, such as glucose.
In this study, we explore the use of Sporosarcina pasteurii to obtain rGO in a green and low-cost way. S. pasteurii is a bacterium widely recognized for its role in microbially induced calcium carbonate precipitation (MICP), a process with significant applications in biocementation, self-healing concrete, and wastewater treatment.37,38 Its effectiveness in these areas is largely attributed to its resilience and ability to function under extreme environmental conditions. A key factor in this resilience is its capacity to form endospores—dormant, nonreproductive structures that allow the bacterium to survive in the absence of nutrients and under harsh stressors such as ultraviolet radiation, desiccation, extreme temperatures, and exposure to chemical agents, including antibiotics. When environmental conditions become unfavorable, S. pasteurii initiates sporulation, forming intracellular spores that are eventually released upon the lysis of the mother cell. These metabolically inactive spores can remain viable for extended periods, reviving when conditions improve. This exceptional survival capability, combined with its urease activity, underpins the bacterium's robustness and versatility in biotechnological and environmental engineering applications.39–41 In this work, we test whether S. pasteurii in an environment lacking nutrients can still interact with GO and mediate its reduction. Remarkably, even under these nutrient-deprived and controlled-temperature conditions, the bacterium remains viable and exhibits a measurable ability to reduce GO, suggesting that extracellular electron transfer and the action of secreted mediators can occur independently of active growth. This observation provides new perspectives for employing S. pasteurii as a biocatalyst in green nanomaterial production, leveraging its intrinsic resilience and metabolic versatility to overcome the constraints that typically limit microbial reduction strategies.
Materials and methods
Reagents were purchased from Sigma-Aldrich, Fluka, Alfa Aesar, VWR, TCI, Cell Signaling, and Enzo Life Sciences and used as received without further purification, unless otherwise stated. Milli-Q water with a resistivity of 18.2 MΩ cm at 25 °C was employed in the experiments. Commercial aqueous dispersions of GO (4 mg mL−1) were provided by Graphenea.Bacterial growth, viability and ROS measurements
Sporosarcina pasteurii (ATCC 11859) was initially cultured overnight in nutrient urea-YE broth under standard growth conditions.42 Following incubation, cells were harvested by centrifugation at 6000 rpm and resuspended in Milli-Q water. The cell number was evaluated by plating samples overnight at 30 °C and counting colony-forming units (CFUs). For growth and viability assessments, aliquots of bacterial suspensions previously exposed to GO were reinoculated in fresh nutrient broth, and the optical density at 600 nm (OD600) was recorded at regular intervals to generate growth curves. In parallel, metabolic activity under nutrient-deprived conditions was evaluated using a resazurin assay according to the manufacturer's instructions (Cell Signaling) using a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 dilution and a 590/640 nm ratio, as reported previously by our group.11 Fluorescence readings were collected using a Cytation 3 (Biotek) plate reader, with untreated bacterial suspensions used as controls. All experiments were performed in triplicate to ensure reproducibility. To obtain several fractions of bacterial byproducts, solutions were filtered using sterile Millipore filters (0.2 μm) combined with centrifugation. NADH quantification was performed by measuring its intrinsic autofluorescence by exciting at 340 nm and recording the fluorescence emission at 460 nm. The nonfluorescent, cell-permeable oxidative stress detection reagent (green, Ex/Em = 490/525 nm) from Enzo Life Sciences was used to quantify reactive species (hydrogen peroxide, peroxynitrite and hydroxyl radicals). Fluorescence signals were quantified employing a UV-transparent 96-well plate using a Cytation 3 (Biotek) plate reader.
10 dilution and a 590/640 nm ratio, as reported previously by our group.11 Fluorescence readings were collected using a Cytation 3 (Biotek) plate reader, with untreated bacterial suspensions used as controls. All experiments were performed in triplicate to ensure reproducibility. To obtain several fractions of bacterial byproducts, solutions were filtered using sterile Millipore filters (0.2 μm) combined with centrifugation. NADH quantification was performed by measuring its intrinsic autofluorescence by exciting at 340 nm and recording the fluorescence emission at 460 nm. The nonfluorescent, cell-permeable oxidative stress detection reagent (green, Ex/Em = 490/525 nm) from Enzo Life Sciences was used to quantify reactive species (hydrogen peroxide, peroxynitrite and hydroxyl radicals). Fluorescence signals were quantified employing a UV-transparent 96-well plate using a Cytation 3 (Biotek) plate reader.
GO reduction protocol, sample characterization by spectroscopy and electrical resistivity measurements
GO suspensions were prepared with a final concentration between 10 and 100 µg mL−1 in sterile deionized water. Bacterial suspensions obtained after overnight inoculation in broth were centrifuged to remove nutrients and resuspended in water to obtain an optical density (OD at 600 nm) of 0.25. Then, bacteria were mixed with GO (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) and incubated under two different temperature conditions: 4 °C for long-term reduction (from 1 to 30 days of incubation) and 30 °C for short-term reduction.
1) and incubated under two different temperature conditions: 4 °C for long-term reduction (from 1 to 30 days of incubation) and 30 °C for short-term reduction.
After the designated incubation times, samples were washed with 1% sodium dodecyl sulfate (SDS) and 70% ethanol and dialyzed (1 kDa membranes) to obtain purified GO or rGO.43 UV-Vis spectra were recorded with a Shimadzu UV-2600i Plus spectrophotometer at room temperature, employing quartz cuvettes with a 10 mm path length. Fourier transform infrared (FTIR) spectroscopy was performed using an ALPHA II compact FTIR spectrometer (Bruker) to assess the surface chemical composition of scaffolds. Samples were directly placed onto the crystal and dried, and spectra were recorded within the wavenumber range of 4000–550 cm−1.
Raman spectra were acquired at room temperature in the back-scattering geometry with an inVia Renishaw micro-Raman spectrometer equipped with an air-cooled CCD detector and super-Notch filters. An Ar+ ion laser (λlaser = 514 nm) was used, coupled to a Leica DLML microscope with a 20× objective. The resolution was set to 2 cm−1, and the spectra were calibrated using the 520.5 cm−1 line of a silicon wafer. Raman spectra were acquired from several6–10 different spots on the surface of the samples. Each spectrum was acquired with 10% power, 10 seconds of spectral acquisition, and 10 scans. All spectra were compared to pristine GO controls to quantify the extent of reduction.
For X-ray photoelectron spectroscopy (XPS), a freshly prepared H-terminated Si(100) surface was used as a support for the drop-casting of aqueous dispersions of pristine GO and bacteria-treated samples. XPS measurements were carried out using an Omicron NanoTechnology (Uppsala, Sweden) Multiprobe MXPS system equipped with a monochromatic Al Kα (hν = 1486.7 eV) X-ray source (Scienta Omicron XM-1000), with a 14 kV accelerating voltage and a 13 mA emission current. The experimental spectra were theoretically reconstructed by fitting the secondary electron background to a Shirley function and the elastic peaks to pseudo-Voigt functions with a 70![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 30 Gaussian–Lorentzian ratio.
30 Gaussian–Lorentzian ratio.
The relative percentage amount of the different oxygenated functional groups (OFGs) of GO (A%OFG) was determined from the area of the OFG peaks (AOFG) within the curve-fitting envelope of the C 1s region (the epoxy area must be divided by 2), according to the following equation:44
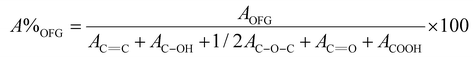 | (1) |
The oxygen content was determined from the RO/C ratio obtained after the curve-fitting of the C 1s region using the following equation:45,46
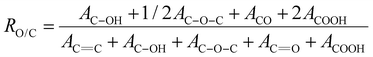 | (2) |
This approach rules out any possible contributions to the oxygen content, either deriving from contaminants or from the support material (Si).
The hydrodynamic size distribution and surface charge of GO nanosheets were characterized using a Zetasizer Nano ZS (Malvern, Herrenberg, Germany). Measurements were performed at a fixed detection angle of 173° with automatic attenuator settings. For each sample, three consecutive readings were taken, and the Z-average size was calculated via the Stokes–Einstein equation using cumulant analysis. Electrophoretic light scattering was employed for ζ-potential determination, with values calculated from electrophoretic mobility by applying the Henry correction within the Smoluchowski model. Data acquisition and processing were conducted using the Malvern Zetasizer software.
Electrical resistivity was measured using a four-terminal configuration in order to minimize contact resistance effects. Samples were mechanically compacted into cylindrical pellets of a known thickness and cross-sectional area before measurement. A current was applied through the outer electrodes, while the voltage drop was measured across the inner electrodes.
The volumetric resistivity (ρ, expressed in Ω cm) was calculated according to the following equation:
| ρ = R·(A/L), |
Scanning electron microscopy (SEM) and field-emission scanning electron microscopy (FE-SEM)
For morphological analysis, bacterial samples exposed to GO were fixed in 2.5% glutaraldehyde in phosphate-buffered saline (PBS) for 2 hours at 4 °C. The samples were then washed with PBS, dehydrated through a graded ethanol series (30%–100%), and air-dried on silicon wafers. Prior to imaging, specimens were sputter-coated with a thin gold layer to ensure conductivity. SEM imaging was performed using a field-emission scanning electron microscope operated at an accelerating voltage of 5–10 kV. Images were acquired at various magnifications to evaluate the bacterial morphology, integrity, and potential interactions with GO sheets. For the field-emission scanning electron microscopy (FE-SEM) investigations, the graphene-based samples were dispersed in Milli-Q water, and 50–100 μL of these aqueous dispersions was dropped on H-terminated n-type Si(100) 1.5 cm × 1.5 cm wafers and allowed to dry at room temperature. A Zeiss Auriga 405 field-emission scanning electron microscope (Zeiss Microscopy GmbH, Jena, Germany), equipped with a Bruker Quantax detector (Bruker, Berlin, Germany), was used to obtain the SEM images, maintaining a pressure from 10−5 to 10−6 mbar. The electron beam operated at a working distance of 5.4 mm, while the acceleration voltage was set at 1.50 keV for imaging. The collected images were processed using ImageJ software.Results
GO reduction mediated by S. pasteurii occurs in ultrapure water without nutrients at different temperatures
In this study, we evaluated the capability of Sporosarcina pasteurii to reduce GO surfaces. Generally, the reduction of GO is known to occur via bacterial metabolism or direct contact with bacteria, but this mechanism has not been confirmed for all bacterial species.24–34 In our experimental setup, S. pasteurii cells were inoculated overnight in nutrient broth to allow cell division and expansion. After overnight inoculation, cells were washed and suspended in Milli-Q water to remove nutrients, left in Milli-Q water for 1 hour and then incubated with GO at 30 °C (24 hours incubation or short-term reduction) or 4 °C (7 day incubation or long-term reduction), as depicted in Fig. 1A. The final GO concentration was set to 100 μg mL−1, and the optical density (OD) of the bacterial suspension was 0.25, corresponding to 2.5 × 107 cells per mL by CFU counting. The exposure of GO to bacteria for varying times and temperatures does not alter the morphology of GO, as evidenced by FE-SEM images (Fig. 1B). After incubation, the nanomaterial was recovered from the medium and washed, and different spectroscopic techniques were used to quantify the mechanism. The UV-Vis spectra after GO incubation with S. pasteurii are shown in Fig. 1C. In particular, the spectrum of GO (Fig. 1C, black line) displays a strong absorption peak at ∼230 nm, attributed to the π–π* electron transitions of the sp2-hybridized carbon domains. Additionally, a shoulder appears around 300 nm, corresponding to the n–π* electron transitions of the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O functional groups. Upon reduction mediated by S. pasteurii, the main absorption peak is red-shifted from ∼230 nm to ∼285 nm (Fig. 1C, orange and dotted lines), indicating the partial restoration of the π-conjugated network as a result of the extensive removal of oxygen-containing functional groups. Simultaneously, the shoulder at 300 nm, associated with C
O functional groups. Upon reduction mediated by S. pasteurii, the main absorption peak is red-shifted from ∼230 nm to ∼285 nm (Fig. 1C, orange and dotted lines), indicating the partial restoration of the π-conjugated network as a result of the extensive removal of oxygen-containing functional groups. Simultaneously, the shoulder at 300 nm, associated with C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O functionalities, disappears, confirming the effective depletion of these functionalities.22 Moreover, the successful reduction of GO is clearly evidenced by a distinct colour change of the medium from brown to black (Fig. 1D), which becomes clearly observable after 24 hours at 30 °C or after 7 days at 4 °C. Besides, rGO samples display high dispersibility without evident aggregation.
O functionalities, disappears, confirming the effective depletion of these functionalities.22 Moreover, the successful reduction of GO is clearly evidenced by a distinct colour change of the medium from brown to black (Fig. 1D), which becomes clearly observable after 24 hours at 30 °C or after 7 days at 4 °C. Besides, rGO samples display high dispersibility without evident aggregation.
To gain deeper insights into the reducing effect of S. pasteurii on GO, Raman analysis was performed to investigate the characteristic spectral features of the graphene-based samples. In particular, the pristine GO employed in the experiments showed its D band localized at ∼1355 cm−1, ascribed to the ring breathing mode of the sp2 carbon rings adjacent to an edge or a defect, and the G band centered at ∼1603 cm−1, attributed to the carbon–carbon stretching mode of the sp2 domains (Fig. 2A and B, black line).47–49 Notably, the ID/IG increases upon the reduction of GO, indicating the activation of pre-existing defects and/or the introduction of additional ones—particularly within the basal plane—resulting from the removal of oxygen-based functionalities, such as epoxide groups.8,22 In particular, the layers of GO treated with S. pasteurii are easily reduced. Specifically, the ID/IG ratio increases from 0.81 ± 0.02 in GO to 1.03 ± 0.02 (Fig. 2A, orange line) and 0.97 ± 0.02 (Fig. 2B, red line) in rGO-based samples obtained after short and long reduction times, respectively. Long-term reduction experiments were conducted for up to 30 days, and the ID/IG ratio does not show any substantial variation over time, indicating that the value of 0.97 reached at day 7 can be considered a plateau within 1 month. According to Lehner et al., who compared microbial reduction by Shewanella oneidensis and chemical reduction using hydrazine,29 the ID/IG ratio is 1 for both methods, which is comparable to the value obtained using our methods. Also, ascorbic acid or N-acetyl cysteine green reduction methods used by our group afforded ID/IG ratios of 0.85 and 0.97, respectively.46 Therefore, here we demonstrate that without a complicated process, rGO can be obtained using bacteria that can replicate easily in water. This method can be scaled up at low temperature, resulting in time and cost savings.
The FT-IR spectra of short-term and long-term reduced samples were acquired and compared to that of pristine GO (Fig. 2C). Specifically, the spectrum of GO (Fig. 2C, black line) exhibits a broad band in the 3600–2400 cm−1 region, mainly attributed to the O–H stretching modes of water molecules adsorbed on the GO surface. In addition, the spectrum displays the diagnostic peak at ∼1723 cm−1 (highlighted by the dashed black line), corresponding to the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O stretching vibrations of ketones, aldehydes, and carboxyl groups. The band located at ∼1619 cm−1 is assigned to the bending vibrations of adsorbed water molecules. The peaks in the fingerprint region (1500–500 cm−1) are difficult to unambiguously assign, in agreement with the literature.47 The FT-IR spectrum of the bacteria (Fig. 2C, magenta line) contains numerous bands arising from different classes of biomolecules, and many of these peaks persist across all spectra. The C–H stretching vibrations of the bacterial membrane lipids appear in the 3000–2800 cm−1 region. Protein- and peptide-related signals dominate the spectral region below 1700 cm−1, with a pronounced amide I band centered at approximately 1650 cm−1. Additional features are observed in the mixed protein-fatty acid region between 1500 and 1200 cm−1, along with bands attributable to nucleic acids and polysaccharides in the 1200–900 cm−1 spectral interval.50 The formation of rGO induced by bacterial activity can be tentatively confirmed by the dampening of the ∼1723 cm−1 band (Fig. 2C, orange and red lines), which indicates the depletion of carbonyl-based functional groups as a result of the reduction process. Regarding the involvement of other functionalities, such as epoxide groups, their potential removal during bioreduction cannot be reliably assessed by FT-IR spectroscopy. This limitation arises from the difficulty in assigning signals in the fingerprint region in the spectrum of GO and the overlapping absorption from bacterial components in that range, which clearly remain absorbed on the GO surface even after extensive washing.
O stretching vibrations of ketones, aldehydes, and carboxyl groups. The band located at ∼1619 cm−1 is assigned to the bending vibrations of adsorbed water molecules. The peaks in the fingerprint region (1500–500 cm−1) are difficult to unambiguously assign, in agreement with the literature.47 The FT-IR spectrum of the bacteria (Fig. 2C, magenta line) contains numerous bands arising from different classes of biomolecules, and many of these peaks persist across all spectra. The C–H stretching vibrations of the bacterial membrane lipids appear in the 3000–2800 cm−1 region. Protein- and peptide-related signals dominate the spectral region below 1700 cm−1, with a pronounced amide I band centered at approximately 1650 cm−1. Additional features are observed in the mixed protein-fatty acid region between 1500 and 1200 cm−1, along with bands attributable to nucleic acids and polysaccharides in the 1200–900 cm−1 spectral interval.50 The formation of rGO induced by bacterial activity can be tentatively confirmed by the dampening of the ∼1723 cm−1 band (Fig. 2C, orange and red lines), which indicates the depletion of carbonyl-based functional groups as a result of the reduction process. Regarding the involvement of other functionalities, such as epoxide groups, their potential removal during bioreduction cannot be reliably assessed by FT-IR spectroscopy. This limitation arises from the difficulty in assigning signals in the fingerprint region in the spectrum of GO and the overlapping absorption from bacterial components in that range, which clearly remain absorbed on the GO surface even after extensive washing.
After purification via dialysis with 1 kDa membranes, the GO samples were characterized with XPS in order to assess the possible chemical composition changes upon exposure to S. pasteurii. In Fig. 2D, the C 1s photoionization spectra of the short- and long-term samples are compared to that of pristine GO. The C 1s XPS spectrum of the reference GO (Fig. 2D, upper panel) can be fit to five distinct components located at binding energies (BEs) of 284.8 eV, 286.5 eV, 287.0 eV, 288.0 eV, and 289.0 eV. These peaks are, respectively, assigned to C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C and C
C and C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C–H sp2-hybridized carbon (red) and hydroxyl (blue), epoxide (shaded green), carbonyl (magenta), and carboxyl (orange) functional groups,22,51,52 and the resulting RO/C atomic ratio calculated using eqn (2) is 0.42 (Table 1). Upon exposure to S. pasteurii, significant changes occur in the chemical composition of GO in both the short- and long-term samples (Fig. 2D, middle and lower panels), implying a decrease in the RO/C values to 0.34 and 0.36, respectively. Additionally, in both these rGO spectra, a drastic decrease in the epoxide component (green shaded curve, 287.0 eV) can be seen, indicating a major increase in the C
C–H sp2-hybridized carbon (red) and hydroxyl (blue), epoxide (shaded green), carbonyl (magenta), and carboxyl (orange) functional groups,22,51,52 and the resulting RO/C atomic ratio calculated using eqn (2) is 0.42 (Table 1). Upon exposure to S. pasteurii, significant changes occur in the chemical composition of GO in both the short- and long-term samples (Fig. 2D, middle and lower panels), implying a decrease in the RO/C values to 0.34 and 0.36, respectively. Additionally, in both these rGO spectra, a drastic decrease in the epoxide component (green shaded curve, 287.0 eV) can be seen, indicating a major increase in the C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C component and a slight increase in the C–OH component (Table 1). It should be noted that in the case of the rGO long-term sample (Fig. 2D, lower panel), an additional asymmetric feature at a low BE (grey, 284.5 eV) was needed to accomplish a satisfactory curve-fitting of experimental data. This component is usually assigned to graphitic C atoms with an extended π-electron conjugation22,45,53,54 and has been found necessary for GO-based materials, aside from the localized C
C component and a slight increase in the C–OH component (Table 1). It should be noted that in the case of the rGO long-term sample (Fig. 2D, lower panel), an additional asymmetric feature at a low BE (grey, 284.5 eV) was needed to accomplish a satisfactory curve-fitting of experimental data. This component is usually assigned to graphitic C atoms with an extended π-electron conjugation22,45,53,54 and has been found necessary for GO-based materials, aside from the localized C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C component, whenever a reduction mechanism, implying an extension of the π-electron conjugation, is at work.55 Therefore, XPS shows that upon the exposure of GO to S. pasteurii, a decrease in the O/C ratio, mostly due to the abatement of epoxide OFGs, occurs under both short- and long-term conditions, with the latter case involving a significant extension of the π-electron conjugation.
C component, whenever a reduction mechanism, implying an extension of the π-electron conjugation, is at work.55 Therefore, XPS shows that upon the exposure of GO to S. pasteurii, a decrease in the O/C ratio, mostly due to the abatement of epoxide OFGs, occurs under both short- and long-term conditions, with the latter case involving a significant extension of the π-electron conjugation.
Bulk electrical measurements reveal a pronounced decrease in the resistivity following bacterial reduction. Pristine GO exhibits a volumetric resistivity of 1.5 × 1013 Ω cm, while after reduction, rGO displays a resistivity of4 × 108 Ω cm. These values are consistent with literature reports for bulk or compacted graphene oxide and reduced graphene oxide materials.56 GO is widely reported to exhibit resistivity in the 1012–1014 Ω cm range due to disruption of the π-conjugated network by oxygen functional groups. Upon chemical or green reduction, rGO typically shows resistivity values in the 107–109 Ω cm range when measured in bulk/pellet configurations, reflecting partial restoration of sp2 domains and improved interflake charge transport.24,57
Therefore, the magnitude of resistivity decrease observed here is fully in agreement with expectations for mild reduction strategies and is consistent with the spectroscopic evidence reported above, including the increase in the Raman ID/IG ratio from 0.81 to 1.03 and the decrease in the oxygen content detected by XPS.
It is important to note that bulk resistivity values are generally higher than those reported for thin percolated films due to the porosity and interflake contact resistance inherent to compacted samples. Nonetheless, the observed multi-order-of-magnitude improvement clearly demonstrates the functional restoration of electrical transport pathways in bacterially reduced graphene oxide.
GO interferes with the S. pasteurii metabolism and allows the immediate reactivation of cells in broth, but GO is not used as a carbon source
Since it is known that interactions between GO and microorganisms can exert either antibacterial or probiotic effects, we quantified cell growth during the process.58 Indeed, an antibacterial effect would stop the reduction process.Fig. 3A illustrates the growth of S. pasteurii cells after incubation with GO at a final concentration of 100 μg mL−1 at a low temperature for 12 hours, 7 days or 30 days. Incubation was performed at 4 °C in Milli-Q water to minimize cell metabolism. Growth was measured by reinoculating GO-exposed samples in nutrient broth and monitoring the OD at 600 nm. Cell replication appears to initiate earlier in the presence of GO. The final OD600 plateau indicates that the total biomass yield is not significantly altered by GO exposure compared to control samples after 12 hours of treatment, as also confirmed by CFU counting (data not shown). Interestingly, while a prolonged incubation in water for 7 or 30 days decreases the ability of S. pasteurii cells to grow immediately after broth reinoculation, this phenomenon does not occur in the presence of GO, demonstrating a positive effect on cells for long-term incubation (Fig. 3A).
Representative images of bacterial suspensions in water, with and without GO, are presented in Fig. 3B. These SEM images confirm the preservation of cellular integrity after GO exposure for 12 hours or 7 days. Control samples display few intact cells after 7 days in ultrapure water compared to samples incubated with GO, which contain a high number of intact cells and a visible GO blanket. These morphological differences are confirmed after 30 days of incubation (images not shown).
The interaction between bacterial cells and GO was further characterized using dynamic light scattering (DLS) and zeta potential measurements, and the results are shown in Fig. 3C. In the number-based distribution shown in Fig. 3C, the GO sample (gray) exhibits a primary mode centred around 200 nm, while the S. pasteurii sample (blue) shows a sharp, narrow peak near 2 µm, reflecting the typical size of individual or small bacterial aggregates. When the two are combined (orange), the resulting profile preserves the graphene-associated peak but displays a noticeable shoulder or minor peak in the bacterial size range. The number distribution clarifies that while the GO nanosheets remain numerically dominant in the mixed sample, a fraction of larger aggregates—likely involving GO–bacteria interactions—also contributes to the size profile. The coexistence of both populations suggests partial association; furthermore, the mixed sample's secondary population with a hydrodynamic diameter smaller than that of the bacteria alone can be explained by the shape and structural features of the aggregates. Indeed, nonspherical or flexible aggregates can appear significantly smaller than their actual geometric dimensions, possibly with partially wrapping or coating, causing compact or flattened structures that diffuse more rapidly in a suspension. The zeta potential of the bacterial suspension is −34.4 mV ± 0.49 mV, whereas both the pristine GO and the GO–bacteria mixture exhibit values around −40 mV, namely, −40.7 mV ± 0.17 mV and −39.5 mV ± 0.63 mV, respectively. This supports the idea that GO dominates the surface characteristics of the mixed system, possibly through adsorption or surface wrapping. However, this wrapping is not detrimental to cell membranes due to the capability of the cells to grow again after broth reinoculation, as evidenced by their appearance in microscopy characterization (Fig. 3A).58 The cell integrity on the GO surface was also confirmed by FESEM imaging after reduction (Fig. 3D).
We investigated whether varying the GO concentration influenced the S. pasteurii cell viability and metabolic activity. As shown in Fig. 3E, the incubation of GO at different concentrations and then reinoculation in broth consistently results in accelerated growth onset under all tested conditions; however, the final cell population, indicated by the plateau in the OD, remains unchanged. This observation suggests that GO does not serve as a nutrient source itself for S. pasteurii, contrary to what has been hypothesized for other microbial species.59 This experiment also indicates that the mechanism is not related to the available GO surface being unchanged at different concentrations of GO down to 25 μg mL−1.
Nonetheless, the modulation of the metabolic routes of S. pasteurii cannot be excluded, particularly given the anticipated growth onset after sub-inoculation (Fig. 3A) and the distinct cell morphologies observed via SEM in Fig. 3B.
To measure metabolically active cells without nutrient-mediated spore reactivation, we evaluated metabolic activity using the resazurin assay on samples incubated for 12 hours in ultrapure water with or without GO at different concentrations. As shown in the representative images of the colorimetric assay in Fig. 3F, cell viability is not compromised after incubation with GO, both in samples with a high initial cell number (5 × 107 cells per mL) and with a low initial cell number (2.5 × 107 cells per mL), across a dilution series of GO concentrations ranging from 250 µg mL−1 to 0.07 µg mL−1 (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 dilutions). In other words, pictures show how the colour shift from blue to pink is comparable under all conditions, indicating similar metabolic activity.
2 dilutions). In other words, pictures show how the colour shift from blue to pink is comparable under all conditions, indicating similar metabolic activity.
The quantitative results reported in Fig. 3G (low cell number) and Fig. 3H (low cell number) confirm these observations. The resazurin reduction, expressed as the 590/640 nm absorbance ratio, shows no significant changes with varying concentrations of GO using both high and low cell numbers.
Mediators of reduction: bacterial membranes, GO – membrane contact, and soluble mediators effects
It has been hypothesized that GO reduction can be mediated by (i) direct contact involving membrane-bound cytochromes, (ii) oxidative stress and production of reactive oxygen species (ROS) and/or (iii) extracellular electron transfer via soluble redox mediators.60Under nutrient-deprived conditions, S. pasteurii is known to initiate sporulation as a survival strategy, forming endospores to withstand environmental stress. S. pasteurii spores can be stably preserved in sterile water for months, whereas vegetative cells lose viability in ultrapure water within days to weeks without nutrients. Upon reintroduction into a nutrient-rich environment, spores germinate and return to a metabolically active state, resuming growth and metabolic activity.61 We also hypothesize a possible spore-mediated mechanism (iv) for this bacterium.
To determine which cellular or extracellular component determines GO reduction, we incubated S. pasteurii cells in water for 12 hours and then separated the suspension by centrifugation and filtration into four fractions: cell pellets, extracellular components (supernatant after centrifugation), extracellular components smaller than 0.22 µm (filtered supernatant), and whole cells. These fractions were subsequently compared for their ability to reduce GO at 100 µg mL−1 (see the scheme in Fig. 4A).
Raman spectroscopy was used to investigate the reduction process of GO. A significant reduction was observed when GO was incubated with both whole cells and supernatants, as indicated by an increase in the ID/IG ratio from 0.86 ± 0.01 (Fig. 4B, red line) to 0.97 ± 0.01 (Fig. 4B, black line). A moderate reduction was also detected in the presence of the supernatant alone, with no noticeable difference between filtered and unfiltered samples, as reflected by an ID/IG ratio of 0.89 ± 0.01 (Fig. 4B, green and magenta lines). When using cell pellets, we observed aggregation along with a modest reduction of GO, as indicated by an ID/IG ratio of 0.89 ± 0.01 in the Raman spectrum (Fig. 4B, blue line). These results suggest that direct contact with the cell pellets alone is insufficient to induce a significant reduction of GO. Raman data are further supported by the UV-Vis spectra (Fig. 4C). The characteristic absorption peaks of GO at ∼230 nm and ∼300 nm (Fig. 4C, red line) underwent significant changes in the whole sample (Fig. 4C, black line). Specifically, the peak at ∼230 nm shifted to ∼265 nm, while the signal at ∼300 nm disappeared. These spectral changes indicate a partial restoration of the π-conjugated network due to the extensive removal of the oxygen-based functional groups, confirming the effectiveness of the S. pasteurii-mediated reduction.22 Concerning the UV-Vis spectra of the cell pellets, the pristine supernatant and its filtered form (<0.22 µm), as displayed in Fig. 5C, the reduction of GO was not cleanly observed, probably due to the absorption of the biological medium in the UV region.
For each fraction, we assessed viability directly via resazurin assays and by monitoring the OD600 after reinoculation in broth to reactivate eventual spores (Fig. 4D and E). In the untreated samples, viability was reduced but not completely lost in the cell pellets and in the unfiltered supernatant, whereas the filtered supernatant (0.22 µm) contained neither cells nor spores, as no growth was detected when nutrients were provided by reinoculation in broth. The dashed lines in Fig. 4D and E represent the blank value. Each of the four fractions was then incubated with GO (100 µg mL−1) in water at 30 °C for 24 hours, and the resazurin and final OD values after broth reinoculation are reported in grey in Fig. 4D and E, respectively. There was no difference in viability between treated and untreated samples, besides a slight reduction in pellets’ ability to metabolize resazurin due to the formation of visible aggregates in the presence of GO. When inoculated in broth (Fig. 4E), all the samples except for the filtered supernatants also grew again in the presence of GO.
Therefore, GO was lightly reduced in the presence of extracellular products, and the reduction was more evident when all mediators were present in the solution, indicating the possible effect of the nanomaterial in the propagation of the soluble signals produced by interacting with both bacteria and extracellular mediators, including spores.
When GO is introduced before mediators can interact with cell pellets, it aggregates, and reduces cell pellets metabolism, supporting a wrapping mechanism.
The viability experiments indicated that the unfiltered supernatants might contain spores that poorly metabolize resazurin (Fig. 4D) but can grow in the presence of nutrients (Fig. 4E).
Since Raman and UV spectroscopy evidenced a slight reduction after GO incubation with filtered supernatants, we quantified NADH and ROS levels in each sample to verify the contribution of soluble mediators. Indeed, these mediators are known to interact with graphene-based materials and are capable of reducing GO by transferring electrons, thereby contributing to its chemical reduction even in the absence of intact cells.62 In Fig. 5A, a schematic of the experimental setup is shown. The figure illustrates how the different fractions obtained from Sporosarcina pasteurii—namely, whole samples, cell pellets, unfiltered supernatants, and filtered supernatants—were incubated with GO under controlled conditions. After incubation, each fraction was processed, and either the whole samples (still containing GO) or the supernatants after centrifugation and removal of GO were used for measurements of NADH and ROS levels. In order to analyse whether the surface of GO had an effect on the probes used to measure ROS, the samples were incubated with the probe before and after the centrifugation of the samples to remove the GO flakes’ signal.
NADH fluorescence (Fig. 5B) revealed that whole-cell samples contained the highest NADH levels, while cell pellets exhibited the lowest. Upon incubation with GO, NADH concentrations decreased markedly in all GO-incubated fractions (in gray) compared to untreated controls. Notably, in whole cells and supernatants, the NADH signal was quenched in unprocessed samples but reappeared after GO removal by centrifugation, indicating that fluorescence suppression was due to GO interaction rather than NADH depletion itself. This effect was absent in untreated controls, which maintained stable fluorescence regardless of centrifugation.
ROS measurements (Fig. 5C) showed comparable ROS concentrations in supernatants and whole cells. In the presence of GO, ROS levels increased across all fractions except pellets; however, this signal diminished in centrifuged samples, suggesting that the elevated fluorescence was linked to direct probe interaction with rGO surfaces rather than to intrinsic ROS accumulation. Together, these data support the role of soluble mediators in GO reduction and point to distinct patterns of NADH consumed on the GO surface and ROS generated across the different fractions.
In bacterial cells, including Sporosarcina pasteurii, when minimal metabolism such as residual respiration continues but no external electron acceptors are available—whether oxygen, nitrate, iron or graphene oxide itself—reduced cofactors like NADH accumulate. The respiratory chain is blocked, energy production collapses, ion gradients can no longer be maintained, and the cell ultimately dies. In spore-forming bacteria such as S. pasteurii, sensor systems detect environmental stress, nutrient depletion, oxidative imbalance or redox imbalance and activate regulatory cascades through factors such as Spo0A.63 If electron acceptors are scarce but a minimal energy reserve is still present, the cell is driven toward sporulation as a protective strategy. If energy is completely depleted, NADH is excessively accumulated, membranes begin to collapse, and cell death becomes unavoidable.
Graphene oxide has been shown to act as an NADH electron shuttle.62 This electron transfer gradually depletes NADH, a phenomenon observed spectrophotometrically. In the presence of GO, S. pasteurii can offload electrons from NADH onto this material, maintaining an internal redox balance instead of allowing NADH to build up. In this way, redox stress and metabolic collapse are avoided. We, therefore, hypothesize that GO does not supply nutrients or induce the growth of S. pasteurii, but it stabilizes cells in the stationary phase by acting as a metabolic anchor.
These concepts are supported by findings on Bacillus subtilis in harsh environments, as described by Chen et al.,64 where extracellular electron transfer persists even at pH 1.5 for years or at 100 °C for hours. Under such conditions, NAD and NADH directly participate in electron transfer, and even under stress, the cells retain the ability to communicate electronically with their environment and link sporulation to redox activity. Taken together, this supports the idea that under challenging conditions, S. pasteurii may use NADH oxidation and extracellular electron transfer to GO to maintain redox balance. We also observed an increase in the production of ROS when the surface of samples containing GO was in contact with the cell-permeable fluorogenic green dye capable of measuring ROS levels (Fig. 5C).
As Zhao et al. (2017) demonstrated, NADH can donate electrons to graphene oxide, which then reduces molecular oxygen and generates ROS at or near its surface.62 Therefore, when the probe diffuses into the vicinity of GO particles where ROS are being formed, it can be oxidized on or near the graphene oxide surface, leading to increased fluorescence. The phenomenon occurs only when the probe is incubated with the whole sample, indicating a GO surface-dependent mechanism. Indeed, whilst the phenomenon occurs in cell-containing samples, we also observed it with the filtered supernatants in the presence of GO.
The reaction with the surface of rGO occurs because the functionalized carbon lattice can adsorb the probe through π–π interactions or electrostatic interactions, bringing the probe into close contact with sites where electron transfer and ROS generation are taking place. As soon as ROS are produced by the redox cycling between NADH and rGO in the presence of oxygen, the probe molecules in that microenvironment are oxidized. Therefore, the observed green fluorescence reflects both intracellular ROS produced by stressed cells and ROS generated on or near the rGO surface due to its catalytic electron shuttle activity.
Thus, the increase in ROS observed in the presence of GO can be explained as a secondary effect of NADH accumulation rather than as the primary driver of GO reduction. In this model, GO acts as an electron shuttle that accepts electrons from NADH and transfers them to molecular oxygen, thereby generating ROS at or near its surface. The resulting green fluorescence arises from both intracellular ROS produced by metabolically stressed cells and extracellular ROS generated through redox cycling between NADH and rGO in the presence of oxygen. Accordingly, ROS formation reflects the catalytic role of GO in maintaining redox balance, rather than serving as the main mechanism responsible for its reduction.
This influence on redox balance reflects on S. pasteurii viability and cell integrity during incubation and consequently might serve as a cell-stabilization process in the absence of nutrients.
Conclusions
Developing sustainable and biologically compatible methods for the production of rGO is of increasing importance, as conventional chemical or thermal reduction processes often require harsh conditions, toxic reagents, or high energy inputs. Several hypotheses have been proposed regarding the bacterium-mediated reduction of GO. One of the proposed mechanisms involves reduction driven by bacterial respiration and metabolism; however, in our experiments, metabolic activity was deliberately minimized, and we demonstrated that Sporosarcina pasteurii is capable of reducing GO under nutrient-depleted conditions, offering a potential route toward rGO production that is mild, scalable, and compatible with biological systems. The recent activity-based cost analysis of green rGO synthetic routes indicates that the total cost of ownership (TCO) of the process is primarily driven by material preparation, energy consumption and labor associated with multistep reduction protocols, rather than by the reducing agent itself.18 In particular, conventional hydrothermal reduction methods can reach costs up to 248.64 € per g, while simplified green routes reduce this value to approximately 19.48 € per g, mainly by decreasing the number of synthesis steps and energy demand. In contrast to chemical green reductants such as ascorbic acid, whose production relies on industrial fermentation, followed by chemical purification, our bacterial reduction protocol is performed as a one-pot biological process in ultrapure water without the addition of nutrients. Moreover, S. pasteurii is non-pathogenic and does not require strictly anaerobic conditions, eliminating additional infrastructure requirements typically associated with microbial reduction processes. As a result, the proposed method simultaneously reduces raw material costs, energy input and labor demand, which represent the dominant contributors to the synthesis cost, according to activity-based costing models. These characteristics suggest a favorable scalability profile and a potential economic advantage compared to existing green reduction strategies.Some microbial species cause oxygen loss from the surface of the material, i.e. the production of rGO, and simultaneously lose cell integrity due to the mechanical action of GO and/or cellular stress caused by the production of ROS.32–34 Indeed, previous observations have reported that the final product, rGO, impairs the viability of species such as E. coli and Pseudomonas spp.because the antibacterial effects of rGO are more pronounced than those of GO. We observed that S. pasteurii viability is not adversely affected during these processes. The absence of the antimicrobial effects of GO suggests that this microorganism can be sustainably employed in repeated or batch production processes without compromising its biological activity. Our data clearly show that GO reduction occurs in the presence of S. pasteurii fractions, with a marked contribution from soluble mediators released by the cells, i.e. NADH, together with direct contact with cell pellets. This is possibly due to signals activated in harsh environments rather than classical bacterial respiration. We propose that S. pasteurii, due to its intrinsic stability in nutrient-limited environments, can utilize GO as an extracellular electron acceptor. By transferring electrons to the GO surface, the cells maintain redox homeostasis and prolong spore dormancy without active nutrient consumption. This ability is consistent with previous evidence that microbial metabolism generates electrons that can be exported to external substrates and with reports that in harsh environments—such as in Bacillus subtilis at low pH—bacteria maintain extracellular electron transfer to stabilize their metabolism. Furthermore, NADH and NAD+ molecules crossing the plasma membrane may serve as intercellular signals, as described in both bacterial and eukaryotic systems64
Taken together, our findings not only reveal a new biological method for reducing GO but also suggest a novel strategy to improve the viability and long-term performance of S. pasteurii and possibly other similar bacilli.
Since Sporosarcina pasteurii is a non-pathogenic bacillus, the rGO produced through its biomediated reduction may represent a promising platform for advanced material applications. In the biomedical field, one possible application lies in drug-delivery systems, where the biocompatibility and low toxicity of the producing microorganism are advantageous during the fabrication stage. In this context, bacterially derived rGO could be synthesized under mild aqueous conditions and subsequently purified for use as a carrier platform, minimizing the need for toxic chemical reductants. Beyond biomedical applications, S. pasteurii is actively involved in microbially induced calcite precipitation (MICP), enabling the integration of rGO within calcium carbonate matrices. Such hybrid bio-mineral composites could reinforce biocement by providing conductive and mechanically resilient rGO domains within the mineralized structure. This approach may support the development of self-healing and potentially self-sensing construction materials for soil stabilization and concrete crack repair. Finally, the ability of S. pasteurii to mediate rGO formation at low temperatures opens opportunities in systems requiring cold-processed graphene derivatives. These include microbial fuel cells operating in harsh or low-temperature environments, remote bioelectrochemical sensors, and in situ energy-harvesting devices designed for extreme conditions. Together, these scenarios illustrate a versatile technological platform spanning biomedical material fabrication, sustainable construction, and bioelectrochemical energy systems.37
Conflicts of interest
There are no conflicts to declare.Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author upon reasonable request.Acknowledgements
This research was supported by EU funding within the NextGeneration EU-MUR PNRR Extended Partnership initiative on Emerging Infectious Diseases (Project no. PE00000007, INF-ACT) CUP B53C20040570005. We acknowledge the contribution of 3D Bioprinting Research Core Facility G-STeP of the Fondazione Policlinico Universitario “A. Gemelli” IRCCS for sample processing and the G-STeP Microscopy Facility of the Fondazione Policlinico Universitario “A. Gemelli” IRCCS for microscopy experiments.References
- V. Palmieri, W. Lattanzi, G. Perini, A. Augello, M. Papi and M. De Spirito, 3D-printed graphene for bone reconstruction, 2D Mater., 2020, 7, 022004 CrossRef.
- F. De Maio, E. Rosa, G. Perini, A. Augello, B. Niccolini and F. Ciaiola, et al., 3D-printed graphene polylactic acid devices resistant to SARS-CoV-2: Sunlight-mediated sterilization of additive manufactured objects, Carbon, 2022, 194, 34–41 CrossRef PubMed.
- R. Di Santo, E. Quagliarini, S. Palchetti, D. Pozzi, V. Palmieri and G. Perini, et al., Microfluidic-generated lipid-graphene oxide nanoparticles for gene delivery, Appl. Phys. Lett., 2019, 114, 233701 CrossRef.
- F. De Maio, V. Palmieri, G. Santarelli, G. Perini, A. Salustri and I. Palucci, et al., Graphene Oxide-Linezolid Combination as Potential New Anti-Tuberculosis Treatment, Nanomaterials, 2020, 10, 1431 CrossRef PubMed.
- J. Liu, L. Cui and D. Losic, Graphene and graphene oxide as new nanocarriers for drug delivery applications, Acta Biomater., 2013, 9, 9243–9257, DOI:10.1016/j.actbio.2013.08.016.
- F. Fauzi, M. M. Musawwa, H. Hidayat, A. Kusumaatmaja and W. S. B. Dwandaru, Nanocomposites based on biocompatible polymers and graphene oxide for antibacterial coatings, Polym. Polym. Compos., 2021, 29, S1609–S1620 Search PubMed.
- F. Bugli, M. Cacaci, V. Palmieri, R. Di Santo, R. Torelli and G. Ciasca, et al., Curcumin-loaded graphene oxide flakes as an effective antibacterial system against methicillin-resistant staphylococcus aureus, Interface Focus, 2018, 8, 20170059 CrossRef PubMed.
- F. Tiberio, F. Amato, C. Desiderio, F. Vincenzoni, G. Perini and I. Moretti, et al., The osteoconductive properties of graphene-based material surfaces are finely tuned by the conditioning layer and surface chemistry, Mater. Adv., 2024, 5, 4772–4785 RSC.
- V. Palmieri, M. Barba, L. Di Pietro, C. Conti, M. De Spirito and W. Lattanzi, et al., Graphene Oxide Induced Osteogenesis Quantification by In situ 2D-Fluorescence Spectroscopy, Int. J. Mol. Sci., 2018, 19, 3336 CrossRef.
- Z. Han, L. Huang, H. Qu, Y. Wang, Z. Zhang and Q. Rong, et al., A review of performance improvement strategies for graphene oxide-based and graphene-based membranes in water treatment, J. Mater. Sci., 2021, 56, 9545–9574 CrossRef.
- V. Palmieri, M. de Spirito and M. Papi, Enhanced Microbial Sensing via Resazurin Reduction Catalyzed by Graphene Oxide, A Versatile Approach for Diagnostics and Electrochemical Applications, Environ. Toxicol., 2025, 24572 Search PubMed.
- H. Kumar, R. Sharma, A. Yadav and R. Kumari, Recent advancement made in the field of reduced graphene oxide-based nanocomposites used in the energy storage devices: A review, J Energy Storage, 2021, 33, 102032 CrossRef.
- A. Magaz, X. Li, J. E. Gough and J. J. Blaker, Graphene oxide and electroactive reduced graphene oxide-based composite fibrous scaffolds for engineering excitable nerve tissue, Mater. Sci. Eng., C, 2021, 119, 111632 CrossRef.
- V. Palmieri, F. Sciandra, M. Bozzi, M. De Spirito and M. Papi, 3D graphene scaffolds for skeletal muscle regeneration: future perspectives, Front. Bioeng. Biotechnol., 2020, 8 DOI:10.3389/fbioe.2020.00383.
- A. Gutiérrez-Cruz, A. R. Ruiz-Hernández, J. F. Vega-Clemente, D. G. Luna-Gazcón and J. Campos-Delgado, A review of top-down and bottom-up synthesis methods for the production of graphene, graphene oxide and reduced graphene oxide, J. Mater. Sci., 2022, 57, 14543–14578 CrossRef.
- I. O. Faniyi, O. Fasakin, B. Olofinjana, A. S. Adekunle, T. V. Oluwasusi and M. A. Eleruja, et al., The comparative analyses of reduced graphene oxide (RGO) prepared via green, mild and chemical approaches, SN Appl. Sci., 2019, 1, 1181 CrossRef.
- A. C. Christudoss, K. K. Sah, R. Vikram, S. Giri, D. Viswanathan and C. O. Dimkpa, et al., Tailoring the Synthesis Route of Reduced Graphene Oxide and Its Toxicological Effects on Allium cepa L, ACS Omega, 2025, 10, 20771–20783 CrossRef.
- D. A. Gkika, K. N. Maroulas and G. Z. Kyzas, Various reduced graphene oxide green synthetic routes: comparing the cost procedures, ACS Omega, 2025, 10, 36221–36237 CrossRef PubMed.
- J. J. J. Zhang, H. Yang, G. Shen, P. Cheng, J. J. J. Zhang and S. Guo, Reduction of graphene oxide vial-ascorbic acid, Chem. Commun., 2010, 46, 1112–1114 RSC.
- M. Kurian, Recent progress in the chemical reduction of graphene oxide by green reductants–A Mini review, Carbon Trends, 2021, 5, 100120 CrossRef.
- V. Palmieri, E. A. Dalchiele, G. Perini, A. Motta, M. De Spirito and R. Zanoni, et al., Biocompatible: N -acetyl cysteine reduces graphene oxide and persists at the surface as a green radical scavenger, Chem. Commun., 2019, 55, 4186–4189 RSC.
- F. Amato, G. Perini, G. Friggeri, A. Augello, A. Motta and L. Giaccari, et al., Unlocking the Stability of Reduced Graphene Oxide Nanosheets in Biological Media via Use of Sodium Ascorbate, Adv. Mater. Interfaces, 2023, 10, 2300105 CrossRef.
- J.-Y. Tee, F.-L. Ng, F. S.-L. Keng, G. Gnana Kumar and S.-M. Phang, Microbial reduction of graphene oxide and its application in microbial fuel cells and biophotovoltaics, Front. Mater. Sci., 2023, 17, 230642 CrossRef.
- E. C. Salas, Z. Sun, A. Lüttge and J. M. Tour, Reduction of graphene oxide via bacterial respiration, ACS Nano, 2010, 4, 4852–4856 CrossRef PubMed.
- G. Wang, F. Qian, C. W. Saltikov, Y. Jiao and Y. Li, Microbial reduction of graphene oxide by Shewanella, Nano Res., 2011, 4, 563–570 CrossRef.
- T. Lin, W. Ding, L. Sun, L. Wang, C.-G. Liu and H. Song, Engineered Shewanella oneidensis-reduced graphene oxide biohybrid with enhanced biosynthesis and transport of flavins enabled a highest bioelectricity output in microbial fuel cells, Nano Energy, 2018, 50, 639–648 CrossRef.
- Y. Yong, Y. Yu, X. Zhang and H. Song, Highly active bidirectional electron transfer by a self–assembled electroactive reduced–graphene–oxide–hybridized biofilm, Angew. Chem., Int. Ed., 2014, 53, 4480–4483 CrossRef PubMed.
- Y. Jiao, F. Qian, Y. Li, G. Wang, C. W. Saltikov and J. A. Gralnick, Deciphering the electron transport pathway for graphene oxide reduction by Shewanella oneidensis MR-1, J. Bacteriol., 2011, 193, 3662–3665 CrossRef PubMed.
- B. A. E. Lehner, V. A. E. C. Janssen, E. M. Spiesz, D. Benz, S. J. J. Brouns and A. S. Meyer, et al., Creation of conductive graphene materials by bacterial reduction using Shewanella oneidensis, ChemistryOpen, 2019, 8, 888–895 CrossRef PubMed.
- Y. Lu, L. Zhong, L. Tang, H. Wang, Z. Yang and Q. Xie, et al., Extracellular electron transfer leading to the biological mediated production of reduced graphene oxide, Chemosphere, 2020, 256, 127141 Search PubMed.
- N. Yoshida, Y. Miyata, K. Doi, Y. Goto, Y. Nagao and R. Tero, et al., Graphene oxide-dependent growth and self-aggregation into a hydrogel complex of exoelectrogenic bacteria, Sci. Rep., 2016, 6, 21867 CrossRef.
- S. Kalathil, K. P. Katuri, A. S. Alazmi, S. Pedireddy, N. Kornienko and P. M. F. J. Costa, et al., Bioinspired synthesis of reduced graphene oxide-wrapped Geobacter sulfurreducens as a hybrid electrocatalyst for efficient oxygen evolution reaction, Chem. Mater., 2019, 31, 3686–3693 CrossRef.
- S. Gurunathan, J. W. Han, A. A. Dayem, V. Eppakayala and J.-H. Kim, Oxidative stress-mediated antibacterial activity of graphene oxide and reduced graphene oxide in Pseudomonas aeruginosa, Int. J. Nanomed., 2012, 7, 5901 CrossRef.
- O. Akhavan and E. Ghaderi, Escherichia coli bacteria reduce graphene oxide to bactericidal graphene in a self-limiting manner, Carbon, 2012, 50, 1853–1860 CrossRef.
- S. Raveendran, N. Chauhan, Y. Nakajima, H. Toshiaki, S. Kurosu and Y. Tanizawa, et al., Ecofriendly route for the synthesis of highly conductive graphene using extremophiles for green electronics and bioscience, Part. Part. Syst. Charact., 2013, 30, 573–578 CrossRef.
- N. K. Rathinam, S. Berchmans, R. K. Sani and D. R. Salem, Rewiring the microbe-electrode interfaces with biologically reduced graphene oxide for improved bioelectrocatalysis, Bioresour. Technol., 2018, 256, 195–200 CrossRef PubMed.
- M. J. Tuttle, B. M. Bradow, R. L. Martineau, M. S. Carter, J. A. Mancini and K. A. Holley, et al., Shelf-Stable Sporosarcina pasteurii Formulation for Scalable Laboratory and Field-Based Production of Biocement, ACS Appl. Mater. Interfaces, 2025, 17, 7251–7261 CrossRef PubMed.
- K. Zhang, C.-S. Tang, N.-J. Jiang, X.-H. Pan, B. Liu and Y.-J. Wang, et al., Microbial-induced carbonate precipitation (MICP) technology: a review on the fundamentals and engineering applications, Environ. Earth Sci., 2023, 82, 229 CrossRef PubMed.
- Q. Fu, Y. Wu, S. Liu, L. Lu and J. Wang, The adaptability of Sporosarcina pasteurii in marine environments and the feasibility of its application in mortar crack repair, Constr. Build. Mater., 2022, 332, 127371 CrossRef.
- F. M. Lapierre, J. Schmid, B. Ederer, N. Ihling, J. Büchs and R. Huber, Revealing nutritional requirements of MICP-relevant Sporosarcina pasteurii DSM33 for growth improvement in chemically defined and complex media, Sci. Rep., 2020, 10, 22448 CrossRef.
- S. Khoshtinat, Advancements in Exploiting Sporosarcina pasteurii as Sustainable Construction Material: A Review, Sustainability, 2023, 15, 13869 CrossRef.
- O. Šovljanski, L. Pezo, J. Grahovac, A. Tomić, A. Ranitović and D. Cvetković, et al., Best-performing Bacillus strains for microbiologically induced CaCO3 precipitation: Screening of relative influence of operational and environmental factors, J. Biotechnol., 2022, 350, 31–41 CrossRef.
- Y. Qu, J. Wang, Q. Ma, W. Shen, X. Pei and S. You, et al., A novel environmental fate of graphene oxide: Biodegradation by a bacterium Labrys sp. WJW to support growth, Water Res., 2018, 143, 260–269 CrossRef.
- F. Amato, M. Fazi, L. Giaccari, S. Colecchia, G. Perini and V. Palmieri, et al., Isolation by dialysis and characterization of luminescent oxidized carbon nanoparticles from graphene oxide dispersions: a facile novel route towards a more controlled and homogeneous substrate with a wider applicability, Nanotechnology, 2025, 36, 185602 CrossRef PubMed.
- A. Kovtun, D. Jones, S. Dell'Elce, E. Treossi, A. Liscio and V. Palermo, Accurate chemical analysis of oxygenated graphene-based materials using X-ray photoelectron spectroscopy, Carbon, 2019, 143, 268–275 CrossRef.
- A. G. Marrani, A. Motta, V. Palmieri, G. Perini, M. Papi and E. A. Dalchiele, et al., A comparative experimental and theoretical study of the mechanism of graphene oxide mild reduction by ascorbic acid and N-acetyl cysteine for biomedical applications, Mater. Adv., 2020, 1, 2745–2754 RSC.
- S. Guo, S. Garaj, A. Bianco and C. Ménard-Moyon, Controlling covalent chemistry on graphene oxide, Nat. Rev. Phys., 2022, 4, 247–262 CrossRef.
- M. S. Dresselhaus, A. Jorio, M. Hofmann, G. Dresselhaus and R. Saito, Perspectives on carbon nanotubes and graphene Raman spectroscopy, Nano Lett., 2010, 10, 751–758 CrossRef PubMed.
- J. Ribeiro-Soares, M. E. Oliveros, C. Garin, M. V. David, L. G. P. Martins and C. A. Almeida, et al., Structural analysis of polycrystalline graphene systems by Raman spectroscopy, Carbon, 2015, 95, 646–652 Search PubMed.
- H. Yang, H. Shi, B. Feng, L. Wang, L. Chen and A. Alvarez-Ordóñez, et al., Protocol for bacterial typing using Fourier transform infrared spectroscopy, STAR Protoc., 2023, 4, 102223 Search PubMed.
- C. Y. Lin, C. E. Cheng, S. Wang, H. W. Shiu, L. Y. Chang and C. H. Chen, et al., Synchrotron Radiation Soft X-ray Induced Reduction in Graphene Oxide Characterized by Time-Resolved Photoelectron Spectroscopy, J. Phys. Chem. C, 2015, 119, 12910–12915 Search PubMed.
- I. Ferrari, A. Motta, R. Zanoni, F. A. Scaramuzzo, F. Amato and E. A. Dalchiele, et al., Understanding the nature of graphene oxide functional groups by modulation of the electrochemical reduction: A combined experimental and theoretical approach, Carbon, 2023, 203, 29–38 CrossRef.
- F. Tiberio, F. Amato, C. Desiderio, F. Vincenzoni, G. Perini and I. Moretti, et al., The osteoconductive properties of graphene-based material surfaces are finely tuned by the conditioning layer and surface chemistry, Mater. Adv., 2024, 5, 4772–4785 RSC.
- R. Larciprete, P. Lacovig, S. Gardonio, A. Baraldi and S. Lizzit, Atomic Oxygen on Graphite: Chemical Characterization and Thermal Reduction, J. Phys. Chem. C, 2012, 116, 9900–9908 CrossRef.
- A. G. Marrani, A. C. Coico, D. Giacco, R. Zanoni, A. Motta and R. Schrebler, et al., Flexible Interfaces between Reduced Graphene Oxide and Indium Tin Oxide/Polyethylene Terephthalate for Advanced Optoelectronic Devices, ACS Appl. Nano Mater., 2019, 2, 5963–5972 CrossRef.
- D. R. Dreyer, A. D. Todd and C. W. Bielawski, Harnessing the chemistry of graphene oxide, Chem. Soc. Rev., 2014, 5288–5301 RSC.
- M. J. Fernández-Merino, L. Guardia, J. I. Paredes, S. Villar-Rodil, P. Solís-Fernández and A. Martínez-Alonso, et al., Vitamin C is an ideal substitute for hydrazine in the reduction of graphene oxide suspensions, J. Phys. Chem. C, 2010, 114, 6426–6432 CrossRef.
- V. Palmieri, F. Bugli, M. C. Lauriola, M. Cacaci, R. Torelli and G. Ciasca, et al., Bacteria Meet Graphene: Modulation of Graphene Oxide Nanosheet Interaction with Human Pathogens for Effective Antimicrobial Therapy, ACS Biomater. Sci. Eng., 2017, 3, 619–627 CrossRef CAS PubMed.
- D. D. Simeonova, K. Pollmann, A. Bianco and D. Lièvremont, Graphene oxide and bacteria interactions: what is known and what should we expect?, mSphere, 2024, 9, e00715–e00723 CrossRef.
- C. Vargas, R. Simarro, J. A. Reina, L. F. Bautista, M. C. Molina and N. González-Benítez, New approach for biological synthesis of reduced graphene oxide, Biochem. Eng. J., 2019, 151, 107331 CrossRef CAS.
- Y. Hu, W. Liu, Q. Zhang and X. Hu, Investigation on mineralization performance and spore germination conditions of calcium carbonate mineralizing bacteria, Mater. Res. Express, 2022, 9, 65403 CrossRef CAS.
- Y. Zhao, H.-S. Hsieh, M. Wang and C. T. Jafvert, Light-independent redox reactions of graphene oxide in water: Electron transfer from NADH through graphene oxide to molecular oxygen, producing reactive oxygen species, Carbon, 2017, 123, 216–222 CrossRef CAS.
- T. G. Villa and T. de Miguel Bouzas, Developmental biology in prokaryotes and lower eukaryotes, Springer, 2021 Search PubMed.
- L. Chen, C. Cao, S. Wang, J. R. Varcoe, R. C. T. Slade and C. Avignone-Rossa, et al., Electron communication of Bacillus subtilis in harsh environments, iScience, 2019, 12, 260–269 CrossRef CAS PubMed.
Footnote |
| † Co-first authors, these authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |





