Influence of gestational diabetes mellitus on the breast milk microbiota and oligosaccharides and their effects on the infant gut microbiota
Feng
Xu
ab,
Bo
Yang
 *abc,
Shumao
Cui
abc,
Zhenyu
Yang
de,
Nan
Dai
f,
Catherine
Stanton
cgh,
R. Paul
Ross
cg,
Jianxin
Zhao
*abc,
Shumao
Cui
abc,
Zhenyu
Yang
de,
Nan
Dai
f,
Catherine
Stanton
cgh,
R. Paul
Ross
cg,
Jianxin
Zhao
 abc,
Jianqiang
Lai
de,
Wei
Chen
abc,
Jianqiang
Lai
de,
Wei
Chen
 abc and
Ye
Wang
*de
abc and
Ye
Wang
*de
aState Key Laboratory of Food Science and Technology, Jiangnan University, Wuxi, China
bSchool of Food Science and Technology, Jiangnan University, 1800 Lihu Avenue, Wuxi, Jiangsu 214122, China. E-mail: bo.yang@jiangnan.edu.cn
cInternational Joint Research Laboratory for Maternal–Infant Microbiota and Health, Jiangnan University, Wuxi 214122, China
dNational Institute for Nutrition and Health, Chinese Center for Disease Control and Prevention, Beijing 100050, China. E-mail: wangye@ninh.chinacdc.cn
eKey Laboratory of Human Milk Science, Chinese Center for Disease Control and Prevention, Beijing 100050, China
fThe People's Hospital of Danyang, Affiliated Danyang Hospital of Nantong University, Danyang, China
gAPC Microbiome Ireland, University College Cork, Cork, Ireland
hTeagasc Food Research Centre, Moorepark, Co., Cork, Ireland
First published on 15th December 2025
Abstract
While the interplay between gestational diabetes mellitus (GDM) and the maternal–infant microbial axis is increasingly recognized, the specific pathways of influence remain unclear. This study comprehensively investigated the impact of GDM on the breast milk microbiota, human milk oligosaccharides (HMOs), and the subsequent development of the infant gut microbiota. We analyzed breast milk and paired infant fecal samples collected from healthy and GDM-affected mothers at two time points (0–7 and 42 days postpartum). The microbiota of both sample types was profiled by metagenomic sequencing, and HMOs in breast milk were quantified via liquid chromatography-tandem mass spectrometry (LC-MS/MS). Our findings revealed that GDM had a strong influence on the infant gut microbiota via reducing HMO concentrations than via direct alterations to the breast milk microbiota. These GDM-associated HMO alterations induced stage-specific shifts in the offspring's gut microbiota. Notably, the correlation between specific HMOs and gut bacteria reversed from the colostrum stage to the mature milk stage. This suggests that HMOs influence microbial colonization not only through direct utilization but also, and perhaps more importantly, via indirect ecological mechanisms such as cross-feeding. Collectively, our results identify maternal HMOs as a critical link between maternal metabolism and infant gut health, highlighting their potential as a promising nutritional target to improve long-term metabolic outcomes in GDM-exposed infants.
1. Introduction
Gestational diabetes mellitus (GDM) is defined as varying degrees of glucose intolerance that first emerge or are recognized during pregnancy. Its core mechanism involves a progressive decline in maternal insulin sensitivity with advancing gestational age, compounded by an inadequate compensatory increase in insulin secretion from pancreatic β-cells. Consequently, the body fails to maintain glucose homeostasis, leading to hyperglycemia.1 Beyond its short- and long-term risk to maternal health, GDM also elevates the offspring's risks of disorders such as obesity, metabolic syndrome, and type 2 diabetes.2,3 This is primarily mediated by increased placental glucose transfer resulting from the intrauterine hyperglycemic environment.4–6Emerging evidence suggests that GDM may influence the early establishment of the offspring's gut microbiota, though findings remain inconsistent and sometimes contradictory. For instance, while some studies report reduced alpha diversity compared to the controls,7–9 others show no significant difference or even increased microbial richness.10 Similarly, alterations in beneficial bacteria such as Bifidobacterium and Lactobacillus are disputed.11–13 These discrepancies may stem from variations in research methodologies, population heterogeneity, and sampling timing. The first 1000 days after birth represent a critical window for the establishment of a healthy gut microbiota, a period during which immune system maturation and gut microbial development proceed synchronously.14 Early-life gut dysbiosis has been linked to an increased risk of diseases later in childhood or adulthood,15 and cohort studies have shown that maternal GDM increases the offspring's susceptibility to conditions such as asthma, atopic dermatitis, and obesity.16–18 Breast milk represents the optimal source of nutrition for newborns, providing a complex array of components that extend beyond basic nutrients such as proteins, fats, and carbohydrates to include numerous bioactive factors.
These bioactive components such as lactoferrin, immunoglobulins, cytokines, growth factors and human milk oligosaccharides (HMOs) collectively contribute to the maturation of the infant's immune system. As the third most abundant solid component in breast milk, HMOs play particularly crucial roles: although not directly metabolized by infants, they serve as key prebiotics that selectively enrich beneficial gut microbiota,19–21 inhibit pathogen adhesion, and modulate intestinal mucosal immune responses, thereby shaping a healthy infant gut microbiome.22,23 Furthermore, breast milk contains a substantial quantity of live microorganisms, which are directly transmitted to the infant during breastfeeding and participate in the initial colonization and establishment of the neonatal gut microbiota.24
It is important to note that the breast milk microbiota is dynamic, and influenced by various maternal and external factors including the lactation stage, maternal health (such as gestational diabetes), gestational age, and diet.25–27 These maternally driven alterations in the milk microbiota can, in turn, be transmitted to the offspring, potentially influencing their microbial and metabolic health.28
Emerging evidence indicates that GDM can alter the content and composition of HMOs. Preliminary studies report that colostrum from mothers with GDM generally contains lower concentrations of HMOs, including key sialylated and fucosylated structures, with a significant reduction in specific components such as 3′-SL compared to the healthy controls.29,30 While some of these HMOs may show a transient increase after the colostrum phase, the differences between GDM and healthy mothers typically cease to be significant by three months postpartum.30 This temporal pattern suggests that GDM may specifically disrupt the timing of HMO synthesis and secretion. In contrast, research on the breast milk microbiota in GDM remains limited and inconsistent. For instance, one study by Gámez-Valdez et al. observed a higher Shannon index in the breast milk microbiota of GDM mothers,31 whereas others found no significant compositional differences between GDM and healthy mothers.11,32
Most existing studies have investigated the breast milk microbiota and HMOs in isolation, creating a significant knowledge gap regarding whether GDM currently alters both components and how they may interact to shape the infant gut microbiota. To address this, our study enrolled GDM mother–infant pairs, and collected breast milk and infant fecal samples at two critical time points: 0–7 days and 42 days postpartum. We employed a comprehensive approach, profiling the breast milk microbiota via metagenomic sequencing and quantifying core HMOs using liquid chromatography-tandem mass spectrometry (LC-MS/MS). By integrating these datasets, this research aims to elucidate the mechanisms underlying gut microecological imbalance in the GDM offspring, thereby providing a scientific foundation for developing early nutritional or microbial interventions.
2. Materials and methods
2.1 Study design and participants
This study, conducted as part of the project “Establishing a database of breast milk nutritional functional components and microbiota with Chinese characteristics”, received ethical approval from the Ethics Committee of the National Institute for Nutrition and Health, Chinese Center for Disease Control and Prevention (approval no.: 2020-023), and written informed consent was obtained from all participants. Between January 2022 and December 2023, we recruited both GDM and healthy control mother–infant pairs from Danyang People's Hospital. GDM diagnosis adhered to the International Association of Diabetes and Pregnancy Study Groups (IADPSG) criteria, which require meeting at least one of the following thresholds: fasting plasma glucose (FPG) between 5.1 and 7.0 mmol L−1, 1-hour oral glucose tolerance test (OGTT) ≥10.0 mmol L−1, or 2-hour OGTT between 8.5 and 11.1 mmol L−1. All enrolled mothers were of Han ethnicity, had delivered a single full-term infant (≥37 weeks of gestation), and intended to exclusively breastfeed for a minimum of 6 weeks postpartum. Infants were excluded from the study if they had congenital diseases, experienced perinatal complications, or required antibiotic treatment.Approximately 10 mL of breast milk was aseptically collected during the second morning feeding session at two time points 0–7 days postpartum (colostrum) and 42 days postpartum (mature milk). Fresh infant fecal samples were collected at the corresponding time points, immediately snap-frozen at −80 °C, and subsequently transported to the laboratory on dry ice for unified analysis.
2.2 DNA extraction and metagenomic sequencing
Genomic DNA extraction and metagenomic sequencing were conducted by Beijing Novogene Bioinformatics Technology Co., Ltd. The brief procedure was as follows: DNA was extracted from samples using the TIANamp stool DNA kit (Tiangen, China); 1 μg of DNA per sample was used for library construction. DNA was fragmented to an average size of 350 bp using Covaris LE220R-plus (Covaris, USA), followed by sequential steps including end repair, A-tailing, adapter ligation, and PCR amplification. The resulting libraries were purified by using the AMPure XP system (Beckman Coulter, USA), and assessed for quality using the Agilent 5400 system (AATI). Quantification was performed by real-time quantitative PCR (1.5 nM), and libraries meeting quality standards were pooled based on the effective concentration and subjected to PE150 sequencing on an Illumina platform.2.3 Metagenomic data analysis
Raw data were demultiplexed and converted to the FASTQ format. Quality control was performed using Trimmomatic to remove adapter sequences, reads with the N ratio exceeding 10%, and low-quality reads. The resulting high-quality sequences were then aligned to the human reference genome (hg38) using BWA. Subsequently, host-derived sequences were removed with data using samtools and bedtools, yielding clean, non-host data for the subsequent metagenomic analysis. Taxonomic profiling was performed using MetaPhlAn4 for broad relative abundance estimation. Microbial diversity was assessed by calculating alpha diversity (Shannon and Chao1 indexes) and beta diversity (PCoA based on Bray–Curtis distances) at the species level.2.4 Human milk oligosaccharide (HMO) analysis
Quantitative analysis of 32 HMOs was performed by Beijing Zhongke Hesheng Glycobiology Technology Co., Ltd. Following sample pretreatment, which included centrifugation for defatting, ethanol precipitation of proteins, and dilution with 50% acetonitrile (resulting in a total dilution factor of 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 60), HMOs were separated using a Waters ACQUITY BEH Amide column (130 Å, 1.7 μm, 2.1 × 150 mm) and detected using an Agilent 6475 triple quadrupole mass spectrometer operated in ESI negative ion mode with multiple reaction monitoring (MRM). Quantification was performed using the external standard method (formula: C = (S − b)/nm × H), with the relative standard deviation (RSD) of quality control samples maintained below 15%.
60), HMOs were separated using a Waters ACQUITY BEH Amide column (130 Å, 1.7 μm, 2.1 × 150 mm) and detected using an Agilent 6475 triple quadrupole mass spectrometer operated in ESI negative ion mode with multiple reaction monitoring (MRM). Quantification was performed using the external standard method (formula: C = (S − b)/nm × H), with the relative standard deviation (RSD) of quality control samples maintained below 15%.
2.5 Statistical analysis
All statistical analyses were performed using R language (v4.2.0) and SPSS software (version 27.0; IBM Corp., USA). Alpha diversity and beta diversity were calculated using the R packages vegan and phyloseq. Inter-group differences were evaluated using the t test and further examined via LEfSe analysis in R language. Associations between variables were assessed by Spearman's correlation analysis using SPSS; all graphical visualizations were generated using R language or GraphPad Prism 9.0. Statistical significance was denoted as follows: *p < 0.05, **p < 0.01, and *p < 0.001.3. Results
3.1 Participant characteristics
This study enrolled 23 healthy mothers and 28 mothers with GDM. From these participants, breast milk and the corresponding infant fecal samples were collected at two time points: 0–7 days and 42 days postpartum. Metagenomic sequencing was successfully performed on a total of 53 breast milk and 50 infant fecal samples. In parallel, HMO concentrations were quantitatively measured in 84 breast milk samples. The demographic and clinical characteristics of the mother–infant pairs, including maternal age, pre-pregnancy BMI, gestational age at delivery, and infant birth weight, are summarized in Table 1.| Characteristic | GDM group | Healthy group | P |
|---|---|---|---|
| Maternal | |||
| Age, years [mean ± SD] | 29.86 ± 2.65 | 28.22 ± 2.65 | 0.032 |
| Smoking status [n (%)] | |||
| Ever | 0 (0%) | 0 (0%) | |
| Never | 28 (100%) | 23 (100)% | |
| Drinking status [n (%)] | |||
| Ever | 0 (0%) | 0 (0%) | |
| Never | 28 (100%) | 23 (100)% | |
| Parity status [n (%)] | |||
| Nulliparae | 14 (50.00%) | 16 (69.57%) | |
| Multiparae | 14 (50.00%) | 7 (30.43%) | |
| Gestational age, days [mean ± SD] | 272.50 ± 6.63 | 276.48 ± 5.50 | 0.026 |
| Pre-pregnancy BMI [mean ± SD] | 24.04 ± 3.29 | 21.49 ± 2.46 | 0.003 |
| Blood glucose during pregnancy [mean ± SD] | |||
| Fasting glucose | 4.85 ± 0.43 | 4.30 ± 0.27 | <0.001 |
| OGTT 1 h | 9.40 ± 1.78 | 7.13 ± 1.18 | <0.001 |
| OGTT 2 h | 8.16 ± 1.90 | 6.33 ± 0.97 | <0.001 |
| Neonatal | |||
| Sex [n (%)] | |||
| Boy | 16 (57.14%) | 13 (56.52%) | |
| Girl | 12 (42.86%) | 10 (43.48%) | |
| Delivery status [n (%)] | |||
| Vaginal | 7 (25.00%) | 10 (43.48%) | |
| Cesarean | 21 (75.00%) | 13 (56.52%) | |
| Birth weight (BW), g [mean ± SD] | 3516.00 ± 427.48 | 3349.57 ± 268.32 | 0.101 |
3.2 Composition and differences in the breast milk microbiota of GDM mothers
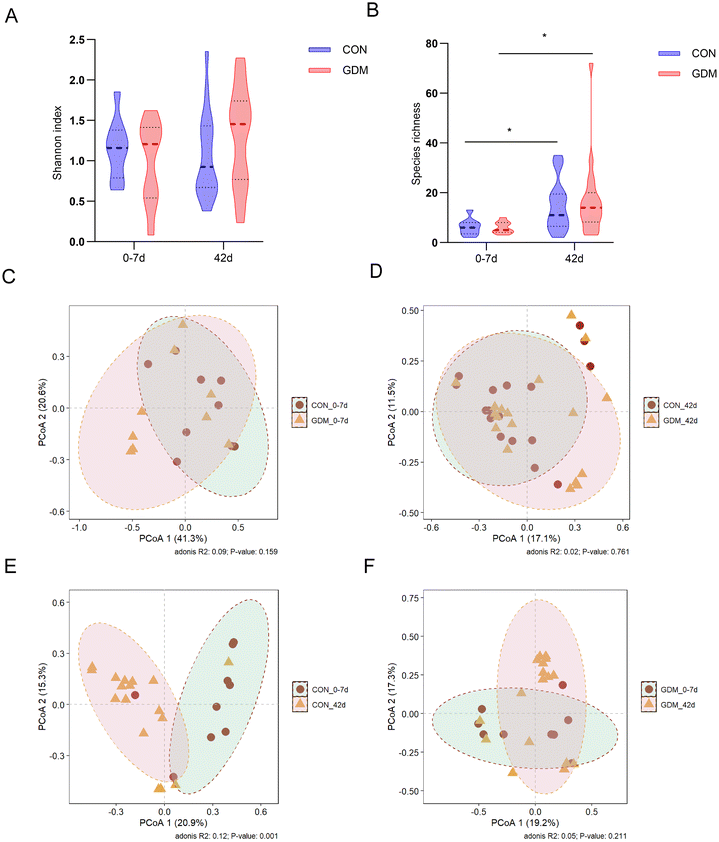
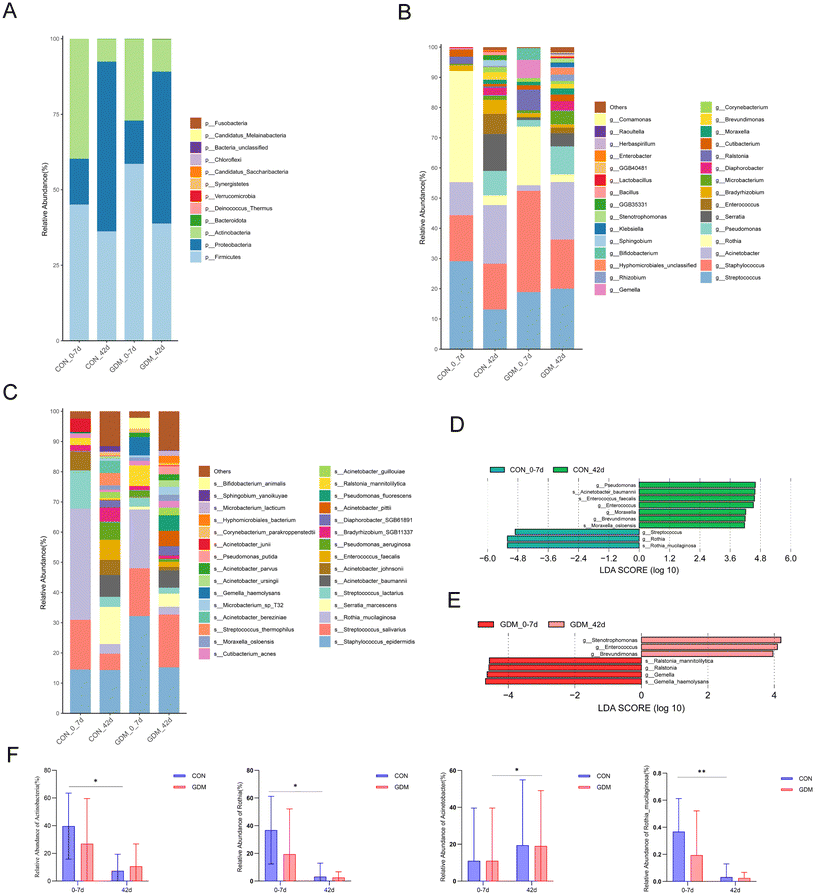
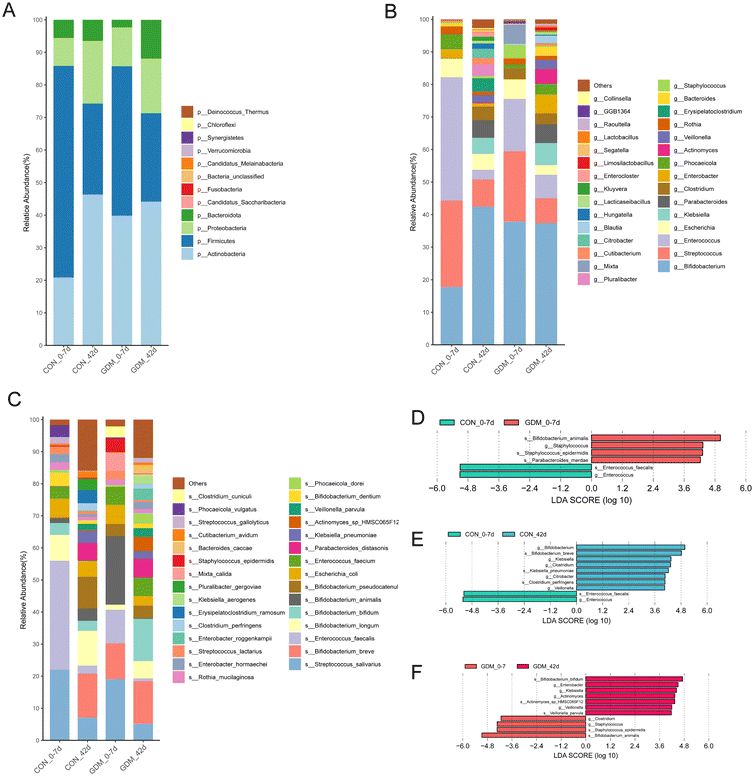
Analysis of the breast milk microbiota revealed no significant differences in alpha diversity (Shannon's index and species richness) between GDM and healthy mothers at either 0–7 days or 42 days postpartum (Fig. 1A and B). Consistent with this, beta diversity assessed by PCoA showed no significant separation between the two groups at both time points (Fig. 1C and D). Longitudinal analysis within each group indicated that species richness significantly increased at 42 days compared to 0–7 days in both groups, though the Shannon index remained unchanged. Notably, beta diversity in the healthy group shifted significantly over time (Fig. 1E), whereas the GDM group exhibited no such temporal variation (Fig. 1F), suggesting a distinct trajectory in microbial community dynamics.At the phylum level, the breast milk microbiota was predominantly composed of Firmicutes, Proteobacteria, and Actinobacteria (Fig. 2A), with no significant influence of the GDM status on the overall composition. Longitudinal analysis within the healthy control (CON) group revealed a significant decrease in the relative abundance of Actinobacteria by 42 days compared to the 0–7 days period (Fig. 2F). At the genus level, dominant genera included Streptococcus, Staphylococcus, Acinetobacter, Rothia, and Pseudomonas (Fig. 2B). While no significant genus-level differences were observed between GDM and the control mothers at either time points, temporal changes were evident within groups. Specifically, the relative abundance of Rothia significantly declined in healthy mothers over time. In contrast, a significant increase in Acinetobacter was observed at 42 days postpartum, but the shift was specific to the GDM group. At the species level, the breast milk microbiota was predominantly constituted by Staphylococcus epidermidis, Streptococcus salivarius, Rothia mucilaginosa, Serratia marcescens, and Streptococcus lactarius (Fig. 2C). Notably, the relative abundance of R. mucilaginosa in breast milk from healthy mothers exhibited a significant decrease at 42 days postpartum compared to the 0–7 days period.
To identify the microbial biomarkers, we performed linear discriminant analysis with effect size (LEfSe) on genus- and species-level taxa (LDA ≥4). No significant biomarkers distinguishing GDM from healthy breast milk were identified at the same lactation stage. From a longitudinal perspective, Enterococcus and Brevundimonas were significantly enriched at 42 days postpartum in both groups (Fig. 2D and E). Group-specific temporal shifts were also observed: the CON_42 d group exhibited enrichment of Pseudomonas, Acinetobacter baumannii, Enterococcus faecalis, Moraxella, and Moraxella osloensis, whereas the CON_0–7 d group was characterized by Streptococcus, Rothia, and Rothia mucilaginosa. In the GDM group, Stenotrophomonas and Brevundimonas were enriched at 0–7 days, while Ralstonia mannitolilytica, Ralstonia, and Gemella haemolysans were enriched at 42 days. This result indicated that while breast milk microbiota exhibited dynamic temporal enrichment patterns, the GDM status did not induce unique taxonomic enrichment at the same lactation stage compared to the healthy group.
3.3 Human milk oligosaccharide composition
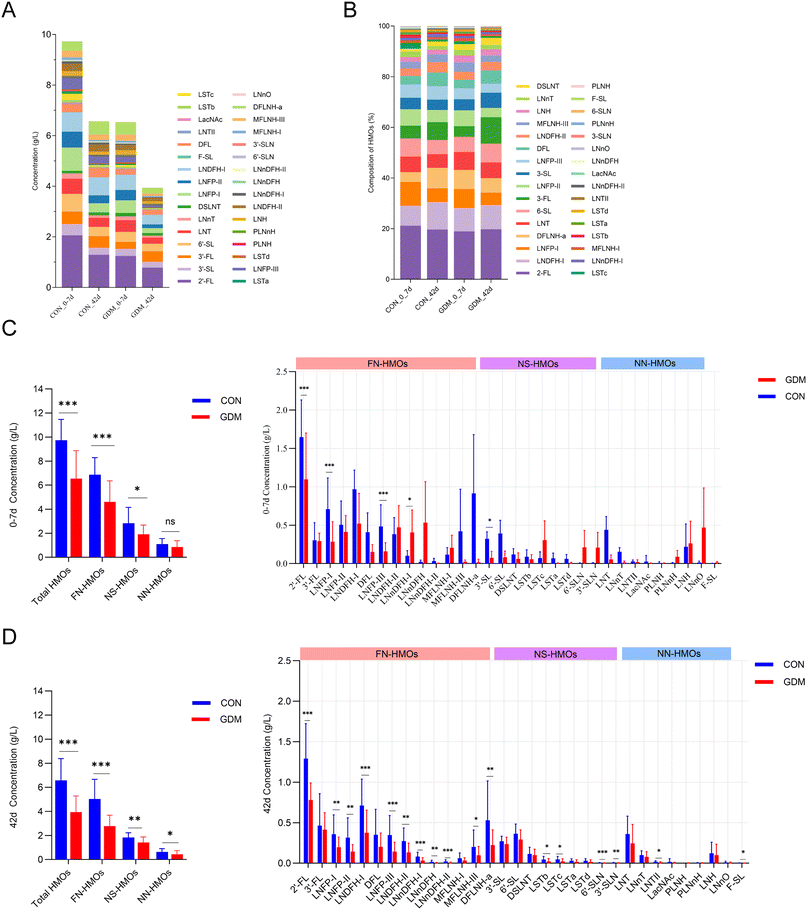
The composition of 32 core HMOs in 91 breast milk samples was quantified using LC-MS/MS. Phenotyping was conducted following the approach as described previously,33 where samples with 2′-FL ≥15 mg kg−1 were classified as secretor-positive (Se+), and those with LNFP-II ≥30 mg kg−1 were classified as Lewis-positive (Le+). This classification yielded four phenotypes: Se+Le+, Se+Le−, Se−Le+, and Se−Le−. The Se+Le+ group accounted for 89% (81/91) of the samples, and further analyses were focused exclusively on this type.In Se+Le+ breast milk, 2′-FL was the most predominant oligosaccharide, followed by LNDFH-I, 3′-FL, DFLNH-a, and LNFP-I (Fig. 3A and B). At comparable lactation stages, the GDM group exhibited a significantly lower total HMO concentration than the control group, with widespread reductions observed across most individual HMOs (Fig. 3C and D). As lactation progressed, total HMO levels declined over time in both groups, decreasing from 9.74 ± 1.74 g L−1 at 0–7 days to 6.57 ± 1.81 g L−1 at 42 days in the control, and from 6.55 ± 2.32 g L−1 to 3.94 ± 1.33 g L−1 in the GDM group. A similar decreasing trend was observed for 2′-FL, which decreased from 2.06 ± 0.50 g L−1 to 1.29 ± 0.43 g L−1 in the control group and from 1.25 ± 0.48 g L−1 to 0.78 ± 0.21 g L−1 in the GDM group.
When classified by the core structure, fucosylated non-sialylated HMOs (FN-HMOs) and non-fucosylated sialylated HMOs (NS-HMOs) were significantly lower in the GDM colostrum than in the control colostrum. These reductions persisted in mature milk, where FN-HMOs, NS-HMOs, and non-fucosylated non-sialylated HMOs (NN-HMOs) were all significantly decreased. Notably, four specific FN-HMOs, including 2′-FL, LNFP-I, LNFP-III, and LNnDFH-I, were significantly lower in the GDM group at both stages. By 42 days postpartum, the control group had significant higher concentrations of 17 individual HMOs comprising 11 FN-HMOs, 4 NS-HMOs, 1 NN-HMOs, and F-SL.
3.4 Alterations in the gut microbiota of the GDM offspring and their association with the breast milk microbiota and HMOs
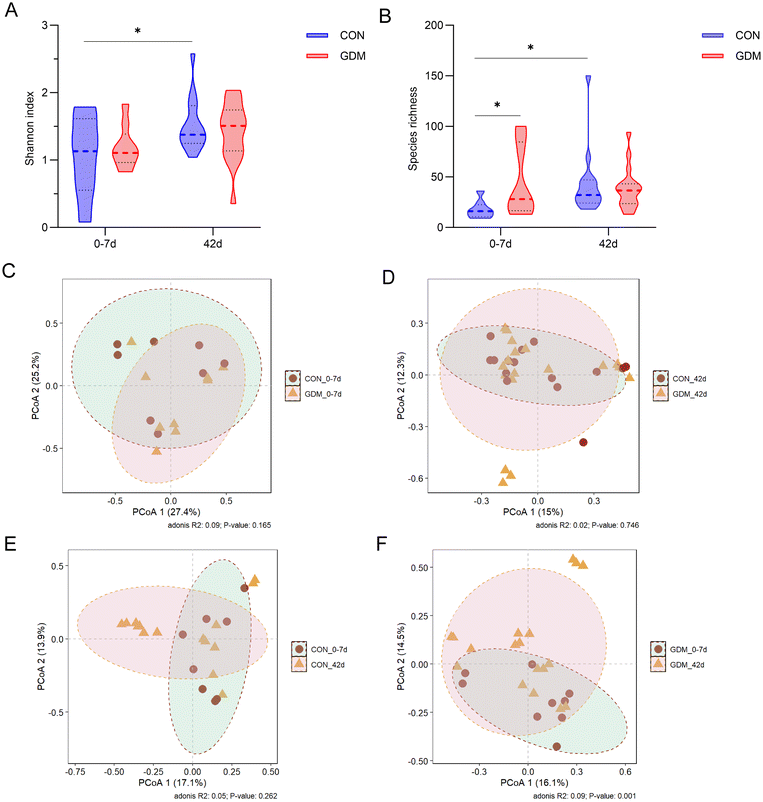
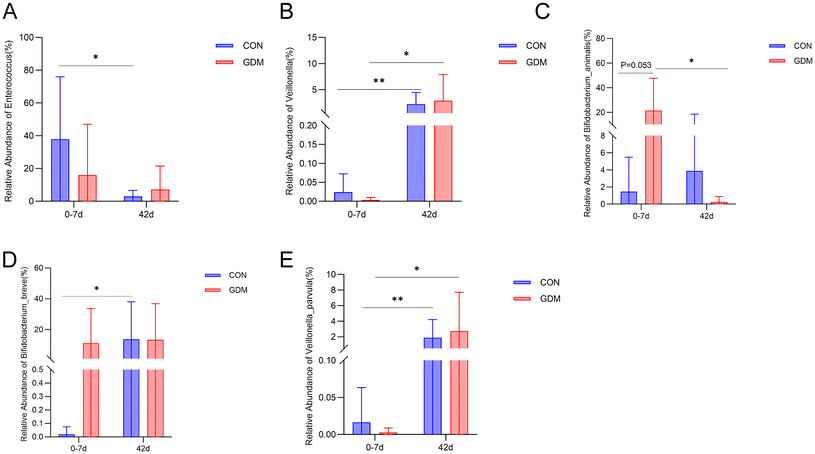
At 0–7 days, the offspring of GDM mothers had significantly higher gut microbiota species richness than those of healthy mothers (Fig. 4A), despite no significant difference in the Shannon index (Fig. 4B). Over time, from 0–7 to 42 days, all infants showed an increased Shannon index and species richness, with the increase being statistically significant in the control offspring. No significant separation was noticed in the beta diversity between the two groups at either 0–7 or 42 days postpartum via PCoA (Fig. 4C and D). Despite significant overall shifts over time in the GDM group, considerable inter-individual heterogeneity persisted (Fig. 4E and F).At the phylum level, the infant gut microbiota was primarily composed of by Actinobacteria, Firmicutes, Proteobacteria, and Bacteroidetes (Fig. 5A). At the genus level, the dominant taxa were Bifidobacterium, Streptococcus, Enterococcus, Escherichia, and Klebsiella (Fig. 5B), with Bifidobacterium being the most abundant. The CON group experienced a significant decline in Enterococcus and a rise in Veillonella over time (Fig. 6A and B). The species-level composition was dominated by core taxa such as Streptococcus salivarius, Bifidobacterium breve, Enterococcus faecalis, and Bifidobacterium longum (Fig. 5C). A trend towards a higher abundance of Bifidobacterium animalis was found in the GDM offspring compared to the control offspring (p = 0.053) at 0–7 days postpartum (Fig. 6C). As lactation progressed, the control offspring showed significant enrichment of Bifidobacterium breve and Veillonella parvula (Fig. 6D and E). LEfSe analysis (LDA score ≥4) revealed that the gut microbiota of GDM and the control offspring exhibited distinct, early-life microbial profiles that converged by day 42, highlighting the transient nature of the initial group differences. At 0–7 days, the GDM offspring was characterized by a significant enrichment of Bifidobacterium animalis, Staphylococcus, and Staphylococcus epidermidis, whereas the control offspring was enriched in Enterococcus and Enterococcus faecalis (Fig. 5D). Examining the temporal succession with each group revealed divergent development trajectories. The gut microbiota of the control offspring developed from an early community were enriched in Bifidobacterium, Bifidobacterium breve, Klebsiella, Citrobacter, Clostridium perfringens, and Veillonella to a later profile dominated by Enterococcus and Enterococcus faecalis (Fig. 5E). In contrast, the GDM offspring transitioned from an initial state enriched in Clostridium, Staphylococcus, Staphylococcus epidermidis, and Bifidobacterium animalis to a later state characterized by Bifidobacterium bifidum, Enterobacter, Klebsiella, Actinomyces, and Veillonella parvula (Fig. 5F).
3.5 Associations between the infant gut microbiota and breast milk microbiota
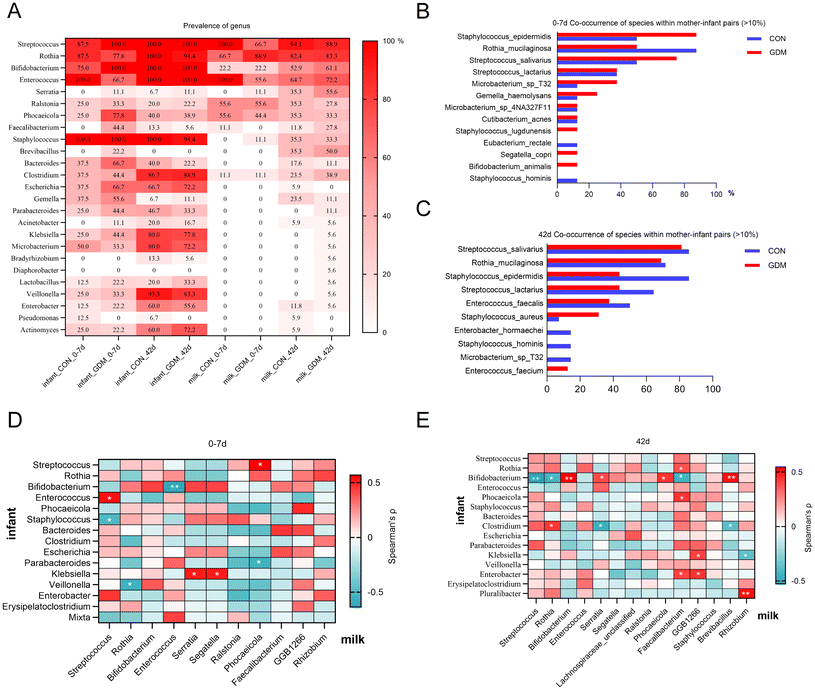
To assess the impact of GDM on microbial vertical transmission from breast milk to the infant gut, we analyzed the prevalence of core and potentially beneficial genera across colostrum, mature milk, and the corresponding infant feces (Fig. 7A). Genera such as Streptococcus, Rothia, and Enterococcus were frequently detected (>60% prevalence) in both breast milk and infant samples across all eight groups. The prevalence of Bifidobacterium in infant feces was notably high, reaching 100% in the GDM_0–7 d, CON_42 d, and GDM_42 d groups. However, its prevalence in breast milk was only 22.2% at 0–7 days in both maternal groups, increasing to over 50% by day 42. A more pronounced discrepancy was observed for Escherichia, Klebsiella, Microbacterium, and Veillonella. These taxa exhibited a high prevalence (>70%) in 42-day-old infants, yet were rarely detected in breast milk (prevalence below 5% and as low as 0%), suggesting a very low probability of direct vertical transmission from breast milk to the infant gut.We next performed a co-occurrence analysis on microbial species simultaneously detected in both breast milk and the corresponding infant gut at different time points, focusing on species present in more than 10% of the mother–infant pairs (Fig. 7B and C). The analysis identified Staphylococcus epidermidis, Rothia mucilaginosa, and Streptococcus salivarius as the most frequently shared species. At 0–7 days postpartum, the co-occurrence rate of R. mucilaginosa was significantly higher in the control mother–infant pairs than in the GDM pairs (87.5% vs. 50%), suggesting that its vertical transmission may be suppressed by GDM. In contrast, S. epidermidis co-occurred more frequently in the GDM group (87.5% vs. 50%) during the colostrum stage. However, the pattern reversed by day 42, with S. epidermidis co-occurrence becoming lower in the GDM group. Meanwhile the co-occurrence rate of R. mucilaginosa converged to a similar level (approximately 70%) in both groups.
To identify the genera potentially important in microbial transmission from breast milk to the infant gut, we performed a correlation analysis on 46 mother–infant pairs, focusing on genera with an average relative abundance exceeding 1% in each environment. At 0–7 days, the abundance of Streptococcus and Rothia in breast milk showed a non-significant negative correlation with those in the infant gut, whereas Bifidobacterium and Enterococcus exhibited non-significant positive correlations (Fig. 7D). By 42 days, the correlations for Streptococcus and Rothia had reversed to positive, and that for Enterococcus to negative. Notably, the correlations for Bifidobacterium remained positive and became statistically significant (Fig. 7E). The overall increase in significant positive correlations from 0–7 to 42 days indicates that the microbial communities in both niches not only change over time but also that their interrelations can reverse as the infant gut ecosystem develops.
3.6 Association between the infant gut microbiota and HMOs
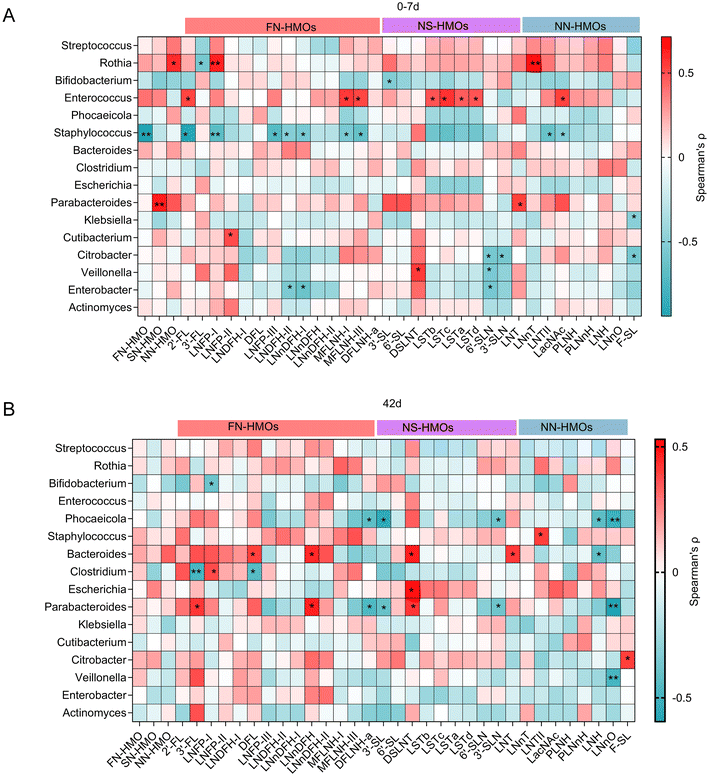
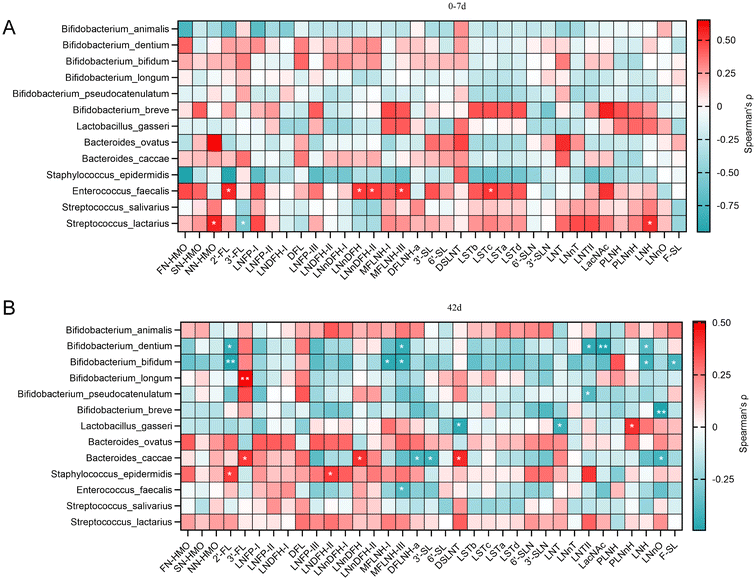
At 0–7 days postpartum, correlation analysis revealed a distinct association between the infant gut microbiota and HMOs. The genera Rothia, Enterococcus, and Parabacteroides were positively correlated with most HMOs, whereas Staphylococcus was negatively correlated with most HMOs (Fig. 8A). Specifically, Rothia exhibited significant positive correlations with NN-HMOs, LNFP-II, and LNnT. Parabacteroides was significantly positively correlated with SN-HMOs and LNT. Enterococcus showed significant positive correlations with eight HMOs including 2′-FL, MFLNH-I, and MFLNH-III. In contrast, Staphylococcus showed significant negative correlation with the FN-HMO group, which encompasses seven specific HMOs. At 42 days postpartum, the correlation profiles had shifted substantially. Neither Rothia nor Enterococcus maintained any significant positive correlation with the 32 measured HMOs (Fig. 8B). The associations for Parabacteroides with several HMOs were also reversed – specifically, its correlations with MFLNH-I and DFLNH – a shifting from negative to positive. Conversely, Staphylococcus exhibited a reversal to a positive correlation with most HMOs, showing a significant positive association with LNnT.To further elucidate the interactions between HMOs and the gut microbiota, we analyzed the correlations of the reported HMO-utilizing bacteria and the core infant gut species with specific HMO structures. In the first week of life, Bifidobacterium animalis and Staphylococcus epidermidis were significantly negatively correlated with multiple fucosylated HMOs, whereas the relative abundance of Enterococcus faecalis was positively associated with specific HMOs such as 2′-FL and LNnDFH (Fig. 9A). By postpartum day 42, the correlation profile had evolved: several Bifidobacterium species, including B. dentium and B. bifidum, showed broad negative correlations with a range of HMOs, reaching statistical significance for structures such as 2′-FL, MFLNH-III, and LNH. In contrast, Bacteroides displayed more positive associations, as exemplified by Bacteroides caccae, which was significantly correlated with 3′-FL, LNnDFH, and DSLNT (Fig. 9B).
4. Discussion
Recent studies suggested that vertical transmission of specific breast milk microorganisms plays a role in the early colonization of the infant gut.34 Since the maternal health status, particularly prenatal metabolic conditions such as GDM, can influence the composition of both breast milk microbiota and HMOs, this study aimed to investigate the specific impact of GDM on these components and to explore their subsequent association with the infant gut microbiota development. The breast milk microbiota is predominantly composed of the phyla Firmicutes, Proteobacteria, and Actinobacteria, with Staphylococcus, Streptococcus, and Pseudomonas identified as the key genera.35Consistent with the previous reports in which no significant alpha diversity difference was observed between the GDM and non-GDM groups at 1–3 weeks and one month postpartum,11,32 our study of the milk microbiota also detected no significant differences in alpha or beta diversity, or in the microbial composition at the phylum, genus, or species levels at either 0–7 or 42 days postpartum. These results collectively indicated that GDM had a limited effect on the breast milk microbiota. A key limitation was the sample size, which, while comparable to other studies,24,36 may have been insufficient to detect subtle GDM-induced shifts due to the inherent challenges of a low microbial biomass and high inter-individual variation in breast milk. Consequently, the conclusion that GDM did not significantly alter the breast milk microbiota warrants verification in larger cohorts. The composition of the breast milk microbiota undergoes dynamic changes throughout lactation.35,37 To assess longitudinal shifts, we directly compared the microbiota profiles between the colostrum and mature milk from the same mothers. While the Shannon index did not differ significantly, the control group showed a significant increase in species richness and distinct beta diversity between the two lactation stages. These findings aligned with a previous report which indicated no difference in the Shannon index between the colostrum (1–5 days postpartum) and mature milk (42 days postpartum).38 Similarly, it was reported that there was no significant change in the Shannon index of the breast milk microbiota within the first 8 weeks, with noticeable differences emerging only after this period.35 This suggested that shifts in the breast milk microbiota occur gradually and may require extended timeframes to become evident. In this study, the relative abundance of Rothia decreased as lactation progressed in both control and GDM mothers, consistent with previous reports.39 Moreover, the relative abundance of Acinetobacter significantly increased in the mature milk of GDM mothers, a trend contrasting with the observations in the healthy mothers,26,35 suggesting a potential association with the influence of the metabolic disease state.
HMOs serve as specific substrates for the infant gut microbiota and selectively promote the colonization and growth of beneficial microorganisms. In this study, the total concentration of the 32 core HMOs was significantly lower in mothers with GDM than in healthy mothers, observed in both colostrum and mature milk. Additionally, the contents of FN-HMOs and NS-HMOs were also significantly reduced at both stages. These findings were consistent with previous reports, which indicated that HMO levels were significantly reduced in the colostrum due to GDM.30,40,41 However, trends in the HMO changes in the mature milk stage vary across studies. Some reports found no significant difference in the HMO levels between the GDM and healthy groups at 42 days postpartum,30,41 whereas our study and that by Zhang et al.40 revealed persistently lower HMO levels in the mature milk of the GDM group. Notably, our study and that of Zhang et al. were conducted in Danyang and Wuxi, respectively, which are located in a nearby region in Southern China, while the other two studies with different conclusions were from Dalian and Beijing in northern China. We speculated that the influence of GDM on the HMO profiles may exhibit regional variations, potentially linked to genetic background, dietary habits, or environmental differences between the southern and northern populations.
Furthermore, the influence of GDM on HMO profiles was observed to be more pronounced at the mature milk stage than in the colostrum. Specifically, the number of HMOs showing a significant reduction increased from only 4 in the colostrum to 17 in mature milk. Notably, all four HMOs that were significantly reduced in the colostrum (2′-FL, LNFP-I, LNFP-III, and LNnDFH-I) remained at lower concentrations in mature milk. This pattern aligns with the report by Li et al. concerning LNFP-I, suggesting that the suppressive effect of GDM on HMO synthesis may intensify as lactation progresses.30 Moreover, consistent with previous studies,41,42 both total HMO concentration and the levels of most individual HMOs gradually declined over the course of lactation, further supporting that HMO synthesis and secretion were strongly modulated by the lactation stage.43
Growing evidence suggests that maternal GDM affects offspring health, yet the association between GDM and the offspring gut microbiota remained incompletely understood. In this study, the species richness of the gut microbiota in the GDM offspring was significantly higher than that in the control offspring at 0–7 days, though this difference was no longer observed at 42 days. Beta diversity, however, did not differ significantly between the two groups at either time point. Consistent with a previous report,8 gut microbiota species richness in the GDM offspring was reduced at 1 week of age but showed no significant difference by 9 months. It was reported that there was no difference in alpha or beta diversity between the GDM and the control offspring.40,44,45 In contrast, other studies have indicated that GDM significantly reduced alpha diversity in the offspring.9,46 This heterogeneity may be attributable to variations in sampling times, given the rapid development of the infant gut microbiota in early life, during which differences at specific time points can be substantial.47 The dominant genera in the infant gut, Bifidobacterium, Streptococcus, Enterococcus, Escherichia, and Klebsiella, were consistent with previous reports.48 Beyond confirming this established profile, our study provides, for the first time, a species-level characterization of gut microbiota differences between GDM and normal offspring. We found that at 0–7 days, Bifidobacterium animalis, Staphylococcus epidermidis, and Parabacteroides merdae were enriched in the GDM group, whereas Enterococcus faecalis was more abundant in the control group. Together with the observed difference in alpha diversity at 0–7 days, these findings suggested that maternal GDM exerts a more pronounced effect on the offspring gut microbiota during the colostrum stage.
To explore the potential vertical transmission of microbial species from breast milk to the infant gut and evaluate the influence of GDM on this process, we examined the prevalence of highly abundant species shared between breast milk and infant feces. Streptococcus, Rothia, and Enterococcus were frequently detected in both breast milk and infant gut (prevalence >60%), consistent with previous reports,24 indicating a high likelihood of vertical transmission. Bifidobacterium, although less prevalent in breast milk, is a dominant genus in the infant gut.49 We observed a positive correlation between its relative abundance in breast milk and the infant gut at both lactation stages, supporting its potential transfer via breastfeeding.50 Correlation analysis revealed dynamic shifts in the mother–infant microbial relationship over time. For instance, Streptococcus and Rothia exhibited a negative correlation at 0–7 days, which shifted to a positive correlation by day 42. Moreover, the number of significant positive correlations between breast milk and infant gut microbes was higher at day 42 than that in the early postpartum period. These findings suggested that both microbial communities undergo temporal development, with their interrelationship strengthening as lactation progresses. Furthermore, this study showed that the effect of GDM on microbial vertical transmission is stage-specific. At 0–7 days, GDM promoted the transmission of S. epidermidis, but this pattern was reversed by day 42, when its co-occurrence rate became lower in the GDM group than that in the controls. This dynamic shift revealed that the influence of GDM on microbial transmission was not uniform but time-dependent, potentially mediated through changes in the breast milk composition.
Correlation analysis between the infant gut microbiota and HMOs revealed that some associations reversed during lactation. For instance, significant negative correlations observed between S. epidermidis and specific HMOs (including LNT-II and 2′-FL) at 0–7 days shifted to positive correlations by day 42. The reversal was closely linked to the dramatic ecological succession in the infant gut. Key driving factors included the coordinated evolution of the intestinal redox status and the host immune environment. Postnatally, the gut transitions from a relatively oxygen-rich state to a strictly anaerobic environment, driving a shift in microbial dominance from aerobic pioneers like Enterobacteriaceae to obligate anaerobes such as Bifidobacterium.51,52 Concurrently, the host immune system matures rapidly, with mechanisms like secretory IgA-targeted binding and regulatory T cell-driven tolerance becoming more refined. This maturation exerts precise selective pressure that favors microbes capable of utilizing breast milk components and promoting immune homeostasis.53
This study observed that Bifidobacterium, a well-established utilizer of HMOs, exhibited limited positive correlations with HMOs, whereas another key utilizer, Bacteroides, showed positive correlations with multiple HMOs. Specifically, Bacteroides ovatus was positively associated with the majority of HMOs at 42 days. Previous reports suggested that in the absence of Bifidobacterium, Bacteroides tends to become dominant, suggesting a potential mutually exclusive relationship mediated by HMO consumption.54 Critically, beyond fueling competition, HMOs also foster cooperative interactions within the gut microbiota. This indicates that HMOs influence the microbial community not only via direct utilization but also through indirect ecological mechanisms like cross-feeding. It is well-established that primary degraders, including Bifidobacterium and Bacteroides, break down complex HMOs into metabolites such as short-chain fatty acids and sialic acid. These products, in turn, serve as growth substrates for other bacteria, thereby creating a trophic network.55 This cross-feeding encompasses complex interactions, from the sharing of core structural sugars like lacto-N-biose by B. bifidum to boost co-colonizing Bifidobacteria,56 to cross-utilization of metabolic intermediates like 1,2-propanediol between species such as B. infantis and Eubacterium hallii.57 Consequently, the GDM-induced reduction in HMOs may disrupt this intricate network, exerting stage-specific effects on the assembly of the offspring gut microbiota and potentially driving microecological dysbiosis.
5. Conclusion
This study shows that GDM only modestly affects the breast milk microbiota but substantially reduces HMO levels, particularly in mature milk. Although GDM did not alter the overall microbial composition of breast milk, it nonetheless modulated the vertical transmission of specific bacteria in a stage-dependent manner. The associated HMO alterations induced stage-specific shifts in the offspring's gut microbiota, as evidenced by the notable reversal of HMO–microbe correlations over time. The emergence of these dynamic interactions suggests that HMOs influenced microbial colonization not only through direct utilization but also, and perhaps more importantly, via indirect ecological mechanisms such as cross-feeding. Overall, GDM-induced alterations in breast milk HMOs, more than microbial transmission, play a key role in shaping the infant gut microbiota. These findings advance our understanding of how maternal metabolic health programs the early-life microbiome and suggest that HMO-based interventions could help improve metabolic outcomes in GDM-exposed infants.Author contributions
Feng Xu: conceptualization, methodology, investigation, formal analysis, data curation, writing – original draft, and visualization. Bo Yang: conceptualization, resources, writing – review & editing, supervision, project administration, and funding acquisition. Shumao Cui: investigation and validation. Zhenyu Yang: investigation and software. Nan Dai: resources. Catherine Stanton: writing – review & editing. R. Paul Ross: writing – review & editing. Jianxin Zhao: supervision. Jianqiang Lai: methodology. Wei Chen: writing – review & editing, supervision and funding acquisition. Ye Wang: writing – review & editing, supervision and funding acquisition.Conflicts of interest
There are no conflicts of interest to declare.Data availability
The data that support the findings of this study are available from the corresponding author Bo Yang upon reasonable request.Acknowledgements
This research was supported by the Research Program of State Key Laboratory of Food Science and Resources, Jiangnan University (no. SKLF-ZZB-202510), the Collaborative Innovation Centre of Food Safety and Quality Control in Jiangsu Province and the Fundamental Research Funds for the Central Universities.References
- R. Mittal, K. Prasad, J. R. N. Lemos, G. Arevalo and K. Hirani, Unveiling Gestational Diabetes: An Overview of Pathophysiology and Management, Int. J. Mol. Sci., 2025, 26, 2320 CrossRef CAS PubMed
.
- M. E. Bianco and J. L. Josefson, Hyperglycemia During Pregnancy and Long-Term Offspring Outcomes, Curr. Diabetes Rep., 2019, 19, 143 CrossRef PubMed
.
- M. M. Pathirana, Z. S. Lassi, A. Ali, M. A. Arstall, C. T. Roberts and P. H. Andraweera, Association between metabolic syndrome and gestational diabetes mellitus in women and their children: a systematic review and meta-analysis, Endocrine, 2021, 71, 310–320 CrossRef CAS PubMed
.
- S. Butalia, L. Donovan, A. Savu, J. Johnson, A. Edwards and P. Kaul, Postpartum Diabetes Testing Rates after Gestational Diabetes Mellitus in Canadian Women: A Population-Based Study, Can. J. Diabetes, 2017, 41, 613–620 CrossRef PubMed
.
- R. Retnakaran and B. R. Shah, Glucose screening in pregnancy and future risk of cardiovascular disease in women: a retrospective, population-based cohort study, Lancet Diabetes Endocrinol., 2019, 7, 378–384 CrossRef PubMed
.
- E. Al Bekai, C. E. Beaini, K. Kalout, O. Safieddine, S. Semaan, F. Sahyoun, H. E. Ghadieh, S. Azar, A. Kanaan and F. Harb, The Hidden Impact of Gestational Diabetes: Unveiling Offspring Complications and Long-Term Effects, Life, 2025, 15, 440 CrossRef CAS
.
- T. Chen, Y. Qin, M. Chen, Y. Zhang, X. Wang, T. Dong, G. Chen, X. Sun, T. Lu, R. A. White, P. Ye, H. M. Tun and Y. Xia, Gestational diabetes mellitus is associated with the neonatal gut microbiota and metabolome, BMC Med., 2021, 19, 120 CrossRef CAS PubMed
.
- M. K. W. Crusell, T. H. Hansen, T. Nielsen, K. H. Allin, M. C. Ruhlemann, P. Damm, H. Vestergaard, C. Rorbye, N. R. Jorgensen, O. B. Christiansen, F. A. Heinsen, A. Franke, T. Hansen, J. Lauenborg and O. Pedersen, Comparative Studies of the Gut Microbiota in the Offspring of Mothers With and Without Gestational Diabetes, Front. Cell. Infect. Microbiol., 2020, 10, 536282 CrossRef CAS PubMed
.
- M. L. Su, Y. Y. Nie, R. C. Shao, S. H. Duan, Y. H. Jiang, M. Y. Wang, Z. C. Xing, Q. Sun, X. H. Liu and W. M. Xu, Diversified gut microbiota in newborns of mothers with gestational diabetes mellitus, PLoS One, 2018, 13, e0205695 CrossRef PubMed
.
- E. Aja and J. P. Jacobs, Mommy's microbes: Gestational diabetes mellitus shapes the maternal and infant gut microbiome, Cell Host Microbe, 2024, 32, 1048–1049 CrossRef CAS PubMed
.
- S. Y. Valencia-Castillo, M. J. Hernández-Beza, I. Powell-Cerda, E. Acosta-Cruz, G. C. Rodríguez-Castillejos, F. Siller-López and H. Martínez-Montoya, Impact of gestational diabetes mellitus in gut and human breast milk microbiome in Colombian women and their infants, Rev. Argent. Microbiol., 2025, 57, 14–23 Search PubMed
.
- L. L. Huang, P. Sililas, C. Thonusin, T. Tongsong, S. Luewan, N. Chattipakorn and S. C. Chattipakorn, Association Between Gut Microbiota and Insulin Therapy in Women With Gestational Diabetes Mellitus, Can. J. Diabetes, 2022, 46, 804–812 CrossRef PubMed
.
- Z. Y. Song, S. M. Li and R. Q. Li, An Investigation into the Correlation of Intestinal Flora with Obesity and Gestational Diabetes Mellitus, Comput. Math. Methods, 2022, 2022, 5677073 Search PubMed
.
- K. Donald and B. B. Finlay, Early-life interactions between the microbiota and immune system: impact on immune system development and atopic disease, Nat. Rev. Immunol., 2023, 23, 735–748 CrossRef CAS PubMed
.
- A. Sarkar, J. Y. Yoo, S. Valeria Ozorio Dutra, K. H. Morgan and M. Groer, The Association between Early-Life Gut Microbiota and Long-Term Health and Diseases, J. Clin. Med., 2021, 10, 459 CrossRef PubMed
.
- O. Dumas, A. C. Arroyo, M. K. Faridi, K. James, S. Hsu, C. Powe and C. A. Camargo Jr., Cohort Study of Maternal Gestational Weight Gain, Gestational Diabetes, and Childhood Asthma, Nutrients, 2022, 14 CrossRef PubMed
.
- R. Kumar, F. X. Ouyang, R. E. Story, J. A. Pongracic, X. M. Hong, G. Y. Wang, C. Pearson, K. Ortiz, H. Bauchner and X. B. Wang, Gestational diabetes, atopic dermatitis, and allergen sensitization in early childhood, J. Allergy Clin. Immunol., 2009, 124, 1031–1038 CrossRef CAS PubMed
.
- C. D. Bendor, A. Bardugo, R. S. Rotem, E. Derazne, H. C. Gerstein, D. Tzur, O. Pinhas-Hamiel, A. M. Tsur, T. Cukierman-Yaffe, Y. Lebenthal, A. Afek, G. Chodick and G. Twig, Glucose Intolerance in Pregnancy and Offspring Obesity in Late Adolescence, Diabetes Care, 2022, 45, 1540–1548 CrossRef CAS PubMed
.
- B. Berger, N. Porta, F. Foata, D. Grathwohl, M. Delley, D. Moine, A. Charpagne, L. Siegwald, P. Descombes, P. Alliet, G. Puccio, P. Steenhout, A. Mercenier and N. Sprenger, Linking Human Milk Oligosaccharides, Infant Fecal Community Types, and Later Risk To Require Antibiotics, mBio, 2020, 11, e03196–19 CrossRef CAS PubMed
.
- A. Marcobal, M. Barboza, E. D. Sonnenburg, N. Pudlo, E. C. Martens, P. Desai, C. B. Lebrilla, B. C. Weimer, D. A. Mills, J. B. German and J. L. Sonnenburg, Bacteroides in the infant gut consume milk oligosaccharides via mucus-utilization pathways, Cell Host Microbe, 2011, 10, 507–514 CrossRef CAS PubMed
.
- T. Katayama, Host-derived glycans serve as selected nutrients for the gut microbe: human milk oligosaccharides and bifidobacteria†, Biosci., Biotechnol., Biochem., 2016, 80, 621–632 CrossRef CAS PubMed
.
- S. Zhang, T. Li, J. Xie, D. Zhang, C. Pi, L. Zhou and W. Yang, Correction to: Gold standard for nutrition: a review of human milk oligosaccharide and its effects on infant gut microbiota, Microb. Cell Fact., 2021, 20, 140 CrossRef
.
- C. Kong, M. Beukema, M. Wang, B. J. de Haan and P. de Vos, Human milk oligosaccharides and non-digestible carbohydrates prevent adhesion of specific pathogens via modulating glycosylation or inflammatory genes in intestinal epithelial cells, Food Funct., 2021, 12, 8100–8119 RSC
.
- C. Qi, H. Y. Tu, J. B. Zhou, R. D. Tu, H. Chang, J. Chen, H. T. Hu, R. Q. Yu and J. Sun, Widespread vertical transmission of secretory immunoglobulin A coated trace bacterial variants from the mother to infant gut through breastfeeding, Food Funct., 2022, 13, 11543–11554 RSC
.
- P. Singh, N. Al Mohannadi, S. Murugesan, F. Almarzooqi, B. S. A. Kabeer, A. K. Marr, T. Kino, T. Brummaier, A. Terranegra, R. McGready, F. Nosten, D. Chaussabel and S. Al Khodor, Unveiling the dynamics of the breast milk microbiome: impact of lactation stage and gestational age, J. Transl. Med., 2023, 21, 784 CrossRef CAS PubMed
.
- H. D. Ge, W. X. Zhu, J. Zhang, Z. J. Wang, H. J. Shi, J. Sun and M. Shi, Human milk microbiota and oligosaccharides in colostrum and mature milk: comparison and correlation, Front. Nutr., 2024, 11, 1512700 CrossRef PubMed
.
- P. Mokhtari, K. A. Schmidt, H. Zamanian, M. Babaei, C. J. Machle, D. Trifonova, T. L. Alderete, E. A. Holzhausen, J. Ottino-González, B. N. Chalifour, R. B. Jones, A. Furst, C. Yonemitsu, L. Bode and M. I. Goran, Maternal Diet Associated with Oligosaccharide Abundances in Human Milk from Latina Mothers, Nutrients, 2024, 16, 1795 CrossRef CAS PubMed
.
- C. Xue, Q. Xie, C. Zhang, Y. Hu, X. Song, Y. Jia, X. Shi, Y. Chen, Y. Liu, L. Zhao, F. Huang and H. Yuan, Vertical transmission of the gut microbiota influences glucose metabolism in offspring of mice with hyperglycaemia in pregnancy, Microbiome, 2022, 10, 122 CrossRef CAS PubMed
.
- X. Wang, J. Liu, C. Li, Y. Xu, X. Wang, Y. Lu, T. Zhang, H. Cao, L. Huang and Z. Wang, Pregnancy-Related Diseases and Delivery Mode can Affect the Content of Human Milk Oligosaccharides: A Preliminary Study, J. Agric. Food Chem., 2022, 70, 5207–5217 CrossRef CAS
.
- X. K. Li, X. X. Ning, B. Q. Rui, Y. S. Wang, Z. J. Lei, D. Yu, F. T. Liu, Y. J. Deng, J. L. Yuan, W. Z. Li, J. Y. Yan and M. Li, Alterations of milk oligosaccharides in mothers with gestational diabetes mellitus impede colonization of beneficial bacteria and development of ROR?t+ Treg cell-mediated immune tolerance in neonates, Gut Microbes, 2023, 15, 2256749 CrossRef PubMed
.
- J. S. Gámez-Valdez, J. F. García-Mazcorro, A. H. Montoya-Rincón, D. L. Rodríguez-Reyes, G. Jiménez-Blanco, M. T. A. Rodríguez, R. P.-C. de Vaca, M. R. Alcorta-García, M. Brunck, V. J. Lara-Díaz and C. Licona-Cassani, Differential analysis of the bacterial community in colostrum samples from women with gestational diabetes mellitus and obesity, Sci. Rep., 2021, 11, 24373 CrossRef PubMed
.
- L. S. Rold, J. M. Guldbæk, C. S. Lindegaard, S. Kirk, L. D. Nygaard, C. Bundgaard-Nielsen, J. N. Holm-Jacobsen, P. Leutscher, A.-C. F. Viuff, S. Hagstrøm and S. Sørensen, A comparison of the breast milk microbiota from women diagnosed with gestational diabetes mellitus and women without gestational diabetes mellitus, BMC Pregnancy Childbirth, 2024, 24, 412 CrossRef CAS
.
- M. Wang, Z. Zhao, A. Zhao, J. Zhang, W. Wu, Z. Ren, P. Wang and Y. Zhang, Neutral Human Milk Oligosaccharides Are Associated with Multiple Fixed and Modifiable Maternal and Infant Characteristics, Nutrients, 2020, 12, 826 CrossRef CAS PubMed
.
- Z. Zhong, H. Tang, T. Shen, X. Ma, F. Zhao, L.-Y. Kwok, Z. Sun, M. Bilige and H. Zhang, Bifidobacterium animalis subsp. lactis Probio-M8 undergoes host adaptive evolution by glcU mutation and translocates to the infant's gut via oral-/entero-mammary routes through lactation, Microbiome, 2022, 10, 197 CrossRef CAS PubMed
.
- K. E. Lyons, C.-A. O. Shea, G. Grimaud, C. A. Ryan, E. Dempsey, A. L. Kelly, R. P. Ross and C. Stanton, The human milk microbiome aligns with lactation stage and not birth mode, Sci. Rep., 2022, 12, 5598 CrossRef CAS PubMed
.
- S. Jiang, J. Qin, L. Shi, J. Feng, J. Mo, W. Su, Y. Cheng, J. Lv, Q. Li, S. Li, L. Zeng, B. Han and J. Zhou, Association among Gestational Weight Gain, Fucosylated Human Milk Oligosaccharides, and Breast Milk Microbiota—An Evidence in Healthy Mothers from Northwest China, J. Agric. Food Chem., 2024, 72, 25135–25145 CrossRef CAS PubMed
.
- P. S. Pannaraj, F. Li, C. Cerini, J. M. Bender, S. Yang, A. Rollie, H. Adisetiyo, S. Zabih, P. J. Lincez, K. Bittinger, A. Bailey, F. D. Bushman, J. W. Sleasman and G. M. Aldrovandi, Association Between Breast Milk Bacterial Communities and Establishment and Development of the Infant Gut Microbiome, JAMA Pediatr., 2017, 171, 647–654 CrossRef PubMed
.
- H. Ge, W. Zhu, J. Zhang, Z. Wang, H. Shi, J. Sun and M. Shi, Human milk microbiota and oligosaccharides in colostrum and mature milk: comparison and correlation, Front. Nutr., 2024, 11, 1512700 CrossRef PubMed
.
- B. Liu, J. Y. Zhao, Y. P. Liu, W. C. Qiao, T. M. Jiang and L. J. Chen, Diversity and temporal dynamics of breast milk microbiome and its influencing factors in Chinese women during the first 6 months postpartum, Front. Microbiol., 2022, 13, 1016759 CrossRef PubMed
.
- L. Zhang, J. Zhang, Y. Yan, S. Duan, X. Wang, I. M.-Y. Szeto, J. Liu, Z. Sun, X. Feng, K. Hettinga and P. Zhou, The changes in HMOs of GDM mothers over lactation: Regulation on the gut microbiota development in offspring, Food Biosci., 2024, 61, 104957 CrossRef CAS
.
- Y. Dou, Y. Luo, Y. Xing, H. Liu, B. Chen, L. Zhu, D. Ma and J. Zhu, Human Milk Oligosaccharides Variation in Gestational Diabetes Mellitus Mothers, Nutrients, 2023, 15, 1441 CrossRef CAS PubMed
.
- J. F. Plows, P. K. Berger, R. B. Jones, T. L. Alderete, C. Yonemitsu, J. A. Najera, S. Khwajazada, L. Bode and M. I. Goran, Longitudinal Changes in Human Milk Oligosaccharides (HMOs) Over the Course of 24 Months of Lactation, J. Nutr., 2021, 151, 876–882 CrossRef CAS PubMed
.
- X. Ren, J. Yan, Y. Bi, P. W. Shuttleworth, Y. Wang, S. Jiang, J. Wang, Y. Duan, J. Lai and Z. Yang, Human Milk Oligosaccharides Are Associated with Lactation Stage and Lewis Phenotype in a Chinese Population, Nutrients, 2023, 15, 1408 CrossRef CAS PubMed
.
- Q. Song, T. Zhou, S. Chen, Y. Liao, H. Huang, B. Xiao, J. V. Zhang, L. Ma and Y. Zhu, Association of Gestational Diabetes With the Dynamic Changes of Gut Microbiota in Offspring From 1 to 6 Months of Age, J. Clin. Endocrinol. Metab., 2023, 108, 2315–2323 CrossRef CAS PubMed
.
- T. K. Soderberg, C. M. Carpenter, R. C. Janssen, T. L. Weir, C. E. Robertson, D. Ir, B. E. Young, N. F. Krebs, T. L. Hernandez, L. A. Barbour, D. N. Frank, M. Kroehl and J. E. Friedman, Gestational Diabetes Is Uniquely Associated With Altered Early Seeding of the Infant Gut Microbiota, Front. Endocrinol., 2020, 11, 603021 CrossRef
.
- T. Chen, Y. F. Qin, M. J. Chen, Y. Q. Zhang, X. Wang, T. Y. Dong, G. L. Chen, X. Sun, T. Lu, R. A. White, P. Ye, H. M. Tun and Y. K. Xia, Gestational diabetes mellitus is associated with the neonatal gut microbiota and metabolome, BMC Med., 2021, 19, 120 CrossRef CAS PubMed
.
- S. Jeong, Factors influencing development of the infant microbiota: from prenatal period to early infancy, Clin. Exp. Pediatr., 2022, 65, 439–447 CAS
.
- Y. Li, L. Ren, Y. Wang, J. Li, Q. Zhou, C. Peng, Y. Li, R. Cheng, F. He and X. Shen, The Effect of Breast Milk Microbiota on the Composition of Infant Gut Microbiota: A Cohort Study, Nutrients, 2022, 14, 5397 CrossRef CAS PubMed
.
- B. Yang, Y. Chen, C. Stanton, R. P. Ross, Y.-K. Lee, J. Zhao, H. Zhang and W. Chen, Bifidobacterium and Lactobacillus Composition at Species Level and Gut Microbiota Diversity in Infants before 6 Weeks, Int. J. Mol. Sci., 2019, 20, 3306 CrossRef CAS PubMed
.
- J. Ding, R. Ouyang, S. Zheng, Y. Wang, Y. Huang, X. Ma, Y. Zou, R. Chen, Z. Zhuo, Z. Li, Q. Xin, L. Zhou, S. Mei, J. Yan, X. Lu, Z. Ren, X. Liu and G. Xu, Effect of Breastmilk Microbiota and Sialylated Oligosaccharides on the Colonization of Infant Gut Microbial Community and Fecal Metabolome, Metabolites, 2022, 12, 1136 Search PubMed
.
- J. E. Koenig, A. Spor, N. Scalfone, A. D. Fricker, J. Stombaugh, R. Knight, L. T. Angenent and R. E. Ley, Succession of microbial consortia in the developing infant gut microbiome, Proc. Natl. Acad. Sci. U. S. A., 2011, 108(Suppl 1), 4578–4585 CrossRef PubMed
.
- C. J. Stewart, N. J. Ajami, J. L. O'Brien, D. S. Hutchinson, D. P. Smith, M. C. Wong, M. C. Ross, R. E. Lloyd, H. Doddapaneni, G. A. Metcalf, D. Muzny, R. A. Gibbs, T. Vatanen, C. Huttenhower, R. J. Xavier, M. Rewers, W. Hagopian, J. Toppari, A.-G. Ziegler, J.-X. She, B. Akolkar, A. Lernmark, H. Hyoty, K. Vehik, J. P. Krischer and J. F. Petrosino, Temporal development of the gut microbiome in early childhood from the TEDDY study, Nature, 2018, 562, 583–588 CrossRef CAS PubMed
.
- A. Rey-Marino and M. P. Francino, Nutrition, Gut Microbiota, and Allergy Development in Infants, Nutrients, 2022, 14, 4316 CrossRef CAS PubMed
.
- C. Lin, Y. Lin, H. Zhang, G. Wang, J. Zhao, H. Zhang and W. Chen, Intestinal ‘Infant-Type’ Bifidobacteria Mediate Immune System Development in the First 1000 Days of Life, Nutrients, 2022, 14, 1498 CrossRef CAS PubMed
.
- R. Díaz and D. Garrido, Screening competition and cross-feeding interactions during utilization of human milk oligosaccharides by gut microbes, Microbiome Res. Rep., 2024, 3, 12 Search PubMed
.
- K. Nishiyama, A. Nagai, K. Uribayashi, Y. Yamamoto, T. Mukai and N. Okada, Two extracellular sialidases from Bifidobacterium bifidum promote the degradation of sialyl-oligosaccharides and support the growth of Bifidobacterium breve, Anaerobe, 2018, 52, 22–28 CrossRef CAS PubMed
.
- C. Schwab, H. J. Ruscheweyh, V. Bunesova, V. T. Pham, N. Beerenwinkel and C. Lacroix, Trophic Interactions of Infant Bifidobacteria and Eubacterium hallii during L-Fucose and Fucosyllactose Degradation, Front. Microbiol., 2017, 8, 95 Search PubMed
.
| This journal is © The Royal Society of Chemistry 2026 |