 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
From bitter to better: optimizing polyphenol extraction from yellow pea (Pisum sativum L.) using natural deep eutectic solvents (NADES) for improved plant protein products
Amy McMackin ,
Gloria Müller,
Joseph Dumpler
,
Gloria Müller,
Joseph Dumpler * and
Alexander Mathys
* and
Alexander Mathys
Sustainable Food Processing Laboratory, Department of Health Sciences and Technology, ETH Zurich, Schmelzbergstrasse 9, 8092 Zürich, Switzerland. E-mail: joseph.dumpler@hest.ethz.ch
First published on 25th March 2026
Abstract
World food systems are shifting toward more sustainable and ethical production, driving demand for plant-based proteins. Yet, consumer acceptance remains limited due to undesirable flavor, color, and texture. Natural deep eutectic solvents (NADES) can be used to sustainably remove bitter and astringent phytochemicals from plants, yielding whitened, odorless, and protein-rich materials. This study examines the selective extraction of polyphenols from yellow split peas (Pisum sativum L.) using two food-grade NADES (betaine or choline chloride with 1,2-propanediol). Kinetic modeling is derived from analytical data from varying pea particle size (fractions of 0.25–0.5, 0.5–1, 1–2, and 2–3.15 mm), temperatures (35, 45, 55, 65, 75 °C) and NADES hydration levels (0, 10, 20% water), revealing how these parameters influence extraction behavior. Comparisons with 1,2-propanediol (0, 10, 20% water) highlight the key role of eutectic solvents' components in determining kinetics and yields. The studied NADES achieve selective polyphenol extraction (around 550 mg kg−1) while retaining proteins (around 8 g kg−1, co-extraction), outperforming conventional ethanol or water as solvents. Increasing NADES hydration enhanced diffusion and extraction rates, demonstrating water's importance as a ternary component in these systems. This work underscores NADES’ potential as an efficient, scalable, and sustainable strategy for purifying plant proteins in a promising step towards improving these products' sensory quality and consumer appeal.
Sustainability spotlightThis study introduces a novel natural deep eutectic solvent-based approach for purifying plant proteins for sustainable food processing. Derived from food-grade, biodegradable components, NADES offer a low-impact alternative to conventional organic solvents, directly supporting the Sustainable Development Goal on Responsible Consumption and Production (SDG 12). This research demonstrates how differences in solubilization potential between proteins and polyphenols from yellow peas can be harnessed to obtain protein materials for foods aligned with consumer preferences. By minimizing chemical waste and enabling efficient, scalable protein recovery from plants, this approach promotes more sustainable, circular food systems. Moreover, the direct application of this technology to plant-based foods promotes Good Health and Well-Being (SDG 3). |
1 Introduction
Consumers are increasingly interested in plant-based proteins as substitutes for animal products, motivated by environmental concerns, animal welfare, health benefits, and the need to secure sustainable food sources for a growing global population under changing climate conditions.1,2 As a result, there is a growing need to diversify and enhance the quality of plant-based foods and nutraceuticals. Many plant-based protein-rich products still struggle with issues related to functionality and sensory quality, such as unpleasant textures and bitter or astringent flavors.3,4 While these attributes may appeal to vegans and vegetarians, they tend to deter flexitarian and omnivorous consumers: groups that account for the majority of global consumers.4–6Innovative plant-protein processing methods that enhance functionality, nutritional value, and taste are key to creating a more sustainable food system aligned with consumer preferences. Expanding research into alternative plant protein sources is also essential for enhancing food security and biodiversity. Currently, most plant proteins are derived from a very limited number of crops, namely soybeans and wheat, although many other sources contain significant, nutritious protein levels (i.e., peas, lupines, rapeseed).7,8 Developing versatile and efficient methods to isolate functional, neutral-tasting proteins from these sources is essential to developing the plant-based protein market.
The yellow field pea (Pisum sativum L.) is known for its high protein content (20–30%), availability, affordability, and adaptability to various climatic conditions.9 Pea proteins are predominantly albumins (lectin, pea albumin 1, pea albumin 2) and globulins (legumin, vicilin, convicilin), represent a low risk of allergenic response, and boast a digestible indispensable amino acid score (DIAAS) of 1.0.9–12 Despite this, pea protein use in dairy, meat, egg, and seafood analogues may be restricted by weak functional properties or met with poor consumer acceptance due to undesirable flavors and colors from co-extracted pea polyphenols.4,10,13 Pea protein purification presents a solution to these limitations.
The most widely used methods for plant protein extraction and purification rely on aqueous alkali or salt-based systems. Isoelectric precipitation after alkaline extraction isolates proteins by solubilizing them at alkaline pH and then precipitating them at their isoelectric point, where they have minimal solubility.14,15 Alternatively, salt-based extraction takes advantage of ionic interactions to solubilize proteins at increased ion concentrations, then encourage their aggregation by reducing ion concentration through subsequent ultrafiltration or dialysis.16,17 These methods often fail to remove antinutritive compounds like polyphenols, which interfere with protein functionality and impart unwanted flavors and taste on protein isolates.3,4,18–20 Indeed, alkaline extraction–isoelectric precipitation may enrich polyphenol concentrations in protein extracts, as polyphenol solubilization also increases under alkaline conditions.21 Organic solvents, often associated with toxicity and hazards, are often used for separating phytochemicals from proteins. However, they may denature proteins, reducing yield and compromising functionality.19,22–24 Alternatively, using supercritical carbon dioxide for phytochemicals extraction from protein-rich plants has been shown to reduce off-aroma compounds in protein isolates but may increase perceived bitterness.25 Elsewhere, exploring enzymatic or mechanical techniques such as ultrasonication reduces the phytochemical content in refined proteins but the likelihood of protein denaturation remains.14,26–28
In contrast, deep eutectic solvents (DES) provide a sustainable approach for protein refining. DES are defined as mixtures of Brønsted or Lewis acids and bases, where specific molar ratios lead to charge delocalization through supramolecular hydrogen bonding networks. This results in the mixture's melting point being lower than that of its individual components.29 Typically, DES are formed by complexing a hydrogen bond acceptor (HBA) (i.e., quaternary ammonium salt), with a hydrogen bond donor (HBD) (i.e., metal salts, sugars, carboxylic acids, amino acids).29,30 Natural deep eutectic solvents (NADES) are a subclass of DES prepared from natural compounds, and are believed to be the missing link in stabilizing enzymatic activity, catalyzing various metabolic reactions, and solubilizing intermediate polarity compounds in plant cells.31 If DES are considered the next generation of ionic liquids (ILs), offering similar properties with improved sustainability and cost-effectiveness,32,33 then NADES can be seen as the next step in sustainability for novel solvents.
NADES' intrinsic non-toxicity, stabilizing nature, and exceptional selectivity predict their extension to plant-based food processing applications.31,32,34 Recent research has validated such applications, showing that NADES can stabilize proteins and enzymes beyond the cellular environment, protecting them from denaturation and facilitating selective molecular interactions.32,34–38 Lores and co-workers pioneered this food science research in 2017 by studying the solubility of wheat gluten in NADES.39 Since then, NADES have been employed to recover small amounts of high-purity proteins from oilseed press cakes, brewery residues, pomegranates, and fava beans.40–44 Elsewhere, research demonstrates NADES' ability to extract specific bioactive compounds (i.e., flavonoids, tannins, terpenoids) from a variety of plant matrices. These include traditional medicinal plants, rattan palms, agrifood by-products, olive oil and related processing wastes, grape skins, grapevine canes, blueberries, and microalgae.33,45–56 When used in protein or phytochemical extraction, NADES compatibility with techniques such as ultrasound- or microwave-assisted extraction can further improve yields or reduce processing times.39,49,50,55–57 However, even when coupled with mechanical techniques, protein extraction using NADES typically results in low yields, as these solvents are not particularly effective at solubilizing proteins. On the other hand, NADES excel at selectively extracting intermediate polarity polyphenols and other phytochemicals from plant materials.58–60 When operating within these solubility differences, NADES pre-processing of plant-based materials could become a strategy to mitigate off-flavors and colors in plant-based, protein-rich foods. This frames NADES as powerful tools to control food properties through selective extraction processes, beyond their participation in Maillard reactions, suspected synergism with extracted compounds to enhance bioactive properties, or inherent distinct characteristics.36,61–63
This work consequently presents the first integrated approach employing natural deep eutectic solvents (NADES) for the extraction of polyphenols from yellow peas, with a particular focus on elucidating the extraction kinetics under various experimental conditions. Understanding the rates at which NADES can extract polyphenols versus proteins from yellow peas is key to optimizing both the efficiency and selectivity of the process, as well as eventually extending the approach to other legumes.
2 Materials and methods
2.1 Materials
Dehulled, dried yellow peas were purchased from Biofarm Genossenschaft (Kleindietwil, Switzerland) in July 2023. Anhydrous betaine (>99%) and crystalline choline chloride (>98%) were obtained from Healthy Husbandry Sci-tech Co., Ltd (Zhejiang, China), while 1,2-propanediol (>99.5%, for synthesis) was sourced from Carl Roth GmbH+ Co. (Karlsruhe, Germany). Quercetin (>99.5%, HPLC grade, solid), anhydrous sodium carbonate (≥99.5%, powder, ACS reagent), sodium bicarbonate, sodium hydroxide, and absolute ethanol (>99.8%, for analysis) were obtained from Sigma-Aldrich Chemie GmbH (Steinheim, Germany). Bovine serum albumin (2.0 mg mL−1), as well as reagents A (bicinchoninic acid in 2% sodium carbonate solution) and B (4% cupric sulfate solution in 0.4 N sodium hydroxide) for the Pierce™ BCA Protein Assay were obtained from ThermoScientific (Rockford, USA). Stearin organic wax granules (melting point 66 °C) were obtained from Glorex AG (Füllinsdorf, Switzerland) and Safranin Coloring Fluid (PB.5295) was from Euromex Microscopen BV (Duiven, the Netherlands). Water used throughout this work was deionized.2.2 NADES selection & preparation
A pre-screening of 16 different NADES according to their respective viscosities identified the most promising hydrogen bond donor (HBD) as 1,2-propanediol of the 16 pre-screened options, NADES prepared with 1,2-propanediol presented the lowest viscosities, and viscosity is recognized as a critical factor in solid–liquid extraction as high viscosities limit the mass transfer rate and overall process efficiency.64 It is also believed that starch swelling in water would improve overall extraction rates,65 as well as the general hydration of the cell wall (thereby opening cellular pores and capillaries),66 but that this would also lead to greater protein co-extraction due greater solubility potential in water or salt solutions.67 These ideas are underlined in previous work, where a maximum of 20% water addition to NADES provided the best compromise between maximum polyphenol extraction with minimal protein co-extraction.60 Consequently, this work only studied NADES or 1,2-propanediol with 10 or 20% water addition. Further rationale behind the NADES pre-screening and selection process is described in previous work.60Specifically, betaine and 1,2-propanediol at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 4 molar ratio (BetaPro_14) and choline chloride and 1,2-propanediol at a 1
4 molar ratio (BetaPro_14) and choline chloride and 1,2-propanediol at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 molar ratio (ChChlPro_13) were selected for this work due to their relatively low viscosities compared to other NADES and neutral pH. To prepare these NADES, the common “heating” method was used.64,68,69 Briefly, the required quantities of betaine or choline chloride and 1,2-propanediol were mixed in a flask, then slowly heated to 80 °C. Under constant stirring at 300 rpm, the mixture was held at 80 °C for 10 minutes, ensuring the establishment of the NADES' particular hydrogen bond networks. Finally, NADES was cooled, transferred to closed containers, and stored at room temperature.
3 molar ratio (ChChlPro_13) were selected for this work due to their relatively low viscosities compared to other NADES and neutral pH. To prepare these NADES, the common “heating” method was used.64,68,69 Briefly, the required quantities of betaine or choline chloride and 1,2-propanediol were mixed in a flask, then slowly heated to 80 °C. Under constant stirring at 300 rpm, the mixture was held at 80 °C for 10 minutes, ensuring the establishment of the NADES' particular hydrogen bond networks. Finally, NADES was cooled, transferred to closed containers, and stored at room temperature.
2.3 Yellow pea grinding and sieving
Solid–liquid extraction experiments involved four different pea particle size fractions derived from ground whole peas. To this end, whole, dehulled peas were milled at room temperature using a commercial grinding mill (Bosch MUZ5GM3) attached to a stand mixer (Bosch MUM 86) at fineness level 3 (coarse) and milled a second time at fineness level 1 (fine). Particle size fractionation (AS 200-digit cA, Retsch, Germany) was performed at an amplitude of 1.7 mm g−1 for 15 min using test sieves. Size fractions, pictured in Fig. 1, were obtained as follows: small (0.25 mm ≤ x < 0.5 mm), medium (0.5 mm ≤ x < 1 mm), large (1 mm ≤ x < 2 mm), and extra-large (2 mm ≤ x < 3.15 mm). Particles smaller than 0.25 mm or larger than 3.15 mm were discarded. Milled pea particle fractions were stored in a dark environment in ambient conditions until further use.2.4 Compositional analysis & reference solid–liquid extraction experiments
Compositional analysis of ground yellow peas (0.25–2 mm particle size) was performed by Amt für Verbraucherschutz (AVS), Steinhausen, Switzerland. Protein was analyzed in triplicate in digested peas using the Kjeldahl method (N-conversion factor of 6.25).70Peas were subjected to exhaustive polyphenol extraction, using a method reported elsewhere.71 Briefly, peas were ground to a fine powder, dried at 50 °C in an oven overnight, and stored in a desiccator until use. Then, 5 g were filled into cellulose extraction thimbles, and loaded into a Soxhlet extraction setup. The samples were extracted with a refluxing 70% aqueous methanol solution at 70 °C for 3 hours. Soxhlet extraction was performed in triplicate.
Moreover, solid–liquid extraction experiments were performed with 60% aqueous ethanol and deionized water. While 60% aqueous ethanol is recognized as an effective solvent for intermediate polarity compounds such as plant polyphenols, deionized water provides a reference for protein extraction, mainly due to the water-soluble albumin fraction of yellow peas.10,67 These experiments provide an opportunity to compare the extractive behaviors of the NADES and 1,2-propanediol against that of conventional solvents. These batch extraction experiments were also performed in triplicate.
2.5 Solid–liquid batch extraction
Yellow pea particles and solvent were mixed at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 (w/w) solid-to-liquid ratio. Samples were placed in a water bath atop a multi-position magnetic stirring plate set to 350 rpm. The water bath was maintained at a constant temperature using a separate recirculating heating bath. Solid–liquid extractions were performed in triplicate.
10 (w/w) solid-to-liquid ratio. Samples were placed in a water bath atop a multi-position magnetic stirring plate set to 350 rpm. The water bath was maintained at a constant temperature using a separate recirculating heating bath. Solid–liquid extractions were performed in triplicate.
Sampling was performed at defined times ranging from 0.16 to a maximum of 96 hours. Samples of the suspensions were taken in a manner ensuring that the 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 10 solid-to-liquid ratio remained constant throughout the experiment. The liquid phase was preserved at −20 °C until analysis, while extracted peas were discarded.
10 solid-to-liquid ratio remained constant throughout the experiment. The liquid phase was preserved at −20 °C until analysis, while extracted peas were discarded.
Solid–liquid extraction experiments involved five variables: solvent type, solvent hydration, yellow pea particle size, extraction temperature, and extraction duration (time). For a given duration, a single variable was changed systematically in each trial to assess its influence on polyphenol and protein extraction from yellow peas. Besides evaluating the extraction efficiency of the two pre-selected NADES, this work also looks at the extraction efficiency of the HBD component of the NADES (1,2-propanediol) alone as a negative control. Quantities of water are added to these solvents, increasing solvent polarity and reducing viscosity. In NADES, up to 40% water can be added as a ternary compound, without disrupting the solvent's unique supramolecular structure.64,72 Table 1 summarizes the experimental parameters.
| Experimental factor | Evaluated conditions |
|---|---|
| Solvent type | BetaPro_14 |
| ChChlPro_13 | |
| 1,2-Propanediol | |
| Solvent hydration (%) | 0, 10, 20 |
| Particle size fraction (mm) | Small (0.25 ≤ x < 0.5) |
| Medium (0.5 ≤ x < 1) | |
| Large (1 ≤ x < 2) | |
| Extra-large (2 ≤ x < 3.15) | |
| Temperature (°C) | 35, 45, 55, 65, 75 |
| Duration (h) | Variable, 0.16 to 96 |
2.6 Bicinchoninic acid (BCA) assay for polyphenol and protein quantification
Centrifuging the liquid extracts from solid–liquid batch extraction and reference solid–liquid extraction experiments (9 min at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300×g, ambient temperature) resulted in a pellet of insoluble plant material (starch and fiber) and a supernatant containing the solvent itself and the analytes (polyphenols and proteins). Since both polyphenols and proteins can reduce Cu2+ and produce a colored compound with BCA,73 a sample preparation procedure that quantitatively separates the two was developed and verified, as described elsewhere.60 Briefly, the supernatant was combined with ethanol at a 1
300×g, ambient temperature) resulted in a pellet of insoluble plant material (starch and fiber) and a supernatant containing the solvent itself and the analytes (polyphenols and proteins). Since both polyphenols and proteins can reduce Cu2+ and produce a colored compound with BCA,73 a sample preparation procedure that quantitatively separates the two was developed and verified, as described elsewhere.60 Briefly, the supernatant was combined with ethanol at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 9 ratio and vortexed for 5 s. These tubes were stored overnight (>14 h) at −21 °C to encourage protein precipitation. Next, the samples were centrifuged for 7 min at 12
9 ratio and vortexed for 5 s. These tubes were stored overnight (>14 h) at −21 °C to encourage protein precipitation. Next, the samples were centrifuged for 7 min at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300×g. Then, 25 µL aliquots of supernatant and 200 µL of BCA working reagent were pipetted into a 96-well plate in triplicate. The working reagent (WR) was prepared immediately prior to plating by mixing a 50
300×g. Then, 25 µL aliquots of supernatant and 200 µL of BCA working reagent were pipetted into a 96-well plate in triplicate. The working reagent (WR) was prepared immediately prior to plating by mixing a 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio of reagent A with reagent B (Pierce™ BCA Protein Assay). Then, samples were incubated for 30 min at 37 °C, and their absorbance at 562 nm was recorded using a microplate reader (TECAN Spark, TECAN, Austria).
1 ratio of reagent A with reagent B (Pierce™ BCA Protein Assay). Then, samples were incubated for 30 min at 37 °C, and their absorbance at 562 nm was recorded using a microplate reader (TECAN Spark, TECAN, Austria).
To analyze polyphenols in samples: absorbance data was related to the polyphenol content in the samples through a quercetin standard series (2, 5, 10, 25, 50, and 75 mg per L quercetin). The average linear regression for the quercetin standards across all plates (n = 76) delivered an R2 of 0.9999. The corresponding eqn (1) relates absorption to total polyphenol content (TPC):
| cTPC,QE = (x − x0) × m × 10 | (1) |
Then, to analyze protein content in samples: the excess ethanol-based supernatant used in TPC quantification was discarded, such that 100 µL liquid and protein pellet remained. The protein pellet was resolubilized by adding 400 µL of an aqueous solution of 2% Na2CO3 and 0.95% NaHCO3 (pH 11.25) and vortexed. An additional centrifuging step for 7 min at 12![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 300×g was applied to settle out any remaining fibrous components. Similarly to TPC analysis, 25 µL aliquots of protein-containing supernatant were mixed with 200 µL of BCA reaction working reagent into a 96-well plate, incubated for 30 min at 37 °C, and analyzed for light absorbance at 562 nm.
300×g was applied to settle out any remaining fibrous components. Similarly to TPC analysis, 25 µL aliquots of protein-containing supernatant were mixed with 200 µL of BCA reaction working reagent into a 96-well plate, incubated for 30 min at 37 °C, and analyzed for light absorbance at 562 nm.
Due to residual polyphenols in the protein phase, protein concentration needed to be calculated in three steps. First, absorbance data was related to apparent protein content with a bovine serum albumin (BSA) standard series (25, 125, 250, 500, 750, 1000, 1500, and 2000 mg per L BSA), with the resulting average second-order polynomial regression across all plates (n = 76, R2 of 0.9962), using eqn (2):
| cPC,apparent = a × (x − x0)2 + b × (x − x0) | (2) |
| cTPC,residual = a × (x − x0)2 + b × (x − x0) | (3) |
Finally, the actual protein content (cPC,BSAE) was calculated using eqn (4), where both 5−1 and 5 represent dilution factors used in sample preparation, as described above:
 | (4) |
Due to the fixed solid-to-liquid mixing ratio used in the extraction experiments, the concentration of TPC and proteins extracted from the peas (provided in mg L−1 solvent through the BCA test) can be converted to mg kg−1 peas by a factor of 11 L kg−1.
2.7 Color analysis of liquid extracts
The major polyphenols in yellow peas, kaempferol and quercetin, are associated with a yellowish hue.74 Throughout the solid–liquid extraction process, these and other yellow-colored polyphenols migrate from the peas to the solvent, marked by a noticeable yellowing of the solvent and whitening of the peas. This process could be quantified through color analysis, adding a numerical value to validate visual observations and associating colorimetric changes to TPC as quantified by BCA.Firstly, a Chroma Meter CR-410 colorimeter (Konica Minolta Inc., Japan) was calibrated against a solid, pure white surface. Liquid extracts from the batch extractions of all four particle size fractions were prepared at 55 °C using water, 60% ethanol, BetaPro_14 and ChChlPro_13 NADES, and 1,2-propanediol. No additional water was added to the NADES or 1,2-propanediol. Samples were taken after 1, 3, and 5 hours, and analyzed (n = 3).
2.8 Kinetic modeling and statistical analysis
The essence of the solid–liquid batch extraction process is the mass transfer of solutes (polyphenols and protein) from their intracellular diffusion to the solvent. This unstable mass transfer process is defined by three main steps: (1) solvent and solid mixing, solvent penetration of the solids, and solute dissolution; (2) internal diffusion of dissolved solutes diffuse to the solids' surfaces; (3) external diffusion of solutes to the bulk solution. Kinetic modeling based on experimental results defines the kinetically limiting factor for this process, relying on several assumptions: spherical and constant solid particle dimensions, constant solid-to-liquid ratios throughout sampling, uniform solute distribution throughout the solid particles, chemical inertia between solvent and solutes, constant temperatures, uniform distribution of solid particles within the liquid, and negligible mass transfer resistance of the solutes on the solids' surfaces.75 With these assumptions, eqn (5) describes a general differential equation, derived from Lagergren's 1898 empirical kinetic equation for solute behavior in solid–liquid systems, and is used to model extraction kinetics:76
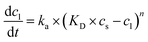 | (5) |
Integrating eqn (5) for a reaction order of 1 provides eqn (6):
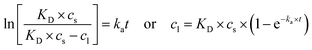 | (6) |
Alternatively, integrating eqn (5) for a reaction order of 2 provides eqn (7), where the apparent rate constant is denoted as Ka (mg L−1 h−1), distinct from the rate constant for the first-order model.77
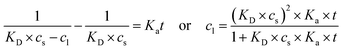 | (7) |
The Bayesian Information Criterion (BIC), chosen for its capacity to balance model complexity with goodness-of-fit, indicated which of the two models was most appropriate to determine the kinetic parameters describing specific extraction experiments (first- vs. second-order functions). All analytical data was processed and analyzed using OriginPro 2025 software (OriginLab Corporation, Northampton, MA, USA).
3 Results and discussion
3.1 Compositional analysis & reference solid–liquid extraction experiments
The total protein content of the peas used in this study was determined according to the Kjeldahl method, revealing a concentration of 181 g per kg protein (nitrogen conversion factor 6.25). The protein content for dried peas aligns with values reported in the literature,10 and is close to the average value reported by the manufacturer of 213 g per kg protein. For reference, protein extraction was targeted with reference water-based extraction experiments.Peas were subjected to Soxhlet exhaustive extractions to determine baseline TPC. Analysis of the methanol-based extracts using the BCA method, revealed a polyphenol yield of 550 mg per kg QE. This value corresponds with TPC values reported in the literature, ranging from 126 to 1286 mg per kg TPC, reported in gallic acid equivalents.78 Crucially, peas of a more intense or darker color are known to contain higher amounts of TPC.78 For batch extraction experiments involving 60% ethanol, an approximate extraction of 473 mg per kg TPC, QE was recorded after extraction at 55 °C, with a 2–3.15 mm particle size, over 96 hours. It is to be expected that a Soxhlet extraction approach yields higher quantities of solute compared to maceration-based batch extraction.79
3.2 Color analysis of liquid extracts
Visual observations of the extraction process indicated the general trend of the solvent's discoloration from colorless to greenish-yellow with prolonged extraction times, as seen in Fig. 2. As the major polyphenols in yellow peas (i.e., kaempferol, quercetin) are associated with a yellowish hue,74 this discoloration is used as a preliminary visual indication of extraction efficiency. Correlating with solvent discoloration, pea particles became noticeable paler compared to their original color. In this sense, measuring solvent discoloration as a proxy of pea whitening is important in terms of overall process efficiency. Consumer studies report that product whiteness is associated with favorable acceptance of protein concentrates and isolates. Hence, color analysis to characterize food ingredients is reported in similar work.80–82General discoloration of the solvents is shown in Fig. 2. Water and 60% ethanol led to strong discoloration of the solvent and whitening of the pea particles. This indicates more exhaustive polyphenol extraction, as validated through BCA analysis of the samples. In contrast, the NADES and pure 1,2-propanediol showed weaker color transfer, which is equally confirmed through quantitative sample analysis. Adding water to the NADES and 1,2-propanediol solvents improved their degree of discoloration. These trends are confirmed quantitatively through the BCA analysis of the extracts and kinetic modeling of the extraction process (Fig. 3 and 4). Crucially, non-hydrated NADES (0% water addition) has a limited capacity to extract polyphenols from peas, seen both through qualitative, colorimetric observations (Fig. 2) and quantitative analysis (Fig. 3).
Water extracts exhibited the highest turbidity, indicating protein solubilization, particularly of water-soluble albumins.67 Although this confirmed water's strong extractive power, it was undesirable in regard to the goal of maximizing polyphenol extraction and minimizing protein co-extraction. Turbidity was therefore a quick qualitative proxy for protein extraction, complementing quantitative methods such as BCA analysis of the extracts.
Regardless of solvent, extract color intensified (yellow, green, or brown tones) with longer extraction times or smaller particles, aligned with the literature.83 Water produced the darkest brown extracts, rich in both polyphenols and proteins, whereas ethanol extracts tended towards greenish hues. Non-hydrated NADES caused the least discoloration, consistent with its lower extraction yield described below. The pattern paralleled known extraction kinetics' associations to color analysis of solvents, such as in coffee brewing, where smaller particle sizes accelerate solute release and lead to a more intensely-colored solution.84 Full color analysis data can be found in the SI.
Importantly, color analysis of liquid extracts quantified discoloration via 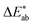 , but this only reflected colored phenolic compound extraction. Some pea polyphenols (i.e., gallic and vanillic acids) are colorless and undetectable by optical methods.78 Moreover, solvatochromism can complicate direct interpretation of colorimetric data.85 This underlies the need for kinetic modeling based on robust chemometric data, such as the BCA assay, as described below.
, but this only reflected colored phenolic compound extraction. Some pea polyphenols (i.e., gallic and vanillic acids) are colorless and undetectable by optical methods.78 Moreover, solvatochromism can complicate direct interpretation of colorimetric data.85 This underlies the need for kinetic modeling based on robust chemometric data, such as the BCA assay, as described below.
3.3 Kinetic modeling
Although an abundance of literature suggests NADES' promise in extracting polyphenols from intact plant tissues, comments on extraction rates as a function of experimental parameters remain limited to select Design of Experiments (DOE) analyses.44,59 Furthermore, this approach considers protein co-extraction, revealing a pathway whereby NADES may be used as a crucial pre-processing step for polyphenols from foods, prior to protein recovery according to conventional methods. Kinetic modeling also allows for the consideration of water content as a ternary component of NADES systems. Previous work applied this methodology to rapeseed as a model oilseed,60 while this work examines peas as a model legume. Compositional differences between oilseeds and legumes may influence their behaviors in NADES-based solid–liquid extraction experiments.Fig. 3 and 4 clearly reveal the impact of adding water as a ternary component to NADES on extraction rates and efficiency. With increasing water content, more TPC and protein are extracted, although extraction rates remain similar. This impact is extended to 1,2-propanediol alone, although 10% water addition is revealed to be more useful for TPC extraction than 20%. Adding 10 or 20% water to NADES or 1,2-propanediol achieves similar TPC extraction efficiencies and rates as demonstrated by conventional solvents (water and 60% ethanol), without the drawbacks of high protein co-extraction (water) or protein denaturation (ethanol).86 These findings are particularly relevant for larger pea particle size fractions. Specifically, 10% water content in 1,2-propanediol or 20% water content in ChChlPro_13 NADES can extract around 550 mg per kg TPC after 6.5 hours, achieving the same exhaustive TPC extraction as Soxhlet extraction or batch extraction in water. These conditions represent between 8 and 13 g per kg protein co-extraction, while water co-extracts around 33 g per kg protein.
Extraction experiments involving different particle size fractions of the ground peas reveals how smaller pea particles generally favor faster extraction rates (Fig. 5 and 6). This is especially relevant for protein extraction, where decreasing particle size is strictly accompanied by a faster extraction rate. Although overall yields remain similar for TPC and protein, independent of particle size, one notable discrepancy is observed for water. Here (Fig. 6e), smaller particle size fractions correlate with higher protein extraction as a function of increasing cellular rupture from more intensive grinding. This is further validated upon experimental analysis of a commercial pea protein concentrate, presented in the SI. Consequently, extraction should find compromise between desirable fast rates and larger particle size, especially when using solvents with increasing water content, to avoid protein co-extraction.
Although extraction experiments were carried out at 35, 45, 55, 65, and 75 °C, the data for 35 °C is omitted from Fig. 7 and 8. For all solvents and sampling times, measured concentrations of TPC and protein were outside of the limit of detection of the methods used in this work. Consequently, it is possible to conclude that 35 °C does not encourage extraction rates that can be reliably captured within the experimental boundaries, although with sufficient time similar overall yields could theoretically be expected. Overall, regardless of solvent type, higher temperatures encourage higher extraction rates which is aligned with the Arrhenius principle. Around 75 °C, starch gelation rates increase. At higher temperatures, extraction kinetics may change due to starch gelation-induced structural modifications of the pea particles.87 It is possible that the onset of starch gelation also explains the discrepancies for TPC extraction in water (Fig. 7e).
Fig. 8e describing protein extraction by water does not feature the first- and second-order models, as the data cannot be described by these functions. Instead, there is a clear competing reaction occurring, whereby enzymes may be hydrolyzing proteins as they are extracted by the water.88 This explains the initial increase of proteins in solution over the first few minutes of the extraction experiment, followed by a decline as the rate of protein hydrolysis exceeds additional extraction.
To have both, a fast and efficient extraction, high rate constants (ka) and high distribution coefficients (KD × cs) are necessary. As this work seeks to maximize polyphenol extraction and minimize protein co-extraction, the ideal operating conditions are defined by the point at which the values of ka and KD are most different between polyphenols and proteins. For 1,2-propanediol and both NADES, these differences are most evident when working with larger particle size fractions or when adding 10–20% water to the solvent. Rate constants and distribution coefficients from the second-order models plotted on Fig. 3 and 4 are summarized in Fig. 9. The BIC analysis indicated that the second-order model provides a more accurate description of TPC and protein extraction from peas (Table 2 and 3). All other plotting rate constants, distribution coefficients, and results of the BIC test can be found in the SI.
| Solvent | % Water | Model | ||
|---|---|---|---|---|
| First order | Second order | Difference | ||
| a A lower BIC value indicates the model that is more likely to be correct. | ||||
| 1,2-Propanediol | 0 | 28.4 | 19.4a | 9.0 |
| 10 | 40.5 | 34.5a | 6.0 | |
| 20 | 24.0 | 17.5a | 6.5 | |
| BetaPro_14 | 0 | 33.7 | 30.7a | 3.0 |
| 10 | 22.4 | 17.0a | 5.4 | |
| 20 | 23.0a | 28.5 | 5.4 | |
| ChChlPro_13 | 0 | 35.1 | 29.3a | 5.8 |
| 10 | 30.1 | 22.9a | 7.2 | |
| 20 | 27.5 | 24.1a | 3.4 | |
| Ethanol | 40 | 16.6 | 13.5a | 3.1 |
| Water | 100 | 44.9 | 42.1a | 2.8 |
| Solvent | % Water | Model | ||
|---|---|---|---|---|
| First order | Second order | Difference | ||
| a A lower BIC value indicates the model that is more likely to be correct.b An absolute BIC difference greater than 10 indicates the model that is decisively correct. | ||||
| 1,2-Propanediol | 0 | 56.3 | 51.1a | 5.1 |
| 10 | 72.9 | 60.6a | 12.3b | |
| 20 | 84.5 | 75.9a | 8.7 | |
| BetaPro_14 | 0 | 66.8 | 65.6a | 1.2 |
| 10 | 84.5 | 72.7a | 11.8b | |
| 20 | 95.2 | 88.4a | 6.8 | |
| ChChlPro_13 | 0 | 85.6 | 84.3a | 1.3 |
| 10 | 82.7 | 79.3a | 3.4 | |
| 20 | 87.0 | 84.6a | 2.4 | |
| Ethanol | 40 | 115.8a | 116.2 | 0.4 |
| Water | 100 | 77.8a | 85.6 | 7.8 |
4 Conclusions
In this study, we mapped out the conditions whereby NADES refining of yellow pea particles can ensure maximal polyphenol extraction and minimal protein co-extraction to minimize protein losses. Moreover, identifying the specific extraction kinetics under different processing conditions provides valuable insights for eventual process up-scaling to meet growing consumer demand for high-quality, plant-based proteins from yellow peas with improved sensory properties. This work achieved both objectives, paving the way for yellow pea's proliferation in the alternative protein market. It also facilitates the method's extension to other legumes—such as soybeans, fava beans, and lupines—thereby enhancing the variety and appeal of plant-based food analogues.Author contributions
Amy McMackin: methodology, validation, formal analysis, investigation, writing – original draft, visualization. Gloria Müller: formal analysis, investigation, writing – original draft, visualization. Joseph Dumpler: conceptualization, methodology, validation, resources, writing – review & editing, supervision, funding acquisition. Alexander Mathys: resources, writing – review & editing, supervision, funding acquisition, project administration. All authors have approved the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
Research data is available on the ETH Research Collection: https://doi.org/10.3929/ethz-c-000790062.Supplementary information (SI): additional experimental details, materials, and methods. See DOI: https://doi.org/10.1039/d6fb00009f.
Acknowledgements
Many thanks are extended to Daniel Kiechl and Peter Bigler of the ETH university workshop for the custom devices they made to support this project. We would also like to acknowledge Christoph Jans and Urs Aschwanden of the Amt für Verbraucherschutz (AVS) in Steinhausen, Switzerland for analysis of the peas' chemical composition. And thank you, Lenja Lemcke and Dylan Zimmermann, for early experimental contributions to the work, completed as bachelor's theses. The graphical abstract was created in BioRender (https://BioRender.com/xaysli7). This project was supported by Bühler AG, Nestec SA, Givaudan International AG via the ETH Zurich Foundation in the framework of the Future Food Initiative 2023 (ETH Zurich Foundation project number: 2023-HS-227).References
- S. R. Hertzler, J. C. Lieblein-Boff, M. Weiler and C. Allgeier, Nutrients, 2020, 12, 1–27 CrossRef PubMed.
- A. Green, C. Blattmann, C. Chen and A. Mathys, Trends Food Sci. Technol., 2022, 124, 250–258 CrossRef CAS.
- C. Nuvoli, L. Fillion, C. Lacoste Gregorutti and D. Labbe, Physiol. Behav., 2023, 262, 114092 CrossRef CAS PubMed.
- V. Caputo, G. Sogari and E. J. Van Loo, Appl. Econ. Perspect. Policy, 2023, 45, 86–105 CrossRef.
- C. Giezenaar, A. J. R. Godfrey, M. Foster and J. Hort, Food Qual. Preference, 2024, 105070 CrossRef.
- D. Hagmann, M. Siegrist and C. Hartmann, Public Health Nutr., 2019, 22, 2448–2459 CrossRef PubMed.
- A. M. De Ron, F. Sparvoli, J. J. Pueyo and D. Bazile, Front. Plant Sci., 2017, 8, 105 Search PubMed.
- M. M. Lucas, F. L. Stoddard, P. Annicchiarico, J. Frías, C. Martínez-Villaluenga, D. Sussmann, M. Duranti, A. Seger, P. M. Zander and J. J. Pueyo, Front. Plant Sci., 2015, 6, 705 Search PubMed.
- C. Tanger, J. Engel and U. Kulozik, Food Hydrocolloids, 2020, 107, 105949 CrossRef CAS.
- Z. X. Lu, J. F. He, Y. C. Zhang and D. J. Bing, Crit. Rev. Food Sci. Nutr., 2020, 60, 2593–2605 CrossRef CAS PubMed.
- M. G. Nosworthy, G. Medina, Z.-H. Lu and J. D. House, Foods, 2023, 12, 2816 CrossRef CAS.
- F. M. Guillin, C. Gaudichon, L. Guérin-Deremaux, C. Lefranc-Millot, G. Airinei, N. Khodorova, R. Benamouzig, P.-H. Pomport, J. Martin and J. Calvez, Am. J. Clin. Nutr., 2022, 115, 353–363 CrossRef CAS PubMed.
- M. Klost and S. Drusch, Food Hydrocolloids, 2019, 94, 622–630 CrossRef CAS.
- I. Cruz-Solis, C. C. Ibarra-Herrera, M. d. R. Rocha-Pizana and D. Luna-Vital, in Green Protein Processing Technologies from Plants: Novel Extraction and Purification Methods for Product Development, ed. A. J. Hernández-Álvarez, M. Mondor and M. G. Nosworthy, Springer International Publishing, Cham, 2023, ch. 1, pp. 1–29 Search PubMed.
- R. M. Kramer, V. R. Shende, N. Motl, C. N. Pace and J. M. Scholtz, Biophys. J., 2012, 102, 1907–1915 CrossRef CAS PubMed.
- K. Ayan, R. M. Boom and C. V. Nikiforidis, J. Food Eng., 2024, 381, 112188 CrossRef CAS.
- J. Yang and L. M. Sagis, Curr. Opin. Colloid Interface Sci., 2021, 56, 101499 CrossRef CAS.
- P. Wilding, P. J. Lillford and J. M. Regenstein, J. Chem. Technol. Biotechnol., 1984, 34, 182–189 CrossRef.
- S. Vatansever, B. Chen and C. Hall, Sustainable Food Proteins, 2024, 2, 33–57 CrossRef CAS.
- S. H. Tan, R. J. Mailer, C. L. Blanchard and S. O. Agboola, Food Res. Int., 2011, 44, 1075–1082 CrossRef CAS.
- S. Peng, L. Zou, W. Zhou, W. Liu, C. Liu and D. J. McClements, J. Agric. Food Chem., 2019, 67, 7506–7511 CrossRef CAS PubMed.
- S. Feyzi, M. Varidi, F. Zare and M. J. Varidi, Int. J. Biol. Macromol., 2017, 105, 27–35 CrossRef CAS PubMed.
- A. Ihsan, Z. Ahmad, J. Zheng, M. Bilal, H. Muhammad Rizwan Abid and A. Hu, Food Biosci., 2024, 57, 103476 CrossRef CAS.
- L. J. Rubin, L. L. Diosady and C. R. Phillips, US Pat., US4460504A, 1984, https://patentimages.storage.googleapis.com/61/62/7b/129ff887964272/US4460504.pdf Search PubMed.
- S. Vatansever, J. Ohm, S. Simsek and C. Hall, Cereal Chem., 2022, 99, 130–143 CrossRef CAS.
- S. B. Alpiger, C. Solet, T. T. Dang and M. Corredig, Foods, 2024, 2423 CrossRef CAS.
- S. B. Alpiger and M. Corredig, Food Res. Int., 2024, 175, 113736 CrossRef.
- N. Yousefi and S. Abbasi, Food Chem. Adv., 2022, 1, 100090 CrossRef.
- E. L. Smith, A. P. Abbott and K. S. Ryder, Chem. Rev., 2014, 114, 11060–11082 CrossRef CAS PubMed.
- H. Bowen, R. Durrani, A. Delavault, E. Durand, J. Chenyu, L. Yiyang, S. Lili, S. Jian, H. Weiwei and G. Fei, Front. Chem., 2022, 10, 912411 CrossRef PubMed.
- Y. H. Choi, J. van Spronsen, Y. Dai, M. Verberne, F. Hollmann, I. W. Arends, G.-J. Witkamp and R. Verpoorte, Plant Physiol., 2011, 156, 1701–1705 CrossRef CAS PubMed.
- M. M. Cajnko, F. A. Vicente, U. Novak and B. Likozar, Green Chem., 2023, 25, 9045–9062 RSC.
- Q. Cao, J. Li, Y. Xia, W. Li, S. Luo, C. Ma and S. Liu, Molecules, 2019, 24, 113 CrossRef PubMed.
- T. Caldas, N. Demont-Caulet, A. Ghazi and G. Richarme, Microbiology, 1999, 145, 2543–2548 CrossRef CAS.
- N. F. Gajardo-Parra, L. Meneses, A. R. C. Duarte, A. Paiva and C. Held, ACS Sustain. Chem. Eng., 2022, 10, 12873–12881 CrossRef CAS PubMed.
- D. M. Hartl, O. Frank, C. Dawid and T. F. Hofmann, Foods, 2023, 12, 1877 CrossRef CAS PubMed.
- Y. Tian, M. Zhu, T. Hu and C. Liu, Int. J. Biol. Macromol., 2023, 247, 125477 CrossRef CAS PubMed.
- A. Kovács, M. Yusupov, I. Cornet, P. Billen and E. C. Neyts, J. Mol. Liq., 2022, 366, 120180 CrossRef.
- H. Lores, V. Romero, I. Costas, C. Bendicho and I. Lavilla, Talanta, 2017, 162, 453–459 CrossRef CAS PubMed.
- A. Grudniewska, E. M. de Melo, A. Chan, R. Gniłka, F. Boratyński and A. S. Matharu, ACS Sustain. Chem. Eng., 2018, 6, 15791–15800 CrossRef CAS.
- E. Hernández-Corroto, S. Olivares-Galván, M. L. Marina and M. C. García, LWT, 2023, 189, 115470 CrossRef.
- M. Guzmán-Lorite, M. Marina and M. García, Food Res. Int., 2022, 161, 111862 CrossRef PubMed.
- E. Hernández-Corroto, M. Plaza, M. L. Marina and M. C. García, Innovative Food Sci. Emerging Technol., 2020, 60, 102314 CrossRef.
- A. Hewage, O. O. Olatunde, C. Nimalaratne, J. D. House, R. E. Aluko and N. Bandara, Food Hydrocolloids, 2024, 147, 109283 CrossRef CAS.
- C. Bakirtzi, K. Triantafyllidou and D. P. Makris, J. Appl. Res. Med. Aromat. Plants, 2016, 3, 120–127 Search PubMed.
- M. d. l. Á. Fernández, M. Espino, F. J. Gomez and M. F. Silva, Food Chem., 2018, 239, 671–678 CrossRef PubMed.
- E. Rodríguez-Juan, C. Rodríguez-Romero, J. Fernández-Bolaños, M. C. Florido and A. Garcia-Borrego, J. Food Sci. Technol., 2021, 58, 552–561 CrossRef.
- A. García, E. Rodríguez-Juan, G. Rodríguez-Gutiérrez, J. J. Rios and J. Fernández-Bolaños, Food Chem., 2016, 197, 554–561 CrossRef PubMed.
- K. M. Jeong, J. Zhao, Y. Jin, S. R. Heo, S. Y. Han, D. E. Yoo and J. Lee, Arch. Pharmacal Res., 2015, 38, 2143–2152 CrossRef CAS.
- M. Cvjetko Bubalo, N. Ćurko, M. Tomašević, K. Kovačević Ganić and I. Radojčić Redovniković, Food Chem., 2016, 200, 159–166 CrossRef CAS PubMed.
- D. T. da Silva, R. Pauletto, S. d. S. Cavalheiro, V. C. Bochi, E. Rodrigues, J. Weber, C. d. B. da Silva, F. D. P. Morisso, M. T. Barcia and T. Emanuelli, J. Food Compos. Anal., 2020, 89, 103470 CrossRef.
- M. Kiene, M. Zaremba, H. Fellensiek, E. Januschewski, A. Juadjur, G. Jerz and P. Winterhalter, Foods, 2023, 12, 4184 CrossRef CAS PubMed.
- M. Kiene, M. Zaremba, E. Januschewski, A. Juadjur, G. Jerz and P. Winterhalter, Foods, 2024, 13, 324 CrossRef CAS PubMed.
- W. M. A. Wan Mahmood, A. Lorwirachsutee, C. Theodoropoulos and M. Gonzalez-Miquel, ACS Sustain. Chem. Eng., 2019, 7, 5018–5026 CrossRef CAS.
- M. Coscarella, M. Nardi, K. Alipieva, S. Bonacci, M. Popova, A. Procopio, R. Scarpelli and S. Simeonov, Antioxidants, 2023, 13, 62 CrossRef PubMed.
- S. Bonacci, M. L. Di Gioia, P. Costanzo, L. Maiuolo, S. Tallarico and M. Nardi, Antioxidants, 2020, 9, 513 CrossRef CAS PubMed.
- E. Gil-Martín, T. Forbes-Hernández, A. Romero, D. Cianciosi, F. Giampieri and M. Battino, Food Chem., 2022, 378, 131918 CrossRef PubMed.
- N. P. E. Hikmawanti, D. Ramadon, I. Jantan and A. Mun'im, Plants, 2021, 10, 2091 CrossRef CAS PubMed.
- L. Duan, L.-L. Dou, L. Guo, P. Li and E.-H. Liu, ACS Sustain. Chem. Eng., 2016, 4, 2405–2411 CrossRef CAS.
- J. Dumpler, A. McMackin, M. Herren, S. H. Zadeh and A. Mathys, ACS Sustain. Chem. Eng., 2025, 13, 17275–17286 CrossRef CAS.
- H. Wang, T. Liu, T. Zhao, G. Su, M. Zhao and J. Zhang, Food Bioprocess Technol., 2024, 17, 2446–2458 CrossRef CAS.
- K. Radošević, N. Ćurko, V. Gaurina Srček, M. Cvjetko Bubalo, M. Tomašević, K. Kovačević Ganić and I. Radojčić Redovniković, LWT, 2016, 73, 45–51 CrossRef.
- P. Manuela, S. Drakula, G. Cravotto, R. Verpoorte, M. Hruškar, I. Radojčić Redovniković and K. Radošević, Innovative Food Sci. Emerg. Technol., 2020, 66, 102514 CrossRef CAS.
- G. Li, J. Lei, S. Li, Y. Jiang, F. Zhang, C. Song, S. Xiao, S. Fu, J. Zhou, F. Wu and G. Wang, RSC Adv., 2022, 12, 26975–26988 RSC.
- R. Jia, C. Cui, L. Gao, Y. Qin, N. Ji, L. Dai, Y. Wang, L. Xiong, R. Shi and Q. Sun, Carbohydr. Polym., 2023, 321, 121260 CrossRef CAS PubMed.
- D. S. Thompson and A. Islam, Plants, 2021, 10, 1263 CrossRef CAS PubMed.
- T. B. Osborne and C. G. Voorhees, J. Am. Chem. Soc., 1894, 16, 778–785 CrossRef CAS.
- V. I. Castro, R. Craveiro, J. M. Silva, R. L. Reis, A. Paiva and A. R. C. Duarte, Cryobiology, 2018, 83, 15–26 CrossRef CAS PubMed.
- Y. Dai, G.-J. Witkamp, R. Verpoorte and Y. H. Choi, Food Chem., 2015, 187, 14–19 CrossRef CAS PubMed.
- J. Kjeldahl, Z. Anal. Chem., 1883, 22, 366–382 CrossRef.
- L. S. Putra, J. Sukweenadhi, C. Nathania, E. S. Wibowo, G. Buschle-Diller and M. G. Marianti Purwanto, Heliyon, 2022, 8, e08951 CrossRef CAS PubMed.
- N. Mustafa, V. Spelbos, G.-J. Witkamp, R. Verpoorte and Y. Choi, Molecules, 2021, 26, 2645 CrossRef CAS PubMed.
- D. Cestari Marino, L. Zuppardo Lacerda Sabino, J. Armando, A. De Andrade Ruggiero and H. Dorigan Moya, J. Agric. Food Chem., 2009, 57, 11061–11066 CrossRef PubMed.
- F. Guo, H. Xiong, X. Wang, L. Jiang, N. Yu, Z. Hu, Y. Sun and R. Tsao, J. Agric. Food Chem., 2019, 67, 11955–11968 CrossRef CAS PubMed.
- Y. Wang, C. Wang, H. Xue, Y. Jin, M. Yang and F. Leng, Biomass Convers. Biorefin., 2023, 13, 12917–12933 CrossRef CAS PubMed.
- S. Lagergren, K. Sven. Vetenskapsakad. Handl., 1898, 24, 1–39 Search PubMed.
- H. A. Harouna-Oumarou, H. Fauduet, C. Porte and Y.-S. Ho, Chem. Eng. Commun., 2007, 194, 537–552 CrossRef CAS.
- D.-T. Wu, W.-X. Li, J.-J. Wan, Y.-C. Hu, R.-Y. Gan and L. Zou, Foods, 2023, 12, 2527 CrossRef CAS PubMed.
- J. F. Osorio-Tobón, J. Food Sci. Technol., 2020, 57, 4299–4315 CrossRef.
- J. Zhao, M. Xu, J. Baker and S. Kinsey, in Flavour and Consumer Perception of Food Proteins, ed. J. Zhao and C. Liu, Royal Society of Chemistry, London, 2023, vol. 41, ch. 2, pp. 20–44 Search PubMed.
- L. Etzbach, S. Gola, F. Küllmer, I.-H. Acir, D. Wohlt, L. M. Ignatzy, S. Bader-Mittermaier and U. Schweiggert-Weisz, Proc. Natl. Acad. Sci. U. S. A., 2024, 121, e2319019121 CrossRef CAS PubMed.
- M. Das Purkayastha, J. Gogoi, D. Kalita, P. Chattopadhyay, K. S. Nakhuru, D. Goyary and C. L. Mahanta, J. Agric. Food Chem., 2014, 62, 7903–7914 CrossRef CAS PubMed.
- S. Chanioti, G. Liadakis and C. Tzia, in Food Engineering Handbook, ed. T. Varzakas and C. Tzia, CRC Press, Boca Raton, FL, 2014, ch. 6, pp. 253–286 Search PubMed.
- R. J. Clarke, in Coffee, ed. R. J. Clarke and R. Macrae, Springer, Dordrecht, 1987, vol. 2, ch. 5, pp. 109–145 Search PubMed.
- A. Marini, A. Muñoz-Losa, A. Biancardi and B. Mennucci, J. Phys. Chem. B, 2010, 114, 17128–17135 CrossRef CAS PubMed.
- Q. Shao, Y. Fan, L. Yang and Y. Qin Gao, J. Chem. Phys., 2012, 136, 115101 CrossRef PubMed.
- W. S. Ratnayake, R. Hoover and T. Warkentin, Starch/Staerke, 2002, 54, 217–234 CrossRef CAS.
- M. Sajib, E. Albers, M. Langeland and I. Undeland, Sci. Rep., 2020, 10, 9590 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |









