Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Valorisation of tomato pomace: bioactive compounds, antimicrobial activity and photodynamic inactivation
Raquel
Nunes da Silva†
 *ab,
Bárbara
Maurício†
*ab,
Bárbara
Maurício†
 abc,
Ana
Fernandes
abc,
Ana
Fernandes
 a,
Iva
Fernandes
a,
Iva
Fernandes
 ab,
Nuno
Mateus
ab,
Nuno
Mateus
 ab and
Victor
de Freitas
ab and
Victor
de Freitas
 ab
ab
aLAQV-REQUIMTE, Chemistry and Biochemistry Department, Faculty of Sciences, University of Porto, Rua do Campo Alegre 689, 4169-007 Porto, Portugal. E-mail: raq.silva@fc.up.pt
bChemistry and Biochemistry Department, Faculty of Sciences, University of Porto, Rua do Campo Alegre 689, 4169-007 Porto, Portugal
cFaculty of Engineering, University of Porto, s/n, R. Dr Roberto Frias, 4200-465 Porto, Portugal
First published on 1st April 2026
Abstract
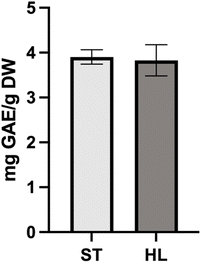
Tomato processing generates large amounts of by-products, particularly tomato pomace, which remains an underutilized source of bioactive compounds. The objective of this study was to investigate the potential valorisation of tomato pomace by characterising its polyphenol and carotenoid composition and evaluating its antimicrobial activity, including under photodynamic inactivation (PDI) conditions. Total polyphenol content determined by the Folin–Ciocalteu method was 3.9 ± 0.2 mg g−1 DW for the standard variety (ST) and 3.8 ± 0.3 mg g−1 DW for the high-lycopene variety (HL). HPLC-DAD-ESI-MS/MS analysis tentatively identified 27 polyphenols, with flavonoids representing the dominant class (82%), particularly naringenin chalcone, rutin, quercetin, and naringenin derivatives. Carotenoid levels reached 0.118 ± 0.003 mg g−1 DW in ST and 0.196 ± 0.002 mg g−1 DW in HL, with lycopene accounting for around 93% of the total carotenoid content. Antimicrobial assays revealed only modest activity overall. Polyphenol extracts showed a slight enhancement in antibacterial effect against Staphylococcus aureus when combined with UVA irradiation, while carotenoid extracts displayed little photodynamic response. These findings highlight tomato pomace as a relevant source of bioactive compounds and support its potential valorisation as a sustainable ingredient in food preservation strategies.
Sustainability spotlightTomato pomace, a major by-product of industrial processing, is typically underutilized and contributes to food waste, representing a lost opportunity for sustainable resource use. This work valorises tomato pomace by optimizing extraction and characterization of polyphenols and lycopene, demonstrating antimicrobial activity and potential as a natural food preservative. By converting processing residues into functional ingredients, the study reduces waste, promotes circular economy strategies, and supports sustainable food production. These findings align with the UN Sustainable Development Goals 12 (Responsible Consumption and Production) and 2 (Zero Hunger), offering a practical approach to enhance food system efficiency while fostering health-oriented, naturally preserved products. |
1 Introduction
In 2022, global production of tomato reached 186 million tonnes over 5 million hectares, making it the most produced vegetable crop.2 Industrial processing converts a large share of this production into sauces, soups, juices, pastes, ketchups, salsas, and canned products.3 The World Processing Tomato Council reports that nearly 39 million tonnes of tomatoes are processed annually, with 83% of this volume concentrated in the ten leading producing countries.4The FAO estimates that about one-third of all food produced for human consumption is lost or discarded.5 Within the tomato industry, this translates into roughly 8.5 million tonnes of by-products annually,6 with tomato pomace representing the largest fraction of this waste.3 The management of such residues is a major sustainability issue, as improper disposal can harm the environment. The most common practice for tomato waste management is to use it as animal feed. Due to its optimal nutrient composition, it has been used for many different feeds such as cattle feed, goat, pig, chicken, cow, sheep and even rabbit feed.7 Revalorisation of pomace presents numerous advantages, including reducing food waste, minimizing environmental impact, and lowering processing costs.8
Tomato by-products are notable sources of both nutrients and biologically active molecules, including carotenoids, polyphenols, dietary fibres, fatty acids (linoleic, oleic, palmitic), tocopherols, sterols, and proteins.6,9 These compounds contribute health-related properties such as antioxidant, antimicrobial, vasodilatory, cardioprotective, anti-inflammatory, and anti-allergenic effects.9 This makes them attractive for valorisation into value-added applications, such as ingredients for functional foods.10 These by-products have been added into a variety of food products for improvement of sensory properties, to extend shelf-life and as a nutritive supplement due to their richness in bioactive compounds.11 Tomato by-products have been incorporated into wheat-flour based foods such as crackers, cookies, bread and pasta; some meat products such as hamburger and sausages, and other products such as tomato paste, soda, ice cream and jams.6,7 Bioethanol and biogas production is also another method for valorisation of by-products, as tomato wastes can be considered as low-cost feedstocks, as a source of lignocellulosic matter for bioethanol and biogas production.12,13 Natural compounds such as vitamins, lycopene, polyphenols, oils, proteins and fibers present in the waste material are also extracted and purified into high value-added products for the pharmaceutical and cosmetic industry, as well as to produce organic fertilizers.6
Food products are highly perishable which facilitates microbial colonization.1 Contamination during cultivation, storage, and transport often leads to spoilage, resulting in flavours, odour, colour, and texture deterioration, as well as a potential risk for human health as some can carry pathogens responsible for food-borne illnesses.14 Preservation methods are therefore essential to ensure safety and shelf life. Biological approaches, including the application of natural antimicrobials – non-thermal endogenous chemical preservation method, are gaining attention as sustainable alternatives to synthetic additives.15
Beyond conventional chemical preservation strategies, antimicrobial photodynamic inactivation (aPDI) has recently emerged as a non-thermal strategy for microbial control in food systems. This approach uses photosensitizers (PS) activated by specific light wavelengths to generate reactive oxygen species (ROS) that inactivate bacteria, fungi, parasites, and viruses.16 Considerable research is now exploring aPDI for reducing foodborne pathogens and extending shelf life in various products.17 Treatment efficiency depends strongly on the choice of PS, which dictates selectivity, activity, and safety.18
In this study, we investigated tomato processing by-products, specifically tomato pomace, as potential sources of natural antimicrobials and photosensitizers, namely polyphenols and carotenoids. Extraction methods were optimized to efficiently recover and characterize these bioactive compounds, improving upon previously reported HPLC techniques. The resulting extracts were evaluated for their antioxidant potential and tested against models of food spoilage, highlighting the promise of tomato residues as functional ingredients and natural preservatives in the food industry.
Although tomato processing by-products have been studied as sources of bioactive compounds, their potential use as natural photosensitizers for antimicrobial photodynamic inactivation remains poorly explored. In particular, studies combining detailed characterization of polyphenols and carotenoids in tomato pomace with the evaluation of their antimicrobial activity under light activation are still limited.
2 Materials and methods
2.1 Materials
Methanol, Folin–Ciocalteu reagent, gallic acid, β-carotene, phenol, Mueller Hinton Broth, were purchased from Sigma-Aldrich (St. Louis, MO, USA). TPTZ from Acros Organics (Geel, Belgium) and acetic acid from Carlo Erba (Milano, Italy). Acetone, hexane, and ethyl acetate from LabChem (Pennsylvania, United States). Acetonitrile and hydrochloridric acid from Chem-Lab (Zedelgem, Belgium). Sulfuric acid and sodium carbonate from PanReac AppliChem ITW reagents (Barcelona, Spain). Potassium dihydrogen phosphate from VWR chemicals (Pennsylvania, USA). Sodium chloride and sodium phosphate dibasic from Honeywell (North Carolina, USA). Mueller Hinton agar from Liofilchem (Roseto degli Abruzzi, Italy).Industrially produced dried tomato pomace, including standard tomato pomace (ST) from conventional varieties and high-lycopene tomato pomace (HL) from hybrid varieties, was kindly provided by Kagome Foods Portugal, S.A., (Castanheira do Ribatejo, Vila Franca de Xira, Portugal) and stored at room temperature. Exact details of the production process cannot be disclosed due to industrial confidentiality. The ST and HL varieties were selected to investigate the pomace derived from pulp and sauce products, with the pomace consisting of skin and seeds. Markedly, the HL variety, used in the production of patented products with health-related prevention claims, was included to assess the composition and bioactivity of its pomace. Samples were collected immediately after industrial pulp extraction, dried, and stored at room temperature until analysis, allowing a comparative evaluation of bioactive compound profiles and antimicrobial properties between the two varieties.
2.2 Polyphenol extraction from tomato pomace
Adapted from a method by Szabo et al.,19 0.5 g of the lyophilized samples of both tomato varieties were extracted with 25 mL of methanol: water (80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20) and ultra-sonicated in a ultrasonic sonicator bath (VWR Ultrasonic Cleaner, Pennsylvania, United States) for 60 min at room temperature. Mixtures were centrifuged (Dynamica Velocity 14 Refrigerated Centrifuge, Dynamic Scientific Ltd; Livingston, UK) at 18
20) and ultra-sonicated in a ultrasonic sonicator bath (VWR Ultrasonic Cleaner, Pennsylvania, United States) for 60 min at room temperature. Mixtures were centrifuged (Dynamica Velocity 14 Refrigerated Centrifuge, Dynamic Scientific Ltd; Livingston, UK) at 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000×g for 10 min at 4 °C and the samples were filtered through a 0.45 µm syringe and evaporated on a rotary evaporator under vacuum (Rotavapor R-114, Büchi, Flawil, Switzerland). Samples were resuspended in 5 mL of methanol: water Milli Q (50
000×g for 10 min at 4 °C and the samples were filtered through a 0.45 µm syringe and evaporated on a rotary evaporator under vacuum (Rotavapor R-114, Büchi, Flawil, Switzerland). Samples were resuspended in 5 mL of methanol: water Milli Q (50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50) and refrigerated at −18 °C.
50) and refrigerated at −18 °C.
2.3 Quantification of the total phenolic content by Folin–Ciocalteu colorimetric assay
In each well of a 96 well plate, 3.75 µL of polyphenol extract, 18.75 µL of Folin–Ciocalteu reagent, and 125 µL of distilled water were added and the mixture was shaken for 30 seconds. 75 µL of 20% Na2CO3 and 152.5 µL of water was added and the plate left at room temperature for 30 min in the dark. The plate was analysed using absorbance at 750 nm in a plate reader (Biotek, Berthold Technologies, Bad Wildbad, Germany).20 A calibration curve was prepared using standard solutions of gallic acid with results expressed in mg of gallic acid equivalents (GAE) per g of dry weight (DW).2.4 Quantification and identification of the polyphenol content by HPLC-DAD/ESI-MS/MS
The polyphenol content of each sample was analysed through High Performance Liquid Chromatography, with a method adapted from Xiao et al.,21 with key modifications, including flow rate, gradient parameters, and duration. A reverse phase C18 column (Purospher STAR RP-18, 150 × 4.6 mm, i.d.: 5 µm) was used at 25 °C. The solvents used were H2O![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) CH3COOH (99
CH3COOH (99![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, v/v) (A) and CH3OH (B), at a flow rate of 0.4 mL min−1 (0–30 min: 95% A, 30–40 min: 70% A, 40–45 min: 60% A, 45–52 min: 50% A, 52–60 min: 30% A, 60–65 min: 60% A, 65–70 min: 95% A). Detection was made from 200 to 400 nm.
1, v/v) (A) and CH3OH (B), at a flow rate of 0.4 mL min−1 (0–30 min: 95% A, 30–40 min: 70% A, 40–45 min: 60% A, 45–52 min: 50% A, 52–60 min: 30% A, 60–65 min: 60% A, 65–70 min: 95% A). Detection was made from 200 to 400 nm.
Mass detection was made on a Finnigan LCQ DECA XP MAX (Finnigan Cor., San Jose, CA, USA) quadrupole ion trap equipped with an atmospheric pressure ionization (API) source using an electrospray ionization (ESI) source. The vaporizer and capillary voltages were 5 kV and 4 V, respectively. The capillary temperature was set at 325 °C. Nitrogen was used as both sheath and auxiliary gas flow rates of 80 and 30, respectively (in arbitrary units). Spectra were recorded in the negative mode between m/z 120 and 2000. The mass spectrometer was programmed to do a series of three scans: a full mass, a zoom scan of the most intense ion in the first scan, and a MS/MS of the most intense ion using relative collision energies of 30 and 60 V.
2.5 Carotenoid extraction from tomato pomace
Carotenoid extraction method, adapted from Choi et al.17 and Szabo et al.,18 consisted of 1 g of tomato pomace sample of both varieties placed into a falcon with 20 mL of an acetone: hexane solution (4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6, v/v). Samples were wrapped in an aluminon foil to keep from light exposure, ultra-sonicated in an ultrasonic sonicator bath (VWR Ultrasonic Cleaner, Pennsylvania, United States) for 10 minutes at 30 °C and then centrifuged at 18
6, v/v). Samples were wrapped in an aluminon foil to keep from light exposure, ultra-sonicated in an ultrasonic sonicator bath (VWR Ultrasonic Cleaner, Pennsylvania, United States) for 10 minutes at 30 °C and then centrifuged at 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) × g for 10 minutes at 4 °C. The pellet was re-extracted, and the supernatants were combined.
× g for 10 minutes at 4 °C. The pellet was re-extracted, and the supernatants were combined.
2.6 Quantification of carotenoids from tomato pomace via UV-vis analysis
Spectrophotometric quantification, based on Szabo et al.18 method, consisted of absorbance measurement at 450 nm in a UV-vis spectrophotometer (PerkinElmer Lambda 365, Massachusetts, US). A standard curve was performed with β-carotene standard, from 0.0005 to 0.005 mg mL−1.2.7 Quantification of carotenoids via HPLC-DAD
The HPLC method was adapted from Sérino et al.19 using a reverse phase C18 column (Purospher STAR RP-18, 150 × 4.6 mm, i.d.: 5 µm) at 30 °C. The solvent used was acetonitrile: water: ethyl acetate (53![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 7
7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40, v/v/v) at a flow rate of 0.4 mL min−1 for 8 minutes. Detection was made from 350–500 nm. Standard curves were performed with concentrations ranging from 0.0005 to 0.005 mg mL−1 and 0.0075 to 0.0275 mg mL−1 for β-carotene and lycopene, respectively.
40, v/v/v) at a flow rate of 0.4 mL min−1 for 8 minutes. Detection was made from 350–500 nm. Standard curves were performed with concentrations ranging from 0.0005 to 0.005 mg mL−1 and 0.0075 to 0.0275 mg mL−1 for β-carotene and lycopene, respectively.
2.8 Antimicrobial activity of polyphenol- and carotenoid-rich extracts
Strains used in this study included Escherichia coli (E. coli) ATCC 25922 and Staphylococcus aureus (S. aureus) ATCC 29213. Bacteria were grown on Mueller Hinton (MH) agar plates from stock cultures, incubated at 37 °C for 24 hours prior to each assay to obtain fresh cultures.Polyphenol and carotenoid-rich extracts were dried in a CentriVap Benchtop Centrifugal Vacuum Concentrator (Labconco, Missouri, United States) and resuspended in 100% dimethyl sulfoxide (DMSO). Samples were filtered with 0.22 µm filter and diluted with phosphate buffered saline (PBS) to achieve 5% DMSO. Optical density at 600 nm of each inoculum was adjusted to 0.1 (approximately 1 × 108 CFU mL−1) in PBS and then diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100. In a 96 well plate, 50 µL of bacterial suspension was mixed with 50 µl of diluted extract. The antimicrobial activity of the samples was tested with and without light irradiation (two groups). For carotenoids' extract, irradiation was performed for 1 hour, using a device composed of light-emitting diodes (LEDs) emitting at 400–700 nm at an irradiance of 25 mW cm−2 measured by a Spectroradiometer USB2000 + RAD (OceanOptics, USA). For polyphenols, a UVA lamp 365 nm with irradiance of 42 mW cm−2 was used. Light sources were chosen by evaluation of the absorption spectrum of the extracts (performed via a spectophotometer) and overall energy dose based on studies with similar doses.20 The samples were drop plated in MH agar for colony forming units (CFU) count, with 6 serial dilutions. Plates were incubated for 24 hours at 37 °C.
100. In a 96 well plate, 50 µL of bacterial suspension was mixed with 50 µl of diluted extract. The antimicrobial activity of the samples was tested with and without light irradiation (two groups). For carotenoids' extract, irradiation was performed for 1 hour, using a device composed of light-emitting diodes (LEDs) emitting at 400–700 nm at an irradiance of 25 mW cm−2 measured by a Spectroradiometer USB2000 + RAD (OceanOptics, USA). For polyphenols, a UVA lamp 365 nm with irradiance of 42 mW cm−2 was used. Light sources were chosen by evaluation of the absorption spectrum of the extracts (performed via a spectophotometer) and overall energy dose based on studies with similar doses.20 The samples were drop plated in MH agar for colony forming units (CFU) count, with 6 serial dilutions. Plates were incubated for 24 hours at 37 °C.
2.9 Statistical analyses
The data was analysed with the statistical program GraphPad Prism 10.3.1 for MacOS (GraphPad Software, La Jolla California, USA). The mean and standard deviations within samples were calculated for all cases. The differences between measurements were obtained by the application of ANOVA with Tukey's post-test (for more than two groups), and the unpaired t-test to analyse the differences between two groups. Statistical calculations were based on a confidence level of >95% (p < 0.05 was considered statistically significant). Three independent extraction replicas were analysed, and three analytical replicates were performed for each.3 Results and discussion
3.1 Polyphenols
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20 methanol–water solvent with UAE at room temperature as the optimal approach, since elevated temperatures promoted polyphenol degradation and oxidation.23
20 methanol–water solvent with UAE at room temperature as the optimal approach, since elevated temperatures promoted polyphenol degradation and oxidation.23
There is a wide range of published values for tomato by-products (including tomato waste which represents any unwanted parts of a tomato that arise during processing, including bruised parts, undeveloped fruits, or any other unusable material),24 with TPC as low as 0.7525 to 18.65 mg GAE g−1 DW, whilst the TPC for this study fits well within this range of values reported in literature.26,27 The wide range of phenolic contents reported in the literature can be attributed to genetic variability, ripening stages, environmental influences, agricultural practices, processing conditions and extraction method.7,26
The HPLC-TPC of the ST and HL tomato pomace was measured at 2.9 ± 0.1 mg g−1 DW and 2.9 ± 0.2 mg g−1 DW, respectively. These values fall within the ranges reported in other studies for different tomato varieties, such as 1.12–4.08 mg g−1 DW![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 22 and 1.93–5.21 mg g−1 DW.23 The TPC values obtained via the Folin method were, as expected, slightly higher than those measured by HPLC analysis, considering the Folin assay can oftentimes overestimate polyphenol content due to interference from other reducing compounds, such as the ascorbic acid present in tomatoes.24
22 and 1.93–5.21 mg g−1 DW.23 The TPC values obtained via the Folin method were, as expected, slightly higher than those measured by HPLC analysis, considering the Folin assay can oftentimes overestimate polyphenol content due to interference from other reducing compounds, such as the ascorbic acid present in tomatoes.24
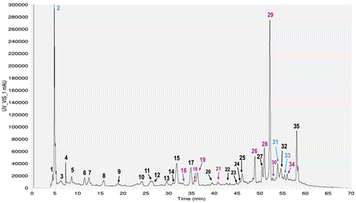
Of the 35 detected peaks, 27 were tentatively identified. For peaks tentative identification and each peak quantification please see Table A1 at SI. The identified compounds included the main classes of phenolic acids (hydroxycinnamic and hydroxybenzoic acids) and flavonoids such as flavanones, flavonols, chalcones, and dihydrochalcones. Phenolic acids accounted for 18% of the total polyphenols, while flavonoids represented 82%, consistent with the findings of Kalogeropoulos et al.,25 who reported flavonoids as the predominant class in tomato by-products, followed by phenolic acids at 23.3%.
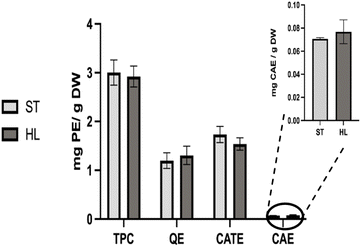
Naringenin chalcone and rutin were the polyphenols detected at the highest concentrations. Consistent with the literature, these compounds, along with quercetin derivatives are the predominant polyphenols in tomato and its by-products.21,22 Rutin concentration (0.86 and 1.10 mg g−1 DW for ST and HL, respectively) were similar to typical values reported for tomato by-products, including averages of 0.84 mg g−1 DW![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 22 and 0.61 mg g−1 DW.23 Its presence reflects the terminal phase of the polyphenols biosynthetic pathway and is often associated with tomato ripening.26 Naringenin chalcone content in the ST variety (0.30 mg g−1 DW) was comparable to literature values of 0.40
22 and 0.61 mg g−1 DW.23 Its presence reflects the terminal phase of the polyphenols biosynthetic pathway and is often associated with tomato ripening.26 Naringenin chalcone content in the ST variety (0.30 mg g−1 DW) was comparable to literature values of 0.40![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 22 and 0.34 mg g−1 DW,19 while the HL variety exhibited a slightly higher level (0.71 mg g−1 DW). During the ripening stages, naringenin chalcone and other flavonoids accumulate predominantly in the peel as a result of the limited expression of flavonoid biosynthetic genes in the flesh of tomatoes.27 This compound also serves as a precursor for most flavonoids, including naringenins, kaempferols, quercetins, and their derivatives.10,27
22 and 0.34 mg g−1 DW,19 while the HL variety exhibited a slightly higher level (0.71 mg g−1 DW). During the ripening stages, naringenin chalcone and other flavonoids accumulate predominantly in the peel as a result of the limited expression of flavonoid biosynthetic genes in the flesh of tomatoes.27 This compound also serves as a precursor for most flavonoids, including naringenins, kaempferols, quercetins, and their derivatives.10,27
Hydroxycinnamic acids were the most abundant subclass among the phenolic acids, with coumaric acid and its derivative detected at the highest level (0.395 mg g−1 DW and 0.150 mg g−1 DW for ST and HL, respectively). This pattern differs from typical reports in the literature,21,25,28 where chlorogenic acid is usually the dominant compound. Although generally abundant in tomato fruits, chlorogenic acid decreases substantially during ripening and the final stages of development,29 which likely explains variations in profiles.
3.2 Carotenoids
Carotenoid accumulation is strongly influenced by tomato ripening, during which biosynthesis is activated mainly in the peel, while inner tissues remain largely inactive.27 Consequently, the peel can contain up to five times more carotenoids than the flesh of ripe fruits.36 By-products, being rich in peel, therefore represent a concentrated source compared to whole tomatoes. Processing steps can also enhance carotenoid extractability and bioavailability, which explains why higher levels are more often detected in purees than in fresh tomatoes.37
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water
water![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ethyl acetate mixture (53
ethyl acetate mixture (53![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 7
7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40) described by Sérino et al.38 was selected, with flow rate adjustments. We identified two main peaks, corresponding to lycopene and β-carotene, the predominant tomato carotenoids.
40) described by Sérino et al.38 was selected, with flow rate adjustments. We identified two main peaks, corresponding to lycopene and β-carotene, the predominant tomato carotenoids.
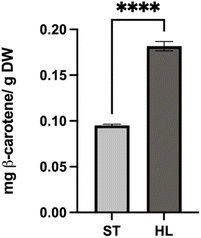
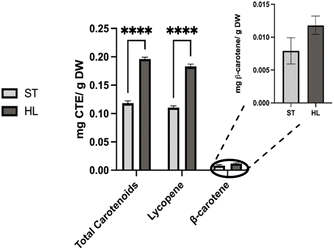
Total carotenoids content in this study was 0.118 ± 0.003 mg g−1 DW for ST and 0.196 ± 0.002 mg g−1 DW for HL (Fig. 5). For chromatogram, please see Fig. A1 at SI.
Although published values in literature vary widely, these findings are within the ranges reported. For instance, Szabo et al.35 found carotenoid concentrations from 0.077 to 0.37 mg g−1 DW in tomato pomace, while their later work22 indicated 0.20–0.38 mg g−1 DW. Strati et al.39 measured 0.004–0.17 mg g−1 DW depending on extraction solvent. Using a hexane: acetone mixture like that applied here, they obtained 0.22 mg g−1 DW, comparable to HL but slightly higher than ST.
These concentrations were only slightly higher than those obtained via spectrophotometric method whichoften overestimate carotenoids content due to absorption at overlapping wavelengths from chlorophyll and degradation products.40
Lycopene concentration in the tomato pomace were 0.110 ± 0.003 mg g−1 DW for ST and 0.183 ± 0.003 mg g−1 DW for HL, with the ST variety falling toward the lower spectrum reported in the literature. Lycopene was the main carotenoid present, at roughly 14 times the β-carotene concentration and accounting for 93% of total carotenoids. These findings align with other reports, where lycopene represented 74–94% of total carotenoids in tomato products.28 Previous studies on tomato pomace have shown a broad distribution of lycopene content. One study22 observed 0.15–0.29 mg g−1 DW, while another report35 indicated a slightly higher average of 0.27 mg g−1 DW. Wider variations have been documented in tomato processing waste, ranging from 0.13 to 0.82 mg g−1 DW,41 and another as low as 0.05 mg g−1 DW.42 β-Carotene concentrations were comparatively low, measured at 0.008 ± 0.002 mg g−1 DW in ST and 0.0129 ± 0.0006 mg g−1 DW in HL, values closer to those found in whole tomatoes (0.0096–0.0703 mg g−1 DW)28 rather than the higher ranges reported for pomace (0.03–0.08 mg g−1 DW).22,35
3.3 Antimicrobial activity of the polyphenol- and carotenoid-rich extracts
E.coli and S. aureus were selected as representative models of Gram-negative and Gram-positive bacteria, respectively, allowing the evaluation of potential differences in susceptibility related to bacterial cell wall structure. In addition, both microorganisms have been previously associated with contamination and spoilage scenarios in tomato, making them relevant initial models for assessing the antimicrobial potential of tomato pomace extracts.13PDI efficacy depends on the wavelength of light, which must align with the absorption spectrum of the PS.17 Absorption spectra of the extracts were first determined to guide light selection. Absorption spectrum of polyphenol and carotenoids extracts of HL variety (as an example) can be found at SI (Fig. A2–A5).
Polyphenol extracts absorbed mainly between 200 and 400 nm. UVA was chosen to minimize bactericidal effects from light alone,17 with previous studies having tested antimicrobial activity also using UVA-based PDI with polyphenols such as gallic acid and caffeic acid.43,44 Carotenoid extracts showed broader absorption between 350 and 550 nm, so a visible light source spanning 400–700 nm was selected. Total energy dose for polyphenols was 150 J cm−2 and for carotenoids 90 J cm−2.
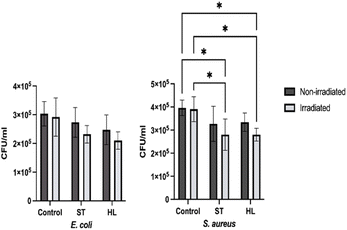
Polyphenol extracts from both ST and HL tomato varieties had no significant effect in CFU mL−1 for E. coli whilst S. aureus showed small, but significant, reductions in the irradiated group, indicating that UVA light enhances polyphenols' antimicrobial effects. Similar trends have been reported in the literature. Studies of tomato by-products19 and tomato seeds45 have demonstrated strong activity of polyphenols against Gram-positive bacteria, with limited effects on Gram-negative species. E. coli's higher resistance is attributed to the Gram-negative outer membrane, composed of lipopolysaccharides, which limits polyphenol penetration.45 Another study,46 reported inhibition of tomato pathogens, including E. coli and S. aureus, with MIC values of 3125–12500 µg mL−1, and noted a concentration-dependent antimicrobial response. Given the average polyphenol concentration in this study was considerably smaller, 386 µg mL−1, the minor, and non-significant, reductions observed align with these findings.
Polyphenols have well documented antimicrobial activity, which is closely related to their chemical structure, including hydroxyl, methoxy, prenyl groups, and glycosylation patterns.47 These can affect microorganisms by disrupting membranes, interfering with metabolic pathways, chelating essential metal ions, damaging nucleic acids, and inhibiting key enzymes.47 Flavonoids such as quercetin, kaempferol, naringenin, isorhamnetin glycosides, and rutin have been shown to actively inhibit bacterial growth.48,49 Chalcones and quinones have also been proven to enhance the effectiveness of antibiotics by modulating efflux pumps and increasing intracellular drug accumulation.50 These findings confirm that the high levels of polyphenols present in these pomaces can contribute directly to their antimicrobial activity.
Evaluating the PDI results, it should be noted that light exposure alone (control samples without polyphenols) did not inhibit bacterial growth. Treatment with polyphenol extracts combined with UVA irradiation resulted in a modest log reduction of 0.14. Although small, this effect aligns with the relatively low polyphenol concentration used, which limits antimicrobial potency. Nevertheless, this decrease can indicate that light exposure enhances the bioactivity of polyphenols. Polyphenols are increasingly investigated as photosensitizers in aPDI. Their aromatic ring systems facilitate intersystem crossing to triplet states, which can generate reactive species via two mechanisms: Type II, where energy transfer to molecular oxygen produces singlet oxygen (as with hypericin and riboflavin), and Type I, involving electron or hydrogen transfer that forms radicals such as superoxide or hydroxyl species. Previous research using quercetin,51 resveratrol,52 gallic acid,44 and caffeic acid,43 with varying wavelengths and concentrations, demonstrated substantial bacterial reductions. Given that the polyphenol concentration in this study is considerably lower than those applied in these reports, the modest reductions observed are consistent with expectations.
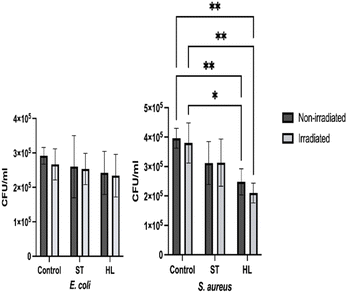
Several studies have highlighted the antimicrobial potential of carotenoid extracts against different bacterial species.53,54 Natividad et al.53 demonstrated that tomato carotenoids inhibited S. aureus growth while having negligible effects on E. coli, consistent with this study's observations. Similarly, carotenoids from Peltophorum petrocarpum55 and Sinularia sp.56 showed strong activity against S. aureus but limited effects on Gram-negative bacteria. In this study, the carotenoid concentration in the extracts was only 3 and 5 µg mL−1, far below the levels reported to achieve complete bacterial inhibition, which explains the modest log reductions observed and the weaker activity of the ST extract, which contained an even lower carotenoid content. These results emphasize that carotenoid antimicrobial effects are dose dependent, requiring higher concentrations to achieve bacterial inactivation.
The main two carotenoids detected in the tomato pomace are likely responsible for the antimicrobial activity observed. Previous studies have shown that lycopene exhibited inhibitory effects against microorganisms such as S. aureus and C. albicans,57 while β-carotene demonstrated significant activity at concentrations of 100–1200 µg mL−1, particularly against Gram-positive bacteria.58 These bioactive compounds can interact with bacterial outer membranes by binding to porin proteins, disrupting their structure, and reducing cell wall permeability, which limits bacterial growth.56 Additionally, lycopene has been reported to induce bactericidal effects through reactive oxygen species, especially hydroxyl radicals, causing DNA damage beyond bacterial repair capacity.59
Carotenoids absorb visible light and are excited to singlet states, which rapidly undergo internal conversion or intersystem crossing, but instead of transferring energy to oxygen, most carotenoids dissipate the energy through vibrations along their polyene chains, effectively limiting singlet oxygen production.60 Consequently, they mainly act as photoprotective molecules, producing minimal ROS during PDI. Although research on carotenoids as PDI agents is scarce, a recent study61 also tested tomato carotenoids against foodborne bacteria under light exposure. β-Carotene and lutein did not show enhanced antimicrobial activity, while lycopene displayed some intrinsic toxicity but limited photodynamic effects under white light. Effective PDI with lycopene likely requires illumination closely matching its absorption peak (475 nm) rather than a wide wavelength range. In this study, the applied irradiation may not have provided sufficient intensity at the optimal wavelength, explaining the negligible photodynamic antimicrobial enhancement observed for carotenoid extracts. However, this study contributes to improving the documentation and consistency in the investigation of these bioactive compounds as antimicrobials and as photosensitizers for the inactivation of bacterial models relevant to foods and their ingredients, while emphasizing new efforts to develop naturally derived antimicrobials obtained from tomato pomace.
It is also worth considering that photoactivation may induce chemical changes in the extracts themselves. Although the antimicrobial effects observed here were primarily attributed to the generation of reactive oxygen species during photodynamic treatment, modifications in the composition of the extracts could also contribute to the antibacterial response. Future work should therefore assess the chemical profile of the extracts before and after irradiation, allowing evaluation of possible photochemical transformations and their contribution to the antimicrobial activity.
4 Conclusions
This study highlights tomato processing pomace as a rich source of bioactive compounds, supporting their valorisation within a circular economy framework. Optimized extraction and quantification methods enabled the characterization of phenolic and carotenoid content in two industrial produced tomato pomace varieties, alongside assessment of their antimicrobial properties. Notably, the HL variety is employed in the production of patented products with health-related prevention claims, further underscoring the relevance of studying its corresponding pomace.The pomaces exhibited a diverse phenolic profile with 27 tentatively identified polyphenols, from rutin and quercetin derivatives to naringenin chalcone and various hydroxycinnamic acids. Carotenoid analysis confirmed these pomaces as significant amounts of lycopene, with the high-lycopene variety nearly doubling the content of the standard variety. Antimicrobial testing indicated moderated effects of high-lycopene carotenoid extracts against Gram-positive models. Photodynamic inactivation experiments demonstrated that polyphenol extracts could potentially act as natural photosensitizers, producing some bacterial reductions (however moderate and not superior to 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) log-reduction), whereas carotenoids did not show enhanced activity under the tested light conditions.
log-reduction), whereas carotenoids did not show enhanced activity under the tested light conditions.
Overall, these findings validate the optimized methodologies for tomato pomace valorisation, providing the first detailed insight into its bioactive composition. Together, these results position tomato pomace as a valuable resource for sustainable food preservation, functional ingredient development, and broader biotechnological applications.
Future studies should focus on antimicrobial evaluation with other bacterial models and a wider range of extract concentrations and irradiation conditions. This should include the addition of a standard positive antimicrobial control to enable broader comparison of extract efficacy. Further work should also examine changes in the chemical profile after photoactivation, optimise irradiation conditions and extract concentrations and evaluate their effectiveness in more complex food systems.
Author contributions
Raquel Nunes da Silva: conceptualization, supervision, formal analysis, writing – review & editing. Bárbara Maurício: investigation, methodology, data curation, writing – original draft. Ana Fernandes: conceptualization, writing – review & editing. Iva Fernandes: conceptualization, writing – review & editing. Nuno Mateus: conceptualization, writing – review & editing. Victor de Freitas: conceptualization, funding acquisition, project administration, writing – review & editing. All authors reviewed and approved the final version of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
Data will be made available on request.Supplementary information (SI): detailed information on polyphenol characterization including HPLC-DAD-ESI-MS/MS retention times, [M − H]− ions, MS/MS fragmentation patterns, tentative identification and individual polyphenol concentrations. Additionally, chromatograms (HPLC-DAD) from both polyphenol and caratenoid extracts are present as well as absorption spectra of both extracts. See DOI: https://doi.org/10.1039/d5fb00729a.
Acknowledgements
This work received financial support from the PT national funds (FCT/MECI, Fundação para a Ciência e Tecnologia and Ministério da Educação, Ciência e Inovação) through the project UID/50006/2025 – Laboratório Associado para a Química Verde – Tecnologias e Processos Limpos. We would like to thank Doctor Ana Silva for her assistance with conducting irradiance measurements at the Physics Department of University of Porto Faculty of Sciences. We also wish to express our gratitude to Kagome Foods Portugal, S.A. for the challenge proposed, to which this research will contribute through the development of innovative solutions, and for kindly providing the samples on which this study is based. AF acknowledges FCT through research contract (https://doi.org/10.54499/CEECIND/00029/2018/CP1545/CT0010). IF acknowledges her assistant professor contract CEECINST/00064/2021.References
- J. Tiwari, T. Behera, N. Rai, S. Yerasu, M. Singh and P. Singh, J. Veg. Sci., 2021, 49, 123–132 Search PubMed.
- D. Kaur, A. A. Wani, D. P. S. Oberoi and D. S. Sogi, Food Chem., 2008, 108, 711–718 CrossRef CAS PubMed.
- World Processing Tomato Council, https://www.wptc.to/production/, accessed 1 March 2025.
- FAO, Global Food Losses and Food Waste- Extent, Causes and Prevention, Rome, 2011 Search PubMed.
- K. Szabo, A.-F. Cătoi and D. Vodnar, Plant Foods Hum. Nutr., 2018, 73, 268–277 CrossRef CAS PubMed.
- Z. Lu, J. Wang, R. Gao, F. Ye and G. Zhao, Trends Food Sci. Technol., 2019, 86, 172–187 CrossRef CAS.
- M. Casa, M. Miccio, G. De Feo, A. Paulillo, R. Chirone, D. Paulillo, P. Lettieri and R. Chirone, Detritus, 2021, 15, 31–39 CrossRef.
- M. C. Coelho, A. S. Rodrigues, J. A. Teixeira and M. E. Pintado, Food Chem., 2023, 410, 135319 CrossRef CAS PubMed.
- B. Farinon, M. Felli, M. Sulli, G. Diretto, D. V Savatin, A. Mazzucato, N. Merendino and L. Costantini, Food Chem., 2024, 452, 139509 CrossRef CAS PubMed.
- I. B. Chabi, O. Zannou, E. S. C. A. Dedehou, B. P. Ayegnon, O. B. Oscar Odouaro, S. Maqsood, C. M. Galanakis and A. Pierre Polycarpe Kayodé, Heliyon, 2024, 10, e25261 CrossRef CAS PubMed.
- Y. Li, Y. Li, D. Zhang, G. Li, J. Lu and S. Li, Bioresour. Technol., 2016, 217, 50–55 CrossRef CAS PubMed.
- M. Lenucci, M. Durante, A. Montefusco, G. Dalessandro and G. Piro, J. Agric. Food Chem., 2013, 61, 3683–3692 CrossRef CAS PubMed.
- A. O. Momoh, A. E. Aladeyelu and D. E. Tuomokeme, Eur. J. Biotechnol. Biosci., 2020, 6, 660–672 Search PubMed.
- P. M. Davidson, H. B. Cekmer, E. A. Monu and C. Techathuvanan, in Woodhead Publishing Series in Food Science, Technology and Nutrition, ed. Q. Taylor, Woodhead Publishing, Oxford, 2015, pp. 1–27 Search PubMed.
- A. Talib, A. Samad, M. J. Hossain, A. Muazzam, B. Anwar, R. Atique, Y.-H. Hwang and S.-T. Joo, FoodLife, 2024, 2024, 19–32 Search PubMed.
- J. Qiu, H. Yang, Y. Zhang, Y. Xiao, L. Wang, Y. Peng, X. Yu, X. Huang and T. Zhong, Trends Food Sci. Technol., 2024, 143, 104295 CrossRef CAS.
- S. Zhu, Y. Song, J. Pei, F. Xue, X. Cui, X. Xiong and C. Li, Food Chem.:X, 2021, 12, 100150 CAS.
- V. S. Ghate, W. Zhou and H.-G. Yuk, Compr. Rev. Food Sci. Food Saf., 2019, 18, 402–424 CrossRef PubMed.
- K. Szabo, Z. Diaconeasa, A. F. Cătoi and D. C. Vodnar, Antioxidants, 2019, 8, 292 CrossRef CAS PubMed.
- K. Zhou and L. Yu, LWT–Food Sci. Technol., 2004, 37, 717–721 CrossRef CAS.
- X. Xiao, F. Liu, M. Sun, Z. Tang, Y. Wu, J. Lyu, K. S. Khan and J. Yu, J. Chromatogr. A, 2024, 1733, 465254 CrossRef CAS PubMed.
- K. Szabo, F. V. Dulf, Z. Diaconeasa and D. C. Vodnar, LWT, 2019, 116, 108558 CrossRef CAS.
- G. Ćetković, S. Savatović, J. Čanadanović-Brunet, S. Djilas, J. Vulić, A. Mandić and D. Četojević-Simin, Food Chem., 2012, 133, 938–945 CrossRef.
- A. Scalbert and G. Williamson, J. Nutr., 2000, 130, 2073S–2085S CrossRef CAS PubMed.
- N. Kalogeropoulos, A. Chiou, V. Pyriochou, A. Peristeraki and V. T. Karathanos, LWT–Food Sci. Technol., 2012, 49, 213–216 CrossRef CAS.
- D. Giuntini, V. Lazzeri, V. Calvenzani, C. Dall'Asta, G. Galaverna, C. Tonelli, K. Petroni and A. Ranieri, J. Agric. Food Chem., 2008, 56, 5905–5915 CrossRef CAS PubMed.
- A.-R. Ballester, J. Molthoff, R. De Vos, B. Hekkert, D. Orzáez, J.-P. Fernandez-Moreno, P. Tripodi, S. Grandillo, C. Martin, J. Heldens, M. Ykema, A. Granell and A. Bovy, Plant Physiol., 2009, 152, 71–84 CrossRef PubMed.
- S. H. Choi, D.-S. Kim, N. Kozukue, H.-J. Kim, Y. Nishitani, M. Mizuno, C. E. Levin and M. Friedman, J. Food Compos. Anal., 2014, 34, 115–127 CrossRef CAS.
- A. Fleuriet and J.-J. Macheix, Phytochemistry, 1981, 20, 667–671 CrossRef CAS.
- N. Baaka, I. El Ksibi and M. F. Mhenni, Nat. Prod. Res., 2017, 31, 196–203 CrossRef CAS PubMed.
- E. Larsen and L. P. Christensen, J. Agric. Food Chem., 2005, 53, 6598–6602 CrossRef CAS PubMed.
- M. Kimura, D. B. Rodriguez-Amaya and H. T. Godoy, Food Chem., 1990, 35, 187–195 CrossRef CAS.
- B. O. Ferrando, N. Baenas, F. Rincón and M. J. Periago, Food Bioprocess Technol., 2024, 17, 3017–3030 CrossRef CAS.
- J. Nagarajan, H. Pui Kay, N. P. Krishnamurthy, N. R. Ramakrishnan, T. M. S. Aldawoud, C. M. Galanakis and O. C. Wei, Biomolecules, 2020, 10, 1019 CrossRef CAS PubMed.
- K. Szabo, B.-E. Teleky, F. Ranga, I. Roman, H. Khaoula, E. Boudaya, A. B. Ltaief, W. Aouani, M. Thiamrat and D. C. Vodnar, Molecules, 2022, 27, 3771 CrossRef CAS PubMed.
- K. Szabo, A.-F. Cătoi and D. Vodnar, Plant Foods Hum. Nutr., 2018, 73, 268–277 CrossRef CAS PubMed.
- A. Bianchi, E. Vitale, V. Guerretti, G. Palumbo, I. Clemente, L. Ottaiano, C. Arena and A. de maio, Antioxidants, 2023, 12, 761 CrossRef CAS PubMed.
- S. Sérino, L. Gomez, G. Costagliola and H. Gautier, J. Agric. Food Chem., 2009, 57, 8753–8760 CrossRef PubMed.
- I. Strati, E. Gogou and V. Oreopoulou, Food Bioprod. Process., 2015, 94, 668–674 CrossRef CAS.
- E. Biehler, F. Mayer, L. Hoffmann, E. Krause and T. Bohn, J. Food Sci., 2010, 75, C55–C61 CrossRef CAS PubMed.
- S. Stajčić, G. Ćetković, J. Čanadanović-Brunet, S. Djilas, A. Mandić and D. Četojević-Simin, Food Chem., 2015, 172, 225–232 CrossRef PubMed.
- M. M. Poojary and P. Passamonti, Food Chem., 2015, 173, 943–950 CrossRef CAS PubMed.
- P. Min-Young and K. Dong-Hyun, Appl. Environ. Microbiol., 2021, 87, e00631–21 Search PubMed.
- A. Cossu, D. Ercan, Q. Wang, W. Peer, N. Nitin and R. Tikekar, Innov. Food Sci. Emerg. Technol., 2016, 37, 44–52 CrossRef CAS.
- M. Taveira, L. R. Silva, L. A. Vale-Silva, E. Pinto, P. Valentão, F. Ferreres, P. Guedes de Pinho and P. B. Andrade, J. Agric. Food Chem., 2010, 58, 9529–9536 CrossRef CAS PubMed.
- N. Silva, S. Ruiz-Cruz, L. Cira, M. Estrada Alvarado, J. Ornelas-Paz, M. López, L. Del Toro, J. F. Ayala-Zavala and E. Marquez Rios, Int. J. Environ. Anal. Chem., 2015, 2015, 1–10 CrossRef PubMed.
- L. De Rossi, G. Rocchetti, L. Lucini and A. Rebecchi, Antioxidants, 2025, 14, 200 CrossRef CAS PubMed.
- A. Vikram, P. R. Jesudhasan, G. K. Jayaprakasha, S. D. Pillai, A. Jayaraman and B. S. Patil, Int. J. Food Microbiol., 2011, 145, 28–36 CrossRef CAS PubMed.
- H. Liu, Y. Mou, J. Zhao, J. Wang, L. Zhou, M. Wang, D. Wang, J. Han, Z. Yu and F. Yang, Molecules, 2010, 15, 7933–7945 CrossRef CAS PubMed.
- M. Makarewicz, I. Drożdż, T. Tarko and A. Duda-Chodak, Antioxidants, 2021, 10, 188 CrossRef CAS PubMed.
- M. A. Olszewska, A. Zimińska, A. Draszanowska and T. Sawicki, Food Microbiol., 2024, 124, 104608 CrossRef CAS PubMed.
- D. P. dos Santos, D. P. Soares Lopes, R. C. de Moraes, C. Vieira Gonçalves, L. Pereira Rosa, F. C. da Silva Rosa and R. A. A. da Silva, Photodiagnosis Photodyn. Ther., 2019, 25, 227–236 CrossRef PubMed.
- L. Natividad and R. Rafael, Res. J. Recent Sci., 2014, 3, 40–45 Search PubMed.
- N. Tao, G. Yumei, L. YueJin and G. Fei, Am. Eurasian J. Agric. Environ. Sci., 2010, 7, 110–115 CAS.
- J. T. J. Amalya, J. Sumathy and I. J. Curr, Trends Pharm. Res., 2015, 3, 748–753 Search PubMed.
- L. Kusmita, A. Nur Prasetyo Edi, Y. Dwi Franyoto, M. Mutmainah, S. Haryanti and A. Dwi Retno Nurcahyanti, Saudi Pharm. J., 2023, 31, 101680 CrossRef CAS PubMed.
- V. Divyadharsini, T. N. Maheswari and R. Shanmugam, Cureus, 2023, 15, e42419 CAS.
- K. Majeed, A. A. L. hashimi and B. Jaber, Annals of the Romanian Society for Cell Biology, 2021, 25, 14006–14028 Search PubMed.
- W. Lee and D. Lee, J. Microbiol. Biotechnol., 2014, 24, 1232–1237 CrossRef CAS PubMed.
- T. Maoka, J. Nat. Med., 2020, 74, 1–16 CrossRef CAS PubMed.
- A. Prasad, E. Wynands, S. M. Roche, C. Romo-Bernal, N. Allan, M. Olson, S. Levengood, R. Andersen, N. Loebel, C. P. Sabino and J. A. Ross, Foods, 2024, 13, 453 CrossRef CAS PubMed.
Footnote |
| † The authors contributed equally to this work. |
| This journal is © The Royal Society of Chemistry 2026 |