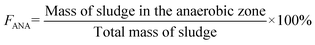Open Access Article
Open Access ArticleDisruption in gene expression cycles of polyphosphate-accumulating organisms is associated with a full-scale enhanced biological phosphorus removal instability event
Jessica A.
Deaver†
 ab,
Thomas
Solon
cd,
Amy M.
Grunden
be and
Douglas F.
Call
ab,
Thomas
Solon
cd,
Amy M.
Grunden
be and
Douglas F.
Call
 *ab
*ab
aDepartment of Civil, Construction, and Environmental Engineering, North Carolina State University, Raleigh, NC 27606, USA. E-mail: dfcall@ncsu.edu
bScience and Technologies for Phosphorus Sustainability (STEPS) Center, North Carolina State University, Raleigh, NC 27606, USA
cDepartment of Environmental Engineering and Earth Sciences, Clemson University, SC 29634, USA
dRenewable Water Resources (ReWa), Greenville, SC 29607, USA
eDepartment of Plant and Microbial Biology, North Carolina State University, Raleigh, NC 27606, USA
First published on 31st March 2026
Abstract
Enhanced biological phosphorus removal (EBPR) enriches polyphosphate-accumulating organisms (PAOs) via alternating anaerobic/aerobic feast–famine cycles to remove phosphorus from wastewater. EBPR can be prone to instability, although the causes are often unclear. Genome-centric metatranscriptomics was used to investigate an EBPR instability event at a full-scale facility that typically experiences a winter instability event to identify changes in microbial community composition and gene expression characteristic of reduced EBPR performance. The facility sampled operates an anaerobic/anoxic/oxic (A2O) EBPR process. Notably, the process monitoring data indicated few process changes beyond increased effluent phosphorus (>3 mg L−1 compared to typical concentrations <0.5 mg L−1) and lower water temperatures during the instability event. Microbial community composition remained consistent before, during, and after the instability event. Two PAO MAGs, Ca. Accumulibacter phosphatis and Ca. Accumulibacter propinquus, were the most abundant and transcriptionally active PAOs. DESeq2 analyses of significantly (adjusted p-value < 0.01) and differentially (|log2(FoldChange)| >1) expressed genes revealed that the expression of key carbon metabolism, energy metabolism, and denitrification genes that typically peak in the anaerobic zone under anaerobic, high carbon conditions shifted to peak in the anoxic zone during the instability. These results demonstrate a shift in PAO activity, and not community composition, associated with a full-scale EBPR instability event.
Water impactUnstable biological phosphorus removal remains an ongoing challenge for many water resource recovery facilities (WRRFs). Improving the stability of the biological process can increase WRRF confidence in the process. We find strong evidence that unstable phosphorus removal at a full-scale WRRF is associated with significant changes in the activity of abundant polyphosphate-accumulating organisms, likely driven by changes in redox conditions across the biological basins. |
Introduction
Water resource recovery facilities (WRRFs) often remove phosphorus from wastewater via enhanced biological phosphorus removal (EBPR). In EBPR, alternating anaerobic and aerobic zones enrich polyphosphate-accumulating organisms (PAOs). PAOs uptake soluble carbon, such as volatile fatty acids (VFAs) and sugars, and store it intracellularly, in forms such as polyhydroxyalkanoates (PHAs), under anaerobic conditions. PAOs generate the energy (i.e., ATP) and reducing power (e.g., NADH) required to internally store carbon by degrading stores of glycogen and polyphosphate (poly-P). Then, under aerobic conditions, PAOs respire on oxygen and oxidize internal carbon for growth and cell division, replenishing their stores of glycogen and poly-P and removing phosphorus from the bulk solution in the process.Phosphorus removal in full-scale EBPR can become unstable, often for reasons not well understood.1 Many factors may contribute to EBPR stability, or lack thereof, including the redox conditions and the concentration of VFAs entering the anaerobic stage.2,3 Side-stream EBPR (S2EBPR) can help with stability by improving VFA supply and enabling a sufficiently reduced redox environment.4 Some process data and EBPR stability correlate, including positive relationships between readily biodegradable chemical oxygen demand (rbCOD) to influent phosphorus ratios and effluent total phosphorus (TP). Other process data, such as biochemical oxygen demand (BOD):TP and COD:TP, are often not directly correlated.3 Consequently, deciphering the root cause of full-scale EBPR instabilities from process data is challenging. Deeper insight into the underlying biology may help reveal the causes of instability.
Our knowledge of the microorganisms involved and the biochemical pathways they use is primarily limited to stable EBPR systems. Full-scale EBPR systems that experience instabilities have received less attention. The most dominant PAOs in stable EBPR systems globally belong to the genera Candidatus (Ca.) Accumulibacter, Azonexus (formerly Dechloromonas), and Ca. Phosphoribacter (formerly Tetrasphaera).5Ca. Accumulibacter is considered the canonical model for PAO metabolism, though it is now known that there can be vast diversity between different Ca. Accumulibacter strains.6,7 In unstable systems, linkages between microbial community and activity dynamics and instability are limited.8–10 Lindner et al.10 used genome-centric metagenomics and metatranscriptomics on two samples collected during stable operation and two samples collected approximately six weeks later during unstable operation. They observed that unstable P removal was associated with changes in mixed liquor suspended solids, decreased VFA loads, and increased primary effluent orthophosphate, and they reported genus level changes in Ca. Accumulibacter relative abundance and carbon metabolism gene expression during unstable EBPR. However, low sample numbers and the length of time between sampling events make it difficult to discern if the shift in community dynamics led to the instability or vice versa.
In this study, we utilized metagenome-based transcriptomics to investigate changes, or lack thereof, to microbial community structure and the regulation of population-specific pathways before versus during an instability event occurring within two solids retention times (SRTs) at a full-scale WRRF with an anaerobic/anoxic/oxic (A2O) process configuration. This facility historically experiences EBPR instability in the winter, so we conducted our sampling campaign during that time. We collected samples throughout two parallel treatment trains on three days before, during, and after an instability event defined by effluent phosphorus exceeding 1 mg P L−1. We used metagenomic de novo assemblies and phylogenomic comparisons to conduct a species-level identification of key PAOs, and mapped RNA reads to metagenome-assembled genomes (MAGs) to identify expression-level gene changes. Our findings show that shifts in PAO activity, and not community composition, can be associated with a full-scale instability event.
Materials and methods
Facility description
The WRRF studied (operated by Renewable Water Resources in Greenville, SC, USA) is in southeastern United States. During the study period, it treated an average daily flow of 16 million gallons per day (MGD) of primarily residential and industrial wastewater from a combined sewer system. The facility performs biological nitrogen and phosphorus removal via three parallel A2O treatment trains that each receive an equal proportion of primary clarifier effluent. The SRT of the A2O process during the study was ∼10 days with a mixed liquor return rate of 6.4 MGD. The anaerobic mass fraction was ∼13.5%, which was calculated using the following equation, where FANA is the fraction of mass in the anaerobic zone.The facility typically adds waste from a juice processing facility as a supplemental carbon source. Influent measurements were taken using a pump station composite sampler that includes side stream flow back to the head of the plant (∼1 MGD). Effluent measurements were collected post deep bed sand filters, UV disinfection, and re-aeration using a composite sampler. The facility historically experiences unstable EBPR in the winter (Fig. S1), which guided decisions on when to collect more comprehensive sets of samples (i.e., samples for DNA/RNA sequencing across all redox zones and two parallel basins).
Sample collection
Fresh activated sludge (AS) samples were collected and preserved for DNA and/or RNA sequencing. Approximately 500 mL of AS was collected at multiple sampling locations along two A2O basins (Fig. 1) and sub-sampled as described in the SI. Samples preserved in 2× DNA/RNA Shield solution (Zymo Research, Irvine, CA, USA) were shipped at room temperature to NC State University (Raleigh, NC, USA), where they were aliquoted into 2 mL tubes and stored at −20 °C until further processing. To collect samples for simultaneous orthophosphate measurements, an 8 mL subsample was immediately filtered with a 0.45 μm filter and shipped overnight on ice to NC State.DNA sequencing using both short read (Illumina) and long read (Oxford Nanopore) technologies was performed on eight samples collected from the end of the aerobic zone (location G in Fig. 1) of two parallel treatment trains on 11/20/2023, 11/29/2023 (unstable EBPR), 12/6/2023, and 2/23/2024 (Table S1). RNA sequencing using short read (Illumina) technology was performed on 30 samples collected from the beginning and end of each zone (locations A through G in Fig. 1) of two parallel treatment trains on 11/20/2023 (n = 12), 11/29/2023 (n = 6), and 12/6/2023 (n = 12). The samples collected from the same location in each of the two parallel treatment trains were considered biological replicates (n = 2 per sample location per date). On 11/29/2023, samples were collected only from the end of each redox zone (locations B, D, and G) due to time constraints. Samples for RNA sequencing were collected on 2/23/2024 (n = 6) as well; however, the facility was experiencing reduced nitrification at that time, so it was decided to exclude those samples from further metatranscriptomic analyses but include them in the metagenomic draft genome reconstruction.
Sample processing and nucleic acid extractions
DNA was extracted using the FastDNA Spin Kit (MP Biomedicals, Santa Ana, CA, USA) with modifications described in the SI. DNA was eluted in 50 μL of 10 mM Tris–HCl at pH 8.0 and stored in multiple aliquots at −20 °C.RNA was extracted using the ZymoBiomics RNA miniprep kit (Zymo Research) following the manufacturer's protocols with modifications described in the SI. RNA was stored in multiple aliquots at −80 °C.
DNA and RNA purity was assessed using a Nanodrop Spectrophotometer ND-1000 (ThermoFisher Scientific, Waltham, MA, USA), and the quantity was assessed using dsDNA BR assays and RNA BR assays with a Qubit 3.0 Fluorometer (ThermoFisher Scientific). DNA and RNA integrity were evaluated using gDNA and RNA ScreenTape Analyses on an Agilent 4150 TapeStation (Agilent Technologies, Inc., Santa Clara, CA, USA).
Library preparation and sequencing
For Oxford Nanopore sequencing, DNA libraries were prepared using the Native Barcoding Kit 24 V14, SQK-NBD114, according to the manufacturer's protocols (Oxford Nanopore Technologies, Oxford, UK). Samples were pooled at equimolar concentrations and run on an Oxford Nanopore MinION Mk1C device with an R14 FLO-MIN114 flow cell. The flow cell was run within one month of delivery and run for 48 hours. The data output equaled 16.5 Gbp with an estimated N50 of 7.2 kbp.For Illumina sequencing, DNA and RNA samples were shipped frozen to Admera Health, LLC (South Plainfield, NJ, USA). DNA libraries were prepared using a KAPA HyperPrep kit (Roche, Indianapolis, IN, USA) according to the manufacturer's protocols and sequenced 2 × 150-bp on an NovaSeqX Plus 10B (Illumina, San Diego, CA, USA). One third of the RNA samples were treated with DNase due to evidence of DNA contamination found during QC. The final RNA Integrity Numbers (RIN) were all >7. All RNA samples were then rRNA depleted using a QIAseq FastSelect Epidemiology kit (for bacteria, humans, mouse, and rat rRNA) (Qiagen, Hilden, Germany), and RNA libraries were prepared using a NEB Ultra II Directional RNA Library Prep kit (New England Biolabs, Ipswich, MA, USA). RNA was sequenced 2 × 150-bp on an NovaSeqX Plus 10B (Illumina). Total reads sequenced equaled approximately 70 million paired-end reads per DNA or RNA sample. DNA and RNA sequencing reads are available in NCBI under BioProject PRJNA1232215.
Metagenomic assembly, binning, and annotation
Detailed information about assembly, binning, and genome annotation is described in the SI and example scripts used for analyses are provided at https://github.com/jadeaver/bioP_MAGs. Briefly, demultiplexed short reads received from Admera were quality checked with FastQC (v0.12.1).11 Raw long-read sequences were basecalled with Oxford Nanopore's basecaller Dorado using the super accurate model version 4.3.0 (https://github.com/nanoporetech/dorado), quality checked with Nanoplot (v1.42.0), and filtered using Filtlong (v0.2.1).12 Long reads were polished with short reads using Ratatosk (v0.9.0) and assembled using Flye (v.2.9.4).13,14 Each sample was individually assembled, samples collected on the same day were co-assembled, and all samples were co-assembled for a total of 13 assemblies (Table S2). Each assembly was binned using both maxbin2 (v2.2.7) and metabat2 (v2.15.2) accounting for differential coverage of all eight samples.15,16 Bins derived from the same assembly were dereplicated using DasTool (v1.1.7)17 and remaining bins were further dereplicated at a 95% average nucleotide identity (ANI) threshold across all assemblies using dRep (v3.5.0) to select the best quality representatives at approximately the species-level.18 Genomes were preliminarily classified using the GTDB-Tk classify workflow (v2.4.0), quality checked with CheckM (v1.1.6) and annotated with Bakta (v1.9.4) using the full Bakta database (v5.1).19–21Phylogenomic analyses and relative abundance
Reference genomes from Petriglieri et al.22 were downloaded using NCBI datasets and phylogenomic trees created with GToTree (v1.8.8) using the Proteobacteria HMM profiles.23 More details are provided in the SI. FastANI (v1.34) was used to perform pairwise average nucleotide identity (ANI) comparisons between reference genomes and PAO MAGs assembled during this study to assign taxonomy.24 Taxonomic profiles of individual metagenomes and MAG recovery were assessed using SingleM (v0.18.3).25 The vegan R package was used to calculate β-diversity based on Bray–Curtis distances, using the SingleM genus-level relative abundance table as input.26 A subsequent PERMANOVA test was performed on the calculated distances based on date (n = 2 per date) and basin (n = 3 per basin). Scripts used for these analyses are available at https://github.com/jadeaver/bioP_MAGs.Metagenome-enabled transcriptomics
Detailed information about gene expression analyses is described in the SI, and scripts used for analyses are provided at https://github.com/jadeaver/bioP_MT. Briefly, RNA reads were quality checked using FastQC, adapters removed using fastp (v0.23.4), and rRNA sequences filtered out with SortMeRNA (v4.3.7) leaving 30–45 million paired end reads per sample remaining.27,28 When possible, MAGs recovered from this study were used as references for transcript quantification. For Ca. Accumulibacter species identified as present in the metagenomes but not recovered during binning, species-level representative MAGs (as defined by Petriglieri et al.22) were used as references. The predicted coding regions were concatenated together to create a mapping index of Ca. Accumulibacter MAGs. Quality processed metatranscriptomic reads from all samples were competitively pseudoaligned to the mapping index and quantified with kallisto (v0.51.0).29 DESeq2 was used for statistical analyses of differentially expressed genes.30 Multiple contrasts were performed between groups defined as the zone and day (i.e., “Anaerobic 11/20/23”, “Anoxic 12/6/23”, etc.) and the results are reported here. As a note, multiple contrasts were performed between location groups defined as the exact sample location and day (i.e., “Location B 11/20/23”, “Location D 12/6/23”, etc.) to examine how considering precise location versus multiple sample locations within the same zone impact results. The results were consistent with each other, and therefore, we present the data from zone/day groups here.Results and discussion
Full-scale EBPR performance
The facility experienced a winter instability event during which peak effluent TP concentrations exceeded 3 mg P L−1 (Fig. 2A). Any measurement over 1 mg P L−1 was considered unstable performance; thus, this event lasted ∼7 days. Table 1 summarizes facility performance during the study period. Notably, the temperature decreased by approximately 3.4 °C from mid-November to mid-December. Supplemental juice waste and acetic acid were added to support EBPR during this period. In response to the instability, alum was added after the final clarifiers and before the sand bed filters from 11/28/23 to 11/30/23, resulting in effluent TP concentrations returning to <0.5 mg P L−1. Alum did not impact orthophosphate concentrations measured in the anaerobic, anoxic, and aerobic zones because it was not recycled back into the biological process. End-of-anaerobic-zone orthophosphate concentrations averaged 25.1 + 10.6 mg P L−1 from November 13 to December 15 (Fig. 2B). Orthophosphate concentrations measured at the end of the anoxic zone for the same time averaged 8.1 + 2.6 mg P L−1 and remained relatively consistent throughout the instability event. End-of-aerobic-zone orthophosphate concentrations measured <0.5 mg P L−1 during stable operation and exceeded 1 mg P L−1, peaking around 3 mg P L−1, during instability.| Week of | Influent TPa | Influent BODa | BOD:P Ratio | Influent pH | Influent Ammoniaa | Temperature | MLSS Basin 2b | MLSS Basin 3b | HRT | RAS Nitrateb | Effluent TPa | Effluent Ammoniac | Effluent Nitratec | Effluent Nitritec |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Units | mg-P L−1 | mg L−1 | — | SU | mg-N L−1 | °C | mg L−1 | mg L−1 | Days | mg-N L−1 | mg-P L−1 | mg-N L−1 | mg-N L−1 | mg-N L−1 |
| a Composite sample measured by a certified lab. b Grab sample measured by an operator. c ChemScan measurement. | ||||||||||||||
| Nov. 13 | 9.02 ± 0.64 | 251.9 ± 14.6 | 18 | 7.14 ± 0.56 | 32.92 ± 3.27 | 21.7 ± 0.99 | 3572 ± 168 | 3437 ± 144 | 0.55 ± 0.06 | 7.12 ± 0.93 | 0.24 ± 0.10 | 0.54 ± 0.01 | 11.06 ± 0.47 | 0.08 ± 0.00 |
| Nov. 20 | 7.84 ± 3.99 | 291.1 ± 10.0 | 37 | Not available | 34.04 ± 3.49 | 20.2 ± 0.55 | 3470 ± 192 | 3366 ± 277 | 0.56 ± 0.08 | 6.80 ± 2.38 | 0.69 ± 0.80 | 0.53 ± 0.01 | 11.38 ± 0.03 | 0.08 ± 0.00 |
| Nov. 27 | 9.63 ± 1.78 | 284.2 ± 19.2 | 30 | 7.03 ± 0.01 | 34.00 ± 4.56 | 19.2 ± 1.02 | 3310 ± 248 | 3174 ± 231 | 0.56 ± 0.02 | 7.21 ± 1.58 | 1.30 ± 1.18 | 0.54 ± 0.00 | 11.06 ± 0.26 | 0.08 ± 0.00 |
| Dec. 4 | 9.84 ± 3.90 | 231.3 ± 78.8 | 24 | 6.92 ± 0.07 | 34.39 ± 6.56 | 19.3 ± 0.89 | 3221 ± 305 | 3167 ± 379 | 0.50 ± 0.08 | 8.14 ± 0.98 | 0.46 ± 0.20 | 0.54 ± 0.00 | 10.88 ± 0.23 | 0.08 ± 0.00 |
| Dec. 11 | 7.07 ± 0.48 | 289.9 ± 152.6 | 41 | 6.88 ± 0.02 | 31.50 ± 4.39 | 18.3 ± 0.37 | 3423 ± 236 | 3344 ± 122 | 0.53 ± 0.02 | 7.23 ± 2.37 | 0.55 ± 0.46 | 0.53 ± 0.01 | 10.76 ± 0.33 | 0.08 ± 0.00 |
MAG assembly and classification
In total, 53 MAGs were recovered after species-level dereplication at 95% ANI. Twenty-two were high quality (HQ) as per MIMAG standards with >90% completion, <5% redundancy, and the presence of 23S, 16S, 5S rRNA genes and >18 tRNA genes (Table S3).31 The remaining MAGs were medium quality (MQ) with >75% completion and <10% redundancy, and most included the presence of all rRNA genes and >18 tRNA genes. Of these, six MAGs (two HQ and four MQ) were taxonomically classified as PAOs belonging to either Ca. Accumulibacter or Azonexus lineages according to GTDB-Tk classification.To further classify the PAO MAGs, we performed phylogenomic analyses to compare the four Ca. Accumulibacter MAGs reported here with the species representatives defined by Petriglieri et al.22 (Fig. S2A, Table S4). Ca. Accumulibacter lineages have been divided into 18 clades based on the comparison of their ppk1 genes. Genome-based comparisons accurately reflect ppk1-based phylogeny.22,32 Pairwise ANI comparisons and phylogenomic tree construction based on Proteobacteria hidden Markov model (HMM) profiles indicated that MAG 010, MAG 12, and MAG 11 belong to the species Ca. Accumulibacter phosphatis (clade IIA), Ca. Accumulibacter propinquus (clade IIB), and Ca. Accumulibacter conexus (clade IIF), respectively. MAG 166 likely belongs to a new species cluster, Ca. Accumulibacter UW 21, proposed by Stewart et al.32 The ANI value between MAG 166 and Ca. Accumulibacter UW 21 is 97.1%, suggesting that MAG 166 tracks to the same species as UW 21.
GTDB-based classifications indicated two MAGs, MAG 109 and MAG 311, belonging to the Azonexus lineage. Pairwise ANI comparisons indicated that neither Azonexus MAG belonged to either Azonexus phosphoritrophus or Azonexus phosphorivorans, the two Azonexus species whose contributions to EBPR have been best characterized.33 Pairwise ANI comparisons to GTDB Azonexus representative MAGs ranged from 78% to 99% with MAG 109 most closely matching Azonexus sp009469585 (GCA_009469585.1) at 92% ANI and MAG 311 most closely matching Azonexus sp020621865 (GCA_020621865.1) at 99% ANI (Fig. S2B, Table S5). Further analyses are necessary to determine what role, if any, these Azonexus species play in phosphorus removal. Considering their low abundance and activity (described below), we did not further investigate these MAGs.
Community composition and transcriptional profiles
To estimate overall microbial community relative abundance and to ensure that the most abundant PAOs were represented by the recovered MAGs, we profiled the raw metagenomes with SingleM. Genus-level profiles demonstrated that Ca. Accumulibacter was the most abundant genus on all days with an average relative abundance of 6.8 + 0.2%, 5.8 + 0.1%, and 6.1 + 0.1% on 11/20/23, 11/29/23, and 12/6/23, respectively (Fig. 3A). Approximately 25% of the community was not classified at the genus level (data not shown), and approximately 30% of the community was represented by a genus with a relative abundance of 1% or more. A diverse, highly redundant microbial community where few individuals dominate is typical of full-scale activated sludge communities. In this study, β-diversity was calculated based on Bray–Curtis distances. A subsequent PERMANOVA test performed on the calculated distances based on date (n = 2 per date) and basin (n = 3 per basin) groupings revealed that date had a greater effect than basin (R2 = 0.829 for date versus R2 = 0.171 for basins) but that overall genus-level community composition changes were not significant over the instability event (p-value = 0.067) (Fig. S3). The community results suggest that unstable phosphorus removal was not caused by changes in genus-level microbial community structure.To explore if changes in PAO population structure were linked to the instability, species-level profiles of Ca. Accumulibacter, Azonexus, and Ca. Phosphoribacter were further assessed using SingleM output. There were nine Ca. Accumulibacter and one Azonexus species each comprising more than 0.25% of the total community composition in at least one sample (Fig. 3B). Ca. Accumulibacter phosphatis and Ca. Accumulibacter propinquus were identified in the highest proportions, comprising about 1/3 and 1/5 of the PAO community, respectively, across all sampling dates. Ca. Accumulibacter contiguus, Ca. Accumulibacter cognatus, Ca. Accumulibacter vicinus, and Ca. Accumulibacter necessarius were identified in the raw metagenomes, but MAGs were not recovered.
The four Ca. Accumulibacter MAGs recovered in this study and the species representative MAGs22 for Ca. Accumulibacter contiguus, Ca. Accumulibacter cognatus, Ca. Accumulibacter vicinus, and Ca. Accumulibacter necessarius were used as reference genomes for further metatranscriptomic analysis. Metatranscriptomic profiling of Ca. Accumulibacter species demonstrated that the most abundant Ca. Accumulibacter MAGs also contributed the most to the transcriptional profile of the Ca. Accumulibacter community with 44–52% of RNA reads mapping to Ca. Accumulibacter phosphatis (MAG 010) or Ca. Accumulibacter propinquus (MAG 12) (Fig. 3C). The lack of changes in composition or transcript profiles of the PAO populations indicated that shifts in microbial community and activity did not contribute to the instability event. The facility's SRT is 10 days, so if a shift did occur, we would expect it to be captured between the first and last samples collected 16 days apart. While this study suggests that community composition changes did not contribute to the instability event, a study by Lindner et al.10 demonstrated a microbial community shift in connection with an EBPR instability event. Microbial community composition shifts might occur in response to an instability, rather than being the cause of the instability itself. Further investigations into instability events at other facilities are necessary to confirm if microbial community composition is more likely to be a lagging indicator of instability, rather than a leading indicator.
Gene expression profiles across redox zones
We performed differential gene expression analyses on Ca. Accumulibacter phosphatis and Ca. Accumulibacter propinquus, the two PAOs with the highest relative abundance and greatest contribution to the PAO transcript profile. DESeq2 analyses between redox zones on 11/29/23 (unstable EBPR) and 12/6/23 suggested that most genes were differentially expressed in the anoxic zone between 11/29/23 (unstable EBPR) and 11/20/23 (Fig. S4). To further explore this phenomenon, we used DESeq2 to compare the number of differentially (log2[FoldChange] >1 or < −1) and significantly (adjusted p-value <0.01) expressed genes across redox zones on each sampling day. For Ca. Accumulibacter phosphatis, before the instability, there were 80 significantly (adjusted p-value <0.01) differentially expressed genes with a log2(FoldChange) >1 or <−1 between the anaerobic and aerobic zones, 38 between the anaerobic and anoxic zones, and only one between the anoxic and aerobic zones (Table 2, Fig. 4). During the instability event, the reverse was true. There were 68 significantly differentially expressed genes with a log2[FoldChange] >1 or <−1 between the anoxic and aerobic zone, but only 11 between the anaerobic and anoxic zones and 1 between the anaerobic and aerobic zones. Ca. Accumulibacter propinquus also exhibited the same gene expression patterns (Fig. S5).| Anaerobic vs. anoxic | Anoxic vs. aerobic | Anaerobic vs. aerobic | |||||||
|---|---|---|---|---|---|---|---|---|---|
| 11/20/23 | 11/29/23 | 12/6/23 | 11/20/23 | 11/29/23 | 12/6/23 | 11/20/23 | 11/29/23 | 12/6/23 | |
| Number of DEG with the adjusted p-value <0.01 | 74 | 14 | 66 | 1 | 80 | 0 | 129 | 1 | 160 |
| Median |L2FC| for DEG with the adjusted p-value < 0.01 | 1.02 | 1.3 | 0.95 | — | 1.60 | — | 1.13 | — | 1.06 |
| Number of DEG with the adjusted p-value < 0.01 and |L2FC| > 1 | 38 | 11 | 29 | 1 | 68 | 0 | 80 | 1 | 89 |
Many genes differentially expressed between the anaerobic and aerobic zones on days 11/20/23 and 12/6/23 were the same genes differentially expressed between the anoxic and aerobic zones on day 11/29/23 (unstable EBPR) (Fig. 5A). Cyclical gene expression has been identified in studies of Ca. Accumulibacter-enriched SBRs, and we found that many of the genes whose expression patterns changed during the instability event deviated from typical patterns identified in lab-scale studies and observed during stable phosphorus removal. For example, Oyserman et al.34 performed metatranscriptomic analyses on a SBR enriched in Ca. Accumulibacter operating under alternating anaerobic/aerobic, feast/famine conditions and identified various expression trends (e.g., upregulation during the redox transition) of genes involved in carbon metabolism, denitrification, and energy production. Genes involved in those same metabolic pathways exhibited cyclical gene expression patterns in this study; however, the timing of their expression shifted during the instability. Genes exhibited peak expression in the anoxic zone, rather than the anaerobic zone, during the instability (Fig. 5B).
Genes exhibiting a shifted expression pattern included several Ca. Accumulibacter phosphatis carbon metabolism genes. These genes were differentially expressed in samples obtained from the anaerobic zone (compared to the aerobic zone) on days 11/20/23 and 12/6/23. Those same genes were differentially expressed in the anoxic zone (compared to the aerobic zone), rather than the anaerobic zone, on day 11/29/23 (unstable EBPR). These genes included PHA synthesis related genes such as phaC and phaR and methylmalonyl mutase genes. For example, the gene phaC was differentially expressed between the anaerobic and aerobic zones on 11/20/23 (log2[FoldChange] = 1.3; adjusted p-value = 2 × 10−5). However, on 11/29/23 (unstable EBPR), it was not significantly differentially expressed between the anaerobic and aerobic zones but rather between the anoxic and aerobic zones (log2[FoldChange] = 2.1; adjusted p-value = 7.6 × 10−7) (Fig. 5). The phaC gene encodes a subunit of PHA synthase, and its anaerobic expression is considered a hallmark feature of Ca. Accumulibacter during phosphorus release.6,34,35 In Oyserman et al.,34 PHA synthesis genes phaC and phaR were upregulated under high acetate, anaerobic conditions. The genes for methylmalonyl mutase and its associated GTPase also exhibited peak expression in the anoxic zone rather than the anaerobic zone on 11/29/23 (unstable EBPR) exhibiting log2[FoldChange] = 0.97 (adjusted p-value = 0.004) and 1.1 (adjusted p-values = 0.003), respectively. Methylmalonyl mutase and its associated GTPase are involved in the conversion of succinyl-CoA to methylmalonyl-CoA in the split mode TCA cycle utilizing the methylmalonyl-CoA pathway. Various metabolic models exist to explain how Ca. Accumulibacter consumes and stores carbon while balancing energy and reducing equivalents, including the full TCA cycle, the split mode TCA cycle, and the glyoxylate shunt models. It is likely that Ca. Accumulibacter possess metabolic flexibility that enables them to thrive under dynamic conditions.7Ca. Accumulibacter is suggested to use the methylmalonyl-CoA pathway in particular during anaerobic PHA synthesis from mixed carbon sources, including the co-utilization of glutamate and acetate, aspartate, lactate, and succinate.36,37 Chen et al.37 noted greater anaerobic methylmalonyl-CoA mutase gene expression in variable carbon source anaerobic/aerobic batch tests performed on SBR enrichment cultures, suggesting its involvement in anaerobic PHA synthesis particularly under mixed carbon feed conditions. We cannot determine the actual pathways used by the Ca. Accumulibacter phosphatis populations in this study, but expression of the TCA cycle, glyoxylate shunt, and methylmalonyl-CoA pathway genes demonstrated capabilities for a dynamic response. The shift in expression dynamics of key carbon metabolism genes suggests that a disruption to carbon cycling under stable phosphorus removal occurred during the instability event.
Denitrification gene expression also changed during the instability event. Ca. Accumulibacter phosphatis (MAG 010) possessed genes encoding enzymes in every step of denitrification (Fig. 6). Denitrification potential has been reported for other Ca. Accumulibacter phosphatis MAGs.38 Both sets of nitrate reductase genes, napAB and narGH were annotated in the Ca. Accumulibacter phosphatis MAG recovered in this study. In addition, norB, encoding the nitric oxide large subunit, was annotated. The gene encoding the nitric oxide small subunit, norC, was not annotated, however, possibly due to gaps in assembly or annotation of the MAG. All annotated genes encoding denitrification enzymes, except for nirS, encoding nitrite reductase, were significantly upregulated in the anoxic zone compared to the aerobic zone on 11/29/23 (unstable EBPR) (log2[FoldChange] >1.5; adjusted p-value <0.004) (Fig. 5A). These genes exhibited a shift in expression where peak expression (in TPM) occurred in the anoxic zone instead of the anaerobic zone on 11/29/23 (Fig. S6). nirS gene expression stayed consistent between the anaerobic and anoxic zones on 11/29/23 rather than peaking in the anaerobic zone even though it was not identified as significantly differentially expressed by DESeq2 analysis. Previous studies exploring Ca. Accumulibacter gene expression cycles during anaerobic/aerobic (feast/famine) cycles have noted an increase in expression of denitrification genes after anaerobiosis.38,39 Stable EBPR on 11/20/23 was characterized by the same expression pattern, with denitrification gene expression peaking in the anaerobic zone samples, and a return to this expression pattern is noted on 12/6/23.
Energy metabolism genes important to denitrification also exhibited shifted gene expression cycles. Genes including several encoding respiratory chain components such as cytochrome b, cytochrome c proteins, cytochrome c biogenesis proteins, F0F1 ATP synthase genes, and electron transfer flavoproteins were significantly upregulated in the anoxic zone (vs. the aerobic zone) rather than the anaerobic zone (vs. the aerobic zone) on 11/29/23 (unstable EBPR) (adjusted p-value <0.01, log2[FoldChange] >1) (Fig. 5A). For example, the gene for the F0F1 ATP synthase B subunit was differentially expressed between the anaerobic and aerobic zones on 11/20/23 (log2[FoldChange] = 1.2; adjusted p-value = 6.1 × 10−8) and differentially expressed between the anoxic and aerobic zones on 11/29/23 (unstable EBPR) (log2[FoldChange] = 1.6; adjusted p-value = 6.1 × 10−7). Aerobic respiration and denitrification depend on the same core respiratory machinery and are thought to have co-evolved.40 Shared machinery includes NADH dehydrogenase (complex I), the cytochrome bc1 complex (complex III), and cytochrome c. The F0F1 ATP synthase genes encode the machinery for generating ATP from the proton motive force generated by the respiratory chain enabling energy yield from either final electron acceptor.41 For PAOs, pit transport of inorganic phosphate hydrolyzed from poly-P is thought to be the primary mode of proton motive force production that in turn drives carbon uptake (e.g., acetate via the ActP transporter) and ATP production via F0F1 ATP synthase.42 Previous studies suggest that clade IIA Ca. Accumulibacter cannot respire on nitrate despite possessing a nap operon.39,43 The presence and expression of narGH in addition to napAB suggest that nitrate respiration may be possible. The delayed upregulation of key denitrification and associated energy production genes suggests that a disruption of typical energy metabolism activity occurred during the instability event.
Ca. Accumulibacter propinquus (MAG 12) exhibited the same shifts in gene expression patterns of similar denitrification and energy metabolism associated genes (Fig. S7). Nitrate reductase gene napA was the only denitrification gene that exhibited a shift in significant differential gene expression (adjusted p-value <0.01, log2[Foldchange] >1). Ca. Accumulibacter propinquus did not contain annotations for nitric oxide reductase (norBC) and nitrous oxide reductase (nosZ) genes. Associated respiratory machinery genes exhibited the shift to anoxic upregulation, including cytochrome c, cbb3-type cytochrome c oxidase subunit I, and two F0F1 ATP synthase subunits. The cbb3-type cytochrome c oxidase subunit I is of note because of its high substrate affinity and tendency to be expressed under microaerobic conditions.44 Camejo et al.39 identified a full denitrification pathway in a clade IC Ca. Accumulibacter MAG (UW-LDO-IC) enriched in an anaerobic/microaerobic SBR. UW-LDO-IC gene expression patterns also suggested coregulation of denitrification and high oxygen affinity cbb3-type cytochrome genes with expression peaking at the anaerobic/microaerobic transition. Delayed expression of Ca. Accumulibacter propinquus genes typically expressed following anaerobic contact further demonstrating that PAO activity changes characterize the instability event.
Implications for full-scale EBPR stability
Our findings indicate that unstable phosphorus removal at the full-scale facility in this study was associated with changes in “what” the PAOs were doing rather than “who” was present. This finding is different from that described by Lindner et al.,10 wherein they found changes in PAO relative abundance between stable and unstable operation, in addition to changes in carbon metabolism gene expression. The major changes we observed were shifts in pathways associated with carbon utilization, denitrification, and energy metabolism across the redox zones. As a result, aerobic uptake of phosphorus was delayed and phosphorus concentrations leaving the aerobic zone were elevated compared to stable conditions.The shifted pathways illustrated in Fig. 6 are largely involved in carbon metabolism, denitrification, and bioenergetics, key metabolic pathways with implications for poly-P accumulation. Though we cannot definitively say what caused the changes in PAO activity, we hypothesize based on our results that either the redox conditions of the anaerobic zone conditions and/or influent VFA composition contributed. The shift in peak expression from the anaerobic to anoxic zone may have occurred due to limited anaerobic contact time to sufficiently induce expression of key carbon storage and energy metabolism genes. Redox conditions in the anaerobic zone are thought to be a significant contributing factor for successful EBPR. Achieving sufficiently low redox potential in the anaerobic zone can be limited due to high mixing energy that traps oxygen, low anaerobic zone SRTs, and introduction of flows with elevated dissolved oxygen (DO), such as from primary effluent during wet weather events.45 Changes in influent VFA composition may have also contributed. Many of the genes whose expression shifted were upregulated under high acetate conditions in lab-scale experiments, suggesting that changes in carbon availability may alter expression patterns.34 If anaerobic-zone carbon availability is insufficient or oxidized, genes that normally peak in the anaerobic zone will peak in the anoxic zone, coincident with EBPR deterioration. Influent VFA concentration and oxidation–reduction potential (ORP) measurements across redox zones are necessary to definitively determine if either of the proposed hypotheses regarding anaerobic disruption is correct. Future studies should evaluate influent VFA concentration and composition and profile basins for ORP, DO, and nitrate in parallel to microbial community and gene expression analyses.
Conclusions
This study evaluated a full-scale EBPR instability event using genome-centric metatranscriptomics and concluded that the instability was associated with changes in the activity rather than the composition of PAOs. The shift in peak expression of key carbon metabolism, denitrification, and energy production genes from the anaerobic zone to the anoxic zone strongly suggests that redox conditions and/or VFA availability in the anaerobic zone were compromised. Future studies of full-scale EBPR systems should couple metatranscriptomics with additional measurements, such as ORP, DO, and VFAs, to further develop the linkages necessary to understand the underlying causes of instability events.Conflicts of interest
There are no conflicts to declare.Data availability
Raw sequencing data and high-quality MAGs are available in NCBI under BioProject PRJNA1232215. Medium quality MAGs are available upon request. Code used for data analyses is available in the following GitHub repositories: https://github.com/jadeaver/bioP_MAGs (metagenomics code) and https://github.com/jadeaver/bioP_MT (metatranscriptomics code).Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ew01105a.
Acknowledgements
This work was supported by the STEPS Center, a US National Science Foundation Science and Technology Center (CBET-2019435). The authors are appreciative of discussions and collaborations with various team members of the Science and Technologies for Phosphorus Sustainability (STEPS) Science and Technology Center. We also acknowledge the computing resources provided by the North Carolina State University High-Performance Computing Services Core Facility (RRID:SCR_022168). We thank our WRRF partner for providing samples, process data, and their knowledge and expertise during this study.References
- A. W. Merck, J. A. Deaver and L. Crane, et al., Stakeholder Views of Science and Technologies for Phosphorus Sustainability: A Comparative Analysis, Soc. Nat. Resour., 2024, 37(11), 1528–1545, DOI:10.1080/08941920.2024.2389806.
- S. He, A. Z. Gu and K. D. McMahon, Progress Toward Understanding the Distribution of Accumulibacter Among Full-Scale Enhanced Biological Phosphorus Removal Systems, Microb. Ecol., 2008, 55(2), 229–236, DOI:10.1007/s00248-007-9270-x.
- J. B. Neethling, B. Bakke and M. Benisch, et al., Factors Influencing the Reliability of Enhanced Biological Phosphorus Removal, Water Environment Research Foundation, 2005, 5 Search PubMed.
- D. Wang, N. B. Tooker and V. Srinivasan, et al., Side-stream enhanced biological phosphorus removal (S2EBPR) process improves system performance - A full-scale comparative study, Water Res., 2019, 167, 115109, DOI:10.1016/j.watres.2019.115109.
- M. K. D. Dueholm, M. Nierychlo and K. S. Andersen, et al., MiDAS 4: A global catalogue of full-length 16S rRNA gene sequences and taxonomy for studies of bacterial communities in wastewater treatment plants, Nat. Commun., 2022, 13, 1908, DOI:10.1038/s41467-022-29438-7.
- E. A. McDaniel, F. Moya-Flores and N. K. Beach, et al., Metabolic Differentiation of Co-occurring Accumulibacter Clades Revealed through Genome-Resolved Metatranscriptomics, mSystems, 2021, 6(4), 1–17, DOI:10.1128/mSystems.00474-21.
- L. Guedes Da Silva, K. Olavarria Gamez and J. Castro Gomes, et al., Revealing the Metabolic Flexibility of “Candidatus Accumulibacter phosphatis” through Redox Cofactor Analysis and Metabolic Network Modeling, Appl. Environ. Microbiol., 2020, 86(24), e00808–e00820, DOI:10.1128/AEM.00808-20.
- Y. Law, R. H. Kirkegaard and A. A. Cokro, et al., Integrative microbial community analysis reveals full-scale enhanced biological phosphorus removal under tropical conditions, Sci. Rep., 2016, 6(1), 25719, DOI:10.1038/srep25719.
- M. V. Pérez, L. D. Guerrero, E. Orellana, E. L. Figuerola and L. Erijman, Time Series Genome-Centric Analysis Unveils Bacterial Response to Operational Disturbance in Activated Sludge, mSystems, 2019, 4(4), 1–16, DOI:10.1128/mSystems.00169-19.
- B. Lindner, B. Layton and B. Lindner, et al., Genome-centric Insights into Full-scale EBPR Captured During a Biomonitoring Campaign. In: Proceedings of the Water Environment Federation. Water Environment Federation, 2023, DOI:10.2175/193864718825159085.
- S. Andrews, FastQC: a quality control tool for high throughput sequence data. Babraham, Bioinformatics, 2010 Search PubMed , http://www.bioinformatics.babraham.ac.uk/projects/fastqc.
- W. De Coster and R. Rademakers, NanoPack2: population-scale evaluation of long-read sequencing data, Bioinformatics, 2023, 39(5), btad311, DOI:10.1093/bioinformatics/btad311.
- G. Holley, D. Beyter and H. Ingimundardottir, et al., Ratatosk: hybrid error correction of long reads enables accurate variant calling and assembly, Genome Biol., 2021, 22, 28, DOI:10.1186/s13059-020-02244-4.
- M. Kolmogorov, D. M. Bickhart and B. Behsaz, et al., metaFlye: scalable long-read metagenome assembly using repeat graphs, Nat. Methods, 2020, 17(11), 1103–1110, DOI:10.1038/s41592-020-00971-x.
- Y. W. Wu, B. A. Simmons and S. W. Singer, MaxBin 2.0: an automated binning algorithm to recover genomes from multiple metagenomic datasets, Bioinformatics, 2016, 32(4), 605–607, DOI:10.1093/bioinformatics/btv638.
- D. D. Kang, F. Li and E. Kirton, et al., MetaBAT 2: an adaptive binning algorithm for robust and efficient genome reconstruction from metagenome assemblies, PeerJ., 2019, 7, e7359, DOI:10.7717/peerj.7359.
- C. M. K. Sieber, A. J. Probst and A. Sharrar, et al., Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy, Nat. Microbiol., 2018, 3(7), 836–843, DOI:10.1038/s41564-018-0171-1.
- M. R. Olm, C. T. Brown, B. Brooks and J. F. Banfield, dRep: a tool for fast and accurate genomic comparisons that enables improved genome recovery from metagenomes through de-replication, ISME J., 2017, 11(12), 2864–2868, DOI:10.1038/ismej.2017.126.
- O. Schwengers, L. Jelonek, M. A. Dieckmann, S. Beyvers, J. Blom and A. Goesmann, Bakta: rapid and standardized annotation of bacterial genomes via alignment-free sequence identification, Microb, Genomics, 2021, 7, 000685, DOI:10.1099/mgen.0.000685.
- D. H. Parks, M. Imelfort, C. T. Skennerton, P. Hugenholtz and G. W. Tyson, CheckM: Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes, Genome Res., 2015, 25(7), 1043–1055, DOI:10.1101/gr.186072.114.
- P. A. Chaumeil, A. J. Mussig, P. Hugenholtz and D. H. Parks, GTDB-Tk v2: memory friendly classification with the genome taxonomy database, Bioinformatics, 2022, 38(23), 5315–5316, DOI:10.1093/bioinformatics/btac672.
- F. Petriglieri, C. M. Singleton and Z. Kondrotaite, et al., Reevaluation of the Phylogenetic Diversity and Global Distribution of the Genus “Candidatus Accumulibacter.”, mSystems, 2022, 7(3), e00016–e00022, DOI:10.1128/msystems.00016-22.
- M. D. Lee, GToTree: a user-friendly workflow for phylogenomics, Bioinformatics, 2019, 35(20), 4162–4164, DOI:10.1093/bioinformatics/btz188.
- C. Jain, L. M. Rodriguez-R, A. M. Phillippy, K. T. Konstantinidis and S. Aluru, High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries, Nat. Commun., 2018, 9(1), 5114, DOI:10.1038/s41467-018-07641-9.
- B. J. Woodcroft, S. T. N. Aroney and R. Zhao, et al., Comprehensive taxonomic identification of microbial species in metagenomic data using SingleM and Sandpiper, Nat. Biotechnol., 2025 DOI:10.1038/s41587-025-02738-1.
- J. Oksanen, F. G. Blanchet and M. Friendly, et al., vegan: Community Ecology Package. Published online, 2019, https://cran.r-project.org/package=vegan.
- S. Chen, Y. Zhou, Y. Chen and J. Gu, fastp: an ultra-fast all-in-one FASTQ preprocessor, Bioinformatics, 2018, 34(17), i884–i890, DOI:10.1093/bioinformatics/bty560.
- E. Kopylova, L. Noé and H. Touzet, SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data, Bioinformatics, 2012, 28(24), 3211–3217, DOI:10.1093/bioinformatics/bts611.
- N. L. Bray, H. Pimentel, P. Melsted and L. Pachter, Near-optimal probabilistic RNA-seq quantification, Nat. Biotechnol., 2016, 34(5), 525–527, DOI:10.1038/nbt.3519.
- M. I. Love, W. Huber and S. Anders, Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2, Genome Biol., 2014, 15, 550, DOI:10.1186/s13059-014-0550-8.
- R. M. Bowers, N. C. Kyrpides and R. Stepanauskas, et al., Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea, Nat. Biotechnol., 2017, 35(8), 725–731, DOI:10.1038/nbt.3893.
- R. D. Stewart, K. S. Myers, C. Amstadt, M. Seib, K. D. McMahon and D. R. Noguera, Refinement of the “Candidatus Accumulibacter” genus based on metagenomic analysis of biological nutrient removal (BNR) pilot-scale plants operated with reduced aeration, mSystems, 2024, 9(3), 1–25, DOI:10.1128/msystems.01188-23.
- F. Petriglieri, C. Singleton, M. Peces, J. F. Petersen, M. Nierychlo and P. H. Nielsen, “Candidatus Dechloromonas phosphoritropha” and “Ca. D. phosphorivorans”, novel polyphosphate accumulating organisms abundant in wastewater treatment systems, ISME J., 2021, 15(12), 3605–3614, DOI:10.1038/s41396-021-01029-2.
- B. O. Oyserman, D. R. Noguera, T. G. Del Rio, S. G. Tringe and K. D. McMahon, Metatranscriptomic insights on gene expression and regulatory controls in Candidatus Accumulibacter phosphatis, ISME J., 2016, 10(4), 810–822, DOI:10.1038/ismej.2015.155.
- S. He and K. D. McMahon, Candidatus Accumulibacter gene expression in response to dynamic EBPR conditions, ISME J., 2011, 5(2), 329–340, DOI:10.1038/ismej.2010.127.
- G. Qiu, X. Liu and N. M. M. T. Saw, et al., Metabolic Traits of Candidatus Accumulibacter clade IIF Strain SCELSE-1 Using Amino Acids As Carbon Sources for Enhanced Biological Phosphorus Removal, Environ. Sci. Technol., 2020, 54(4), 2448–2458, DOI:10.1021/acs.est.9b02901.
- L. Chen, G. Wei and Y. Zhang, et al., Candidatus Accumulibacter use fermentation products for enhanced biological phosphorus removal, Water Res., 2023, 246, 120713, DOI:10.1016/j.watres.2023.120713.
- E. A. McDaniel, J. J. M. Van Steenbrugge and D. R. Noguera, et al., TbasCO: trait-based comparative ‘omics identifies ecosystem-level and niche-differentiating adaptations of an engineered microbiome, ISME Commun., 2022, 2(1), 1–20, DOI:10.1038/s43705-022-00189-2.
- P. Y. Camejo, B. O. Oyserman, K. D. McMahon and D. R. Noguera, Integrated Omic Analyses Provide Evidence that a “Candidatus Accumulibacter phosphatis” Strain Performs Denitrification under Microaerobic Conditions, mSystems, 2019, 4(1), 1–23, DOI:10.1128/mSystems.00193-18.
- J. Chen and M. Strous, Denitrification and aerobic respiration, hybrid electron transport chains and co-evolution, Biochim. Biophys. Acta, Bioenerg., 2013, 1827(2), 136–144, DOI:10.1016/j.bbabio.2012.10.002.
- T. V. Zharova, V. G. Grivennikova and V. B. Borisov, F1·Fo ATP Synthase/ATPase: Contemporary View on Unidirectional Catalysis, Int. J. Mol. Sci., 2023, 24(6), 5417, DOI:10.3390/ijms24065417.
- A. M. Saunders, A. N. Mabbett, A. G. McEwan and L. L. Blackall, Proton motive force generation from stored polymers for the uptake of acetate under anaerobic conditions, FEMS Microbiol. Lett., 2007, 274(2), 245–251, DOI:10.1111/j.1574-6968.2007.00839.x.
- J. J. Flowers, S. He, S. Yilmaz, D. R. Noguera and K. D. McMahon, Denitrification capabilities of two biological phosphorus removal sludges dominated by different ‘Candidatus Accumulibacter’ clades, Environ. Microbiol. Rep., 2009, 1(6), 583–588, DOI:10.1111/j.1758-2229.2009.00090.x.
- R. S. Pitcher and N. J. Watmough, The bacterial cytochrome cbb3 oxidases, Biochim. Biophys. Acta, 2004, 1655, 388–399, DOI:10.1016/j.bbabio.2003.09.017.
- J. L. Barnard, P. Dunlap and M. Steichen, Rethinking the Mechanisms of Biological Phosphorus Removal, Water Environ. Res., 2017, 89(11), 2043–2054, DOI:10.2175/106143017X15051465919010.
Footnote |
| † Hazen and Sawyer, 4011 Westchase Blvd, Suite 500, Raleigh, NC 27607. |
| This journal is © The Royal Society of Chemistry 2026 |