Development of models and a tool (DBPCytoGenoTOX Predictor) for predicting the cytotoxicity and genotoxicity of disinfection byproducts
M. Y.
Liu
,
H. H.
Liu
 and
X. H.
Yang
and
X. H.
Yang
 *
*
School of Environmental and Biological Engineering, Nanjing University of Science and Technology, Nanjing 210094, China. E-mail: xhyang@njust.edu.cn
First published on 25th November 2025
Abstract
Disinfection byproducts (DBPs) have been receiving global attention because they may have detrimental effects on organisms. Cytotoxicity and genotoxicity are two toxicity endpoints that are of wide concern in the field of DBP hazard assessment. Hitherto, only around one hundred out of thousands of identified DBPs had available experimental cytotoxicity and genotoxicity data. It is important to fill this data gap of thousands of DBPs by employing an efficient and high-throughput method. Herein, we first summarized the extensive and heterogeneous cytotoxicity (184-DBPs) and genotoxicity (105-DBPs) data sets related to DBPs. Then, quantitative and qualitative models with acceptable internal and external prediction performance were developed for cytotoxicity and genotoxicity, respectively. Next, a user-friendly tool named “DBPCytoGenoTOX Predictor” was developed using the optimal models. This tool was further applied to fill the missing cytotoxicity and genotoxicity data gaps of an updated DBP inventory with 1816 substances. Finally, the high-priority DBPs were screened from the DBP inventory based on the experimental and predicted cytotoxicity and genotoxicity data as well as the previously reported endocrine-disrupting effects and aquatic toxicity data. As a result, 385 high-priority DBPs were identified. More efforts should be taken to confirm the potential adverse effects of these high-priority DBPs on organisms in the future.
Environmental significanceAs a group of toxic emerging chemicals (ECs) that exist widely in disinfected drinking water and wastewater, disinfection byproducts (DBPs) have been drawing global attention. Among the more than 6000 identified DBPs, only a very small proportion have available experimental health and ecological toxicity data. This has limited the ability of authorities to screen the DBPs of high concern and take appropriate management measures for high-priority DBPs. In response to these issues, we selected the cytotoxicity and genotoxicity of DBPs as an example and developed the corresponding models and a tool for assessing the cytotoxicity and genotoxicity of DBPs. The study provides a tool for filling the missing cytotoxicity and genotoxicity data gap of DBPs efficiently and with high-throughput. |
1 Introduction
Water disinfection using chlorine, chloramines, ozone, etc. has become one of the indispensable units in modern water treatment systems,1,2 dramatically reducing the outbreaks of historically common waterborne diseases and observably improving the longevity and living quality of the population.3,4 However, besides killing pathogenic microorganisms, disinfectants can also react with natural organic matter (NOM) or anthropogenic organic substances (e.g. organic pollutants), and/or halides in water, and form various disinfection byproducts (DBPs).5,6 Apart from drinking water disinfection, other processes such as swimming pool, indoor and outdoor disinfection, making tea, and even daily cooking have been found to generate DBPs,7–14 which means that the population may be continuously exposed to various DBPs in their daily lives. Thus, it is vital to distinguish which DBPs could elicit deleterious health and/or ecological effects.To date, extensive epidemiology and animal testing have revealed that DBPs might cause diverse cancers (e.g. bladder and rectum cancers), malformation or miscarriages in humans,15–18 induce cytotoxicity, genotoxicity, mutagenicity, etc. in rodents, and elicit harmful effects on aquatic organisms (e.g. fish).3,19–23 We conducted a cluster analysis based on the keywords of available references from the Web of Science to illustrate the research focus in the field of DBP hazard assessment (Text S1 and Fig. 1). The analysis results show that cytotoxicity and genotoxicity are two toxicity endpoints that are of wide concern because many articles are related to cytotoxicity (#2) and genotoxicity (#3). Although the potential cytotoxicity and genotoxicity of DBPs have been the focus of extensive investigation and the corresponding test protocols, i.e. the Chinese hamster ovary (CHO) cell-based cytotoxicity and genotoxicity assay, have been established for several decades,24 DBPs with available cytotoxicity and genotoxicity data remain limited.25 For instance, Wagner and Plewa summarized the cytotoxicity and genotoxicity data up to 2017,26 and they found that there were 98 and 62 CHO cell-based cytotoxicity and genotoxicity data available at that time, respectively. Nevertheless, about 1187 DBPs (all contained CAS numbers and SMILES code) and 6310 DBPs (610 of them have CAS numbers and SMILES code) were reported recently by Sui et al.27 and Chen et al.,28 respectively, indicating that the number of DBPs with available cytotoxicity and genotoxicity data only accounts for a relatively small proportion of the total identified DBPs. The huge data gap of toxicity data related to DBPs has restricted the systematic assessment of DBPs, resulting in only 14 and 11 DBPs being regulated by the authorities of China and U.S., respectively (Table S1).3,29 Thus, efficient and high-throughput methods should be employed to fill the data gap of the thousands of DBPs so as to perform a systematic hazard assessment of the DBPs, and screen priority substances of high concern so that appropriate management action can be taken.
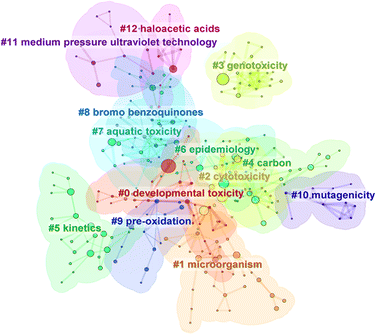 | ||
| Fig. 1 Cluster analysis results related to the keywords of available references focus on the DBPs hazard assessment. A lower label number indicates that the corresponding group is more important. CiteSpace (v.6.3R1 (64-bit)) was employed to perform the cluster analysis.30 The references were retrieved from the Web of Science database using “disinfection byproduct toxicity” as the topic. | ||
Computational toxicology models and tools can be used to fill the data gap efficiently and with high throughput.31–34 Available models related to the cytotoxicity and genotoxicity of DBPs are summarized and listed in Text S2, Tables S2, and S3. As shown, many teams have attempted to develop models for predicting the cytotoxicity and genotoxicity of DBPs, and their models are also of great quality.35–38 However, it is worth pointing out that most of those models only focus on a small number of DBP data sets that belong to one DBP subgroup or several numbered DBP subgroups, which may limit the application of those models to predict the cytotoxicity and genotoxicity of other DBP subgroups (Table S2). In addition, most of those models were not integrated into any tool, which hinders their further use by others. Furthermore, although some existing software contained genotoxicity models on CHO cells, the models did not consider the genotoxicity data from Single Cell Gel Electrophoresis (comet) Assays, which are generally employed in the research field of DBP hazard assessment (Table S3). Considering that the predictive ability and applicability domain of models depend on the quality and number of model compounds used in the model development process,5 new cytotoxicity and genotoxicity models and tools for DBP studies should be further constructed by employing extensive and heterogeneous cytotoxicity and genotoxicity data sets to extend the model's scope of application.
Hence, the primary purpose of this study is to derive new cytotoxicity and genotoxicity models and tools with more extensive and heterogeneous cytotoxicity and genotoxicity data sets, and to fill the missing cytotoxicity and genotoxicity data gap of thousands of identified DBPs. First, we performed an in-depth literature survey to obtain the extensive and heterogeneous cytotoxicity and genotoxicity data sets; second, the cytotoxicity and genotoxicity models were constructed by employing typical machine learning algorithms. Then, a tool was developed based on the optimal cytotoxicity and genotoxicity models. In addition, the cytotoxicity and genotoxicity data gap of the DBP inventory with 1816 substances was filled by employing the tool. Finally, the high-priority DBPs were identified, and their current control status worldwide was assessed.
2 Materials and methods
2.1 Cytotoxicity and genotoxicity data sets and the DBP inventory
In 2017, Wagner and Plewa reviewed and reported the CHO cell-based cytotoxicity and genotoxicity data sets with 98 and 62 DBPs, respectively.26 To obtain more comprehensive CHO cell-based cytotoxicity and genotoxicity data sets of DBPs, we performed an in-depth literature survey using the Web of Science (https://webofscience.clarivate.cn/wos/woscc/basic-search) with several keywords, including but not limited to “DBP toxicity CHO”, “DBP cytotoxicity”, “DBP genotoxicity”, and “DBP toxicity mammalian”. All data was checked by tracing back to the original articles. In the data collection process, the following terms for each data were recorded, i.e. the basic identifying information of a given DBP (e.g. chemical name, CAS NO, SMILES code, molecular weight), cell line, experimental method, exposure duration, 50% lethal concentration (LC50) and/or 50% Tail DNA (TDNA or midpoint of Tail moment) values. The experimental methods used to determine the toxicity data incorporated into the dataset (LC50 for cytotoxicity and TDNA for genotoxicity) were detailed by Wagner et al.26 The CAS NO and SMILES code of these DBPs were obtained from CAS Finder (https://scifinder-n.cas.org), OECD QSAR Toolbox v4.7 (https://qsartoolbox.org/download) and/or PubChem (https://pubchem.ncbi.nlm.nih.gov). The duplicate substances, substances with no data (i.e. data marked as “Not Available”), and mixtures were carefully cross-checked and treated. Multiple verifications for each substance in the raw database of cytotoxicity and genotoxicity were carried out to ensure there were no duplicate substances. Mixtures were removed. The final cytotoxicity and genotoxicity data sets are presented in Tables S4 and S5, respectively. All references to the toxicity data are listed after the corresponding tables. In the modelling, the final cytotoxicity and genotoxicity data sets were randomly split into training and validation sets, respectively.Usually, the cytotoxicity and genotoxicity data obtained from the literature were based on molar concentrations. However, mass concentrations are commonly used in the case of assignment to categories of hazardous substances. For example, the cytotoxicity and genotoxicity could be divided into the following 5 hazardous categories based on the corresponding mass concentration of cytotoxicity and genotoxicity values:39 relatively harmless (>1000 mg L−1), low toxicity (100–1000 mg L−1), moderately toxic (10–100 mg L−1), highly toxic (1–10 mg L−1), extremely toxic (≤1 mg L−1).
To fill the cytotoxicity and genotoxicity data gap of known DBPs with unequivocal CAS NO and SMILES codes, the DBP inventory containing specific CAS NO and SMILES codes was updated. The nominated DBPs were obtained from three sources, namely (a) 1187 DBPs summarized by our lab;27 (b) 580 DBPs with CAS NO from the DBPs list reported by Yang Xiaoqiu's lab of Jianghan University;28 (c) 49 DBPs with CAS NO searched from the Web of Science (https://webofscience.clarivate.cn/wos/woscc/basic-search) or combined from the cytotoxicity and genotoxicity data sets. The updated DBP inventory contains 1816 DBPs (Table S6). To our knowledge, this is the largest DBP inventory with explicit identifying information, i.e. CAS NO and SMILES code, available to date.
2.2 Calculation of molecular descriptors
We imported the substances into the PaDEL Descriptors tool using SMILES code as the input term to obtain their 1D and 2D molecular descriptors and Pubchem Fingerprints.40 The 1D and 2D molecular descriptors and Pubchem Fingerprints were cleaned by dislodging parameters with missing values and removing parameters with constant values as well as with cross-correlations >0.95.41 In total, 770 and 705 PaDEL descriptors were retained in the later cytotoxicity and genotoxicity modelling process, respectively.2.3 Machine learning modelling
Initially, we attempted to derive the quantitative predictive models for both cytotoxicity and genotoxicity. However, no acceptable quantitative models could be obtained for genotoxicity (Table S7). Thus, we further developed qualitative predictive models for genotoxicity. The machine learning (ML) algorithms used to establish the quantitative models included the multiple linear regression (MLR), k nearest neighbor (kNN), multi-layer perceptron neural network (MLP), decision tree (DT), gradient boosting decision tree (GBDT), random forest (RF), and eXtreme gradient boosting (XGBoost). The algorithms employed to construct the qualitative models included kNN, Logistic Regression (LR), support vector machine (SVM), DT, RF, and GBDT. In the quantitative and qualitative modelling, the StandardScaler method was used to scale the descriptors. The forward stepwise approach, belonging to one of the wrapper-based feature selection methods, was employed to choose the final critical predictive variables.41 The hyperparameters of the quantitative and qualitative models were searched using the grid search method as well as 5-fold cross-validation. The importance of predictive variables was illustrated by the Shapley additive explanation (SHAP).42 The statistical parameters related to goodness-of-fit, robustness, and predictive ability were calculated to characterize the internal and external predictive performance. The Euclidean distance and leverage-based method were used to determine the application domain. The methods used to obtain the above parameters were detailed in our previous studies,41,43 and are also briefly presented in Text S3.2.4 Development and application of a predictive tool
Based on the optimal cytotoxicity and genotoxicity models, a user-friendly cytotoxicity and genotoxicity of DBPs prediction tool (named DBPCytoGenoTOX Predictor) was constructed. The tool was developed in Python 3.12.7 (https://www.python.org). The development method was described in our recent study.43After deriving the tool, the cytotoxicity and genotoxicity data gaps of the substances without experimental cytotoxicity and genotoxicity data in the updated DBP inventory were filled in batch mode using the SMILES code as input format. Then, the profiles of the predicted cytotoxicity and genotoxicity data were analysed.
2.5 Identification of high-priority DBPs
The following high-priority DBP lists were identified: (a) DBPs with experimental or high reliability predicted cytotoxicity values <100 mg per L (cytotoxicity-based high priority list); (b) DBPs with experimental or high reliability predicted positive genotoxicity values (genotoxicity-based high priority list). In our recent studies, the DBPs were set as a priority in view of their aquatic toxicity and endocrine-disrupting effects, respectively.27,44 Herein, both the aquatic toxicity-based and endocrine-based high-priority lists were also employed to identify high-priority DBPs. In this regard, other high-priority lists were further analysed; namely, (c) DBPs shared in two endpoint lists, and (d) DBPs shared in three or four endpoint lists.3 Results and discussion
3.1 Data distribution
The updated cytotoxicity and genotoxicity data sets contained 184 and 105 data points, respectively, which represent an increase of 87.8% and 69.4%, respectively, compared to the data sets reported by Wagner and Plewa (i.e. 98 and 62).26 To find the characteristics of the toxicity distribution, frequency plots toward these data sets were generated (Fig. 2). For cytotoxicity, the 50% lethal concentration (LC50) values of 184 DBPs on CHO cells ranged from 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 mg L−1 (5-bromocytosine) to 0.164 mg L−1 (diiodoacetamide). The cytotoxicity LC50 of 57.6% (106 substances) DBPs were ≤100 mg L−1 (moderate or higher cytotoxicity). Among these 106 DBPs, 45 (24.4%) and 15 (8.2%) DBPs belonged to highly toxic (1–10 mg L−1) and extremely toxic (<1 mg L−1) groups, respectively. For genotoxicity, 41.9% (14 positive (13.3%) and 30 negative (28.6%)) data were qualitative. Among the 61 quantitative genotoxicity data, the 50% TDNA values of 8.6% (9 DBPs), 31.4% (33 DBPs) and 18.1% (19 DBPs) DBPs ranged from 1–10 mg L−1, 10–100 mg L−1, and 100–1000 mg L−1, respectively.
400 mg L−1 (5-bromocytosine) to 0.164 mg L−1 (diiodoacetamide). The cytotoxicity LC50 of 57.6% (106 substances) DBPs were ≤100 mg L−1 (moderate or higher cytotoxicity). Among these 106 DBPs, 45 (24.4%) and 15 (8.2%) DBPs belonged to highly toxic (1–10 mg L−1) and extremely toxic (<1 mg L−1) groups, respectively. For genotoxicity, 41.9% (14 positive (13.3%) and 30 negative (28.6%)) data were qualitative. Among the 61 quantitative genotoxicity data, the 50% TDNA values of 8.6% (9 DBPs), 31.4% (33 DBPs) and 18.1% (19 DBPs) DBPs ranged from 1–10 mg L−1, 10–100 mg L−1, and 100–1000 mg L−1, respectively.
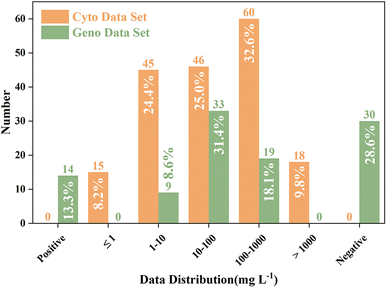 | ||
| Fig. 2 Frequency plots of the distribution of the cytotoxicity and genotoxicity data values of DBPs. | ||
3.2 Quantitative cytotoxicity model
Hundreds of quantitative cytotoxicity models were obtained using different algorithms. It was recommended that the statistical parameters of acceptable quantitative models should be R2 > 0.700, Q2 > 0.600, QEXT2 > 0.700, CCC > 0.850, rm2 > 0.500, and Δrm2 < 0.200.45 We found that only the GBDT cytotoxicity model met all the acceptable threshold values. The hyperparameters of this optimal model are listed in Table S8. The optimal GBDT model contains 8 predictive variables (nC, ATS2s, ETA_dPsi_B, RotBFrac, AATS8s, SRW2, BCUTw-1h, and GATS6s); their definitions are presented in Table S9. Fig. S1 illustrates the relationship between the observed vs. predicted log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) LC50 values for this model. The statistical parameters of this optimal GBDT model are presented in Table 1. As shown, the model has good goodness-of-fit and robustness, given that the values of all the statistical parameters were higher than the corresponding acceptable threshold values. The model also shows good external prediction accuracy, since the QEXT2 (0.793 > 0.700), CCC (0.888 > 0.850), rm2 (0.714 > 0.500), and Δrm2 (0.148 < 0.200) values are also greater or lower than the corresponding acceptable threshold values.
LC50 values for this model. The statistical parameters of this optimal GBDT model are presented in Table 1. As shown, the model has good goodness-of-fit and robustness, given that the values of all the statistical parameters were higher than the corresponding acceptable threshold values. The model also shows good external prediction accuracy, since the QEXT2 (0.793 > 0.700), CCC (0.888 > 0.850), rm2 (0.714 > 0.500), and Δrm2 (0.148 < 0.200) values are also greater or lower than the corresponding acceptable threshold values.
| Statistical parametersa | Acceptable threshold valueb | GBDTc |
|---|---|---|
| a m is the number of predictive variables; k is the number of nearest neighbors; ntrain and nEXT are the numbers of model compounds in the training and validation sets, respectively; Rtrain2 is the squared correlation coefficient of the observed and predicted values for the training set; QLOO2, QLMO2 and QBOOT2 are leave-one out cross validation Q2, leave-many out cross validation Q2 and bootstrapping coefficient, respectively; QEXT2 is the externally explained variance; CCC is the concordance correlation coefficient; rm2 is the external validation metric; Δrm2 is the absolute difference of rm2; and RMSEtrain and RMSEEXT, strain and sEXT, MAEtrain and MAEEXT are the root mean square errors, standard errors, mean absolute errors for the training and validation sets, respectively. b The acceptable threshold values of these statistical parameters were from Chirico.45 c GBDT: gradient boosting decision tree. | ||
| Training set | ||
| m | — | 8 |
| n train | — | 138 |
| R train 2 | >0.700 | 1 |
| Q LOO 2 | >0.600 | 0.752 |
| Q LMO 2 | >0.600 | 0.654 |
| Q BOOT 2 | >0.600 | 0.633 |
| Q CV 2 | >0.600 | 0.677 |
| RMSEtrain | — | 0.00139 |
| s train | — | 0.00144 |
| MAEtrain | — | 0.00112 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) |
||
| Validation set | ||
| n EXT | — | 46 |
| Q EXT 2 | >0.700 | 0.793 |
| CCC | >0.850 | 0.888 |
| r m 2 | >0.500 | 0.714 |
| Δrm2 | <0.200 | 0.148 |
| RMSEEXT | — | 0.511 |
| s EXT | — | 0.569 |
| MAEEXT | — | 0.379 |
3.3 Qualitative genotoxicity model
Many quantitative genotoxicity models were also obtained using different algorithms; however, none of them met the acceptable threshold values of a quantitative model (Table S7). We further developed qualitative genotoxicity models. As both quantitative and qualitative genotoxicity data were obtained, in the classification modelling, the quantitative genotoxicity data were assigned as positive data if the corresponding 50% TDNA value was ≤1000 mg L−1; in contrast, the genotoxicity data were grouped as negative data if the corresponding 50% TDNA value was >1000 mg L−1. In total, there were 75 positive and 30 negative data points, respectively. Upon comparison, the RF-based classification model was considered to be the optimal model; its hyperparameters are listed in Table S8. The optimal model contains 3 predictor variables (GATS3s, Sare and ETA_Shape_P). Their definitions are also presented in Table S9. Table 2 presents the statistical parameters of the RF model. For the training set, the basic parameters, such as sensitivity (Sn), specificity (Sp), accuracy (Q) and precision, are all quite high, which indicates that this model has excellent internal predictive ability. Meanwhile, the sensitivity, accuracy, MCC, precision, and F1-score values of the validation set were 1, which implied that this RF model has good generalization ability to new data. The ROC curve of this RF model is plotted in Fig. S2.| Data seta | n | S n | S p | Q | MCC | AUC | Mean_acc | Precision | F 1-score | Balanced accuracy |
|---|---|---|---|---|---|---|---|---|---|---|
| a T and V indicate the training set and validation set, respectively. n is the number of DBPs in the training and validation sets; Sn is sensitivity; Sp is specificity; Q is the predictive accuracy; MCC is the Matthews Correlation Coefficient; AUC is the area under the receiver operating characteristics (ROC) curve; F1-score is the average of precision and recall; and balanced accuracy is the arithmetic mean of true positive rate and false positive rate. | ||||||||||
| T | 81 | 0.982 | 1 | 0.988 | 0.971 | 0.991 | 0.727 | 1 | 0.991 | 0.991 |
| V | 24 | 1 | 1 | 1 | 1 | 1 | — | 1 | 1 | 1 |
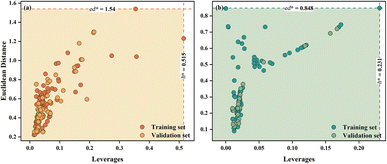
3.4 Applicability domain (AD)
The AD of the quantitative cytotoxicity and qualitative genotoxicity models was evaluated by the Euclidean distance vs. leverage-based method (Fig. 3). For both models, no compounds from the training or validation sets fell outside the domain, which indicates that the training sets of both quantitative cytotoxicity and qualitative genotoxicity models had great representativeness.3.5 Mechanistic implications
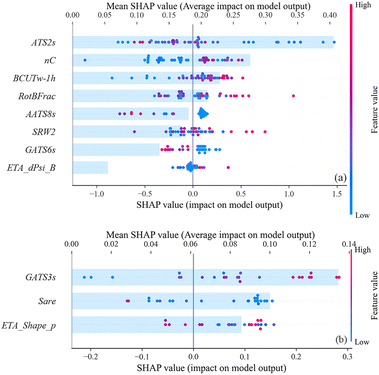
Previous studies have documented that the cytotoxicity of a given substance usually relates to two major processes. Firstly, the substance was transported from the solution to the cytomembrane, and then crossed the cell membrane and entered the cytoplasm (i.e. cell permeability). The molecular properties that could be used to characterize the aforementioned process are usually those descriptors related to hydrophobicity or lipophilicity, such as the n-octanol/water partition coefficient, dipole moment, molecular weight, and carbon chain length.38,46–50 Another factor may be the interaction between small molecules and key biomolecules in cells (e.g., structural proteins, enzymes, and DNA).51–53 In this case, this process may relate to the molecular properties and could be described by the noncovalent and covalent interactions between small molecules and the key biomolecules in cells. The descriptors that could be used to characterize these molecular properties may include atomic properties or functional groups, electronegativity, charge distribution, polarizability, dissociation energy, and molecular connectivity.47,51,53,54 Similarly, the genotoxicity of a given substance may also be associated with the cell permeability and the reactivity between small molecules and DNA.Fig. 4 displays all the predictive variables selected in the cytotoxicity and genotoxicity models. The importance of the 8 predictive variables in the cytotoxicity GBDT model is listed as ATS2s > nC > BCUTw-1h > RotBFrac > AATS8s > SRW2 > GATS6s > ETA_dPsi_B; while the importance of the 3 descriptors in the genotoxicity RF model is GATS3s > Sare > ETA_Shape_P.
Three of the 8 predictive variables (ATS2s, AATS8s and GATS6s) in the cytotoxicity model and one of the 3 predictive variables in the genotoxicity model (GATS6s) belong to the autocorrelation descriptor. They were weighted by intrinsic state (I-state, s). Taking ATS2s as an example, it means Broto–Moreau autocorrelation – lag 2/weighted by I-state. Lag 2 indicates the number of bonds separating the atoms compared to the autocorrelation function, which considers pairs of atoms that are separated by two bonds,55 while the I-state describes the intrinsic state (e.g. electronic structure, topological environment) of atoms, which is related to electronegativity.56 These descriptors may characterize the interaction between model compounds and the key biomolecules in cells. As shown in Fig. 4, these descriptors usually exhibit a clear positive correlation with the output. Autocorrelation descriptors have also been used to develop many toxic QSAR models, especially for acute toxicity or cytotoxicity.57–59
Sare is the second most important variable of the RF model, and it belongs to the constitutional descriptors. Sare can reflect the molecular composition of compounds,60 and describes the sum of atomic Allred-Rochow electronegativities. Constitutional descriptors were widely used to develop QSAR models for toxicity (e.g. oral LD50 dose, mutagenic, carcinogenic, aquatic toxicity).61–63 Similar to GATS3s, Sare also showed a positive effect on the dependent variable, i.e. genotoxicity.
The nC is the second most important predictive variable of the cytotoxicity model. The number of carbon atoms might affect the cell permeability of substances by influencing their hydrophobicity or their reactivity.50 Studies have shown that the toxicity of compounds can be enhanced if the length of the carbon chain is increased.50 In Fig. 4, the cytotoxicity may increase with the increase in the number of nC.
BCUTw-1h belongs to the eigenvalue-based descriptor and is weighted by atomic weight (w); it is incorporated into the connectivity (according to actual bonding or interatomic distances) and atomic properties. Both BCUTw-1h and RotBFrac reflect the number of rotatable bonds within a molecule, which is considered to be a critical factor for cytotoxicity.64 Both descriptors were widely used in building QSAR models for toxicity prediction.57,65,66 In the graph, BCUTw-1h shows a positive impact, while RotBFrac shows a fuzzy effect.
Extended topological chemical-atomic (ETA) indices are often used for modelling toxicity.31,67,68 ETA_dPsi_B is a measure of the hydrogen-bonding propensity of molecules, which may be used to describe the possibility of molecules forming hydrogen bond interactions with biomolecules in cells.69 ETA_Shape_P indicated the molecular shape, especially the P-shaped structural fragment of the molecule.67Fig. 4 illustrates that ETA_dPsi_B has a relatively negative contribution to the output of cytotoxicity, while ETA_Shape_P has a positive effect on genotoxicity.
3.6 Comparison of optimal model with previous quantitative/qualitative models
A comparison between the optimal cytotoxicity model developed here and previous predictive models is shown in Table S2. Among these CHO cell-based cytotoxicity models, our model contained more model compounds, but the statistical parameters of our new model are still comparable to or slightly better than those of the other models. For the NGBoost-RF model, more data were used because its dataset contained many cytotoxic data from other cell lines.For the genotoxicity model, no model was available for the genotoxicity data from the same assay as that mentioned in this study. In QSAR Toolbox v4.7, 8 genotoxicity models were also derived based on the CHO cells-based genotoxicity; however, the assay methods differ from those mentioned in this study. More information about the assay methods of those models is presented in Text S2 and Table S3. In order to make a comparison, the validation dataset (24 DBPs) used in the optimal genotoxicity RF model was employed to determine the predictive performance of those 8 models. The parameters comparison results between the model developed here and the other 8 models are listed in Table S3. The predicted genotoxicity data are presented in Table S10. The results show that our RF model exhibits better predictive ability for those 24 DBPs.
3.7 Development of the DBPCytoGenoTOX predictor
Based on the quantitative cytotoxicity GBDT model and qualitative genotoxicity RF model, a user-friendly software called “DBPCytoGenoTOX Predictor” was developed. Fig. S3 and S4 show the input and the display and save interfaces of the tool. In brief, it supports five input formats, i.e. both CAS NO and SMILES code in single and batch modes, as the structure file. After inputting valid data, information on the entered DBPs will be visualized in the input interface. The backend of the tool would then automatically predict their genotoxicity and cytotoxicity, along with an assessment of the AD and reliability. In the display and save interface, the basic information of the input substance (SMILES code and/or CAS NO), the experimental values of cytotoxicity/genotoxicity (if possible), the predicted values of cytotoxicity/genotoxicity, as well as the assessment results of the AD and reliability, would also be visualized. The results can also be saved in CSV format.3.8 Fill the data gaps of the DBP inventory
Among the 1816 substances in the updated DBP inventory, there were 1632 and 1711 DBPs without experimental cytotoxicity or genotoxicity values, respectively. The missing cytotoxicity and genotoxicity data for those 1632 and 1711 DBPs were estimated by employing the “DBPCytoGenoTOX Predictor” (Fig. 5 and Table S11). As shown, 1012 (59.15%) out of 1711 DBPs were classified as positive genotoxic substances. For cytotoxicity, the number of DBPs that belonged to extremely toxic, highly toxic, moderately toxic, low toxicity, and relatively harmless were 35 (2.15%), 128 (7.84%), 612 (37.5%), 740 (45.34%) and 117 (7.17%), respectively.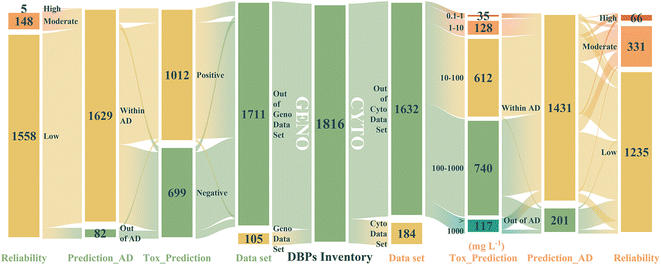 | ||
| Fig. 5 Alluvial plots of the distribution of values and reliability of the predictive results of the DBPCytoGenoTOX predictor for the updated DBP inventory. | ||
It has been recommended that the assessment results of both AD and reliability should be taken into account to decide whether a predicted value is reliable or not.41,70 The AD assessment results of two models for the corresponding predictive set are illustrated in Fig. S5. For cytotoxicity, 1431 (87.7%) out of 1632 DBPs were within both the threshold value of the Euclidean distance (ed*) and the threshold value of leverage (h*) of the quantitative cytotoxicity GBDT model. Among those 1431 DBPs, the number of predicted values labelled as low, moderate, and high reliability were 1034, 331 and 66, respectively. Taking AD and reliability into consideration, we believe that the predicted cytotoxicity of 66 DBPs on CHO cells is reliable. Further analysis results implied that 5, 33, 27, and 1 out of those 66 DBPs belonged to high, moderate, low toxicity and relatively harmless groups, respectively. Specifically, the predicted cytotoxicity LC50 of N-bromo-2,2,2-trichloroacetamide (4.78 mg L−1), 2-chloro-4-nitrophenol (5.59 mg L−1), 2-iodocyclohexa-2,5-diene-1,4-dione (5.78 mg L−1), 4-iodo-2,6-dimethylphenol (6.15 mg L−1), and 2-chlorophenol (8.57 mg L−1) were <10 mg L−1, and further tests should be performed to determine their potential harmful effects.
For genotoxicity, 1629 (95.21%) out of 1711 DBPs were within both the threshold value of the Euclidean distance (ed*) and the threshold value of leverage (h*) of the qualitative genotoxicity RF model. Within that, the number of predicted values labelled as low, moderate, and high reliability were 1476, 148 and 5, respectively. All 5 DBPs marked high reliability (dichloroacetate, N-bromo-2,2,2-trichloroacetamide, 2-iodocyclohexa-2,5-diene-1,4-dione, N,2,2,2-tetrachloroacetamide, and N,2,2-trichloroacetamide) were predicted as positive on genotoxicity.
3.9 Identification of high-priority DBPs
The number of DBPs with experimental or high reliability predicted cytotoxicity values ≤100 mg L−1 was 106 and 38, respectively, indicating that there were 144 DBPs in the cytotoxicity-based high-priority list. The number of DBPs with experimental genotoxicity values ≤1000 mg L−1 or experimental positive genotoxicity was 75, while the number of DBPs with high reliability predicted positive genotoxicity was 5. Thus, there were 80 DBPs in the genotoxicity-based high-priority list.Furthermore, the aquatic toxicity-based and endocrine-based high-priority lists were also employed to identify high-priority DBPs. Specifically, the endocrine-based high-priority list contained 216 DBPs;27 among them, 23 and 193 DBPs belonged to high and moderate endocrine-disrupting priority, respectively. For the aquatic toxicity-based list, there were 50 DBPs;44 19 and 31 DBPs out of 50 were the high and moderate aquatic toxicity priority, respectively. After merging the four lists, 385 high-priority DBPs were obtained. The distribution of those 385 DBPs is illustrated in Fig. 6 and listed in Table S12. In Fig. 6(a), the numbers of DBPs on the diagonal of the graph represent the number of DBPs belonging to a specific single-priority list, while the cross-point grid represents the number of DBPs appearing simultaneously in the priority list of two endpoints. Fig. 6(b) shows those DBPs that appeared simultaneously in the priority list of three endpoints. None of the DBPs were included in four endpoint high-priority lists simultaneously.
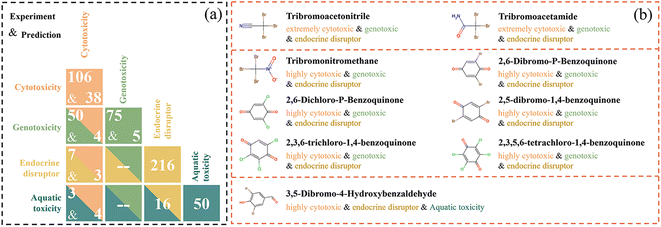 | ||
| Fig. 6 Distribution of 385 identified high-priority DBPs. The number of DBPs shared in the priority list of one or two endpoints (a) and three endpoints (b). | ||
As shown in Fig. 6(a), there were 87 DBPs found in two endpoint high-priority lists simultaneously; 54 out of 87 DBPs were in the “cytotoxicity and positive genotoxicity” lists. Of those 54 DBPs, the cytotoxicity of 11, 29 and 14 substances exhibited extreme, high and moderate cytotoxicity, respectively. There were 10 DBPs in the cytotoxicity (one high and 9 moderate cytotoxicity) and medium endocrine-disruptor lists, while 7 and 16 DBPs were in the “cytotoxicity and aquatic toxicity” and “aquatic toxicity and endocrine disruptor” lists, respectively.
Nine DBPs appeared simultaneously in three endpoint high-priority lists (Fig. 6(b)). Specifically, 2 DBPs (tribromoacetonitrile and tribromoacetamide) exhibited extreme cytotoxicity, positive genotoxicity and medium endocrine-disrupting priority; while another 6 DBPs (tribromonitromethane, 2,6-dibromo-P-benzoquinone, 2,6-dichloro-P-benzoquinone, 2,5-dibromo-1,4-benzoquinone, 2,3,6-trichloro-1,4-benzoquinone, and 2,3,5,6-tetrachloro-1,4-benzoquinone) have high cytotoxicity, positive genotoxicity, and medium endocrine-disrupting priority. 3,5-Dibromo-4-hydroxybenzaldehyde had moderate cytotoxic, medium endocrine-disrupting effects, and medium aquatic toxicity. The molecular structures of the 87 and 9 DBPs identified above are listed in Table S13.
In addition, the high-priority DBPs were analysed by only considering the DBPs in the high-toxicity groups in the four lists; i.e., DBPs with experimental and predicted cytotoxicity values ≤10 mg L−1 (65 DBPs), experimental genotoxicity values ≤10 mg L−1, and DBPs with high reliability predicted positive genotoxicity values (14 DBPs), and the 23 high endocrine-disrupting priority and 19 high aquatic toxicity priority. In total, 110 DBPs were recognized in this case (Fig. S6). Especially, there were only 11 DBPs shared in the priority list of two endpoints; i.e. in the “cytotoxicity and positive genotoxicity” lists.
Finally, the control status of the DBPs in the regulatory authorities was analysed. To date, there are 35 regulated DBPs (Table S1) in the standards and regulations issued by China, the United States Environmental Protection Agency (U.S. EPA), the World Health Organization (WHO), and the European Union (EU).3,25,27,71 We compared the 385 identified high-priority DBPs with the 35 regulated DBPs, and found that there are 10 substances identified as high-priority DBPs that are regulated by the authorities. Specifically, 2 (one also in WHO's list), 3, 3 and 3 DBPs were found under the regulation of China, U.S. EPA, WHO, and EU (Table S12). This result highlighted that many potentially highly toxic DBPs have still not received enough attention from the authorities to date (Fig. S7).
4 Conclusions
In this study, we summarized the currently largest CHO cell-based cytotoxicity (184 DBPs) and genotoxicity (105 DBPs) dataset, and developed cytotoxicity and genotoxicity ML models with good predictive ability. A user-friendly predictive tool called “DBPCytoGenoTOX Predictor” was developed, which could be employed to fill the missing cytotoxicity and genotoxicity data gap of the DBPs efficiently and with high-throughput.In addition, we obtained the largest DBP inventory to date, with clear CAS NO and SMILES codes (1816 DBPs). The missing cytotoxicity and genotoxicity values of those DBPs in the inventory without experimental cytotoxicity and genotoxicity values were predicted by “DBPCytoGenoTOX Predictor”. The predicted cytotoxicity and genotoxicity values for 66 and 5 DBPs were reliable, respectively. The priority setting results indicated that 385 substances were identified as high-priority DBPs. Among those 385 high-priority DBPs, only ten DBPs have been regulated by the authorities to date. In conclusion, this study provides an efficient and high-throughput tool for predicting the potential cytotoxicity and genotoxicity of a given DBP. Moreover, high-priority DBPs were identified, which may help others to select substances for further testing to determine their potential adverse effects on organisms.
It also deserves mentioning that the cytotoxicity and genotoxicity data used in this study were limited to one group of emerging contaminants, i.e. DBPs. To extend the predictive ability and the scope of application for the cytotoxicity and genotoxicity models, it is important to develop other predictive models for cytotoxicity and genotoxicity by modelling extensive and heterogeneous datasets with more groups of emerging contaminants in the future.
Author contributions
M. Y. Liu: investigation; methodology, data curation, formal analysis, writing – original draft. H. H. Liu: writing – review & editing. X. H. Yang: conceptualization, data curation, formal analysis, resources, funding acquisition, project administration, visualization, writing – review & editing.Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information: detailed information for cluster analysis, selection of previous cytotoxicity and genotoxicity models, statistical parameters of the models developed here and obtained from others, list of regulated DBPs, cytotoxicity and genotoxicity data set of DBP, list of updated DBPs inventory, hyperparameters of the optimal models, predicted values of cytotoxicity and genotoxicity data for DBPs inventory, list of high priority DBPs, molecular structures of high priority DBPs, interface of DBPCytoGenoTOX Predictor; the tool will be made available on request. See DOI: https://doi.org/10.1039/d5em00552c.Acknowledgements
We thank Xiaoqiu Yang from Jianghan University for providing their inventory of the identified DBPs. The study was supported by the National Natural Science Foundation of China (22176097); the Postgraduate Research & Practice Innovation Program of Jiangsu Province (No. SJCX25_0221); and the Fundamental Research Funds for the Central Universities (No. 30923011032).References
- S. D. Richardson and T. Manasfi, Water Analysis: Emerging Contaminants and Current Issues, Anal. Chem., 2024, 96, 8184–8219 CrossRef CAS PubMed.
- Y. Feng, S. S. Lau, W. A. Mitch, C. Russell, G. Pope and A. Z. Gu, Impacts of disinfection methods in a granular activated carbon (GAC) treatment system on disinfected drinking water toxicity, J. Hazard. Mater., 2025, 490, 137737 CrossRef CAS PubMed.
- D. M. DeMarini, A review on the 40th anniversary of the first regulation of drinking water disinfection by-products, Environ. Mol. Mutagen., 2020, 61, 588–601 CrossRef CAS PubMed.
- G. Sun, P. Guo, R. Liu, H. Y. Kaw and W. Wang, Fragmentation pattern-based nontargeted screening strategy uncovered novel halogenated nucleotides in drinking water, J. Hazard. Mater., 2025, 490, 137797 CrossRef CAS PubMed.
- R. Sikder, H. Zhang, P. Gao and T. Ye, Machine learning framework for predicting cytotoxicity and identifying toxicity drivers of disinfection byproducts, J. Hazard. Mater., 2024, 469, 133989 CrossRef CAS PubMed.
- T. Wang, Q. Tang, L. Deng, C. Tan, Y. Fu, J. Hu and R. P. Singh, Formation of halonitromethanes, dichloroacetonitrile, and trichloromethane in the presence of E. coli and nitrophenols during UV/post-chlorination, J. Hazard. Mater., 2025, 488, 137499 CrossRef CAS PubMed.
- D. Li, L. Long, W. Xia, W. Zhao, L. Feng, X. Xia, S. He, Y. Liu, S. You and L. Wei, Utilization of UV/VUV irradiation for removal of human body fluids related pollutants in swimming pool water, J. Hazard. Mater., 2025, 489, 137549 CrossRef CAS PubMed.
- D. Zhang, S. Dong, L. Chen, R. Xiao and W. Chu, Disinfection byproducts in indoor swimming pool water: detection and human lifetime health risk assessment, J. Environ. Sci., 2023, 126, 378–386 CrossRef CAS PubMed.
- M. Wang, Y. Tang, S. Mi, J. Pan, C. Guo, J. Li and B. Han, Difference in accumulation of five phthalate esters in different elite tea cultivars and their correlation with environment factors, Agriculture, 2022, 12, 1516 CrossRef CAS.
- L. Xu, S. Song, N. J. D. Graham and W. Yu, Direct generation of DBPs from city dust during chlorine-based disinfection, Water Res., 2024, 248, 120839 CrossRef CAS PubMed.
- M. Wu, S. Ding, Z. Cao, Z. Du, Y. Tang, X. Chen and W. Chu, Insights into the formation and mitigation of iodinated disinfection by-products during household cooking with laminaria japonica (Haidai), Water Res., 2022, 225, 119177 CrossRef CAS PubMed.
- H. Ding, L. Meng, H. Zhang, J. Yu, W. An, J. Hu and M. Yang, Occurrence, profiling and prioritization of halogenated disinfection by-products in drinking water of China, Environ. Sci.: Processes Impacts, 2013, 15, 1424–1429 RSC.
- S. Dong, N. Masalha, M. J. Plewa and T. H. Nguyen, Toxicity of wastewater with elevated bromide and iodide after chlorination, chloramination, or ozonation disinfection, Environ. Sci. Technol., 2017, 51, 9297–9304 CrossRef CAS PubMed.
- S. S. Lau, X. Wei, K. Bokenkamp, E. D. Wagner, M. J. Plewa and W. A. Mitch, Assessing Additivity of Cytotoxicity Associated with Disinfection Byproducts in Potable Reuse and Conventional Drinking Waters, Environ. Sci. Technol., 2020, 54, 5729–5736 CrossRef CAS PubMed.
- J. R. Wilkins, N. A. Reiches and C. W. Kruse, Organic chemical contaminants in drinking water and cancer, Am. J. Epidemiol., 1979, 110, 420–448 CrossRef PubMed.
- M. Koivusalo, J. J. Jaakkola, T. Vartiainen, T. Hakulinen, S. Karjalainen, E. Pukkala and J. Tuomisto, Drinking water mutagenicity and gastrointestinal and urinary tract cancers: an ecological study in Finland, Am. J. Public Health, 1994, 84, 1223–1228 CrossRef CAS PubMed.
- M. Koivusalo, T. Vartiainen, T. Hakulinen, E. Pukkala and J. J. Jaakkola, Drinking water mutagenicity and leukemia, lymphomas, and cancers of the liver, pancreas, and soft tissue, Arch. Environ. Health, 1995, 50, 269–276 CrossRef CAS PubMed.
- S. Kali, M. Khan, M. S. Ghaffar, S. Rasheed, A. Waseem, M. M. Iqbal, M. Bilal Khan Niazi and M. I. Zafar, Occurrence, influencing factors, toxicity, regulations, and abatement approaches for disinfection by-products in chlorinated drinking water: a comprehensive review, Environ. Pollut., 2021, 281, 116950 CrossRef CAS PubMed.
- Y. Du, W.-L. Wang, Z.-W. Wang, C.-J. Yuan, M.-Q. Ye and Q.-Y. Wu, Overlooked cytotoxicity and genotoxicity to mammalian cells caused by the oxidant peroxymonosulfate during wastewater treatment compared with the sulfate radical-based ultraviolet/peroxymonosulfate process, Environ. Sci. Technol., 2023, 57, 3311–3322 CrossRef CAS PubMed.
- National Toxicology Program, Report on the carcinogenesis bioassay of chloroform (CAS No. 67-66-3), National Cancer Institute Carcinogenesis Technical Report Series, 1976, pp. 1–60 Search PubMed.
- V. F. Simmon, K. Kauhanen, K. Mortelmans and R. Tardiff, Mutagenic activity of chemicals identified in drinking water, Mutat. Res., Environ. Mutagen. Relat. Subj., 1978, 53, 262 Search PubMed.
- S. Zhao, X. Yang, H. Liu, Y. Xi and J. Li, Potential disrupting effects of wastewater-derived disinfection byproducts on Chinese rare minnow (gobiocypris rarus) transthyretin: an in vitro and in silico study, Environ. Sci. Technol., 2023, 57, 3228–3237 CrossRef CAS PubMed.
- J. Liu and X. Zhang, Comparative toxicity of new halophenolic DBPs in chlorinated saline wastewater effluents against a marine alga: halophenolic DBPs are generally more toxic than haloaliphatic ones, Water Res., 2014, 65, 64–72 CrossRef CAS PubMed.
- S. D. Richardson, M. J. Plewa, E. D. Wagner, R. Schoeny and D. M. DeMarini, Occurrence, genotoxicity, and carcinogenicity of regulated and emerging disinfection by-products in drinking water: a review and roadmap for research, Mutat. Res., Rev. Mutat. Res., 2007, 636, 178–242 CrossRef CAS PubMed.
- E. McKenna, K. A. Thompson, L. Taylor-Edmonds, D. L. McCurry and D. Hanigan, Summation of disinfection by-product CHO cell relative toxicity indices: sampling bias, uncertainty, and a path forward, Environ. Sci.: Processes Impacts, 2020, 22, 708–718 RSC.
- E. D. Wagner and M. J. Plewa, CHO cell cytotoxicity and genotoxicity analyses of disinfection by-products: An updated review, J. Environ. Sci., 2017, 58, 64–76 CrossRef CAS PubMed.
- S. Sui, N. Zhou, H. Liu, P. Watson and X. Yang, Recognizing high-priority disinfection byproducts based on experimental and predicted endocrine disrupting data: Virtual screening and in vitro study, Chemosphere, 2024, 358, 142239 CrossRef CAS PubMed.
- H. Chen, J. Xie, C. Huang, Y. Liang, Y. Zhang, X. Zhao, Y. Ling, L. Wang, Q. Zheng and X. Yang, Database and review of disinfection by-products since 1974: constituent elements, molecular weights, and structures, J. Hazard. Mater., 2024, 462, 132792 CrossRef CAS PubMed.
- General Administration of Quality Supervision Standardization Administration of the People’s Republic of China, Standards for drinking water quality (GB 5749-2022), Standards Press of China, 2022, https://openstd.samr.gov.cn/bzgk/std/newGbInfo?hcno=99E9C17E3547A3C0CE2FD1FFD9F2F7BE (accessed Apr. 4, 2025) Search PubMed.
- C. Chen, CiteSpace II: detecting and visualizing emerging trends and transient patterns in scientific literature, J. Am. Soc. Inf. Sci. Technol., 2006, 57, 359–377 CrossRef.
- A. Seth, P. K. Ojha and K. Roy, QSAR modeling with ETA indices for cytotoxicity and enzymatic activity of diverse chemicals, J. Hazard. Mater., 2020, 394, 122498 CrossRef CAS PubMed.
- B. Chen, T. Zhang, T. Bond and Y. Gan, Development of quantitative structure activity relationship (QSAR) model for disinfection byproduct (DBP) research: a review of methods and resources, J. Hazard. Mater., 2015, 299, 260–279 CrossRef CAS PubMed.
- S. Cui, Y. Gao, Y. Huang, L. Shen, Q. Zhao, Y. Pan and S. Zhuang, Advances and applications of machine learning and deep learning in environmental ecology and health, Environ. Pollut., 2023, 335, 122358 CrossRef CAS PubMed.
- W. Liu, J. Chen, H. Wang, Z. Fu, W. J. G. M. Peijnenburg and H. Hong, Perspectives on advancing multimodal learning in environmental science and engineering studies, Environ. Sci. Technol., 2024, 58, 16690–16703 CAS.
- L.-T. Qin, X. Zhang, Y.-H. Chen, L.-Y. Mo, H.-H. Zeng, Y.-P. Liang, H. Lin and D.-Q. Wang, Predicting the cytotoxicity of disinfection by-products to Chinese hamster ovary by using linear quantitative structure-activity relationship models, Environ. Sci. Pollut. Res., 2019, 26, 16606–16615 CrossRef CAS PubMed.
- Z. Zhang, Q. Zhu, C. Huang, M. Yang, J. Li, Y. Chen, B. Yang and X. Zhao, Comparative cytotoxicity of halogenated aromatic DBPs and implications of the corresponding developed QSAR model to toxicity mechanisms of those DBPs: binding interactions between aromatic DBPs and catalase play an important role, Water Res., 2020, 170, 115283 CrossRef CAS PubMed.
- X. Wei, M. Yang, Q. Zhu, E. D. Wagner and M. J. Plewa, Comparative quantitative toxicology and QSAR modeling of the haloacetonitriles: forcing agents of water disinfection byproduct toxicity, Environ. Sci. Technol., 2020, 54, 8909–8918 CrossRef CAS PubMed.
- Z. Zhang, S. Hu, G. Sun and W. Wang, Target analysis, occurrence and cytotoxicity of halogenated polyhydroxyphenols as emerging disinfection byproducts in drinking water, Water Res., 2024, 248, 120883 CrossRef CAS PubMed.
- K. S. Egorova and V. P. Ananikov, Toxicity of ionic liquids: eco(cyto)activity as complicated, but unavoidable parameter for task-specific optimization, ChemSusChem, 2014, 7, 336–360 CrossRef CAS PubMed.
- C. W. Yap, PaDEL-descriptor: an open source software to calculate molecular descriptors and fingerprints, J. Comput. Chem., 2011, 32, 1466–1474 CrossRef CAS PubMed.
- X. Yang, Y. Yang, P. Watson and H. Liu, Development of quantitative structure property relationship models and tool for predicting the soil adsorption coefficient (logKOC), Environ. Pollut., 2025, 368, 125703 CrossRef CAS PubMed.
- S. M. Lundberg and S.-I. Lee, in Proceedings of the 31st International Conference on Neural Information Processing Systems, Curran Associates Inc., Red Hook, NY, USA, 2017, pp. 4768–4777 Search PubMed.
- X. Li, H. Liu, S. Zhao, P. Watson and X. Yang, Binding interaction of typical emerging contaminants on Gobiocypris rarus transthyretin: an in vitro and in silico study, Front. Environ. Sci. Eng., 2024, 18, 135 CrossRef CAS.
- N. Zhou, S. Sui, H. Liu, X. Yang, H. Hong and T. A. Patterson, Determining high priority disinfection byproducts based on experimental aquatic toxicity data and predictive models: virtual screening and in vivo study, Sci. Total Environ., 2024, 951, 175489 CrossRef CAS PubMed.
- N. Chirico and P. Gramatica, Real external predictivity of QSAR models. Part 2. New intercomparable thresholds for different validation criteria and the need for scatter plot inspection, J. Chem. Inf. Model., 2012, 52, 2044–2058 CrossRef CAS PubMed.
- C.-Y. Chen and J.-H. Lin, Toxicity of chlorophenols to pseudokirchneriella subcapitata under air-tight test environment, Chemosphere, 2006, 62, 503–509 CrossRef CAS PubMed.
- A. A. Toropov, B. F. Rasulev and J. Leszczynski, QSAR modeling of acute toxicity by balance of correlations, Bioorg. Med. Chem., 2008, 16, 5999–6008 CrossRef CAS PubMed.
- T. Akabli, T. Hamid and F. Lamchouri, In silico modeling studies of N9-substituted harmine derivatives as potential anticancer agents: combination of ligand-based and structure-based approaches, J. Biomol. Struct. Dyn., 2022, 40, 3965–3978 CrossRef CAS PubMed.
- S. Wang, L. C. Yan, S. S. Zheng, T. T. Li, L. Y. Fan, T. Huang, C. Li and Y. H. Zhao, Toxicity of some prevalent organic chemicals to tadpoles and comparison with toxicity to fish based on mode of toxic action, Ecotoxicol. Environ. Saf., 2019, 167, 138–145 CrossRef CAS PubMed.
- L.-L. Uppgård, Å. Lindgren, M. Sjöström and S. Wold, Multivariate quantitative structure-activity relationships for the aquatic toxicity of technical nonionic surfactants, J. Surfactants Deterg., 2000, 3, 33–41 CrossRef.
- S. Wang, D. Tian, W. Zheng, S. Jiang, X. Wang, M. E. Andersen, Y. Zheng, G. He and W. Qu, Combined exposure to 3-chloro-4-dichloromethyl-5-hydroxy-2(5H)-furanone and microsytin-LR increases genotoxicity in Chinese hamster ovary cells through oxidative stress, Environ. Sci. Technol., 2013, 47, 1678–1687 CAS.
- P. V. AshaRani, G. Low Kah Mun, M. P. Hande and S. Valiyaveettil, Cytotoxicity and genotoxicity of silver nanoparticles in human cells, ACS Nano, 2009, 3, 279–290 CrossRef CAS PubMed.
- J. Wu, C. Bian, X. Yang and G. Su, CytoToxLCM: a software to predict cytotoxicity of emerging contaminant liquid crystal monomers, Environ. Sci. Technol., 2025, 59, 7028–7038 CrossRef CAS PubMed.
- M. J. Plewa, E. D. Wagner, S. D. Richardson, A. D. Thruston, Y.-T. Woo and A. B. McKague, Chemical and biological characterization of newly discovered iodoacid drinking water disinfection byproducts, Environ. Sci. Technol., 2004, 38, 4713–4722 CrossRef CAS PubMed.
- S. Yang and S. Kar, How safe are wild-caught salmons exposed to various industrial chemicals? First ever in silico models for salmon toxicity data gaps filling, J. Hazard. Mater., 2024, 477, 135401 CrossRef CAS PubMed.
- L.-T. Qin, S.-S. Liu and H.-L. Liu, QSPR model for bioconcentration factors of nonpolar organic compounds using molecular electronegativity distance vector descriptors, Mol. Diversity, 2010, 14, 67–80 CrossRef CAS PubMed.
- N.-X. Tan, P. Li, H.-B. Rao, Z.-R. Li and X.-Y. Li, Prediction of the acute toxicity of chemical compounds to the fathead minnow by machine learning approaches, Chemom. Intell. Lab. Syst., 2010, 100, 66–73 CrossRef CAS.
- Z. Liu, K. Dang, J. Gao, P. Fan, C. Li, H. Wang, H. Li, X. Deng, Y. Gao and A. Qian, Toxicity prediction of 1,2,4-triazoles compounds by QSTR and interspecies QSTTR models, Ecotoxicol. Environ. Saf., 2022, 242, 113839 CrossRef CAS PubMed.
- M. Iman and A. Davood, QSAR and QSTR study of pyrimidine derivatives to improve their therapeutic index as antileishmanial agents, Med. Chem. Res., 2013, 22, 5029–5035 CrossRef CAS.
- Danishuddin and A. U. Khan, Descriptors and their selection methods in QSAR analysis: paradigm for drug design, Drug Discovery Today, 2016, 21, 1291–1302 CrossRef CAS PubMed.
- M. H. Keshavarz and A. R. Akbarzadeh, A simple approach for assessment of toxicity of nitroaromatic compounds without using complex descriptors and computer codes, SAR QSAR Environ. Res., 2019, 30, 347–361 CrossRef CAS PubMed.
- M. Chatterjee and K. Roy, Prediction of aquatic toxicity of chemical mixtures by the QSAR approach using 2D structural descriptors, J. Hazard. Mater., 2021, 408, 124936 CrossRef CAS PubMed.
- Y. Hao, G. Sun, T. Fan, X. Sun, Y. Liu, N. Zhang, L. Zhao, R. Zhong and Y. Peng, Prediction on the mutagenicity of nitroaromatic compounds using quantum chemistry descriptors based QSAR and machine learning derived classification methods, Ecotoxicol. Environ. Saf., 2019, 186, 109822 CrossRef CAS PubMed.
- H.-J. Huang, Y.-H. Lee, C.-L. Chou, C.-M. Zheng and H.-W. Chiu, Investigation of potential descriptors of chemical compounds on prevention of nephrotoxicity via QSAR approach, Comput. Struct. Biotechnol. J., 2022, 20, 1876–1884 CrossRef CAS PubMed.
- D. Zarini, A. Sangion, E. Ferri, E. Caruso, S. Zucchi, A. Orro and E. Papa, Are In silico approaches applicable As a first step for the prediction of e-liquid toxicity in e-cigarettes?, Chem. Res. Toxicol., 2020, 33, 2381–2389 Search PubMed.
- S. Wu, S.-X. Li, J. Qiu, H.-M. Zhao, Y.-W. Li, N.-X. Feng, B.-L. Liu, Q.-Y. Cai, L. Xiang, C.-H. Mo and Q. X. Li, Accurate prediction of rat acute oral toxicity and reference dose for thousands of polycyclic aromatic hydrocarbon derivatives based on chemometric QSAR and machine learning, Environ. Sci. Technol., 2024, 58, 15100–15110 CAS.
- K. Roy and R. N. Das, QSTR with extended topochemical atom (ETA) indices. 16. Development of predictive classification and regression models for toxicity of ionic liquids towards daphnia magna, J. Hazard. Mater., 2013, 254–255, 166–178 CrossRef CAS PubMed.
- K. Roy and G. Ghosh, QSTR with extended topochemical atom (ETA) indices. 11. Comparative QSAR of acute NSAID cytotoxicity in rat hepatocytes using chemometric tools, Mol. Simul., 2009, 35, 648–659 CrossRef CAS.
- M. Haritha, M. Sreerag and C. H. Suresh, Quantifying the hydrogen-bond propensity of drugs and its relationship with lipinski's rule of five, New J. Chem., 2024, 48, 4896–4908 RSC.
- X. Yang, W. Ou, S. Zhao, Y. Xi, L. Wang and H. Liu, Rapid screening of human transthyretin disruptors through a tiered in silico approach, ACS Sustainable Chem. Eng., 2021, 9, 5661–5672 CrossRef CAS.
- European Chemicals Agency, Candidate list of substances of very high concern for authorisation (Article 59(10) of the REACH Regulation), ECHA, 2025, https://echa.europa.eu/candidate-list-table (accessed Mar. 20, 2025) Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |