Interaction of copper(II) (3 + 2) N-chelating complexes with DNA and BSA: hydrolytic DNA cleavage and ROS-mediated apoptosis†
Mariappan
Murali
 *a,
Anbarasu
Kanchana Mala
b,
Somasundaram
Sangeetha
ac,
Balasubramaniam
Selvakumaran
*a,
Anbarasu
Kanchana Mala
b,
Somasundaram
Sangeetha
ac,
Balasubramaniam
Selvakumaran
 a,
Pitchan
Arul Prakash
d,
Selvaraj
Shanmugavadivel
a,
Tamilarasan
Ajay Kamal
a,
Pitchan
Arul Prakash
d,
Selvaraj
Shanmugavadivel
a,
Tamilarasan
Ajay Kamal
 ef and
Mohamed Sultan
Mohamed Jaabir
d
ef and
Mohamed Sultan
Mohamed Jaabir
d
aCoordination and Bioinorganic Chemistry Research Laboratory, Department of Chemistry, National College (Autonomous), Tiruchirappalli 620 001, Tamil Nadu, India. E-mail: murali@nct.ac.in; ma66mu@gmail.com; Fax: +91-431-2481997; Tel: +91-431-2482995
bDepartment of Pharmacology, Saveetha Institute of Medical and Technical Sciences, Chennai 600 124, Tamil Nadu, India
cDepartment of Chemistry, Tamilavel Umamaheswaranar Karanthai Arts College, Thanjavur 613 002, Tamil Nadu, India
dDepartment of Biotechnology and Microbiology, National College (Autonomous), Tiruchirappalli 620 001, Tamil Nadu, India
eCenter for Advanced Energy Materials, SRM TRP Engineering College, Tiruchirapalli-621 105, Tamil Nadu, India
fCentre for Research, Trichy SRM Medical College Hospital and Research Center, Tiruchirappalli-621105, India
First published on 7th April 2026
Abstract
Mixed-ligand copper(II) complexes of the form [Cu(L)(phen)](ClO4)2, where L is (6-methylpyridin-2-ylmethylene)-(pyridin-2-ylmethyl)amine (L1) and (6-methylpyridin-2-ylmethylene)-(pyridin-2-ylethyl)amine (L2), are obtained and characterized using analytical, spectral, and electrochemical methods. A distorted square pyramidal coordination geometry is observed in the single-crystal X-ray structure of [Cu(L2)(phen)](ClO4)2. DFT calculations of copper(II) complexes in solution offered insights into the molecular geometry, electronic properties, and quantum chemical descriptors. Complexes interact hydrophobically with calf thymus (CT) DNA through partial intercalation, as demonstrated by absorption spectral titration, EthBr displacement assay, and circular dichroic spectral and electrochemical investigations. They undergo a static process of quenching when exposed to bovine serum albumin (BSA) protein. Both complexes cleave pUC19 supercoiled (SC) DNA in the absence of an activator at concentrations of 130–140 μM, degrading SC DNA into nicked circular (NC) DNA and thereby indicating that they are effective hydrolytic chemical nucleases. They are notable for exhibiting cytotoxicity against human cervical carcinoma (HeLa) and human lung epithelial adenocarcinoma (A549) cell lines with potency greater than cisplatin, demonstrating that they have the potential to be effective anticancer drugs. They preferentially target cancer cells because they are non-toxic to human lung epithelial (L132) cells and normal mouse embryonic fibroblasts (NIH 3T3). AO/EB and DAPI staining, annexin-V-FITC staining assay, propidium iodide assay, and metal-assisted reactive oxygen species generation indicated an apoptotic mode of cell death, possibly by causing membrane instability and initiating apoptosis. These complexes can also stop the proliferation of A549 cancer cells by causing cell cycle arrest at the S-phase. Their efficiency as promising drugs for targeted cancer treatment is highlighted in this study.
Introduction
Many transition-metal complexes have been identified as potential chemotherapeutic drugs, and some of them are in various stages of clinical trials.1 Among them, certain dinuclear platinum complexes have been found to be more effective anticancer drugs than cisplatin.2–4 However, the widespread use of platinum-based drugs in medical applications has resulted in a variety of side effects, including nephrotoxicity, ototoxicity, and neurotoxicity, as well as intrinsic or acquired drug resistance.5,6 As a result, the research community has been inspired to develop new low-toxicity compounds with improved pharmacological properties.7,8 Among them, the metal ion selected also plays an outstanding role in chemotherapeutic drug development. Here, we opted for one of the most abundant transition metals, specifically the less toxic and biocompatible copper. Copper, a bio-essential metal ion, is involved in many biological processes, most notably the regular functioning of cells. Copper(II) complexes with well-defined redox activity can be effective antioxidants, antimicrobials, antiparasitics, and anticancer agents.9,10 Some families of copper complexes have been studied as prospective anticancer agents in recent years,8 and our new copper complexes are considered better alternatives to platinum complexes.The significance of copper complexes in anticancer research has grown since the discovery of their antiproliferative activity. Researchers have examined the structure–activity relationship of known copper complexes to develop therapeutic molecules for cancer therapy.11 Due to its importance in tumour growth, angiogenesis, and apoptosis, Cu has become increasingly essential in cancer treatment.12 In chemotherapeutic applications, the anticancer activity of copper complexes is greatly affected by the ligand environment, which also determines the reduction potential of the Cu(II)/Cu(I) system.13,14 According to Ruiz-Azuara and coworkers, Casiopeínas®, a copper(II) compound, exhibits selective cytotoxic, genotoxic, and antineoplastic effects. It interacts with DNA to promote mitochondrial toxicity and the production of reactive oxygen species (ROS), which damage cells and trigger apoptosis via the intrinsic pathway.15,16 Based on recent reports, copper(II) complexes containing Schiff bases9c have shown substantial antiproliferative activity and selectivity for lung adenocarcinoma cells, while copper(II) complexes containing 1,10-phenanthroline17 have demonstrated excellent inhibition activity on A549 tumour cells and induced cell death through apoptosis. Cubisbel (copper-bis-belinostat), as described by Marmion and coworkers,18 dramatically slowed the development of colon cancer cells by inhibiting histone deacetylase (HDAC) and inducing apoptosis. Interfering with important master regulators (MRs) in colon cancer improves metabolic stability in vitro and targets anticancer pathways. Tsvetkov et al. demonstrated that the mechanism of copper toxicity is different from all other known methods of controlled cell death, such as ferroptosis, pyroptosis, necroptosis, and apoptosis. Cuproptosis, a copper-dependent form of apoptosis, is induced by the direct binding of copper to the lipoylated components of the tricarboxylic acid (TCA) cycle. This results in the loss of iron–sulfur cluster proteins and the aggregation of lipoylated proteins, which ultimately leads to proteotoxic stress and cell death.19 Because their mechanisms of action are different from those of other metal complexes (especially platinum drugs), copper complexes have garnered attention and raised hopeful expectations.20,21 Due to their strong DNA-binding/cleaving ability, mixed-ligand copper(II) complexes have recently been discovered to be good anticancer agents via inducing apoptosis.22–24 Therefore, considering the advantages of processes that produce fragments similar to those formed by restriction enzymes,25,26 designing suitable copper complexes for DNA binding and cleavage under both oxidative and hydrolytic conditions, depending on the recognition elements in the ligand, is of importance.
We synthesized and characterized two mixed ligand copper(II) complexes of the type [Cu(L1/L2)(phen)](ClO4)2, where L1 or L2 is a tridentate 3N Schiff base ligand derived from the condensation of 6-methylpyridine-2-carboxaldehyde with 2-aminomethylpyridine (L1) or 2-aminoethylpyridine (L2) (Scheme 1). Different degrees of flexibility can be imposed upon these complexes by alterations in the spacers between the pyridine and imine nitrogen of the ligands. The L1 was modified in order to investigate the impact of the number of chelate ring members on biological activity when the methyl spacer (L1, five-membered) was substituted with an ethyl spacer (L2, six-membered). Copper(II) complexes with the simple or methylated 3N tridentate ligand have attracted the most attention among the various coordination molecules due to their strong hydrolytic cleavage of genomic DNA and potent anticancer properties.27,28 As a result, the biological activity of 3N Schiff base is linked to its capacity to chelate transition metal ions; coordinating with the metal ion via nitrogen atoms may increase biological activity by acting as a strong DNA cleaver and efficient cytotoxic substance.25,29 It has been reported that combining a single phen moiety can increase activity; however, combining two phen moieties in the same complex can retard biological processes, especially under physiological conditions.30 The interactions of these complexes with CT DNA and BSA were studied using various spectral and electrochemical techniques, and the intent was to explore binding properties. Further, the cleavage activities of the complexes with pUC19 supercoiled plasmid DNA was investigated. The chemotherapeutic efficacy of the complexes was studied in terms of cytotoxicity against human cervical carcinoma (HeLa) and human lung epithelial adenocarcinoma (A549) cells and non-cancerous mouse embryonic fibroblast (NIH 3T3) and human lung epithelial (L132) cell lines. The copper(II) complexes are more sensitive to cancerous A549 cells; hence, this cell line was selected to detect the mode of cell death, such as (i) cell cycle arrest, (ii) the production of reactive oxygen species (ROS), and (iii) the induction of apoptosis. Interestingly, the present mixed ligand copper(II) complexes of tridentate 3N Schiff base ligands with phen are involved in partial intercalative binding with DNA, display hydrolytic DNA cleavage, exhibit greater anticancer activities, induce apoptosis, and are predominantly non-toxic to normal cells.
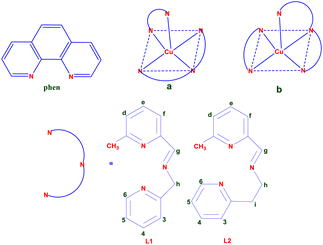 | ||
| Scheme 1 Schematic of the copper(II) complex (a and b) and structure of ligands (L1, L2, and phen) and the proton numbering scheme (L1 and L2). | ||
Experimental
Materials
Copper(II) acetate monohydrate, 1,10-phenanthroline (Merck), 2-aminomethylpyridine, 2-aminoethylpyridine, 6-methyl-pyridine-2-carboxaldehyde, ethidium bromide (EthBr), sodium azide (NaN3), potassium iodide (KI), sodium hydrogen phosphate monohydrate, di-sodium hydrogen phosphate dehydrate, (Sigma-Aldrich), L-ascorbic acid (Fisher Scientific), hydrogen peroxide solution (30% w/v), tetra-N-butylammonium bromide, perchloric acid, sodium perchlorate (Sigma Aldrich), pUC19 supercoiled DNA and agarose (Genei). Calf thymus (CT) DNA (highly polymerized, stored at 4 °C), superoxide dismutase (SOD) (stored at 4 °C), bovine serum albumin (BSA) (stored at 4 °C), catalase (stored at 4 °C), and T4 DNA ligase enzyme (stored at 4 °C) (Sigma Aldrich) were used as received without further purification. Methanol, ethanol, diethyl ether, acetone, HPLC-grade dimethylformamide (DMF), dimethyl sulfoxide (DMSO), and acetonitrile were purchased from Merck. Ultrapure Milli Q water (18.2 μΩ) was used for all the experiments.Physical measurements
The elemental analyses (C, H, N) were carried out using a PerkinElmer 2400 series II analyzer. The electrical conductivity was obtained with a Systronic 305 conductivity bridge using 1 × 10−3 M solution of the complex in DMF. FTIR spectra were recorded using a PerkinElmer Paragon 1000 FTIR spectrophotometer, equipped with a Golden Gate Diamond ATR device, applying the reflectance technique (4000–300 cm−1), and the peaks are reported in cm−1. PerkinElmer Lambda 35 and PerkinElmer Lambda 365 were used to record diffuse reflectance spectra, and a UV-VIS spectrophotometer was attached to a refrigerated circulating water bath (25 ± 0.2 °C) using cuvettes (path length, 1 cm) to capture solution electronic absorption spectra in the 200–1100 nm range. The X-band EPR spectra of the solid at ambient temperature and the solution at 77 K were acquired using a JEOL JES-FA200 ESR spectrometer at an X-band frequency (9–10 GHz) with 100 kHz field modulation. 1H NMR spectra were recorded using a Bruker DPX 300 (300 MHz) spectrometer. Chemical shifts are reported as δ (ppm) values (multiplicity, integration, and assignment) relative to the DMSO-d6 solvent peak. Emission intensity measurements were carried out using a Shimadzu RF-5301PC spectrofluorophotometer equipped with a thermostatic bath. A circular dichroic spectrum of DNA or BSA was obtained using a JASCO J-716 spectropolarimeter. A CHI 620C electrochemical analyzer operating at 25 ± 0.2 °C was used to evaluate the redox potentials of copper(II) complexes 1 and 2 (0.001 M) in methanol with TBAP (0.1 M) as the supporting electrolyte. Cyclic voltammetry (CV) and differential pulse voltammetry (DPV) were employed. The standard configurations of three electrodes, such as the working electrode (glassy carbon; A, 0.0707 cm2), counter electrode (platinum wire), and reference electrode (saturated calomel), were employed. The potential varied between −1 and +1 V at scan speeds of 0.01–0.1 V s−1. Before measurements, N2 was quickly bubbled into the solution to eliminate oxygen. Using the formula E1/2 = (Epc + Epa)/2 and ΔEp = Epa − Epc, redox data were calculated.Synthesis of ligands
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1525 ν(C
N), 1525 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 982 νC–H(H–C
C), 982 νC–H(H–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 726, and 782 ν(C–H). HRMS value in CH2Cl2: m/z, 211.98 (M + H)+. 1H NMR (300 MHz, DMSO-d6) δ/ppm: 7.45 (d, 1H, Hd), 7.06 (t, 1H, He), 7.21 (d, 1H, Hf), 8.91 (s, 1H, Hg), 3.79 (s, 2H, Hh), 7.36 (d, 1H, H3), 7.56 (t, 1H, H4), 7.14 (t, 1H, H5), 8.53 (d, 1H, H6), 2.46 (s, 3H, Py-CH3).
N), 726, and 782 ν(C–H). HRMS value in CH2Cl2: m/z, 211.98 (M + H)+. 1H NMR (300 MHz, DMSO-d6) δ/ppm: 7.45 (d, 1H, Hd), 7.06 (t, 1H, He), 7.21 (d, 1H, Hf), 8.91 (s, 1H, Hg), 3.79 (s, 2H, Hh), 7.36 (d, 1H, H3), 7.56 (t, 1H, H4), 7.14 (t, 1H, H5), 8.53 (d, 1H, H6), 2.46 (s, 3H, Py-CH3).
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1529 ν(C
N), 1529 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 981 νC–H(H–C
C), 981 νC–H(H–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 730, and 783 ν(C–H). HRMS value in CH2Cl2: m/z, 226.22 (M + H)+. 1H NMR (300 MHz, DMSO-d6) δ/ppm: 7.43 (d, 1H, Hd), 7.07 (t, 1H, He), 7.25 (d, 1H, Hf), 8.95 (s, 1H, Hg), 3.81 (t, 2H, Hh), 3.62 (t, 2H, Hi), 7.41 (d, 1H, H3), 7.63 (t, 1H, H4), 6.98 (t, 1H, H5), 8.62 (d, 1H, H6), 2.48 (s, 3H, Py-CH3).
N), 730, and 783 ν(C–H). HRMS value in CH2Cl2: m/z, 226.22 (M + H)+. 1H NMR (300 MHz, DMSO-d6) δ/ppm: 7.43 (d, 1H, Hd), 7.07 (t, 1H, He), 7.25 (d, 1H, Hf), 8.95 (s, 1H, Hg), 3.81 (t, 2H, Hh), 3.62 (t, 2H, Hi), 7.41 (d, 1H, H3), 7.63 (t, 1H, H4), 6.98 (t, 1H, H5), 8.62 (d, 1H, H6), 2.48 (s, 3H, Py-CH3).
Synthesis of copper(II) complexes
The copper(II) complexes were prepared using the general procedure as follows: copper(II) acetate monohydrate (0.2 g, 1 mmol) in ethanol (15 mL) was slowly added to 1,10-phenanthroline monohydrate (0.20 g, 1 mmol) in ethanol (15 mL), and the mixture was stirred for 1 h to ensure complete dissolution. To this mixture, ligand L1 (0.21 g, 1 mmol) or L2 (0.23 g, 1 mmol) was added with stirring to the reaction mixture. The resulting green solution was refluxed at 70 °C for 3 h. After the complexation, it was filtered while hot, and an ethanolic solution of NaClO4 (0.12 g, 1 mmol) was added. The solution was cooled to room temperature, and green colored crystalline solids of 1 or 2 were formed after three days. They were filtered off, washed with a small amount of cold ethanol, and dried under a vacuum over P4O10.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1519 ν(C
N), 1519 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 852 νC–H(H–C
C), 852 νC–H(H–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 724, 778 ν(C–H), 1090, 622 ν(ClO4−), 530, and 562 ν(Cu–N). UV-Visible data in DMF [λmax/nm (εmax/dm3 mol−1 cm−1)]: 684 (100), 896 sh, 323 (4485), 294 (24
N), 724, 778 ν(C–H), 1090, 622 ν(ClO4−), 530, and 562 ν(Cu–N). UV-Visible data in DMF [λmax/nm (εmax/dm3 mol−1 cm−1)]: 684 (100), 896 sh, 323 (4485), 294 (24![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 670), 268 (46
670), 268 (46![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 420). EPR data: in solid (RT) giso = 2.147; in DMF solution (77 K), g∥ = 2.233, g⊥ = 2.045, A∥ = 188 × 10−4 cm−1, g∥/A∥ = 119 cm, G = 5.2. Electrochemical data (DMF, 0.1 M TBAP): cyclic voltammetry, E1/2 = −0.077 V, ΔEp = 100 mV, ipa/ipc = 0.7, D = 7.7 × 10−6 cm2 s−1, and differential pulse voltammetry, E1/2 = −0.063 V.
420). EPR data: in solid (RT) giso = 2.147; in DMF solution (77 K), g∥ = 2.233, g⊥ = 2.045, A∥ = 188 × 10−4 cm−1, g∥/A∥ = 119 cm, G = 5.2. Electrochemical data (DMF, 0.1 M TBAP): cyclic voltammetry, E1/2 = −0.077 V, ΔEp = 100 mV, ipa/ipc = 0.7, D = 7.7 × 10−6 cm2 s−1, and differential pulse voltammetry, E1/2 = −0.063 V.
![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 1519 ν(C
N), 1519 ν(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) C), 852 νC–H(H–C
C), 852 νC–H(H–C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N), 724, 779 ν(C–H), 1090, 622 ν(ClO4−), 538, 571 ν(Cu–N). UV-Visible data in DMF [λmax/nm (εmax/dm3 mol−1 cm−1)]: 675 (280), 856 sh, 321 (7895), 293 (32
N), 724, 779 ν(C–H), 1090, 622 ν(ClO4−), 538, 571 ν(Cu–N). UV-Visible data in DMF [λmax/nm (εmax/dm3 mol−1 cm−1)]: 675 (280), 856 sh, 321 (7895), 293 (32![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 420), 268 (49
420), 268 (49![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 330). EPR data: in solid (RT) giso = 2.172; in DMF solution (77 K), g∥ = 2.233, g⊥ = 2.042, A∥ = 189 × 10−4 cm−1, g∥/A∥ = 118 cm, G = 5.5. Electrochemical data (DMF, 0.1 M TBAP): cyclic voltammetry, E1/2 = −0.072 V, ΔEp = 86 mV, ipa/ipc = 0.9, D = 8.2 × 10−6 cm2 s−1, and differential pulse voltammetry, E1/2 = −0.059 V.
330). EPR data: in solid (RT) giso = 2.172; in DMF solution (77 K), g∥ = 2.233, g⊥ = 2.042, A∥ = 189 × 10−4 cm−1, g∥/A∥ = 118 cm, G = 5.5. Electrochemical data (DMF, 0.1 M TBAP): cyclic voltammetry, E1/2 = −0.072 V, ΔEp = 86 mV, ipa/ipc = 0.9, D = 8.2 × 10−6 cm2 s−1, and differential pulse voltammetry, E1/2 = −0.059 V.
X-ray structure crystallography
The single crystal of [Cu(L2)(phen)](ClO4)2 (2) (CCDC no. 2143125) in a green color suitable for the determination of X-ray structure was acquired upon slow evaporation at 5 °C after a couple of weeks in a MeOH![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeCN (2
MeCN (2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 v/v) mixture. A single crystal of the size 0.350 × 0.320 × 0.210 mm3 was chosen under the polarizing microscope and then mounted on the glass fiber. Diffraction data for the complex were collected at 298 K using a Bruker AXS-KAPPA APEX II diffractometer (Mo-Kα radiation, λ = 0.71073 Å) in the θ range 2–25°. Unit cell parameters were determined using the least-squares method based on all reflections with F2 > 2σ(F2) with Bruker SMART software. Cell refinement, correction for Lorentz and polarization effects, integration of intensities, and absorption corrections were performed using Bruker SAINT and SADABS. The structure was solved using the direct method (SIR-92)31 and refined using a full-matrix least-squares method (SHELXL-2014).32 All the calculations were carried out using the programs in the WinGX module.33 Non-hydrogen atoms were refined anisotropically. Cl1 and O1, O2, O3, and O4 were found to be disordered over two positions, and the occupancy factors were assigned to 0.775(10) and 0.225(10). O5, O6, O7, and O8 were also disordered over two positions, and the occupancy factors were 0.506(8) and 0.494(8). All the hydrogen atoms were placed geometrically and refined with the riding model. The final difference Fourier maps displayed no peaks of chemical importance. The graphic representation of the molecule was prepared using the ORTEP3 program for Windows34 with 40% probability displacement ellipsoids.
1 v/v) mixture. A single crystal of the size 0.350 × 0.320 × 0.210 mm3 was chosen under the polarizing microscope and then mounted on the glass fiber. Diffraction data for the complex were collected at 298 K using a Bruker AXS-KAPPA APEX II diffractometer (Mo-Kα radiation, λ = 0.71073 Å) in the θ range 2–25°. Unit cell parameters were determined using the least-squares method based on all reflections with F2 > 2σ(F2) with Bruker SMART software. Cell refinement, correction for Lorentz and polarization effects, integration of intensities, and absorption corrections were performed using Bruker SAINT and SADABS. The structure was solved using the direct method (SIR-92)31 and refined using a full-matrix least-squares method (SHELXL-2014).32 All the calculations were carried out using the programs in the WinGX module.33 Non-hydrogen atoms were refined anisotropically. Cl1 and O1, O2, O3, and O4 were found to be disordered over two positions, and the occupancy factors were assigned to 0.775(10) and 0.225(10). O5, O6, O7, and O8 were also disordered over two positions, and the occupancy factors were 0.506(8) and 0.494(8). All the hydrogen atoms were placed geometrically and refined with the riding model. The final difference Fourier maps displayed no peaks of chemical importance. The graphic representation of the molecule was prepared using the ORTEP3 program for Windows34 with 40% probability displacement ellipsoids.
Theoretical methods
The coordination geometries of copper(II) mixed ligand complexes in the ground state with doublet spin state were optimized using density functional theory (DFT) at the B3LYP level of theory by employing the Gaussian 09 program package.35 The calculations were administered with a mixed basis set (B1) of LANL2DZ for the copper metal atom, which contains a relativistic effective core potential with a valence basis set and 6-31G* for the remaining atoms.36–38 The normal mode analyses were performed to examine the minimal energy nature of the geometry. Copper(II) complexes 1 and 2 were considered mono-positive cations, as the copper metal was considered in the +2 oxidation state and doublet spin state, while the ligand was considered a positive ligand. The salvation of the complexes was achieved by employing the CPCM method using methanol as the solvent.Charge distribution and molecular orbital analyses
We used the natural bond orbital (NBO)39 approach to calculate the atomic charge distribution on each complex atom. From the NBO analysis, we can predict electron delocalization, atomic charge distribution, and intra-atomic interaction of atoms within a compound.39 The energies of the frontier molecular orbitals (FMOs),40 the highest and second-highest occupied molecular orbitals (EHOMO and EHOMO−1), and the lowest and second-lowest unoccupied molecular orbitals (ELUMO and ELUMO+1) were calculated. Some quantum chemical descriptors, including ionization potential (IP),41 electron affinity (EA),41 band gap (ΔE), chemical hardness (η),42 global softness (S),42 electrochemical potential (μ),43 electrophilicity index (ω),44 and electronegativity (χ),43 were estimated as outlined in eqn (1)–(8).| IP = −EHOMO | (1) |
| EA = −ELUMO | (2) |
| ΔE = ELUMO − EHOMO | (3) |
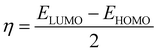 | (4) |
| S = 1/η | (5) |
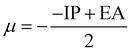 | (6) |
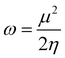 | (7) |
| χ = −μ | (8) |
DNA and protein binding experiments
Solutions of DNA in 5 mM Tris HCl/50 mM NaCl buffer gave a ratio of UV absorbances at 260 and 280 nm, A260/A280, of 1.9,45 indicating that the DNA was sufficiently free of protein. The stock solution of protein (1.0 × 10−4 mol L−1) was prepared by dissolving the solid BSA in 0.05 M phosphate buffer at pH 7.4, stored at 0–4 °C in the dark for about a week, and then diluted to 1.0 × 10−6 mol L−1 using phosphate buffer (pH 7.4, 0.05 M) when used. The concentration of BSA was determined from optical density measurements using the value of molar absorptivity of ε280 = 44![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 720 M−1 cm−1.46 The detailed procedure for the spectroscopic experiments of DNA and protein binding, as well as the electrochemical experiments for DNA binding, is described in the SI.
720 M−1 cm−1.46 The detailed procedure for the spectroscopic experiments of DNA and protein binding, as well as the electrochemical experiments for DNA binding, is described in the SI.
DNA cleavage experiments
The cleavage of DNA in the absence of activating agents was monitored using agarose gel electrophoresis. A typical reaction mixture containing pUC19 supercoiled phage DNA (form I, 20 µM) and copper(II) complex in 2% DMF/5 mM Tris-HCl/50 mM NaCl buffer (pH 7.1) was incubated at 37 °C for 1 h. After the incubation period, the reaction was quenched by keeping the samples at −20 °C, followed by the addition of loading buffer (0.025 mg bromophenol blue, 1 mL glycerol, and 1 mL MilliQ water). This was then loaded on a 1% agarose gel containing ethidium bromide (2.54 µM in the gel and buffer). The gels were run at a constant voltage of 40 V for 3 h in the 1 × TBE buffer (TBE = Tris-Borate-EDTA buffer) containing ethidium bromide. After washing with distilled water, the gels were visualized under a UV transilluminator, and the bands were documented and quantified using a BioRad Gel Doc 1000 apparatus interfaced with a computer. The cleavage efficiency was measured by determining the ability of the complex to convert the supercoiled DNA (SC) to a nicked circular form (NC) and linear form (LC). To identify the reactive oxygen species (ROS) involved in the cleavage reaction, radical scavengers such as hydroxyl radical (DMSO, 20 μM), singlet oxygen (NaN3, 100 μM), superoxide (SOD, 0.5 units), and catalase (0.5 units) were introduced. Additionally, methyl green (100 μM) is used to determine the groove binding nature of copper(II) complexes.Religation studies
An enzymatic study was carried out using T4 DNA ligase to determine whether the cleaved products were consistent with the hydrolysis of the phosphodiester linkages in DNA. For the religation tests, the solution was incubated for 18 h at 16 °C prior to gel electrophoresis. The nicked circular (NC) DNA obtained from the hydrolytic cleavage reaction was recovered from the agarose gel using a phenol extraction method and purified by ethanol precipitation. This was followed by the addition of a 10× ligation buffer and T4 DNA ligase (4 units) to the purified NC DNA.Cell line
The human cervical cancer cell line (HeLa), human lung epithelial adenocarcinoma cell line (A549), normal mouse embryonic fibroblast cells (NIH 3T3), and human lung epithelial normal cell line (L132) were acquired from the National Centre for Cell Science (NCCS), Pune. They were grown in Eagle's minimum essential medium containing 10% fetal bovine serum (FBS). The cells were preserved at 37 °C, 5% CO2, 95% air, and 100% relative humidity.47 Maintenance cultures were channelized weekly, and the culture medium was altered twice a week.Cell culture
To make single-cell suspensions, the monolayer cells were detached with trypsin-ethylenediaminetetraacetic acid (EDTA), and viable cells were counted using a hemocytometer. They were diluted with a medium containing 5% FBS to give a final density of 1 × 105 cells per mL. The cell suspension of one hundred microlitres per well was seeded into 96-well plates at a plating density of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per well. They were incubated to allow for cell attachment at 37 °C, 5% CO2, 95% air, and 100% relative humidity. The cells were treated with serial concentrations of the test samples after 24 h. They were dissolved in dimethylsulfoxide (DMSO), and an aliquot of the sample solution was diluted twice to the desired final maximum test concentration with a serum-free medium. Additionally, four serial dilutions were made to give a total of five sample concentrations. Aliquots of 100 µl of different sample dilutions were added to the suitable wells already containing 100 µl of the medium, resulting in the required final sample concentrations. The plates were incubated, followed by sample addition at 37 °C, 5% CO2, 95% air, and 100% relative humidity for an additional 48 h. The medium alone served as a control, and triplicate was maintained for all concentrations.
000 cells per well. They were incubated to allow for cell attachment at 37 °C, 5% CO2, 95% air, and 100% relative humidity. The cells were treated with serial concentrations of the test samples after 24 h. They were dissolved in dimethylsulfoxide (DMSO), and an aliquot of the sample solution was diluted twice to the desired final maximum test concentration with a serum-free medium. Additionally, four serial dilutions were made to give a total of five sample concentrations. Aliquots of 100 µl of different sample dilutions were added to the suitable wells already containing 100 µl of the medium, resulting in the required final sample concentrations. The plates were incubated, followed by sample addition at 37 °C, 5% CO2, 95% air, and 100% relative humidity for an additional 48 h. The medium alone served as a control, and triplicate was maintained for all concentrations.
Cell viability assay
The cell viability was carried out using the MTT assay.48 Complex in the concentration range of 0.25–10 μM (in HeLa), 0.2–1.0 μM (in A549), 25–500 μM (in NIH 3T3) or 25–250 μM (in L132) dissolved in 2% DMF/5 mM Tris-HCl/50 mM NaCl buffer at pH 7.1 was added to the wells 24 h after seeding of 1 × 105 cells per well in 100 μL of fresh culture medium. After 48 h, 15 µL of 3-[4,5-dimethylthiazol-2-yl]2,5-diphenyltetrazolium bromide (MTT, 5 mg mL−1) in phosphate-buffered saline (PBS) was added to each well and incubated at 37 °C for 4 h. The formed formazan crystals were solubilized in 100 µL of DMSO after the medium with MTT was flicked off. The microplate reader was used to measure the absorbance at 570 nm. Data were collected in three replicates, and the percentage of cell viability and the percentage of cell inhibition were calculated using the following formulas:| % Cell viability = [As]/[Ac] × 100 |
| % Cell inhibition = [100 − ([As]/[Ac])] × 100. |
A nonlinear regression graph was plotted between % cell inhibition and log concentration, and IC50 was calculated using GraphPad Prism software.
Studies on intracellular reactive oxygen species (ROS)
Studies on cell cycle arrest
The A549 cells were seeded in six-well plates with high glucose DMEM after the cells had grown, and the cells were treated with two copper(II) complexes in their respective IC50 concentrations in the medium for 24 h. After 24 h of incubation, the cells were trypsinized and resuspended in complete media. The cells were collected and centrifuged at 1000 rpm for 5 min. The cell pellet was washed with phosphate-buffered saline (PBS) twice and subsequently fixed with 1 mL of 70% ice-cold ethanol overnight at 4 °C. Following the ice-cold ethanol, the cell pellet was washed twice with cold PBS and added to 10 μL of RNase A at 10 g mL−1 of concentration, which was then incubated for 30 min and washed with PBS at the end. The cells were then incubated in 1 mL of PBS with 50 μL of propidium iodide (PI) (1 mg mL−1 of stock) for 30 min in the dark. Then, the cells were analyzed to check the cell cycle phase using the FACSVerse flow cytometer (Becton Dickinson).50Studies on the induction of apoptosis
Results and discussion
Synthesis and general properties
Schiff base ligands L1 and L2 were synthesized by condensing 2-aminomethylpyridine or 2-aminoethylpyridine and 5-methylpyridine-2-carboxaldehyde. The copper(II) complexes of the ligands were prepared by the reaction of copper(II) acetate monohydrate, 1,10-phenanthroline, and L1 or L2 in equimolar quantities using ethanol as a solvent. Both complexes 1 and 2 were isolated as green-colored crystalline compounds in good yields. Based on the elemental analysis, the complexes were formulated as [Cu(L1/L2)(phen)](ClO4)2, which was confirmed by a single-crystal X-ray structure determination of 2. The ESI mass spectral data of 1 and 2 display base peaks at m/z 391.25 and 405.27, respectively, which can be attributed to the complex cations [Cu(L1)(phen)]2+ and [Cu(L2)(phen)]2+, respectively. This is supported by the values of molar conductivity in DMF (ΛM/Ω−1 cm2 mol−1: 1, 203; 2, 205), which falls in the range52 for 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2 electrolytes. The formation of the dicationic complexes was evidenced by the intense absorption of the ClO4− stretches at 1090 and 622 cm−1 in the IR spectrum of the complexes. As expected, the band belonging to νimine(C
2 electrolytes. The formation of the dicationic complexes was evidenced by the intense absorption of the ClO4− stretches at 1090 and 622 cm−1 in the IR spectrum of the complexes. As expected, the band belonging to νimine(C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N) (L1, 1610; L2, 1611 cm−1) was shifted to 1585 cm−1 due to the coordination of imine nitrogen with copper(II). The sharp bands observed at 530–571 cm−1 can be assigned to ν(Cu–N), probably originating from the coordination of L1 (1) or L2 (2) and phen (1 and 2) ligands.
N) (L1, 1610; L2, 1611 cm−1) was shifted to 1585 cm−1 due to the coordination of imine nitrogen with copper(II). The sharp bands observed at 530–571 cm−1 can be assigned to ν(Cu–N), probably originating from the coordination of L1 (1) or L2 (2) and phen (1 and 2) ligands.
Single crystal X-ray structure of [Cu(L2)(phen)](ClO4)2
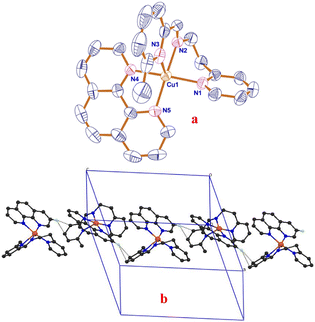
The ORTEP representation of the complex, [Cu(L2)(phen)](ClO4)22, including the coordinated atom numbering scheme, is shown in Fig. 1a. The crystal structure refinement data, selected bond lengths and bond angles are listed in Table S1 and Table 1, respectively. The asymmetric unit of 2 contains one dicationic complex molecule and two perchlorate anions. The copper atom in the complex is coordinated by all three nitrogen atoms (N1, N2, and N3) of tridentate ligand L2 and both nitrogen atoms (N4 and N5) of phen. The value of the structural index τ of 0.06 [τ = (β − α)/60, where α = N2–Cu1–N5 = 165.67° and β = N1–Cu1–N4 = 169.43°] reveals that the coordination geometry around copper(II) is best described as distorted square pyramidal.53 The corners of the square plane of the geometry are occupied by N1 and N2 nitrogen atoms of the facially coordinated L2 and N4 and N5 nitrogen atoms of phen. The N3 nitrogen atom of L2 occupies the apical position at a distance (2.250(3) Å) longer than the equatorial nitrogen atoms (1.991–2.022 Å), as a consequence of the presence of two electrons in the dz2 orbital of copper(II). It is interesting to note that the τ value of 2 is very much lower than that of its dipica analogue [Cu(dipica)(phen)]2+ (τ, 0.77),54 indicating that both the apical coordination of 6-methypyridine (mpy) nitrogen atom (N3) of L2 at a longer distance and the six-membered chelate ring formed by L2 decrease the steric congestion at copper(II) and change (i) the meridional fashion to facial and (ii) coordination geometry from trigonal bipyramidal towards square pyramidal. Additionally, the incorporation of the six-membered chelate ring and axial coordination of mpy facilitates stronger coordination of pyridyl N1 nitrogen atom (Cu–Npy: 2, 2.007(2); [Cu(dipica)(phen)]2+, 2.058(5) Å) and phen N4 and N5 nitrogen atoms (Cu–Nphen: 2, 2.022(2) and 2.012; [Cu(dipica)(phen)]2+, 2.157(5) Å). Notably, the incorporation of phen in the coordination sphere of 2 (Scheme 1) displaces the axially coordinated imine nitrogen of L2 (Scheme 1a) to a more strongly bound equatorial position (Scheme 1b; 1.991(3) Å) and decreases the steric congestion at copper(II) resulting in the distorted square pyramidal geometry, as revealed by the increase in trans bond angles (N1–Cu1–N4 = 169.43° and N2–Cu1–N5 = 165.67°). Another important structural observation is the inter-pair C–H⋯π anti-parallel non-covalent interactions (Fig. 1b) involving C(23)–H of the pyridine ring of phen and the centroid (Cg) of the mpy ring of L2 (C(23)–H⋯Cg(mpy) = 2.942 Å), which is probably responsible for the stabilization of the distorted square pyramidal geometry.55| X-ray | DFTa | ||
|---|---|---|---|
| 2 | 2 | 1 | |
| a Calculated from the B3LYP level of theory with mixed basis sets LANL2DZ/6-31G* using methanol as a solvent employing the CPCM method. | |||
| Cu(1)–N(1) | 2.007(2) | 2.067 | 2.132 |
| Cu(1)–N(2) | 1.991(3) | 2.045 | 1.977 |
| Cu(1)–N(3) | 2.250(3) | 2.358 | 2.224 |
| Cu(1)–N(4) | 2.022(2) | 2.073 | 2.022 |
| Cu(1)–N(5) | 2.012(3) | 2.075 | 2.236 |
| N(5)–Cu(1)–N(1) | 96.15(11) | 97.28 | 95.34 |
| N(5)–Cu(1)–N(2) | 165.67(12) | 166.31 | 171.67 |
| N(5)–Cu(1)–N(3) | 116.18(11) | 116.95 | 104.48 |
| N(5)–Cu(1)–N(4) | 81.64(11) | 80.96 | 79.44 |
| N(1)–Cu(1)–N(2) | 83.26(10) | 81.58 | 78.77 |
| N(1)–Cu(1)–N(3) | 101.57(10) | 101.93 | 149.54 |
| N(1)–Cu(1)–N(4) | 169.43(11) | 163.71 | 103.76 |
| N(2)–Cu(1)–N(3) | 77.82(11) | 76.49 | 78.78 |
| N(2)–Cu(1)–N(4) | 96.30(10) | 96.28 | 107.56 |
| N(3)–Cu(1)–N(4) | 88.62(10) | 93.19 | 102.47 |
DFT-optimized structure and quantum chemical descriptors
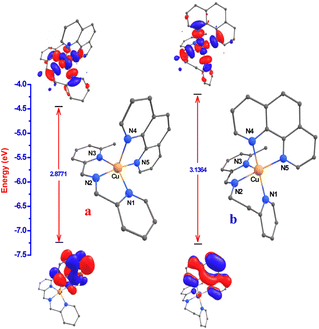
The theoretical calculations help validate the experimental results and provide further insight into the complexes. In the present study, DFT calculations were incorporated to provide a comparison between experimental and theoretical values.56 Geometry optimization for 1 and 2 in methanol was carried out by employing the B3LYP, 6-31G*/LANL2DZ levels, and the Gaussian 09 computer software.36 The bond lengths, bond angles, optimized energies, frontier molecular orbital (FMO), and highest occupied molecular orbital (HOMO)–lowest unoccupied molecular orbital (LUMO) energy gap are also determined. The optimized structures of 1 and 2 (Fig. 2) in methanol show the donor sites N1, N2, and N3 of L1 and N5 of phen (1) or N1 and N2 of L2 and N4 and N5 of phen (2) coordinated to Cu(II) in the four corners of the square plane. Consequently, the N4 of phen (1) or the N3 of L2 (2) occupies the axial position in the coordination sphere of the Cu(II) ion with square pyramidal geometry. The geometrical parameters, like calculated bond lengths and bond angles, are in good agreement with the single-crystal X-ray data of 2 (Table 1). All strain energy minimized structures reproduced the observed X-ray structure to a maximum tolerance of 0.108 Å for 2 in the Cu–N bond lengths. The maximum deviation in the N–Cu–N bond angle for 2 is 5.72°. The small discrepancies in bond lengths and bond angles are attributable to non-covalent C–H⋯π inter-pair anti-parallel interactions within the lattice in the solid-state, which are not observed in the solution. Generally, in such instances, the normal discrepancy is higher than the mean between the observed and calculated structures,57 but the average structure is sufficient to represent the observed structure.Interestingly, the tridentate 3N L1 ligand coordinated meridionally, while L2 coordinated facially to copper(II) due to a rigid five-membered (L1) or flexible six-membered (L2) chelate ring. As a result, the geometry of the optimized structure in 1 differed from that in 2. Two longer estimated bond angles (1, N5–Cu–N2 and N1–Cu–N3; 2, N5–Cu–N2 and N1–Cu–N4) were used to determine the trigonality index (τ) for 1 (0.37) and 2 (0.04). The geometry of the copper center in 1 is severely distorted, while 2 is slightly distorted square pyramidal, and the τ for 2 is identical to the experimental result (τ, 0.06). In addition, the difference in the twist angle between the N–Cu–N (L1 or L2) and N–Cu–N (phen) planes estimated for 1 (1.3) and 2 (1.9) is due to the higher aromatic ring delocalization in the phen ring of 2 compared to the phen ring of 1, which enforces the planarity around the Cu(II) center in 2 because of the flexible six-membered chelate ring. This has beneficial effects on the DNA binding ability, mode of binding, and the DNA cleavage efficiency of the complex (cf. below).
The HOMO and LUMO are the main orbitals among FMOs that take part in chemical reactions. The HOMO acts as an electron donor, while the LUMO acts as an electron acceptor, and the gap between HOMO and LUMO characterizes the ability of electron transfer or chemical reactivity of the molecule.58 It has been found that HOMO is largely distributed over the phen ligand in 1, on the metal, the methylpyridine (mpy) nitrogen atom of L2, and the phen ligand in 2. LUMO is distributed over the metal center, imine, pyridine (py), and mpy nitrogen atoms of L1 or L2 and phen nitrogen atoms in 1 or 2 (Fig. 2). The EHOMO of 1 (−7.199) > 2 (−7.260) reveals that L1 releases more electrons; hence, these are involved in the strongest σ-bonding to Cu(II). Thus, the σ-donor capability of the L1 is tuned by the disparity in the planarity of phen because of the rigid 555 chelate rings in 1 rather than the flexible 655 chelate rings in 2, which reflects the presence of synergy between L1 or L2 and phen ligands when bound to the metal. The ELUMO of 1 (−4.323) < 2 (−4.124) indicates that the π* orbital of phen in 2 is delocalized more than that of phen in 1. Therefore, the more delocalized π* orbital of phen in 2 is involved in a stronger π back-bonding with Cu(II) to stabilize the Cu(I) oxidation state more than that observed with the phen in 1. This is in good agreement with the observed trend in E1/2 values and illustrates that the delocalization of the π* orbital of phen is important in stabilizing the lower oxidation of Cu(I) (cf. below). There is a significant energy difference between ELUMO+1 and ELUMO; LUMO has around 1.4 (1) or 1.5 (2) times the value of LUMO+1, indicating a high-energy transition from LUMO+1 to LUMO. However, EHOMO−1 and EHOMO values are comparable, indicating that these orbital transitions differ very little (Table 2).
| Parameters | 1 | 2 | Cisplatin |
|---|---|---|---|
| a In eV. b In eV−1. | |||
| Optimized energy (×104)a | −3.905 | −4.012 | |
| HOMOa | −7.199 | −7.260 | −6.618 |
| HOMO−1a | −7.205 | −7.270 | |
| LUMOa | −4.323 | −4.124 | −1.970 |
| LUMO+1a | −3.110 | −2.824 | |
| Energy gap, ΔEa | 2.877 | 3.136 | 4.647 |
| IPa | 7.193 | 7.260 | 6.618 |
| EAa | 4.323 | 4.124 | 1.970 |
| η | 1.438 | 1.568 | 2.324 |
| σ | 0.695 | 0.637 | 0.430 |
| μ | −5.761 | −5.692 | −4.294 |
| S | 0.347 | 0.319 | 0.215 |
| ω | 11.537 | 10.330 | 3.967 |
| χ | 5.761 | 5.692 | 4.294 |
| N | 0.086 | 0.096 | 0.252 |
| ΔNmax | 4.006 | 3.630 | 1.848 |
In most stable compounds, the HOMO–LUMO gap, viz. 1 (2.877 eV) < 2 (3.136 eV), defines the properties of the electrons and explains the eventual charge transfer interactions taking place within the molecules. This reflects the ability of electrons to move from occupied orbitals to unoccupied orbitals, thereby providing a fundamental basis for judging the chemical reactivity, selectivity, and stability of the compounds. As a result, the energies of the FMOs were examined to study some quantum chemical descriptors,59–63 such as ionization potential (IP = −EHOMO), electron affinity (EA = −ELUMO), energy gap (ΔE = ELUMO − EHOMO), chemical hardness (η = ELUMO − EHOMO/2), absolute softness (σ = 1/η), global softness (S = 1/2η), electrochemical potential (μ = −(IP + EA)/2), electrophilicity index (ω = μ2/2η), nucleophilicity index (N = 1/ω), additional electronic charges (ΔNmax = σ/η), and electronegativity (χ = −μ) during excitation of 1 and 2. The quantum chemical descriptors of 1 and 2 are compared with the special parameters of cisplatin64 (Table 2) to determine the anticancer properties. A high EHOMO denotes the donation of an electron by a suitable molecule, while a low ELUMO implies electron acceptance. As EHOMO increases and ELUMO decreases, this indicates an increase in biological activity. Electron freedom, or the energy gap between HOMO and LUMO, is crucial in determining biological reactivity since activity increases as ΔE decreases. The ΔE of soft molecules (σ) is small, while that of hard molecules (η) is large. Consequently, soft molecules have more biological activity than hard molecules. A molecule can readily donate electrons to coordinate the proper structure if the global electronegativity (χ) is small or the chemical potential (μ) is large. This indicates that the electron delocalizes the entire structure. As the nucleophilicity index (N) increases and the electrophilicity index (ω) decreases, biological activity also increases. The greater the ΔNmax value of a compound, the greater its biological activity. The increasing biological activity of the compound is implied by its increasing global softness (S) value. The computed values indicate the possible biological activity of 1 and 2, and the most advantageous parameters are ΔE, σ, η, ΔNmax, and S. For the present copper(II) complexes, their anticancer activities are greater than those of cisplatin. It is observed from the above parameters that they are good candidates for cancer drugs.
Electronic and EPR spectral and redox properties
The electronic spectra of 1 and 2 are very similar to each other (Fig. S1 and S2) and show a broad low energy ligand field (LF) band (λmax: 1, 684; 2, 675 nm), followed by a shoulder (λmax: 1, 896; 2, 856) in the visible region revealing a square-derived coordination geometry around Cu(II) with slight distortion.65 The observed molar absorptivity of 2 (εmax, 280 M−1 cm−1) is higher than that of 1 (εmax, 100 M−1 cm−1) due to the presence of a flexible six-membered chelate ring, which increases the distortion in square-based Cu(II) coordination geometry of 2. The intense absorption bands observed in the range of 268–323 nm are attributed to the intraligand π → π* transitions.The polycrystalline EPR spectra of 1 and 2 are isotropic (Fig. S3 and S4). The frozen solution EPR spectra of both the complexes exhibit spectral features typical of mononuclear Cu(II) species (Fig. S5 and S6). They are axial with g∥ > g⊥ > 2.0 and G = [(g∥ − 2)/(g⊥ − 2)] = 5.2–5.5, suggesting the presence of a dx2−y2 ground state in copper(II) located in square-based geometries.66 A square-based CuN4 chromophore is expected67,68 to show a g∥ value of 2.200 and A∥ value in the range of 180–200 × 10−4 cm−1, and the distortion or incorporation of strong axial interaction is expected to enhance the g∥ value and decrease the A∥ value.67,68 Therefore, the observed values of g∥ (2.233) and A∥ (188–189 × 10−4 cm−1) are consistent with the CuN5 chromophore of the complexes in the solution. The g∥/A∥ values (118–119 cm) fall within the range (105–135 cm) expected for complexes with square planar geometry,69 suggesting that there is a slight distortion from planarity, which is evident from the X-ray crystal structure of 2.
The cyclic (CV) and differential pulse voltammetric (DPV) techniques were employed to investigate the redox behavior of 1 and 2 in the DMF solution on stationary glassy carbon electrodes (Fig. S7 and S8). They display well-defined cathodic and anodic waves revealing the reversible CuII/CuI redox couple owing to the linearity of the plots of ipcversus ν1/2 (D: 1, 7.7; 2, 8.2 × 10−6 cm2 s−1) and the value of the peak current ratio (ipa/ipc: 1, 0.7; 2, 0.9). However, the value of the difference in peak potentials (ΔEp: 1, 100; 2, 86 mV) is close to the Fc/Fc+ couple (ΔEp, 86 mV; E1/2, 0.406 V vs. SCE) under similar experimental conditions. Interestingly, the less negative redox potential, E1/2 of 2, is higher, and peak potential separation, ΔEp, is lower than that of 1 (E1/2 and ΔEp: 1, −0.077 V and 100 mV; 2, −0.072 V and 86 mV) for the CuII/CuI couple. This indicates that the smallest structural reorganization between copper(II) and copper(I) species renders the Cu(II) to Cu(I) electron transfer more facile. This is due to the presence of a flexible six-membered chelate ring forming a boat conformation, which is attracted by the steric hindrance between the 6-methyl group in the pyridine moiety of L2 and phen ligands, thereby distorting the Cu(II) coordination geometry.70 The redox potentials of the CuII/CuI couple from the DPV (Fig. S9 and S10) are −0.063 V (1) and −0.059 V (2) vs. SCE.
Stability studies
Both complexes are soluble in DMSO and 2% DMF/5 mM Tris-HCl/50 mM NaCl buffer (pH 7.1), stable in air, and non-hygroscopic. The complexes in DMSO, buffer, and cell growth medium (DMEM) were subjected to UV-visible spectral analyses at various time intervals (0, 24, and 48 h). After 0, 24, and 48 h in solution, no new absorption peaks emerged for 1 and 2, and there was no discernible red or blue shift in the position of the absorption peaks (Fig. S11 and S12). However, after 24 h, the intensity of the absorption peaks for 1 and 2 decreased, indicating a decrease in the concentration. Consequently, the findings show that the structures of 1 and 2 did not change significantly in DMSO, buffer, or DMEM, which establishes the basis to conduct further biological studies.DNA binding properties
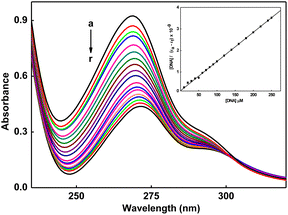
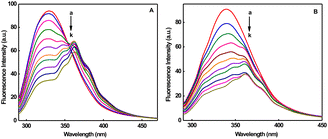
Upon incremental addition of CT DNA to 1 (Fig. 3) and 2 (Fig. S13), the ligand-centered band (268 nm) shows a decrease in molar absorptivity (hypochromism: 1, ∼60; 2, ∼32%) with red shift (1, 3; 2, 2 nm) at R = 25 (R = [DNA]/[Cu]). The observed hypochromic effect, accompanied by a slight red shift, strongly indicates the binding of 1 and 2 to CT DNA through a partial intercalative interaction (Table S2).71 The intrinsic binding constant (Kb) value (Fig. 3 and Fig. S13) demonstrates the strong DNA binding affinity of the complexes and follows the order 1 (1.36 × 105 M−1) > 2 (1.06 × 105 M−1). Compared to the partially intercalating metal complex {[Ru(bipy)2(dppz)]2+ (Kb, 1.35 × 106 M−1)}, the Kb values of 1 and 2 are 10 times lower.72 However, relative to the previously published ternary copper(II) complexes of phen bases, [Cu(pmdt)(phen)](ClO4)2 (Kb, 3.1 × 103 M−1)73 and [Cu(dipica)(phen)](ClO4)2 (Kb, 2.7 × 103 M−1),74 the noted DNA binding affinity of these complexes is 100 times greater. In contrast, 1 and 2 exhibit comparable DNA binding affinities when compared to the established intercalator (EthBr, Kb, 4.94 × 105 M−1)72 and mixed ligand copper(II) complexes that contain phen, [Cu(cur)(phen)(ClO4)] (Kb, 7.69 × 105 M−1) and [Cu(acac)(phen)(ClO4)] (Kb, 3.18 × 105 M−1).75 The higher binding strength in 1 is attributed to the presence of a rigid five-membered chelate ring. The rigidity enhances the partial intercalative interaction of the phen ligand between the base pairs of DNA. The hydrophobic interaction of the 6-methyl group with the hydrophobic DNA surface further supports an increase in the DNA binding affinity56,76 in 1. The lower binding affinity of 2 is due to the presence of a flexible six-membered chelate ring. The flexibility reduces the efficiency of the partial intercalative interaction of the phen ligand, and the decrease in the hydrophobic interaction of the 6-methyl group in 2.The fluorescence intensity of EthBr bound to DNA at 594 nm (λex, 520 nm) shows a remarkable decrease (Fig. S14 and S15) with the addition of 1 (78%) or 2 (77%), indicating that DNA-bound EthBr molecules are released into solution.77 Such fluorescence quenching may be caused by the strong partial intercalative interaction of 1 or 2 (cf. above), leading to the displacement of DNA-bound EthBr. The slope of the plot of I0/I versus [complex] gives KSV (Fig. S14 and S15, inset). The fluorescence quenching follows Stern–Volmer behavior (KSV: 1, 4.26; 2, 4.12 × 105 M−1). The apparent DNA binding constant78 values (Kapp: 1, 5.20; 2, 5.00 × 106 M−1) suggest that 1 intercalates more strongly than 2. Compared to copper(II) complexes based on phen such as [Cu(L)(phen)](ClO4)2 (Kapp, ∼105 M−1),79 [Cu(L1–L5)(phen)-(H2O)](ClO4)2 (Kapp, 3.60–4.18 × 105 M−1),80 [Cu(phdpa)(phen)-(H2O)](ClO4)2 (Kapp, 9.59 × 104 M−1)81 and [Cu(ETDPA)-(phen)](ClO4)2 (Kapp, 3.40 × 103 M−1),82 the Kapp values of 1 and 2 are 10–1000 times higher.
The observed circular dichroic (CD) spectrum of CT DNA (2 × 10−5 M) consists of a positive band (270 nm) due to base stacking and a negative band (240 nm) due to helicity (Fig. S16 and S17), which is characteristic of DNA in the right-handed B form.56,83 When 1 or 2 is incubated with DNA at the 1/R (= [Cu complex]/[DNA]) value of 3, the CD spectrum of DNA (Fig. S16 and S17) changes both positive (vast increase in intensity; red-shift: 1 and 2, 3 nm) and negative bands (slight decrease in intensity; red-shift: 1 and 2, 4 nm), which is consistent with partial intercalative interaction84 of phen ring.
Both complexes display a well-defined cathodic and anodic wave within the time scale of the cyclic voltammetry (CV) experiments (2% DMF/5 mM Tris-HCl/50 mM NaCl buffer, pH 7.1). Redox-active complexes 1 and 2 show the reversible Cu(II)/Cu(I) couple (E1/2: 1, −0.079; 2, −0.073 V/ΔEp: 1, 99; 2, 89 mV/ipa/ipc: 1, 0.8; 2, 0.6). Upon the addition of CT DNA (R = [DNA]/[Cu] = 5), a significant reduction in cathodic and a slight decrease in anodic peak currents (Fig. S18 and S19) are observed due to the slow diffusion of the DNA-bound complex to the electrode surface.85 The observed redox potential (E1/2) shift for 1 is less positive (+2 mV) and for 2 is less negative (−4 mV), which implies that the binding of 1 and 2 to DNA is through partial intercalative interaction. Additionally, the E1/2 values from differential pulse voltammetry (DPV: 1, −0.063; 2, −0.058 V) for the Cu(II)–Cu(I) redox couple of the complexes (Fig. S20 and S21) are shifted (1, +1; 2, −6 mV), revealing a stronger DNA binding affinity. The shift in the value of the formal potential (ΔE°′) can be used to estimate the ratio of the equilibrium binding constant (K+/K2+) values of the Cu(I) and Cu(II) forms to DNA, which are calculated to be 1.0 (1) and 1.3 (2), respectively, suggesting their stabilization in the Cu(I) oxidation state upon binding to CT DNA.86
DNA cleavage properties
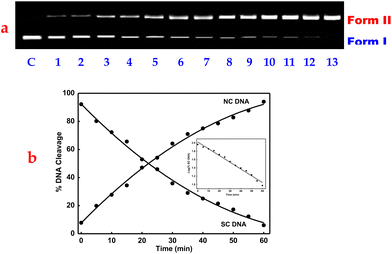
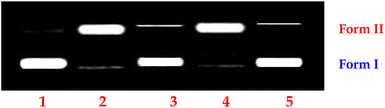
Upon incubation of the pUC19 plasmid DNA (20 μM in base pairs) and 1 or 2 (10–100 μM) in the absence of external additives for 1 h at 37 °C in 2% DMF/5 mM Tris-HCl/50 mM NaCl buffer (pH = 7.1), followed by electrophoresis, the fastest migration is observed for the SC DNA (form I). A single strand of SC DNA relaxes upon cleavage, resulting in slower-moving nicked circular (NC) DNA (form II; 43%), and no linear DNA (LC, form III) is discernible, indicating that neither of the DNA strands is cleaved by 1 or 2 (Fig. S22). The amount of form I decreases whereas that of form II increases when the concentrations of 1 or 2 are increased (100–160 μM) while keeping the DNA concentration constant (20 μM), causing the complete conversion to form II [97%; conc. 140 (1) or 130 μM (2)], which demonstrates that the cleavage is concentration-dependent. Due to the absence of form III in either 1 or 2, single-strand DNA scission was interpreted (Fig. 4).87The time-dependent cleavage of DNA by 1 or 2 was investigated {[complex] = 140 (1) or 130 (2) μM and [DNA] = 20 μM in bp} as a function of time (0–60 minutes). The amount of NC DNA increases as the reaction time is extended, while the amount of SC DNA steadily decreases (Fig. 5 and S23), and the rate of DNA cleavage reaches its maximum within 55 minutes. The appearance of NC DNA and the disappearance of SC DNA follow pseudo-first-order kinetic profiles and fit well into a single-exponential decay curve (Fig. 5 and S23). The copper(II) center may be crucial in fostering Lewis acid–base activity, which would increase the rate at which DNA cleaves.
When supercoiled pUC19 DNA was treated with methyl green, the cleavage reaction mediated by the complexes was not quenched (Fig. S24, lane 7), while it was completely cleaved to form II. This suggests that the complexes prefer to bind through partial intercalation rather than groove binding (cf. above).88 The mechanism by which 1 or 2 cleaves pUC19 DNA was examined (Fig. S24) and explained in the presence of singlet oxygen quencher NaN3 (100 μM), hydroxyl radical scavenger DMSO (20 μM), superoxide anion generator SOD (0.5 units), and H2O2 producer catalase (0.5 units) in an aerobic environment.89 As illustrated in Fig. S24, the mechanism of DNA cleavage by 1 or 2 is as follows: DNA cleavage activity is not affected by NaN3, DMSO, SOD, or catalase (lanes 3–6), ruling out the potential of cleavage by singlet oxygen, hydroxyl radical, superoxide, and H2O2, respectively.
This finding suggests that a hydrolytic mechanism may be the cause of the DNA cleavage response by 1 or 2. In general, two main processes are involved in DNA cleavage: oxidative cleavage and hydrolytic cleavage. In the presence of external reducing agents, the mixed-ligand Cu(II) complexes, including phen as a coligand with the CuN5 chromophore, like [Cu(L)(phen)](ClO4)2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 81 and [Cu(dipica)(phen)](ClO4)2,55 cleave plasmid DNA oxidatively. In most cases, this involves a photoactive or redox-active metal center and produces reactive singlet oxygen or hydroxyl radical species. This damages the sugar and/or base and produces fragmented species that are not religated. There are no disadvantages to hydrolytic cleavage without external additives because it is possible to religate the cleaved products enzymatically. Therefore, to ascertain the hydrolytic nature of the cleavage reaction, the NC form obtained from the cleavage of SC DNA was reacted with a T4 ligase enzyme.90,91 We observed 90% conversion of the NC DNA to its original SC form (Fig. 6, lanes 3 and 5), providing direct evidence in favor of the hydrolytic mechanism. Consequently, the hydrophobic interaction of the 6-methyl moiety supports the partial intercalation of planar phen with rigid 555 (1) or flexible 556 chelate rings (2). This leads to the distortion of the DNA double helix due to the increased DNA binding of 2 compared to 1, resulting in significant hydrolytic DNA cleavage activity92 at lower concentrations of 2 (130 μM) rather than 1 (140 μM).
81 and [Cu(dipica)(phen)](ClO4)2,55 cleave plasmid DNA oxidatively. In most cases, this involves a photoactive or redox-active metal center and produces reactive singlet oxygen or hydroxyl radical species. This damages the sugar and/or base and produces fragmented species that are not religated. There are no disadvantages to hydrolytic cleavage without external additives because it is possible to religate the cleaved products enzymatically. Therefore, to ascertain the hydrolytic nature of the cleavage reaction, the NC form obtained from the cleavage of SC DNA was reacted with a T4 ligase enzyme.90,91 We observed 90% conversion of the NC DNA to its original SC form (Fig. 6, lanes 3 and 5), providing direct evidence in favor of the hydrolytic mechanism. Consequently, the hydrophobic interaction of the 6-methyl moiety supports the partial intercalation of planar phen with rigid 555 (1) or flexible 556 chelate rings (2). This leads to the distortion of the DNA double helix due to the increased DNA binding of 2 compared to 1, resulting in significant hydrolytic DNA cleavage activity92 at lower concentrations of 2 (130 μM) rather than 1 (140 μM).
BSA interaction
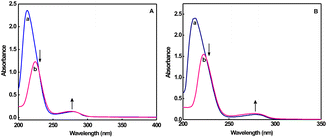
Tryptophan emission93 quenching experiments were carried out to measure the protein binding affinity of complexes by adding increasing concentrations (0–4 mM) of complexes to BSA (1 mM). At 300 K, the fluorescence intensity of BSA decreases to 63 (1), 62% (2), and at 310 K, it decreases to 69% (1, 2), with a red shift of 4–10 nm (Fig. 7 and S25). The observed Stern–Volmer plots (Fig. S26–S28) show a good linear relationship94 in which KSV (Table S3) increases as temperature rises (300 K: 1, 4.31; 2, 4.08 × 105 M−1 and 310 K: 1, 5.43; 2, 5.62 × 105 M−1), indicating that the most likely source of fluorescence quenching is a dynamic quenching mechanism. The obtained bimolecular quenching rate constant (kq) is in the order of 1013 M−1 s−1, which is 1000 times greater than the diffusion-controlled quenching maximum limit (2.0 × 1010 M−1 s−1).95 Moreover, when the temperature is increased (300 K, 1, 1.50; 2, 3.23 × 105 M−1 and 310 K, 1, 2.26; 2, 3.88 × 105 M−1), the effective quenching constant, Ka (Table S3), derived using the modified Stern–Volmer equation,96 increases.When the complexes are added, the 210 nm absorbance peak of BSA97 is dramatically reduced, with a red-shift (1, from 210 to 220 nm; 2, from 210 to 222 nm). Meanwhile, the absorption intensity at 280 nm has slightly increased (Fig. 8), indicating that additional aromatic acid residues have extended into the aqueous environment and destroyed the tertiary structure of BSA.98 These findings indicate that the interaction between copper(II) complexes and BSA is primarily a static quenching process caused by the formation of a ground-state complex (BSA-1 or BSA-2). The binding constant, Kb, increases with temperature (300 K: 1, 1.10; 2, 2.19 × 106 M−1 and 310 K: 1, 1.34; 2, 2.62 × 106 M−1), indicating that a stable BSA-1 or BSA-2 ground-state complex is formed99 (Table S3). Additionally, the value of n, which is close to 1 (Table S3), shows that the binding site in BSA is unique and that the complexes have easy access to the hydrophobic environment of the tryptophan residue.100 Since the values of ΔG° are negative, it is clear that the binding process is spontaneous, as shown in Table S3. The presence of significant hydrophobic forces of interaction between the ligands of copper(II) complexes and BSA is also supported by the positive values reported for both ΔH° and ΔS°.101 Covalent bonding could be considered in addition to the hydrophobic interaction. However, the obtained value for ΔH° (1, 70; 2, 74 kJ mol−1) is significantly lower than what would be predicted for the formation of covalent bonds, which should be ≥120 kJ mol−1.102
Cytotoxicity of copper(II) complexes
The antiproliferative activity in vitro against human cervical carcinoma (HeLa) and human lung epithelial adenocarcinoma (A549) cell lines, mouse embryonic fibroblasts (NIH 3T3) and human lung epithelial (L132) normal cells was carried out for 1, 2, cisplatin, free ligands, and copper-acetate using the MTT assay method. The activities of 1 and 2 are much higher than those of the corresponding free ligands in HeLa or A549 cells. They have exhibited a significant inhibitory potency against the proliferation of the HeLa (IC50: 24 h, 1.164 ± 0.085 (1); 1.372 ± 0.104 μM (2) and 48 h, 0.853 ± 0.046 (1); 1.023 ± 0.078 μM (2)) or A549 (IC50: 24 h, 0.496 ± 0.043 (1); 0.494 ± 0.041 μM (2) and 48 h, 0.199 ± 0.015 (1); 0.201 ± 0.016 μM (2)) cell line at a very low concentration in dose- and time-dependent manners. Since the phen co-ligand possesses higher lipophilicity103 to penetrate the cell membrane and act as a recognition element for non-covalent interaction with DNA, the IC50 values of 1 and 2 against the A549 cell line are marginally lower (∼2 to 3 times) than those of copper(II)-phen complexes [(phdpa)Cu(phen)(H2O)](ClO4)2 (IC50, 3.10 μM),84 [Cu(L)(phen)], and [Cu(sal)(phen)] (IC50, 1.42 μM).83 They are lower (∼6 to 16 times) than those of [(phdpa)Cu(phen)(H2O)](ClO4)2 (IC50, 3.10 μM),84 [Cu(L)(phen)](ClO4)2 (IC50, 3.55 μM),104 [Cu(phen)2(CH3CN)](ClO4)2 (IC50, 4.00 μM),105 [Cu(Aldpa)(phen)](ClO4)2 (IC50, 7.60 μM),106 [Cu(pabt)(phen)]ClO4 (IC50, 7.60 μM)107 and [Cu(ETDPA)(phen)](ClO4)2 (IC50, 8.00 μM).108 Interestingly, compared to [Cu(L1–L5)(phen)(H2O)](ClO4)2 (IC50, 3.85–46.75 μM),82 these complexes are likely to be significantly lower (∼8 to 95 times).82 They are remarkably much lower (∼170 to 200 times) than those of [Cu(cur)(phen)(ClO4)] (IC50, 85.30 μM) and [Cu(acac)(phen)(ClO4)] (IC50, >100 μM).77 Further, the cytotoxicity of 1 and 2 against the HeLa and A549 cancer cell lines was comparable to that of the prevalent benchmark metallodrug, cisplatin, under the same experimental conditions. The IC50 values of 1 and 2 were ∼90 (HeLa) or ∼200 (A549) times lower than that of cisplatin (HeLa: 48 h, IC50 = 16.4 μM; A549: 48 h, IC50 = 40 μM),117,118 which is certainly a beneficial factor for considering these complexes as promising candidates for new antitumor agents. In addition, 1 and 2 do not cause any damage to the normal mouse embryonic fibroblast cells (NIH 3T3: 48 h, IC50 > 500 μM) or normal human lung epithelial cells (L132: 48 h, IC50 > 250 μM), indicating that they are non-toxic to healthy cells, which is expected for a better drug.Reactive oxygen species (ROS) detection
The ROS levels in the A549 cancer cell line are assessed in this study by employing an oxidant-sensitive fluorescent dye. Cellular esterase can alter the non-polar fluorescent dye dichlorodihydrofluorescein diacetate (DCFH-DA) into the polar DCFH. However, this DCFH is not visible under a microscope and requires intracellular ROS to change into a fluorescent DCF. Our findings propose that complexes treated with A549 exhibit fluorescence rather than the absence of fluorescence observed in the control cells (Fig. 9). This suggests that both complexes accelerate the apoptosis caused by ROS in A549 cells.109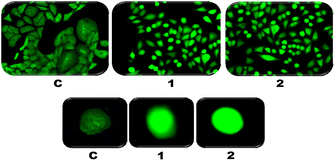 | ||
| Fig. 9 Reactive oxygen species (ROS) generation in the absence (C) and presence of Cu(II) complexes (1 and 2) and visualized using a fluorescence microscope. | ||
Further, we assessed DCF fluorescence intensity using flow cytometry to investigate the influence of 1 or 2 on ROS levels. The DCF fluorescence intensity is 100% in the control. Hydrogen peroxide (H2O2) is next to the superoxide anion and hydroxyl radical, a key member of the class of ROS. It is an important ROS produced by the mitochondrial respiratory chain in many chemically reactive molecules derived from oxygen;110 hence, it is used as a positive control. The A549 cells are treated at 250 μM concentrations of H2O2, indicating a higher DCF fluorescence intensity of 149.12 ± 0.49% at the cellular level of ROS. The DCF fluorescence intensity increases to 140.34 ± 0.93% (1) or 148.42 ± 0.62% (2) when A549 cells are treated with 1 or 2. A greater shift denotes higher fluorescence intensity due to the formation of more DCF, resulting in the production of more ROS for 2 than for 1, which is comparable to the DCF fluorescence intensity of H2O2 ROS (Fig. 10). This demonstrates that 1 and 2 activate the ROS-generating machinery and lead to increased levels of ROS when exposed to cancer cells, causing apoptosis. Therefore, it is clear that oxidative stress contributes to the effective anticancer properties of Cu(II) complexes.
Cell cycle arrest studies
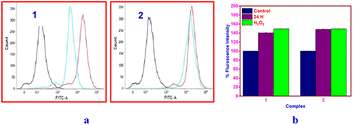
By inducing apoptosis and/or cell cycle arrest, which are important mechanisms of action for anticancer metal complexes, cytotoxic drugs may prevent the growth of cancer cells.111 This was clarified using flow cytometry to examine the profiles of PI-stained A549 cells treated with IC50 doses of 1 or 2 for 24 h.112 It is possible to quantify the populations in the G0/G1, S, and G2/M phases, which can be connected to the quantity of DNA copies in the single-cell suspension. The percentage of control cells at S phase is 18.48 ± 0.34% (Fig. 11). When exposed to synchronized cells in the S phase, 1 (28.76% ± 0.29%) or 2 (30.91% ± 0.40%) started a startling rise of 10.28% (1) or 12.43% (2). Both the G0/G1 and G2/M phases decreased in tandem with the increase in the S phase. The S phase, which occurs before cell division, is when histone and DNA synthesis occur.113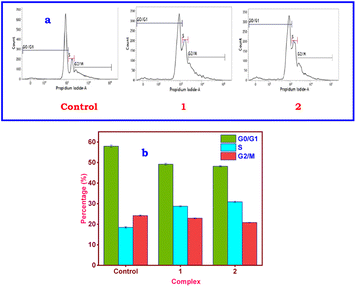 | ||
| Fig. 11 (a) Flow cytometric analysis showing the S phase cell cycle arrest by copper(II) complexes in A549 cells and (b) bar diagram showing % S phase arrest obtained from the FACSverse analysis. | ||
However, DNA replication is halted if DNA breaks occur during the process. According to the flow cytometry data, 1 and 2 cause DNA damage that leads to A549 cells undergoing apoptosis, which arrests the cells in the S-phase. It is clear that, under the same conditions, 2 produces a greater apoptotic effect on cell cycle arrest than 1 (Fig. 11). The impact on cell cycle arrest does not match the IC50 values of the complexes. Given that A549 cancer cells have shorter doubling times than healthy cells, S-phase cell cycle arrest in cancer cells may help deliver differential toxicity. Numerous anticancer drugs, such as cisplatin,114 mitomycin C,115 hydroxyurea,116 and the polyphenolic anticancer agent resveratrol,117 have been shown to cause S-phase arrest in various human cancer cell types. Numerous anticancer drugs are known to suppress the activity of topoisomerase and other essential enzymes involved in DNA duplication.118 Further study into the processes underlying the S-phase arrest brought on by 1 or 2 will be intriguing.
Apoptosis studies
To determine whether the inhibition of cell proliferation and cytotoxicity is caused by apoptosis induction or nonspecific necrosis, the most active cytotoxic complexes 1 and 2 were examined. Apoptosis, also known as programmed cell death, is a type of cell death that is considered when developing anticancer drugs. In order to evaluate the anticancer drug, an apoptosis study that uses acridine orange (AO) and ethidium bromide (EB) staining is introduced. Although EB only stains cells that have lost their membrane integrity, AO is a vital dye that can stain both live and dead cells.119 Viable cells have green fluorescing nuclei with a highly organized structure when viewed under a fluorescence microscope. Early apoptotic cells also have green fluorescing nuclei, but perinuclear chromatin condensation is visible as bright green patches or fragments.Late apoptotic cells have orange to red fluorescing nuclei with condensed or fragmented chromatin. Necrotic cells have consistently orange to red fluorescing nuclei and have expanded to large sizes; there is no sign of chromatin fragmentation. The complexes promoted the apoptosis of A549 cells, as evidenced by the microphotographs of AO- and EB-stained A549 cells pretreated for 24 hours with the copper(II) complexes. Apoptosis, which is characterized by extremely condensed chromatin that marinates into a horseshoe-shaped structure, can be induced by both complexes (Fig. 12). As a result, the observed morphological changes demonstrate that while both complexes cause cell death, they induce only apoptosis and not necrosis.120
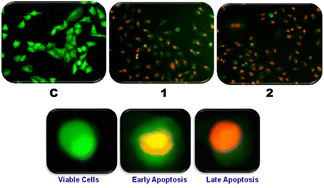 | ||
| Fig. 12 A549 cells stained with acridine orange (AO)/ethidium bromide (EB) in the absence (C) and presence of copper(II) complexes (1 or 2, 0.50 μM) and visualized under a fluorescence microscope. | ||
4′,6′-Diamidino-2-phenylindole (DAPI) staining was also performed to help establish the apoptotic phase. The copper(II) complexes treated with A549 cells and the control were stained with DAPI and observed under a confocal microscope (Fig. 13). In contrast to the treated cells, which exhibit typical indicators of cells undergoing apoptosis, the nuclei of the control cells are permeabilized with detergent (0.1% Triton X-100) and are light and uniformly marked. The treated cells are found to have fragmented or highly condensed nuclei; bright field images indicate cell shrinkage and membrane blebbing, which are characteristics of apoptotic cells. Therefore, the DAPI staining with both complexes121 shows an apoptotic mode of cell death.
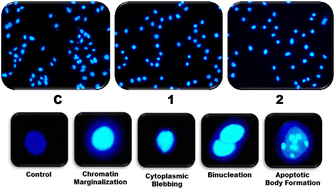 | ||
| Fig. 13 A549 cells stained with DAPI in the absence (C) and presence of copper(II) complexes (1 or 2, 0.50 μM) and visualized under a fluorescence microscope. | ||
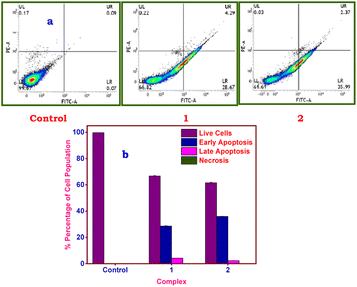
Annexin V and propidium iodide (PI) assays, along with flow cytometry, were used to examine the induction of apoptosis in A549 cells. The IC50 concentrations determined from the cytotoxicity assessment were used to measure the induction of apoptosis after 24 h of treatment with 1 and 2. The assay of annexin V and PI selectively labels cells, allowing for the differentiation of live, apoptotic, and necrotic cells. Annexin V binds to phosphatidylserines externalized to the outer leaflet of the plasma membrane during apoptosis, while PI, a membrane-impermeable dye, binds to the DNA of necrotic cells with damaged plasma membranes.122 The analysis of the flow cytometry data indicates (Fig. 14) that the viable cells (annexin V−/PI− cells) in the lower left quadrant comprise 99.67% of the total; the viable cells in the treated ones decrease to 66.82 ± 0.52% (1) or 61.61 ± 0.48% (2). The early apoptotic cells (annexin V+/PI− cells) in the lower right quadrant increased from 0.07% in the untreated cells to 28.67% ± 0.25% (1) or 35.99% ± 0.21% (2), while the late apoptotic cells (annexin V+/PI+) in the upper right quadrant abruptly increased from 0.09% in the untreated cells to 4.29% ± 0.07% (1) or 2.37% ± 0.05% (2) in the treated cells.
The absence of necrotic cells is shown by the dearth of such a large fluctuation [untreated, 0.17%; treated, 0.22 ± 0.015% (1) or 0.03 ± 0.001% (2)] in the upper left quadrant (annexin V−/PI+ cells). The overall percentages of apoptotic cells in 1 and 2 treated cells at IC50 concentration were 33 and 38%, respectively. This indicates that 2 is more potent than 1 at inducing apoptosis in A549 cells. The fact that the lower right quadrant comprised the highest percentage of cells made it clear that both complexes caused early apoptosis in A549 cells. One of the primary mechanisms of action for copper complexes is generally acknowledged to be their capacity to induce apoptosis, which has been extensively reported in the literature.27,123 These findings demonstrated that 1 or 2 produced apoptosis alone in treated A549 cells in contrast to the control, suggesting that 1 or 2 induced toxicity involving apoptotic cell death.124,125
Conclusions
Novel mixed-ligand copper(II) complexes of the type [Cu(L1)/(L2)(phen)](ClO4)2 are synthesized and characterized. The geometry of [Cu(L2)(phen)](ClO4)2 was analyzed through single-crystal X-ray diffraction and was shown to have a CuN5 chromophore. The coordination environment of the Cu(II) atom can be described as a distorted square pyramidal. A three-dimensional infinite network is formed and dominated by the inter-pair C–H⋯π anti-parallel non-covalent interactions. The chemical reactivity, stability, and selectivity of both complexes in the solution were understood using electronic parameter calculations and structural optimization using DFT. The optimized structures show that the coordination geometry of the copper center is a distorted square pyramidal. They are promising candidates for cancer drugs corresponding to the computed quantum chemical descriptors. Absorption, EPR, and ESI-MS spectral investigations revealed that both complexes preserve their geometry in solution. The binding properties of the complexes with CT DNA and BSA are investigated. The results of electronic absorption and circular dichroic spectroscopy, as well as electrochemistry, indicate that both complexes bind to DNA in a partial intercalative mode with a high affinity for DNA. The binding studies with EthBr have revealed a significant decrease in the fluorescence intensity of the EthBr–DNA system in the presence of both complexes, implying that the complexes can compete for DNA-binding sites with EthBr and displace EthBr from the EthBr–DNA system. Interestingly, both complexes show reversible redox behavior. They cause a considerable decrease in current when they bind to CT DNA, shift to a less negative or less positive Cu(II)/Cu(I) redox potential, and stabilize the Cu(I) oxidation state, resulting in more evident chemical nuclease activity. The mechanism of quenching in BSA is believed to occur through the static mode, implying that complexes bind to BSA via hydrophobic interaction based on protein binding studies. Both complexes show hydrolytic DNA cleavage activity in the absence of an activator (97%, 130–140 μM complex concentration). Their affinity for BSA (Kb, 106 M−1) shows that the complexes are delivered in the blood by serum albumin. The two copper(II) complexes are non-toxic to NIH 3T3 and L132 healthy cells, yet they are more cytotoxic in vitro than cisplatin against HeLa and A549 cancer cells. These results showed that the complexes were particularly detrimental to cancer cells probably due to their higher hydrophobicity. Both complexes were shown to accumulate preferentially in the A549 cells. Live cell confocal microscopy demonstrated ROS generation, crumpling and fragmentation of cell nuclei, elevated ROS levels comparable to H2O2, flow cytometry demonstrating early apoptosis (LR quadrant) in A549 cells, and confirmation of apoptotic cell death by AO/EB and DAPI staining and annexin V-FITC/PI assays. Additionally, complexes in the cell cycle analysis prevent the proliferation of A549 cancer cells by causing cell cycle arrest at the S-phase. As a result, early apoptosis, ROS production, and S-phase arrest show that the complexes prevent cancer cell proliferation and trigger apoptosis. The biological properties of the newly developed mixed-ligand copper(II) complexes outlined here provided a substantial anticancer activity and are suitable for in vivo research and further investigation, which can facilitate the discovery of new anticancer agents with enhanced efficacy.Author contributions
Mariappan Murali: conceptualization, funding acquisition, investigation, methodology, project administration, resources, supervision, writing – original draft, writing – review and editing; Anbarasu Kanchana Mala: formal analysis, investigation, methodology; Somasundaram Sangeetha: data curation, formal analysis, investigation, methodology; Balasubramaniam Selvakumaran: data curation, formal analysis, investigation, methodology; Pitchan Arul Prakash: formal analysis, investigation, methodology; Selvaraj Shanmugavadivel: data curation, investigation; Tamilarasan Ajay Kamal: methodology, software; Mohamed Sultan Mohamed Jaabir: conceptualization, funding acquisition, investigation, project administration, resources, supervision.Conflicts of interest
There are no conflicts to declareData availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5dt02476e.CCDC 2143125 contains the supplementary crystallographic data for this paper.126
Acknowledgements
We thank STIC, Cochin University of Science and Technology, for X-ray crystal structure data. We thank the SAIF, Indian Institute of Technology Madras, for structure solution and refinement and for recording EPR spectra. Author B. S. thanks the Council of Scientific and Industrial Research (CSIR), New Delhi, India, for giving the CSIR SRF (Direct) Fellowship [file no. 08/1365(25505)/2025-EMR-I]. Author TA acknowledges support from SRM TRP Engineering College, India (SRM/TRP/RI/005).References
- S. Adhikari, P. Nath, A. Das, A. Datta, N. Baildya, A. K. Duttaroy and S. Pathak, Biomed. Pharmacother., 2024, 171, 116211 CrossRef CAS PubMed.
- T. C. Johnstone, K. Suntharalingam and S. J. Lippard, Chem. Rev., 2016, 116, 3436–3486 CrossRef CAS PubMed.
- S. Zornić, B. Simović Marković, A. A. Franich, M. D. Živković, B. Luković, N. N. Arsenijević, G. D. Radosavljević, S. Rajković and J. Pantić, J. Biol. Inorg. Chem., 2024, 29, 51–73 CrossRef.
- T. Arai, M. Oshima, M. Uemura, T. Matsunaga, T. Ashizawa, Y. Suhara, M. Morii, H. Yoneyama, Y. Usami, S. Harusawa, S. Komeda and Y. Hirota, Inorg. Chem., 2024, 63, 20951–20963 CrossRef CAS PubMed.
- A. I. Matesanz, C. Hernández, A. Rodríguez and P. Souza, Dalton Trans., 2011, 40, 5738–5745 RSC.
- R. Oun, Y. Moussa and N. Wheate, Dalton Trans., 2018, 47, 6645–6653 RSC.
- S. Dhar, D. Senapati, P. K. Das, P. Chatopadhyay, M. Nethaji and A. R. Chakravarty, J. Am. Chem. Soc., 2003, 125, 12118–12124 CrossRef CAS PubMed.
- B. Rogalewicz and A. Czylkowska, Eur. J. Med. Chem., 2025, 292, 117702 CrossRef CAS PubMed.
- (a) C. S. Novoa-Ramírez, R. I. López-Palma, K. De la Mora-Zarco, M. E. Maya-Yescas, L. Rascón-Valenzuela, A. Espinoza-Guillén, J. S. Delgado-Domínguez, V. Gómez-Vidales, M. Flores-Alamo, R. E. Robles-Zepeda, I. Becker and L. Ruiz-Azuara, J. Inorg. Biochem., 2026, 275, 113113 CrossRef PubMed; (b) A. Hussain, M. F. AlAjmi, M. T. Rehman, S. Amir, F. M. Husain, A. Alsalme, M. A. Siddiqui, A. A. AlKhedhairy and R. A. Khan, Sci. Rep., 2019, 9, 5237 CrossRef PubMed; (c) L. Santa Maria de la Parra, A. I. B. Romo, J. Rodríguez-López, O. R. Nascimento, G. A. Echeverría, O. E. Piro and I. E. León, Inorg. Chem., 2024, 63, 4925–4938 CrossRef CAS PubMed.
- (a) D. V. Szalóki, M. Szabó, D. Bonczidai-Kelemen, A. Udvardy, N. V. May, K. Kovács, L. Virág, I. Fábián and N. Lihi, Inorg. Chem., 2025, 64, 21718–21729 CrossRef PubMed; (b) Y. A. Gur'eva, O. A. Zalevskaya, O. G. Shevchenko, P. A. Slepukhin, V. A. Makarov and A. V. Kuchin, RCS Adv., 2022, 12, 8841–8851 Search PubMed.
- S. Tardito, I. Bassanetti, C. Bignardi, L. Elviri, M. Tegoni, C. Mucchino, O. Bussolati, R. Franchi-Gazzola and L. Marchio, J. Am. Chem. Soc., 2011, 133, 6235–6242 CrossRef CAS PubMed.
- D. Denoyer, S. Masaldan, S. La Fontaine and M. A. Cater, Metallomics, 2015, 7, 1459–1476 CrossRef CAS PubMed.
- H. Arif, A. Sohail, M. Farhan, A. A. Rehman, A. Ahmad and S. M. Hadi, Int. J. Biol. Macromol., 2018, 106, 569–578 CrossRef CAS PubMed.
- U. Jungwirth, C. R. Kowol, B. K. Keppler, C. G. Hartinger, W. Berger and P. Heffeter, Antioxid. Redox Signaling, 2011, 15, 1085–1127 CrossRef CAS PubMed.
- L. R. Azuara and M. E. Bravo-Gómez, Curr. Med. Chem., 2010, 17, 3606–3615 CrossRef PubMed.
- A. R. Barbosa, K. R. Caleffi-Ferracioli, C. Q. Leite, J. C. García-Ramos, Y. Toledano-Magaña, L. Ruiz-Azuara, V. L. D. Siqueira, F. R. Pavan and R. F. Cardoso, Chemotherapy, 2016, 61, 249–255 CrossRef CAS PubMed.
- L. Chavelas-Hernandez, L. G. Hernandez-Vazquez, J. R. Valdez-Camacho, A. Espinoza-Guillen, C. A. Tavira-Montalvan, A. Meneses-Acosta, E. Juaristi, L. Ruiz-Azuara and J. Escalante, Molecules, 2025, 30, 634 CrossRef CAS PubMed.
- E. Finnegan, W. Ding, Z. Ude, S. Terer, T. McGivern, A. M. Blümel, G. Kirwan, X. Shao, F. Genua, X. Yin, A. Kel, S. Fattah, P. A. Myer, S.-A. Cryan, J. H. M. Prehn, D. P. O'Connor, L. Brennan, G. Yochum, C. J. Marmion and S. Das, Cell. Oncol., 2024, 47, 533–553 CrossRef CAS PubMed.
- P. Tsvetkov1, S. Coy, B. Petrova, M. Dreishpoon, A. Verma, M. Abdusamad, J. Rossen, L. Joesch-Cohen, R. Humeidi, R. D. Spangler, J. K. Eaton, E. Frenkel, M. Kocak, S. M. Corsello, S. Lutsenko, N. Kanarek, S. Santagata and T. R. Golub, Science, 2022, 375, 1254–1261 CrossRef PubMed.
- S. Kathiresan, S. Mugesh, M. Murugan, F. Ahamed and J. Annaraj, RSC Adv., 2016, 6, 1810–1825 RSC.
- V. Gandin, M. Porchia, F. Tisato, A. Zanella, E. Severin, A. Dolmella and C. Marzano, J. Med. Chem., 2013, 56, 7416–7430 CrossRef CAS PubMed.
- K. Malakar, B. Selvakumaran, M. Murali, P. Arul Prakash, S. Sangeetha, W. P. Sohtun, M. S. Mohamed Jaabir and M. Velusamy, New J. Chem., 2023, 47, 20070–20087 RSC.
- D.-H. Cai, C.-L. Zhang, Q.-Y. Liu, L. He, Y.-J. Liu, Y.-H. Xiong and X.-Y. Le, Eur. J. Med. Chem., 2021, 213, 113182 CrossRef CAS PubMed.
- A. Mamindla, D. Murugan, M. Varadhan, T. Ajaykamal, L. Rangasamy, M. Palaniandavar and V. Rajendiran, RSC Adv., 2024, 14, 31704–31722 RSC.
- P. Anitha, N. Chitrapriya, Y. J. Jang and P. Viswanathamurthi, J. Photochem. Photobiol., B, 2013, 129, 17–26 CrossRef CAS PubMed.
- W.-J. Lian, X.-T. Wang, C.-Z. Xie, H. Tian, X.-Q. Song, H.-T. Pan, X. Qiao and J.-Y. Xu, Dalton Trans., 2016, 45, 9073–9087 RSC.
- C. Santini, M. Pellei, V. Gandin, M. Porchia, F. Tisato and C. Marzano, Chem. Rev., 2014, 114, 815–862 CrossRef CAS PubMed.
- L. J. H. Borges, E. S. Bull, C. Fernandes, A. Horn Jr., N. F. Azeredo, J. A. L. C. Resende, W. R. Freitas, E. C. Q. Carvalho, L. S. Lemos, H. Jerdy and M. M. Kanashiro, Eur. J. Med. Chem., 2016, 123, 128–140 CrossRef CAS.
- A. Basu, D. Thiyagarajan, C. Kar, A. Ramesh and G. Das, RSC Adv., 2013, 3, 14088–14098 RSC.
- (a) C. Evans, M. Ahmed, D. F. Beirne, M. McCann, K. Kavanagh, M. Devereux, D. Rooney and F. Heaney, BioMetals, 2023, 36, 1241–1256 CrossRef CAS PubMed; (b) C. Lüdtke, S. Sobottka, J. Heinrich, P. Liebing, S. Wedepohl, B. Sarkar and N. Kulak, Chem. Eur. J., 2021, 27, 3273–3277 CrossRef PubMed.
- A. Altomare, G. Cascarano, C. Giacovazzo and A. Guagliardi, J. Appl. Crystallogr., 1993, 26, 343–350 CrossRef.
- G. M. Sheldrick, Acta Crystallogr., Sect. C: Struct. Chem., 2015, 71, 3–8 Search PubMed.
- L. J. Farrugia, J. Appl. Crystallogr., 1999, 32, 837–838 CrossRef CAS.
- L. J. Farrugia, J. Appl. Crystallogr., 1997, 30, 565 CrossRef CAS.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, B. Mennucci, G. A. Petersson, H. Nakatsuji, M. Caricato, X. Li, H. P. Hratchian, A. F. Izmaylov, J. Bloino, G. Zheng, J. L. Sonnenberg, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Staroverov, R. Kobayashi, J. Normand, K. Raghavachari, A. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, N. Rega, J. M. Millam, M. Klene, J. E. Knox, J. B. Cross, V. Bakken, C. Adamo, J. Jaramillo, R. Gomperts, R. E. Stratmann, O. Yazyev, A. J. Austin, R. Cammi, C. Pomelli, J. W. Ochterski, R. L. Martin, K. Morokuma, V. G. Zakrzewski, G. A. Voth, P. Salvador, J. J. Dannenberg, S. Dapprich, A. D. Daniels, O. Farkas, J. B. Foresman, J. V. Ortiz, J. Cioslowski and D. J. Fox, Gaussian 09, Revision D.01, Gaussian Inc., Wallingford CT, 2009, 1–720 Search PubMed.
- A. D. Becke, J. Chem. Phys., 1993, 98, 1372–1377 CrossRef CAS.
- P. J. Hay and W. R. Wadt, J. Chem. Phys., 1985, 82, 270–283 CrossRef CAS.
- P. J. Hay and W. R. Wadt, J. Chem. Phys., 1985, 82, 299–310 CrossRef CAS.
- F. Weinhold and C. R. Landis, Chem. Educ. Res. Pract., 2001, 2, 91–104 RSC.
- J. D. Bradley and G. C. Gerrans, J. Chem. Educ., 1973, 50, 463–467 CrossRef CAS.
- C.-G. Zhan, J. A. Nichols and D. A. Dixon, J. Phys. Chem. A, 2003, 107, 4184–4195 CrossRef CAS.
- W. T. Yang and R. G. Parr, Proc. Natl. Acad. Sci. U. S. A., 1985, 82, 6723–6726 CrossRef CAS PubMed.
- M. Berkowitz, J. Am. Chem. Soc., 1987, 109, 4823–4826 CrossRef CAS.
- S. B. Liu, J. Chem. Sci., 2005, 117, 477–481 CrossRef CAS.
- J. Marmur, J. Mol. Biol., 1961, 3, 208–218 CrossRef CAS.
- N. S. Ng, P. Leverett, D. E. Hibbs, Q. Yang, J. C. Bulanadi, M. J. Wu and J. R. Aldrich-Wright, Dalton Trans., 2013, 42, 3196–3209 RSC.
- P. Patel, M. V. Nadar, D. Umapathy, J. V. Antony, S. Pappu, Y. Rathinam, P. A. Prakash and P. Kumar, Appl. Organomet. Chem., 2020, 34, e5740 CrossRef.
- P. Kumar, A. Nagarajan and P. D. Uchil, Cold Spring Harb. Protoc., 2018, 6, 469–471 Search PubMed.
- D. Figueroa, M. Asaduzzaman and F. Young, J. Pharmacol. Toxicol. Methods, 2018, 94, 26–33 CrossRef CAS PubMed.
- R. M. Roy, P. Pratheeshkumar, Y.-O. Son, L. Wang, J. A. Hitron, S. P. Divya, Z. Zhang and X. Shi, Toxicol. Appl. Pharmacol., 2016, 307, 81–90 CrossRef CAS PubMed.
- R. N. Jadeja, K. M. Vyas, K. K. Upadhyay and R. V. Devkar, RSC Adv., 2017, 7, 17107–17116 RSC.
- P. Jaividhya, M. Ganeshpandian, R. Dhivya, M. A. Akbarsha and M. Palaniandavar, Dalton Trans., 2015, 44, 11997–12010 RSC.
- B. Selvakumaran, M. Murali and V. Sathya, Inorg. Chim. Acta, 2023, 553, 121514 CrossRef CAS.
- N. Z. Fantoni, Z. Molphy, S. O'Carroll, G. Menounou, G. Mitrikas, M. G. Krokidis, C. Chatgilialoglu, J. Colleran, A. Banasiak, M. Clynes, S. Roche, S. Kelly, V. McKee and A. Kellett, Chem. – Eur. J., 2021, 27, 971–983 CrossRef CAS PubMed.
- B. Selvakumaran and M. Murali, Inorg. Chim. Acta, 2022, 534, 120819 CrossRef CAS.
- S. Sangeetha, T. Ajaykamal and M. Murali, New J. Chem., 2021, 74, 7578–7593 RSC.
- R. Kumar, S. Obrai, A. Kaur, G. Hundal, H. Meehnian and A. K. Jana, Polyhedron, 2013, 56, 55–61 CrossRef CAS.
- V. C. Ravisankar, B. Selvakumaran, S. Shanmugavadivel, T. Ajaykamal and M. Murali, Inorg. Chem. Commun., 2026, 185, 116060 CrossRef CAS.
- A. Z. El-Sonbati, I. M. El-Deen and M. A. El-Bindary, J. Mol. Liq., 2016, 215, 612–624 CrossRef CAS.
- L. Dinparast, H. Valizadeh, E. Vessally and M. B. Bahadori, J. Mol. Struct., 2016, 1105, 118–127 CrossRef CAS.
- R. Shah, J. Mol. Liq., 2016, 220, 939–953 CrossRef CAS.
- A. Y. Al-Dawood, N. M. El-Metwaly and H. A. El-Ghamry, J. Mol. Liq., 2016, 220, 311–323 CrossRef CAS.
- K. Sayin, N. Kurtoglu, M. Kose, D. Karakas and M. Kurtoglu, J. Mol. Struct., 2016, 1119, 413–422 CrossRef CAS.
- S. Koray and Ü. Ayhan, Spectrochim. Acta, Part A, 2018, 193, 147–155 CrossRef PubMed.
- M. Murali, V. Sathya and B. Selvakumaran, J. Biol. Inorg. Chem., 2021, 26, 67–79 CrossRef CAS PubMed.
- V. Sathya and M. Murali, Inorg. Chim. Acta, 2019, 496, 119016 CrossRef CAS.
- R. Loganathan, S. Ramakrishnan, E. Suresh, A. Riyasdeen, M. A. Akbarsha and M. Palaniandavar, Inorg. Chem., 2012, 51, 5512–5523 CrossRef CAS PubMed.
- V. Sathya and M. Murali, Inorg. Chem. Commun., 2018, 92, 55–59 CrossRef CAS.
- C. Rajarajeswari, R. Loganathan, M. Palaniandavar, E. Suresh, A. Riyasdeen and M. A. Akbarsha, Dalton Trans., 2013, 42, 8347–8355 RSC.
- S. Usha and M. Palaniandavar, J. Chem. Soc., Dalton Trans., 1994, 15, 2277–2283 Search PubMed.
- A. Erxleben, Coord. Chem. Rev., 2018, 360, 92–111 CrossRef CAS.
- M. Ganeshpandian, R. Loganathan, S. Ramakrishnan, A. Riyasdeen, M. A. Akbarsha and M. Palaniandavar, Polyhedron, 2013, 52, 924–938 CrossRef CAS.
- S. Ramakrishnan and M. Palaniandavar, J. Chem. Sci., 2005, 117, 179–186 CrossRef CAS.
- A. Banaspati, V. Ramu, M. K. Raza and T. K. Goswami, RSC Adv., 2022, 12, 30722–30733 RSC.
- S. Ramakrishnan, V. Rajendiran, M. Palaniandavar, V. S. Periasamy, B. S. Srinag, H. Krishnamurthy and M. A. Akbarsha, Inorg. Chem., 2009, 48, 1309–1322 CrossRef CAS PubMed.
- J. Manivel, S. Sangeetha and M. Murali, J. Sci. Res., 2020, 12, 111–133 CrossRef CAS.
- M. Ganeshpandian, R. Loganathan, E. Suresh, A. Riyasdeen, M. A. Akbarsha and M. Palaniandavar, Dalton Trans., 2014, 43, 1203–1211 RSC.
- S. Jain, K. Bhar, S. Kumar, S. Bandyopadhyaya, S. Tapryal, C. C. Mandal and A. K. Sharma, Dalton Trans., 2020, 49, 4100–4113 RSC.
- K. Radhakrishnan, T. Khamrang, K. Sambantham, V. K. Sali, U. Chitgupi, J. F. Lovell, A. A. Mohammad and R. Venugopal, Polyhedron, 2021, 194, 114886 CrossRef CAS.
- Q.-Y. Chen, H.-J. Fu, W.-H. Zhu, Y. Qi, Z.-P. Ma, K.-D. Zhao and J. Gao, Dalton Trans., 2011, 40, 4414–4420 RSC.
- M. Devereux, D. O'Shea, M. O'Connor, H. Grehan, G. Connor, M. McCann, G. Rosair, F. Lyng, A. Kellett, M. Walsh, D. Egan and B. Thati, Polyhedron, 2007, 26, 4073–4084 CrossRef CAS.
- N. Shahabadi, M. Falsafi and N. H. Moghadam, J. Photochem. Photobiol., B, 2013, 122, 45–51 CrossRef CAS PubMed.
- E. Grueso, G. López-Pérez, M. Castellano and R. Prado-Gotor, J. Inorg. Biochem., 2012, 106, 1–7 CrossRef CAS PubMed.
- (a) F. Jozefíková, Ľ. Šab, S. Perontsis, M. Šimunková, Z. Barbieriková, D. Švorc, M. Valko, G. Psomas and J. Moncol’, New J. Chem., 2020, 44, 12827–12842 RSC; (b) V. Vinayagam, S. Prabu, S. Rajalakshmi and M. N. Arumugam, J. Mol. Struct., 2023, 1287, 135649 CrossRef CAS.
- M. Hazra, T. Dolai, A. Pandey, S. K. Dey and A. Patra, Bioinorg. Chem. Appl., 2014, 1, 104046 Search PubMed.
- S. Parveen, F. Arjmand and I. Ahmad, J. Photochem. Photobiol., B, 2014, 130, 170–178 CrossRef CAS PubMed.
- M. S. Deshpande, S. Junedi, H. Prakash, S. Nagao, M. Yamanaka and S. Hirota, Chem. Commun., 2014, 50, 15034–15037 RSC.
- (a) S. U. Parsekar, M. Singh, D. P. Mishra, P. K. S. Antharjanam, A. P. Koley and M. Kumar, J. Biol. Inorg. Chem., 2019, 24, 343–363 CrossRef CAS PubMed.
- S. Sangeetha and M. Murali, Inorg. Chem., 2022, 61, 2864–2882 CrossRef CAS PubMed.
- K. Paliwal, P. Haldar, P. K. S. Antharjanam and M. Kumar, ACS Omega, 2022, 7, 21961–21977 CrossRef CAS PubMed.
- S. Parveen, M. Usman, S. Tabassum and F. Arjmand, RSC Adv., 2015, 5, 72121–72131 RSC.
- R. Starosta, F. C. Santos and R. F. M. de Almeida, J. Mol. Struct., 2020, 1221, 128805 CrossRef CAS.
- J. R. Lakowicz, Principles of Fluorescence Spectroscopy, Springer Science + Business Media, New York, 3rd edn, 2006 Search PubMed.
- X. Zhao, R. Liu, Z. Chi, Y. Teng and P. Qin, J. Phys. Chem. B, 2010, 114, 5625–5631 CrossRef CAS PubMed.
- M. H. Gehlen, J. Photochem. Photobiol., C, 2020, 42, 100338 CrossRef CAS.
- R. Patel, N. Maurya, M. U. D. Parray, N. Farooq, A. Siddique, K. L. Verma and N. Dohare, J. Mol. Recognit., 2018, 31, e2734 CrossRef PubMed.
- M. A. Qureshi and S. Javed, ACS Omega, 2021, 6, 18054–18064 Search PubMed.
- M. Aharma, M. Ganeshpandian, M. Majumder, A. Tamilarasan, M. Sharma, R. Mukhopadhyay, N. S. Islam and M. Palaniandavar, Dalton Trans., 2020, 49, 8282–8297 RSC.
- L. Abdel-Aziz, L. Abdel-Fattah, A. El-Kosasy and M. Gaied, J. Appl. Spectrosc., 2015, 82, 620–627 Search PubMed.
- O. A. Chaves, V. A. da Silva, C. M. R. Sant'Anna, A. B. B. Ferreira, T. A. N. Ribeiro, M. G. de Carvalho, D. Cesarin-Sobrinho and J. C. Netto-Ferreira, J. Mol. Struct., 2017, 1128, 606–611 Search PubMed.
- C. N. Lunardi, A. C. Tedesco, T. L. Kurth and I. M. Brinn, J. Photochem. Photobiol. Sci., 2003, 2, 954–959 Search PubMed.
- (a) A. Gandioso, A. Vidal, P. Burckel, G. Gasser and E. Alessio, ChemBioChem, 2022, 23, e202200398 Search PubMed; (b) C. M. J. Pisani, P. D. Fromm, Y. Mulyana, R. J. Clarke, H. Körner, K. Heimann, J. G. Collins and F. R. Keene, ChemMedChem, 2011, 6, 84–858 Search PubMed.
- P. Levín, M. C. Ruiz, A. I. B. Romo, O. R. Nascimento, A. L. Di Virgilio, A. G. Oliver, A. P. Ayala, I. C. N. Diógenes, I. E. León and L. Lemus, Inorg. Chem. Front., 2021, 8, 3238–3252 RSC.
- Q.-Y. Chen, H.-J. Fu, J. Huang and R.-X. Zhang, Spectrochim. Acta Part A, 2010, 75, 355–360 CrossRef PubMed.
- M. Kumar, S. U. Parsekar, N. Duraipandy, M. S. Kiran and A. P. Koley, Inorg. Chim. Acta, 2019, 484, 219–226 CrossRef CAS.
- Q.-Y. Chen, J. Huang, W.-J. Guo and J. Gao, Spectrochim. Acta, Part A, 2009, 72, 648–653 CrossRef PubMed.
- S. Sangeetha and M. Murali, Inorg. Chem. Commun., 2015, 59, 46–49 CrossRef CAS.
- L. Pathaw, T. Khamrang, B. Selvakumaran, M. Murali, P. Arul Prakash, M. S. Mohamed Jaabir and M. Velusamy, Appl. Organomet. Chem., 2020, 43, e6025 Search PubMed.
- C. Qian, J.-Q. Wang, C.-L. Song, L.-L. Wang, L.-N. Ji and H. Chao, Metallomics, 2013, 5, 844–854 CrossRef CAS PubMed.
- W. Li, L. Cao, L. Han, Q. Xu and Q. Ma, Int. J. Oncol., 2015, 46, 2613–2620 CrossRef CAS PubMed.
- R. Cao, J. Jia, X. Ma, M. Zhou and H. Fei, J. Med. Chem., 2013, 56, 3636–3644 CrossRef CAS PubMed.
- L. C. Crowley, B. J. Marfell, A. P. Scott and N. Waterhouse, J. Cold Spring Harb. Protoc., 2016, 11, 953–957 Search PubMed.
- (a) D. M. Nelson, X. Ye, C. Hall, H. Santos, T. Ma, G. D. Kao, T. J. Yen, J. W. Harper and P. D. Adams, Mol. Cell. Biol., 2002, 22, 7459–7472 CrossRef CAS PubMed; (b) Y. Anno, T. Kubo, R. Ueki, M. Yano, K. Sasaki, H. Ohba and M. Fujii, Nucleosides, Nucleotides Nucleic Acids, 2003, 22, 1451–1453 CrossRef CAS PubMed.
- G. He, J. Kuang, A. R. Khokhar and Z. H. Siddik, Gynecol. Oncol., 2011, 122, 402–409 CrossRef CAS PubMed.
- T. Nakayama, N. Nozawa, C. Kawada, S. Yamamoto, T. Ishii, M. Ishizuka, T. Namikawa, S.-I. Ogura, K. Hanazaki, K. Inoue and T. Karashima, Photodiagn. Photodyn. Ther., 2020, 31, 101893 CrossRef CAS PubMed.
- A. E. Shaw, M. N. Mihelich, J. E. Whitted, H. J. Reitman, A. J. Timmerman, M. Tehseen, S. M. Hamdan and G. D. Schauer, Proc. Natl. Acad. Sci. U. S. A., 2024, 121, e2404470121 CrossRef CAS PubMed.
- M. Zielińska-Przyjemska, M. Kaczmarek, V. Krajka-Kuźniak, M. Łuczak and W. Baer-Dubowska, Toxicol. In Vitro, 2017, 43, 69–75 CrossRef PubMed.
- M. Gaikwad, V. B. Konkimalla and S. Salunke-Gawali, Inorg. Chim. Acta, 2022, 542, 121089 CrossRef CAS.
- S. M. Smith, D. Ribble, N. B. Goldstein, D. A. Norris and Y. G. Shellman, Methods Cell Biol., 2012, 112, 361–368 CAS.
- U. Basu, I. Khan, A. Hussain, B. Gole, P. Kondaiah and A. R. Chakravarty, Inorg. Chem., 2014, 53, 2152–2162 CrossRef CAS PubMed.
- J. P. Johnpeter, G. Gupta, J. M. Kumar, G. Srinivas, N. Nagesh and B. Therrien, Inorg. Chem., 2013, 52, 13663–13673 CrossRef CAS PubMed.
- A. Costigan, E. Hollville and S. J. Martin, Curr. Protoc., 2023, 3, e951 CrossRef CAS PubMed.
- S. Abdolmaleki, A. Aliabadi and S. Khaksar, J. Cancer Res. Clin. Oncol., 2024, 150, 213 CrossRef CAS PubMed.
- A. Lekshmi, S. N. Varadarajan, S. S. Lupitha, D. Indira, K. A. Mathew, A. C. Nair, M. Nair, T. Prasad, H. Sekar, A. K. Gopalakrishnan, A. Murali and T. R. Santhoshkumar, Cell Death Discovery, 2017, 3, 16101 CrossRef CAS PubMed.
- P. D. Pattanayak, A. Banerjee, G. Sahu, S. Das, S. Lima, O. Akintola, A. Buchholz, H. Görls, W. Plass, H. Reuter and R. Dinda, Inorg. Chem., 2024, 63, 19418–19438 CrossRef CAS PubMed.
- CCDC 2143125: Experimental Crystal Structure Determination, 2026, DOI:10.5517/ccdc.csd.cc29y32g.
Footnote |
| † This paper is in honor of Professor S. Natarajan, Solid State and Structural Chemistry Unit, Indian Institute of Science, Bangalore, India, for his lifetime of dedicated service and profound contributions to the field of Inorganic Solid State Chemistry. |
| This journal is © The Royal Society of Chemistry 2026 |