Open Access Article
Open Access ArticleDiscovery of hydrogen storage molecules using large language models and machine learning
Hassan
Harb
 *a,
Magali S.
Ferrandon
*a,
Magali S.
Ferrandon
 b,
Timothy A.
Goetjen
b,
Timothy A.
Goetjen
 c,
Seryeong
Lee
c,
Seryeong
Lee
 bc,
Omar K.
Farha
bc,
Omar K.
Farha
 c,
Massimiliano
Delferro
c,
Massimiliano
Delferro
 b and
Rajeev Surendran
Assary
b and
Rajeev Surendran
Assary
 *a
*a
aMaterials Science Division, Argonne National Laboratory, Lemont, IL 60439, USA. E-mail: hharb@anl.gov; assary@anl.gov
bChemical Sciences and Engineering Division, Argonne National Laboratory, Lemont, IL 60439, USA. E-mail: ferrandon@anl.gov; delferro@anl.gov
cDepartment of Chemistry, Northwestern University, Evanston, Illinois 60208, USA. E-mail: tim.goetjen@u.northwestern.edu; SeryeongLee2026@u.northwestern.edu; o-farha@northwestern.edu
First published on 28th April 2026
Abstract
Accelerating the discovery of new molecules with targeted properties is a central challenge in molecular design. In this contribution, we present an AI-driven molecular discovery framework that integrates Large Language Models (LLMs) for generative molecular design with Machine Learning (ML)-based screening to identify novel Liquid Organic Hydrogen Carrier (LOHC) candidates. Using the developed framework, LOHC molecules were systematically generated, evaluated, and refined iteratively, combining LLM-guided molecular generation and ML-predicted hydrogenation enthalpies (ΔH), under physicochemical property constraints such as optimal melting points (MP), desired hydrogen storage capacity (wt% H2), and synthetic accessibility (SA) scores. This approach enabled the discovery of 42 new LOHC candidates in two distinct campaigns, one seeded with experimentally known and another with previously computationally identified LOHCs, respectively. Although we began with different numbers of starting molecules (31 vs. 7 seed molecules), both runs yielded a comparable number of viable candidates, suggesting an influence of chemically intuitive seed molecule selection for success. Selected LOHC molecules, such as 3-methyl pyridine, 1-ethylnapthalene, 1,1-diphenylethane, and benzofuran, were experimentally tested and compared with benchmark LOHCs (toluene and 9-ethylcarbazole) for hydrogenation using a series of commercial supported metal catalysts. The order of conversion into fully hydrogenated products at 200 °C was 3-methyl pyridine (100%) > 9-ethyl carbazole (86.4%) > 2,3-benzofuran (74%) > 1,1-diphenylethane (66.9%) > 1-ethylnapthalene (66.7%) > toluene (57%), further validating the AI-guided molecular design. This study demonstrates promise of LLM-driven molecular design in conjunction with ML-based screening for accelerated discovery and design of molecules.
1 Introduction
Generative molecular discovery is rapidly emerging as a transformative approach to navigating the immense molecular space (>1060).1–6 Traditional brute-force molecular screening trial-and-error synthesis is infeasible due to the vast chemical space.6–14 To overcome these limitations, generative artificial intelligence (AI), particularly Large Language Models (LLMs), has gained prominence.15–20 Originally developed for natural language tasks, pre-trained LLMs can be adapted to generate and refine molecular structures from simple text prompts, enabling efficient exploration of chemical space beyond the constraints of conventional methods.1,21,22 In molecular discovery, predictive and generative AI play complementary roles.23–25 Predictive AI, employing techniques such as random forest regression or graph neural networks (GNNs), forecasts molecular properties and interactions based on historical data, enabling rapid screening without resource-intensive experimental validation.7,9,26–30 In contrast, generative AI, including LLMs and diffusion models, creates novel molecular structures by learning patterns from training data and generating candidates that may not exist in known databases.2,3,31–35 These generative models explore uncharted regions of chemical space, guided by learned design principles or optimization rules.2,3 While predictive models excel at accurately estimating properties within the boundaries of known structure–property relationships, generative approaches transcend these boundaries by proposing entirely new molecules. This synergy between generative and predictive AI holds the potential to revolutionize materials discovery, dramatically accelerating the design process by expanding the search space and efficiently filtering candidates, far surpassing the capabilities of traditional screening methods (Fig. 1 and 2). | ||
| Fig. 1 Overview of the molecular discovery workflow depicting the synergy between generative AI, predictive AI, and validation. | ||
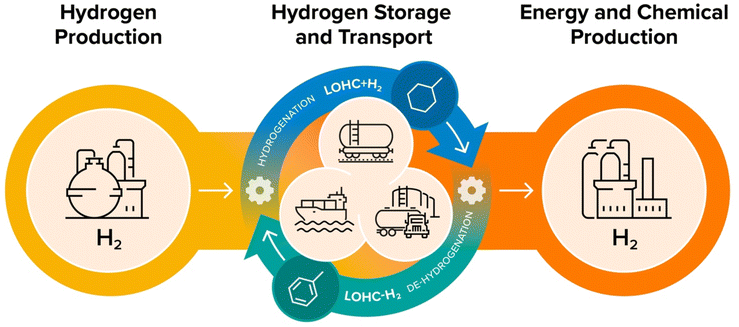 | ||
| Fig. 2 Schematic representation of hydrogen production, transportation, and use in chemical manufacturing using toluene/methylcyclohexane cycle as an LOHC system. | ||
As a representative case, AI-driven discovery of Liquid Organic Hydrogen Carrier (LOHC)36–39 molecules was chosen, which provides a tractable and well-defined system for demonstrating LLM-driven molecular design and discovery.36,38,40–45 LOHCs are unsaturated organic molecules that chemically bind and release hydrogen via reversible hydrogenation/dehydrogenation cycles, facilitated by catalysts.38,39,46 Their liquid-phase stability, safety, and compatibility with existing fuel infrastructure make them a practical and scalable energy storage solution.37,39,47,48 Unlike compressed hydrogen, LOHCs eliminate boil-off losses and enable long-term storage and transport without degradation.49,50 In terms of research and development, comprehensive overviews of potential LOHC systems have been widely reported in the literature and exemplar systems include benzene, toluene, N-ethyl carbazole, and dibenzyl toluene, among others.38,47,48,51,52 At present, the LOHCs face significant challenges and limitations that hinder their widespread application.48,53,54 An optimal LOHC molecule should possess a combination of properties that ensure efficiency, stability, and practicality for hydrogen storage and transport.36,37,40 LOHCs must exhibit a gravimetric hydrogen capacity (wt% H2) above 5.5% to ensure sufficient energy density for transportation applications36,48 and an optimal enthalpy range (40–70 kJ mol−1 per H2) for efficient low-temperature cycling between the hydrogenated and dehydrogenated forms.36,39 In addition, both the hydrogen-lean and hydrogen-rich states should remain liquid at room temperature to facilitate handling and storage.36,39,48 LOHCs must undergo hydrogenation and dehydrogenation without molecular degradation36,37,48,55 and have low toxicity to ensure safe and practical implementation.56,57 These challenges highlight the continuous need for novel LOHC molecules with improved stability, thermodynamics, and catalytic performance, motivating the search for an accelerated and data-driven molecular design approaches to LOHC discovery.36
Recently, we have established a computational screening approach to accelerate LOHC discovery.40 In this massive in silico molecular screening, we systematically processed 160 billion molecules from the ZINC15 (ref. 58) and GDB-17 (ref. 59) molecular databases, leveraging cheminformatics-based selection criteria and accurate quantum chemical calculations.40 This approach led to the identification of 41 novel LOHC candidates with enhanced hydrogen storage capacity and favorable thermodynamic properties.40 To address the accurate data deficiency of LOHCs, we have developed the QM9-LOHC dataset that includes 10 k dehydrogenation reactions calculated using the high-accuracy G4MP2 quantum chemical method.60 This dataset supports ML-based prediction of hydrogenation enthalpies and facilitates data-driven LOHC molecular discovery. Paragian and colleagues42 screened over one million PubChem61 molecules as potential LOHC candidates using RING62 approach for structure generation, OPERA63 for phase property predictions, and ML models for dehydrogenation enthalpies. They identified 14![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 feasible LOHC pairs and selected 37 promising candidates based on hydrogen capacity, synthetic accessibility, and key molecular features analyzed via sparse linear discriminant analysis.42 Despite these advances, many viable LOHC candidates may still be overlooked due to the scale of chemical space and constraints of the applied screening methods.40,42,64
000 feasible LOHC pairs and selected 37 promising candidates based on hydrogen capacity, synthetic accessibility, and key molecular features analyzed via sparse linear discriminant analysis.42 Despite these advances, many viable LOHC candidates may still be overlooked due to the scale of chemical space and constraints of the applied screening methods.40,42,64
Building upon this foundation, this present study integrated generative AI (LLMs) and predictive ML using Random Forest (RF) regression into an iterative molecular discovery framework, where LLMs generated new molecular candidates from seed structures, and an ML model trained on the QM9-LOHC dataset60 predicted the enthalpy of hydrogenation (ΔH) for rapid screening. This AI-driven design process enables efficient generation, evaluation, and refinement of candidate molecules without the need for the costly step of finetuning the LLM. Despite differences in the seed sets used, both approaches converged on a similar number of viable molecules, leading to the discovery of 42 distinct new LOHC structures that satisfy key thermodynamic and MP criteria. From these, 4 LOHC molecules (3-methyl pyridine, 1-ethylnapthalene, 1,1-diphenylethane, and 2,3-benzofuran) were selected and compared with benchmark LOHCs (9-ethylcarbazole and toluene) for the hydrogenation reaction at 150 °C and 200 °C using commercial catalysts. Based on the experimental studies, the order of conversion into fully hydrogenated products is consistent with computational predictions. These results demonstrate that a well-curated data set of molecules (seed set) is more critical than its size, permitting computationally efficient molecular generation. By combining LLM-driven molecular ideation with ML-based screening, this framework accelerates in silico discovery and materials selection processes for targeted experimentation to complete the discovery loop.
2 Results and discussion
2.1 Generation of LOHC molecules
In Fig. 3, the computational molecular discovery workflow that combines generative and predictive AI tools in an iterative loop to develop new molecules from system prompt is shown. We begin with carefully selected seed molecules, either experimentally known or computationally discovered LOHCs, that guide the LLM to generate chemically valid and structurally diverse Simplified Molecular Input Line Entry System (SMILES) strings. The seeds define the initial chemical space and help the model focus on structures that are realistic for hydrogen storage. The LLM samples multiple variations for each seed to explore substitutions, ring patterns, and functional group changes.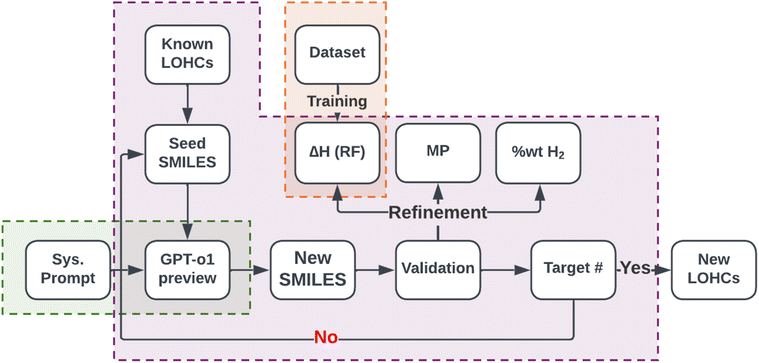 | ||
| Fig. 3 Workflow implemented in this study. The workflow has three main components, shown with the green, purple, and orange shaded regions. The green region is the LLM agent which contains the GPT-o1preview model (accessed via Argo API) and the custom system prompt. The orange region shows the ML part, which contains the dataset and the ML predictive model trained on that dataset and used to evaluate the generated Simplified Molecular Input Line Entry System (SMILES) strings. The purple box shows a generative loop that starts the process with the seed SMILES strings, prompted into LLM agent, and then generates, validates, and refines new SMILES strings. ‘Sys. Prompt’ refers system prompt, ΔH (RF) denotes reaction enthalpies predicted by Random Forest model, MP denotes melting point (predicted using the OPERA63 model), ‘wt% H2’ represents hydrogen storage capacity, and Target # represents the user-defined target number of SMILES (set at a maximum of 200). | ||
Each generated molecule is screened using an ML model, trained on QM9-LOHC dataset to predict hydrogenation enthalpies (ΔH). Details on the ML model are provided in the methods section. The screening step also filters invalid structures. It assigns priority to molecules that stay within known LOHC chemical classes while expanding the space with new motifs. Only molecules that fall within the desired thermodynamic window (40–70 kJ mol−1 per H2) and meet physicochemical criteria (melting point ≤40 °C, wt H2 ≥ 5.5%, synthetic accessibility (SA) score ≤3.3) are retained. The workflow then feeds these retained structures back into the generator. This feedback step helps steer the next round of sampling toward chemistries with improved ΔH values and higher hydrogen capacity. Each cycle expands the pool of candidates while keeping the search focused on feasible LOHC designs. This refinement process repeats for up to ten iterations or 200 unique molecules. These unique molecules and their enthalpies (ΔH) are validated using first principles calculations, ensuring efficiency and accuracy in identifying LOHCs.
2.2 Selection of LOHC molecules
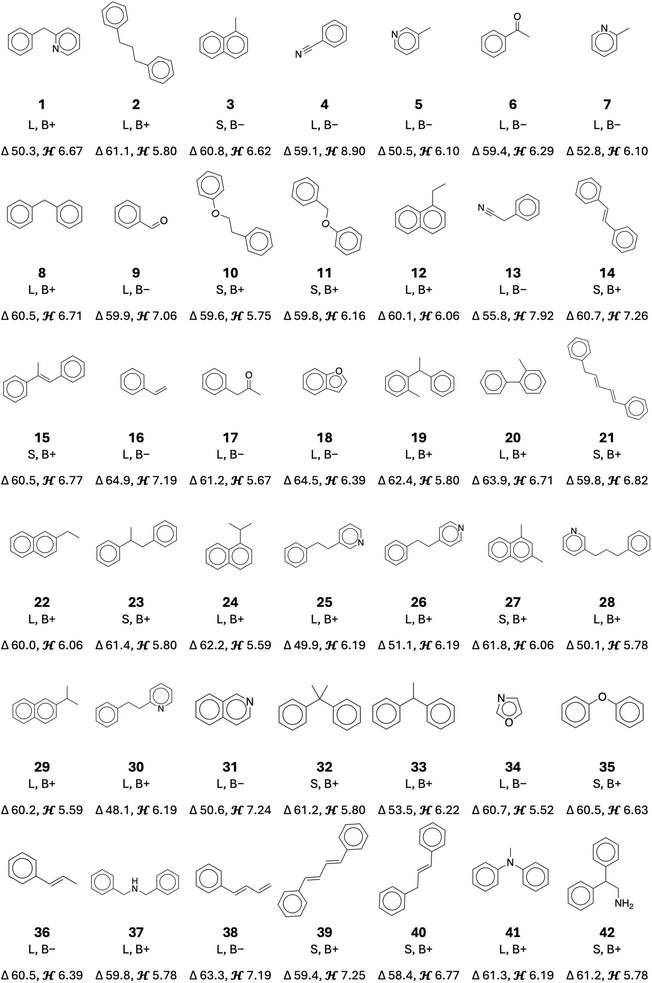
To down select the promising LOHC molecules, we imposed additional criteria for filtering, including practicality and synthesizability. First, both the hydrogen-lean (HL) and hydrogen-rich (HR) forms were required to have MPs below 40 °C. Second, we imposed a synthesizability filter to gauge the likelihood that these molecules can be feasibly prepared in a laboratory setting. We retained only those molecules that either (a) demonstrated an SA65 score below 3.3 based on our previously established cutoff for practical synthesis65 or (b) were already in the PubChem database.61 The SA score factor reflects complexity and likely number of synthetic steps required, while a PubChem listing indicates prior knowledge or availability of the compound. Additionally, all halogen-bearing structures were removed due to their susceptibility to undesired reactions (e.g., elimination) under LOHC operating conditions.40 By combining these two criteria, we prioritized LOHC structures that were both thermally suitable and synthetically feasible.Using the molecular selection workflow (Fig. 3), 42 new LOHC molecules were identified, schematically shown in Fig. 4. Note that additional details of these molecules, including their melting and boiling points and SA scores for the hydrogen-rich and -lean forms, are presented in the SI (Table S4). These molecules exhibit a broad spectrum of physical and thermodynamic properties, emphasizing how structural variations can profoundly impact key performance indicators. In most cases, the MPs of both hydrogen-lean (HL) and hydrogen-rich (HR) forms are below 40 °C, indicating that they remain liquid under ambient conditions. Among these, 11 molecules have MPs below 0 °C. Notably, 7 (2-methylpyridine), 16 (styrene), and 36 (propenyl benzene) melt at −54.9 °C, at −30.1 °C, at −28.3 °C, respectively, thereby minimizing solidification risks in colder environments. Although their boiling points (BP) span a wide range (∼130–150 °C to >300 °C), each of these candidates remains liquid at room temperature, further underscoring their use as an LOHC.
Beyond phase behavior, all 42 compounds shown in Fig. 4 feature ΔH in the desired 40–70 kJ mol−1 H2 range. The entries in Fig. 4, 16 (styrene) and 18 (benzofuran), sit near the upper end (∼64–65 kJ mol−1 H2). Despite this higher ΔH, both remain attractive: benzofuran has previously been shown to undergo efficient hydrogenation, even before its relevance to LOHC applications was established.66 Benzofuran is also a key structural motif in bio-derived compounds.67 Styrene's very low MP potentially operates at sub ambient temperatures, compensating for its higher enthalpy requirement by. By contrast, entries 30, 25, and 28 (all belonging to the phenylethyl- or phenylpropyl-pyridine family) exhibit lower ΔH values (∼48–50 kJ mol−1 H2), potentially enabling easier dehydrogenation at moderate temperatures. Meanwhile, gravimetric hydrogen capacities (wt% H2) run from about 5.5% to nearly 9%, with 4 (benzonitrile) standing out at 8.9%. H2, the highest among this set.
Finally, synthesizability and availability were assessed using PubChem listings and SA scores, indicating that all 42 molecules are likely available in known databases or accessible via standard synthetic routes. Notably, each hydrogen-lean (HL) and hydrogen-rich (HR) form meets our practical synthesis cutoff (SA < 3.3, as discussed in our prior work40,65) or appears in PubChem, minimizing potential barriers to laboratory or pilot-scale validation. This combination of low MPs, balanced ΔH values, and synthetic accessibility underscores their commercial potential as next-generation LOHCs.
2.3 Experimental validation
Five LOHC molecules, 4 (benzonitrile), 5 (3-methyl pyridine), 12 (1-ethylnapthalene), 18 (2,3-benzofuran), and 33 (1,1-diphenylethane) were selected for experimental validation and compared to toluene and 9-ethylcarbazole as benchmark LOHCs. These molecules were selected based on their high wt% H2 (above 6%), low MPs (all below 0 °C), and their availability in PubChem. These molecules were chosen as representative candidates spanning different chemical classes to demonstrate the viability of the AI-driven framework; other promising candidates, (e.g. benzonitrile; 8.9 wt% H2), remain targets for future experimental validation. One of the most studied LOHCs is 9-ethylcarbazole.68 This heterocyclic compound can theoretically take up 6 moles of equivalent hydrogen (5.7%). However, it converts into 4 partially hydrogenated products.69 Schemes of the hydrogenation reactions for all the LOHC molecules are shown in Fig. S5. Hydrogenation catalysts were chosen according to literature.70 10 wt% Pd on carbon and alumina, 5 wt% Pt on NU-1000, as an example of metal–organic framework, and 5 wt% Rh on alumina were employed as hydrogenation catalysts at 150 °C and 200 °C for 12 h at 300 psi of H2. Table 1 shows the conversion of 3-methyl pyridine into 3-methyl piperidine and toluene into methyl cyclohexane at 150 °C and 200 °C for 12 h, respectively. All 4 catalysts exhibited greater conversion of 3-methyl pyridine compared to toluene to fully hydrogenated product, with Rh/Al2O3 being the most active catalyst at 150 °C. At 200 °C, full conversion is achieved for 3-methyl pyridine using Pd/C and Rh/Al2O3 and almost full conversion for Pd/Al2O3. Comparison with other substrates (1-ethylnapthalene, 2,3-benzofuran, and 1,1-diphenylethane) using the most performant catalyst, Rh/Al2O3, at 200 °C is listed in Table 2. All three substrates are fully converted to either fully or partially hydrogenated products, with 74% selectivity for 8H-benzofuran. 9-Ethylcarbazole converts into fully hydrogenated product (86.4%). However, the catalyst also converts into the de-ethylated product (9.2%). The hydrogenation of benzonitrile leads to the formation of dibenzylamine with high conversion using the Rh/Al2O3 catalyst in comparison with the Pt/NU-1000 (Table 3). The order of conversion into fully hydrogenated products at 200 °C is 3-methyl pyridine > 9-ethyl carbazole > 2,3-benzofuran > 1,1-diphenylethane > 1-ethylnapthalene > toluene. Recycling experiments show that Rh/Al2O3 maintain a high activity (Table 3).| Catalyst | 3-Methyl pyridine conversion (%) | Toluene conversion (%) | ||
|---|---|---|---|---|
| 150 °C | 200 °C | 150 °C | 200 °C | |
| 10 wt% Pd/C | 35.8 | 100.0 | 16.2 | 45.3 |
| 5 wt% Rh/Al2O3 | 58.9 | 100.0 | 24.9 | 57.0 |
| 10 wt% Pd/Al2O3 | 38.5 | 96.8 | 7.0 | 45.6 |
| 5 wt% Pt/NU-1000 | 29.1 | 75.4 | 5.8 | 39.7 |
| None | 0 | 0 | 0.5 | 9.0 |
| Substrate | Fully hydrogenated (%) | Partiallyhydrogenated (%) | Other (%) |
|---|---|---|---|
| 1-Ethylnapthalene | 66.7 | 31.3 | — |
| 2,3-Benzofuran | 74.0 | 26.0 | — |
| 1,1-Diphenylethane | 66.9 | 33.1 | — |
| 9-Ethyl carbazole | 86.4 | 4.4 | 9.2 |
3 Conclusions
This study demonstrates that combining LLM-driven molecular generation with ML-based screening provides an efficient path for discovering new LOHC molecules. The workflow identified chemically meaningful structures, evaluated them with rapid predictive models, and refined the candidates through an iterative loop that focuses on thermodynamic and physicochemical targets. Using this approach, 42 new LOHC candidates were discovered and 39 of them were validated using high-accuracy G4MP2 calculations. Four of these molecules were tested experimentally and showed hydrogenation performance that aligns with the AI predictions. These results show that a small but well-chosen seed set can guide LLMs toward novel and viable LOHC designs. By combining pretrained generative models with modular screening, this framework enables efficient exploration of chemical space and can be readily extended to diverse molecular discovery tasks.In the present work, the LLM was guided solely through a system prompt and seed molecules represented as SMILES strings, without access to an external knowledge base. An alternative and potentially complementary strategy is retrieval-augmented generation (RAG), in which the LLM is provided with relevant chemical context retrieved from a structured database at generation time.71,72 For example, Zhang et al. recently demonstrated a RAG-based framework for solid-state hydrogen storage materials, where a knowledge base of over 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 literature-extracted entries was used to inform LLM-driven candidate generation and iterative refinement.73 Incorporating a similar RAG approach could enrich the chemical context available to the LLM, potentially improving the diversity, novelty, and relevance of generated candidates, and represents a promising direction for future work.
000 literature-extracted entries was used to inform LLM-driven candidate generation and iterative refinement.73 Incorporating a similar RAG approach could enrich the chemical context available to the LLM, potentially improving the diversity, novelty, and relevance of generated candidates, and represents a promising direction for future work.
Looking ahead, several directions can further strengthen and extend this framework. The workflow is inherently modular and not restricted to the lohc candidates presented here. By adjusting the seed molecules, filtering criteria, and ml models trained on the relevant target property, the same protocol can be adapted to discover functionalized lohcs, alternative hydrogen carriers, or molecules for entirely different applications such as electrolytes, solvents, or organic semiconductors. Furthermore, the iterative design loop implicitly steers the llm toward favorable regions of chemical space, as accepted molecules from one cycle serve as seeds for the next, progressively narrowing the search toward structures that satisfy the physicochemical constraints. While this does not constitute direct molecular optimization, it provides a practical mechanism for discovering structures with targeted property combinations, and the filtering thresholds for properties such as melting point and hydrogen capacity can be systematically tightened or relaxed to further guide this process. Additionally, integrating catalyst prediction into the workflow would provide a more complete picture of the lohc design process. Training ml models on catalyst performance data could enable simultaneous optimization of both the carrier molecule and its catalytic system, bridging molecular discovery with process-level design. This includes composition, metal–support interactions, and reaction conditions. Future efforts should also account for toxicity, environmental impact, and scalability of synthesis to ensure that discovered candidates are not only thermodynamically favorable but also safe and industrially viable.
4 Methods
4.1 Selection of seed molecules
To initiate the LLM-guided molecular generation, selected seed molecules based on the design rules established in previous work40 were chosen (Fig. 5; Text S3; see SI). Note that this group contains experimentally evaluated LOHC candidates (Expt-31; known ΔH values) as well as molecules identified from prior computational screening (LOHC-7).40 The latter consists of the molecules discovered by using high-throughput screening, quantum chemical calculations, and practical down-selection. This dual selection strategy balances experimentally validated structures with computationally curated discoveries, allowing us to assess whether the LLM can generate novel candidates that align with empirical and theoretical insights in LOHC chemistry (Fig. 5).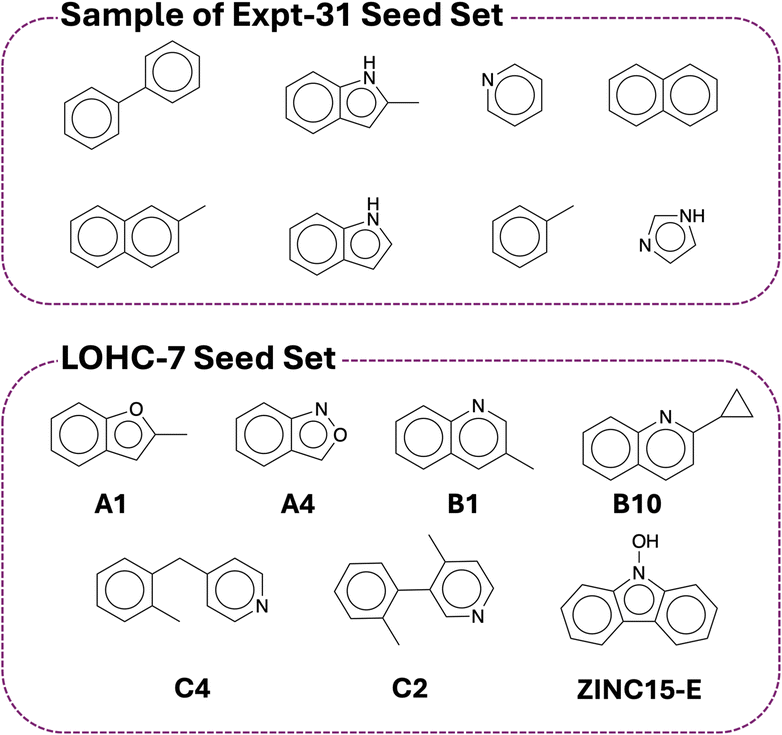 | ||
| Fig. 5 Overview of the seed molecules used to prompt the LLM. The top panel displays eight molecules selected from the Expt-31 dataset, which consists of experimentally known LOHCs. The bottom panel shows seven molecules chosen from previous work on LOHC design.40 | ||
4.2 Machine learning (ML)
As part of the workflow, a ML model capable of predicting hydrogenation enthalpy (ΔH) directly from molecular structures (e.g.: SMILE strings) was developed (Fig. 3, orange box). This ML model was trained QM9-LOHC,60 a high-fidelity computational dataset containing 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 373 dehydrogenation reactions.60 The dataset includes reactions with hydrogen storage capacities of 5.5 wt% H2 or higher, and ΔH values computed at the G4MP2 (ref. 74) level of theory (Fig. 6a). For model training, each molecule was represented using 2048-bit Morgan fingerprints (ECFP4).75 The dataset was randomly split (80
373 dehydrogenation reactions.60 The dataset includes reactions with hydrogen storage capacities of 5.5 wt% H2 or higher, and ΔH values computed at the G4MP2 (ref. 74) level of theory (Fig. 6a). For model training, each molecule was represented using 2048-bit Morgan fingerprints (ECFP4).75 The dataset was randomly split (80![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 20), with 80% used for training and 20% reserved for testing. Then RF regressor76 to predict ΔH was trained using default hyperparameters, and 100 trees were used in the model. The trained model exhibited good predictive accuracy, achieving mean absolute error (MAE) = 4.67 kJ mol−1, root mean squared deviation (RMSD) = 7.35 kJ mol−1, and coefficient of determination (R2) = 0.93, with a high correlation between predicted and computed ΔH values. A parity plot shown in Fig. 6b, comparing G4MP2 computed vs. ML predicted ΔH confirmed its reliability for screening newly generated LOHC candidates. This ML model was subsequently integrated into the iterative molecular generation process to prioritize molecules within the optimal ΔH range of 40–70 kJ mol−1.
20), with 80% used for training and 20% reserved for testing. Then RF regressor76 to predict ΔH was trained using default hyperparameters, and 100 trees were used in the model. The trained model exhibited good predictive accuracy, achieving mean absolute error (MAE) = 4.67 kJ mol−1, root mean squared deviation (RMSD) = 7.35 kJ mol−1, and coefficient of determination (R2) = 0.93, with a high correlation between predicted and computed ΔH values. A parity plot shown in Fig. 6b, comparing G4MP2 computed vs. ML predicted ΔH confirmed its reliability for screening newly generated LOHC candidates. This ML model was subsequently integrated into the iterative molecular generation process to prioritize molecules within the optimal ΔH range of 40–70 kJ mol−1.
4.3 LLM agent
To implement the generative component of the workflow, an LLM-driven molecular design agent capable of proposing new LOHC candidates was established (Fig. 3, green box). This agent was built using Argonne's Argo interface to GPT-o1preview model, to systematically generate molecular structures based on provided seed molecules. The LLM agent was constructed with a system prompt that explicitly defines its role, instructing it to act as an expert in molecular design specializing in LOHC. This prompt ensures that the model focuses on functionally relevant molecular motifs rather than arbitrarily generating structures. The core of the agent consists of three steps.4.4 Seeding the prompt with known molecules
The LLM is provided with an initial list of known LOHC SMILES strings, serving as guiding examples to ground the generation process in known chemistry. These molecules establish the structural space in which the LLM operates and generate chemically consistent candidates.4.5 Generating novel molecules
The LLM is explicitly prompted to produce 30 SMILE strings, that are unique, chemically valid, and distinct from the provided list. This step ensures that the agent expands molecular diversity while maintaining relevance to LOHC chemistry.4.6 Structured output collection
The generated molecules are returned in a JSON format, containing only a structured list of SMILES strings. This was followed by parsing and integration into downstream validation and screening steps. If the LLM deviates from this structure, its response is discarded, and it is re-prompted to ensure compliance with the required format.The generated molecules serve as inputs for further refinement and validation in subsequent steps of the workflow. The full system prompt is given below:
4.7 Iterative generation and refinement
The LLM-generated molecules go through a multi-stage filtering process to ensure chemical feasibility and suitability as LOHCs (Fig. 2, purple box). This iterative workflow was designed to generate and refine molecular candidates during ten iterations. The process continued until either 200 valid SMILES were collected or 10 iterations were completed, whichever came first. To account for variability in the LLM responses, each iteration was given up to three attempts. If no new molecules were generated within an iteration, the LLM was re-prompted to ensure that the search space was sufficiently explored. The first filtering step involved a chemical validity verification, where each generated SMILE string was processed using RDKit to confirm their chemically valid molecular structure. Chemically invalid molecules suggested during this step was discarded to maintain structural integrity in the dataset. Next, we applied hydrogen storage capacity screening, where the wt% H2 of each molecule was computed using eqn (1), which quantifies the hydrogen content as a percentage of the molecule's total mass before (MWH-lean) and after full hydrogenation (MWH-rich):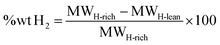 | (1) |
At this stage of screening, only molecules with wt% H2 ≥ 5.5% were retained. Candidates that passed this threshold were then evaluated for hydrogenation enthalpy (ΔH) using a trained ML model, with only those falling within the desired 40 ≤ ΔH ≤ 70 kJ mol−1 per H2 ranges being selected.36,39 Further, MP predictions were incorporated, as LOHCs must remain in the liquid phase under ambient conditions for ease of handling and transport. Using Leruli's MP prediction platform based on OPERA77 model, molecules with MP > 40 °C were filtered out. Duplicate removal was enforced at multiple stages, including after generation, filtering, and before final dataset compilation ensuring that the final LOHC set contained unique and chemically distinct molecules from initial seed set.
At the end of each iteration of the workflow, the molecules that successfully passed filtering were merged with those from previous iterations and used as an updated prompt for the LLM in the subsequent cycle. This process allowed the LLM to iteratively refine and expand the chemical space, generating increasingly viable LOHC candidates. Upon completion of ten iterations, the final dataset was compiled, containing SMILES representations of all validated LOHC candidates, alongside their predicted ΔH values, wt% H2, and MP. The details of the workflow and generated data are described in the GitHub repository: https://github.com/HydrogenStorage/LLM_LOHC.
4.8 Computation of enthalpies
In order to validate the ΔH predictions using ML, quantum chemical calculations using Gaussian 16 software78 using accurate G4MP2 (ref. 74) method were performed. The G4MP2 is a composite method based on the G4 theory that utilizes MP2 perturbation theory to enhance computational efficiency. The minimum energy molecular conformers were first determined using the Universal Force Field (UFF) method in RDKit. The G4MP2 approach relies on geometries optimized at the B3LYP/6-31G(2df,p) level of theory,79–82 followed by a series of high-level single-point energy calculations. The Zero-point energy (EZPE) corrections are derived from the B3LYP/6-31G(2df,p) computed vibrational frequencies, scaled by a factor of 0.984 to account for anharmonicity.74 We confirmed the nature of each located stationary point on the potential energy surface by the absence of imaginary frequencies. Additional details about the G4MP2 method is described in the SI (Text S1).4.9 Catalytic hydrogenation of LOHCs
Four catalysts were tested for the hydrogenation of 3-methyl pyridine and toluene. Two commercial catalysts, 10 wt% Pd/C (Sigma Aldrich), and 5 wt% Rh/Al2O3 (Degussa) and two home-made catalysts, 10 wt% Pd/Al2O3 and 5 wt% Pd/NU-1000. Pd/Al2O3 was prepared using the incipient wetness technique by adding a sufficient amount of a solution containing palladium nitrate solution (Sigma Aldrich) to Al2O3 support (Sigma Aldrich) and Pd/NU-100 was synthesized as described elsewhere.83 Screening of all the catalysts and control (no catalyst) were carried out in 48-well plate using the Screening Pressure Reactor (SPR, Unchained Labs Inc). 10 mg of catalysts was dispensed in 1/2dr shell vials with 100 µL of either 3-methylpyridne (#5, Fig. 5) (≥99.5%, Sigma Aldrich) 1-ethylnapthalene (#12) (≥95%, Oakwood), 1,1-diphenylethane (#32) (≥97%, Ambeed), 9-ethylcarbazole (≥97%, Sigma Aldrich), benzofuran (>99%, Sigma Aldrich), or toluene (≥99.5%, Sigma Aldrich) in 1 mL dodecane (≥99, Sigma Aldrich). The multiwell plate with vials was then covered with a pinhole graphite gasket and a stainless-steel pinhole plate to ensure gas diffusion but to minimize cross-contamination between the vials. Initially, the SPR was flushed with 500 mL min−1 N2 for 15 min at room temperature, at an orbital shaking of 150 rpm and pressurized with H2 initially to minimize evaporation of the reagents and solvent. The reactor was then heated up slowly (10 °C min−1 ramp rate) to either 150 °C or 200 °C. Under the given conditions, the pressure of the reactor reached around 300 psi or 600 psi, respectively. After 12 h, the shaking was stopped, the reactor was cooled down to room temperature and was flushed with 100 mL min−1 N2 for 15 min. Aliquots were transferred into filter vials (Whatman Mini-UniPrep Syringeless Filter, 0.2 µm). Aliquots were analysed sequentially by a GC Ultra Gas Chromatograph system equipped with a Tri Plus RSH autosampler, an ISQ MS detector, and a FID (Thermo Scientific). The column used for the MS detector was an Agilent J&W DB-5 column (30 m × 0.25 mm × 0.25 µm film thickness) while the column used for the FID was an Agilent J&W DB-5MS column (30 m × 0.25 mm × 0.25 µm film thickness). GC data were analysed using the Thermo Xcalibur 2.2 SP1.48 software. The following method was used: a 0.5 µL split injection S5 with a split ratio of 100 run under a constant gas flow of 1 mL min−1. The oven temperature profile was as follows: initial temperature = 30 °C, hold for 10 minutes, ramp at 20 °C min−1, final temperature = 250 °C. The conversions of the substrates were determined based on the sum of the peaks. For the recycling experiments, the spent catalysts (Rh/Al2O3 and Pt/NU-1000) were washed 3 times with toluene and one time with pentane. After drying the catalysts were re-used.Author contributions
H. H. and R. S. A. conceived the project and performed all computational, machine learning, and LLM-based molecular discovery work. T. A. G., S. L., and O. K. F. contributed expertise on catalyst design, synthesis, and mechanistic analysis. M. S. F. and M. D. carried out experimental validation and characterization studies. All authors discussed the results and contributed to the final manuscript.Conflicts of interest
The authors declare no competing financial interest.Data availability
All data associated with this study are publicly available. The repository includes SMILES strings of the starting materials, the trained Random Forest model, workflow scripts, and representative output files. The code and data can be accessed at https://github.com/HassanHarb92/LLM_LOHC and are archived at https://doi.org/10.5281/zenodo.18853735.Supplementary information (SI): including an overview of the G4MP2 method, seed sets with calculated and experimental hydrogenation enthalpies, comparisons for down-selected molecules, chemical and physical properties of new LOHCs, LLM and ML performance analyses, design rules, hydrogenation schemes, and synthesis details for Pd/NU-1000. All code used in this study is present on GitHub: https://github.com/HydrogenStorage/LLM_LOHC. See DOI: https://doi.org/10.1039/d6dd00102e.
Acknowledgements
This material is based upon work supported by Laboratory Directed Research and Development (LDRD) funding from Argonne National Laboratory, provided by the Director, Office of Science, U.S. Department of Energy, under Contract No. DE-AC02-06CH11357. We acknowledge computing resources provided by “BEBOP,” a cluster operated by the Laboratory Computing Resource Center (LCRC) at Argonne National Laboratory. Prompting was conducted using Argo, Argonne's internal generative AI chatbot, operated by the Business and Information Services (BIS) division. Experimental work was supported by the Catalyst Design for Decarbonization Center (CD4DC), an Energy Frontier Research Center funded by the DOE Office of Science, Basic Energy Sciences (BES), under Award No. DE-SC0023383. ChatGPT v5.1 was used to assist with editing this manuscript.References
- D. Bhowmik, P. Zhang, Z. Fox, S. Irle and J. Gounley, Enhancing Molecular Design Efficiency: Uniting Language Models and Generative Networks with Genetic Algorithms, Patterns, 2024, 5(4), 100947, DOI:10.1016/j.patter.2024.100947.
- D. Menon and R. Ranganathan, A Generative Approach to Materials Discovery, Design, and Optimization, ACS Omega, 2022, 7(30), 25958–25973, DOI:10.1021/acsomega.2c03264.
- C. Bilodeau, W. Jin, T. Jaakkola, R. Barzilay and K. F. Jensen, Generative Models for Molecular Discovery: Recent Advances and Challenges, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2022, 12(5), e1608, DOI:10.1002/wcms.1608.
- B. Sanchez-Lengeling and A. Aspuru-Guzik, Inverse Molecular Design Using Machine Learning: Generative Models for Matter Engineering, Science, 2018, 361(6400), 360–365, DOI:10.1126/science.aat2663.
- Y. Zimmermann, A. Bazgir, A. Al-Feghali, M. Ansari, J. Bocarsly, L. C. Brinson, Y. Chiang, D. Circi, M.-H. Chiu, N. Daelman, M. L. Evans, A. S. Gangan, J. George, H. Harb, G. Khalighinejad, S. Takrim Khan, S. Klawohn, M. Lederbauer, S. Mahjoubi, B. Mohr, S. Mohamad Moosavi, A. Naik, A. Beste Ozhan, D. Plessers, A. Roy, F. Schöppach, P. Schwaller, C. Terboven, K. Ueltzen, Y. Wu, S. Zhu, J. Janssen, C. Li, I. Foster and B. Blaiszik, 32 Examples of LLM Applications in Materials Science and Chemistry: Towards Automation, Assistants, Agents, and Accelerated Scientific Discovery, Mach. Learn. Sci. Technol., 2025, 6(3), 030701, DOI:10.1088/2632-2153/ae011a.
- O. A. von Lilienfeld, K.-R. Müller and A. Tkatchenko, Exploring Chemical Compound Space with Quantum-Based Machine Learning, Nat. Rev. Chem., 2020, 4(7), 347–358, DOI:10.1038/s41570-020-0189-9.
- N. K. Dandu, L. Ward, R. S. Assary, P. C. Redfern and L. A. Curtiss, Accurate Prediction of Adiabatic Ionization Potentials of Organic Molecules Using Quantum Chemistry Assisted Machine Learning, J. Phys. Chem. A, 2023, 127(28), 5914–5920, DOI:10.1021/acs.jpca.3c00823.
- J. Vamathevan, D. Clark, P. Czodrowski, I. Dunham, E. Ferran, G. Lee, B. Li, A. Madabhushi, P. Shah, M. Spitzer and S. Zhao, Applications of Machine Learning in Drug Discovery and Development, Nat. Rev. Drug Discovery, 2019, 18(6), 463–477, DOI:10.1038/s41573-019-0024-5.
- M. M. Rashidi, M. Alhuyi Nazari, C. Harley, E. Momoniat, I. Mahariq and N. Ali, Applications of Machine Learning Methods for Boiling Modeling and Prediction: A Comprehensive Review, Chem. Thermodyn. Therm. Anal., 2022, 8, 100081, DOI:10.1016/j.ctta.2022.100081.
- J. A. Keith, V. Vassilev-Galindo, B. Cheng, S. Chmiela, M. Gastegger, K.-R. Müller and A. Tkatchenko, Combining Machine Learning and Computational Chemistry for Predictive Insights Into Chemical Systems, Chem. Rev., 2021, 121(16), 9816–9872, DOI:10.1021/acs.chemrev.1c00107.
- M. Sajjan, J. Li, R. Selvarajan, S. H. Sureshbabu, S. S. Kale, R. Gupta, V. Singh and S. Kais, Quantum Machine Learning for Chemistry and Physics, Chem. Soc. Rev., 2022, 51(15), 6475–6573, 10.1039/D2CS00203E.
- L. Patel, T. Shukla, X. Huang, D. W. Ussery and S. Wang, Machine Learning Methods in Drug Discovery, Molecules, 2020, 25(22), 5277, DOI:10.3390/molecules25225277.
- J. F. Gaviria, G. Narváez, C. Guillen, L. F. Giraldo and M. Bressan, Machine Learning in Photovoltaic Systems: A Review, Renewable Energy, 2022, 196, 298–318, DOI:10.1016/j.renene.2022.06.105.
- W. Yang, T. T. Fidelis and W.-H. Sun, Machine Learning in Catalysis, From Proposal to Practicing, ACS Omega, 2020, 5(1), 83–88, DOI:10.1021/acsomega.9b03673.
- E. Kasneci, K. Sessler, S. Küchemann, M. Bannert, D. Dementieva, F. Fischer, U. Gasser, G. Groh, S. Günnemann, E. Hüllermeier, S. Krusche, G. Kutyniok, T. Michaeli, C. Nerdel, J. Pfeffer, O. Poquet, M. Sailer, A. Schmidt, T. Seidel, M. Stadler, J. Weller, J. Kuhn and G. Kasneci, ChatGPT for Good? On Opportunities and Challenges of Large Language Models for Education, Learn. Individ. Differ., 2023, 103, 102274, DOI:10.1016/j.lindif.2023.102274.
- Y. Zimmermann, A. Bazgir, Z. Afzal, F. Agbere, Q. Ai, N. Alampara, A. Al-Feghali, M. Ansari, D. Antypov, A. Aswad, J. Bai, V. Baibakova, D. D. Biswajeet, E. Bitzek, J. D. Bocarsly, A. Borisova, A. M. Bran, L. C. Brinson, M. M. Calderon, A. Canalicchio, V. Chen, Y. Chiang, D. Circi, B. Charmes, V. Chaudhary, Z. Chen, M.-H. Chiu, J. Clymo, K. Dabhadkar, N. Daelman, A. Datar, M. L. Evans, M. G. Fard, G. Fisicaro, A. S. Gangan, J. George, J. D. C. Gonzalez, M. Götte, A. K. Gupta, H. Harb, P. Hong, A. Ibrahim, A. Ilyas, A. Imran, K. Ishimwe, R. Issa, K. M. Jablonka, C. Jones, T. R. Josephson, G. Juhasz, S. Kapoor, R. Kang, G. Khalighinejad, S. Khan, S. Klawohn, S. Kuman, A. N. Ladines, S. Leang, M. Lederbauer, S.-L. M. Liao, H. Liu, X. Liu, S. Lo, S. Madireddy, P. R. Maharana, S. Maheshwari, S. Mahjoubi, J. A. Márquez, R. Mills, T. Mohanty, B. Mohr, S. M. Moosavi, A. Moßhammer, A. D. Naghdi, A. Naik, O. Narykov, H. Näsström, X. V. Nguyen, X. Ni, D. O'Connor, T. Olayiwola, F. Ottomano, A. B. Ozhan, S. Pagel, C. Parida, J. Park, V. Patel, E. Patyukova, M. H. Petersen, L. Pinto, J. M. Pizarro, D. Plessers, T. Pradhan, U. Pratiush, C. Puli, A. Qin, M. Rajabi, F. Ricci, E. Risch, M. Ríos-García, A. Roy, T. Rug, H. M. Sayeed, M. Scheidgen, M. Schilling-Wilhelmi, M. Schloz, F. Schöppach, J. Schumann, P. Schwaller, M. Schwarting, S. Sharlin, K. Shen, J. Shi, P. Si, J. D'Souza, T. Sparks, S. Sudhakar, L. Talirz, D. Tang, O. Taran, C. Terboven, M. Tropin, A. Tsymbal, K. Ueltzen, P. A. Unzueta, A. Vasan, T. Vinchurkar, T. Vo, G. Vogel, C. Völker, J. Weinreich, F. Yang, M. Zaki, C. Zhang, S. Zhang, W. Zhang, R. Zhu, S. Zhu, J. Janssen, I. Foster and B. Blaiszik, Reflections from the 2024 Large Language Model (LLM) Hackathon for Applications in Materials Science and Chemistry, arXiv, 2024, preprint, arXiv.2411.15221, DOI:10.48550/ARXIV.2411.15221.
- K. M. Jablonka, Q. Ai, A. Al-Feghali, S. Badhwar, J. D. Bocarsly, A. M. Bran, S. Bringuier, L. C. Brinson, K. Choudhary, D. Circi, S. Cox, W. A. De Jong, M. L. Evans, N. Gastellu, J. Genzling, M. V. Gil, A. K. Gupta, Z. Hong, A. Imran, S. Kruschwitz, A. Labarre, J. Lála, T. Liu, S. Ma, S. Majumdar, G. W. Merz, N. Moitessier, E. Moubarak, B. Mouriño, B. Pelkie, M. Pieler, M. C. Ramos, B. Ranković, S. G. Rodriques, J. N. Sanders, P. Schwaller, M. Schwarting, J. Shi, B. Smit, B. E. Smith, J. Van Herck, C. Völker, L. Ward, S. Warren, B. Weiser, S. Zhang, X. Zhang, G. A. Zia, A. Scourtas, K. J. Schmidt, I. Foster, A. D. White and B. Blaiszik, 14 Examples of How LLMs Can Transform Materials Science and Chemistry: A Reflection on a Large Language Model Hackathon, Digit. Discov., 2023, 2(5), 1233–1250, 10.1039/D3DD00113J.
- G. Marvin, N. Hellen, D. Jjingo and J. Nakatumba-Nabende, Prompt Engineering in Large Language Models, in Data Intelligence and Cognitive Informatics; Algorithms for Intelligent Systems, I. J. Jacob, S. Piramuthu and P. Falkowski-Gilski, ed. Springer Nature Singapore, Singapore, 2024, pp. 387–402, DOI:10.1007/978-981-99-7962-2_30.
- A. Birhane, A. Kasirzadeh, D. Leslie and S. Wachter, Science in the Age of Large Language Models, Nat. Rev. Phys., 2023, 5(5), 277–280, DOI:10.1038/s42254-023-00581-4.
- J. Kaddour, J. Harris, M. Mozes, H. Bradley, R. Raileanu and R. McHardy, Challenges and Applications of Large Language Models, arXiv, 2023, preprint, arXiv.2307.10169, DOI:10.48550/ARXIV.2307.10169.
- A. Malusare and V. Aggarwal, Improving Molecule Generation and Drug Discovery With a Knowledge-Enhanced Generative Model, IEEE Trans. Comput. Biol. Bioinforma., 2025, 22(1), 375–381, DOI:10.1109/TCBB.2024.3477313.
- M. Han, J. F. Joung, M. Jeong, D. H. Choi and S. Park, Generative Deep Learning-Based Efficient Design of Organic Molecules with Tailored Properties, ACS Cent. Sci., 2025, 11(2), 219–227, DOI:10.1021/acscentsci.4c00656.
- V. Dorna, D. Subhalingam, K. Kolluru, S. Tuli, M. Singh, S. Singal, N. M. A. Krishnan and S. Ranu, TAGMol: Target-Aware Gradient-Guided Molecule Generation, arXiv, 2024, preprint, arXiv.2406.01650, DOI:10.48550/ARXIV.2406.01650.
- J. Westermayr, J. Gilkes, R. Barrett and R. J. Maurer, High-Throughput Property-Driven Generative Design of Functional Organic Molecules, Nat. Comput. Sci., 2023, 3(2), 139–148, DOI:10.1038/s43588-022-00391-1.
- R. R. Kotkondawar, S. R. Sutar, A. W. Kiwelekar, V. J. Kadam and S. M. Jadhav, A Generative Framework for Enhancing Drug Target Interaction Prediction in Drug Discovery, Sci. Rep., 2025, 15(1), 35588, DOI:10.1038/s41598-025-01589-9.
- F. Bhuiyan, H. Harb, R. Assary and Á. Vázquez-Mayagoitia, Redox Potential Prediction of Fe(II)/Fe(III) Complexes: A Density Functional Theory and Graph Neural Network Approach, ChemRxiv, 2025, preprint, DOI:10.26434/chemrxiv-2025-t9kmf.
- A. Chowdhury, H. Harb, R. Egele, C. Alves, F. H. Bhuyan, H. A. Doan; A. Vazquez-Mayagoitia, R. S. Assary and P. Balaprakash, Automated Learning of GNN Ensembles for Predicting Redox Potentials with Uncertainty, ChemRxiv, 2025, preprint, DOI:10.26434/chemrxiv-2025-0tq7j-v2.
- H. A. Doan, C. Li, L. Ward, M. Zhou, L. A. Curtiss and R. S. Assary, Accelerating the Evaluation of Crucial Descriptors for Catalyst Screening via Message Passing Neural Network, Digit. Discov., 2023, 2(1), 59–68, 10.1039/D2DD00088A.
- L. Ward, G. Sivaraman, J. G. Pauloski, Y. Babuji, R. Chard, N. Dandu, P. C. Redfern, R. S. Assary, K. Chard, L. A. Curtiss, R. Thakur and I. C. Foster, Scalable Machine-Learning-Based Steering of Ensemble Simulations for High Performance Computing, in 2021 IEEE/ACM Workshop on Machine Learning in High Performance Computing Environments (MLHPC), IEEE, St. Louis, MO, USA, 2021, pp. 9–20, DOI:10.1109/MLHPC54614.2021.00007.
- H. A. Doan, G. Agarwal, H. Qian, M. J. Counihan, J. Rodríguez-López, J. S. Moore and R. S. Assary, Quantum Chemistry-Informed Active Learning to Accelerate the Design and Discovery of Sustainable Energy Storage Materials, Chem. Mater., 2020, 32(15), 6338–6346, DOI:10.1021/acs.chemmater.0c00768.
- D. P. Kingma and M. Welling, Auto-Encoding Variational Bayes, arXiv, 2013, preprint, arXiv.1312.6114, DOI:10.48550/ARXIV.1312.6114.
- A. Brock, J. Donahue and K. Simonyan, Large Scale GAN Training for High Fidelity Natural Image Synthesis, arXiv, 2018, preprint, arXiv.1809.11096, DOI:10.48550/ARXIV.1809.11096.
- C. Zang and F. Wang, MoFlow: An Invertible Flow Model for Generating Molecular Graphs, in Proceedings of the 26th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining; ACM, Virtual Event CA USA, 2020, pp. 617–626, DOI:10.1145/3394486.3403104.
- H. H. Loeffler, S. Wan, M. Klähn, A. P. Bhati and P. V. Coveney, Optimal Molecular Design: Generative Active Learning Combining REINVENT with Precise Binding Free Energy Ranking Simulations, J. Chem. Theory Comput., 2024, 20(18), 8308–8328, DOI:10.1021/acs.jctc.4c00576.
- X. Yan, N. Hudson, H. Park, D. Grzenda, J. G. Pauloski, M. Schwarting, H. Pan, H. Harb, S. Foreman, C. Knight, T. Gibbs, K. Chard, S. Chaudhuri, E. Tajkhorshid, I. Foster, M. Moosavi, L. Ward, E. A. Huerta, MOFA: Discovering Materials for Carbon Capture with a GenAI- and Simulation-Based Workflow, arXiv, 2025, preprint, arXiv.2501.10651, DOI:10.48550/ARXIV.2501.10651.
- P. Preuster, C. Papp and P. Wasserscheid, Liquid Organic Hydrogen Carriers (LOHCs): Toward a Hydrogen-Free Hydrogen Economy, Acc. Chem. Res., 2017, 50(1), 74–85, DOI:10.1021/acs.accounts.6b00474.
- T. He, Q. Pei and P. Chen, Liquid Organic Hydrogen Carriers, J. Energy Chem., 2015, 24(5), 587–594, DOI:10.1016/j.jechem.2015.08.007.
- C. Chu, K. Wu, B. Luo, Q. Cao and H. Zhang, Hydrogen Storage by Liquid Organic Hydrogen Carriers: Catalyst, Renewable Carrier, and Technology - A Review, Carbon Resour. Convers., 2023, S2588913323000248, DOI:10.1016/j.crcon.2023.03.007.
- P. T. Aakko-Saksa, C. Cook, J. Kiviaho and T. Repo, Liquid Organic Hydrogen Carriers for Transportation and Storing of Renewable Energy – Review and Discussion, J. Power Sources, 2018, 396, 803–823, DOI:10.1016/j.jpowsour.2018.04.011.
- H. Harb, S. N. Elliott, L. Ward, I. T. Foster, S. J. Klippenstein, L. A. Curtiss and R. S. Assary, Uncovering Novel Liquid Organic Hydrogen Carriers: A Systematic Exploration of Chemical Compound Space Using Cheminformatics and Quantum Chemical Methods, Digit. Discov., 2023, 2(5), 1233–1250, 10.1039/D3DD00123G.
- G. Vishwakarma and J. Hachmann, Liquid Organic Hydrogen Carriers: High-Throughput Screening of Homogeneous Catalysts, ChemRxiv, 2023, preprint, DOI:10.26434/chemrxiv-2023-s8pkf.
- K. Paragian, B. Li, M. Massino and S. Rangarajan, A Computational Workflow to Discover Novel Liquid Organic Hydrogen Carriers and Their Dehydrogenation Routes, Mol. Syst. Des. Eng., 2020, 5(10), 1658–1670, 10.1039/D0ME00105H.
- P. Modisha and D. Bessarabov, Aromatic Liquid Organic Hydrogen Carriers for Hydrogen Storage and Release, Curr. Opin. Green Sustain. Chem., 2023, 42, 100820, DOI:10.1016/j.cogsc.2023.100820.
- D. Teichmann, K. Stark, K. Müller, G. Zöttl, P. Wasserscheid and W. Arlt, Energy Storage in Residential and Commercial Buildings via Liquid Organic Hydrogen Carriers (LOHC), Energy Environ. Sci., 2012, 5(10), 9044, 10.1039/c2ee22070a.
- F. Valentini, A. Marrocchi and L. Vaccaro, Liquid Organic Hydrogen Carriers (LOHCs) as H-Source for Bio-Derived Fuels and Additives Production, Adv. Energy Mater., 2022, 12(13), 2103362, DOI:10.1002/aenm.202103362.
- M. Niermann, S. Drünert, M. Kaltschmitt and K. Bonhoff, Liquid Organic Hydrogen Carriers (LOHCs) – Techno-Economic Analysis of LOHCs in a Defined Process Chain, Energy Environ. Sci., 2019, 12(1), 290–307, 10.1039/C8EE02700E.
- M. Niermann, S. Timmerberg, S. Drünert and M. Kaltschmitt, Liquid Organic Hydrogen Carriers and Alternatives for International Transport of Renewable Hydrogen, Renew. Sustain. Energy Rev., 2021, 135, 110171, DOI:10.1016/j.rser.2020.110171.
- P. C. Rao and M. Yoon, Potential Liquid-Organic Hydrogen Carrier (LOHC) Systems: A Review on Recent Progress, Energies, 2020, 13(22), 6040, DOI:10.3390/en13226040.
- L. Mulky, S. Srivastava, T. Lakshmi, E. R. Sandadi, S. Gour, N. A. Thomas, S. Shanmuga Priya and K. Sudhakar, An Overview of Hydrogen Storage Technologies – Key Challenges and Opportunities, Mater. Chem. Phys., 2024, 325, 129710, DOI:10.1016/j.matchemphys.2024.129710.
- D. Mori and K. Hirose, Recent Challenges of Hydrogen Storage Technologies for Fuel Cell Vehicles, Int. J. Hydrog. Energy, 2009, 34(10), 4569–4574, DOI:10.1016/j.ijhydene.2008.07.115.
- M. Brodt, K. Müller, J. Kerres, I. Katsounaros, K. Mayrhofer, P. Preuster, P. Wasserscheid and S. Thiele, The 2-Propanol Fuel Cell: A Review from the Perspective of a Hydrogen Energy Economy, Energy Technol., 2021, 9(9), 2100164, DOI:10.1002/ente.202100164.
- J.-Y. Cho, H. Kim, J.-E. Oh and B. Y. Park, Recent Advances in Homogeneous/Heterogeneous Catalytic Hydrogenation and Dehydrogenation for Potential Liquid Organic Hydrogen Carrier (LOHC) Systems, Catalysts, 2021, 11(12), 1497, DOI:10.3390/catal11121497.
- P. M. Modisha, C. N. M. Ouma, R. Garidzirai, P. Wasserscheid and D. Bessarabov, The Prospect of Hydrogen Storage Using Liquid Organic Hydrogen Carriers, Energy Fuels, 2019, 33(4), 2778–2796, DOI:10.1021/acs.energyfuels.9b00296.
- D. Wei, X. Shi, R. Qu, K. Junge, H. Junge and M. Beller, Toward a Hydrogen Economy: Development of Heterogeneous Catalysts for Chemical Hydrogen Storage and Release Reactions, ACS Energy Lett., 2022, 7(10), 3734–3752, DOI:10.1021/acsenergylett.2c01850.
- D. Dean, B. Davis and P. G. Jessop, The Effect of Temperature, Catalyst and Sterics on the Rate of N-Heterocycledehydrogenation for Hydrogenstorage, New J. Chem., 2011, 35(2), 417–422, 10.1039/C0NJ00511H.
- M. Markiewicz, Y.-Q. Zhang, M. T. Empl, M. Lykaki, J. Thöming, P. Steinberg and S. Stolte, Hazard Assessment of Quinaldine-, Alkylcarbazole-, Benzene- and Toluene-Based Liquid Organic Hydrogen Carrier (LOHCs) Systems, Energy Environ. Sci., 2019, 12(1), 366–383, 10.1039/C8EE01696H.
- M. Markiewicz, Y. Q. Zhang, A. Bösmann, N. Brückner, J. Thöming, P. Wasserscheid and S. Stolte, Environmental and Health Impact Assessment of Liquid Organic Hydrogen Carrier (LOHC) Systems – Challenges and Preliminary Results, Energy Environ. Sci., 2015, 8(3), 1035–1045, 10.1039/C4EE03528C.
- T. Sterling and J. J. Irwin, ZINC 15 – Ligand Discovery for Everyone, J. Chem. Inf. Model., 2015, 55(11), 2324–2337, DOI:10.1021/acs.jcim.5b00559.
- L. Ruddigkeit, R. van Deursen, L. C. Blum and J.-L. Reymond, Enumeration of 166 Billion Organic Small Molecules in the Chemical Universe Database GDB-17, J. Chem. Inf. Model., 2012, 52(11), 2864–2875, DOI:10.1021/ci300415d.
- H. Harb, S. N. Elliott, L. Ward, I. T. Foster, S. J. Klippenstein, L. A. Curtiss and R. S. Assary, Accurate Dehydrogenation Enthalpies Dataset for Liquid Organic Hydrogen Carriers, Sci. Data, 2025, 12(1), 171, DOI:10.1038/s41597-025-04468-0.
- PubChem, PubChem, https://pubchem.ncbi.nlm.nih.gov/, accessed 2023-10-09.
- S. Rangarajan, T. Kaminski, E. Van Wyk, A. Bhan and P. Daoutidis, Language-Oriented Rule-Based Reaction Network Generation and Analysis: Algorithms of RING, Comput. Chem. Eng., 2014, 64, 124–137, DOI:10.1016/j.compchemeng.2014.02.007.
- K. Mansouri, C. M. Grulke, R. S. Judson and A. J. Williams, OPERA Models for Predicting Physicochemical Properties and Environmental Fate Endpoints, J. Cheminform., 2018, 10(1), 10, DOI:10.1186/s13321-018-0263-1.
- B. Huang and O. A. von Lilienfeld, Ab Initio Machine Learning in Chemical Compound Space, Chem. Rev., 2021, 121(16), 10001–10036, DOI:10.1021/acs.chemrev.0c01303.
- A. S. Lee, S. Elliott, H. Harb, L. Ward, I. Foster, L. Curtiss and R. S. E. Assary, A First-Principles Thermochemical Descriptor for Predicting Molecular Synthesizability, J. Chem. Inf. Model., 2024, 3c01583, DOI:10.1021/acs.jcim.3c01583.
- . A. Karakhanov and E. A. Viktorova, Hydrogenation and Dehydrogenation Reactions of Benzofuran and Its Derivatives, Chem. Heterocycl. Compd., 1976, 12(4), 367–375, DOI:10.1007/BF00480416.
- I. F. Teixeira, B. T. W. Lo, P. Kostetskyy, L. Ye, C. C. Tang, G. Mpourmpakis and S. C. E. Tsang, Direct Catalytic Conversion of Biomass-Derived Furan and Ethanol to Ethylbenzene, ACS Catal., 2018, 8(3), 1843–1850, DOI:10.1021/acscatal.7b03952.
- K. M. Eblagon, D. Rentsch, O. Friedrichs, A. Remhof, A. Zuettel, A. J. Ramirez-Cuesta and S. C. Tsang, Hydrogenation of 9-Ethylcarbazole as a Prototype of a Liquid Hydrogen Carrier, Int. J. Hydrog. Energy, 2010, 35(20), 11609–11621, DOI:10.1016/j.ijhydene.2010.03.068.
- F. Sotoodeh and K. J. Smith, Kinetics of Hydrogen Uptake and Release from Heteroaromatic Compounds for Hydrogen Storage, Ind. Eng. Chem. Res., 2010, 49(3), 1018–1026, DOI:10.1021/ie9007002.
- S. Ramadhani, Q. N. Dao, Y. Imanuel, M. Ridwan, H. Sohn, H. Jeong, K. Kim, C. W. Yoon, K. H. Song and Y. Kim, Advances in Catalytic Hydrogenation of Liquid Organic Hydrogen Carriers (LOHCs) Using High-Purity and Low-Purity Hydrogen, ChemCatChem, 2024, 16(24), e202401278, DOI:10.1002/cctc.202401278.
- P. Lewis, E. Perez, A. Piktus, F. Petroni, V. Karpukhin, N. Goyal, H. Küttler, M. Lewis, W. Yih, T. Rocktäschel, S. Riedel and D. Kiela, Retrieval-Augmented Generation for Knowledge-Intensive NLP Tasks, arXiv, 2021, preprint, arXiv.2005.11401, DOI:10.48550/arXiv.2005.11401.
- Y. Feng, J. Wang, R. He, L. Zhou and Y. A. Li, Retrieval-Augmented Knowledge Mining Method with Deep Thinking LLMs for Biomedical Research and Clinical Support, arXiv, 2025, preprint, arXiv.2503.23029, DOI:10.48550/arXiv.2503.23029.
- D. Zhang, X. Jia, H. B. Tran, S. H. Jang, L. Zhang, R. Sato, Y. Hashimoto, T. Sato, K. Konno, S. Orimo and H. Li, “DIVE” into Hydrogen Storage Materials Discovery with AI Agents, Chem. Sci., 2026, 17(6), 3031–3042, 10.1039/D5SC09921H.
- L. A. Curtiss, P. C. Redfern and K. Raghavachari, Gaussian-4 Theory Using Reduced Order Perturbation Theory, J. Chem. Phys., 2007, 127(12), 124105, DOI:10.1063/1.2770701.
- H. L. Morgan, The Generation of a Unique Machine Description for Chemical Structures-A Technique Developed at Chemical Abstracts Service, J. Chem. Doc., 1965, 5(2), 107–113, DOI:10.1021/c160017a018.
- F. Pedregosa, G. Varoquaux, A. Gramfort, V. Michel, B. Thirion, O. Grisel, M. Blondel, P. Prettenhofer, R. Weiss, V. Dubourg, J. Vanderplas, A. Passos and D. Cournapeau, Scikit-Learn: Machine Learning in Python, 2012 Search PubMed.
- D. Lemm, G. F. von Rudorffvon Lilienfeld, https://www.leruli.com/, accessed 2023-03-13 Search PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery Jr., J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian 16, Gaussian, Inc., Wallingford CT. 2016 Search PubMed.
- A. D. Becke, Density-Functional Thermochemistry. III. The Role of Exact Exchange, J. Chem. Phys., 1993, 98(7), 5648–5652, DOI:10.1063/1.464913.
- M. J. Frisch, J. A. Pople and J. S. Binkley, Self-consistent Molecular Orbital Methods 25. Supplementary Functions for Gaussian Basis Sets, J. Chem. Phys., 1984, 80(7), 3265–3269, DOI:10.1063/1.447079.
- R. Krishnan, J. S. Binkley, R. Seeger and J. A. Pople, Self-consistent Molecular Orbital Methods. XX. A Basis Set for Correlated Wave Functions, J. Chem. Phys., 1980, 72(1), 650–654, DOI:10.1063/1.438955.
- A. D. Becke, Density-Functional Exchange-Energy Approximation with Correct Asymptotic Behavior, Phys. Rev. A, 1988, 38(6), 3098–3100, DOI:10.1103/PhysRevA.38.3098.
- K. E. McCullough, D. S. King, S. P. Chheda, M. S. Ferrandon, T. A. Goetjen, Z. H. Syed, T. R. Graham, N. M. Washton, O. K. Farha, L. Gagliardi and M. Delferro, High-Throughput Experimentation, Theoretical Modeling, and Human Intuition: Lessons Learned in Metal–Organic-Framework-Supported Catalyst Design, ACS Cent. Sci., 2023, 9(2), 266–276, DOI:10.1021/acscentsci.2c01422.
| This journal is © The Royal Society of Chemistry 2026 |