Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
WeChemSynOntology: semantic modeling of wet chemical syntheses in a self-driving lab for nano- and advanced materials
Markus
Schilling
 a,
Harald
Bresch
a,
Harald
Bresch
 b,
Bernd
Bayerlein
b,
Bernd
Bayerlein
 *a and
Bastian
Ruehle
*a and
Bastian
Ruehle
 *b
*b
aFederal Institute for Materials Research and Testing (BAM), Unter den Eichen 87, 12205 Berlin, Germany. E-mail: bernd.bayerlein@bam.de; bastian.ruehle@bam.de
bFederal Institute for Materials Research and Testing (BAM), Richard-Willstaetter-Str. 11, 12489 Berlin, Germany
First published on 20th April 2026
Abstract
Representing experimental procedures in an unambiguous way that can be understood and reproduced by other scientists is at the heart of scientific progress. For centuries, these descriptions were made by humans and for humans, often assuming implicit or tacit knowledge. However, when Materials Acceleration Platforms (MAPs) and Self-Driving Labs (SDLs) are used for the autonomous discovery and optimization of materials, sharing knowledge, and workflows that were designed and executed by machines becomes increasingly important. These machines require an explicit, precise and accurate description and modeling of all process parameters and steps that need to be executed. To address these needs, especially in the domain of materials science and nano and advanced materials synthesis, we developed the Wet Chemical Synthesis Ontology (WCSO), which is based on the Platform MaterialDigital core ontology (PMDco) and the Basic Formal Ontology (BFO). The ontology contains recurring concepts from millions of wet chemical synthesis procedures in the scientific literature. We discuss the design considerations, concepts, and architecture of our ontology in detail, and demonstrate how it can be applied to the construction and querying of semantically annotated knowledge graphs from wet chemical nano- and advanced materials synthesis workflows that were previously designed for and then executed on an SDL. Using such formal representations and semantic annotations for describing synthesis procedures and workflows facilitates the reproducibility, sharing, and execution of synthesis procedures across different labs around the world that use different orchestrators for their robotic hardware.
Introduction
The accurate, precise, and unambiguous description of an experimental procedure is of paramount importance in all experimental sciences. It allows scientists around the world as well as future generations to reproduce and build upon the knowledge and progress that has been achieved in the past. It is hence very concerning that some scientific disciplines report a substantial reproducibility crisis.1In (synthetic) chemistry and materials science, almost all modern journals require an “Experimental section” or “Methods Sections” in which the experiments that lead to the compounds, observations, and conclusions that are discussed in the contribution need to be described in a free text format. This approach has worked for decades (or even centuries) for experimental procedures described by human scientists for human scientists, but with the advent of Self-Driving Labs (SDLs) and Materials Acceleration Platforms (MAPs), as well as AI accelerated materials discovery,2–4 some new challenges with this approach become apparent. These experimental procedures are in general in free text form that do not follow a unified format or have a precise, controlled vocabulary. Together with the inherent entropy of language that allows expressing the same facts or procedures in different ways (or the fact that some concepts might not even have exact correspondences in other languages), this approach makes the readability, interchangeability, and interoperability of experimental procedures more challenging for machines, even when using modern tools for Natural Language Processing (NLP) such as Large Language Models (LLMs).5–10
Moreover, in a human-centric workflow that is typically described in these experimental sections, sometimes imprecise language is used; or at least language that relies on tacit knowledge or concepts that are foreign to a robotic platform or a piece of automated equipment. Examples include describing an addition as “dropwise” or “slow” rather than giving a precise flow rate (e.g., 1 mL min−1), a reaction time as “overnight” rather than giving a precise time (e.g., 16 h), a stirring speed as “vigorous” rather than giving a precise value (e.g., 800 rpm), or referring to a temperature as “room temperature” rather than giving the exact temperature (e.g., 23 °C). While the use of such subjective descriptions might indicate that the respective process parameters were not strictly controlled or optimized during the experiment, a machine still needs to make assumptions regarding their values when building an automated synthesis workflow,8,9 and the exact values that are used during the synthesis might still be important for reproducing the experiment.
Especially in workflows that were carried out – and in some cases even designed – by machines, the representation of the experimental processes and parameters in a more structured, unambiguous, and semantically annotated way can enhance the interoperability and reusability of experimental procedures.11 This approach supports adherence to the FAIR principles12 and advances the idea of democratizing MAPs and accelerated discovery workflows globally. One way to achieve this is by modelling these processes as knowledge graphs that conform to a well-defined underlying ontology.
SDLs increase experimental throughput and enable closed-loop optimization, yet they also amplify long-standing challenges that concern data harmonization across instruments, software stacks, and research groups. In distributed SDL settings, architectures that rely on knowledge transfer, graphs, and ontologies have been proposed because they enable an explicit representation of experimental workflows, material flows, and provenance across sites, even highlighting potential economic benefits.13–15 Such approaches align with the FAIR Guiding Principles, which emphasize that data should be findable, accessible, interoperable, and reusable, particularly in ways that support machine-actionable reuse at scale.12 In many SDL deployments, however, data management practices still evolve in a project-specific manner, which yields heterogeneous definitions, ad-hoc annotation properties, and inconsistent terminology across data exports and documentation. The resulting “vocabulary drift” reduces interoperability and complicates both automated analysis and cross-experiment comparison, which motivates standardization based on semantic technologies as a foundational step toward robust, machine-actionable research data management, and interoperable data sharing.16
A practical response to these challenges typically relies on semantic frameworks that separate domain-neutral concepts from domain-specific content, which supports reuse and promotes semantic interoperability across communities. The ISO/IEC 21838 (ref. 17) family formalizes this idea by defining requirements for top-level ontologies that can function as hub ontologies and can support exchange, retrieval, discovery, integration, and analysis across heterogeneous information systems. Within this landscape, the Basic Formal Ontology (BFO) plays a prominent role because it provides a domain-neutral set of abstract concepts that support suites of interoperable ontologies, and it is standardized as ISO/IEC 21![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 838-2.18 BFO offers a high-level modeling vocabulary that distinguishes, amongst others, continuants from occurrents and supports systematic representations of material entities, processes, and dependent qualities, which also fits laboratory-centric use cases that combine chemistry, materials, process engineering, measurements, and automation (Fig. 1).
838-2.18 BFO offers a high-level modeling vocabulary that distinguishes, amongst others, continuants from occurrents and supports systematic representations of material entities, processes, and dependent qualities, which also fits laboratory-centric use cases that combine chemistry, materials, process engineering, measurements, and automation (Fig. 1).
 | ||
| Fig. 1 Self-driving lab platform for autonomous wet chemical (nano-)material syntheses with depiction of important concepts. | ||
In Materials Science and Engineering (MSE), the PMD core ontology (PMDco), as developed in the frame of the project Platform MaterialDigital (PMD),19 provides a mid-level semantic framework that builds on BFO and reuses established BFO-aligned ontologies, while it also targets MSE-specific needs that concern processes, material structures/states, properties, and performance.20 PMDco is explicitly supposed to be an intermediate layer that connects domain-neutral top-level categories to extendable, application-focused ontologies, which supports interoperability across MSE subdomains and supports consistent modeling of experiments and workflows.21 This design rationale matches the architectural distinction that many ontology engineering communities describe as a separation between top-level ontologies (TLOs), mid-level ontologies (MLOs), and domain/application ontologies that provide increasingly specific terminology for distinct use cases. The PMDco user guidance also recommends that application ontologies describing aspects from the field of MSE are to be built on PMDco and de facto standardized practices advocated by the groundwork laying Open Biological and Biomedical Ontology (OBO) Foundry and the technically implementing Ontology Development Kit (ODK). This supports a workflow that remains compatible with community conventions and reuse of existing ontological assets is encouraged.
This ontology work targets wet chemical syntheses in an SDL context, which requires representations that cover laboratory processes (planning, execution, and reporting), materials and material transformations, instruments and devices, measurands, quantities and units, and domain-specific qualities that describe products and outcomes during chemical syntheses. In addition, the measurands, quantities and units must also conform to the usual worldwide standards, such as the SI system22 and the standards from the ISO expert committees, in the case of the nanomaterials targeted here with ISO TC 229,23 but also with the globally recognized test methods of the OECD and the specifications contained in the test guidelines.24 Such breadth motivates a BFO-aligned approach because BFO is intended to support integration across multiple domains and because it provides stable high-level categories that can host more specific terms from chemistry, materials science, and laboratory automation. It also motivates the selection of PMDco as the mid-level anchor because PMDco already targets the MSE paradigm of processing–structure–properties and because it supports extensions through application ontologies that remain interoperable with a broader MSE semantic framework.21,25–27 In parallel, SDL-oriented work has demonstrated that ontologies can represent design–make–test–analyze cycles and can record provenance in a way that supports FAIR goals, which provides an external validation of ontology-centric design for automated experimentation pipelines.28,29
Recent work has further underscored the relevance of semantic modelling and knowledge graphs as enabling infrastructure for laboratory automation and SDLs. In particular, a recent perspective on data integration and fusion in SDLs highlights ontology-driven frameworks and knowledge graphs as key mechanisms for harmonizing heterogeneous experimental, computational, and literature-derived data and for making such data machine-actionable at scale.30 In a complementary direction, connected digital-twin visions argue for comprehensive, distributed laboratory digital twins grounded in dynamic knowledge graphs and a universal knowledge model that supports orchestration, interoperability, and reasoning beyond platform-specific “island solutions”.31 At the interface of synthesis reporting and machine-readable representations, LLM-assisted pipelines have been demonstrated that extract synthesis procedures from unstructured literature and integrate them into larger knowledge ecosystems via dedicated synthesis ontologies and semantic agents.32–34 Finally, BFO-aligned efforts have been reported that construct knowledge graphs from electronic lab notebook (ELN) environments by harvesting metadata via APIs and transforming them into BFO-compliant RDF graphs using SPARQL-based transformation pipelines.35,36
In this landscape, WCSO is positioned as a lightweight, BFO/PMDco-anchored application ontology that focuses on the procedural, recipe-level representation of wet-chemical synthesis workflows as executed in SDL environments. Compared to broader digital-twin or ELN-centric knowledge graph efforts, the primary emphasis of WCSO lies in modelling temporally orchestrated action sequences (including concurrency and ordered subprocess structure) and in capturing process parameters through reusable process-attribute patterns. This design directly supports cross-workflow comparison and retrieval of “recipe fragments” across heterogeneous protocols and automation stacks while remaining interoperable with the broader BFO-aligned ecosystem.
Application and domain ontology design typically rely on (selective) reuse of established ontologies and concepts that provide mature vocabularies for methods, investigations, process chemistry, qualities, units, and laboratory-centric representations. Some of these usable resources relevant in the regarded field are summarized in Table S.1 (see SI).
To facilitate the discovery and verification of candidate vocabularies and ontologies for semantic modeling, a number of freely accessible online tools provide curated collections, term-level browsing, and programmatic access, such as, e.g.: (i) Ontobee which is a linked-data server and browser that dereferences ontology term URIs and exposes them both as human-readable HTML pages and machine-readable RDF, facilitating term inspection and navigation across many ontologies.37 (ii) The TIB Terminology Service offers a single point of access to scientific and technical terminologies with web-based browsing and a REST API (Representational State Transfer Application Programming Interface) to retrieve term information (e.g., identifiers, definitions, relations) for integration into tools and workflows.38 (iii) MatPortal is an ontology repository dedicated to materials science that supports publishing, searching, and comparing relevant ontologies, and provides additional portal functions such as annotation and mappings.39
Moreover, a common ontology can help making workflows interoperable between SDLs. Over the past years, numerous orchestrating frameworks for SDLs have been developed to fit the specific needs of the individual platforms they were built for, for example MinervaOS,4 IvoryOS,40 ChemOS,41 AlabOS,42 FINALES,43 NIMS-OS,44 HELAO,45 XDL,14 MADSci,46 and numerous others. While there are clearly parallels and similarities in their architectures and the underlying concepts, there is no universally adopted naming convention. Hence, the functions that are called for executing the exact same experimental steps on the individual hardware (e.g., heating, stirring, adding) and their corresponding process parameters (e.g., temperature, stirring speed, flow rate) can have different names in the different orchestrators, even though they semantically refer to the same concepts or operations. Having to analyze the architecture and naming conventions of the orchestrators that ran the experimental procedures or workflows for a specific synthesis every time an experiment from a different lab is adapted so that they can be mapped to one's own orchestrator poses an unnecessary burden on sharing, comparing, and expanding existing experimental knowledge. Abstracting the concepts and operations away from the underlying hardware and software that orchestrates the execution and mapping them to a semantically well annotated ontology on the other hand can accelerate this process, make it significantly less tedious and error prone, and enable true and easy interoperability between labs running different SDL/MAP platforms with different hardware and software backends. While such a semantic layer can substantially reduce technical barriers, interoperability at scale ultimately remains dependent on community uptake, mappings to existing platform conventions, and continued alignment across stakeholders.
If existing recipes and (meta)data, such as information on chemicals, synthesis routes, and parameters, are systematically structured and integrated according to ontological concepts and subsequently extended through automated and dedicated data integration workflows, the resulting semantic representations enable flexible and machine-readable access to experimental knowledge. This provides a foundation for data-driven discovery approaches. As the knowledge base grows, such frameworks support the identification of optimal batch parameters with respect to economic efficiency and sustainability, as well as the exploration of previously unexplored parameter regimes and design spaces. Furthermore, the approach presented here, in which natural language synthesis descriptions and an easy-to-use graphical user interface are used for constructing workflows and ontology-aligned knowledge graphs can also help with community uptake by reducing the complexity of using the correct ontological concepts for mainly synthetically trained chemists and material scientists.
Using unambiguous ontology-based (meta)data descriptions, the resulting gain in interoperability enables hardware-independent synthesis workflows and supports the international reproduction of (reference) materials on demand.
Despite these advantages, the adoption of semantic technologies in materials science and laboratory automation remains limited due to steep learning curves and the need for specialized expertise.47 Recent developments have nevertheless demonstrated the long-term value of mature and community driven ontologies. The recognition of the AlphaFold developers indirectly highlighted how BFO aligned resources such as the Gene Ontology and ChEBI enable systematic annotation of structural data and binding sites and support the semantic linking of instances across large datasets.48–50
Motivated by these advances, we present an SDL use case that demonstrates the benefits of semantic interoperability enabled by ontologies and knowledge graphs in a real laboratory context. By employing established tools and methods, our approach aims to support the convergence of technology, expertise, and community acceptance. The BFO and PMDco compliant ontology patterns developed in this work provide a transferable foundation for further SDL and MAP applications. Furthermore, the presented ontology “WeChemSyn-Ontology (WCSO)” supports a federated and collaborative approach in which both the ontology itself and the associated nanomaterial synthesis knowledge base can be continuously extended and refined by the community.
Results and discussion
We recently presented an approach for using natural language processing to automate the generation of workflows and knowledge graphs8 in our SDL.4 In this approach, a rule-based algorithm5 and LLMs51 were used to process experimental descriptions and create “action graphs” that represent – in a standardized way – the individual steps that were described in the synthetic procedures in natural language. For describing the actions, we used the 21 “action tags” that were originally suggested by Hawizy et al. for their work on ChemicalTagger5 (and that were the result of a literature survey with human domain experts) and extended them with 5 additional tags. The resulting set of 26 “action tags” could be used to describe the experimental procedures that were extracted from over 1.5 million patents. They are also very similar to the 28 action tags that were used by Vaucher et al. in their work on automated extraction of chemical synthesis actions from experimental procedures9 for describing 6.2 million experimental descriptions from the Pistachio dataset, as well as 1764 hand annotated actions from the chemical literature. This illustrates the universality of these actions across several millions of experimental procedures and demonstrates the feasibility of describing the vast space of wet chemical syntheses with only around two dozen individual actions.To operationalize the above action vocabulary for nanomaterials synthesis, we developed the WeChemSyn Ontology (WCSO) as a lightweight, use-case-driven application ontology that provides a semantic framework for representing nanomaterial production and characterization workflows from a chemical perspective. In contrast to broad, domain-spanning nanomaterial ontologies, our primary design target was the procedural description of wet chemical syntheses as they are executed in a SDL: i.e., representing “recipes” as machine-actionable, modular process descriptions that can be transferred across experimental setups and executed (or at least interpreted) by heterogeneous automation systems. This emphasis directly addresses a key bottleneck in the nanomaterial community: while the scientific literature offers enormous “big-data” coverage of synthesis procedures, the operational knowledge is often embedded in narrative descriptions, requiring tacit expertise and manual interpretation. This represents a kind of “recipe detour” that hinders reproducibility, automation, and sustainable-by-design decision making across materials and processes.
WCSO is explicitly grounded in the PMD core ontology (PMDco), a mid-level ontology for Materials Science and Engineering (MSE) designed to bridge application-specific models to domain-neutral top-level categories. PMDco follows the canonical MSE paradigm (processing–structure–properties) and reuses established BFO-aligned resources such as RO, IAO, and OBI to represent processes, materials, devices, roles, functions, and information artifacts in a modular and interoperable manner.
By importing and extending PMDco, WCSO inherits (i) a well-tested semantic backbone for representing experimental process chains and their participants, and (ii) an interoperability strategy aligned with BFO that enables in particular integration with other BFO-conformant ontologies and knowledge graphs in the broader MSE-related semantic ecosystem. This anchoring is especially beneficial in the SDL setting, in which synthesis, characterization, and data-driven optimization need to be connected seamlessly. PMDco provides stable modeling primitives (e.g., planned process, plan specification, objective specification, material entities, devices) that allow for the representation of nanomaterial synthesis steps consistently alongside subsequent analytics and metadata.
Building on the action tags introduced above, WCSO extends PMDco with a controlled vocabulary of chemistry-centric laboratory actions, including terms such as, e.g., add, mix, stir, heat, cool, quench, centrifuge, wash, filter, precipitate, extract, purify, recover, and remove, each modeled as a process type with domain-appropriate definitions and usage examples, as well as accompanied by consistent labelling. This coverage is crucial for nanomaterial syntheses, in which the “same” target material can be obtained by multiple procedural variants, and where the ability to modularize and recombine action sequences is a practical prerequisite for automation, comparison, and optimization at scale. In addition, WCSO introduces or refines domain terms relevant for synthesis outcomes and characterization descriptors (e.g., yield and yield variants, concentration, dispersive system) to enable the representation of both what is done and what is obtained/observed as structured knowledge that may be easily queried.
The level of granularity of the 26 action classes was selected to balance (i) interoperability and broad retrieval across heterogeneous protocols with (ii) method-specific distinctions that matter for reproducibility, automation constraints, and outcome interpretation. Actions that may appear similar in everyday laboratory language are intentionally kept separate when they differ in their underlying physical mechanism and typical parameterization, because these differences affect both the feasible device realizations and the expected effects on materials and mixtures. For example, stirring primarily induces macroscopic convective mixing through mechanical agitation (typically parameterized by rotational speed and geometry), whereas sonication introduces acoustic energy that can drive dispersion, deagglomeration, and localized heating (typically parameterized by power/amplitude, duty cycle, and sonication time). Similarly, filtration and centrifugation are both separation steps, yet they rely on different separation principles and operational constraints (e.g., filter medium/porosity and pressure or vacuum versus centrifugal force and rotor settings), which leads to different measurable process attributes and often to different process outcomes and failure modes.
At the same time, the action vocabulary is intended to support queries at multiple abstraction levels. Where method-specific distinctions are not required, broader retrieval and cross-protocol comparison can be performed by targeting higher-level groupings (e.g., “agitation/mixing” or “separation/purification”) and retrieving all corresponding subclasses. When detailed comparison is desired, the more specific action classes enable stratified analyses (e.g., separating “sonicate” from “stir” or “centrifuge” from “filter”) to examine how procedural variants correlate with outcomes such as dispersion state, yield, or downstream characterization results. This multi-level querying capability supports both general competency questions (e.g., “Which protocols use any separation step?” (see also Methods)) and controlled cross-protocol comparisons (e.g., “How do protocols differ when a separation step is performed by filtration versus centrifugation?”).
As essential for the creation of fully-fledged semantic representations cast in an ontology, a central design goal of WCSO was to move beyond a purely taxonomic “term list” and provide axioms that support consistent instantiation, validation, and inference. Such axioms formally constrain class membership and property usage and thereby enable automated reasoning and consistency checking. Specifically, WCSO leverages BFO semantic patterns around planned processes, plan specifications, and objective specifications to represent procedural actions as realizations of explicit recipe intentions. This is essential for SDL contexts because it preserves the provenance of “what was intended” versus “what was executed”, allowing workflows to diverge adaptively (e.g., substitutions or adjustments during execution) while still remaining semantically traceable as instances of a planned process whose realized plan encompasses relevant modifications. As an example, the class heat is modeled as a defined class in the ontology using three equivalence axioms that jointly characterize when a process instance qualifies as being a heat process (Fig. 2).
 | ||
| Fig. 2 Equivalent classes axioms (rendered as labels only) for process class “heat”, given in description logic (DL) notation. | ||
The first axiom constrains heating to processes that realize (object property) at least one relevant function being subclass to realizable entity, either a heating function or a heat treatment function, thereby capturing the functional/teleological aspect of applying heat in a laboratory or processing context. The second axiom requires that a heating process has participant (object property) some temperature, ensuring that the process is explicitly linked to the thermodynamic quantity that is being manipulated or tracked. Finally, the third axiom specifies an additional participation constraint: the process must involve at least one participant that is either a heated magnetic stirrer, a heat treatment device, or a more general temperature change device. This disjunctive modeling (union) accommodates multiple realizations of heating, ranging from combined stir-and-heat lab equipment to generic devices designed to induce temperature change, while keeping them under the same formally defined process type.
To maintain modeling consistency while scaling to dozens of actions and many mapped procedures, we relied on semantic (ontology design) patterns. This approach aligns with PMDco's development strategy, which explicitly supports pattern libraries and reusable modeling templates to ensure consistent axiomatization across modules and contributors. For WCSO, patterns play two crucial roles: (i) they provide repeatable “blueprints” for representing common synthesis structures (e.g., action with specified inputs/outputs; action executed by device; action achieving objective), and (ii) they keep the ontology maintainable as new nanomaterial systems, action variants, or characterization steps are added. Furthermore, the visual representation of patterns provides an advantageous documentation for easy perceptible human understanding of the specific semantic modeling regarded.
A practical modeling challenge in laboratory workflows is to connect process-level descriptions (e.g., “anneal at 1000 °C”) to qualities that inhere in physical participants (e.g., the temperature of a sample or an oven). In BFO-aligned modeling, many such qualities are specifically dependent continuants (SDCs) and must inhere in an independent continuant (IC) rather than in a process. Moreover, widely reused relations such as has characteristic are functional, i.e., a specific SDC instance can only be the characteristic of one bearer; attaching the same SDC instance to both a process and a participant is therefore invalid according to the first-order modeled logic. WCSO addresses this by extensively making use of the process attribute pattern: processes have process attributes (rates, setpoints, etc.), and these attributes can refer to SDCs of process participants, thereby linking process conditions to their appropriate bearers without violating functional constraints (Fig. 3). In addition, WCSO distinguishes relational qualities (qualities inhering in two or more bearers) via a dedicated non-functional relation (relational quality of), avoiding incorrect inferences that would follow from using functional “quality of” relations in cases like proportions or shared relational measures (Fig. 4). For our use case, this choice is beneficial because synthesis protocols frequently express relational and contextual quantities (e.g., ratios, concentrations, rates) whose correct interpretation is essential for reproducibility and for transferring recipes between different laboratory configurations.
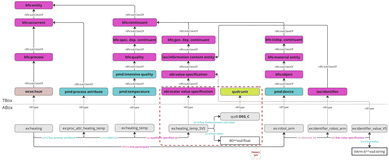
Fig. 3 shows a semantic pattern that visualizes how the ontology (general semantic representation) and an assertion graph (knowledge graph) work together to represent a heating process, an associated process attribute (heating temperature), and the respective measurement/value encoding (numeric value and measurement unit). Moreover, an example of an identification for a device is given. Hence, there are two layers represented: the terminological box (“TBox”) denoting the schema that consists of classes and their subclass hierarchy (upper half, pink/teal/yellow boxes) as modeled in the ontology and the assertional box (“ABox”) referring to the instance data, which are individuals (example instances) and the relations between them (lower half, grey boxes). In the pattern, a particular heating process is instantiated as an instance of the class wcso:heat and linked to an initially indefinite process attribute. Here, the process attribute “heating temperature” was selected as an example; to further characterize the heating process, several other parameters are specified as process attributes in a similar form, with the heating temperature being given as a representative. The process attribute (which is process-dependent) is connected to the relevant quality-type (pmd:temperature) indirectly by using the object property pmd:refers to, rather than pretending the process bears the temperature quality. Hence, the temperature quality is still conceptually a quality of some bearer (often a participant like the sample or the environment), but the process attribute can “point at” the relevant quality dimension using pmd:refers to. This is the intended approach of the modeled representation in the ontology for linking process attributes to a process or process step, which is highly relevant for modeling recipes in wet chemical syntheses.
Furthermore, the scalar value specification pattern (SVS) is introduced, which is intended to be reused whenever a (measured) value and a measurement unit need to be specified and linked, representing a building block or puzzle piece. This is indicated (and visually encapsulated) by the red dashed box. Hence, the diagram shows a small relation involving iao:quality is specified as between the temperature quality instance and the SVS (ex:heating_temp is linked to ex:heating_temp_SVS using iao:quality is specified as). The intention of the pattern is that the temperature quality is modeled as being associated with a scalar value specification that encodes its measured value (80) and its measurement unit (°C).
Moreover, a similar pattern is used to demonstrate how a device gets an identifier: The robot arm (example device) is denoted by (iao:denoted by) an identifier, and that identifier has a value specification whose literal value is ‘XArm-6’. This mirrors the SVS approach, but here, the value is a string identifier rather than a numeric measurement.
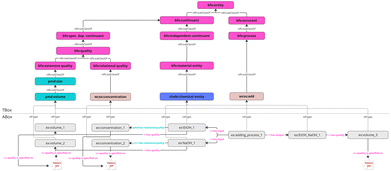
Fig. 4 illustrates an extension of the semantic pattern to the example case of an adding process and demonstrates how process inputs/outputs and composition-like descriptors can be modeled in a BFO-aligned knowledge graph. In the terminological part, the pattern positions wcso:add as a subclass of bfo:process and chebi:chemical entity as a subclass of bfo:material entity, while following the logical hierarchy of BFO, PMDco and WCSO. On the quality side, the diagram explicitly differentiates between extensive qualities and relational qualities: pmd:volume (subclass of pmd:size) is modeled as an extensive quality (i.e., scaling with system size), whereas wcso:concentration is subclass of bfo:relational quality which reflects that concentrations inherently encode a relationship between at least two relata (e.g., part and whole/total). In the instance layer, the central individual ex:adding_process_1 is typed as an instance of wcso:add and is linked to two input participants – ex:EtOH_1 and ex:NaOH_1 – via the ro:has input object property, expressing the intended reading: there is an adding process that has EtOH_1 and NaOH_1 as inputs. Correspondingly, where feasible, the process can be linked by ro:has output to an output chemical entity ex:EtOH_NaOH_1 which captures the transformation result: the adding process produces an output chemical entity. This mirrors the process-centric modeling principle illustrated in Fig. 3, but now, material transformation via input/output is emphasized rather than process attributes such as heating temperature. The pattern then annotates both the inputs and the output with quantities that are relevant for compositional interpretation. On the output side, ex:EtOH_NaOH_1 is connected via ro:has quality to a volume instance ex:volume_3 (typed as pmd:volume), indicating that the resulting entity is associated with a measurable extensive quality. On the input side, the diagram shows volume instances as well (ex:volume_1, ex:volume_2), each typed as pmd:volume, while making use of the SVS pattern introduced beforehand. This conveys the modeling intention that the input chemical entities have their own volumes (measured/recorded) which are treated as the inputs' volume qualities.
In addition to absolute volumes, the figure introduces concentrations as relational qualities borne by the input entities. Specifically, the individuals ex:concentration_1 and ex:concentration_2 are typed as instances of wcso:concentration and are connected to ex:EtOH_1 and ex:NaOH_1 via pmd:has relational quality. This choice aligns with the PMDco modeling rationale that relational qualities (such as concentrations or proportions) require a dedicated relation distinct from the standard “has quality” pattern, because they are not naturally constrained to a single bearer in the same way as ordinary qualities. Semantically, these edges are meant to be read as: each component is associated with a relational quality capturing its concentration as a solute relative to a relevant carrier/solution (i.e., the total solution). Thus, besides the intended general usage of inputs and outputs, the figure demonstrates how to represent both (i) absolute extensive measures (volumes) and (ii) relative, part-whole descriptors (concentrations) within a coherent ontological framework. Finally, the small “SVS” puzzle-piece annotations mark the reuse of the SVS pattern (see Fig. 3) to create a structured value representation.
While the example in Fig. 4 highlights a process-centric input/output view to make material transformation explicit, this should be understood as an illustrative modeling option rather than a mandatory requirement for every (sub-)step in a workflow. In practice, experimental protocols often comprise many micro-actions whose primary role is temporal orchestration (e.g., priming/purging, waiting, cleaning) or container handling, without a need to introduce an explicitly named intermediate (virtual) “chemical entity” after each action. Accordingly, subprocesses may be connected simply via temporal relations or via their part-whole structure, including cases of concurrency, without asserting a fully instantiated chain of intermediate material outputs. Intermediate entities should be introduced when they are semantically or analytically relevant (e.g., when a distinct fraction is selected, when multiple outputs matter, or when an intermediate is measured/characterized and thus warrants explicit identity and attributes). The overall process can still consistently specify its main inputs and outputs at the level of scientific interest, even if many internal steps do not “pass on” explicitly instantiated inputs and outputs.
The orchestration patterns discussed can be situated within a broader landscape of temporal modelling strategies in ontology engineering. In addition to BFO-aligned approaches that distinguish continuants from occurrents and represent temporal coordination through process part structure and temporal relations, prominent four-dimensional (4D) approaches exist in which entities are treated as spatiotemporal extents and change is captured via temporal parts. A well-known example is ISO 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 926,52 whose life-cycle integration ontology was developed for industrial asset and process-plant information integration and explicitly supports temporal part modelling as a 4D mechanism. In the present work, the key requirement concerns recipe-level coordination of laboratory actions (e.g., sequential ordering, concurrency, waiting, and device-mediated handling), rather than long-horizon industrial asset life-cycle integration, which are fully representable within the BFO/PMDco framework through occurrent decomposition (e.g., has temporal part, has occurrent part) and temporal relations (e.g., precedes, simultaneous with), while retaining compatibility with reused BFO-aligned resources (e.g., RO/IAO and other OBO-ecosystem ontologies) that support modular integration of provenance and information-artifact representations.
926,52 whose life-cycle integration ontology was developed for industrial asset and process-plant information integration and explicitly supports temporal part modelling as a 4D mechanism. In the present work, the key requirement concerns recipe-level coordination of laboratory actions (e.g., sequential ordering, concurrency, waiting, and device-mediated handling), rather than long-horizon industrial asset life-cycle integration, which are fully representable within the BFO/PMDco framework through occurrent decomposition (e.g., has temporal part, has occurrent part) and temporal relations (e.g., precedes, simultaneous with), while retaining compatibility with reused BFO-aligned resources (e.g., RO/IAO and other OBO-ecosystem ontologies) that support modular integration of provenance and information-artifact representations.
To further illustrate the relevant modeling of process steps, the following patterns show processes in relation to their subprocesses. The first demonstrates how a compound experimental step can be represented as a single process that is structured into coordinated temporal parts, each of which is typed by a different process kind and parameterized by its own process attributes (Fig. 5). Concretely, the individual ex:infuse_while_heating is modeled as a process that has temporal parts corresponding to (i) a heating subprocess (ex:infuse_while_heating_heating_part, typed as wcso:heat), (ii) a stirring subprocess (ex:infuse_while_heating_stirring_part, typed as wcso:stir), and (iii) an adding/infusion subprocess (ex:infuse_while_heating_adding_part, typed as wcso:add). It is made explicit that these subprocesses are not merely sequential “steps” but are intended to be co-occurring: the heating part is linked to the stirring part (and likewise the relevant parts are linked among each other) via ro:simultaneous with. This expresses that the overall procedure is executed while heating and stirring are happening concurrently with the infusion/addition action. In parallel to the earlier process-attribute pattern (Fig. 3), each temporal part is then equipped with one characteristic process attribute as an example, respectively, via pmd:has process attribute: the heating part is associated with a time-related attribute (ex:heating_time), the stirring part with a rotational attribute (ex:stirring_speed), and the adding part with an addition-rate attribute (ex:addition_rate). The further modeling of process attributes is indicated exemplarily for wcso:addition rate which is a subclass of pmd:process attribute. Moreover, the familiar SVS pattern is reused to encode the concrete parameter values for each attribute to ensure that the numeric value-and-unit representation remains uniform across different subprocess parameters. Overall, in this manner, a higher-level laboratory instruction such as “infuse while heating and stirring” can be represented as a single process with simultaneous temporal components, each carrying its own typed process attributes (duration, stirring speed, addition rate) expressed through the same value-specification mechanism.
Building directly thereon, Fig. 6 illustrates the complementary case in which a compound experimental step is structured into ordered subprocesses that follow one another. Here, the individual ex:redisperse is modeled as an instance of a generic bfo:process that is explicitly composed of three occurrent parts: ex:redisperse_removing_part (typed as wcso:remove), ex:redisperse_adding_part (typed as wcso:add), and ex:redisperse_dissolving_part (typed as wcso:dissolve). In contrast to the previous “infuse while heating” pattern, the critical intention is to capture a sequential workflow rather than concurrency: the removing part is asserted to precede the adding part (bfo:precedes), and the adding part in turn precedes the dissolving part. Thereby, an ordered protocol fragment of the form remove → add → dissolve is represented. The overall process inherits its internal structure from these occurrent parts via repeated use of bfo:has occurrent part, which allows the higher-level “redisperse” instruction to be queried either as a single step or unpacked into its constituent operations. Taken together, these two patterns provide a coherent modeling repertoire for experimental procedures: complex steps can be represented either as (i) simultaneous subprocess bundles connected by ro:simultaneous with and bfo:has temporal part, or as (ii) sequential subprocess chains connected by bfo:precedes and bfo:has occurrent part, depending on whether the protocol semantics require concurrency or explicit temporal ordering.
 | ||
| Fig. 6 Schematic semantic pattern illustrating a “redisperse” process that consists of three parts, each modeled as separate processes that run sequentially (“preceding” each other). | ||
The ontology was designed to support the downstream transformation of extracted “action graphs” into an executable, queryable knowledge graph representation. In practice, this means that each step in a procedure can be instantiated as a planned process (or a specialization thereof), connected to its specified inputs/outputs and, if applicable, to devices, process attributes, and objectives. The resulting instance graph enables competency-question-driven retrieval, for example: (i) identification of procedures that employ a given purification strategy (e.g., centrifugation + washing cycles), (ii) extraction of parameterized “recipe fragments” (e.g., quench conditions, heating/cooling rates, or solvent removal steps), and (iii) aggregation of outcomes such as yields and concentrations across batches or nanomaterial systems. Such querying is a core benefit for SDL operation because it enables automated comparison between runs, supports systematic exploration of alternative synthesis pathways, and facilitates the reuse of successful “motifs” of actions as templates for future campaigns. The combination of (i) an action-centric vocabulary, (ii) PMDco-anchored mid-level semantics, and (iii) pattern-based axiomatization yields several practical benefits relevant in the regarded context. First, interoperability is improved: by aligning to PMDco and BFO, the modeled procedures can be integrated with other MSE knowledge graphs and tooling ecosystems that adopt the same foundational commitments which lowers integration costs and increases community uptake. Second, reproducibility across setups is enhanced: instead of sharing recipes as prose, WCSO supports a standardized representation of procedural intent, participants, and conditions; this makes “what to do” explicit for machines and humans, and reduces ambiguity when migrating protocols between instruments, labs, or automation tech stacks. Third, the aspect of data becoming AI-ready is supported: structured, formally constrained, and semantically annotated representations reduce noise and ambiguity for downstream learning systems that may include both machine learning (ML)- and large language model (LLM)-based components which benefit from consistent, well-typed data and context as well as from robust linking between actions, parameters, and outcomes. This is particularly relevant for future “neuro-symbolic” approaches, where LLMs generate or adapt candidate procedures while symbolic constraints and pattern-grounded knowledge graphs provide verification, normalization, and explainability.
Methods
The ontology development and maintenance process combined (i) a domain-expert-friendly elicitation phase that relied on spreadsheets and laboratory workflow descriptions with (ii) a standards-oriented engineering phase that relied on Protégé for OWL authoring and on ODK-driven repository automation for builds, imports, quality control, and releases. The integration strategy followed the MaterialDigital approach that couples PMDco-based modeling with ODK-based engineering conventions, which ensured that WCSO evolved as a domain ontology that remained interoperable with a broader materials-science semantic framework. GitHub Actions orchestrate the execution of the toolchain so that each revision undergoes automated checks and produces updated artifacts and documentation that remains aligned with the repository state. Particularly, the automation implemented by using GitHub Actions workflows shortens the feedback loop that domain experts face because each merged update of the ontology triggered documentation updates that reviewers could inspect in a browser without local tooling. The same automation strategy supports lightweight curation because it reduces the manual steps that otherwise accompany ontology publishing, a concern that is identified as a recurring barrier that limits broad community participation.Competency questions, ontology development workflow and integration
The development of the ontology was guided by representative competency questions reflecting typical information needs in SDL-based wet-chemical synthesis, including:(i) Which entities (e.g., materials, instruments, processes, process steps) are involved in a wet-chemical synthesis conducted in a SDL?
(ii) How can wet-chemical synthesis recipes be represented in a structured, machine-interpretable form that allows for interoperable querying and reuse across experiments and systems?
(iii) What precursor materials, solvents, and processing conditions were used to produce a specific intermediate/material?
(iv) What instruments have been utilized across workflows for specific operations?
(v) How are process steps temporally orchestrated (sequential vs. concurrent), and which process attributes (e.g., setpoints, rates, durations) are associated with those steps?
(vi) Which parameterized “recipe fragments” recur across workflows (e.g., addition while heating followed by a separation step) and how do they correlate with recorded outcomes (e.g., yield variants or downstream characterization descriptors)?
The ontology development process followed an established roadmap for domain ontology engineering that has been described in prior work, which outlines a progression from terminology collection and conceptual structuring to a formal OWL/RDF representation.27 Rather than explicitly reproducing that roadmap, the present work aligned with its overall intent while it adapted specific steps to the practical constraints of a rapidly evolving laboratory setting. In particular, the workflow combined (i) a spreadsheet-centered phase that supported rapid review by domain experts and (ii) an OWL-centered phase that supported logical validation, modularization, and release automation.
Knowledge acquisition, term curation, and conceptual structuring
Domain knowledge collection relied on two primary sources: process workflows that originated from the SDL environment and built upon previous collections of process steps from domain experts that proved to be applicable to millions of example reactions from the literature,5,8,9 and a structured terminology inventory that was maintained in a spreadsheet tool. The spreadsheet format supported early-stage consensus building because it exposed candidate entities, process parameters, and measurement-related descriptors in a form that domain experts could inspect without ontology tooling. This stage corresponded to the “information gathering” step of the referenced roadmap, although the present work emphasized iterative refinement that reflected updates in laboratory workflows and reporting practices. From these sources, the development team derived an initial set of classes and properties and assigned them to categories that reflected process phases, material-related entities, and measured qualities. This categorization established the taxonomic backbone that later guided module boundaries and import decisions within the OWL implementation. Conceptual modeling and formal editing occurred in the established Protégé tool,53,54 which served as the primary ontology editor for class hierarchies, property axioms, and annotation management. Protégé provides an OWL-focused environment that supports navigation across imported ontologies and direct inspection of entity annotations, which facilitates consistent labeling and definition practices across the growing term set. The workflow maintained a strict separation between human-oriented term elicitation (which remained spreadsheet-based during early iterations) and machine-oriented axiomatization (which occurred in the editable OWL source). This separation reduced friction during review cycles because spreadsheet revisions did not require OWL expertise, while OWL revisions preserved semantic precision that required downstream tooling.Repository-centered lifecycle management with ODK and PMDco alignment
For publication and long-term curation, GitHub was adopted as the primary platform for version control, issue tracking, and review that it comes with inherently. This choice reflected the desire for a development process that supported frequent, minor updates and that enabled traceability across ontology versions. The repository structure and automation strategy followed the Ontology Development Kit (ODK),55 which provides standardized and configurable workflows for ontology lifecycle tasks that include continuous integration, quality control, release preparation, and import management. ODK also provides a curated toolchain as a Docker image, which reduces local setup differences across contributors and which supports reproducible builds that remain stable across operating systems.The specific repository template that served as a starting point originated from the Platform MaterialDigital (PMD) initiative,21 which offers an ODK-based “application ontology template”20 that has been configured for domain and application ontologies to build upon the PMD core ontology (PMDco). That template includes predefined GitHub workflows and an approach that places the editable ontology file under src/ontology/*-edit.owl, which is consistent with ODK conventions, and which supports collaborative editing through pull requests. This template was adopted as a foundation and extended substantially to match WCSO-specific scope, imports, and release outputs. The alignment with PMDco ensured that domain-level terms remained interoperable with a mid-level semantic framework that has been designed for materials science and engineering (MSE) and that itself relies on ODK-based modular development practices (cf. Introduction).
As with most applied ontologies, the WCSO TBox is expected to evolve over time as laboratory practice, terminology, and modelling requirements change. To support reproducible reuse, changes are tracked through version control and release artifacts, and backward-compatible evolution is preferred where possible (e.g., stable IRIs and deprecation rather than renaming). When TBox changes affect instance data, ABox updates are treated as a controlled migration problem: the workflow-to-RDF export step and repository automation provide a natural integration point for SPARQL-based transformation or regeneration of ABox datasets so that released knowledge graph snapshots remain aligned with a given ontology version.
Automated builds, quality control, and release artifacts via GitHub actions
Automation of routine curation tasks relies on GitHub Actions, which provides repository-integrated workflow execution for continuous integration and deployment tasks. For this, multiple workflows are maintained that address distinct lifecycle stages. One workflow focuses on formal quality checks that detect syntactic and structural issues in the ontology before changes enter a release branch. Another workflow refreshes ontology imports to ensure that external dependencies remain current, which supports consistent reasoning behavior and prevents silent drift that can occur when imported artifacts become outdated. These workflows reflect ODK's emphasis on continuous quality control and dependency management, which the ODK literature highlights as recurring sources of complexity in ontology lifecycle management.55Release generation and file-format production relies on the ROBOT toolchain, which has been developed within the OBO community56 as a command-line system that automates common ontology tasks such as merging, reasoning, querying, validation, and format conversion.57 ROBOT provides explicit commands for converting an OWL edit file into multiple distribution formats, which supports downstream consumers that expect OWL, RDF serializations, or OBO-style artifacts. Within the ODK context, ROBOT acts as a core build engine that the ODK workflows call when they materialize release products and apply standardized checks, typically based on simple SPARQL querying, e.g., for the detection of missing semantic concept annotations that could be violating best practices. In this project, that arrangement allows ontology editors to commit changes to the editable source while automated workflows produce a consistent set of release artifacts that remain synchronized with the repository state. In addition to SPARQL-based quality checks, constraint-based validation could also be expressed using SHACL (SHApes Constraint Language) shapes, which are well suited for standardized validation of RDF instance graphs and for pre-execution checks of workflow graphs. In the present version, the SPARQL-based approach was retained because it integrates seamlessly with the established ODK/ROBOT workflow, while SHACL-based validation is considered a promising complementary option.
Documentation
Furthermore, the workflow treats documentation as a release artifact that requires the same level of automation as ontology files. For that purpose, WIDOCO tooling is used, which generates enriched technical HTML documentation for ontologies, and which supports command-line execution via a downloadable JAR. WIDOCO supports documentation generation that relies on ontology metadata and term annotations, which encourages documentation that remains consistent with the ontology itself, also if versions may change. In practice, GitHub Actions execute documentation workflows after ontology updates, which produces a static website that exposes both the ontology overview and the term-level cross-reference pages. On top of that, another documentation supposed to be easier digestible by human readers is automatically created by a workflow making use of the Mkdocs tool58 that is based on web pages specifically designed by the authors in markdown format. Both the technical WIDOCO and Mkdocs based documentations are independently accessible and dereferenceable.Adding terms and definitions from ISO and other standards
Some terms used in the description of the synthesis and several domain-specific terms are already defined in ISO standards. This is especially the case for overarching terms. Ideally, such established terminology could be reflected in ontologies to support alignment across communities. In practice, however, the reuse of standard text (including formal definitions) is often subject to licensing and copyright constraints imposed by standardization bodies. For this reason, the ontology presented here does not directly reproduce ISO definitions verbatim. Where appropriate, a small number of definitions were instead formulated in close accordance with ISO-based descriptions to avoid unnecessary divergence while still respecting these constraints. More broadly, this highlights a recurring challenge at the interface of standards and ontologies: both aim to promote shared and reliable terminology, yet legal and licensing conditions may limit direct textual reuse. Consequently, ontology developers may need to provide paraphrases or newly authored definitions for concepts that are already standardized. This is not ideal from an interoperability perspective, as it may increase terminological heterogeneity across resources. Endeavoring clarity and improved pathways for standards-aware reuse (e.g., via licensing, or standardized mechanisms for definition referencing) would help communities to converge on shared terminology and be beneficial for long-term interoperability.Knowledge graph generation
There are several excellent, open and commercial tools that can be used for generating knowledge graphs. Here, we implemented a simple exporter for our node-based workflow editor that allows exporting the entire workflow including all its process parameters as a knowledge graph that follows the WCSO in the turtle file format. For each “high level” node in the node graph that represents the entire workflow, the script looks up the corresponding concepts and their IRIs from the WCSO and automatically applies them, modelling compound nodes that comprise several simultaneous or sequential steps (e.g., “Infuse while Heating”, “Centrifuge and Redisperse”) as described in the paper. The process parameters for each high-level node are also extracted and linked to their corresponding concepts of the WCSO, automatically constructing “scalar value representations” for storing the values and their units in accordance with the WCSO and underlying PMDco. To demonstrate the versatility and broad applicability of the WCSO, we chose seven different synthesis procedures that were previously executed on our SDL.4 The example ttl files for these syntheses are provided in the GitHub repository accompanying this article and serve as the knowledge base for the example SPARQL queries in the Jupyter notebook demonstrator.We also included a simple, end-to-end example of how an (imprecise and incomplete) sentence from a synthesis procedure (“The solution was heated at 80 C for 12 hours, while reagentX was added dropwise.”) can be exported as a knowledge graph that follows the WCSO using a simple, graphical tool (node editor), without requiring any deep knowledge of the underlying ontological concepts (see Fig. 1 and Table 2 in the SI).
The example also discusses how heuristics and error correction measures are implemented in the node editor prior to knowledge graph creation to handle imprecise experimental descriptions or missing parameters from the synthesis protocol. For example, imprecise time (e.g., overnight), temperature (room temperature), addition rate (dropwise) or stirring speed (vigorous stirring) specifications can be replaced with default values, such as 16 hours, 25 °C, 1 mL min−1, and 600 rpm, while missing process parameters can be substituted with pre-defined default values (e.g., always using 300 rpm as the stirring speed or 8000 rpm as the centrifugation speed if no values are specified). Further information on these heuristics and substitutions, as well as a comparison of the accuracy of the extraction of synthesis actions by the different models and their performance when being applied to longer and more complex synthetic procedures from different scientific domains can be found in our previous publication.8 It should be noted that the large language models sometimes fail to capture the intended sequence of operations (e.g., should an addition and heating be performed sequentially or in parallel), or to assign qualitative process parameter descriptions correctly to the individual process steps. In that case, the user can manually correct the node setup before exporting it as a knowledge graph, ensuring that the semantic annotation in the knowledge graph will be correct when querying the knowledge base later on.
Jupyter Notebook demonstrator
For presenting the use case and implementing a lightweight demonstrator around the wet-chemical (nano)material synthesis data, Jupyter Notebooks were used. These are particularly well suited for demonstration because they combine human-readable narrative (provided in Markdown format and rendered, accordingly) with executable code cells which allows computational workflows to be documented, explained, and reproduced in a step-by-step manner. This literate-programming style supports transparent communication of assumptions, intermediate results, and parameter choices. Additionally, readers are enabled to rerun and adapt single code blocks without requiring a full software deployment pipeline.Notebook-based SPARQL information retrieval demonstrator
To demonstrate data access and information retrieval over the created knowledge graph, a dedicated Jupyter Notebook is provided. The notebook establishes a local triple store using the OWLready2 Python package by instantiating an owlready2.World() object and loading (i) the PMD Core Ontology (PMDco), (ii) the WCSO, and (iii) several example A-Box datasets (RDF knowledge graphs created from Turtle files), based on previously described and published wet-chemical synthesis workflows,4 into the same in-memory store. This setup yields a self-contained querying environment in which SPARQL queries can be executed directly against the loaded graph.It has to be noted that the use of Owlready2 in the demonstrator is intended as a pragmatic, self-contained way to load OWL/RDF artifacts into a local environment and execute SPARQL queries without additional infrastructure. In larger software-engineering settings, e.g., pydantic-based59 object-graph mappers can provide a more object-oriented developer experience by synchronizing typed Python class hierarchies with RDF knowledge graphs, as illustrated by recent tooling.32,60 Schema-first approaches can further treat a higher-level schema as a “single source of truth” and generate multiple artifacts (e.g., pydantic models, JSON-LD contexts, and OWL/Turtle renderings) as needed; representative examples include LinkML's OWL and pydantic generators61 and OO-LD's combined JSON-Schema/JSON-LD62 approach. These alternatives are compatible with the OWL-first ontology design present in this study and are considered promising integration options for future, larger-scale deployments. Furthermore, dedicated benchmarking of reasoning performance or query complexity was not performed in the present work, as the focus lies on ontology design, modelling patterns, and demonstrator-scale querying rather than on evaluating large-scale deployment scenarios. During ontology development, standard OWL reasoning was used for routine consistency checking and (where applicable) classification of the TBox. Example queries in a demonstrator were executed to validate expressivity and competency-question coverage; practical runtime performance is expected to depend on the selected triple store implementation, indexing strategy, and dataset scale which may be assessed in detail when larger cross-laboratory knowledge bases become available. In general, an ontology-grounded representation is intended to support more complex classes of cross-workflow competency questions that rely on (i) temporally structured process parts (including concurrency), (ii) subclass-aware retrieval over action categories, and (iii) parameter extraction via reusable process-attribute patterns. As an illustrative example of such a query scenario, workflows can be retrieved in which an addition step is performed concurrently with heating (and optionally stirring), followed by any separation step (e.g., filtration or centrifugation), while returning associated parameter values (e.g., addition rate, temperature setpoint, separation duration) together with the produced material and recorded outcome descriptors (e.g., yield variants). Such retrieval can be expressed as a graph-pattern match over temporal relations and variable-length process-part structures, while remaining extensible to new subclasses of “separation” without schema redesign. In conventional relational database schemas, implementing the same retrieval robustly typically requires rigid, predeclared tables for each step type, extensive join logic across workflows of variable length, and ad-hoc schema changes when additional procedural variants or relations are introduced.
Using the self-contained querying environment setup in this work, we demonstrate 5 example queries of varying complexity that can be interesting in real-life settings when (re-)using the data from the knowledge graphs to run the described syntheses on an SDL backend. The first example demonstrates how to retrieve some information about the knowledge base comprising all the examples that are provided by building a simple SPARQL query that counts all the entities and another query for retrieving the descriptions of all the output materials. The next example demonstrates how to extract the centrifugation times (values and measurement units) of all the steps that were used in the workflow describing the Au@MSN Core-Shell nanoparticle synthesis. The third example demonstrates how to query the amount of a certain solvent (here simply water) that is required for a synthesis workflow describing the synthesis of gold nanoparticles. Besides helping with planning the actual synthesis before executing it, queries like this, when applied to a large collection of different synthesis workflows, can be used to analyze, compare, and rank the individual procedures based on sustainability metrics such as solvent consumption or filter for “green” solvents. The fourth example demonstrates how to retrieve the set of chemicals required for the CuO nanoparticle (CuO-NP) synthesis from the knowledge graph, together with commonly used chemical identifiers. To produce a compact and human-readable output, the query leverages the identifier naming scheme encoded in the identifier IRIs while referring to the corresponding chemical name, CAS Registry Number, and SMILES representation, respectively. Distinct tuples of name, CAS, SMILES are returned which yields a clean table of the chemicals needed for the CuO-NP synthesis that is directly suitable for downstream tasks such as reagent planning, procurement, and inventory checks. This also enables cross-workflow comparisons of chemical inputs when applied to larger collections of synthesis protocols. The fifth example demonstrates how to list all the chemicals and their amounts that are required across all the syntheses that are loaded in the triple store. Again, besides helping with inventory management, queries like this can be used to gain knowledge about, e.g., the use of toxic or critical raw materials and can be used in the context of a Safe and Sustainable by Design (SSbD) evaluation.
Within the notebook, small helper functions are defined to facilitate inspection and subsequent analysis. For a convenient presentation of the results, the notebook shows the outputs as readable tables from pandas dataframes.
Artificial intelligence tools
Microsoft Copilot was used during the proof-reading of the manuscript to enhance the language, clarity, and readability of some parts of the text. The authors carefully checked the generated sentences and ensured their correctness and scientific rigor, and take full responsibility for the content of this article.Conclusions
This work introduces the WeChemSyn Ontology (WCSO) and demonstrates its applicability to wet chemical nanomaterial syntheses in SDLs. Knowledge graphs based on the WCSO were automatically constructed for seven different nanoparticle syntheses by exporting their corresponding workflows for the MinervaOS backend. This direct export of knowledge graphs using an intuitive graphical user interface (such as a node editor) that represents the executed workflow and automatically takes care of applying the correct ontological concepts (such as the “SVS-patterns”, annotating process steps as sequential or concurrent, etc.) can help making the application of the ontology to constructing knowledge graphs much more approachable for mainly synthetically trained chemists and materials scientists. At the same time, it significantly reduces the amount of “boilerplate” users need to take care of when creating knowledge graphs aligned with the underlying ontological concepts.In addition, natural language (NL) interfaces based on large language models (LLMs) can further reduce barriers for non-experts by assisting with query formulation and information retrieval over ontology-grounded knowledge graphs. In particular, NL-to-SPARQL support (e.g., via LLM-assisted query generation combined with retrieval over ontology documentation) represents a promising direction to enable ad-hoc cross-workflow questions without requiring users to write SPARQL manually. Such approaches are considered complementary to the GUI-based workflow abstraction and will become increasingly practical as robust constrained-output and validation strategies mature.
Example queries for such knowledge graphs are presented in an executable Jupyter Notebook and demonstrate how these knowledge graphs, once they form a comprehensive knowledge base, can be used for obtaining information about the automated syntheses workflows that is crucial for planning, executing, and reproducing these syntheses on other SDLs or MAPs. Moreover, information about the use and required amount of certain chemicals, e.g., critical raw materials or hazardous or toxic chemicals, can readily be obtained by querying the knowledge graphs and can contribute to informed decisions on, e.g., Safe and Sustainable by Design (SSbD) criteria when planning and executing new material syntheses in the future.
Grounded in the PMDco-BFO semantic framework, the WCSO represents a first step toward a reusable and interoperable semantic foundation for the domain of materials science in general and nano and advanced materials synthesis in MAPs/SDLs in particular. It formalizes synthesis procedures, materials, process parameters, and outcomes in a manner that supports consistent data integration across automated experimentation, analysis, and control workflows, which are central to SDL operation. This can also help with the interoperability and reproducibility of workflows across different MAPs and SDL platforms that use different orchestrator backends. However, broad interoperability depends on adoption and alignment within the community; WCSO is therefore presented as a foundational contribution and reference implementation rather than as a complete, universal solution.
Beyond reusing and defining domain concepts, the WCSO provides explicit usage patterns that enable systematic reuse and semantic interoperability across heterogeneous SDL components. By establishing standardized terminology and process representations, it facilitates the integration of wet chemical synthesis knowledge into material synthesis, optimization, automated reasoning, and data-driven decision-making. Using SHACL (SHApes Constraint Language), it will be possible to directly validate workflows before their execution. This would also help to abstract, formalize, and share heuristics (e.g., check whether the available volume in a container is large enough before attempting the addition of a chemical) or, in the long run, even implicit or tacit knowledge (e.g., which pipette and which aspiration and dispension rate to use in an addition step) that is often encoded in the orchestrator backends or workflows.
The ontology is openly maintained and designed for community-driven extension, while already offering adaptable patterns aligned with the requirements of MAPs. The resulting semantically structured datasets make the workflows “AI-ready”, which positions this work as an enabling infrastructure for scalable, autonomous discovery in wet chemical nanomaterial syntheses.
Author contributions
Conceptualization: MS, BB, BR; data curation: MS, HB, BB, BR; investigation: MS, HB, BB, BR; methodology: MS, BB, BR; project administration: MS, BB, BR; software: MS, BR; supervision: BB, BR; validation: MS, HB, BB, BR; visualization: MS, BB, BR; writing – original draft: MS, BB, BR; writing – review & editing: MS, HB, BB, BR.Conflicts of interest
There are no conflicts to declare.Data availability
Data for this article, including the data related to the ontology and its development, as well as specific sample data sets (created with the workflow node editor found at https://github.com/BAMresearch/MAPz_at_BAM/tree/main/Minerva-Workflow-Generator) and a demonstrator for data retrieval in the form of a Jupyter Notebook, are available at a freely accessible, dedicated GitHub repository entitled BAMresearch/wcso: https://github.com/BAMresearch/wcso. All these data are also available on Zenodo (DOI: https://doi.org/10.5281/zenodo.19609498) at https://zenodo.org/records/19609499. The technical documentation on the ontology is also accessible via its namespace (persistent URL), https://w3id.org/wcso, as well as viahttps://bamresearch.github.io/wcso/and https://bamresearch.github.io/wcso/docs/. Further data can also be found in the supplementary information. Supplementary information: table with selected reusable ontologies and their relevance for wet chemical nanomaterial synthesis; GitHub repository including the data related to the ontology and its development, as well as specific sample data sets and a demonstrator for data retrieval as a Jupyter Notebook. See DOI: https://doi.org/10.1039/d6dd00058d.Acknowledgements
The Federal Institute for Materials Research and Testing (Bundesanstalt für Materialforschung und-prüfung (BAM)) and the Materials Acceleration Platform Center at BAM (MAPz@BAM) are gratefully acknowledged.Notes and references
- M. Baker, Nature, 2016, 533, 452–454 CrossRef CAS PubMed.
- S. P. Stier, C. Kreisbeck, H. Ihssen, M. A. Popp, J. Hauch, K. Malek, M. Reynaud, T. P. M. Goumans, J. Carlsson, I. Todorov, L. Gold, A. Rader, W. Wenzel, S. T. Bandesha, P. Jacques, F. Garcia-Moreno, O. Arcelus, P. Friederich, S. Clark, M. Maglione, A. Laukkanen, I. E. Castelli, J. Carrasco, M. C. Cabanas, H. S. Stein, O. Ozcan, D. Elbert, K. Reuter, C. Scheurer, M. Demura, S. S. Han, T. Vegge, S. Nakamae, M. Fabrizio and M. Kozdras, Adv. Mater., 2024, 36, e2407791 CrossRef PubMed.
- J. Wagner, C. G. Berger, X. Du, T. Stubhan, J. A. Hauch and C. J. Brabec, J. Mater. Sci., 2021, 56, 16422–16446 CrossRef CAS.
- M. Zaki, C. Prinz and B. Ruehle, ACS Nano, 2025, 19, 9029–9041 CrossRef CAS PubMed.
- L. Hawizy, D. M. Jessop, N. Adams and P. Murray-Rust, J. Cheminf., 2011, 3, 17 CAS.
- M. Krallinger, O. Rabal, A. Lourenco, J. Oyarzabal and A. Valencia, Chem. Rev., 2017, 117, 7673–7761 CrossRef CAS PubMed.
- S. Mysore, E. Kim, E. Strubell, A. Liu, H.-S. Chang, S. Kompella, K. Huang, A. McCallum and E. Olivetti, Automatically Extracting Action Graphs from Materials Science Synthesis Procedure, arXiv, 2017, preprint, arXiv:1711.06872, DOI:10.48550/arXiv.1711.06872.
- B. Ruehle, Digital Discovery, 2025, 4, 1534–1543 RSC.
- A. C. Vaucher, F. Zipoli, J. Geluykens, V. H. Nair, P. Schwaller and T. Laino, Nat. Commun., 2020, 11, 3601 CrossRef CAS PubMed.
- N. Yoshikawa, M. Skreta, K. Darvish, S. Arellano-Rubach, Z. Ji, L. Bjørn Kristensen, A. Z. Li, Y. Zhao, H. Xu, A. Kuramshin, A. Aspuru-Guzik, F. Shkurti and A. Garg, Autonomous Robots, 2023, 47, 1057–1086 CrossRef.
- G. A. J. Zia, T. Hanke, B. Skrotzki, C. Völker and B. Bayerlein, ntegr. Mater. Manuf. Innov., 2024, 13, 257–271 CrossRef.
- M. D. Wilkinson, M. Dumontier, I. J. Aalbersberg, G. Appleton, M. Axton, A. Baak, N. Blomberg, J. W. Boiten, L. B. da Silva Santos, P. E. Bourne, J. Bouwman, A. J. Brookes, T. Clark, M. Crosas, I. Dillo, O. Dumon, S. Edmunds, C. T. Evelo, R. Finkers, A. Gonzalez-Beltran, A. J. Gray, P. Groth, C. Goble, J. S. Grethe, J. Heringa, P. A. t Hoen, R. Hooft, T. Kuhn, R. Kok, J. Kok, S. J. Lusher, M. E. Martone, A. Mons, A. L. Packer, B. Persson, P. Rocca-Serra, M. Roos, R. van Schaik, S. A. Sansone, E. Schultes, T. Sengstag, T. Slater, G. Strawn, M. A. Swertz, M. Thompson, J. van der Lei, E. van Mulligen, J. Velterop, A. Waagmeester, P. Wittenburg, K. Wolstencroft, J. Zhao and B. Mons, Sci. Data, 2016, 3, 160018 CrossRef PubMed.
- J. Bai, S. Mosbach, C. J. Taylor, D. Karan, K. F. Lee, S. D. Rihm, J. Akroyd, A. A. Lapkin and M. Kraft, Nat. Commun., 2024, 15, 462 CrossRef CAS PubMed.
- S. Steiner, J. Wolf, S. Glatzel, A. Andreou, J. M. Granda, G. Keenan, T. Hinkley, G. Aragon-Camarasa, P. J. Kitson, D. Angelone and L. Cronin, Science, 2019, 363(642), eaav2211 CrossRef CAS PubMed.
- D. P. Tabor, L. M. Roch, S. K. Saikin, C. Kreisbeck, D. Sheberla, J. H. Montoya, S. Dwaraknath, M. Aykol, C. Ortiz, H. Tribukait, C. Amador-Bedolla, C. J. Brabec, B. Maruyama, K. A. Persson and A. Aspuru-Guzik, Nat. Rev. Mater., 2018, 3, 5–20 CrossRef CAS.
- T. Tüccar, J. Knisz, R. Eckert and T. L. Skovhus, npj Mater. Degrad., 2025, 10(1), 6 CrossRef.
- ISO/IEC 21838-1, 2021.
- ISO/IEC 21838-2, 2021.
- Material Digital, https://materialdigital.de/, accessed 25.01, 2026 Search PubMed.
- Material Digital Application Ontology Template, https://github.com/materialdigital/application-ontology-template, accessed 25.01. 2026 Search PubMed.
- B. Bayerlein, M. Schilling, H. Birkholz, M. Jung, J. Waitelonis, L. Mädler and H. Sack, Mater. Des., 2024, 237 Search PubMed.
- SI Brochure: The International System of Units (SI), https://www.bipm.org/en/publications/si-brochure/, accessed 26.01. 2026.
- ISO 80004-1, 2023.
- OECD, OECD Guidelines for the Testing of Chemicals, Section 1, 2023, DOI:10.1787/af5f9bda-en.
- B. Bayerlein, M. Schilling, P. von Hartrott and J. Waitelonis, Sci. Data, 2024, 11, 434 CrossRef PubMed.
- B. Bayerlein, J. Waitelonis, H. Birkholz, M. Jung, M. Schilling, P. v. Hartrott, M. Bruns, J. Schaarschmidt, K. Beilke, M. Mutz, V. Nebel, V. Königer, L. Beran, T. Kraus, A. Vyas, L. Vogt, M. Blum, B. Ell, Y. F. Chen, T. Waurischk, A. Thomas, A. R. Durmaz, S. Ben Hassine, C. Fresemann, G. Dziwis, H. Beygi Nasrabadi, T. Hanke, M. Telong, S. Pirskawetz, M. Kamal, T. Bjarsch, U. Pähler, P. Hofmann, M. Leemhuis, Ö. L. Özçep, L. P. Meyer, B. Skrotzki, J. Neugebauer, W. Wenzel, H. Sack, C. Eberl, P. D. Portella, T. Hickel, L. Mädler and P. Gumbsch, Adv. Eng. Mater., 2024, 27, 2401092 CrossRef.
- M. Schilling, B. Bayerlein, P. von Hartrott, J. Waitelonis, H. Birkholz, P. Dolabella Portella and B. Skrotzki, Adv. Eng. Mater., 2024, 27, 2400138 CrossRef.
- M. Schilling, S. Bruns, B. Bayerlein, J. Kryeziu, J. Schaarschmidt, J. Waitelonis, P. Dolabella Portella and K. Durst, Adv. Eng. Mater., 2024, 27, 2401527 CrossRef.
- M. Schilling, N. Marschall, U. Niebergall and M. Böhning, Comput. Mater. Sci., 2025, 259, 114085 CrossRef CAS.
- A. V. Gulyuk, N. Abu Zaid, R. Chirkova and Y. G. Yingling, APL Mach. Learn., 2025, 3, 040901 CrossRef.
- S. D. Rihm, J. Bai, A. Kondinski, S. Mosbach, J. Akroyd and M. Kraft, Nexus, 2024, 1, 100004 CrossRef.
- S. D. Rihm, F. Saluz, A. Kondinski, J. Bai, P. W. V. Butler, S. Mosbach, J. Akroyd and M. Kraft, Digital Discovery, 2025, 4, 2893–2909 RSC.
- X. Zhou, P. Bulter, C. Yang, S. D. Rihm, T. Angkanaporn, J. Akroyd, S. Mosbach and M. Kraft, Ontology-to-tools compilation for executable semantic constraint enforcement in LLM agents, arXiv, 2026, preprint, arXiv:2602.03439, DOI:10.48550/arXiv.2602.03439.
- J. Bai, A. Aldossary, T. Swanick, M. Müller, Y. Kang, Z. Zhang, J. W. Lee, T. W. Ko, M. G. Vakili, V. Bernales and A. Aspuru-Guzik, El Agente Gráfico: Structured Execution Graphs for Scientific Agents, arXiv, 2026, preprint, arXiv:2602.17902, DOI:10.48550/arXiv.2602.17902.
- E. Norouzi, N. Jung, A. M. Jacyszyn, J. Waitelonis and H. Sack, AI4DiTraRe: Building the BFO-Compliant Chemotion Knowledge Graph, 2025 Search PubMed.
- M. Schilling, S. Bruns, B. Bayerlein, J. Kryeziu, J. Schaarschmidt, J. Waitelonis, P. D. Portella and K. Durst, Adv. Eng. Mater., 2025, 27, 13 Search PubMed.
- Ontobee, https://ontobee.org/, accessed 25.01., 2026.
- TIB Terminology Service, https://terminology.tib.eu/ts, accessed 25.01., 2026 Search PubMed.
- MatPortal, https://matportal.org/, accessed 25.01., 2026 Search PubMed.
- W. Zhang, L. Hao, V. Lai, R. Corkery, J. Jessiman, J. Zhang, J. Liu, Y. Sato, M. Politi, M. E. Reish, R. Greenwood, N. Depner, J. Min, R. El-Khawaldeh, P. Prieto, E. Trushina and J. E. Hein, Nat. Commun., 2025, 16, 5182 CrossRef CAS PubMed.
- M. Sim, M. G. Vakili, F. Strieth-Kalthoff, H. Hao, R. J. Hickman, S. Miret, S. Pablo-García and A. Aspuru-Guzik, Matter, 2024, 7, 2959–2977 CrossRef CAS.
- Y. Fei, B. Rendy, R. Kumar, O. Dartsi, H. P. Sahasrabuddhe, M. J. McDermott, Z. Wang, N. J. Szymanski, L. N. Walters, D. Milsted, Y. Zeng, A. Jain and G. Ceder, Digital Discovery, 2024, 3, 2275–2288 RSC.
- M. Vogler, S. K. Steensen, F. F. Ramírez, L. Merker, J. Busk, J. M. Carlsson, L. H. Rieger, B. Zhang, F. Liot, G. Pizzi, F. Hanke, E. Flores, H. Hajiyani, S. Fuchs, A. Sanin, M. Gaberšček, I. E. Castelli, S. Clark, T. Vegge, A. Bhowmik and H. S. Stein, Adv. Energy Mater., 2024, 14(46), 2403263 CrossRef CAS.
- R. Tamura, K. Tsuda and S. Matsuda, Sci. Technol. Adv. Mater.:Methods, 2023, 3, 2232297 Search PubMed.
- F. Rahmanian, J. Flowers, D. Guevarra, M. Richter, M. Fichtner, P. Donnely, J. M. Gregoire and H. S. Stein, Adv. Mater. Interfaces, 2022, 9(8), 2101987 CrossRef.
- MADSci, https://github.com/AD-SDL/MADSci, accessed 25.01., 2026.
- A. Valdestilhas, B. Bayerlein, B. Moreno Torres, G. A. J. Zia and T. Muth, Adv. Intell. Syst., 2023, 5(8), 2300051 CrossRef.
- E. Callaway, Nature, 2024, 634, 525–526 CrossRef CAS PubMed.
- C. Gene Ontology, S. A. Aleksander, J. Balhoff, S. Carbon, J. M. Cherry, H. J. Drabkin, D. Ebert, M. Feuermann, P. Gaudet, N. L. Harris, D. P. Hill, R. Lee, H. Mi, S. Moxon, C. J. Mungall, A. Muruganugan, T. Mushayahama, P. W. Sternberg, P. D. Thomas, K. Van Auken, J. Ramsey, D. A. Siegele, R. L. Chisholm, P. Fey, M. C. Aspromonte, M. V. Nugnes, F. Quaglia, S. Tosatto, M. Giglio, S. Nadendla, G. Antonazzo, H. Attrill, G. Dos Santos, S. Marygold, V. Strelets, C. J. Tabone, J. Thurmond, P. Zhou, S. H. Ahmed, P. Asanitthong, D. Luna Buitrago, M. N. Erdol, M. C. Gage, M. Ali Kadhum, K. Y. C. Li, M. Long, A. Michalak, A. Pesala, A. Pritazahra, S. C. C. Saverimuttu, R. Su, K. E. Thurlow, R. C. Lovering, C. Logie, S. Oliferenko, J. Blake, K. Christie, L. Corbani, M. E. Dolan, H. J. Drabkin, D. P. Hill, L. Ni, D. Sitnikov, C. Smith, A. Cuzick, J. Seager, L. Cooper, J. Elser, P. Jaiswal, P. Gupta, P. Jaiswal, S. Naithani, M. Lera-Ramirez, K. Rutherford, V. Wood, J. L. De Pons, M. R. Dwinell, G. T. Hayman, M. L. Kaldunski, A. E. Kwitek, S. J. F. Laulederkind, M. A. Tutaj, M. Vedi, S. J. Wang, P. D’Eustachio, L. Aimo, K. Axelsen, A. Bridge, N. Hyka-Nouspikel, A. Morgat, S. A. Aleksander, J. M. Cherry, S. R. Engel, K. Karra, S. R. Miyasato, R. S. Nash, M. S. Skrzypek, S. Weng, E. D. Wong, E. Bakker, T. Z. Berardini, L. Reiser, A. Auchincloss, K. Axelsen, G. Argoud-Puy, M. C. Blatter, E. Boutet, L. Breuza, A. Bridge, C. Casals-Casas, E. Coudert, A. Estreicher, M. Livia Famiglietti, M. Feuermann, A. Gos, N. Gruaz-Gumowski, C. Hulo, N. Hyka-Nouspikel, F. Jungo, P. Le Mercier, D. Lieberherr, P. Masson, A. Morgat, I. Pedruzzi, L. Pourcel, S. Poux, C. Rivoire, S. Sundaram, A. Bateman, E. Bowler-Barnett, A. J. H. Bye, P. Denny, A. Ignatchenko, R. Ishtiaq, A. Lock, Y. Lussi, M. Magrane, M. J. Martin, S. Orchard, P. Raposo, E. Speretta, N. Tyagi, K. Warner, R. Zaru, A. D. Diehl, R. Lee, J. Chan, S. Diamantakis, D. Raciti, M. Zarowiecki, M. Fisher, C. James-Zorn, V. Ponferrada, A. Zorn, S. Ramachandran, L. Ruzicka and M. Westerfield, Genetics, 2023, 224(1), iyad031 CrossRef PubMed.
- A. Malik, M. Arsalan, C. Moreno, J. Mosquera, E. Felix, T. Kiziloren, V. Muthukrishnan, B. Zdrazil, A. R. Leach and N. M. O'Boyle, Nucleic Acids Res., 2026, 54, D1768–D1778 CrossRef PubMed.
- Huggingface, https://huggingface.co/bruehle, accessed 25.01., 2026.
- ISO 15926-10:2019, 2019, 41.
- Protege, https://protege.stanford.edu/, accessed 26.01., 2026.
- H. Knublauch, R. W. Fergerson, N. F. Noy and M. A. Musen, The Semantic Web – ISWC 2004, 2004 Search PubMed.
- The Ontology Development Kit (ODK), https://incatools.github.io/ontology-development-kit/, accessed 25.01., 2026 Search PubMed.
- Open Biological and Biomedical Ontology Foundry, https://obofoundry.org/, accessed 25.01., 2026 Search PubMed.
- ROBOT, https://github.com/ontodev/robot, accessed 25.01., 2026.
- MkDocs, https://www.mkdocs.org/, accessed 25.01., 2026.
- pydantic, https://github.com/pydantic/pydantic, accessed 20.03., 2026.
- J. Bai, S. D. Rihm, A. Kondinski, F. Saluz, X. Deng, G. Brownbridge, S. Mosbach, J. Akroyd and M. Kraft, Digital Discovery, 2025, 4, 2123–2135 RSC.
- Link-ML generators, DOI:10.48550/arXiv.2511.16935, accessed 20.03., 2026.
- OO-LD schema, DOI:10.5281/zenodo.17205005, accessed 20.03., 2026.
| This journal is © The Royal Society of Chemistry 2026 |