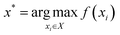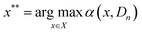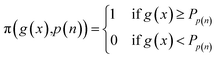Open Access Article
Open Access ArticleDistilling and exploiting quantitative insights from large language models for enhanced Bayesian optimization of chemical reactions
Roshan A.
Patel
a,
Mingxuan
Li
b,
Chin-Fei
Chang
a,
Louis
De Lescure
a,
Paul
Chauvin
c,
Alan
Cherney
a,
Saeed
Moayedpour
b,
Sven
Jager
d and
Yasser
Jangjou
 *a
*a
aCMC Synthetics Platform, Sanofi, 350 Water St, Cambridge, MA 02141, USA. E-mail: Yasser.Jangjou@sanofi.com
bDigital R&D, Sanofi, 450 Water St, Cambridge, MA 02141, USA
cDigital R&D, Sanofi, 58-60 Avenue de la Grande Armée, Paris, France
dDigital R&D, Sanofi, Frankfurt 65929, Germany
First published on 6th April 2026
Abstract
Machine learning and Bayesian optimization (BO) algorithms can significantly accelerate the optimization of chemical reactions. Transfer learning can bolster the effectiveness of BO algorithms in low-data regimes by leveraging pre-existing chemical information or data outside the direct optimization task (i.e., source data). Large Language Models (LLMs) have demonstrated that chemical information present in foundation training data can give them utility for processing chemical data. Furthermore, they can be augmented with and help synthesize potentially multiple modalities of source chemical data germane to the optimization task. In this work, we examine how chemical information from LLMs can be elicited and used for transfer learning to accelerate the BO of reaction conditions to maximize yield. Specifically, we show that a survey-like prompting scheme and preference learning can be used to infer a utility function which models prior chemical information embedded in LLMs over a chemical parameter space; we find that the utility function shows modest correlation to true experimental measurements (yield) over the parameter space despite operating in a zero-shot setting. Furthermore, we show that the utility function can be leveraged to focus BO efforts in promising regions of the parameter space, improving the yield of the initial BO query and enhancing optimization in a majority of the datasets studied. Overall, we view this work as a step towards bridging the gap between the chemistry knowledge embedded in LLMs and the capabilities of principled BO methods to accelerate reaction optimization.
1 Introduction
Machine learning and data-driven approaches can significantly accelerate the optimization of chemical processes.3,9,10,52 In applications where data is insufficient for comprehensive predictive modeling (e.g., high-throughput screening), Bayesian optimization (BO) algorithms stand out as data-efficient methods to iteratively navigate the chemical and process parameter space to target desired properties from the chemical product.7,34,35,45 For example, Shields et al.35 show that BO can work well to identify chemicals (e.g., base, solvent, catalyst ligands) and reaction conditions (e.g., temperature, chemical concentration) to maximize the yields of Buchwald–Hartwig coupling, Suzuki–Miyaura coupling, and direct arylation reactions. We refer readers to a recent review by Guo and Rankovic et al.15 for comprehensive discussion on successful applications of BO for chemical process development.Transfer learning can significantly accelerate BO-led workflows by leveraging (source) information or data outside of the direct domain of a given optimization task.4,12,38 For example, source datasets can be used to better inform model development for the domain task.33,39,42 In addition, source data can be used to identify and focus optimization efforts on promising regions of the parameter space through modification of the acquisition function.1,4,17,19,40,41,48 Overall, though, the application of these and other transfer learning strategies for BO heavily rely on the identification, curation, and numerical encoding of relevant source datasets which are often difficult and laborious to accomplish in practice. Furthermore, qualitative information outside organized datasets (e.g., insights/conclusions found as text in research articles) are typically not leveraged for transfer learning despite representing a large volume of information pertinent for new chemical design tasks.
In recent years, large language models (LLMs) have demonstrated that their ability to model natural language can help perform challenging tasks in disparate chemical domains.16,43 For example, with in-context learning, LLMs have been used as regression and classification models to predict chemical properties.20–22,29 In addition, LLMs have shown promise for designing experiments in the context of chemical optimization problems or making meta decisions about the optimization process (e.g., integrating several systems/software needed to execute optimization or deciding a suitable stopping condition).6,24,27,29,30,32 Given these observations, we posit that LLMs can be queried to transfer pertinent chemical information from source data (e.g., foundation model training data, fine-tuning data) to target Bayesian optimization campaigns and accelerate process development.
In this study, we examine how information from LLMs can be distilled and used to accelerate Bayesian reaction optimization through transfer learning. Specifically, we show that preference learning8,13 can be used to infer a utility function over the reaction parameter space from LLM-answered surveys that shows modest correlation to measured reaction yields; promisingly, we accomplish this despite operating in a zero-shot setting with no in-context learning11,20,22,29 or fine-tuning.18,20,24 Furthermore, we show that when incorporated in the acquisition function, the utility function can be used to focus BO queries in promising regions of the parameter space. We observe that this significantly improves the yield of the initial query to seed BO and enhances optimization in several of the datasets studied. Overall, we view this work as a step towards bridging the gap between the chemistry knowledge embedded in LLMs and the capabilities of principled BO methods to accelerate reaction optimization.
2 Methods
2.1 Datasets
We explore our approach with six chemical reaction datasets compiled by Shields et al.35–37 (accessed date: June 2024). Datasets 1–5 correspond to Buchwald–Hartwig (BH) reactions and each contain 792 recorded experiments. Experiments in these datasets are characterized by four reaction parameters: the identity of a specific aryl halide reactant, the palladium precatalyst, the additive, and the base used for the reaction and are labeled by a measured product yield. Dataset 6 corresponds to a direct arylation (DA) reaction and contains 1728 experiments. Experiments in this dataset are characterized by five reaction parameters: the identity of a palladium catalyst ligand, the base, the solvent, temperature, and concentration and are also labeled with a measured product yield. The objective for all datasets is to identify the experiment, characterized by a specific set of reaction parameters, that will give the maximum product yield.Finally, to evaluate the method's generalization capabilities and rule out data contamination, we utilize three Amide Coupling datasets (AC1–3, accessed date: Oct. 2025) from a study published in 2025.50,51 As these datasets post-date the training cutoff of the LLMs investigated in this study, they serve as a rigorous test of the models' ability to apply chemical reasoning to truly unseen reaction spaces. These datasets involve the optimization of coupling reagents, solvents, and bases for distinct amide bond formation reactions.
2.2 Formulation and implementation of approach
The overall approach taken for each dataset is qualitatively presented in Fig. 1. Step 1 of the approach aims at distilling chemical insights from the LLM in the form of a utility function g(x). In step 1a, we formulate a survey in which each question presents two experiments, each characterized by a different set of experimental parameters. In step 1b, we prompt the LLM to answer the survey, selecting which experiment (A or B) it predicts will give the higher yield for each question. In step 1c, we use preference learning to infer a utility function g(x) based on the preferences expressed as choices in the survey. Since we prompt the LLM to prefer experiments with higher predicted yields, we expect g(x) to correlate to yield and thus represent useful, quantitative prior information from the LLM over the set of experiments in a dataset.Step 2 of the approach aims at leveraging g(x) to expedite BO of reaction parameters. As discussed more in Section 2.2.3, we use g(x) to identify promising regions of the parameter space and constrain optimization to these experiments only. Thus in step 2a, we begin optimization by randomly selecting an experiment in the promising set of experiments. In step 2b, we perform BO, gradually removing restrictions on the design space imposed by g(x) as more data is available for surrogate modeling. The pseudo code for the approach is provided at the end of Section 2.2.3.
where xi is a variable that represents a single candidate experiment, X represents the full set of candidate experiments considered as possible designs, and f(xi) represents the noiseless output quantity (e.g., yield) that should result from performing experiment xi; typically, we can access noisy measurements of f(xi) through experiments: yi = f(xi) + ε. In our work, we represent an experiment xi as a concatenation of one-hot-encoded categorical variables (e.g., base, ligand, solvent identity) and continuous variables (e.g., concentration and temperature). Given that obtaining measurements can be time consuming/expensive, the goal is to identify the x* among all candidate experiments that gives the maximum value of f(x) with as few experiments as possible. Bayesian optimization (BO) is an iterative approach utilizing probabilistic modeling that can be used to solve this class of optimization tasks. At iteration n of BO, we have measured the output of n experiments giving us dataset Dn = {(xi,yi)}i=1n. Following, the dataset is used to develop a surrogate model
![[f with combining circumflex]](https://www.rsc.org/images/entities/i_char_0066_0302.gif) used to predict the output of a given experiment and estimate an uncertainty in that prediction. Gaussian process regression models31 and Bayesian neural networks23 are both common surrogate modeling strategies, offering principled methods to estimate a predictive posterior distribution p(ŷi∣Dn,xi). We employ the modeling strategy developed by Shields et al.,35 which leverages GPR with specific priors on kernel parameters suitable for the experiment representation strategy described in this work. All the details of the surrogate model, including kernel specification, prior mean definition and hyperparameter priors, are provided in the SI (section S3). The surrogate model is used to compute terms in an acquisition function α(x, Dn), the maximizing argument of which is selected as the next best experiment to obtain a measurement for:
used to predict the output of a given experiment and estimate an uncertainty in that prediction. Gaussian process regression models31 and Bayesian neural networks23 are both common surrogate modeling strategies, offering principled methods to estimate a predictive posterior distribution p(ŷi∣Dn,xi). We employ the modeling strategy developed by Shields et al.,35 which leverages GPR with specific priors on kernel parameters suitable for the experiment representation strategy described in this work. All the details of the surrogate model, including kernel specification, prior mean definition and hyperparameter priors, are provided in the SI (section S3). The surrogate model is used to compute terms in an acquisition function α(x, Dn), the maximizing argument of which is selected as the next best experiment to obtain a measurement for:The expected improvement (EI) function is a popular choice among studied acquisition functions and has been applied extensively for chemical design:25,34
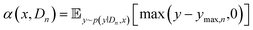 | (1) |
 ) is provided in Fig. S3 of the SI. To design the survey questions Sunanswered for a given dataset, we created two identical arrays, where each array contains L instances of every experiment in a dataset. Then, we randomly paired elements between each array to form questions, removing repeated questions and questions where the paired experiments were identical. For datasets BH1–BH5 we set L to 10 and for DA, L was set to 5 (to keep the total number of questions roughly similar to the BH surveys). This resulted in 7792, 7842, 7788, 7825, 7804, and 8610 survey questions designed for the BH1–5 and DA datasets respectively. Overall, we hypothesized that surveys generated using this procedure would facilitate expressing hierarchical preferences over the full set of experiments and subsequent preference learning. In Section 4.1, we briefly evaluate the performance of several commonly used foundation LLM models for answering survey questions correctly and select the most accurate model to complete our surveys.
) is provided in Fig. S3 of the SI. To design the survey questions Sunanswered for a given dataset, we created two identical arrays, where each array contains L instances of every experiment in a dataset. Then, we randomly paired elements between each array to form questions, removing repeated questions and questions where the paired experiments were identical. For datasets BH1–BH5 we set L to 10 and for DA, L was set to 5 (to keep the total number of questions roughly similar to the BH surveys). This resulted in 7792, 7842, 7788, 7825, 7804, and 8610 survey questions designed for the BH1–5 and DA datasets respectively. Overall, we hypothesized that surveys generated using this procedure would facilitate expressing hierarchical preferences over the full set of experiments and subsequent preference learning. In Section 4.1, we briefly evaluate the performance of several commonly used foundation LLM models for answering survey questions correctly and select the most accurate model to complete our surveys.
A completed survey is represented as 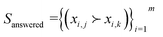 where experiment j is preferred over experiment k in question i of the survey with m total questions. Following, we leverage preference learning to infer a utility function g(x) that aligns with LLM predictions made in the survey: namely, g(xj) should be greater than g(xk) if
where experiment j is preferred over experiment k in question i of the survey with m total questions. Following, we leverage preference learning to infer a utility function g(x) that aligns with LLM predictions made in the survey: namely, g(xj) should be greater than g(xk) if 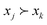 Since the LLM was prompted to prefer experiments with higher expected yield based on its chemical reasoning, we expect g(x) to correlate to the true experimental yield measured for experiments in a dataset. We follow the approach of Chu and Gharamani,8 who model g(x) as a Gaussian process and define a function to model the likelihood of observing a preference among pairs of options given their values from the utility function (assumed to contain noise). To tune hyperparameters (e.g., parameters of the kernel), they use the Laplace approximation to define an expression of the posterior density over utility functions conditioned on the data and optimize it (MAP estimate). For our work, we employ the BoTorch5 implementation of Chu and Gharamani's approach using the PairwiseGP module. Details of the model implementation are provided in the SI (section S4). Upon training, we take the mean of the GP posterior conditioned on the survey data as the utility function g(x) and to be a representation of prior chemical knowledge embedded in the LLM over the chemical parameter space.
Since the LLM was prompted to prefer experiments with higher expected yield based on its chemical reasoning, we expect g(x) to correlate to the true experimental yield measured for experiments in a dataset. We follow the approach of Chu and Gharamani,8 who model g(x) as a Gaussian process and define a function to model the likelihood of observing a preference among pairs of options given their values from the utility function (assumed to contain noise). To tune hyperparameters (e.g., parameters of the kernel), they use the Laplace approximation to define an expression of the posterior density over utility functions conditioned on the data and optimize it (MAP estimate). For our work, we employ the BoTorch5 implementation of Chu and Gharamani's approach using the PairwiseGP module. Details of the model implementation are provided in the SI (section S4). Upon training, we take the mean of the GP posterior conditioned on the survey data as the utility function g(x) and to be a representation of prior chemical knowledge embedded in the LLM over the chemical parameter space.
| απ,n(x,Dn,n) = α(x,Dn)π(g(x),p(n)) | (2) |
where Pp is pth percentile value of the set G = {g(x)∣x ∈ X}. This binary weighting to the acquisition allows optimization to focus on promising regions of the chemical space highlighted by g(x), without further biasing candidate selection with potentially noisy utility values. Our approach can also be viewed as design space pruning,14,28,46 where unpromising portions of the design space are excluded from the set X of candidate experiments. Given that our weighting/pruning approach may adversely impact optimization if g(x) is negatively correlated with f(x) (or by excluding the true maximizing argument of f(x)), we recommend setting percentile p(n) as a decaying function of BO iterations n such that p → 0 as n → ∞. In effect, this relaxes the constraint on the design space imposed by g(x) for candidate selection as more experiments are performed and the surrogate model
![[f with combining circumflex]](https://www.rsc.org/images/entities/i_char_0066_0302.gif) (x) becomes increasingly reliable. In our work, we select p(n) as a simple 2-step function; Sections 4.2 and S1 of the SI provide additional details on how parameters for p(n) were selected in our work.
(x) becomes increasingly reliable. In our work, we select p(n) as a simple 2-step function; Sections 4.2 and S1 of the SI provide additional details on how parameters for p(n) were selected in our work.3 Related works
3.1 Transfer learning via targeted modifications to the acquisition function
In one paradigm of transfer learning, prior information is leveraged to make judicious modifications to the acquisition function to accelerate Bayesian optimization in a target domain 4,17,44,47. For example, Souza et al.40 and Hvarfner et al.19 weight the acquisition function with a prior π(x) on the function's maxima, biasing the BO algorithm to focus early optimization efforts on regions of the parameter space with high probability mass. As mentioned, our approach of leveraging information encoded in g(x) follows a similar framework. In their work, however, π(x) is typically encoded as a parameterized probability function; choosing the type of function or specific parameter values to match source data or information can be non-trivial. Along with formulating a new acquisition function to incorporate π(x), Adachi et al.1 propose using preference learning to distill insights from human experts and obtain π(x). Our own experiments suggested that the quality and quantity of data collected in surveys completed by human experts was not sufficient to apply this method for our domain application. We posited that LLMs offer a promising alternative to human experts: they can answer orders of magnitude more questions in a fraction of the time and could leverage chemical information in source data to accurately answer questions. An alternative approach is presented by Aglietti et al.,2 who introduce FunBO, a framework that leverages LLM-driven program search to generate new acquisition functions expressed in code.3.2 LLM-augmented Bayesian optimization in chemical systems
A few recent works have explored how LLMs can be used to accelerate BO in chemical systems; predominantly, these works have leveraged LLMs to inform the development of the surrogate model. One strategy is to use the LLM as the surrogate model itself. For example, Ramos et al.29 show that in-context learning and specific prompting strategies (and interpretation of token probabilities) could be used to develop a regressor capable of uncertainty quantification, which they then use for BO. Another approach is to use the LLM to process some description of the chemical system/experiment and produce an embedding from which a surrogate model can be trained to make a prediction. For example, Ranković and Schwaller30 show that these LLM embeddings are competitive with (and can outperform) those obtained from more sophisticated and domain-informed pre-training procedures. Kristiadi et al.24 show that the performance of this approach can be further improved when using domain specific and fine-tuned LLMs. Furthermore, they show that parameter-efficient fine-tuning and Bayesian neural networks can offer a principled way to use the LLM as a surrogate model and allow it to further learn informative embeddings of reactions. Overall, these are promising developments in leveraging source information in LLMs to expedite BO in a target domain for chemical systems. Our work differs from these approaches in that we separate the modeling of the target information (GPR surrogate model![[f with combining circumflex]](https://www.rsc.org/images/entities/i_char_0066_0302.gif) (x)) and the source information (LLM-derived utility function g(x)), which is included at the point of defining the acquisition function. Overall, the binary weighting scheme we use to adjust the acquisition function accomplishes a similar purpose to what is presented by Liu et al.,26 who use the LLM to first pre-select which points are considered for initialization and optimization at a given iteration. We suggest that obtaining and exploiting quantitative information present in a utility function can provide finer control for experiment selection strategies. In another promising approach, Zeng et al.49 propose an LLM-enabled multi-task BO framework that uses fine-tuned LLMs to transfer knowledge across tasks via strong initialization points and show their method works well for the design of antimicrobial peptides.
(x)) and the source information (LLM-derived utility function g(x)), which is included at the point of defining the acquisition function. Overall, the binary weighting scheme we use to adjust the acquisition function accomplishes a similar purpose to what is presented by Liu et al.,26 who use the LLM to first pre-select which points are considered for initialization and optimization at a given iteration. We suggest that obtaining and exploiting quantitative information present in a utility function can provide finer control for experiment selection strategies. In another promising approach, Zeng et al.49 propose an LLM-enabled multi-task BO framework that uses fine-tuned LLMs to transfer knowledge across tasks via strong initialization points and show their method works well for the design of antimicrobial peptides.
4 Results and discussion
4.1 Survey grades and preference learning outcomes
We first evaluated multiple LLMs on their ability to distill chemistry insights based on their performance on short surveys designed for the BH1-BH5 and DA datasets. A set of 1000 question pairs was generated for each dataset by randomly pairing distinct, non-identical experiment conditions. Specifically, for each question in a given survey, the question was marked “correct” if the LLM preference (its prediction for whether experiment A or experiment B has the higher yield) aligned with the ground truth; the percentage of questions answered correctly in a given survey is defined as the accuracy. The same fixed set of 1000 question pairs for each dataset was used across all evaluated LLMs (Sonnet-3.5, Sonnet-3, haiku-3 and GPT-4) and the resulting accuracies are shown in Fig. S2 of the SI. Except for a few points that drop below 50% on the BH 4 and BH 5 datasets, we observe that the overall accuracies for all LLMs surveys exceed 50% with Sonnet-3.5 consistently outperforming the rest. Based on this result, Sonnet-3.5 was selected and used for all the subsequent analyses in this study. This suggests that the LLMs could leverage chemical knowledge trained in the foundation model to make informed decisions about which of two experiments would result in a higher yield.Next, we applied Sonnet 3.5 to complete the full length surveys (described in Section 2.2.2) constructed for each dataset. Fig. S3 in the SI gives an example of the typical reasoning provided by the LLM in answering survey questions; we observe that decisions are made from relatively simple chemical reasonings (e.g., polarity of the solvent, strength of base, stereochemistry of ligand). Overall though, we find that this is enough to achieve survey accuracies above 50% (one-tailed binomial test statistically significant for all surveys, p < 0.01), again suggesting that despite the simplicity, the chemical information embedded in the LLM is pertinent enough to help make (on average) informed decisions.
Following, for a given dataset, we use the LLM-completed survey and preference modeling to infer the utility function; we compare its output for experiments to their true measured yields. Overall, we observe a positive correlation between utility function outputs and true experimental yields for each dataset (Fig. 2), indicating that preference modeling could be used to infer a utility function that aligned with the chemically informed, LLM-completed surveys. Importantly, the outputs are not on the same scale as measured yield given that they only encode the utility of a given experiment and not the yield directly. We compared our approach to directly asking the LLM to predict the yield from descriptions of the reaction parameters and for the given reaction (i.e., zero-shot regression), which we observe gives output values that do not correlate positively to yield (Fig. S4). Overall, this suggests that the LLM survey + preference modeling approach presented herein is a promising way to distill quantitative insights from LLMs in the zero-shot setting.
Interestingly, for several of the datasets we observe distinct clusters where the utility function gives similar output values for different experiments (Fig. 2), likely reflecting the prior observation that (for datasets that show clustering) the LLM is largely leveraging simple chemical reasoning (i.e., based on 1 or 2 features) to rank one experiment over another. For datasets BH1–4 we observe three distinct clusters, dataset BH5 has two loosely defined clusters, and dataset DA shows no clustering. Notably, while the mean yield of experiments increases with the mean utility value of each cluster (giving rise to the overall positive correlation), the yield of experiments within a given cluster correlate relatively poorly to corresponding preference model outputs. We posit that for experiments within a cluster, the LLM was not able to apply sound chemical reasoning to predict why one experiment should result in a higher yield than another resulting in random predictions in surveys, and manifesting as overfit noise in the preference model. This observation motivated the formulation of the approach detailed in Section 2.2.3, where we essentially attempt to restrict the BO algorithm to query experiments found in clusters with the highest mean preference value and forgo the precise value due to the apparent noise within the cluster. We suspect that future workflows may benefit from identifying questions in the survey where the LLM is uncertain (e.g., “hallucinating”) in its response (e.g., through repeated questioning) and removing uncertain responses from the preference model training data.
4.2 BO experiments
Next, we aimed to compare the performance of the expected improvement (EI) acquisition function (eqn (1)) to the LLM utility function-modified EI acquisition function (LLM-EI) (eqn (2)) for Bayesian reaction parameter optimization in the datasets included in this study. We perform 50 independent optimization runs for each dataset and acquisition function. Optimization runs using the EI acquisition function are initialized by randomly selecting a single experiment from the given dataset. Optimization runs for the LLM-EI acquisition function are seeded by randomly selecting an experiment from the set {x∣x ∈ X ∧ π(g(x), p(n = 0)) = 1} within a given dataset. For each run, we track the maximum yield observed at a given iteration of BO.Before performing our comparison, however, the precise functional form of p(n) required specification. In general, we expect that the functional form of p(n) best suited for an optimization task will depend, in part, on topological features of the true property surface (e.g., modality in f(x)) and on the quality of the optimization prior g(x) (i.e., its correlation with the true property surface f(x)). Since neither of these are known a priori, we sought to develop a form of p(n) that performs well across several datasets (BH1–5) empirically and to subsequently evaluate its performance on additional reaction optimization datasets not used during parameter tuning (DA, AC1–3). Section S1 of the SI provides additional details of the procedure used to develop p(n) and specifies the functional form used for all optimization results presented henceforth.
Overall, we observe that the LLM-EI acquisition function either significantly outperforms or performs comparably to the EI acquisition function (with no statistically significant differences) across all measured metrics for BH1–5, which were used to tune p(n), as well as DA, which was not. Specifically, Fig. 3a shows that for BH1, BH2, BH4, and DA, LLM-EI often achieves a higher average maximum yield at a given number of experiments in the optimization campaign compared to EI, whereas datasets BH3 and BH5 exhibit comparable performance between the two acquisition functions. Furthermore, Fig. 3b shows that the mean number of experiments required to identify reaction parameters yielding the maximum outcome decreases significantly when using LLM-EI compared to EI, from 32 to 13 (59% decrease), 22 to 11 (50% decrease), and 89 to 33 (63% decrease) for datasets BH1, BH2, and DA, respectively. Due to convergence difficulties in dataset BH4, in which near-optimal yields (>99% of the maximum) are reached early but additional experiments are required to locate the absolute maximum (potentially due to GP noise or a multimodal objective landscape), the mean number of experiments needed to reach the maximum yield shows no statistically significant difference between the two acquisition functions. However, Fig. S5 shows that the mean number of experiments required to identify reaction parameters achieving 99% of the maximum attainable yield decreases significantly from 19 to 9 (53%) for dataset BH4; LLM-EI is similarly advantageous for identifying 99% of the maximum yield for BH1, BH2, and DA. For BH3 and BH5, the mean number of experiments required to identify either 99% or 100% of the maximum yield is similar between the two acquisition functions and does not show statistically significant differences. Additionally, Fig. 3c shows that seed experiments selected using LLM-EI, on average, have significantly higher outcomes than those selected at random across all datasets. This may be particularly advantageous in applications where moderate yields or property values are sufficient to advance development, rather than requiring near-maximum values. Overall, these results suggest that utility functions inferred from LLM-completed surveys can help identify promising regions of chemical space and improve the efficiency of Bayesian optimization. Furthermore, they demonstrate that the optimized p(n) performs well across BH1-5 and generalizes effectively to DA (which was not included in the optimization process) despite differences between the datasets, such as the quality of g(x) shown in Fig. 2.
4.3 Validation on newer datasets: amide Coupling
A critical concern in using LLMs for scientific tasks is data contamination—the possibility that the model performs well simply because it has seen the optimization landscape in its training data. To address this and further validate the elements of our methodology, we applied our method to three Amide Coupling datasets (AC 1–3) published in 2025,51 which are temporally disjoint from the LLM's training data.Similar to what was observed for BH1–5 and DA datasets, Fig. 4 shows that the LLM-EI acquisition function outperforms EI for the AC1–3 datasets. Specifically, Fig. 4a shows that LLM-EI identifies high-yielding conditions significantly faster than EI, resulting in a higher average best yield for a majority of the optimization campaigns across all three datasets. Fig. 4b further shows that LLM-EI significantly reduces the mean number of experiments required to reach the maximum yield across all datasets; this effect is especially pronounced for AC1 and AC2, where the mean number of experiments is reduced by more than half. Furthermore, Fig. 4c shows that LLM-EI identifies better experimental conditions than random selection for AC1 and AC2 at the start of the optimization campaign. For AC3, however, the difference in average yield between initial experiments selected with LLM-EI and those selected at random is small and statistically insignificant. Overall, these results provide strong evidence that the performance gains observed across the datasets in this study arise from chemical reasoning by the LLM rather than memorization and regurgitation of literature data. Furthermore, they indicate that the methodological choices in this study (e.g., design of survey questions, selection of a foundational LLM, and specification of p(n)) can generalize well to other reaction optimization tasks.
 | ||
| Fig. 4 Performance on “unseen” Amide Coupling datasets (AC1–3). See caption of Fig. 3 for additional details on plotted quantities in panels (a–c) and statistical testing. | ||
5 Discussion and conclusion
In this study, we presented an approach to distill and use quantitative insights from LLMs to accelerate Bayesian reaction optimization with transfer learning. Specifically, we prompted the LLM to complete surveys in which each question of a survey asks the LLM to predict which of two experiments is expected to provide the higher yield. We find that the LLM typically employs simple chemical logic to make predictions which led to (on average) correct predictions in surveys. Following, for each dataset, we used preference learning to infer a utility function g(x) which quantitatively models LLM preferences expressed in a survey. We found that the outputs of utility functions show modest correlation to the true yield measured for experiments in a given dataset; thus we interpret g(x) as an expression of prior information provided by the LLM over the chemical parameter space. Lastly, we show that the outputs of g(x) can be used to focus BO queries on promising regions of the parameter space, leading to significantly enhanced optimization in several of the datasets examined and higher experimental yields for initial BO queries.Moving forward we anticipate several avenues of investigation to improve the performance of the method presented in this work. In the first line of investigation, we posit that working to maximize the correlation between g(x) and f(x) for a given optimization task would enable the pruning algorithm to better focus optimization efforts on promising regions of the design space and further accelerate discovery. We imagine several areas of improvement in our algorithm to target this goal. First, we suspect that fine-tuning the LLM with domain-specific literature or using in-context learning (possibly identified via document retrieval systems) could be used to refine the information used to answer survey questions, improving the chemical knowledge encoded in completed surveys. In addition, we suspect that parameters surrounding the survey itself can be further optimized to better encode the chemical knowledge/reasoning of the LLM. For example, it may be advantageous to explore alternative formulations of the survey (e.g., ranking several experiments at the same time) and corresponding preference modeling strategies. In addition, it may be possible in some capacity to remove survey questions where the LLM is very uncertain about a response (e.g., repeated queries, specific prompting), which would remove noisy responses from the dataset used to infer the preference model. Lastly, in any application, it would be important to explore sensitivity to the precise wording used to elicit responses from the LLM.
In another line of work, we posit that it may be advantageous to estimate the quality of g(x), for example, by using the first few labeled experiments obtained during an optimization campaign to validate the LLM's reasoning. As the primary benefit, this could allow the user to avoid failure modes of the method, i.e., when g(x) is negatively correlated to f(x) because the LLM consistently expresses incorrect chemical reasoning in survey responses, which we do not observe but could in principle occur. In such situations, it may be necessary to remove the influence of g(x) in optimization efforts by reverting to the baseline acquisition function. In other cases, additional information about g(x) could enable informed modifications to the optimized p(n) used for pruning in this work. For example, in cases where g(x) is estimated to be close to 1, it may be advantageous to adapt p(n) such that it more aggressively prunes experiments at smaller n. Overall, however, we suspect that such informed modifications to p(n) will require characterizing how improvements depend on the combined effects of the quality of g(x), the characteristics of f(x), and the choice of p(n).
Conflicts of interest
The authors declare no competing financial interest.Data availability
Supplementary information (SI): additional information about the developement of p(n), the performances of LLMs on test surveys, an example of an LLM-generated- response to a survey question, a comparison of our method to zero-shot regression, analysis of additional optimization metrics, and details related to the surrogate and preference modeling methods used in the study. See DOI: https://doi.org/10.1039/d6dd00052e.The code in this article can be found in the GitHub repositories : https://github.com/Sanofi-Public/Pref-BO.
Acknowledgements
The authors would like to thank Jason Tedrow, Shawn Walker, and Christian Airiau from Sanofi's CMC Synthetics Platform for their valuable discussions and support of this project.References
- M. Adachi et al., Looping in the Human Collaborative and Explainable Bayesian Optimization”, arXiv, 2023, preprint arXiv:2310.17273, DOI:10.48550/arXiv.2310.17273.
- V. Aglietti et al., Funbo: Discovering acquisition functions for bayesian optimization with funsearch”, arXiv, 2024, preprint arXiv:2406.04824, DOI:10.48550/arXiv.2406.04824.
- M. Aldeghi, et al., Golem: an algorithm for robust experiment and process optimization, Chem. Sci., 2021, 12.44, 14792–14807 RSC.
- T. Bai et al., Transfer learning for Bayesian optimization: A survey”, arXiv, 2023, preprint arXiv:2302.05927, DOI:10.48550/arXiv.2302.05927.
- M. Balandat, et al., BoTorch: A Framework for Efficient Monte-Carlo Bayesian Optimization, Adv. Neural Inf. Process. Syst., 2020, 33, 21524–21538 Search PubMed.
- K. Chen et al., Chemist-X: Large language model-empowered agent for reaction condition recommendation in chemical synthesis”, arXiv, 2023, preprint arXiv:2311.10776, DOI:10.48550/arXiv.2311.10776.
- M. Christensen, et al., “Data-science driven autonomous process optimization”. en. In, Commun. Chem., 2021, 4(1), 112 CrossRef PubMed.
- W. Chu and Z. Ghahramani. “Preference learning with Gaussian processes”, in Proceedings of the 22nd international conference on Machine learning ( 2005) Search PubMed.
- C. W. Coley, W. H. Green and K. F. Jensen, Machine Learning in Computer-Aided Synthesis Planning, Acc. Chem. Res., 2018, 51.5, 1281–1289 CrossRef PubMed.
- A. F. De Almeida, R. Moreira and T. Rodrigues, Synthetic organic chemistry driven by artificial intelligence, Nat. Rev. Chem., 2019, 3(10), 589–604 CrossRef.
- Q. Dong et al., A Survey on In-context Learning”, in Proceedings of the 2024 Conference on Empirical Methods in Natural Language Processing. ed. Y. Al-Onaizan, M. Bansal, and Y.-N. Chen, Association for Computational Linguistics, Miami, Florida, USA, 2024, pp. 1107–1128 Search PubMed.
- M. Feurer et al., Practical transfer learning for bayesian optimization”, arXiv, 2018, preprint arXiv:1802.02219, DOI:10.48550/arXiv.1802.02219.
- J. Fürnkranz and E. Hüllermeier. “Preference Learning: An Introduction”, in Preference Learning. ed. J. Fürnkranz and E. Hüllermeier, Springer Berlin Heidelberg, Berlin, Heidelberg, 2011, pp. 1–17. isbn: 978-3-642-14125-6 Search PubMed.
- D. E. Graff, et al., Self-Focusing Virtual Screening with Active Design Space Pruning, J. Chem. Inf. Model., 2022, 62(16), 3854–163862 CrossRef CAS PubMed.
- J. Guo, B. Ranković and P. Schwaller, Bayesian optimization for chemical reactions, Chimia, 2023, 77(1–2), 31–38 CrossRef CAS PubMed.
- T. Guo, et al., What can large language models do in chemistry? a comprehensive benchmark on eight tasks, Adv. Neural Inf. Process. Syst., 2023, 36, 59662–59688 Search PubMed.
- R. J. Hickman, et al., Equipping data-driven experiment planning for Self-driving Laboratories with semantic memory: case studies of transfer learning in chemical reaction optimization, React. Chem. Eng., 2023, 8(9), 2284–2296 RSC.
- J. Howard and S. Ruder. “Universal Language Model Fine-tuning for Text Classification”, in Proceedings of the 56th Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers). ed. I. Gurevych and Y. Miyao, Association for Computational Linguistics, Melbourne, Australia, July 2018, pp. 328–339 Search PubMed.
- C. Hvarfner et al., πBO: Augmenting Acquisition Functions with User Beliefs for Bayesian Optimization”, arXiv, 2022, preprint arXiv:2204.11051, DOI:10.48550/arXiv.2204.11051.
- K. M. Jablonka, P. Schwaller, A. Ortega-Guerrero and B. Smit, Is GPT-3 all you need for low-data discovery in chemistry?, ChemRxiv, 2023, preprint, DOI:10.26434/chemrxiv-2023-fw8n4.
- K. M. Jablonka, et al., Leveraging large language models for predictive chemistry, Nat. Mach. Intell., 2024, 6(2), 161–169 CrossRef.
- R. Jacobs et al., Regression with large language models for materials and molecular property prediction”, arXiv, 2024, preprint arXiv:2409.06080 DOI:10.48550/arXiv.2409.06080.
- L. V. Jospin, et al., Hands-On Bayesian Neural Networks—A Tutorial for Deep Learning Users, IEEE Comput. Intell. Mag., 2022, 17.2, 29–48 Search PubMed.
- A. Kristiadi et al., A sober look at LLMs for material discovery: Are they actually good for Bayesian optimization over molecules?, arXiv, 2024, preprint arXiv:2402.05015, DOI:10.48550/arXiv.2402.05015.
- Q. Liang, et al., “Benchmarking the performance of Bayesian optimization across multiple experimental materials science domains”. en, npj Comput. Mater., 2021, 7(1), 188 CrossRef.
- T. Liu et al., Large language models to enhance bayesian optimization”, arXiv, 2024, preprint arXiv:2402.03921 DOI:10.48550/arXiv.2402.03921.
- A. M. Bran, et al., “Augmenting large language models with chemistry tools”. en, Nat. Mach. Intell., 2024, 6(5), 525–535 CrossRef PubMed.
- V. Nguyen, et al., “Filtering Bayesian optimization approach in weakly specified search space”. en. In, Knowl. Inf. Syst., 2019, 60.1, 385–413 CrossRef.
- M. C. Ramos et al., Bayesian optimization of catalysts with in-context learning”, arXiv, 2023, preprint arXiv:2304.05341, DOI:10.48550/arXiv.2304.05341.
- B. Ranković and P. Schwaller. “BoChemian: Large language model embeddings for Bayesian optimization of chemical reactions”, in NeurIPS 2023 Workshop on Adaptive Experimental Design and Active Learning in the Real World. 2023 Search PubMed.
- C. E. Rasmussen. “Gaussian Processes in Machine Learning”. in Advanced Lectures on Machine Learning: ML Summer Schools 2003, Canberra, Australia, February 2 - 14, 2003, Tübingen, Germany, August 4 - 16, 2003, Revised Lectures. ed. O. Bousquet, U. von Luxburg, and G. Rätsch, Springer Berlin Heidelberg, Berlin, Heidelberg, 2004, pp. 63–71. isbn: 978-3-540-28650-9 Search PubMed.
- Y. Ruan, et al., “An automatic end-to-end chemical synthesis development platform powered by large language models”. en, Nat. Commun., 2024, 15(1), 10160 CrossRef CAS PubMed.
- P. Schwaller, et al., Molecular Transformer: A Model for Uncertainty-Calibrated Chemical Reaction Prediction, ACS Cent. Sci., 2019, 5.9, 1572–1583 CrossRef PubMed.
- B. Shahriari, et al., Taking the Human Out of the Loop: A Review of Bayesian Optimization, Proc. IEEE, 2016, 104(1), 148–175 Search PubMed.
- B. J. Shields, et al., “Bayesian reaction optimization as a tool for chemical synthesis”. en, Nature, 2021, 590.7844, 89–96 CrossRef PubMed.
- B. J. Shields et al. Buchwald-Hartwig Dataset, https://github.com/b-shields/edbo/blob/master/experiments/data/aryl_amination/experiment_index.csv, 2021 Search PubMed.
- B. J. Shields et al. Direct-Arylation Dataset. https://github.com/b-shields/edbo/blob/master/experiments/data/direct_arylation/experiment_index.csv, 2021 Search PubMed.
- E. Shim, et al., Machine Learning Strategies for Reaction Development: Toward the Low-Data Limit, J. Chem. Inf. Model., 2023, 63(12), 3659–123668 CrossRef CAS PubMed.
- S. Singh and R. B. Sunoj, “A transfer learning protocol for chemical catalysis using a recurrent neural network adapted from natural language processing”. en. In, Digital Discovery, 2022, 1.3, 303–312 RSC.
- A. Souza et al., Bayesian optimization with a prior for the optimum”, in Joint European Conference on Machine Learning and Knowledge Discovery in Databases. Springer. 2021, pp. 265–296 Search PubMed.
- K. Swersky, J. Snoek and R. P. Adams, Multi-task bayesian optimization, Adv. Neural Inf. Process. Syst., 2013, 26(26) Search PubMed.
- P. Tighineanu et al., Transfer learning with gaussian processes for bayesian optimization”, in International conference on artificial intelligence and statistics. PMLR. 2022, pp. 6152–6181 Search PubMed.
- J. Van Herck, et al., “Assessment of fine-tuned large language models for real-world chemistry and material science applications”. en, Chem. Sci., 2025, 16(2), 670–684 RSC.
- M. Volpp et al., Meta-learning acquisition functions for transfer learning in bayesian optimization”, arXiv, 2019, preprint arXiv:1904.02642, DOI:10.48550/arXiv.1904.02642.
- K. Wang and A. W. Dowling, Bayesian optimization for chemical products and functional materials, Curr. Opin. Chem. Eng., 2022, 36, 100728 CrossRef.
- M. Wistuba, N. Schilling, and L. Schmidt-Thieme. “Hyperparameter search space pruning – A new component for sequential model-based hyperparameter optimization”, in Machine Learning and Knowledge Discovery in Databases. Lecture notes in computer science. Springer International Publishing, Cham, 2015, pp. 104–119 Search PubMed.
- M. Wistuba, N. Schilling and L. Schmidt-Thieme, Scalable gaussian process-based transfer surrogates for hyperparameter optimization, Mach. Learn., 2018, 107(1), 43–78 CrossRef.
- W. Xu, et al., Principled Bayesian optimization in collaboration with human experts, Adv. Neural Inf. Process. Syst., 2024, 37, 104091–104137 Search PubMed.
- Y. Zeng et al., Large Scale Multi-Task Bayesian Optimization with Large Language Models”, arXiv, 2025, preprint arXiv:2503.08131, DOI:10.48550/arXiv.2503.08131.
- C. Zhang et al., Amide Coupling Datasets. https://github.com/aichemeco/amide_coupling/blob/main/data/all_HTE_with_condition.csv, 2025.
- C. Zhang, et al., Intermediate knowledge enhanced the performance of the amide coupling yield prediction model, Chem. Sci., 2025, 16(26), 11809–11822 RSC.
- Z. Zhou, X. Li and R. N. Zare, Optimizing Chemical Reactions with Deep Reinforcement Learning, ACS Cent. Sci., 2017, 3.12, 1337–1344 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |