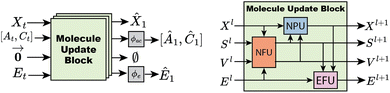Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
FlowMol3: flow matching for 3D de novo small-molecule generation
Ian
Dunn
 a and
David R.
Koes
a and
David R.
Koes
 *b
*b
aDepartment of Computational and Systems Biology, University of Pittsburgh, Pittsburgh, Pennsylvania 15260, USA. E-mail: ian.dunn@pitt.edu
bDepartment of Computational and Systems Biology, University of Pittsburgh, Pittsburgh, Pennsylvania 15260, USA. E-mail: dkoes@pitt.edu
First published on 7th April 2026
Abstract
A generative model capable of sampling realistic molecules with desired properties could accelerate chemical discovery across a wide range of applications. Toward this goal, significant effort has focused on developing models that jointly sample molecular topology and 3D structure. We present FlowMol3, an open-source, multi-modal flow matching model that advances the state of the art for all-atom, small-molecule generation. Its substantial performance gains over previous FlowMol versions are achieved without changes to the graph neural network architecture or the underlying flow matching formulation. Instead, FlowMol3's improvements arise from three architecture-agnostic techniques that incur negligible computational cost: self-conditioning, fake atoms, and train-time geometry distortion. FlowMol3 achieves nearly 100% molecular validity for drug-like molecules with explicit hydrogens, more accurately reproduces the functional group composition and geometry of its training data, and does so with an order of magnitude fewer learnable parameters than comparable methods. We hypothesize that these techniques mitigate a general pathology affecting transport-based generative models, enabling detection and correction of distribution drift during inference. Our results highlight simple, transferable strategies for improving the stability and quality of diffusion- and flow-based molecular generative models.
1 Introduction
Deep generative models that can directly sample molecular structures with desired properties have the potential to accelerate chemical discovery by reducing or eliminating the need to engage in resource-intensive, screening-based discovery paradigms. Moreover, generative models may improve chemical discovery by enabling multi-objective design of chemical matter. In pursuit of this idea, there has been recent interest in developing generative models for the design of small-molecule therapeutics,1–10 proteins,11–15 and materials.16–18In this work we focus on unconditional generation of 3D, organic, small, drug-like molecules. Building models capable of accurate 3D molecule generation is a necessary precursor to accelerating chemical discovery. From a practical perspective, if a model cannot produce valid and synthetically accessible molecules, then it would be difficult to use it for real-world applications. Moreover, if a model struggles to produce reasonable molecules, it calls into question the ability of the generative model to learn even the most basic rules of chemistry. How can a generative model fulfill complex conditioning signals, such as the formation of hydrogen-bonding networks in binder design or selectivity and toxicity constraints for drug design, if that model cannot produce realistic molecules in the first place? Any improvements made to unconditional molecular generative models will have impacts in the design and performance of conditional generative models.
Deep generative models have delivered great advances in the de novo design of molecules. Early attempts focused on generating either textual representations (SMILES strings)19–21 or 2D molecular graphs:22–25 molecular representations that exclude all information about 3D structure. Subsequent approaches were developed for 3D molecule generation using a variety of molecular representations and generative paradigms.26–30
The emergence of diffusion models31–33 significantly changed this landscape, following their success in computer vision. Hoogeboom et al.34 applied the diffusion framework to an attributed point-cloud representation of molecules and showed a substantial improvement over existing methods. Subsequent works demonstrated that using discrete diffusion for categorical data, jointly modeling bond orders, and reparameterizing the denoising objective lead to further performance gains.35–39
In parallel to the development of molecular diffusion models, flow matching emerged as a novel generative modeling framework.40–43 Flow matching generalizes diffusion models, offering simpler implementation and greater flexibility in model design. These advantages enabled Flow Matching to surpass the performance of diffusion models across various applications.9,10,14,15,17,18,44–46 Most relevant to our work here, the application of flow matching to de novo molecule generation produced dramatic improvements in the capability of 3D molecular generative models.44,45
Despite rapid progress in 3D de novo molecule design, it remains apparent that generated molecules differ substantially from “real” molecules. Molecular generative models still produce invalid molecules, unrealistic geometries, and functional group compositions that deviate significantly from their training data.45,47,48
More recent works have argued that relying on simplified, well-tested transformer-style architectures and scaling the size of the model will be essential to building powerful molecular generative models. This was the thesis of Joshi et al.49 and Reidenbach et al.46. While scaling appears to have benefits, we argue there are other pathologies that cannot be remedied by architecture choice and scale.
Both diffusion and flow matching (which we refer to collectively as transport-based generative models) prescribe methods to transport samples between two distributions qsource(x) and qtarget(x) by constructing a time-dependent process pt(x), which has the property that p0 = qsource and p1 = qtarget. Samples are drawn from the target distribution by iteratively sampling transition distributions pt+dt|t, which are implicitly parameterized by a neural network. The learned process pt will only perfectly sample the target distribution in the limit of infinite data and a perfectly trained neural network. In reality, at every integration step the model imperfectly approximates pt+dt|t, incurring drift from the desired marginal process that may accumulate through the sampling procedure. We note that directly quantifying this drift is fundamentally intractable, as it requires access to the marginal probability path or vector field—quantities that cannot be computed exactly for non-trivial data distributions (see Section S15 for a detailed mathematical treatment). Consequently, we assess drift indirectly through its effect on terminal sample quality: discrepancies between the distributions of molecular properties in generated samples versus training data.
We propose that transport-based generative models for de novo molecule generation encounter difficulties primarily due to inference-time distribution drift that degrades the performance of the denoiser, and also an inability to correct distribution drift once it has occurred. Furthermore, we demonstrate several model features that we believe impart robustness to distribution drift and substantially improve molecule quality. These additional features are self-conditioning, modeling an extra “fake” atom type that enables the model to add or remove atoms from the system, and applying distortions to molecular structure on top of the interpolant.
We present FlowMol3 (Fig. 1), a flow matching model for small molecule generation that substantially improves the state of the art in unconditional molecular generation. FlowMol3 is named so because it builds upon our previous iterations of this model.8,45 The primary difference between FlowMol2 and FlowMol3 is the addition of features that, we argue, impart robustness to inference-time distribution drift. We show that the addition of these features alone dramatically alters model performance. Moreover, these changes do not significantly impact model size and introduce minimal computational overhead. These features in combination with a bespoke geometric graph neural network architecture enable FlowMol3 to achieve state-of-the-art performance while being substantially smaller than existing models.
2 Model design
2.1 Problem setting
We represent a 3D molecule with N atoms as a fully-connected graph. Each atom is a node in the graph. Every atom has a position in space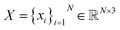 , an atom type (in this case the atomic element) A = {ai}i=1N, and a formal charge C = {ci}i=1N. Additionally, every pair of atoms has a bond order E = {eij∣i, j ∈ {1, …, N}, i ≠ j}. Atom types, charges, and bond orders are categorical variables. For brevity, we denote a molecule by the symbol g, which can be thought of as a tuple of constituent data types g = (X, A, C, E).
, an atom type (in this case the atomic element) A = {ai}i=1N, and a formal charge C = {ci}i=1N. Additionally, every pair of atoms has a bond order E = {eij∣i, j ∈ {1, …, N}, i ≠ j}. Atom types, charges, and bond orders are categorical variables. For brevity, we denote a molecule by the symbol g, which can be thought of as a tuple of constituent data types g = (X, A, C, E).
We refer to these data types as “modalities.” We seek to build a flow matching model that can jointly sample the modalities that form our molecular graph g. FlowMol3 can thus be characterized as a “multi-modal flow matching model.”
We sample atomic coordinates using Euclidean conditional flow matching40–43 and the remaining categorical modalities using discrete flow matching.50,51 In the following sections we briefly summarize the continuous and discrete flows used in FlowMol3. Then, we describe how we train one model to jointly sample interdependent modalities.
2.2 Flow matching on atomic coordinates
Flow matching40–43 is a method for learning to transport samples in a continuous space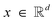 between two distributions qsource(x) and qtarget(x) by constructing a time-dependent process pt(x) having the property that p0 = qsource and p1 = qtarget and an ordinary differential equation dx = ut(x)dt which, when numerically integrated from initial positions x0 ∼ p0(x) up to time t, will produce samples distributed according to pt(x).
between two distributions qsource(x) and qtarget(x) by constructing a time-dependent process pt(x) having the property that p0 = qsource and p1 = qtarget and an ordinary differential equation dx = ut(x)dt which, when numerically integrated from initial positions x0 ∼ p0(x) up to time t, will produce samples distributed according to pt(x).
After defining a time-independent conditioning variable z ∼ p(z), the marginal process pt is constructed as an expectation over conditional probability paths:
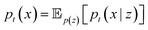 | (1) |
The conditioning variable is generally taken to be either the final value z = x1 or pairs of initial and final points z = (x0, x1). A vector field ut(x) that produces the marginal process pt(x) can be approximated by regressing onto conditional vector fields:
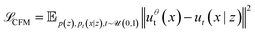 | (2) |
In FlowMol, the conditioning variable for atomic coordinate flows is paired initial and final atomic positions z = (X0, X1). The distribution of our conditioning variable, p(X0, X1), also known as the coupling distribution, is similar to the equivariant optimal transport coupling.52 Essentially, we first obtain the t = 0 atom coordinates as independent samples from a standard Gaussian 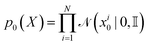 . We then perform a rigid-body alignment and distance minimizing permutation of assignments between the prior positions X0 and target positions X1. This procedure can be seen in Algorithm 1 and is discussed in detail in Section S3.
. We then perform a rigid-body alignment and distance minimizing permutation of assignments between the prior positions X0 and target positions X1. This procedure can be seen in Algorithm 1 and is discussed in detail in Section S3.
The conditional probability path for atomic coordinates is a Dirac density placed on a straight line connecting the terminal states.
| pt(X|X0,X1) = δ(X − (1 − t)X0 − tX1) | (3) |
This is equivalent to a deterministic interpolant:42
| Xt = (1 − t)X0 + tX1 | (4) |
The conditional probability path (4) is produced by the conditional vector field:41,51,53
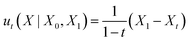 | (5) |
We apply “endpoint parameterization” as described previously.53 Rather than letting the learned vector field uθt(x) be the output of our neural network, we define our learned vector field as a function of the neural network output ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1(Xt):
1(Xt):
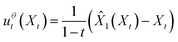 | (6) |
By substituting the true conditional vector field (5) and our chosen form of the approximate marginal vector field (6) into the conditional flow matching loss (2), we obtain the endpoint flow matching loss (7).
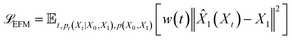 | (7) |
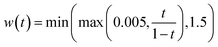 .
.
2.3 Discrete flow matching on molecular identity
Campbell et al.50 and Gat et al.51 develop a flow matching method for sequences of discrete tokens, which we refer to as Discrete Flow Matching (DFM). In this section we will briefly summarize DFM using atom types as our sequences‡ of interest, but this framework applies equivalently to the generation of atom formal charges and bond orders.In DFM, data are sequences of discrete tokens A = {ai}Ni=1 where each sequence element ai ∈ {1, 2, …, D} (i.e., the atom type of a single atom) belongs to one of D possible states. Each atom's type evolves not by an ODE but by a continuous-time Markov chain (CTMC). A CTMC is the stochastic process where the sequence alternates between resting in its current state and jumping to another discrete state.
DFM defines a CTMC on the interval t ∈ [0, 1] that transforms a sample from a simple prior distribution A0 ∼ p0(A) to a complex data distribution A1 ∼ p1(A). Jumps are governed by a probability velocity 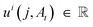 . This object describes the instantaneous flow of probability towards atom type j for atom i, given the current sequence At. This is analogous to the vector field in continuous flow matching. The marginal process pt(A) is simulated by iterative sampling of transition distributions that factorize over atoms:
. This object describes the instantaneous flow of probability towards atom type j for atom i, given the current sequence At. This is analogous to the vector field in continuous flow matching. The marginal process pt(A) is simulated by iterative sampling of transition distributions that factorize over atoms:
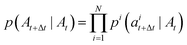 | (8) |
| pi(ait+Δt = j|At) = δ(j, ait)+ui(j,At)Δt | (9) |
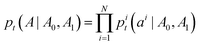 | (10) |
| pit(ai|A0,A1) = tδ(Ai1,ai) + (1 − t)δ(M,ai) | (11) |
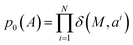 . The conditional path can be interpreted as follows: at time t, the i-th atom has probability t of being in the masked state and probability 1 − t of being in its final state ai1.
. The conditional path can be interpreted as follows: at time t, the i-th atom has probability t of being in the masked state and probability 1 − t of being in its final state ai1.
Campbell et al.50 show that a marginal process constructed as such can be sampled with the following probability velocity for j ≠ ait:
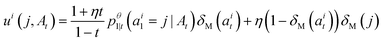 | (12) |
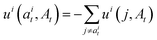 . The only “unknown” quantity in the marginal probability velocity is a probability denoiser
. The only “unknown” quantity in the marginal probability velocity is a probability denoiser 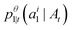 : the distribution of final states for atom i given the current sequence At. We train a neural network to approximate this distribution by minimizing the negative log-likelihood or, in practice, a standard cross-entropy loss. Note that this loss is applied only to atoms that are in the masked state at time t.
: the distribution of final states for atom i given the current sequence At. We train a neural network to approximate this distribution by minimizing the negative log-likelihood or, in practice, a standard cross-entropy loss. Note that this loss is applied only to atoms that are in the masked state at time t.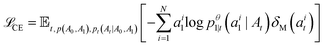 | (13) |
Substituting our choice of ui(j, At) (12) into the transition distribution (8) yields sampling dynamics that can be described simply. If ati = M, the probability of unmasking is 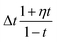 . If we do unmask, the unmasked state is selected according to our learned model
. If we do unmask, the unmasked state is selected according to our learned model 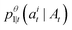 . If we are currently in an unmasked state, the probability of switching to the masked state is ηΔt.
. If we are currently in an unmasked state, the probability of switching to the masked state is ηΔt.
Hyperparameters and additional sampling techniques that we find important to DFM performance are described in Section S5.
2.4 Multi-modal flow matching
We have outlined how we can build flow matching models on the individual modalities in a molecule: continuous flow matching for atomic coordinates and DFM for atom types, charges, and bond orders. Now we describe how to train one model to sample all of these modalities simultaneously. We follow the theoretical formulation of multi-modal flow matching as developed by Campbell et al.50. We can think of the problem of sampling the real distribution of molecules, pdata(g), as sampling the joint distribution over modalities, pdata(X, A, C, E). The conditional probability path for a molecule is designed to factorize over modalities. That is, modalities are conditionally independent given pairs of initial and final molecular graphs g0 and g1.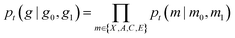 | (14) |
At training time, we obtain gt by sampling the conditional paths of each modality independently. We train one neural network fθ that takes gt as input and has separate “prediction heads” for each modality fθ(gt) = ĝ1(gt) = (![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1,Â1,Ĉ1,Ê1). The atomic coordinate prediction head
1,Â1,Ĉ1,Ê1). The atomic coordinate prediction head 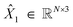 is subjected to the continuous flow matching endpoint loss (7); it is trained to approximate the final coordinates for the trajectory. The categorical outputs (Â1,Ĉ1,Ê1) contain logits over the possible discrete states for each atom/bond; these are subjected to the cross-entropy loss from DFM (13). The overall loss function for training FlowMol3 is a weighted sum of per-modality flow matching losses:
is subjected to the continuous flow matching endpoint loss (7); it is trained to approximate the final coordinates for the trajectory. The categorical outputs (Â1,Ĉ1,Ê1) contain logits over the possible discrete states for each atom/bond; these are subjected to the cross-entropy loss from DFM (13). The overall loss function for training FlowMol3 is a weighted sum of per-modality flow matching losses:
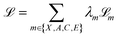 | (15) |
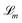 are defined as in Algorithm 1.
are defined as in Algorithm 1.
After training, the neural network outputs (![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1,Â1,Ĉ1,Ê1) parameterize a vector field on atomic coordinates and CTMCs on each of the categorical modalities that, when sampled simultaneously, will produce molecules from the target data distribution as t → 1. We provide simplified training and sampling procedures in Algorithm 1 and 2, respectively.
1,Â1,Ĉ1,Ê1) parameterize a vector field on atomic coordinates and CTMCs on each of the categorical modalities that, when sampled simultaneously, will produce molecules from the target data distribution as t → 1. We provide simplified training and sampling procedures in Algorithm 1 and 2, respectively.
2.5 Model architecture
FlowMol3 is a message-passiated as fully-connected, directed graphs where every atom is a node. The GNN operates on graphs where nodes have Cartesian positions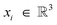 , scalar features
, scalar features 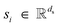 , and vector features
, and vector features  . We also model edges as having scalar features
. We also model edges as having scalar features 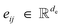 . The operations applied to node positions and vector features are SE(3)-equivariant while the operations on node scalar and edge scalar features are SE(3)-invariant. Node vector features are geometric vectors (vectors with rotation order 1) that are relative to the node position.
. The operations applied to node positions and vector features are SE(3)-equivariant while the operations on node scalar and edge scalar features are SE(3)-invariant. Node vector features are geometric vectors (vectors with rotation order 1) that are relative to the node position.
The coordinates of atoms X correspond to node positions. The initial scalar node features are produced by passing atom type, charge, and time embeddings through a shallow MLP. Similarly the initial edge features are obtained by embedding the bond order along each edge. Node vector features are initialized to zeros and are learned through the message-passing routine as functions of the relative positions between atoms.
Graph features are iteratively updated within the neural network architecture by passing through a chain of Molecule Update Blocks (MUB). After passing through MUB layers, the final positions are the predicted final positions of the molecule. The node scalar features are decoded to atom type and charge logits (Â1,Ĉ1) via shallow MLPs. Similarly, edge features are decoded to bond order logits Ê1via a shallow MLP. The model architecture is visualized in Fig. 2.
A molecule update block contains three components: a node feature update (NFU), node position update (NPU) and edge feature update (EFU). The NFU performs a message-passing graph convolution to update the scalar and vector features on each node. The NPU and EFU blocks are node and edge-wise operations, respectively.
Geometric Vector Perceptrons (GVPs)54 are used to parameterize learnable functions that operate on equivariant features, such as the message-generating functions in graph convolutions. A GVP can be thought of as a single-layer neural network that applies linear and point-wise non-linear transformation to its inputs. The difference between a GVP and a conventional feed-forward neural network is that GVPs operate on two distinct data types: scalar (rotation order 0) and vector (rotation order 1) features. These features exchange information but the operations on scalars are E(3)-invariant and the operations on vector features are E(3)-equivariant. We introduce a variant of GVP that is made SE(3)-equivariant by the addition of cross product operations. FlowMol3 is therefore capable of assigning different likelihoods to stereoisomers. Our cross product variant of GVP is described in Section S6.
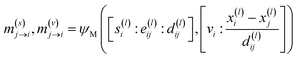 | (16) |
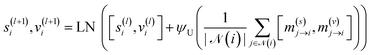 | (17) |
2.5.1.1 Node position update block. The purpose of this block is to update node positions xi. Node positions are updated:
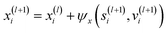 | (18) |
2.5.1.2 Edge feature update block. Edge features, e, are updated by the following equation:
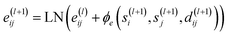 | (19) |
2.6 Enabling self-correction
Based on our experience with FlowMol2,53 which has a similar architecture to FlowMol3, we developed the hypothesis that transport-based models produce poor-quality samples due to an inability to self-correct and avoid distribution drift. An imperfect denoising model may take the sampling procedure out of distribution. Then, once slightly out of distribution, the performance of the denoising model may further degrade; causing more drift from the target distribution. To address the need for self-correction, we incorporate three additional features into FlowMol3. Through ablation experiments we show that these three features dramatically improve the quality of generated molecules.Self-conditioning is viewed by some as a recycling technique;14,56 effectively adding more layers to the network without adding additional parameters. To our knowledge, limited effort has been made to explain why self-conditioning improves performance. We offer the perspective that self-conditioning enables the model to detect and correct inaccurate predictions. At training time, the model must evaluate its own outputs and determine how to improve them; the model can only improve upon its past predictions by having some ability to find fault in them.
The denoising neural network takes in a noisy molecule gt and outputs a predicted final molecule ĝ1(gt). To implement self-conditioning, we modify our network so that it can optionally take a past prediction as an additional input, ĝ1(gt,ĝ1(gs)) where s = t − Δt.
At training time, we first predict a denoised molecule using only the current system state ĝ1(gt). In 50% of training steps we compute losses and take a gradient step on this prediction. For the other 50% of training steps, we pass the prediction back through the neural network, ĝ1(gt,ĝ1(gt)), and then compute losses on this quantity. In the latter case, the first pass through the neural network is done without keeping gradients in the computation graph, so the training-time overhead is minimal. We follow prior work in using a 50% self-conditioning rate; ablations over this proportion are provided in Section S9.
At inference time, we always pass in the network's prediction from the previous integration step along with the current step; ĝ1(gt,ĝ1(gs)) where s = t − Δt; this incurs minimal overhead compared to inference without self-conditioning.
Our network is able to “optionally” take a past prediction as input because we implement a simple self-conditioning module as a residual layer similar to Reidenbach et al.46. The self-conditioning module produces residual node and edge embeddings that quantify the difference between gt and ĝ1(gs). These embeddings are added to the node and edge embeddings of gt just before passing through the molecule update blocks.
In transport-based models, the predicted final molecule does not change very much at t > 0.4 (see Fig. 3). From observing ĝ1 inference trajectories from FlowMol2, we noticed instances where there were not enough atoms in a region of a molecule to form typical topological structures. In these cases, the atom cannot be moved to a completely different region of the molecule in the limited number of timesteps remaining. The model would instead adjust the local topological configuration to produce functional groups that, while technically valid, were unstable or rare in the data distribution. A functional group analysis showed an over-representation of heteroatom-containing functional groups such as epoxides, peroxides, and two heteroatoms separated by a single carbon (see Fig. 4 and Section S8 in the SI). We hypothesized that equipping the model with the ability to adjust the number of atoms would alleviate these issues.
At training time, a random number of “fake atoms” is added to the ground-truth molecule g1. The number of fake atoms is sampled from a uniform distribution 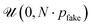 where N is the number of real atoms and pfake is a hyperparameter we set to 0.3. Each fake atom is assigned an “anchor” atom. The positions of fake atoms are Gaussian offsets from their anchor atoms
where N is the number of real atoms and pfake is a hyperparameter we set to 0.3. Each fake atom is assigned an “anchor” atom. The positions of fake atoms are Gaussian offsets from their anchor atoms 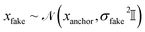 , where σfake = 1.0. Ablations over both pfake and σfake are provided in Section S12.
, where σfake = 1.0. Ablations over both pfake and σfake are provided in Section S12.
At training time, to correctly identify a fake atom, the model must essentially identify that the fake atom position cannot be re-arranged to form a realistic molecule. At inference time, if atoms enter an arrangement that is out-of-distribution, the model may recognize the system state as one where “fake atoms” are present, and thus use that mechanism to propose something that is in distribution. Including fake atoms prevents the model from truly seeing out-of-distribution structures at inference time, even if drift occurs, and imparts the model with a mechanism of correction for these instances.
Concurrently to our work, Schneuing et al.10 proposed the addition of a removable atom type for receptor-conditioned de novo design. Although their implementation differs slightly from ours (different numbers of fake atoms are added to the system, and fake atom positions are placed at the ligand center of mass), they show that this feature improves the quality of designed molecules.
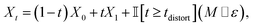 | (20) |
 is the indicator function, ⊙ is the Hadamard product, M ∈ [0,1]N is a binary mask over atoms having the property Mi ∼ Bernoulli(pdistort), and
is the indicator function, ⊙ is the Hadamard product, M ∈ [0,1]N is a binary mask over atoms having the property Mi ∼ Bernoulli(pdistort), and 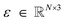 is a per-atom displacement having the property
is a per-atom displacement having the property 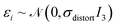 . Geometry distortion is controlled by three hyperparameters that are set to pdistort = 0.7, tdistort = 0.25, and σdistort = 0.5.
. Geometry distortion is controlled by three hyperparameters that are set to pdistort = 0.7, tdistort = 0.25, and σdistort = 0.5.
The motivation for this feature is that transport-based models produce suboptimal geometries during inference despite never observing suboptimal geometries at training time. With geometry distortion, the model should be able to propose corrections after distribution drift has occurred in order to bring the system back in-distribution.
If we choose pdistort = 1.0 and tdistort = 0, then we recover Gaussian conditional probability paths proposed in seminal flow matching works. Theoretical42 and empirical56 arguments have suggested that, when the base distribution is not Gaussian, adding Gaussian noise on top of the interpolants may improve performance by smoothing the learned vector field. SemlaFlow44 uses a conditional probability path where Gaussian noise is added to all atoms at all times, but presents no ablations to quantify the effect of this.
The distortion hyperparameters significantly affect the behavior of the generative model. Setting pdistort = 1.0 and tdistort = 0 yields validity approaching 100%, but causes a substantial increase in functional group deviation and an apparent collapse in sample diversity. We therefore chose pdistort = 0.7 and tdistort = 0.25 to balance geometric quality and validity while maintaining reasonable functional group fidelity (see Section ?? for ablations). By applying distortion only to a subset of atoms, the model observes both valid and perturbed geometries, potentially encouraging it to distinguish between them.
3 Experiments
3.1 Training data
We train models on the GEOM-Drugs dataset57 using explicit hydrogens. GEOM-Drugs contains drug-like molecules with up to 30 conformers per molecule. Each conformer has undergone a relaxation with respect to the GFN2-xTB semi-empirical quantum chemical potential58 and sits at a local minima of this potential. We use the same dataset splits as Vignac et al.35 which, to our knowledge, are purely random splits. The training dataset contains 243![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 480 unique molecules and 5
480 unique molecules and 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 741
741![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 535 conformers; the mean and median number of conformers per molecule is 23.6 and 30.0, respectively. As suggested in Nikitin et al.,47 we kekulize all molecules in the dataset and do not explicitly model aromatic bonds.
535 conformers; the mean and median number of conformers per molecule is 23.6 and 30.0, respectively. As suggested in Nikitin et al.,47 we kekulize all molecules in the dataset and do not explicitly model aromatic bonds.
3.2 Metrics
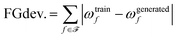 | (21) |
 is the set of functional groups analyzed, ωtrainf and ωgeneratedf are the frequency of functional group f in the training data and generated molecules, respectively. Here we define a functional group frequency as the number of instances of the functional group divided by the number of molecules in the sample.
is the set of functional groups analyzed, ωtrainf and ωgeneratedf are the frequency of functional group f in the training data and generated molecules, respectively. Here we define a functional group frequency as the number of instances of the functional group divided by the number of molecules in the sample.
In addition, we count all of the unique ring systems observed in a batch of molecules. We then record how many times each unique ring system is observed in ChEMBL,66 a database of 2.4 M bio-active compounds. We report the rate at which ring systems occur that are never observed in ChEMBL; we refer to this metric as the out-of-distribution (OOD) Ring Rate. Examples of OOD ring systems are provided in Section S7. Ring system and structural alert counting are implemented using the useful_rdkit_utils repository.67
3.3 Sampling
To generate molecules from FlowMol3 for analysis, we use evenly-spaced timesteps and Euler integration. All sampling of FlowMol models in this work is done with 250 integration steps.3.4 Ablations
To demonstrate the importance of our self-correcting features, we train four ablated versions of FlowMol3: three variants where one of the three self-correction features is removed, and a fourth variant where all three features are removed.3.5 Comparison to existing methods
We compare FlowMol3 to four diffusion models, MiDi,35 JODO,37 EQGAT-Diff,68 and Megalodon,46 and to two flow matching models, SemlaFlow44 and ADiT.49SemlaFlow is the most similar method to FlowMol3; it is also a multi-modal flow matching model with a geometric GNN-based architecture. Megalodon and ADiT opted for the well-tested diffusion-transformer architecture that can be readily scaled to large parameter counts.69 ADiT also notably discards other components that have become somewhat common: equivariance, multi-modal flows, and explicit bond modeling.
For all baseline models except ADiT we sampled 5000 molecules from the trained models using default settings provided by the authors. For ADiT we used a collection of 10 thousand molecules provided by the authors. Metrics on the sampled molecules were then computed using the same script in the FlowMol repository. Sampled molecules are split randomly into 5 subsets before computing metrics; we obtain 95% confidence intervals on the mean metric values by assuming the sample mean is normally distributed with the standard deviation in means obtained from the five subsets.
4 Results
4.1 FlowMol3 achieves state-of-the-art performance
FlowMol3 achieves substantial improvements in validity and PoseBusters validity. Nearly 100% of generated molecules are valid; to our knowledge this is the first all-atom generative model to achieve this. Interestingly FlowMol3 achieves a higher PoseBusters validity rate than that of the training data (96% generated vs. 93% training data). This may come at the cost of not completely matching the training data distribution in other areas such as functional group composition. In additional experiments, we found we could tune (pdistort, tdistort) to achieve superior functional group composition and worse for worse validity. We can even achieve 100% PB-validity but this also induces a collapse in diversity of functional group composition. These additional experiments are included in Section S10. The performance of select models on each individual test run by the PoseBusters suite is included in Section S1.Beyond validity, FlowMol3 matches the training distribution well on some higher-order chemical structure metrics: it achieves the lowest OOD ring rate (matching the training data in Table 1) and a functional group deviation that is comparable to the best-performing baseline. We note that functional group deviation can be further reduced by tuning the geometry distortion parameters (pdistort, tdistort), but this comes with trade-offs in other metrics; the corresponding ablations are reported in Section S10.
| Model | % Valid (↑) | % PB-valid (↑) | FG Dev. (↓) | OOD ring rate (↓) | Med. ΔErelax (↓) | Med. RMSD (↓) | Params (M) |
|---|---|---|---|---|---|---|---|
| Training data | 100.00 | 93.2 ± 0.1 | 0.00 | 0.05 | 0.00 | 0.00 | NaN |
| FlowMol3 | 100.0 ± 0.0 | 95.9 ± 0.2 | 0.37 ± 0.01 | 0.05 ± 0.00 | 4.50 ± 0.07 | 0.28 ± 0.01 | 6 |
| SemlaFlow | 95.5 ± 0.5 | 88.5 ± 1.3 | 0.35 ± 0.02 | 0.20 ± 0.02 | 31.92 ± 2.30 | 0.24 ± 0.03 | 40 |
| Megalodon | 94.8 ± 0.3 | 86.6 ± 0.7 | 0.39 ± 0.03 | 0.17 ± 0.00 | 3.17 ± 0.11 | 0.41 ± 0.01 | 60 |
| ADiT | 99.9 ± 0.0 | 82.7 ± 0.8 | 0.41 ± 0.03 | 0.16 ± 0.01 | 79.32 ± 1.00 | 1.30 ± 0.02 | 150 |
| EQGAT-Diff | 86.0 ± 0.9 | 77.6 ± 0.8 | 0.58 ± 0.03 | 0.28 ± 0.01 | 6.51 ± 0.16 | 0.60 ± 0.01 | 12 |
| JODO | 78.1 ± 0.9 | 65.8 ± 0.8 | 0.43 ± 0.02 | 0.22 ± 0.00 | 10.11 ± 0.19 | 0.73 ± 0.01 | 6 |
| Midi | 72.9 ± 2.5 | 59.1 ± 2.1 | 0.54 ± 0.02 | 0.33 ± 0.01 | 19.63 ± 0.65 | 0.86 ± 0.01 | 24 |
On the relaxation-based metrics (median ΔErelax and median RMSD), FlowMol3 remains competitive with the best-performing baselines. FlowMol3 achieves a median ΔErelax of 4.50 kcal mol−1, which is close to the best value from Megalodon (3.17 kcal mol; a 1.33 kcal mol−1 gap). SemlaFlow attains the lowest median RMSD from relaxation, while FlowMol3 is similar; however, SemlaFlow's median ΔErelax is 31.92 kcal mol−1 (≈7.1× higher than FlowMol3), suggesting that small geometric deviations can correspond to substantial energetic errors under GFN2-xTB relaxation.
To our knowledge, ADiT49 is the only model that achieves a similar validity level to FlowMol3, but its molecules deviate more substantially from training data in terms of the topological (0.4 FG Dev., 0.16 OOD ring rate) and energetic/geometric composition (79 kcal mol−1 ΔErelax, 83% PB-valid).
FlowMol3 is also highly parameter efficient; delivering state of the art performance while being substantially smaller than comparable models. The most comparable models in terms of performance – SemlaFlow, Megalodon, and ADiT – have 6.7, 10, and 25 times more learnable parameters, respectively.
4.2 Effects of self-correction on molecule quality
We introduce three features to FlowMol that we argue enable the model to better perform error detection and correction at inference time: self-conditioning, fake atoms, and geometry distortion. The impact of these features on molecule quality is quantified in Table 2.| Self-Cond. | Fake atoms | Distortion | % Valid (↑) | % PB-valid (↑) | FG Dev. (↓) | OOD rings (↓) | Med. ΔErelax (↓) |
|---|---|---|---|---|---|---|---|
| ✓ | ✓ | ✓ | 100.0 ± 0.0 | 95.9 ± 0.2 | 0.37 ± 0.01 | 0.05 ± 0.00 | 4.50 ± 0.07 |
| ✗ | ✓ | ✓ | 99.7 ± 0.1 | 97.0 ± 0.2 | 0.41 ± 0.02 | 0.06 ± 0.00 | 6.82 ± 0.06 |
| ✓ | ✗ | ✓ | 99.9 ± 0.0 | 94.5 ± 0.6 | 0.24 ± 0.02 | 0.12 ± 0.00 | 4.71 ± 0.10 |
| ✓ | ✓ | ✗ | 98.6 ± 0.2 | 90.6 ± 0.3 | 0.33 ± 0.02 | 0.20 ± 0.01 | 6.40 ± 0.09 |
| ✗ | ✗ | ✗ | 95.1 ± 0.4 | 78.1 ± 1.1 | 0.91 ± 0.04 | 0.32 ± 0.01 | 14.75 ± 0.34 |
While removing any one of these features results in a modest performance degradation, removing all three produces a dramatic difference. The data in Table 2 suggests there are positive interaction effects between these features. For example, removing just one feature causes (+2, −1, −3) percentage point changes to % PB-valid but removing all 3 features causes a 14 point reduction.
While each single-feature removal has mixed effects across metrics, removing all three features simultaneously leads to large degradations across all evaluated metrics (e.g., % PB-valid drops from 95.9 ± 0.2 to 78.1 ± 1.1, and median ΔErelax increases from 4.50 ± 0.07 to 14.75 ± 0.34). This gap is substantially larger than any single ablation (for % PB-valid, the single-feature changes range from a ∼ + 1.1 point increase when removing self-conditioning to a ∼ −5.3 point decrease when removing distortion), suggesting that the three features complement each other.
A potentially useful analysis of the single-feature removal ablations is to identify which feature most affects each metric. Geometry distortion has the most consistent impact on validity and chemical/topological plausibility: removing distortion reduces % Valid (from 100.0 ± 0.0 to 98.6 ± 0.2), reduces % PB-valid (to 90.6 ± 0.3), and substantially increases the OOD ring rate (to 0.20 ± 0.01). Self-conditioning most strongly affects relaxation: removing self-conditioning increases median ΔErelax (from 4.50 ± 0.07 to 6.82 ± 0.06), while having relatively small effects on validity and OOD rings. Fake atoms most strongly affect ring plausibility: removing fake atoms increases the OOD ring rate (from 0.05 ± 0.00 to 0.12 ± 0.00), with comparatively smaller effects on % Valid and ΔErelax; interestingly, FG deviation is slightly lower without fake atoms (from 0.37 ± 0.01 to 0.24 ± 0.02), indicating a trade-off between these chemistry metrics.
Overall, these ablations suggest that distortion primarily improves robustness and validity-related behavior during sampling, self-conditioning improves geometric/energetic refinement, and fake atoms help the model avoid implausible ring/topology artifacts; using all three together yields the best balanced performance.
4.3 Mechanistic interpretation of feature effects
To provide a mechanistic explanation of the effects of our self-correction enabling features, we analyze trajectories of denoiser outputs {ĝ1(gt)}t ∈ [0, 1]. After obtaining such a trajectory, we compute the change in each atom's predicted final coordinates between sequential integration steps: ‖![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1i(gt + Δt) −
1i(gt + Δt) − ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1i(gt)‖. We refer to this quantity as
1i(gt)‖. We refer to this quantity as ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1 movement.
1 movement. ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1 movement can be interpreted as how much the denoiser is updating its estimated endpoint throughout the trajectory. We average this quantity over atoms and across 100 sampled trajectories. Mean
1 movement can be interpreted as how much the denoiser is updating its estimated endpoint throughout the trajectory. We average this quantity over atoms and across 100 sampled trajectories. Mean ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1 movement trajectories for each of the ablated version of FlowMol3 are shown in Fig. 3.
1 movement trajectories for each of the ablated version of FlowMol3 are shown in Fig. 3.
We then fit a linear additive effect model on ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1 movement as a function of whether or not the three ablated features are present in the model; this allows us to isolate the effect of self-conditioning, fake atoms, and geometry distortion on
1 movement as a function of whether or not the three ablated features are present in the model; this allows us to isolate the effect of self-conditioning, fake atoms, and geometry distortion on ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1 movement as a function of integration time by inspecting the coefficients of the additive effects model. The results are shown in Fig. 3.
1 movement as a function of integration time by inspecting the coefficients of the additive effects model. The results are shown in Fig. 3.
Self-conditioning substantially reduces ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1 motion throughout trajectories. This suggests that atoms move more directly towards their final state. In each forward pass, the model is making more accurate estimates of the final molecule.
1 motion throughout trajectories. This suggests that atoms move more directly towards their final state. In each forward pass, the model is making more accurate estimates of the final molecule.
Under the linear additive effect model, it appears that both fake atoms and geometry distortion causes a significant increase in ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1 movement where t > tdistort. The magnitude of the increases in relative
1 movement where t > tdistort. The magnitude of the increases in relative ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1 movement increases with time, hovering around a 25% increase for most of the trajectory and asymptotically increasing at the very end. The increase in
1 movement increases with time, hovering around a 25% increase for most of the trajectory and asymptotically increasing at the very end. The increase in ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1 movement can be interpreted as the model making more updates/corrections to nearly complete molecules when fake atoms and geometry distortion are used.
1 movement can be interpreted as the model making more updates/corrections to nearly complete molecules when fake atoms and geometry distortion are used.
Interestingly, the bump in ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) 1 motion over the t > 0.25 regime due to fake atoms and geometry distortion is more substantial when self-conditioning is not present than when it is. An interpretation is that geometry distortion and fake atoms do indeed enable late-stage corrections but the system is simply less likely to move out of distribution in the first place when self-conditioning is implemented.
1 motion over the t > 0.25 regime due to fake atoms and geometry distortion is more substantial when self-conditioning is not present than when it is. An interpretation is that geometry distortion and fake atoms do indeed enable late-stage corrections but the system is simply less likely to move out of distribution in the first place when self-conditioning is implemented.
4.4 Functional group analysis
We find that visualizing the prevalence of individual functional groups in generated samples relative to their prevalence in the training data reveals interesting pathologies. We include a few notable examples in Fig. 4 as well as all functional groups analyzed in Section S8.Three-membered heterocycles (such as epoxides) are rare in the training data (one instance per 1220 molecules) as they are generally considered a reactive/unstable functional group. Our analysis reveals that this functional group is produced relatively frequently by some of the models evaluated. Our data suggest self-conditioning may dramatically reduce the over-representation of three-membered heterocycles. MiDi, EQGAT-Diff, and FlowMol2, which do not use self-conditioning, produce three-membered heterocycles 44×, 43×, and 56× more frequently than the training data. All but one of the models using self-conditioning produce significantly fewer instances of these functional groups. FlowMol3 produces three-membered heterocycles at only 0.1× the rate of the training data. JODO and SemlaFlow produce three-membered heterocycles at 2–3× the rate of the training data. The exception to this trend is Megalodon which implements self-conditioning yet produces three-membered heterocycles 33× more frequently than the training data; ADiT, which also uses self-conditioning, produces them at 14× the training rate.
Another functional group that is relatively rare due to instability is the occurrence of two heteroatoms separated by a carbon outside of a ring. This functional group occurs in the training data once per 289 molecules. The evaluated models produce this functional group at 8–21× the frequency of the training data except for FlowMol3, which comes in at only 1.3× the frequency of the training data. This demonstrates that there are features of FlowMol3 that uniquely enable it to match this functional group representation in a way existing models have not been able to, and that this must necessarily be caused by fake atoms or geometry distortion.
Cumarine and phenol ester provide interesting examples of how self-correcting features affect functional group representation. For cumarine, FlowMol3 comes closest to the training data rate among all evaluated models at 0.8×, though the match is not exact. For phenol esters, FlowMol3 and ADiT produce them at rates closest to the training data (1.2×), while other models produce them at 1.6–2.8× the training rate. These examples suggest that geometry distortion and fake atoms help the model better capture the distribution of certain functional groups, though there is obviously still room for improvement on the front of matching the chemistry of generated molecules to that of the training data.
Notably, FlowMol3's underproduction of three-membered heterocycles (0.1× the training rate) represents an inversion of the behavior observed in other models and in ablations of FlowMol3 with different geometry distortion parameters, which produce these functional groups at 2-3x the training rate. The same pattern—approaching or undershooting the training rate rather than overshooting—is observed for het-C-het motifs (1.3×) and cumarine (0.8×). This systematic underproduction of rare functional groups, combined with FlowMol3's % PB-valid exceeding that of the training data itself, suggests a possible mode concentration effect: the self-correcting features may cause the model to sample more heavily from high-density regions of the molecular distribution while underrepresenting low-but-non-zero density regions corresponding to rare but valid chemical motifs.
5 Discussion
FlowMol3 demonstrates state-of-the-art performance across a broad range of molecular quality metrics. These improvements arise largely from the addition of three techniques: self-conditioning, fake atoms, and geometry distortion, which we hypothesize mitigate inference-time distribution drift in transport-based generative models.Beyond increasing the validity of generated molecules, these features have measurable effects on molecular geometry, energetic states, and functional group composition. Their impact is particularly notable given that they do not alter model size or training cost. This suggests that they alleviate an existing pathology that prevented the model from making use of its available computational power.
Our analysis of denoiser trajectories supports this interpretation. Self-conditioning reduces the magnitude of atomic updates during sampling, implying that the model converges to its final predictions more directly. Geometry distortion and fake atoms, meanwhile, appear to improve the model's ability to maintain or recover from off-distribution states. Together, these features promote sampling dynamics that stay much closer to the desired marginal process.
Despite these advances, FlowMol3 does not eliminate all gaps between generated and real molecules. Functional group composition, in particular, remains imperfectly matched, and certain classes of functional groups continue to be over- or under-represented. While FlowMol3 corrects some of these discrepancies compared to prior models, others persist or are even exacerbated. Intriguingly, we observe that FlowMol3 systematically underproduces certain rare functional groups (such as three-membered heterocycles at 0.1× the training rate), inverting the overproduction observed in other models. This underproduction, combined with FlowMol3's % PB-Valid exceeding that of the training data, raises the possibility of a mode concentration effect: the self-correcting features may improve validity by biasing sampling toward high-density regions of the molecular distribution at the expense of faithfully reproducing low-density tails. Future work should investigate whether this trade-off can be tuned or whether alternative approaches can achieve both high validity and accurate representation of rare motifs. This hypothesis is further supported by the collapse in diversity and 100% PB-validity observed when tuning geometry distortion parameters to maximize validity (Section S10). Future work should investigate whether this trade-off can be tuned or whether alternative approaches can achieve both high validity and accurate representation of rare motifs.
Interestingly, FlowMol3 achieves its performance with substantially fewer parameters than ADiT and other large transformer-based models. This suggests that architectural scale alone may not be sufficient to address the specific pathologies of transport-based generative modeling. Careful study of and improvement in the underlying generative modeling framework is likely necessary as well.
Looking ahead, future research should aim to (1) develop a deeper theoretical understanding of distribution drift in transport-based models, (2) explore whether the self-correction paradigm can be formalized and incorporated at the level of the generative modeling framework itself, and (3) evaluate whether these mechanisms extend to conditional generation tasks.
6 Conclusions
We have presented FlowMol3, a flow-matching-based generative model for 3D de novo molecule generation that incorporates three design features—–self-conditioning, fake atoms, and geometry distortion—–aimed at improving robustness to inference-time distribution drift. These additions significantly improve the geometric, energetic, and topological quality of generated molecules without increasing model size or training cost.Our findings suggest that the limitations of prior transport-based generative models may stem less from insufficient model capacity and more from an inability to recover from distribution drift. By addressing this issue directly, FlowMol3 achieves performance that approaches the quality of the training distribution across widely used metrics.
Because the proposed features are architecture-agnostic and inexpensive to implement, they may be readily transferable to other models. More broadly, the principle of designing transport-based generative models to explicitly resist or recover from distribution drift may provide a general strategy for improving the reliability of transport-based generative modeling in molecular design and beyond.
All code, trained models, and evaluation scripts are available at https://github.com/dunni3/FlowMol to facilitate reproducibility and to support further work in this area.
Author contributions
Ian Dunn conceptualized the work presented, performed investigations, developed methodology, wrote software, wrote the original draft of the manuscript as well as reviewed and edited the manuscript. David Koes conceptualized the work presented, oversaw the computational experiments and methodology and software development, wrote, reviewed and edited multiple drafts of the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
All code and trained models for FlowMol3 are publicly available at https://github.com/dunni3/FlowMol. The release of the FlowMol repository associated with this publication is v3.1.0 (DOI: https://doi.org/10.5281/zenodo.19151766). The GEOM-Drugs dataset used for model training is archived at Zenodo (DOI: https://doi.org/10.5281/zenodo.19150583) and is also available at https://bits.csb.pitt.edu/files/geom_raw/; instructions for downloading and processing this data are provided in the FlowMol repository README.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00363f.
Acknowledgements
The authors would like to thank Filipp Nikitin, Gabriella Gerlach, and Rishal Aggarwal for helpful discussions in the development of this work. This work is funded through R35GM140753 from the National Institute of General Medical Sciences. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institute of General Medical Sciences or the National Institutes of Health.Notes and references
- L. Huang, T. Xu, Y. Yu, P. Zhao, X. Chen, J. Han, Z. Xie, H. Li, W. Zhong, K.-C. Wong and H. Zhang, Nat. Commun., 2024, 15, 2657 CrossRef CAS PubMed
.
- J. Guan, W. W. Qian, X. Peng, Y. Su, J. Peng and J. Ma, 3D Equivariant Diffusion for Target-Aware Molecule Generation and Affinity Prediction, arXiv, 2023, preprint, arXiv:2303.03543, DOI:10.48550/arXiv.2303.03543, https://arxiv.org/abs/2303.03543.
- A. Schneuing, Y. Du, C. Harris, A. Jamasb, I. Igashov, W. Du, T. Blundell, P. Lió, C. Gomes, M. Welling, M. Bronstein and B. Correia, Structure-based Drug Design with Equivariant Diffusion Models, arXiv, 2023, preprint, arXiv:2210.13695, DOI:10.48550/arXiv.2210.13695, https://arxiv.org/abs/2210.13695.
- X. Peng, S. Luo, J. Guan, Q. Xie, J. Peng and J. Ma, Pocket2Mol: Efficient Molecular Sampling Based on 3D Protein Pockets, arXiv, 2022, preprint, arXiv:2205.07249, DOI:10.48550/arXiv.2205.07249, https://arxiv.org/abs/2205.07249.
- M. Liu, Y. Luo, K. Uchino, K. Maruhashi and S. Ji, Generating 3D Molecules for Target Protein Binding, arXiv, 2022, preprint, arXiv:2204.09410, DOI:10.48550/arXiv.2204.09410, https://arxiv.org/abs/2204.09410.
- J. Torge, C. Harris, S. V. Mathis and P. Lio, DiffHopp: A Graph Diffusion Model for Novel Drug Design via Scaffold Hopping, arXiv, 2023, preprint, arXiv:2308.07416, DOI:10.48550/arXiv.2308.07416, https://arxiv.org/abs/2308.07416.
- I. Igashov, H. Stärk, C. Vignac, A. Schneuing, V. G. Satorras, P. Frossard, M. Welling, M. Bronstein and B. Correia, Nat. Mach. Intell., 2024, 1–11 Search PubMed
.
-
I. Dunn and D. Koes, Accelerating Inference in Molecular Diffusion Models with Latent Representations of Protein Structure, 2023, https://openreview.net/forum?id=Z4ia7s2tpV Search PubMed
.
- J. Cremer, R. Irwin, A. Tibo, J. P. Janet, S. Olsson and D.-A. Clevert, FLOWR: Flow Matching for Structure-Aware De Novo, Interaction- and Fragment-Based Ligand Generation, arXiv, 2025, preprint, arXiv:2504.10564, DOI:10.48550/arXiv.2504.10564, https://arxiv.org/abs/2504.10564.
-
A. Schneuing, I. Igashov, A. W. Dobbelstein, T. Castiglione, M. M. Bronstein and B. Correia, The Thirteenth International Conference on Learning Representations, 2025 Search PubMed
.
- J. L. Watson, D. Juergens, N. R. Bennett, B. L. Trippe, J. Yim, H. E. Eisenach, W. Ahern, A. J. Borst, R. J. Ragotte, L. F. Milles, B. I. M. Wicky, N. Hanikel, S. J. Pellock, A. Courbet, W. Sheffler, J. Wang, P. Venkatesh, I. Sappington, S. V. Torres, A. Lauko, V. De Bortoli, E. Mathieu, S. Ovchinnikov, R. Barzilay, T. S. Jaakkola, F. DiMaio, M. Baek and D. Baker, Nature, 2023, 620, 1089–1100 CrossRef CAS PubMed
.
-
N. R. Bennett, J. L. Watson, R. J. Ragotte, A. J. Borst, D. L. See, C. Weidle, R. Biswas, E. L. Shrock, P. J. Y. Leung, B. Huang, I. Goreshnik, R. Ault, K. D. Carr, B. Singer, C. Criswell, D. Vafeados, M. G. Sanchez, H. M. Kim, S. V. Torres, S. Chan and D. Baker, Atomically accurate de novo design of single-domain antibodies, 2024, https://www.biorxiv.org/content/10.1101/2024.03.14.585103v1, p. 585103 Search PubMed
.
- J. B. Ingraham, M. Baranov, Z. Costello, K. W. Barber, W. Wang, A. Ismail, V. Frappier, D. M. Lord, C. Ng-Thow-Hing, E. R. Van Vlack, S. Tie, V. Xue, S. C. Cowles, A. Leung, J. V. Rodrigues, C. L. Morales-Perez, A. M. Ayoub, R. Green, K. Puentes, F. Oplinger, N. V. Panwar, F. Obermeyer, A. R. Root, A. L. Beam, F. J. Poelwijk and G. Grigoryan, Nature, 2023, 623, 1070–1078 CrossRef CAS PubMed
.
- J. Yim, A. Campbell, A. Y. K. Foong, M. Gastegger, J. Jiménez-Luna, S. Lewis, V. G. Satorras, B. S. Veeling, R. Barzilay, T. Jaakkola and F. Noé, Fast protein backbone generation with SE(3) flow matching, arXiv, 2023, preprint, arXiv:2310.05297, DOI:10.48550/arXiv.2310.05297, https://arxiv.org/abs/2310.05297.
- A. J. Bose, T. Akhound-Sadegh, G. Huguet, K. Fatras, J. Rector-Brooks, C.-H. Liu, A. C. Nica, M. Korablyov, M. Bronstein and A. Tong, SE(3)-Stochastic Flow Matching for Protein Backbone Generation, arXiv, 2024, preprint, arXiv:2310.02391, DOI:10.48550/arXiv.2310.02391, https://arxiv.org/abs/2310.02391.
- C. Zeni, R. Pinsler, D. Zügner, A. Fowler, M. Horton, X. Fu, S. Shysheya, J. Crabbé, L. Sun, J. Smith, B. Nguyen, H. Schulz, S. Lewis, C.-W. Huang, Z. Lu, Y. Zhou, H. Yang, H. Hao, J. Li, R. Tomioka and T. Xie, MatterGen: a generative model for inorganic materials design, arXiv, 2024, preprint, arXiv:2312.03687, DOI:10.48550/arXiv.2312.03687, https://arxiv.org/abs/2312.03687.
- B. K. Miller, R. T. Q. Chen, A. Sriram and B. M. Wood, FlowMM: Generating Materials with Riemannian Flow Matching, arXiv, 2024, preprint, arXiv:2406.04713, DOI:10.48550/arXiv.2406.04713, https://arxiv.org/abs/2406.04713.
- P. Hoellmer, T. Egg, M. M. Martirossyan, E. Fuemmeler, Z. Shui, A. Gupta, P. Prakash, A. Roitberg, M. Liu, G. Karypis, M. Transtrum, R. G. Hennig, E. B. Tadmor and S. Martiniani, Open Materials Generation with Stochastic Interpolants, arXiv, 2025, preprint, arXiv:2502.02582, DOI:10.48550/arXiv.2502.02582, https://arxiv.org/abs/2502.02582.
- F. Grisoni, M. Moret, R. Lingwood and G. Schneider, J. Chem. Inf. Model., 2020, 60, 1175–1183 CrossRef CAS PubMed
.
- R. Gómez-Bombarelli, J. N. Wei, D. Duvenaud, J. M. Hernández-Lobato, B. Sánchez-Lengeling, D. Sheberla, J. Aguilera-Iparraguirre, T. D. Hirzel, R. P. Adams and A. Aspuru-Guzik, ACS Cent. Sci., 2018, 4, 268–276 CrossRef PubMed
.
- H. Dai, Y. Tian, B. Dai, S. Skiena and L. Song, Syntax-Directed Variational Autoencoder for Structured Data, arXiv, 2018, preprint, arXiv:1802.08786, DOI:10.48550/arXiv.1802.08786, https://arxiv.org/abs/1802.08786.
- W. Jin, R. Barzilay and T. Jaakkola, Junction Tree Variational Autoencoder for Molecular Graph Generation, arXiv, 2019, preprint, arXiv:1802.04364, DOI:10.48550/arXiv.1802.04364, https://arxiv.org/abs/1802.04364.
- Q. Liu, M. Allamanis, M. Brockschmidt and A. L. Gaunt, Constrained Graph Variational Autoencoders for Molecule Design, arXiv, 2019, preprint, arXiv:1805.09076, DOI:10.48550/arXiv.1805.09076, https://arxiv.org/abs/1805.09076.
- C. Shi, M. Xu, Z. Zhu, W. Zhang, M. Zhang and J. Tang, GraphAF: a Flow-based Autoregressive Model for Molecular Graph Generation, arXiv, 2020, preprint, arXiv:2001.09382, DOI:10.48550/arXiv.2001.09382, https://arxiv.org/abs/2001.09382.
- J. You, B. Liu, R. Ying, V. Pande and J. Leskovec, Graph Convolutional Policy Network for Goal-Directed Molecular Graph Generation, arXiv, 2019, preprint, arXiv:1806.02473, DOI:10.48550/arXiv.1806.02473, https://arxiv.org/abs/1806.02473.
- M. Ragoza, T. Masuda and D. R. Koes, Learning a Continuous Representation of 3D Molecular Structures with Deep Generative Models, arXiv, 2020, preprint, arXiv:2010.08687, DOI:10.48550/arXiv.2010.08687, https://arxiv.org/abs/2010.08687.
- M. Ragoza, T. Masuda and D. R. Koes, Chem. Sci., 2022, 13, 2701–2713 RSC
.
- N. W. A. Gebauer, M. Gastegger and K. T. Schütt, Symmetry-adapted generation of 3d point sets for the targeted discovery of molecules, arXiv, 2020, preprint, arXiv:1906.00957, DOI:10.48550/arXiv.1906.00957, https://arxiv.org/abs/1906.00957.
-
Y. Luo and S. Ji, An Autoregressive Flow Model for 3D Molecular Geometry Generation from Scratch, 2021, https://openreview.net/forum?id=C03Ajc-NS5W Search PubMed
.
- V. G. Satorras, E. Hoogeboom, F. B. Fuchs, I. Posner and M. Welling, E(n) Equivariant Normalizing Flows, arXiv, 2022, preprint, arXiv:2105.09016, DOI:10.48550/arXiv.2105.09016, https://arxiv.org/abs/2105.09016.
- J. Sohl-Dickstein, E. A. Weiss, N. Maheswaranathan and S. Ganguli, Deep Unsupervised Learning using Nonequilibrium Thermodynamics, arXiv, 2015, preprint, arXiv:1503.03585, DOI:10.48550/arXiv.1503.03585, https://arxiv.org/abs/1503.03585.
- J. Ho, A. Jain and P. Abbeel, Denoising Diffusion Probabilistic Models, arXiv, 2020, preprint, arXiv:2006.11239, DOI:10.48550/arXiv.2006.11239, https://arxiv.org/abs/2006.11239.
- Y. Song, J. Sohl-Dickstein, D. P. Kingma, A. Kumar, S. Ermon and B. Poole, Score-Based Generative Modeling through Stochastic Differential Equations, arXiv, 2021, preprint, arXiv:2011.13456, DOI:10.48550/arXiv.2011.13456, https://arxiv.org/abs/2011.13456.
- E. Hoogeboom, V. G. Satorras, C. Vignac and M. Welling, Equivariant Diffusion for Molecule Generation in 3D, arXiv, 2022, preprint, arXiv:2203.17003, DOI:10.48550/arXiv.2203.17003, https://arxiv.org/abs/2203.17003.
- C. Vignac, N. Osman, L. Toni and P. Frossard, MiDi: Mixed Graph and 3D Denoising Diffusion for Molecule Generation, arXiv, 2023, preprint, arXiv:2302.09048, DOI:10.48550/arXiv.2302.09048, https://arxiv.org/abs/2302.09048.
- C. Vignac, I. Krawczuk, A. Siraudin, B. Wang, V. Cevher and P. Frossard, DiGress: Discrete Denoising diffusion for graph generation, arXiv, 2023, preprint, arXiv:2209.14734, DOI:10.48550/arXiv.2209.14734, https://arxiv.org/abs/2209.14734.
- H. Huang, L. Sun, B. Du and W. Lv, Learning Joint 2D & 3D Diffusion Models for Complete Molecule Generation, arXiv, 2023, preprint, arXiv:2305.12347, DOI:10.48550/arXiv.2305.12347, https://arxiv.org/abs/2305.12347.
- X. Peng, J. Guan, Q. Liu and J. Ma, MolDiff: Addressing the Atom-Bond Inconsistency Problem in 3D Molecule Diffusion Generation, arXiv, 2023, preprint, arXiv:2305.07508, DOI:10.48550/arXiv.2305.07508, https://arxiv.org/abs/2305.07508.
- C. Hua, S. Luan, M. Xu, R. Ying, J. Fu, S. Ermon and D. Precup, MUDiff: Unified Diffusion for Complete Molecule Generation, arXiv, 2024, preprint, arXiv:2304.14621, DOI:10.48550/arXiv.2304.14621, https://arxiv.org/abs/2304.14621.
- Y. Lipman, R. T. Q. Chen, H. Ben-Hamu, M. Nickel and M. Le, Flow Matching for Generative Modeling, arXiv, 2023, preprint, arXiv:2210.02747, DOI:10.48550/arXiv.2210.02747, https://arxiv.org/abs/2210.02747.
- A. Tong, N. Malkin, G. Huguet, Y. Zhang, J. Rector-Brooks, K. Fatras, G. Wolf and Y. Bengio, Improving and generalizing flow-based generative models with minibatch optimal transport, arXiv, 2023, preprint, arXiv:2302.00482, DOI:10.48550/arXiv.2302.00482, https://arxiv.org/abs/2302.00482.
- M. S. Albergo, N. M. Boffi and E. Vanden-Eijnden, Stochastic Interpolants: A Unifying Framework for Flows and Diffusions, arXiv, 2023, preprint, arXiv:2303.08797, DOI:10.48550/arXiv.2303.08797, https://arxiv.org/abs/2303.08797.
- X. Liu, C. Gong and Q. Liu, Flow Straight and Fast: Learning to Generate and Transfer Data with Rectified Flow, arXiv, 2022, preprint, arXiv:2209.03003, DOI:10.48550/arXiv.2209.03003, https://arxiv.org/abs/2209.03003.
- R. Irwin, A. Tibo, J. P. Janet and S. Olsson, Efficient 3D Molecular Generation with Flow Matching and Scale Optimal Transport, arXiv, 2024, preprint, arXiv:2406.07266, DOI:10.48550/arXiv.2406.07266, https://arxiv.org/abs/2406.07266.
- I. Dunn and D. R. Koes, Exploring Discrete Flow Matching for 3D De Novo Molecule Generation, arXiv, 2024, preprint, arXiv:2411.16644, DOI:10.48550/arXiv.2411.16644, https://arxiv.org/abs/2411.16644.
- D. Reidenbach, F. Nikitin, O. Isayev and S. G. Paliwal, 2024.
- F. Nikitin, I. Dunn, D. R. Koes and O. Isayev, GEOM-Drugs Revisited: Toward More Chemically Accurate Benchmarks for 3D Molecule Generation, arXiv, 2025, preprint, arXiv:2505.00169, DOI:10.48550/arXiv.2505.00169, https://arxiv.org/abs/2505.00169.
- M. Buttenschoen, Y. Ziv, G. M. Morris and C. M. Deane, An evaluation of unconditional 3D molecular generation methods, arXiv, 2025, preprint, arXiv:2505.00518, DOI:10.48550/arXiv.2505.00518, https://arxiv.org/abs/2505.00518.
- C. K. Joshi, X. Fu, Y.-L. Liao, V. Gharakhanyan, B. K. Miller, A. Sriram and Z. W. Ulissi, All-atom Diffusion Transformers: Unified generative modelling of molecules and materials, arXiv, 2025, preprint, arXiv:2503.03965, DOI:10.48550/arXiv.2503.03965, https://arxiv.org/abs/2503.03965.
- A. Campbell, J. Yim, R. Barzilay, T. Rainforth and T. Jaakkola, Generative Flows on Discrete State-Spaces: Enabling Multimodal Flows with Applications to Protein Co-Design, arXiv, 2024, preprint, arXiv:2402.04997, DOI:10.48550/arXiv.2402.04997, https://arxiv.org/abs/2402.04997.
- I. Gat, T. Remez, N. Shaul, F. Kreuk, R. T. Q. Chen, G. Synnaeve, Y. Adi and Y. Lipman, Discrete Flow Matching, arXiv, 2024, preprint, arXiv:2407.15595, DOI:10.48550/arXiv.2407.15595, https://arxiv.org/abs/2407.15595.
- L. Klein, A. Krämer and F. Noé, Equivariant flow matching, arXiv, 2023, preprint, arXiv:2306.15030, DOI:10.48550/arXiv.2306.15030, https://arxiv.org/abs/2306.15030.
- I. Dunn and D. R. Koes, Mixed Continuous and Categorical Flow Matching for 3D De Novo Molecule Generation, arXiv, 2024, preprint, arXiv:2404.19739, DOI:10.48550/arXiv.2404.19739, https://arxiv.org/abs/2404.19739.
- B. Jing, S. Eismann, P. N. Soni and R. O. Dror, Equivariant Graph Neural Networks for 3D Macromolecular Structure, arXiv, 2021, preprint, arXiv:2106.03843, DOI:10.48550/arXiv.2106.03843, https://arxiv.org/abs/2106.03843.
- T. Chen, R. Zhang and G. Hinton, Analog Bits: Generating Discrete Data using Diffusion Models with Self-Conditioning, arXiv, 2023, preprint, arXiv:2208.04202, DOI:10.48550/arXiv.2208.04202, https://arxiv.org/abs/2208.04202.
- H. Stärk, B. Jing, R. Barzilay and T. Jaakkola, Harmonic Self-Conditioned Flow Matching for Multi-Ligand Docking and Binding Site Design, arXiv, 2024, preprint, arXiv:2310.05764, DOI:10.48550/arXiv.2310.05764, https://arxiv.org/abs/2310.05764.
- S. Axelrod and R. Gómez-Bombarelli, Sci. Data, 2022, 9, 185 CrossRef CAS PubMed
.
- C. Bannwarth, S. Ehlert and S. Grimme, J. Chem. Theory Comput., 2019, 15, 1652–1671 CrossRef CAS PubMed
.
- M. Buttenschoen, G. M. Morris and C. M. Deane, Chem. Sci., 2024, 15, 3130–3139 RSC
.
- S. Wang, J. Witek, G. A. Landrum and S. Riniker, J. Chem. Inf. Model., 2020, 60, 2044–2058 CrossRef CAS PubMed
.
- A. K. Rappe, C. J. Casewit, K. S. Colwell, W. A. I. Goddard and W. M. Skiff, J. Am. Chem. Soc., 1992, 114, 10024–10035 CrossRef CAS
.
- W. P. Walters, A. A. Murcko and M. A. Murcko, Curr. Opin. Chem. Biol., 1999, 3, 384–387 CrossRef CAS PubMed
.
- P. W. Walters, Generative Molecular Design Isn't As Easy As People Make It Look, 2024, https://practicalcheminformatics.blogspot.com/2024/05/generative-molecular-design-isnt-as.html Search PubMed.
- R. Brenk, A. Schipani, D. James, A. Krasowski, I. H. Gilbert, J. Frearson and P. G. Wyatt, ChemMedChem, 2008, 3, 435–444 CrossRef CAS PubMed
.
- M. Hann, B. Hudson, X. Lewell, R. Lifely, L. Miller and N. Ramsden, J. Chem. Inf. Comput. Sci., 1999, 39, 897–902 CrossRef CAS PubMed
.
- B. Zdrazil, E. Felix, F. Hunter, E. J. Manners, J. Blackshaw, S. Corbett, M. de Veij, H. Ioannidis, D. M. Lopez, J. Mosquera, M. Magarinos, N. Bosc, R. Arcila, T. Kizilören, A. Gaulton, A. Bento, M. Adasme, P. Monecke, G. Landrum and A. Leach, Nucleic Acids Res., 2024, 52, D1180–D1192 CrossRef CAS PubMed
.
- P. Walters, PatWalters/useful_rdkit_utils, 2024, https://github.com/PatWalters/useful˙rdkit˙utils, original-date: 2021-12-31T00, vol. 24, p. , p. 33Z Search PubMed.
- T. Le, J. Cremer, F. Noé, D.-A. Clevert and K. Schütt, Navigating the Design Space of Equivariant Diffusion-Based Generative Models for De Novo 3D Molecule Generation, arXiv, 2023, preprint, arXiv.2309.17296, DOI:10.48550/arXiv.2309.17296, https://arxiv.org/abs/2309.17296.
- W. Peebles and S. Xie, Scalable Diffusion Models with Transformers, arXiv, 2023, preprint, arXiv:2212.09748, DOI:10.48550/arXiv.2212.09748, https://arxiv.org/abs/2212.09748.
Footnotes |
| † Meaning that for any t ∈ [0, 1], we sample the conditional density pt(·|z) in closed-form. To sample via simulation would entail sampling the prior p0(·|z) and then performing numerical integration with the conditional velocity field ut(·|z) up to time t, which makes training prohibitively expensive. |
| ‡ The word “sequence” isn't quite correct, as it implies the existence of an inherent order. A more accurate conception of our data is as unordered or permutation-invariant sets of tokens. Nevertheless, the theory here applies seamlessly as DFM makes no explicit requirement that there be an order to sequences. |
| § Determining whether a functional group is problematic in the context of a drug discovery campaign is subjective. However, measuring the presence of functional groups can still quantify similarity to training data at higher-order levels of organization than chemical valency. |
| This journal is © The Royal Society of Chemistry 2026 |