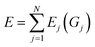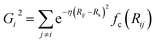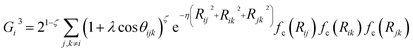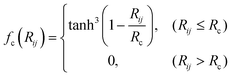Open Access Article
Open Access ArticleEnhancing high-dimensional neural network potential accuracy in OLED systems via element relabeling
Yonghwan
Yun
a,
Dongmin
Park
a,
Junyoung
Choi
a,
Dong Shin
Choi
b and
Yousung
Jung
 *ac
*ac
aDepartment of Chemical and Biological Engineering, Institute of Chemical Processes, Seoul National University, 1 Gwa-nak-ro, Gwanak-gu, Seoul 08826, Republic of Korea. E-mail: yousung.jung@snu.ac.kr
bVirtual Design Engineering Division, LG Display, 245 LG-ro, Wollongong-myeon, Paju-si, Gyeonggi-do 10845, Republic of Korea
cInstitute of Engineering Research, Seoul National University, 1 Gwa-nak-ro, Gwanak-gu, Seoul 08826, Republic of Korea
First published on 22nd March 2026
Abstract
Accurate atomistic simulations are essential for understanding organic light-emitting diode (OLED) materials in complex condensed-phase environments. However, ab initio methods are computationally demanding, whereas classical force fields often lack sufficient accuracy to represent complex molecular behavior. In this work, we present a high-dimensional neural network potential (HDNNP) framework specifically optimized for complex OLED systems. Detailed atomic-level analysis reveals that local environments, measured by intramolecular neighbor density, serve as a key factor in force prediction heterogeneity. To reduce force prediction errors arising from intrinsic molecular complexity, we introduce a conceptually simple yet effective strategy: relabeling atoms exhibiting large force errors as distinct pseudo-elements. By providing more environment-specific local descriptors, this targeted refinement improves force and total-energy prediction accuracy. Such accuracy gains lead to higher fidelity in the reproduction of key simulation observables. To demonstrate the transferability of the element-relabeling strategy, we applied it to chemically diverse benchmark systems, specifically aspirin and double-walled carbon nanotubes. Our approach substantially reduces the accuracy gap with state-of-the-art graph-based models while preserving the computational efficiency of conventional HDNNPs, enabling large scale simulations, exemplified by accurate glass transition temperature prediction.
Introduction
Organic light-emitting diode (OLED) technology has emerged as a prominent innovation in the modern display and lighting industries.1 Over the past few decades, atomistic simulations have significantly advanced the understanding of key aspects of OLED materials, including molecular configurations and charge carrier mobility.2,3 These simulations play a crucial role in bridging microscopic properties with macroscopic behavior, thus providing an efficient pathway for the design and optimization of OLED devices.4 The accuracy and relevance of insights derived from these simulations fundamentally depend on the precision and reliability of the interatomic potentials employed.5,6 Accordingly, ongoing efforts to refine these potentials are essential to enhance the predictive capability of simulation-based approaches.Classical molecular dynamics (MD) has long served as a foundational tool for investigating molecular behavior.7 However, the use of general organic force fields, such as the all-atom optimized potential for liquid simulations (OPLS-AA),8,9 in the study of OLED materials can lead to deficiencies in accurately capturing the specific interactions and electronic properties essential for OLED functionality. These limitations necessitate an extensive parameterization and validation process, which must be benchmarked against experimental or high-level computational data.2,9–11 Even with rigorous parameterization, accuracy may still be constrained by the intrinsic functional forms of the force field (e.g., bonds, angles, and torsions). On the other hand, ab initio molecular dynamics (AIMD) simulations,12 based on first principles, offer higher accuracy in describing the intricate behavior of molecular systems.13 Nevertheless, the considerable computational cost associated with AIMD often restricts its applicability to smaller systems and shorter timescales. Predicting certain properties, such as the glass transition temperature, remains prohibitive using ab initio methods.2,14
Machine learning interatomic potentials (MLIPs) offer a promising hybrid approach that integrates the accuracy of ab initio methods with the computational efficiency of classical MD.15,16 By learning the potential energy surface (PES) with quantum chemical accuracy, these potentials enable energy evaluations at speeds that are orders of magnitude faster than conventional methods.17 MLIPs can be classified based on how they encode atomic environments and propagate that information to approximate the PES. Local MLIPs—such as the Behler–Parrinello high-dimensional neural-network potential (HDNNP),18 Gaussian Approximation Potential (GAP),19 the spectral neighbor analysis potential (SNAP),20 atomic-cluster expansion (ACE)21 and moment-tensor potentials (MTP)22—encode atomic neighborhoods via compact, symmetry-invariant basis expansions. Their inherent computational efficiency enables large-scale and long-timescale molecular dynamics simulations to be performed on standard computing architectures.23 In contrast to local-descriptor models, graph-based MLIPs represent molecular structures as atomic graphs, where node embeddings are iteratively refined via message-passing neural networks (MPNNs).24 Widely adopted examples include SchNet,25 DimeNet,26 GemNet,27 SpookyNet,28 M3GNet29 and the E(3)-equivariant family comprising NequIP,30 MACE31 and Equiformer.32 Iterative message passing significantly improves predictive accuracy by capturing higher-order many-body correlations within the interaction cutoff. However, repeated neighbor communications introduce substantial computational overhead. Recent equivariant architectures that reduce the depth of message passing (e.g., two-layer higher-order MACE31,33,34) or eliminate it entirely (Allegro35) have achieved substantial computational speedups while maintaining state-of-the-art accuracy. Despite these advances, comprehensive benchmarks indicate that, on standard CPUs, graph-based MLIPs generally remain slower than local-descriptor models, and comparable throughput typically requires specialized GPU acceleration—thereby underscoring the fundamental trade-off between accuracy and efficiency.36–38
Although local-descriptor MLIPs have successfully modeled small gas-phase molecular systems,39–41 their applicability to large π-conjugated molecules, especially in condensed-phase environments, has not been thoroughly explored. In this work, we demonstrate that the predictive accuracy of local descriptor-based HDNNPs can be substantially improved for structurally complex molecular systems, such as condensed-phase OLED molecules, without compromising their intrinsic computational efficiency. Atomic-level analyses reveal a strong correlation between force prediction accuracy and the local density of intra-molecular neighboring atoms, indicating systematically higher force errors for atoms situated in densely coordinated environments. Such structural heterogeneity requires advanced approaches capable of accurately capturing subtle atomic-scale variations. To address this, atoms with large prediction errors were reassigned to distinct pseudo-element labels, allowing descriptors to better capture their unique chemical environments. This strategy overcomes a fundamental limitation of conventional local-descriptor methods, which typically assign a uniform descriptor to all atoms of a given element, irrespective of significant variations in their local chemical environments. With improved atomic force and total-energy predictions, the proposed approach reliably reproduces key structural and thermodynamic properties. Additionally, we demonstrate the generality and broad applicability of the element-relabeling strategy by validating its performance across chemically and structurally diverse molecular systems. The element-relabeled HDNNP substantially narrows the accuracy gap with state-of-the-art graph-based models, while its simpler architectural design facilitates superior parallelization and enhanced computational scalability. Such computational efficiency is particularly advantageous for accurately predicting properties that necessitate extensive sampling over nanosecond timescales and large molecular ensembles, exemplified by calculations of the glass transition temperature.
Methods
Workflow for OLED HDNNP development
The construction of an HDNNP for OLED materials involves several sequential steps. Fig. 1 provides a schematic overview of the workflow, which includes the following key processes: | ||
| Fig. 1 Schematic workflow for constructing an HDNNP for OLED materials, comprising sampling, data selection, reference calculations, model training, and validation. | ||
1. Enhanced sampling method: to efficiently explore a diverse ensemble of condensed-phase configurations, replica exchange molecular dynamics (REMD) simulations are performed using the OPLS-AA classical force field.8,9
2. Database selection: a representative subset of structures is selected using the farthest point sampling (FPS) algorithm42 to ensure configurational diversity while minimizing redundancy.
3. Ab initio molecular dynamics: short AIMD simulations (1 ps) based on density functional theory (DFT) are employed to augment the dataset and mitigate limitations inherent to configurations generated via classical MD force fields.
4. Reference forces and energies: for each configuration obtained from AIMD simulations, the reference forces and energies are recomputed using a more rigorous computational framework to ensure high-fidelity training data.
5. Neural network training: the neural network is trained to reproduce the reference forces and energies using a composite loss function that balances force and energy contributions.
6. Validation: finally, model accuracy and generalization are evaluated against independent test sets and reference calculations to validate the predictive performance of the constructed HDNNP. To further assess possible train/test leakage arising from short AIMD trajectories, we also examined a trajectory-wise split in which paired endpoint configurations from the same AIMD trajectory were always assigned to the same subset.
Training data generation for OLED HDNNPs via REMD and AIMD
The construction of a robust dataset is a fundamental step in developing any machine learning model. However, collecting training data for HDNNPs tailored to OLED molecules in the condensed phase poses unique challenges. OLED molecules are typically large, conjugated systems with many degrees of freedom, enabling them to adopt a wide range of conformations. In the amorphous condensed phase characteristic of OLED devices, this diversity is further amplified by intermolecular interactions. As a result, the configuration space that needs to be explored expands substantially, making it increasingly difficult to generate a training dataset that captures all relevant molecular conformations.OLED material configurations are typically simulated using molecular dynamics (MD), where atomistic interactions are represented through classical force fields. However, achieving high accuracy with these force fields demands extensive parameterization.2,9–11 Even with careful parameterization, the attainable accuracy is still constrained by the inherent limitations of predefined functional forms such as bonds, angles, and torsions. In contrast, AIMD computes forces and energies on-the-fly without relying on predefined potentials, making it well suited for systems with complex, dynamic interactions. While this approach captures true physical behavior more accurately, it is computationally intensive, with each ab initio calculation generating a single, often highly correlated data point that limits sampling efficiency and introduces redundancy.
In this study, we employ methods from different theoretical levels to construct a comprehensive dataset. First, we utilize REMD using the classical OPLS-AA force field (with only atomic partial charges reparameterized43) to broadly sample the configuration space of OLED materials. Also known as parallel tempering, REMD enhances phase-space sampling by enabling exchanges between replicas simulated at different temperatures.44,45 In this work, REMD simulations were conducted with LAMMPS (2 Aug 2023), and the corresponding initial molecular packings were generated with Packmol (v20.14.2). For details on the REMD methodology, refer to the Computational details section of the SI.
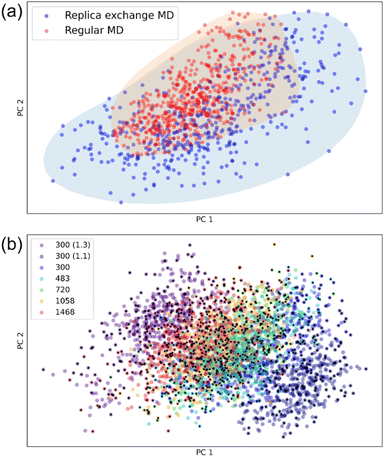
To assess the difference in sampling regions explored by regular MD and REMD, we projected the resulting structures into a low-dimensional space. The low-dimensional map provides a visually interpretable representation, simplifying the identification of a diverse set of structures.41,46–48 This approach provides an efficient way for conducting comparative analysis. Each structure is encoded by a vector of atom-centered symmetry functions (ACSF), and a two-dimensional projection of the data was obtained through principal component analysis (PCA) implemented using the ASAP48 code. As shown in Fig. 2a, REMD samples a significantly broader conformational space than regular MD. Moreover, the dihedral angle distributions of a representative OLED molecule, Bathocuproine (BCP), further confirm this enhanced diversity (Fig. S1). Fig. 2b illustrates that REMD explores a broader range of structural variations across diverse thermodynamic states. To minimize structural redundancy, we employ the FPS algorithm42 to select a representative subset of 1000 snapshots from a pool of 3500 initial configurations. Fig. S2 presents the number of snapshots selected by FPS under various temperature and density conditions.
Due to the high degrees of freedom in large OLED molecules, traditional force field parameter fitting becomes increasingly challenging. To overcome potential deficiencies in the dataset obtained from classical MD, we performed short AIMD simulations (1 ps) for each selected snapshot CP2K (v9.1). These AIMD runs capture a broad range of energy distributions during equilibration (Fig. S3) and yield diverse geometric structures, which together enrich the structural diversity of the dataset (Fig. S4). The use of structures obtained from REMD enables the parallel execution of AIMD simulations, with simulation temperatures naturally determined by the REMD protocol. However, due to the high computational cost of performing AIMD on all 1000 selected structures, we adopted a cost-effective strategy: after a 1 ps AIMD run, the reference energies and forces were recalculated for the initial and final snapshots of each trajectory using a more rigorous computational setup. Because both endpoint structures from each short AIMD trajectory were incorporated into the reference dataset, we additionally examined whether a configuration-level random split could introduce train/test correlation. To address this possibility, we performed an additional trajectory-wise split in which the paired initial and final configurations from each AIMD trajectory were always assigned to the same subset. The resulting force and energy prediction errors were comparable to those obtained from repeated random splits (Fig. S7), indicating that the reported model performance is not driven by inadvertent separation of paired AIMD endpoint structures across the training and test sets. The use of approximately 1 ps-scale structure saving is also consistent with prior dataset-generation practice; for example, Williams et al. reported sampling structures every 1 ps for reference-dataset construction and noted that longer MD trajectories help reduce temporal correlation between sampled structures.49 Further details regarding the AIMD simulations, recalculation procedure, and split validation are provided in the Computational Details section of the SI.
Behler–Parrinello symmetry functions
We employed the HDNNP framework, originally introduced by Behler and Parrinello.18 This section provides a concise overview of the functional forms and parameterization of the two primary families of ACSFs employed in this study. Further details regarding the construction and theoretical background of ACSFs are available in previous literature.50,51 Within this framework, the total energy of a system is expressed as the sum of atomic contributions:where N is the number of atoms and Gj is the descriptor that characterizes the local environment of atom j. ACSFs, which are rotationally, translationally, and permutationally invariant, serve as local structural descriptors for the input layer of the HDNNP. In this study, both radial (Gi2) and angular (Gi3) ACSFs were employed, defined as follows:
where Rij is the distance between atoms i and j, and θijk is the angle formed by atoms i, j, and k with i as the vertex. The shape of the radial symmetry functions is controlled by the parameters η and Rs, whereas the angular symmetry functions depend on η, ξ, and λ. A cutoff function fc(Rij), defined with a cutoff radius Rc is given by
An initial pool of 138 ACSFs was generated per element. A subset of ACSFs was selected to optimally represent the diverse atomic environments encountered in the training dataset. Specifically, 113, 111, and 79 functions were selected for hydrogen, carbon, and nitrogen atoms, respectively. Further details on symmetry function selection are provided in the Computational Details section of the SI. A cutoff radius of 12 Bohr (6.35 Å) was applied to define the local atomic environment.
Results and discussion
Force prediction error evaluation at the atomic level in HDNNPs
Neural network parameters were optimized using the n2p2 package (v2.2.0)52 to accurately reproduce reference forces and energies. Further details regarding the neural network training protocol are presented in the Computational Details section of the SI. Throughout this study, an ensemble of four HDNNP models with distinct random seeds was employed to ensure robustness and reduce variance in predictions for the OLED system.Due to their larger size compared to typical organic molecules, OLED molecules frequently contain atoms of the same element occupying distinctly different local atomic environments. The structural heterogeneity inherent to OLED systems presents challenges for HDNNPs based on local descriptors. Accordingly, force predictions were used to assess atomic-level accuracy rather than global accuracy. Analysis of force errors at the individual-atom level provides detailed insight into the capability of the HDNNP to capture subtle variations in forces across diverse atomic environments. This type of analysis helps pinpoint where the HDNNP systematically underperforms, thereby enabling targeted refinements. Fig. 3 shows the distribution of per-atom force root mean square errors (RMSEs) within the carbon framework of a BCP molecule. The per-atom force RMSE distribution spans a broad range (99–160 meV Å−1), reflecting pronounced site-to-site heterogeneity. Our analysis reveals that carbon atoms located in sterically congested regions, such as the central rings of the polyaromatic core, exhibit markedly higher force RMSEs. Conversely, terminal or peripheral atoms situated within sparsely populated environments exhibit comparatively modest errors. In a recent study,53 specific atomic environments were found to exhibit significantly elevated force errors. Notably, this heterogeneity was consistently observed across multiple MLIP frameworks, irrespective of whether the underlying models employed kernel-based methods or neural networks.
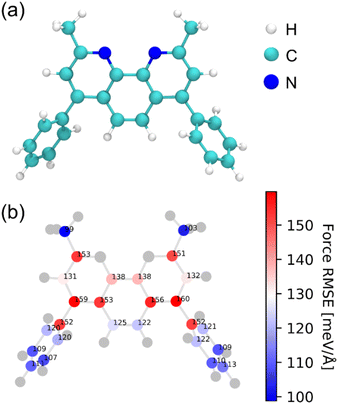 | ||
| Fig. 3 (a) Chemical structure of the BCP molecule. (b) Spatial distribution of per-atom force RMSEs for carbon atoms. Nitrogen and hydrogen atoms are shown in gray for clarity. | ||
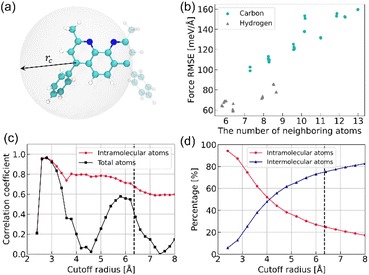
Local structure–error correlation in OLED molecules
The present analysis examines the correlation between force RMSE and the number of atoms within a defined cutoff sphere (Fig. 4a). This correlation analysis aims to uncover the underlying factors responsible for the observed variations in force RMSE distributions. At a 3 Å cutoff radius, RMSE values increase monotonically with the number of neighboring atoms for both carbon and hydrogen atoms (Fig. 4b), suggesting a strong positive correlation between local atomic density and force prediction error. This observation reveals that local structural complexity, as measured by neighbor density, is a key factor in force prediction errors, with highly coordinated atomic sites exhibiting higher RMSE values. To quantify this relationship more systematically, Pearson correlation coefficients were computed as a function of cutoff distance, with specific focus on carbon atoms (Fig. 4c). In this analysis, atoms within each cutoff sphere were classified into two distinct groups: intramolecular atoms (belonging to the same molecule) and total atoms (including both intra- and intermolecular neighbors). This classification enables a more detailed assessment of how each neighbor group contributes to force prediction accuracy. As shown in Fig. 4c, both correlation curves reach their maximum near 3 Å, indicating that short-range neighborhoods are most informative for predicting force errors. Notably, intramolecular neighbor density exhibits a consistently stronger correlation with force errors compared to the total atomic density within the same cutoff radius. This stronger correlation observed for intramolecular neighbors likely results from inherently stronger covalent interactions compared to weaker intermolecular forces. These findings imply that, even in the condensed phase, force-prediction heterogeneity is primarily determined by the intrinsic complexity of molecular structure. These observations further suggest that simple structural descriptors, such as intramolecular coordination number or local neighbor density, may serve as practical predictive criteria for identifying candidate atoms for relabeling. In this study, however, we retained the diagnostic error-based procedure, as it directly identifies the atomic environments associated with the largest residual errors in the baseline model and thus provides the most conservative basis for atom selection. Fig. 4d presents the relative proportions of intra- and intermolecular neighbors as a function of cutoff distance, revealing the expected decay of intramolecular influence at longer distances.Element relabelling for capturing structural complexity in HDNNPs
Addressing the structural-complexity-driven heterogeneity in force-prediction errors requires descriptors that accurately capture subtle variations within local atomic environments. However, modifying the descriptor architecture on a per-atom basis is infeasible, as the standard neural network imposes a fixed functional form and uniform parameterization across all atoms of a given element. Although each descriptor value changes with geometry, the use of a fixed functional form may inadequately represent the diversity of local atomic environments. A straightforward yet effective approach is to reclassify as a pseudo-element “X,” thereby enabling the design of descriptors that capture environment-specific features. As illustrated in Fig. 5, relabeling specific carbon atoms as pseudo-element X requires modifications to their symmetry function vectors and corresponding atomic neural network parameters. Specifically, introducing pseudo-element X requires defining an additional atomic neural network and constructing symmetry functions to explicitly encode interactions between X and hydrogen, carbon, and nitrogen atoms. In practice, this approach involves simply relabeling selected atoms in the dataset by changing their elemental labels in the position and force datasets and incorporating the newly defined symmetry functions. This method was applied to the ten carbon atoms within the BCP molecule that exhibited the highest force RMSE values. These atoms are characterized by a high density of intramolecular neighbors and a lack of covalent bonds to hydrogen (Fig. 3b), representing structurally challenging environments for force prediction. Here, the pseudo-element X is not intended to represent a new chemical species, but rather a label for a distinct local-environment class among atoms that remain chemically carbon. This relabeling therefore provides a separate descriptor-to-network mapping for these crowded carbon environments, rather than forcing all carbon atoms into a single shared representation. Initially, standard sets of symmetry functions were utilized for each relevant elemental interaction. To mitigate the combinatorial increase in feature dimensionality resulting from additional element types, symmetry functions exhibiting negligible variability across the dataset were eliminated. Specifically, symmetry functions with dataset ranges (maximum–minimum values) below 0.01 were discarded. Further methodological details are provided in the Computational Details section of the SI.Data-driven element-relabeling improves HDNNP accuracy
Fig. 6 shows that element relabeling markedly improves atomic force predictions: the mean force RMSE for carbon reduced from 129.5 to 98.0 meV Å−1, while its standard deviation reduced from 33.5 to 23.4 meV Å−1. The reduction in RMSE variability reflects decreased heterogeneity in force predictions, indicating that the element-relabeled HDNNP attains more uniform accuracy across chemically diverse atomic environments. Such improved uniformity is essential for ensuring energy conservation and maintaining stability in long-timescale MD simulations. These gains are transferable, lowering the mean force RMSE for hydrogen (69.3 → 53.7 meV Å−1) and nitrogen (137.4 → 97.0 meV Å−1) even though only carbon atoms were relabeled. This transferability arises because relabeling carbon atoms as the pseudo-element X modifies not only the descriptors of the relabeled sites themselves, but also those of neighboring atoms through the introduction of X-containing interaction channels in the ACSF representation. Consequently, hydrogen and nitrogen atoms located near relabeled carbon sites are described by more environment-specific descriptors, and this improved local representation is further reflected in the force predictions because atomic forces are obtained as derivatives of the total HDNNP energy. Because the HDNNP represents the total potential energy as an explicit sum of atom-centered contributions, improvements in per-atom force predictions naturally propagate to the global scale. Accordingly, the total energy RMSE decreased from 419.5 to 293.6 meV, confirming that the relabeled model yields more accurate and internally consistent energetics. This targeted, data-driven relabeling strategy, guided by atomic-level force-error analysis, proved more effective than conventional heuristic approaches, such as differentiation based on hybridization states (e.g., sp2vs. sp3), typically employed in force-field parameterizations (Fig. S5). These results underscore the advantages of employing data-driven, error-based strategies to systematically enhance the predictive accuracy of HDNNPs.Validation beyond force and energy: RDF and PMF comparisons
While improvements in atomic force and total-energy predictions are beneficial, such refinements alone may not ensure overall fidelity in MD simulations. Thus, rigorous validation of MLIPs should extend beyond force and energy accuracy to include their ability to accurately reproduce physically relevant simulation observables.54,55 Such comprehensive validation requires assessing the accuracy of simulation observables predicted by HDNNPs with respect to AIMD reference data. To this end, we conducted a comparative analysis of structural and thermodynamic properties derived from 60 ps trajectories generated by AIMD, baseline HDNNP, and element-relabeled HDNNP. Specifically, we employed a quantitative evaluation framework that measures deviations from AIMD reference trajectories using percentage-based accuracy metrics.56 Structural accuracy was evaluated by computing radial distribution functions (RDFs) with respect to the OLED molecule's center of mass, while molecular flexibility was characterized through potentials of mean force (PMFs) as a function of the dihedral angle φ. The results of these validation analyses are summarized in Fig. 7. Our analysis demonstrates that the element-relabeling strategy enhances the reproduction of key simulation-derived properties, such as RDFs and PMFs, in addition to improving the accuracy of both atomic force and total energy predictions.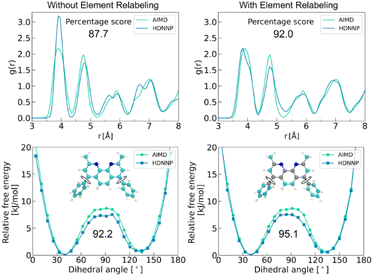 | ||
| Fig. 7 RDFs and PMFs obtained from MD trajectories using the baseline HDNNP (left) and the element-relabeled HDNNP (right). For quantitative assessment, these properties were converted to numerical scores using established difference metrics (see ref. 46 for details). The resulting evaluation scores are shown in the figure for direct comparison. | ||
Application to other systems
To evaluate the transferability and robustness of the element-relabeling strategy, we applied it to two chemically and structurally distinct systems: aspirin (MD17 dataset57) and double-walled carbon nanotube (MD22 dataset58). This extension demonstrates that our approach is not limited to specific molecular structures but can be generalized across diverse chemical environments and molecular systems. The accuracy improvements for these two systems are presented in Fig. 8. To benchmark our approach against graph-based models, all prediction errors were recalculated in terms of mean absolute error (MAE) for direct comparison with EGraFFBench results.55 For aspirin, the baseline HDNNP yielded energy and force MAEs of 12.31 meV and 31.45 meV Å−1, respectively. Analysis of the force error distribution identified the three carbon atoms with high MAEs, which were subsequently relabeled. Application of the element-relabeling strategy lowered the MAEs, 12.31 → 6.61 meV for energy and 31.45 → 19.79 meV Å−1 for force. When compared with the median error values of eight equivariant GNNs in EGraFFBench (7.4 meV and 13.95 meV Å−1, respectively55), the relabeled HDNNP achieved comparable accuracy, indicating that element relabeling markedly enhances the performance of conventional HDNNPs and narrows the performance gap relative to state-of-the-art GNNs (Table 1). Similarly, for the double-walled carbon nanotube, energy and force MAEs were reduced from 146.0 meV and 88.93 meV Å−1 to 113.5 meV and 61.76 meV Å−1, respectively, although a different strategy was used. We first confirmed that a force-error-based selection also reduces the error in the nanotube system, indicating that it provides a consistent and effective starting criterion across the systems examined here. However, for the highly symmetric DWCNT system, relabeling the carbon atoms in the outer shell proved even more effective, as it provides a cleaner structural partition of distinct local environments (see Fig. S8). The consistent accuracy improvements observed across chemically and structurally diverse systems indicate that the element-relabeling strategy offers a generalizable framework for systematically improving the performance of local-descriptor-based HDNNPs.| NequIP | Allegro | BOTNet | MACE | Equiformer | TorchMDNet | PaiNN | DimeNET++ | Baseline HDNNP | Relabeled HDNNP | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| E | F | E | F | E | F | E | F | E | F | E | F | E | F | E | F | E | F | E | F |
| 6.84 | 13.89 | 5.00 | 9.17 | 7.99 | 14.06 | 8.53 | 14.01 | 6.15 | 15.29 | 5.33 | 8.97 | 41.49 | 12.41 | 133.24 | 22.07 | 12.31 | 31.45 | 6.61 | 19.79 |
Local descriptors vs. graph-based models: trade-offs in accuracy and efficiency
Graph-based MLIPs achieve remarkable accuracy by explicitly propagating information across neighboring atomic environments.24 However, this often entails computational overhead from message passing26,27 and from equivariant tensor-product contractions.28,59 Message-passing architectures employing angular/triplet interactions (e.g., DimeNet26 and GemNet27) scale nearly quadratically with the number of neighbors, since each central atom forms k(k − 1)/2 triplets, yielding an overall O(Nk2) complexity (N: total atoms; k: average neighbors within cutoff rc.) that limits their applicability to large condensed-phase systems. Although Allegro reduces some overhead by enforcing strict cutoffs and eliminating explicit message passing, it remains substantially slower than descriptor-based HDNNPs on equivalent CPU hardware.35,36In contrast, the local descriptor-based HDNNP is inherently more computationally efficient. By leveraging ACSFs, the framework enables efficient parallelization and exhibits near-linear scaling with system size, facilitating simulations at the nanosecond timescale or at the billion-atom scale.60,61 The present element-relabeling strategy should be situated within this local-descriptor paradigm rather than among hybrid or equivariant architectures. Specifically, it does not introduce message passing or equivariant tensor operations, but instead increases representational flexibility at the descriptor level by assigning distinct pseudo-element labels to atoms associated with systematically underrepresented local environments. In this sense, the method may be viewed as a descriptor-level refinement of the HDNNP framework that improves the resolution of local structural heterogeneity while retaining the practical efficiency advantages of descriptor-based models. We further mitigated ACSF combinatorial growth by eliminating symmetry functions with negligible variability and retaining informative descriptors. Our approach reduces the accuracy gap with state-of-the-art graph-based models while maintaining high computational efficiency, which is particularly advantageous for simulations over extended timescales and large system sizes. To make this point more explicit, we summarize the descriptor growth and the effect of pruning in Table S3 of the SI; in the present case, the total number of symmetry functions increases from 414 to 928 upon introducing X, but pruning reduces this to 619, indicating that a substantial fraction of the formally introduced channels is not informative in practice. At the same time, this mitigation does not remove the intrinsic combinatorial growth of ACSF-based descriptors.
Leveraging the efficiency of local descriptors for large-scale molecular simulations
To evaluate the practicality and scalability of the element-relabeling HDNNP in OLED applications, we applied it to predict the glass transition temperature (Tg). Predicting Tg of amorphous OLED materials represents a demanding task, requiring nanosecond-scale sampling and extensive molecular ensembles. Previous studies have primarily relied on classical MD simulations to meet these computational demands.2,43 The present approach aims to deliver DFT-level accuracy by leveraging the computational efficiency of HDNNPs. The initial dataset consisted exclusively of high-density configurations (1.1, 1.2, and 1.3 g cm−3). To broaden configurational diversity required for Tg prediction, additional low-density (0.7 and 0.9 g cm−3) and high-temperature (825 and 1202 K) structures were incorporated. Each density–temperature combination contributed 100 structural snapshots, resulting in a total of 400 configurations. Glass transition temperatures were predicted using the computational protocol established by Lin et al.62 The simulation protocol began with classical equilibration of 128 BCP molecules (6144 atoms), followed by a 2.5 ns NPT equilibration at 800 K and 1 atm performed using the element-relabeled HDNNP. The equilibrated system was subsequently subjected to linear cooling from 800 K to 0 K at a rate of 50 K ns−1, during which density and temperature were continuously recorded. To ensure robust Tg estimation, linear regressions were performed on density–temperature data across three temperature intervals (200, 250, and 300 K). Despite pronounced density fluctuations attributable to the small system size (128 molecules), high coefficients of determination (R2 > 0.9) confirmed the robustness of the linear regressions within the selected temperature intervals (Fig. S6). Although the estimated glass transition temperatures varied slightly depending on the regression interval (200, 250, and 300 K), the overall trends remained consistent. The resulting Tg values were 116.6 ± 12.7 °C for HDNNP-MD and 152.3 ± 7.2 °C for classical MD, reported as the mean and standard deviation of estimates obtained from three distinct temperature intervals (200, 250, and 300 K). The Tg predicted by HDNNP-MD showed closer agreement with the experimental value (89 °C),63 indicating improved accuracy over classical force fields. Throughout the cooling process, HDNNP-MD trajectories consistently exhibited higher bulk densities by 0.02 to 0.25 g cm−3 than those obtained from classical MD simulations. The higher densities observed in HDNNP-MD trajectories suggest tighter molecular packing relative to classical MD. One possible contributor is differences in the treatment of short-range exchange (Pauli) repulsion: HDNNP inherits this from the underlying electronic-structure data, whereas many classical force fields rely on Lennard-Jones-type repulsion, whose short-range behavior can be inadequate for some systems.64 These findings demonstrate that the element-relabeled HDNNP can be effectively utilized for predicting properties derived from large-scale MD simulations, with improved accuracy over classical potentials (Fig. 9).Conclusions
In this study, we presented a HDNNP framework specifically designed to address the computational challenges associated with structurally complex OLED systems. Atomic force analysis reveals that force errors are strongly correlated with intramolecular neighbor density, increasing consistently with higher coordination numbers. These insights motivated us to implement an element-relabeling strategy, relabeling high-error atoms as pseudo-elements to provide environment-specific descriptors. The accuracy of the HDNNP was improved using an element-relabeling strategy designed to overcome the limitations inherent to single-descriptor approaches when applied to diverse local environments. Relative to conventional approaches, the developed HDNNP framework demonstrated superior accuracy in predicting atomic forces and total-energy and in reproducing essential structural and thermodynamic characteristics of OLED materials. Application across chemically and structurally diverse systems confirmed the generality of the proposed framework. This strategy preserves the computational efficiency of HDNNPs while achieving accuracy comparable to graph-based MLIPs, enabling large-scale materials simulations such as glass transition temperature prediction. This represents a significant advancement in the modeling of OLED materials. Nonetheless, one important limitation should be noted. Although descriptor pruning substantially reduces the number of formally introduced symmetry functions, the present approach does not eliminate the intrinsic combinatorial growth of ACSF-based descriptors. As additional relabeled classes and elemental species are introduced, the resulting increase in descriptor dimensionality, memory demand, and training cost will become increasingly significant. The method is therefore best regarded as a targeted and practically useful refinement strategy when only a limited number of distinct environment classes require separate treatment.Author contributions
Y. Y. and Y. J. conceptualized the project and acquired funding. Y. Y. curated the dataset, developed the neural network potential models, conducted molecular dynamics simulations, and analysed the results. Y. Y. also drafted the initial manuscript and generated the figures. D. P., J. C., and D. C. reviewed and edited the manuscript. Y. J. provided critical revisions and supervised the project.Conflicts of interest
There are no conflicts to declare.Data availability
The data and code relevant to this study are deposited in Zenodo [DOI: https://doi.org/10.5281/zenodo.19078768]. The deposit includes the raw HDNNP reference datasets in the n2p2 input.data format; the explicit train/test split files corresponding to each trained model; the deposited input.nn files specifying the random_seed values used for the internal n2p2 split; the trained model artifacts (scaling.data and trained weight files) for the OLED reference and relabeled committees and for the aspirin and DWCNT reference/relabeled models; the FPS selection script; additional dataset-preparation scripts; and representative CP2K input files (aimd.inp and endpoint_recalculation.inp). The external benchmark datasets used in this study are publicly available. The aspirin benchmark was taken from the revised MD17 (rMD17) dataset on Figshare (DOI: https://doi.org/10.6084/m9.figshare.12672038.v3), and the double-walled carbon nanotube benchmark was obtained from the MD22 dataset portal hosted by sGDML (accessed March 10, 2026).Supplementary information (SI): additional methodological details and SI figures. See DOI: https://doi.org/10.1039/d5dd00579e.
Acknowledgements
This work was supported by the New Faculty Startup Fund from Seoul National University, LG Display (C2021004543), and IITP grant (RS-2021-II211343).Notes and references
- B. Geffroy, P. Le Roy and C. Prat, Polym. Int., 2006, 55(6), 572–582 CrossRef CAS
.
- A. Mondal, L. Paterson, J. Cho, K. Lin, B. van der Zee, G. Wetzelaer, A. Stankevych, A. Vakhnin, J.-J. Kim and A. Kadashchuk, Chem. Phys. Rev., 2021, 2(3), 031304 CrossRef
.
- L. Paterson, F. May and D. Andrienko, J. Appl. Phys., 2020, 128(16), 160901 CrossRef CAS
.
- Y. Qiu, L. Wang, L. Chen, Y. Tian, Z. Zhou and J. Wu, Chem. Sci., 2025, 16, 17450–17460 RSC
.
- F. Vitalini, A. Mey, F. Noé and B. G. Keller, J. Chem. Phys., 2015, 142(8), 084101 CrossRef CAS PubMed
.
- T. Lewis-Atwell, P. Townsend and M. N. Grayson, Tetrahedron, 2021, 79, 131865 CrossRef CAS
.
- S. Hollingsworth and R. O. Dror, Neuron, 2018, 99(6), 1129–1143 CrossRef CAS PubMed
.
- W. Jorgensen, D. S. Maxwell and J. Tirado-Rives, J. Am. Chem. Soc., 1996, 118(45), 11225–11236 CrossRef CAS
.
- G. Kaminski, R. A. Friesner, J. Tirado-Rives and W. L. Jorgensen, J. Phys. Chem. B, 2001, 105(28), 6474–6487 CrossRef CAS
.
- M. Moral, W. Son, J. Sancho-Garcia, Y. Olivier and L. Muccioli, J. Chem. Theory Comput., 2015, 11(7), 3383–3392 CrossRef CAS PubMed
.
- C. Tonnelé, M. Stroet, B. Caron, A. Clulow, R. C. Nagiri, A. K. Malde, P. L. Burn, I. R. Gentle, A. E. Mark and B. J. Powell, Angew. Chem., Int. Ed., 2017, 56(29), 8402–8406 CrossRef PubMed
.
-
D. Marx and J. Hutter, Ab Initio Molecular Dynamics: Basic Theory and Advanced Methods, Cambridge University Press, 2009, pp. 1–520 Search PubMed
.
- Y. Fu, L. Bernasconi and P. Liu, J. Am. Chem. Soc., 2021, 143(3), 1577–1589 CrossRef CAS PubMed
.
- Y. Qiu, X. Zhang, Y. Tian and Z. Zhou, Chin. J. Struct. Chem., 2023, 42, 100118 CAS
.
- P. Friederich, F. Häse, J. Proppe and A. Aspuru-Guzik, Nat. Mater., 2021, 20(6), 750–761 CrossRef CAS PubMed
.
- O. Unke, S. Chmiela, H. E. Sauceda, M. Gastegger, I. Poltavsky, K. T. Schütt, A. Tkatchenko and K.-R. Müller, Chem. Rev., 2021, 121(16), 10142–10186 CrossRef CAS PubMed
.
- V. Deringer, M. A. Caro and G. Csányi, Adv. Mater., 2019, 31(46), 1902765 CrossRef CAS PubMed
.
- J. Behler and M. Parrinello, Phys. Rev. Lett., 2007, 98(14), 146401 CrossRef PubMed
.
- A. Bartók, M. C. Payne, R. Kondor and G. Csányi, Phys. Rev. Lett., 2010, 104(13), 136403 CrossRef PubMed
.
- A. Thompson, L. P. Swiler, C. R. Trott, S. M. Foiles and G. J. Tucker, J. Comput. Phys., 2015, 285, 316–330 CrossRef CAS
.
- R. Drautz, Phys. Rev. B, 2019, 99, 014104 CrossRef CAS
.
- A. Shapeev, Multiscale Model. Simul., 2016, 14(3), 1153–1173 CrossRef
.
- A. P. Thompson, H. Aktulga, R. Berger, D. Bolintineanu, W. Brown, P. Crozier, P. Veld, A. Kohlmeyer, S. Moore, T. Nguyen, R. Shan, M. Stevens, J. Tranchida, C. Trott and S. Plimpton, Comput. Phys. Commun., 2021, 271, 108171 CrossRef
.
-
J. Gilmer, S. Schoenholz, P. Riley, O. Vinyals and G. Dahl, Proceedings of the 34th International Conference on Machine Learning, 2017, 70, pp. 1263–1272 Search PubMed
.
- K. Schütt, P. Kindermans, H. Sauceda Felix, S. Chmiela, A. Tkatchenko and K.-R. Müller, Schnet: A continuous-filter convolutional neural network for modeling quantum interactions, Adv. Neural Inf. Process. Syst., 2017, 30, 992–1002 Search PubMed
.
-
J. Klicpera, J. J. Groß and S. Günnemann, International Conference on Learning Representations, 2020, pp. 1–13 Search PubMed
.
- J. Gasteiger, F. Becker and S. Günnemann, Adv. Neural Inf. Process. Syst., 2021, 34, 6790–6802 Search PubMed
.
- O. Unke, S. Chmiela, M. Gastegger, K. T. Schütt, H. E. Sauceda and K.-R. Müller, Nat. Commun., 2021, 12(1), 7273 CrossRef CAS PubMed
.
- C. Chen and S. Ong, Nat. Comput. Sci., 2022, 2(11), 718–728 CrossRef PubMed
.
- S. Batzner, A. Musaelian, L. Sun, M. Geiger, J. Mailoa, M. Kornbluth, N. Molinari, T. E. Smidt and B. E. Kozinsky, Nat. Commun., 2022, 13(1), 2453 CrossRef CAS PubMed
.
- I. Batatia, D. Kovacs, G. Simm, C. Ortner and G. Csányi, Adv. Neural Inf. Process. Syst., 2022, 35, 11423–11436 Search PubMed
.
-
Y.-L. Liao and T. Smidt, in The Eleventh International Conference on Learning Representations, 2023 Search PubMed
.
- D. P. Kovacs, I. Batatia, E. Arany and G. Csyani, J. Chem. Phys., 2023, 159, 044118 CrossRef CAS PubMed
.
- D. Kovacs, J. Moore, N. Browning, I. Batatia, J. Horton, Y. Pu, V. Kapil, W. Witt, I.-B. Magdau, D. Cole and G. Csyani, J. Am. Chem. Soc., 2025, 147(21), 17598–17611 CrossRef CAS PubMed
.
- A. Musaelian, S. Batzner, A. Johansson, L. Sun, C. Owen, M. Kornbluth and B. Kozinsky, Nat. Commun., 2023, 14(1), 579 CrossRef CAS PubMed
.
- N. Leimeroth, L. Erhard, K. Albe and J. Rohrer, Model. Simulat. Mater. Sci. Eng., 2025, 33, 065012 CrossRef
.
- G. Wang, C. Wang, X. Zhang, Z. Li, J. Zhou and Z. Sun, iScience, 2024, 27, 109673 CrossRef CAS PubMed
.
- W. Stark, C. Oord, I. Batatia, Y. Zhang, B. Jiang, G. Csyani and R. Maurer, Mach. Learn.: Sci. Technol., 2024, 5, 030501 Search PubMed
.
- J. Smith, O. Isayev and A. Roitberg, Chem. Sci., 2017, 8(4), 3192–3203 RSC
.
- M. Gastegger, J. Behler and P. Marquetand, Chem. Sci., 2017, 8(10), 6924–6935 RSC
.
- S. Manzhos and T. Carrington Jr, Chem. Rev., 2020, 121(16), 10187–10217 CrossRef PubMed
.
- A. Bartók, S. De, C. Poelking, N. Bernstein, J. R. Kermode, G. Csányi and M. Ceriotti, Sci. Adv., 2017, 3(12), e1701816 CrossRef PubMed
.
- K. Lin, L. Paterson, F. May and D. Andrienko, npj Comput. Mater., 2021, 7(1), 179 CrossRef CAS
.
- D. Earl and M. W. Deem, Phys. Chem. Chem. Phys., 2005, 7(23), 3910–3916 RSC
.
- A. Patriksson and D. van der Spoel, Phys. Chem. Chem. Phys., 2008, 10(15), 2073–2077 RSC
.
- O. Isayev, D. Fourches, E. Muratov, C. Oses, K. Rasch, A. Tropsha and S. Curtarolo, Chem. Mater., 2015, 27(3), 735–743 CrossRef CAS
.
- M. Ceriotti, J. Chem. Phys., 2019, 150, 150901 CrossRef PubMed
.
- B. Cheng, R. Griffiths, S. Wengert, C. Kunkel, T. Stenczel, B. Zhu, V. Deringer, N. Bernstein, J. T. Margraf and K. Reuter, Acc. Chem. Res., 2020, 53(9), 1981–1991 CrossRef CAS PubMed
.
- C. D. Williams, J. Kalayan, N. A. Burton and R. A. Bryce, Chem. Sci., 2024, 15, 12780–12795 RSC
.
- J. Behler, Chem. Rev., 2021, 121(16), 10037–10072 CrossRef CAS PubMed
.
- J. Behler, J. Chem. Phys., 2011, 134(7), 074106 CrossRef PubMed
.
- A. Singraber, T. Morawietz, J. Behler and C. Dellago, J. Chem. Theory Comput., 2019, 15(5), 3075–3092 CrossRef CAS PubMed
.
- I. Poltavsky, A. Charkin-Gorbulin, M. Puleva, G. Fonseca, I. Batatia, N. Browning, S. Chmiela, M. Cui, J. T. Frank and S. Heinen, Chem. Sci., 2025, 16, 3738–3754 RSC
.
- X. Fu, Z. Wu, W. Wang, T. Xie, S. Keten, R. Gomez-Bombarelli and T. Jaakkola, arXiv, 2022, preprint arXiv:2210.07237, DOI: DOI:10.48550/arXiv.2210.07237.
- V. Bihani, S. Mannan, U. Pratiush, T. Du, Z. Chen, S. Miret, M. Micoulaut, M. Smedskjaer, S. Ranu and N. A. Krishnan, Digital Discovery, 2024, 3(4), 759–768 RSC
.
- C. Schran, F. Thiemann, P. Rowe, E. A. Müller, O. Marsalek and A. Michaelides, Proc. Natl. Acad. Sci. U. S. A., 2021, 118(38), e2110077118 CrossRef CAS
.
- S. Chmiela, A. Tkatchenko, H. Sauceda, I. Poltavsky, K. T. Schütt and K.-R. Müller, Sci. Adv., 2017, 3(5), e1603015 CrossRef PubMed
.
- S. Chmiela, V. Vassilev-Galindo, O. Unke, A. Kabylda, H. E. Sauceda, A. Tkatchenko and K.-R. Müller, Sci. Adv., 2023, 9(2), eadf0873 CrossRef PubMed
.
- S. Luo, T. Chen and A. Krishnapriyan, ICLR, 2024, 899 Search PubMed
.
-
Z. Guo, D. Lu, Y. Yan, S. Hu, R. Liu, G. Tan, N. Sun, W. Jiang, L. Liu, and Y. Chen, Proceedings of the 27th ACM SIGPLAN Symposium on Principles and Practice of Parallel Programming, 2022, pp. 205–218 Search PubMed
.
-
J. Li, B. Li, Z. Guo, M. Li, E. Li, L. Liu, G. Yuan, Z. Wang, G. Tan, and W. Jia, SC24: International Conference for High Performance Computing, Networking, Storage and Analysis, 2024, pp. 1–15 Search PubMed
.
- M. Gastegger, L. Schwiedrzik, M. Bittermann, F. Berzsenyi and P. Marquetand, J. Chem. Phys., 2018, 148(24), 241709 CrossRef CAS PubMed
.
- Y. Zhao, M. Schwab, A. Kiersnowski, W. Pisula, M. Baumgarten, L. Chen, K. Müllen and C. Li, C. Trifluoromethyl-functionalized bathocuproine for polymer solar cells, J. Mater. Chem. C, 2016, 4(21), 4640–4646 RSC
.
- M. Van Vleet, A. Misquitta, A. Stone and J. Schmidt, J. Chem. Theory Comput., 2016, 12, 3851–3870 CrossRef CAS PubMed
.
| This journal is © The Royal Society of Chemistry 2026 |