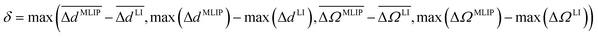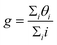Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Evaluation of foundational machine learned interatomic potentials for migration barrier predictions
Achinthya Krishna
Bheemaguli
ab,
Penghao
Xiao
*c and
Gopalakrishnan
Sai Gautam
 *b
*b
aDepartment of Metallurgical and Materials Engineering, National Institute of Technology Karnataka, Surathkal 575025, India
bDepartment of Materials Engineering, Indian Institute of Science, Bengaluru 560012, India. E-mail: saigautamg@iisc.ac.in
cDepartment of Physics and Atmospheric Science, Dalhousie University, Halifax B3H 4R2, Nova Scotia, Canada. E-mail: penghao.xiao@dal.ca
First published on 30th March 2026
Abstract
Fast, and accurate prediction of ionic migration barriers (Em) is crucial for designing next-generation battery materials that combine high energy density with facile ion transport. Given the computational costs associated with estimating Em using conventional density functional theory (DFT) based nudged elastic band (NEB) calculations, we benchmark the accuracy in Em and geometry predictions of five foundational machine learned interatomic potentials (MLIPs), which can potentially accelerate predictions of ionic transport. Specifically, we assess the accuracy of MACE-MP-0, MACE-OMAT-medium, Orb-v3, SevenNet, CHGNet, and M3GNet models, coupled with the NEB framework, against DFT-NEB-calculated Em across a diverse set of battery-relevant chemistries and structures. Notably, MACE-MP-0 and Orb-v3 exhibit the lowest mean absolute errors in Em predictions across the entire dataset and over data points that are not outliers, respectively. Importantly, Orb-v3, MACE-OMAT-medium, and SevenNet classify ‘good’ versus ‘bad’ ionic conductors with an accuracy of >82%, based on a threshold Em of 500 meV, indicating their utility in high-throughput screening approaches. Notably, intermediate images generated by MACE-MP-0 and SevenNet provide better initial guesses relative to conventional interpolation techniques in >71% of structures, offering a practical route to accelerate subsequent DFT-NEB relaxations. Finally, we observe that accurate Em predictions by MLIPs are not correlated with accurate (local) geometry predictions. Our work establishes the use-cases, accuracies, and limitations of foundational MLIPs in estimating Em and should serve as a base for accelerating the discovery of novel ionic conductors for batteries and beyond.
1 Introduction
Developing next-generation batteries is essential for our transition into sustainable energy usage, given that the state-of-the-art lithium-ion batteries (LIBs), while already delivering excellent performance, are approaching their fundamental limits,1,2 necessitating the discovery of novel beyond-LIB materials and chemistries. One key materials property that governs battery performance is the ionic diffusivity (D) of the electroactive ion in electrodes and (solid) electrolytes. D is exponentially influenced by the energy barrier each ion must overcome, commonly referred to as the migration barrier (Em), to hop from its initial lattice site to a symmetrically equivalent final site.3–5 Each ionic hop is often mediated through vacancies in the lattice with the ion overcoming transition state(s) along the hop. D and Em are thus related via the Arrhenius expression, D = D0 exp(−Em/kBT).6Accordingly, materials with a low Em – in both electrodes and (solid) electrolytes exhibit higher ionic conductivity and enable faster charge/discharge rates.7 In particular, emerging multivalent battery chemistries, such as Mg or Ca-based systems that promise higher volumetric energy densities,8 often suffer from poor rate performance.9–14 Some Na-ion cathodes such as marcite-Na(Mn/Fe)PO4, phosphate alluaudite-NaxMnFe2(PO4)3 and sulfate sodium superionic conductors (NaSICONs) that offer lower costs compared to LIB cathodes also suffer from poor rate performance.15,16 Therefore, understanding and minimizing the Em in candidate materials is crucial for advancing the next-generation of high-performance batteries.
Experimental techniques such as quasi-elastic neutron scattering,17 electrochemical impedance spectroscopy,18 nuclear magnetic resonance measurements,19 and galvanostatic intermittent titration techniques20 are commonly employed to study ion dynamics in solids.21 However, these methods often require access to large-scale facilities, and can exhibit chemistry or material-specific constraints/requirements, limiting their accessibility. As a result, computational approaches, particularly density functional theory (DFT22,23)-based nudged elastic band (NEB24) calculations are commonly used for computationally estimating Em with reasonable precision.25 While ab initio molecular dynamics (AIMD) simulations can also be used for estimating Em, such simulations are computationally expensive since they require the sampling over large length and long time scales across different temperatures to provide reasonable Em.26,27 Moreover, AIMD simulations can be unreliable for systems exhibiting high Em (i.e., the ‘true negatives’ among materials that can conduct ions) due to insufficient sampling of ion dynamics, resulting in DFT-NEB being the usual technique deployed for Em predictions.
Calculating Em using DFT-NEB requires an initial guess for the minimum energy path (MEP), which is typically constructed by linearly interpolating the coordinates of the initial and final configurations of the moving ion. Each ‘image’ generated by linear interpolation is subsequently connected via an auxiliary spring force. Note that the initial interpolated guess is often far from the true MEP, increasing the computational expense of DFT-NEB calculations and making them prone to convergence difficulties.25 Alternative approaches such as ‘ApproxNEB’,28 have been proposed to reduce computational intensity, with limited efficacy.
Recently, foundational machine learned interatomic potentials (MLIPs29–31), also referred to as universal potentials, have emerged as a new paradigm in computational materials science. The foundational MLIPs, pre-trained on large and diverse datasets, can generalize to a wide range of downstream tasks31 and are transferable across different materials and property prediction tasks,32–35 unlike classical MLIPs or force-fields that are constrained by a specific chemistry or property. Thus, foundational MLIPs are attractive candidates for accelerating atomistic simulations, including NEB calculations, by potentially improving initial MEP guesses and reducing the need for extensive DFT-based refinement or optimization, which can enable high-throughput screening of materials based on their Em. Indeed, a recent work by Kang et al.36 proposed an alternate to traditional DFT-based NEB calculations for Em estimations by using an MLIP for generating the potential energy surface on a spatial grid and extracting the MEP without the need for pre-defined NEB-images.
Several studies have benchmarked the performance of foundational MLIPs on diverse material properties,37–40 but not on predicting Em in solids. For instance, the ‘Matbench discovery’ platform41 provides a standardized framework for ranking universal potentials, but does not yet evaluate their integration with NEB workflows for Em predictions. Other MLIP benchmarking studies include the work by Zhao et al.42 that evaluated the MLIPs on transition state search for chemical reactions involving molecules. Bihani et al.43 benchmarked the performance of equivariant MLIPs on their generalizability to higher temperature simulations and unseen compositions, while Mannan et al. evaluated the performance of universal potentials against experimental measurements of elastic properties and structural accuracy among minerals.39 So far, there has been no benchmark of the performance of state-of-the-art universal potentials in predicting Em across a wide range of (battery) chemistries and materials, especially by integrating them with NEB workflows.
Here, we assess the performance of foundational MLIPs, namely, MACE-MP-0,44,45 MACE-OMAT-medium,45 SevenNet,46,47 Orb-v3,48,49 CHGNet,50 and M3GNet51 in predicting Em with NEB calculations. Using the dataset DFT-calculated Em compiled and curated by Devi et al.52,53 that spans a wide range of materials and compositions, we benchmark the Em predictions of the foundational MLIPs against conventional DFT-NEB values at the generalized gradient approximation (GGA54) level of exchange–correlation accuracy for 574 migration paths. Additionally, we introduce a metric to assess the similarity of MLIP-NEB relaxed structures with the ground truth of DFT-NEB computed MEPs from our previous works.25,55–57 Finally, we examine the correlation between accuracies in Em and geometry predictions by the MLIPs considered.
Notably, we find that M3GNet and CHGNet (invariant models) tend to underestimate Em and exhibit a high degree of confidence in predicting low Em over a narrow range of possible Em values, while the other potentials (equivariant models) exhibit no clear bias and deliver consistent accuracy over a wide range of Em values. Importantly, we observe that Orb-v3, MACE-OMAT-medium, and SevenNet classify systems as ‘good’ (Em < 500 meV) or ‘bad’ ionic conductors with 82% accuracy. Performing an MLIP-NEB, using any of the potential considered, does result in improved interpolated paths representing the MEP in over 66% of cases, indicating their utility in high-throughput screening workflows. Significantly, we find no evident correlation between the accuracy of Em and geometry predictions, with MLIPs yielding higher accuracy in Em predictions for systems with low Em values, while demonstrating better geometry predictions in systems with large Em. We hope that our study establishes use-cases and quantifies the reliability of using foundational MLIPs in predicting Em over a diverse set of chemistries and crystal structures, which in turn should accelerate materials discovery for novel battery applications and beyond.
2 Methods
2.1 Datasets
We utilize two distinct subsets of the Em dataset in this work, as schematically shown in Fig. 1. The smaller dataset, referred to as ‘Dataset-1’, comprises 60 DFT-NEB calculations, including multiple possible migration pathways for select structures. Each datapoint consists of GGA-calculated Em and the relaxed structures of all intermediate images across a migration pathway. We constructed this dataset using the DFT-NEB results from our previous works,25,55–57 encompassing crystal structures explored primarily as battery materials, such as layered structures, Weberites, spinels, olivines, perovskites, and NaSICONs. The Em in Dataset-1 ranges from 0.06 eV to 2.88 eV. | ||
| Fig. 1 Overview of the methodology, indicating the use of two subsets of the Em dataset that were created for examining geometry predictions, geometry–barrier correlations, and barrier predictions. | ||
The larger dataset, referred to as ‘Dataset-2’, is a subset of a literature-derived collection of Em,52 which comprises of 621 DFT-calculated Em and the initial and final configurations for each migration pathway. Among the 621 datapoints, we excluded systems exhibiting Em > 2.5 eV, since such high Em values would not correspond to any tangible rate performance under battery operating conditions. We also excluded systems that presented significant convergence difficulties during NEB calculations using any of the foundational MLIPs considered (∼10 datapoints), so that a fair and quantitative comparison can be made across the MLIPs. Thus, the final subset that forms our Dataset-2 consists of 574 systems. The systems comprising both datasets are compiled in our https://github.com/sai-mat-group/mlips-migration-barriers repository, while Dataset-2 is also available as a json file on Zenodo.
2.2 Model details
We used publicly available universal potentials that have demonstrated good performance for bulk properties, are constructed on graph-based neural network (GNN) architectures, and are compatible with the atomic simulation environment (ASE58). We leveraged ASE calculators to integrate the MLIPs with the NEB implementation available within ASE. The specific MLIPs we used include (i) the MACE-MP-0 ‘large’ foundation model, trained on approximately 1.6 million materials project59 bulk-crystal relaxation trajectories (i.e., the ‘MPtrj’ dataset60) with maximal message equivariance (L = 2), (ii) the SevenNet-MF-ompa model, which incorporates multifidelity learning with a core architecture based on the neural equivariant interatomic potential (NequIP61) and is trained on MPtrj, ‘OMat24’ trajectories,62 and the subsampled Alexandria (sAlex63) datasets, (iii) the Orb-v3-conservative-inf-omat that is trained on the MPtrj, OMat24 and Alexandria (Alex) dataset with learned forces being conservative by construction and effectively unlimited neighbor lists, (iv) CHGNet v0.3.0, which is trained on MPtrj, and (v) M3GNet MP-2021.2.8-EFS version, which is trained on MPtrj up to February 8, 2021. In addition to MACE-MP-0, we also considered the MACE-OMAT-medium model,45 which has been trained on the OMat24 trajectories, to examine the influence of training data on the performance of a model architecture. We used all models as provided, without any additional fine-tuning or hyperparameter optimizations. A summary of the specific details for each model is provided in Table 1.| Model | Training data | Model type and key features |
|---|---|---|
| MACE-MP-0 | MPtrj dataset | E(3)-equivariant GNN that captures many-body interactions |
| MACE-OMAT-medium | OMat24 | |
| SevenNet-MF-ompa | MPtrj, OMat24, and sAlex | Equivariant GNN incorporating multifidelity learning with efficient parallelization |
| Orb-v3 | MPtrj, OMat24, and Alex | Roto-equivariance inducing regularized GNN with analytical energy gradients (conservative forces) and (effectively) infinite neighbors |
| CHGNet | MPtrj dataset | Invariant GNN including magnetic moment inputs, thus incorporating information on atomic charges |
| M3GNet | MPtrj dataset | Includes three-body interactions within its GNN (invariant) |
2.3 NEB calculations
Typically, DFT-NEB calculations employ linear interpolation (LI) of atomic coordinates between the initial and final endpoints for generating the initial guess for the images. Whereas, the image dependent pair potential (IDPP) interpolation technique, developed by Smidstrup and coworkers64 utilizes a distance-matching objective to generate the initial guess for MEP. Given a preliminary benchmark of MACE-MP-0 based NEB calculations initialized with LI and IDPP interpolation, as detailed in Section S1 and Fig. S1 of the SI, we find IDPP interpolation to provide marginally better initial guesses for the eventual NEB calculations.For NEB calculations of materials in Dataset-1 using all universal potentials, we generated seven intermediate images, mirroring the number used in the corresponding DFT-NEB calculations. The initial interpolated images were connected by a spring constant, k = 5 eV Å−2, and we utilized the NEB implementation following the elastic band (EB65) method with full spring force, given our benchmarking with MACE-MP-0 (see Section S1). We did not include the climbing image technique24 in any of our MLIP-NEBs, as we did not see significantly different results with or without climbing image in our previous work.25 We deemed the NEB converged when the band forces fell below 0.05 eV Å−1, while using the Broyden–Fletcher–Goldfarb–Shanno optimizer.66–69 In the case of Dataset-2, we employed only three intermediate images for all foundational MLIPs considered, to reduce computational costs. Note that employing seven intermediate images with MACE-MP-0 NEB calculations on a random subset of 100 structures did not significantly change the Em predictions (average deviations of ∼75 meV, which is similar to typical DFT-NEB Em errors), indicating that the data and trends reported in our work based on three intermediate images should be robust. Also, we used the set of optimized NEB parameters that were used for calculations on Dataset-1 (i.e., k = 5 eV Å−2, IDPP interpolation, and the EB method) for all calculations involving Dataset-2.
2.4 Geometry metrics
To quantitatively assess the similarity of local geometries between structures, we introduce the geometric similarity classification metric, θ for a given image structure between the endpoints and an averaged geometric similarity score, g, across a migration path. θ evaluates whether the local geometry of an intermediate image obtained via a MLIP-NEB calculation is a better approximation of the reference structure (i.e., DFT-NEB relaxed) compared to the image generated using simple LI. Thus, θ is useful to determine whether intermediate images relaxed using MLIP-NEBs provide a superior initial guess for DFT-NEB calculations than typical LI, based on local geometric features. Note that the ground truth for all our geometric comparisons is the relaxed geometry, as obtained with a DFT-NEB calculation. We define θ using the following steps:| dDFTxy, dMLIPxy, dLIxy |
| ΔdMLIP = |dDFTxy − dMLIPxy| |
| ΔdLI = |dDFTxy − dLIxy| |
for all {x, y} ⊂ {i, j, k, l, m, n, c}, where x ≠ y.
| ΩDFTx, ΩMLIPx, ΩLIx |
In the above notation, two Ω, say ΩDFT, ΩMLIP, having the same x indicates that the polyhedral faces correspond to the same set of neighboring atoms. The absolute differences with the DFT-NEB relaxed structures are then calculated and stored as six-dimensional vectors:
| ΔΩMLIP = |ΩDFTx − ΩMLIPx| |
| ΔΩLI = |ΩDFTx − ΩLIx| |
for all x ∈ {a, b, c, d, e, f}.
We expect the local geometry of an MLIP-NEB relaxed structure to be a poorer approximation of the DFT-NEB relaxed structure than the corresponding LI structure, if at least one of the following conditions is met: (i) one of the 21 pairwise distances or 6 solid angles of the MLIP-NEB relaxed structure deviates significantly more from the DFT-NEB geometry than the corresponding LI structure, or (ii) the average difference in pairwise distances or solid angles of the MLIP-NEB relaxed structure with the DFT-NEB reference is significantly higher compared to LI. To quantify these two conditions, we calculate δ, which represents the maximum value among the differences in the mean and maximum errors of distances and angles between the MLIP-NEB and LI structures:
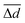 and
and 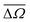 represent the mean of the absolute errors in distances and solid angles, respectively. max(Δd) and max(ΔΩ) represent the maximum absolute errors.
represent the mean of the absolute errors in distances and solid angles, respectively. max(Δd) and max(ΔΩ) represent the maximum absolute errors.
Finally, the metric θ would classify the structure as:
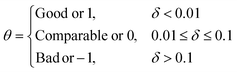 | (1) |
Thus, δ quantifies the difference between the deviations of the MLIP-NEB and LI structures with respect to the DFT-NEB reference based on key local geometric features. A smaller (ideally negative) δ value signifies that the MLIP-NEB structure exhibits consistently lower errors, indicating it's a better approximation of the true DFT-NEB pathway. Conversely, larger (more positive) δ suggests that LI performed as well or even better than the MLIP-NEB for at least one of the local geometric attributes. Therefore, we numerically represent the ‘good’, ‘comparable’ and ‘bad’ structure as 1, 0 and −1 with θ. Finally, for a given system containing i intermediate images, we define g as,
In the case where all the i image local geometries are better (worse) predicted by MLIP-NEB compared to LI, g will take the value of 1 (−1).
3 Results
3.1 Barrier prediction performance
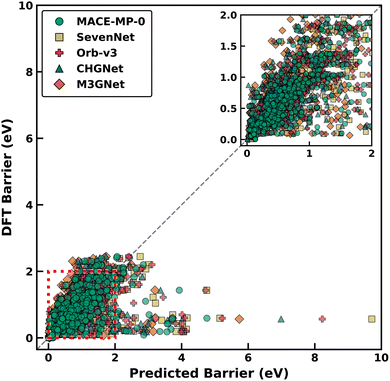
Fig. 2 presents a comparison of Em predictions on Dataset-2 across different foundational MLIPs (x-axis) with their corresponding DFT-NEB calculated Em (y-axis). Green circles, yellow squares, pink pluses, blue triangles, and orange rhombuses represent Em predictions using MACE-MP-0, SevenNet, Orb-v3, CHGNet, and M3GNet, respectively. Individual parity plot for each MLIP considered is compiled in Fig. S2–S6 of the SI and in Fig. S8 for MACE-OMAT-medium. Overall, MACE-MP-0 demonstrates the best performance with an MAE of 0.310 eV, while M3GNet records the highest MAE of 0.349 eV. The MAEs of Orb-v3, CHGNet, and SevenNet are 0.336, 0.343, and 0.344 eV, respectively. To provide a numerical context to the MAEs, DFT-NEB calculations typically carry a ∼60 meV error in their predictions, and a change of 60 meV in Em at 298 K corresponds to an order-of-magnitude change in D.72 However, the calculation of MAEs is influenced by extreme outliers that affect all MLIPs.To obtain a more representative picture of MLIP performance, we exclude 17 systems that act as common outliers across all MLIPs, with each outlier exhibiting absolute errors exceeding 1 eV. Notably, excluding the common outliers also reveals a similar performance hierarchy as with retaining the entire dataset: MACE-MP-0 emerges with the best MAE of 0.239 eV, followed closely by Orb-v3 with 0.245 eV. The remaining MLIPs, namely SevenNet, CHGNet, and M3GNet show MAEs of 0.251, 0.275, and 0.290 eV, respectively. Specific details about the outliers of respective MLIPs can be found in Tables S3–S7 of the SI, while the distribution of outliers across crystal classes is compiled in Fig. S7.
Besides accuracy, we analyze the distribution of datapoints relative to the ideal parity line to determine whether the MLIPs exhibit systematic prediction biases (i.e., under- or over-estimation of Em). Interestingly, we observe MACE-MP-0, SevenNet, and Orb-v3 to demonstrate a relatively balanced prediction behavior with fairly symmetric distributions of under and over-estimated datapoints. Represented as (number of under-estimated datapoints, number of over-estimated datapoints) pairs, MACE-MP-0, SevenNet, and Orb-v3 exhibit distributions of (299,![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 275), (244,
275), (244,![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 330), and (242,
330), and (242,![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 332), respectively. In contrast, CHGNet and M3GNet show a bias toward under-estimating barriers, with under-estimated datapoints accounting for 73.1% and 78.2% of all predictions, respectively. Represented as (under-estimated, over-estimated) pairs, CHGNet and M3GNet exhibit distributions of (420,
332), respectively. In contrast, CHGNet and M3GNet show a bias toward under-estimating barriers, with under-estimated datapoints accounting for 73.1% and 78.2% of all predictions, respectively. Represented as (under-estimated, over-estimated) pairs, CHGNet and M3GNet exhibit distributions of (420,![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 154) and (449,
154) and (449,![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 125), respectively.
125), respectively.
To further understand individual MLIP capabilities, we examined each potential's performance after excluding the outliers specific to each potential (i.e., systems with absolute errors >1 eV as predicted by a given potential) to gain insight into the ‘best-case’ scenario of Em predictions. Notably, despite having 37 outliers, Orb-v3 achieves the lowest MAE of 0.198 eV on its remaining (non-outlier) predictions. With 35 outliers, MACE-MP-0 is a close second with an MAE of 0.202 eV, while SevenNet, with 37 outliers, displays an MAE of 0.203 eV. CHGNet and M3GNet show higher MAE values of 0.248 eV and 0.257 eV with 31 and 36 outliers, respectively. Also, varying training data seems to have a marginal impact on the performance of the model, with MACE-OMAT-medium exhibiting an MAE of 0.35 eV on the entire dataset and an MAE of 0.20 eV excluding its specific outliers. Thus, we find that Orb-v3 can achieve higher accuracies on systems that it describes well while MACE-MP-0 achieves a better balance of both low errors and fewer outliers compared to other MLIPs.
3.2 Predictions over different barrier ranges
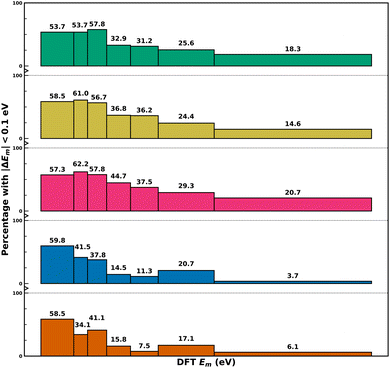
To understand the performance of MLIPs across various ranges of Em, we divided the 574 datapoints into seven equal-sized distributions based on their DFT-NEB Em values, as illustrated in Fig. 3. While the x-axis in Fig. 3 represents DFT-NEB Em ranges, with bar widths indicating the span of Em values within each bin, the y-axis shows the percentage of predictions within each bin that achieve absolute errors <0.1 eV (signifying an “acceptable” degree of accuracy). The exact DFT barrier range of the data points present in each bin can be found in Table S1 of the SI. Fig. S9 compiles the performance of the MACE-OMAT-medium model over different Em ranges.Trends in Fig. 3 indicate that all MLIPs struggle with high Em predictions, with only small percentage of systems exhibiting the acceptable accuracy in the highest barrier range (∼1.31–2.50 eV). Specifically, the percentage of predictions with acceptable accuracy in the highest Em range are 20.7% for Orb-v3, 18.3% for MACE-MP-0, 14.6% for SevenNet, 6.1% for M3GNet, and 3.7% for CHGNet. To examine whether the strategy used for binning influenced the trends we observe in Fig. 3, we performed an identical exercise while keeping the width of each bin a constant and compiled the results in Fig. S10 (bin width of 0.5 eV) and S11 (0.25 eV) of the SI. Importantly, we find no qualitative change in the performance of the models, with the accuracy in Em predictions declining with increasing Em values for all MLIPs considered.
Importantly, we identify a “sweet spot” of Em values where all MLIPs perform reasonably well. For example, in the low-barrier range (∼0.0025–0.25 eV), more than 50% of predictions achieve acceptable accuracy across all MLIPs. Within this range, CHGNet shows the highest success rate (59.8%), followed by M3GNet and SevenNet (both 58.5%), while MACE-MP-0 and Orb-v3 achieve 53.7% and 57.3%, respectively. Additionally, Orb-v3 and SevenNet achieve their best performance (i.e., highest fraction of predicted datapoints with acceptable accuracy) in the 0.25–0.36 eV range, achieving 62.2% and 61% acceptable predictions, respectively. MACE-MP-0 performs best in the slightly higher 0.36–0.50 eV range with 57.8% accuracy. Meanwhile, CHGNet and M3GNet perform best in the lowest Em range (∼0.0025–0.25 eV) with 59.8% and 58.5% accuracy, respectively.
While all MLIPs show declining accuracy with increasing Em, Orb-v3 exhibits the slowest degradation, maintaining better performance across a broader range of Em values compared to other potentials. Thus, we find that ‘simpler’ graph models such as CHGNet and M3GNet demonstrate superior performance for materials with intrinsically low Em values but lack consistency in their predictions over a wider range of Em. On the other hand, increasing complexity among the graph models, such as in Orb-v3 or MACE-MP-0 allows for a more robust performance across a wide range of Em values while sacrificing ‘peak’ performance for materials with low Em, making them better suited for Em predictions in novel materials. This variation in the performance of ‘simple’ and ‘complex’ MLIPs also reveals the general trade-off between building specialized and generalized models in the field of machine learning.
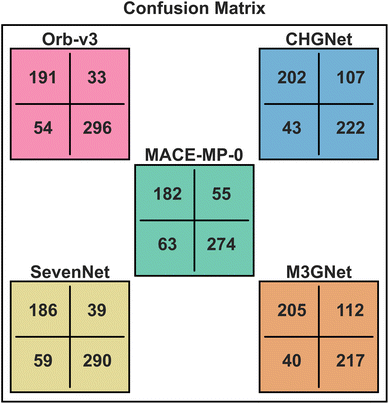
3.3 Barrier classification performance
To quantify the ability of the MLIPs considered to classify a material as a ‘good’ versus a ‘bad’ ionic conductor, which can be used for high-throughput identification of promising candidates, we present the confusion matrices for all MLIPs in Fig. 4 (see Table S8 for the confusion matrix of MACE-OMAT-medium). Each potential in Fig. 4 is represented using a distinct color, such as green (MACE-MP-0), yellow (SevenNet), pink (Orb-v3), blue (CHGNet), and orange (M3GNet). For the classification task, we use a threshold Em of 500 meV, i.e., materials that exhibit a calculated/predicted Em < 500 meV are labeled good ionic conductors, while materials that show higher values of Em are labeled bad ionic conductors. Within each confusion matrix, the true positive (TP), the true negative (TN), the false positive (FP) and the false negative (FN) numbers are listed on the top left, bottom right, top right, and bottom left cells, respectively.From Fig. 4, we observe that Orb-v3 achieves the highest combined number of TP and TN, correctly classifying 487 out of 574 systems (i.e., an accuracy of 84.84%). In comparison, M3GNet yields the lowest TP + TN count of 422 systems (73.52%). MACE-OMAT-medium, SevenNet, MACE-MP-0 and CHGNet correctly classify 477 (83.1%), 476 (82.93%), 456 (79.44%), and 424 (73.87%) systems, respectively. These results highlight Orb-v3 as the most reliable model for distinguishing good and poor ionic conductors, followed closely by SevenNet (accuracy >80%), making both reliable for high-throughput classification tasks.
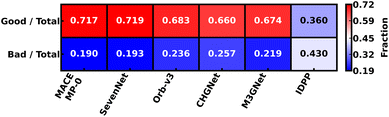
3.4 Geometry prediction performance
It is important for any MLIP to not only get the Em correct but also get the underlying geometries that constitute the MEP (and hence yield the Em value) correct for the MLIP to be truly accurate. Thus, we quantify the performance of the MLIPs considered in their predictions of local geometry of the intermediate image structures in Dataset-1 (using θ of eqn (1)) as a heatmap in Fig. 5. Note that we performed an NEB calculation with EB, k = 5 eV Å−2, and IDPP interpolation with each MLIP and for each material in Dataset-1 to generate the statistics displayed in Fig. 5. For each MLIP (x-axis), we denote the fraction of structures with ‘good’ (top row) and ‘bad’ (bottom) local geometries in Fig. 5. Ideally, the MLIPs should exhibit a high (low) fraction of structures with good (bad) local geometry. Since IDPP (without any subsequent MLIP-based relaxation) also behaves like a potential for generating a guess for the MEP, we include IDPP's statistics in Fig. 5.Among all the MLIPs, SevenNet exhibits the highest fraction of good geometries (0.719), indicating that it frequently generates accurate local geometries. On the other hand, MACE-MP-0 exhibits the lowest fraction of bad geometries (0.190), indicating that it frequently avoids generating inaccurate structures. The difference between the fraction of good and bad geometry predictions for both MACE-MP-0 and SevenNet are similar (0.527 and 0.526, respectively), indicating that both models perform equally well in generating good local geometries.
Other MLIPs show poorer geometry predictions, with Orb-v3, M3GNet, and CHGNet displaying good (bad) fractions of 0.683 (0.236), 0.674 (0.219), and 0.660 (0.257), respectively, with CHGNet showing the smallest difference between the good and bad geometry fractions (0.403). Thus, MACE-MP-0 and SevenNet show significantly better local geometry predictions upon relaxation with NEB compared to Orb-v3, M3GNet and CHGNet, while all MLIPs provide better initial guesses to the MEP than LI in at least 66% of structures (i.e., intermediate images). Also, we note that IDPP generated structures are statistically much farther from DFT than MLIPs, with LI being better than IDPP in 43% of the cases. Given our definition of θ and the specific systems present in Dataset-1, we find that IDPP does not make a significant difference in enhancing the initial guess for the MEP as compared to LI across all MLIPs.
3.5 Geometry–barrier correlation
To investigate whether there is a correlation between geometry and Em prediction performance, i.e., whether a precise Em prediction by a given MLIP is due to its precise geometry prediction, we divide Dataset-1 into five bins based on their DFT-NEB Em, ensuring that each bin contains an equal number of data points. Bins with indices 1, 2, 3, 4, and 5 correspond to Em ranges of [0.058, 0.64], (0.64, 0.97], (0.97, 1.24], (1.24, 1.81], and (1.81, 2.88] eV, respectively. Note that the number of bins in Fig. 6 is different from that of Fig. 3 since the datasets considered for both figures are different. Within each bin, we estimate the fraction of migration paths with ‘good’ local geometry (g > 0.85, blue points) and low absolute errors in Em (ΔE ≤ 0.1 eV, orange points), and plot the statistics for each MLIP in Fig. 6. Note that g > 0.85 signifies cases where the predicted geometry is better than unrelaxed LI for at least six out of the seven intermediate images.Overall, Fig. 6 reveals the absence of any positive correlation between barrier and geometry prediction performance, and more strikingly, an inverse relationship. For example, all models perform poorly in predicting high Em (bin 5), which is consistent with our observations in Fig. 4. However, all models also achieve their best geometry predictions for bin 5. In other words, the best geometry predictions are coincident with the worst Em predictions. The geometry prediction success rates within bin-5 are 66.7%, 75.0%, 66.7%, 66.7%, and 58.3% for MACE-MP-0, SevenNet, Orb-v3, CHGNet, and M3GNet, respectively, and the corresponding Em prediction success rates (i.e., ΔE ≤ 0.1 eV) are 16.7%, 0%, 16.7%, 8.3%, and 0%, respectively.
To further assess geometry–barrier correlation, we examine instances where MLIPs perform well in both metrics. Note that, we term a model to exhibit a ‘good performance in both metrics’ if both the fractions in a given bin are >0.5. Only two potentials show this good performance, and only in a single bin (bin-1), namely, MACE-MP-0 with a success rate of 58.3% in Em prediction and 50.0% in geometry prediction, and M3GNet with a 50.0% success rate for both metrics. SevenNet, Orb-v3, and CHGNet do not achieve this good performance in any bin. Moreover, we find no consistent pattern across all bins and all MLIPs and no instances where good Em predictions coincide with good geometry prediction. Instead, the data suggests that these two performance metrics are largely independent, and that a good Em prediction does not necessarily arise from a good local geometry prediction (and vice versa).
4 Discussion
Given the critical role of Em in battery materials design and the high computational costs associated with DFT-NEB calculations, we have evaluated the performance of foundational MLIPs including MACE-MP-0, SevenNet, Orb-v3, CHGNet, and M3GNet, for Em predictions upon integration with the NEB framework over two data subsets containing Em and structural data (Fig. 1). Specifically, we investigated (i) the ability of MLIPs to predict Em accurately, (ii) the likelihood of generating MLIP-NEB-relaxed image geometries that are close to the ground truth (DFT-NEB), and (iii) whether any correlation exists between the accuracy in Em prediction and geometry relaxation.Analyzing Em predictions across the entire Dataset-2, we find that MACE-MP-0 exhibits the lowest MAE (Fig. 2), followed in order by Orb-v3, CHGNet, SevenNet, and M3GNet. On excluding outliers that are common to all models, we observe SevenNet to exhibit a slightly lower MAE than CHGNet, with the rest of the performance order being the same. Interestingly, when assessing each model independently after removing their respective outliers, Orb-v3 demonstrates the best MAE of 0.198 eV, marginally outperforming MACE-MP0 (0.202 eV), with the other models exhibiting larger errors (0.203–0.257 eV). Thus, Orb-v3 provides the best prediction errors for Em, among MLIPs considered, in systems with a robust description of the corresponding potential energy surface.
Based on the distribution of outliers across different crystal systems (Fig. S7), we observe that while certain outliers are model-specific, systems containing orthosilicates and phosphates consistently pose challenges for all MLIPs, which may be attributed to the inherently complex potential energy surface of these polyanionic frameworks. Specifically, the intricate ionic migration paths within these structures may be difficult for MLIPs to capture accurately, likely due to spurious ‘smoothing’ of the potential energy surface during model training.
While minor inconsistencies in Hubbard U73,74 values between the datapoints in Dataset-2 and the calculation scheme of Materials Project do exist, we expect such inconsistencies to be unlikely as the primary source of error among the identified outliers for each MLIP considered. Indeed, GGA-calculated Em values are the predominant contributor to Dataset-2, accounting for 88.05% of the datapoints, while calculations including a Hubbard U correction only contribute 7.27% of the datapoints. GGA-calculated Em values dominate the literature so far, since the U corrections are frequently omitted in NEB calculations due to significant convergence difficulties and electronic metastability along the migration pathway, as noted by Liu et al.75 Furthermore, benchmarks by Devi et al.25 indicate that for GGA + U calculations, even a substantial change in the U parameter (≈1 eV) typically results in a Em variation of only 15 meV – a value well within the acceptable error margin for DFT-NEB calculations.
Notably, during endpoint relaxations for Orb-v3, 153 systems failed to converge within the threshold forces over 1000 optimization steps despite attempting multiple optimization algorithms, namely limited-memory Broyden–Fletcher–Goldfarb–Shanno (L-BFGS)76 and fast inertial relaxation engine (FIRE).77 While the unconverged structures did not satisfy our rigorous force convergence threshold of |0.05| eV Å−1, the residual forces were marginally above the threshold (typically ∼|0.08| eV Å−1). This suggests that the convergence issues may stem from the inherent features of the learned Orb-v3 potential energy surface, such as noise or shallow local minima, rather than a failure of the optimization algorithm itself. However, to maintain consistency with the other models in the study, we did not modify the obtained results and included the Orb-v3 results as is.
We observe that M3GNet and CHGNet exhibit a systematic bias toward underestimating Em, whereas MACE-MP-0, SevenNet, and Orb-v3 do not display such a tendency. A more granular analysis (Fig. 3) reveals that all models struggle with accurately predicting high Em values. Among them, Orb-v3 shows a relatively slow decay in prediction accuracy as the Em value increases. Interestingly, the simpler (invariant) models CHGNet and M3GNet outperform their more complex (equivariant) counterparts within a very narrow range of low Em but exhibit a rapid decline in performance as the range expands.
The systematic biases observed in Em predictions likely arise from the interplay between training data distribution and architectural inductive biases, such as the level of feature equivariance. Architectural differences, including the handling of many-body interactions and local environment cutoffs, further influence how a model learns the potential energy surface. In terms of the influence of training data on the model performance, we observe MACE-OMAT-medium to exhibit quite similar quantitative performance (in terms of MAEs) and a marginally better classification accuracy compared to MACE-MP-0. Thus, we conclude that architectural choices influence the performance of a model more significantly compared to the choice and size of the training data itself, at least for Em predictions.
Using a threshold Em of 500 meV to categorize structures as ‘good’ or ‘bad’ conductors of ions (Fig. 4), we find that all MLIPs are able to identify good conductors with reasonable accuracy (>73%). Orb-v3 and SevenNet display the highest accuracies in classifying good (or bad) conductors, with ∼85% and ∼83% accuracy, respectively, making them highly suitable for high-throughput screening of candidate battery materials.
Our study on Dataset-1 indicates that MLIP-NEB relaxations tend to produce image geometries that are as close as (or closer to) DFT-NEB structures than those obtained through simple LI or IDPP interpolation in the majority (∼66%, Fig. 5) of cases. Among the considered models, MACE-MP-0 and SevenNet stand out in geometry predictions, relaxing to geometries that are worse than LI or IDPP ones in only 19% of migration paths, suggesting that employing MACE-MP-0 or SevenNet NEB-relaxed images as initial guesses for DFT-NEB calculations could significantly accelerate convergence and reduce computational costs.
Although our metric, θ (see eqn (1)), captures critical local geometric features, it can be improved further to decisively quantify local structural similarity. Nevertheless, we performed DFT-NEB calculations using initial path guesses derived from MACE-MP-0-based NEB for a subset of structures exhibiting high geometric similarity. With all other DFT parameters held constant, we observed a reduction in the number of ionic and electronic steps required to achieve convergence in 5 out of 6 cases compared to LI initialization, as documented in Table S2. This reduction in computational cost provides empirical evidence that MLIP-based path initialization has the potential to accelerate subsequent DFT-NEB calculations.
Finally, when simultaneously evaluating the likelihood of accurate barrier prediction and better geometry initialization (Fig. 6), we observe no evident correlation between the two among all MLIPs considered. Thus, we find that accurate barrier predictions do not necessarily imply better geometry predictions, and vice versa. One possible explanation for this counterintuitive trend is that for systems with low Em, the potential energy surfaces are likely ‘flat’ with variations in local geometries, indicating that even large errors in local bond distances or local bond angles made by the MLIPs do not significantly change the predicted Em, thus leading to accurate Em even with inaccurate geometries. On the other hand, for systems with large Em, the potential energy surfaces should exhibit ‘deep’ minima associated with the ‘stable’ sites occupied by the migrating ion, signifying that even small errors in predicting local bond distances or angles by the MLIPs can cause large errors in the Em predictions, thus resulting in inaccurate Em even with mostly accurate geometries.
5 Conclusion
Given the importance of accurate and swift predictions of Em in materials for battery applications, we systematically evaluated six foundational MLIPs (MACE-MP-0, MACE-OMAT-medium, SevenNet, Orb-v3, CHGNet, and M3GNet) integrated with the NEB framework across a diverse set of battery-relevant chemistries. We found MACE-MP-0 to achieve the lowest overall MAE of 0.310 eV, while Orb-v3 demonstrated superior performance with an MAE of 0.198 eV when excluding outliers. Additionally, we discovered no direct correlation between barrier prediction accuracy and geometric similarity. Based on our results, we propose that Orb-v3, MACE-OMAT-medium, and SevenNet are preferable for high-throughput screening due to their >82% classification accuracy, while simpler (invariant) models such as CHGNet and M3GNet are better suited for specialized applications with low Em (<0.25 eV) requirements. For general-purpose accuracy across unknown barrier ranges, MACE-MP-0 and Orb-v3 remain the most robust choices. Importantly, for studies aiming to accelerate high-fidelity DFT-NEB calculations, we recommend using MACE-MP-0 or SevenNet to generate initial MEP guesses, since these models produce relaxed geometries that are closer to the DFT ground truth in over 71% of cases, significantly outperforming conventional LI. We hope our work establishes the specific use-cases and limitations of foundational MLIPs, facilitating the accelerated discovery of novel ionic conductors for next-generation energy storage.Author contributions
Achinthya Krishna Bheemaguli: methodology, software, investigation, data curation, writing – original draft, visualization. Penghao Xiao: resources, writing – review & editing, supervision. Gopalakrishnan Sai Gautam: conceptualization, methodology, resources, writing – review & editing, supervision, project administration, funding acquisition.Conflicts of interest
There are no conflicts to declare.Data availability
Both datasets and associated python scripts used in this work are compiled and available for free at our https://github.com/sai-mat-group/mlips-migration-barriersrepository. Dataset-2 is available as a json file on Zenodo.Supplementary information (SI): optimal nudged elastic band parameters, parity plots, analysis of outliers and binning strategies, performance of the MACE-OMAT-medium model, nudged elastic band calculations post MLIP relaxation, and range of migration barrier values. See DOI: https://doi.org/10.1039/d5dd00534e.
Acknowledgements
G. S. G. acknowledges financial support from the Science and Engineering Research Board (SERB) of the Department of Science and Technology, Government of India, under sanction number IPA/2021/000007. A. K. B. thanks the Ministry of Human Resource Development, Government of India, for financial assistance. The authors gratefully acknowledge the super-computing facility offered by ACENET and the Digital Research Alliance of Canada. The authors acknowledge the computational resources of the super computer ‘PARAM Pravega’ provided by the super computer education and research centre (SERC) at IISc. The authors also thank the Jülich Supercomputing Centre (at Forschungszentrum Jülich), Germany for the use of the ‘JURECA’ supercomputer, under projects ‘hpc-prf-emdft’ and ‘hpc-prf-desal’.References
- M. Li, J. Lu, Z. Chen and K. Amine, Adv. Mater., 2018, 30, 1800561 CrossRef PubMed.
- M. S. Whittingham, Chem. Rev., 2014, 114, 11413 CrossRef CAS PubMed.
- M. Park, X. Zhang, M. Chung, G. B. Less and A. M. Sastry, J. Power Sources, 2010, 195, 7904–7929 CrossRef CAS.
- J. C. Bachman, S. Muy, A. Grimaud, H.-H. Chang, N. Pour, S. F. Lux, O. Paschos, F. Maglia, S. Lupart and P. Lamp, et al. , Chem. Rev., 2016, 116, 140–162 CrossRef CAS PubMed.
- E. Flores, C. Wölke, P. Yan, M. Winter, T. Vegge, I. Cekic-Laskovic and A. Bhowmik, Digital Discovery, 2022, 1, 440–447 RSC.
- G. H. Vineyard, J. Phys. Chem. Solids, 1957, 3, 121–127 CrossRef CAS.
- A. Van der Ven, Z. Deng, S. Banerjee and S. P. Ong, Chem. Rev., 2020, 120, 6977–7019 CrossRef CAS PubMed.
- P. Canepa, G. Sai Gautam, D. C. Hannah, R. Malik, M. Liu, K. G. Gallagher, K. A. Persson and G. Ceder, Chem. Rev., 2017, 117, 4287–4341 CrossRef CAS PubMed.
- Y. Orikasa, T. Masese, Y. Koyama, T. Mori, M. Hattori, K. Yamamoto, T. Okado, Z.-D. Huang, T. Minato and C. Tassel, et al. , Sci. Rep., 2014, 4, 5622 CrossRef CAS PubMed.
- Q. Liu, H. Wang, C. Jiang and Y. Tang, Energy Storage Mater., 2019, 23, 566–586 CrossRef.
- Y. Liang, H. Dong, D. Aurbach and Y. Yao, Nat. Energy, 2020, 5, 646–656 CrossRef CAS.
- R. D. Bayliss, B. Key, G. Sai Gautam, P. Canepa, B. J. Kwon, S. H. Lapidus, F. Dogan, A. A. Adil, A. S. Lipton and P. J. Baker, et al. , Chem. Mater., 2019, 32, 663–670 CrossRef.
- G. S. Gautam, X. Sun, V. Duffort, L. F. Nazar and G. Ceder, J. Mater. Chem. A, 2016, 4, 17643–17648 Search PubMed.
- D. Wang, Z. Zhang, Y. Hao, H. Jia, X. Shen, B. Qu, G. Huang, X. Zhou, J. Wang and C. Xu, et al. , Adv. Funct. Mater., 2024, 34, 2410406 CrossRef CAS.
- L. Zhu, J.-Y. Xie, G.-M. Zhou, D.-A. Zhang and A. Du, Solid State Ionics, 2023, 398, 116274 CrossRef CAS.
- D. Deb and G. Sai Gautam, J. Mater. Res., 2022, 37, 3169–3196 CrossRef CAS.
- B. Schwaighofer, M. A. Gonzalez, M. R. Johnson, J. S. Evans and I. R. Evans, Chem. Mater., 2025, 37, 3575–3593 CrossRef CAS.
- S. Wang, J. Zhang, O. Gharbi, V. Vivier, M. Gao and M. E. Orazem, Nat. Rev. Methods Primers, 2021, 1, 41 CrossRef CAS.
- R. J. Clément, P. G. Bruce and C. P. Grey, J. Electrochem. Soc., 2015, 162, A2589 Search PubMed.
- S. D. Kang and W. C. Chueh, J. Electrochem. Soc., 2021, 168, 120504 CrossRef.
- P. Heitjans and J. Kärger, Diffusion In Condensed Matter: Methods, Materials, Models, Springer Science & Business Media, 2006 Search PubMed.
- P. Hohenberg and W. Kohn, Phys. Rev., 1964, 136, B864 Search PubMed.
- W. Kohn and L. J. Sham, Phys. Rev., 1965, 140, A1133 CrossRef.
- G. Henkelman, B. P. Uberuaga and H. Jónsson, J. Chem. Phys., 2000, 113, 9901–9904 Search PubMed.
- R. Devi, B. Singh, P. Canepa and G. Sai Gautam, npj Comput. Mater., 2022, 8, 160 Search PubMed.
- D. Frenkel and B. Smit, Understanding Molecular Simulation: From Algorithms To Applications, Elsevier, 2023 Search PubMed.
- X. He, Y. Zhu, A. Epstein and Y. Mo, npj Comput. Mater., 2018, 4, 18 Search PubMed.
- Z. Rong, D. Kitchaev, P. Canepa, W. Huang and G. Ceder, J. Chem. Phys., 2016, 145, 074112 CrossRef PubMed.
- A. Duval, S. V. Mathis, C. K. Joshi, V. Schmidt, S. Miret, F. D. Malliaros, T. Cohen, P. Lio, Y. Bengio and M. Bronstein, arXiv, 2023, preprint arXiv:2312.07511, DOI:10.48550/arXiv.2312.07511.
- O. T. Unke, S. Chmiela, H. E. Sauceda, M. Gastegger, I. Poltavsky, K. T. Schutt, A. Tkatchenko and K.-R. Muller, Chem. Rev., 2021, 121, 10142–10186 CrossRef CAS PubMed.
- J. Choi, G. Nam, J. Choi and Y. Jung, JACS Au, 2025, 5, 1499–1518 CrossRef CAS PubMed.
- F. L. Thiemann, N. O’neill, V. Kapil, A. Michaelides and C. Schran, J. Phys.: Condens. Matter, 2024, 37, 073002 CrossRef PubMed.
- X. Fu, Z. Wu, W. Wang, T. Xie, S. Keten, R. Gomez-Bombarelli and T. Jaakkola, arXiv, 2022, preprint arXiv:2210.07237, DOI:10.48550/arXiv.2210.07237.
- R. Jacobs, D. Morgan, S. Attarian, J. Meng, C. Shen, Z. Wu, C. Y. Xie, J. H. Yang, N. Artrith and B. Blaiszik, et al. , Curr. Opin. Solid State Mater. Sci., 2025, 35, 101214 CrossRef CAS.
- S. Ju, J. You, G. Kim, Y. Park, H. An and S. Han, Digital Discovery, 2025, 4, 1544–1559 RSC.
- H. Kang, T. Lu, Z. Qi, J. Guo, S. Meng and M. Liu, AI for Sci., 2025, 1, 015004 CrossRef.
- B. Focassio, L. P. M. Freitas and G. R. Schleder, ACS Appl. Mater. Interfaces, 2024, 17, 13111–13121 CrossRef PubMed.
- A. Loew, D. Sun, H.-C. Wang, S. Botti and M. A. Marques, npj Comput. Mater., 2025, 11, 178 CrossRef CAS.
- S. Mannan, V. Bihani, C. Gonzales, K. L. K. Lee, N. N. Gosvami, S. Ranu, S. Miret and N. Krishnan, arXiv, 2025, preprint arXiv:2508.05762, DOI:10.48550/arXiv.2508.05762.
- H. Yu, M. Giantomassi, G. Materzanini, J. Wang and G.-M. Rignanese, Mater. Genome Eng. Adv., 2024, 2, e58 CrossRef CAS.
- J. Riebesell, R. E. Goodall, P. Benner, Y. Chiang, B. Deng, G. Ceder, M. Asta, A. A. Lee, A. Jain and K. A. Persson, Nat. Mach. Intell., 2025, 7, 836–847 CrossRef.
- Q. Zhao, Y. Han, D. Zhang, J. Wang, P. Zhong, T. Cui, B. Yin, Y. Cao, H. Jia and C. Duan, Adv. Sci., 2025, 12(34), e06240 CrossRef CAS PubMed.
- V. Bihani, S. Mannan, U. Pratiush, T. Du, Z. Chen, S. Miret, M. Micoulaut, M. M. Smedskjaer, S. Ranu and N. A. Krishnan, Digital Discovery, 2024, 3, 759–768 RSC.
- I. Batatia, P. Benner, Y. Chiang, A. M. Elena, D. P. Kovács, J. Riebesell, X. R. Advincula, M. Asta, M. Avaylon and W. J. Baldwinet al., arXiv, 2023, preprint arXiv:2401.00096, DOI:10.48550/arXiv.2508.05762.
- I. Batatia, S. Batzner, D. P. Kovács, A. Musaelian, G. N. Simm, R. Drautz, C. Ortner, B. Kozinsky and G. Csányi, Nat. Mach. Intell., 2025, 7, 56–67 CrossRef PubMed.
- Y. Park, J. Kim, S. Hwang and S. Han, J. Chem. Theor. Comput., 2024, 20, 4857–4868 CrossRef CAS PubMed.
- J. Kim, J. Kim, J. Kim, J. Lee, Y. Park, Y. Kang and S. Han, J. Am. Chem. Soc., 2024, 147, 1042–1054 CrossRef PubMed.
- M. Neumann, J. Gin, B. Rhodes, S. Bennett, Z. Li, H. Choubisa, A. Hussey and J. Godwin, arXiv, 2024, preprint arXiv:2410.22570, DOI:10.48550/arXiv.2410.22570.
- B. Rhodes, S. Vandenhaute, V. Šimkus, J. Gin, J. Godwin, T. Duignan and M. Neumann, arXiv, 2025, preprint arXiv:2504.06231, DOI:10.48550/arXiv.2504.06231.
- B. Deng, P. Zhong, K. Jun, J. Riebesell, K. Han, C. J. Bartel and G. Ceder, Nat. Mach. Intell., 2023, 5, 1031–1041 CrossRef.
- C. Chen and S. P. Ong, Nat. Comput. Sci., 2022, 2, 718–728 CrossRef PubMed.
- R. Devi, A. Balasubramanian, K. T. Butler and G. S. Gautam, Sci. Data, 2025, 1922 CrossRef CAS PubMed.
- R. Devi, A. Balasubramanian, K. T. Butler and G. Sai Gautam, DFT-Neb-Migration-Barrier-Dataset V1.0, 2025, DOI:10.5281/zenodo.17240095.
- J. P. Perdew, K. Burke and M. Ernzerhof, Phys. Rev. Lett., 1996, 77, 3865 CrossRef CAS PubMed.
- D. B. Tekliye, A. Kumar, X. Weihang, T. D. Mercy, P. Canepa and G. Sai Gautam, Chem. Mater., 2022, 34, 10133–10143 CrossRef CAS.
- D. B. Tekliye and G. S. Gautam, J. Mater. Chem. A, 2024, 12, 18993–19007 RSC.
- D. Deb and G. Sai Gautam, Chem. Mater., 2024, 36, 11892–11904 CrossRef CAS.
- A. H. Larsen, J. J. Mortensen, J. Blomqvist, I. E. Castelli, R. Christensen, M. Dułak, J. Friis, M. N. Groves, B. Hammer and C. Hargus, et al. , J. Phys.: Condens. Matter, 2017, 29, 273002 CrossRef PubMed.
- A. Jain, S. P. Ong, G. Hautier, W. Chen, W. D. Richards, S. Dacek, S. Cholia, D. Gunter, D. Skinner and G. Ceder, et al. , APL Mater., 2013, 1, year Search PubMed.
- B. Deng, Z. Peichen, K. Jun, R. Janosh, K. Han, C. Bartel and C. Gerbrand, Figshare, 2023, DOI:10.6084/m9.figshare.23713842.v2.
- S. Batzner, A. Musaelian, L. Sun, M. Geiger, J. P. Mailoa, M. Kornbluth, N. Molinari, T. E. Smidt and B. Kozinsky, Nat. Commun., 2022, 13, 2453 CrossRef CAS PubMed.
- L. Barroso-Luque, M. Shuaibi, X. Fu, B. M. Wood, M. Dzamba, M. Gao, A. Rizvi, C. L. Zitnick and Z. W. Ulissi, arXiv, 2024, preprint arXiv:2410.12771, DOI:10.48550/arXiv.2410.12771.
- J. Schmidt, T. F. Cerqueira, A. H. Romero, A. Loew, F. Jäger, H.-C. Wang, S. Botti and M. A. Marques, Mater. Today Phys., 2024, 48, 101560 CrossRef.
- S. Smidstrup, A. Pedersen, K. Stokbro and H. Jónsson, J. Chem. Phys., 2014, 140, 214106 CrossRef PubMed.
- E. L. Kolsbjerg, M. N. Groves and B. Hammer, J. Chem. Phys., 2016, 145, 094107 CrossRef PubMed.
- C. Broyden, J. Inst. Math. Appl., 1970, 6, 75–90 Search PubMed.
- R. Fletcher, Comput. J., 1970, 13, 317–322 CrossRef.
- D. Goldfarb, Math. Comput., 1970, 24, 23–26 CrossRef.
- D. F. Shanno, Math. Comput., 1970, 24, 647–656 CrossRef.
- M. O'Keeffe, Acta Crystallogr., Sect. A, 1979, 35, 772–775 CrossRef.
- S. P. Ong, W. D. Richards, A. Jain, G. Hautier, M. Kocher, S. Cholia, D. Gunter, V. L. Chevrier, K. A. Persson and G. Ceder, Comput. Mater. Sci., 2013, 68, 314–319 CrossRef CAS.
- Z. Rong, R. Malik, P. Canepa, G. Sai Gautam, M. Liu, A. Jain, K. Persson and G. Ceder, Chem. Mater., 2015, 27, 6016–6021 CrossRef CAS.
- V. I. Anisimov, J. Zaanen and O. K. Andersen, Phys. Rev. B: Condens. Matter Mater. Phys., 1991, 44, 943 CrossRef CAS PubMed.
- S. L. Dudarev, G. A. Botton, S. Y. Savrasov, C. Humphreys and A. P. Sutton, Phys. Rev. B: Condens. Matter Mater. Phys., 1998, 57, 1505 CrossRef CAS.
- M. Liu, Z. Rong, R. Malik, P. Canepa, A. Jain, G. Ceder and K. A. Persson, Energy Environ. Sci., 2015, 8, 964–974 RSC.
- J. Nocedal, Math. Comput., 1980, 35, 773–782 CrossRef.
- E. Bitzek, P. Koskinen, F. Gähler, M. Moseler and P. Gumbsch, Phys. Rev. Lett., 2006, 97, 170201 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |