Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Artificial intelligence in the discovery and design of molecular semiconductors: a systematic review
Malin
Zollner
 a,
Yashar
Moshfeghi
a,
Yashar
Moshfeghi
 b and
Tahereh
Nematiaram
b and
Tahereh
Nematiaram
 *a
*a
aDepartment of Pure and Applied Chemistry, University of Strathclyde, 295 Cathedral Street, Glasgow G1 1XL, UK. E-mail: tahereh.nematiaram@strath.ac.uk
bDepartment of Computer and Information Sciences, University of Strathclyde, 26 Richmond Street, Glasgow G1 1XH, UK
First published on 25th February 2026
Abstract
Artificial intelligence (AI) is rapidly transforming the discovery and design of molecular semiconductors by linking chemical structure to electronic function with unprecedented speed and accuracy. These materials underpin flexible, lightweight, and sustainable optoelectronic technologies, yet their optimisation has been limited by the immense chemical search space and the cost of exhaustive experimentation and quantum-chemical calculations. This systematic review presents a comprehensive, PRISMA-guided analysis of 237 studies published between 2010 and 2025 that apply AI and machine learning to molecular semiconductor research. The literature is organised into four interconnected domains: electronic structure and spectroscopic properties, photoactive materials, emissive materials, and charge transport. Across these areas, AI models have achieved near quantum-level precision in predicting key electronic and optical properties, enabled the generative design of high-efficiency photoactive and emissive compounds, and accelerated multiscale simulations of charge mobility. The review identifies major trends toward hybrid, data-efficient, and physics-informed learning frameworks while highlighting persistent barriers related to data quality, benchmark inconsistency, and limited interpretability. By consolidating diverse methodologies and findings, this work establishes a unified perspective on how AI can drive reproducible, scalable, and autonomous discovery of molecular semiconductors for next-generation electronic and photonic technologies.
1 Introduction
Molecular semiconductors (MSCs) form the materials foundation of flexible, lightweight, and solution-processable optoelectronic technologies by enabling precise control over electronic structure at the level of individual molecules. These π-conjugated organic compounds support frontier-orbital-mediated charge transport and optoelectronic excitation, making them central to organic photovoltaics (OPVs),1 organic light-emitting diodes (OLEDs),2 and organic field-effect transistors (OFETs).3 Compared with inorganic semiconductors, MSCs can be processed at low temperatures, are compatible with large-area printing, and require reduced material consumption, collectively supporting scalable and potentially more sustainable device architectures.4,5In this review, the term MSC refers specifically to chemically discrete organic semiconductors with well-defined molecular structures and molecular weights, whose optoelectronic behaviour is governed by individual molecular units rather than by indefinitely repeating polymeric chains. This definition encompasses small molecules and finite oligomers when treated as isolated, countable chemical entities, while excluding extended polymeric or crosslinked systems whose properties are dominated by chain-length distributions, polydispersity, or macromolecular disorder.
This chemical discreteness distinguishes MSCs from polymeric systems and enables systematic interrogation of structure–property–performance relationships with molecular-level resolution.6–8 Such resolution is essential because device performance emerges from a highly non-linear interplay between molecular structure, solid-state organisation, and interfacial energetics, where even subtle chemical modifications can strongly influence energy-level alignment, packing motifs, exciton dynamics, and charge-transport behaviour.9–11 Elucidating and controlling these relationships remains a central challenge in MSC design.12
Despite decades of progress, rational optimisation of MSCs remains constrained by the vastness of chemical space and the complexity of structure–function coupling. Historically, advances have relied on chemically intuitive strategies such as extending π-conjugation, introducing donor–acceptor architectures, or modifying side chains to tune packing, solubility, and mobility.13–17 These approaches have delivered OPV power conversion efficiencies exceeding 19%18 and OFET mobilities approaching those of amorphous silicon,19 yet discovery remains slow and resource-intensive. The combinatorial explosion of synthetically accessible molecules severely limits exhaustive experimental exploration.20 Predictive and computationally efficient design frameworks are therefore essential.
Computational chemistry has long provided mechanistic insight into MSC electronic structure and optoelectronic behaviour. Quantum-chemical methods, including density functional theory (DFT)21 and post-Hartree–Fock approaches,22 enable accurate prediction of frontier orbital energies, excited states, and charge-transport descriptors.23–25 High-throughput virtual screening has extended these capabilities to large molecular libraries,26–28 but the computational cost of quantum methods remains prohibitive for comprehensive chemical-space coverage. This limitation has driven a shift toward data-driven discovery paradigms.
Artificial intelligence (AI) now plays a central role in MSC research by enabling rapid and scalable prediction of electronic and optoelectronic properties at near-quantum accuracy and with orders-of-magnitude reduced computational cost.29–31 Machine-learning (ML) models, including neural networks,32 tree-based ensembles,33 and kernel methods,34 have demonstrated reliable prediction of HOMO–LUMO gaps, reorganisation energies, excitation energies, and charge mobilities when trained on experimental or computational datasets.35–37 Advances in molecular representation learning, particularly graph-based and message-passing neural networks,38,39 have further improved data efficiency and model transferability. Beyond forward prediction, generative models and optimisation frameworks increasingly enable inverse molecular design, active learning, and closed-loop discovery pipelines that couple ML with computation and experiment.40–42
Despite these advances, the maturation of AI-driven MSC design is constrained by persistent challenges in data quantity and quality, standardisation, and interpretability. Available datasets are often sparse, biased toward high-performing systems, and inconsistently curated, with limited reporting of negative results or experimental metadata.43–45 The absence of unified benchmarks and validation protocols complicates cross-study comparison,46 while many high-performing neural architectures offer limited physical interpretability.47–49 Physics-informed learning strategies and explainable representations offer promising directions but remain under active development.50,51
Motivated by these challenges, this review provides a systematic and molecular-semiconductor-focused synthesis of AI-driven research in small-molecule organic semiconductors. Using a Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA)52 framework, we analyse 237 peer-reviewed studies published between January 2010 and April 2025. The review is organised into four interconnected domains reflecting the progression from molecular electronic structure to device-level function: (1) electronic structure and spectroscopic properties; (2) photoactive materials; (3) emissive materials; and (4) charge transport. By critically consolidating methodologies, datasets, validation practices, and experimental outcomes across these domains, this review establishes a coherent perspective on the current capabilities and limitations of AI in MSC discovery and highlights pathways toward reproducible, interpretable, and scalable molecular design paradigms.
2 Methodology
2.1 Search strategy and inclusion criteria
A systematic literature search was conducted in accordance with PRISMA guidelines.52 The objective was to identify studies employing AI in the discovery or design of MSCs. The search covered publications from January 2010 to April 2025, reflecting the period during which data-driven methods gained prominence in materials science.Multiple bibliographic databases and publisher platforms were queried, including Web of Science, Scopus, PubMed, and major publisher repositories (ACS, RSC, Elsevier, Nature Publishing Group, and Wiley Online Library). Search strings combined keywords such as “organic semiconductor*”, “molecular semiconductor*”, “small molecule”, “machine learning”, “artificial intelligence”, “deep learning”, “neural network”, “reinforcement learning”, “data-driven”, “organic photovoltaic*”, “organic light-emitting diode*”, “organic electronic*”, “OPV*”, “OFET*”, and “OLED*”. To capture emerging work, we also searched major preprint archives (arXiv) for relevant unpublished studies. The exact search strings are available in the SI (Section 1).
To maintain focus on MSCs, we applied exclusion filters to remove studies dealing exclusively with extended polymeric systems. Studies addressing finite oligomers were retained when these systems were treated as discrete, well-defined molecular entities. Works covering both polymeric and molecular systems were included only when separate analyses or explicit discussions relevant to the molecular or finite-oligomer regime were provided.
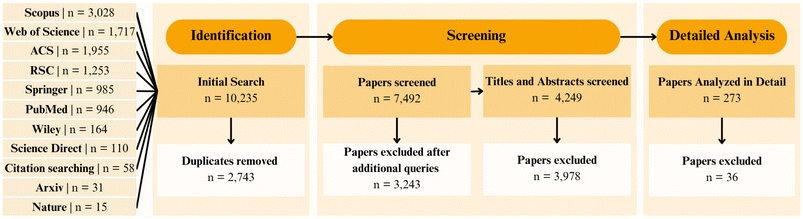
As summarised in Fig. 1, the initial search identified 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 235 records, including journal articles, conference proceedings, and preprints. After removing 2743 duplicates, 7492 unique entries remained. Title-level screening excluded 3243 clearly irrelevant works, such as those focused on inorganic semiconductors, purely theoretical investigations lacking ML components, or device engineering studies unrelated to materials design. The remaining 4249 records were advanced to abstract screening. Abstracts and titles were evaluated against the following inclusion criteria:
235 records, including journal articles, conference proceedings, and preprints. After removing 2743 duplicates, 7492 unique entries remained. Title-level screening excluded 3243 clearly irrelevant works, such as those focused on inorganic semiconductors, purely theoretical investigations lacking ML components, or device engineering studies unrelated to materials design. The remaining 4249 records were advanced to abstract screening. Abstracts and titles were evaluated against the following inclusion criteria:
• Focus on organic semiconductors composed of chemically discrete small molecules or finite oligomers, excluding extended polymeric systems.
• Explicit use of AI or ML methods in materials discovery, screening, design, or property prediction.
• Investigation of properties or performance metrics relevant to optoelectronic functionality (e.g. charge transport, optical absorption, frontier orbital energies, device efficiency, or stability).
• Publication in English.
All records meeting these criteria were retained for full-text review and data extraction.
2.2 Study selection and data extraction
Full-text screening was performed on 273 articles that passed the initial inclusion criteria. Screening and data extraction were carried out independently by two reviewers to minimise selection bias; discrepancies were resolved through discussion, and unresolved cases were adjudicated by a third reviewer. Each article was examined to confirm eligibility and to extract key metadata, including:• Application domain (e.g., OPV, OFET, OLED).
• AI/ML techniques employed (algorithms, descriptors, and model architectures).
• Dataset size and provenance (experimental, computational, or hybrid).
• Target properties or performance metrics predicted or optimised.
• Validation approaches and reported limitations.
Particular attention was given to the nature of the data (experimental vs. simulated) and the extent of experimental validation for AI-generated candidates. During this stage, 36 studies were excluded for insufficient relevance (e.g., works that mentioned ML only superficially or lacked substantive implementation). The final dataset comprised 237 studies, which form the analytical foundation of this review.
2.3 Quality assessment
Given the rapid pace of research in AI-driven materials discovery, several included manuscripts, particularly those from 2025, were still preprints at the time of writing. Each study, whether peer-reviewed or preprint, was critically assessed for methodological soundness and reproducibility. Reported claims were cross-checked wherever possible, particularly when multiple studies presented comparable findings or when later work superseded earlier “state-of-the-art” results.Due to the diversity of AI methodologies and application targets, this review does not conduct a formal quantitative meta-analysis. Instead, it adopts a qualitative and comparative framework that identifies consensus trends, recurring challenges, and divergent findings across the literature. This approach offers a rigorous yet flexible synthesis, well-suited to the evolving and interdisciplinary nature of AI-driven MSC research.
To ensure transparency and reproducibility of the systematic review process, the SI includes a PRISMA flow diagram detailing study identification, screening, and inclusion; a completed PRISMA 2020 checklist; and a comprehensive database summarising all 237 included studies with extracted metadata, including application domain, AI methodology, target properties, data provenance, validation strategy, and experimental verification where available.
3 Landscape of AI applications in molecular semiconductor research
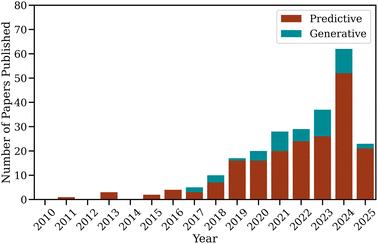
To contextualise the domain-specific analyses that follow, this section surveys the overall landscape of AI applications in MSC research. It outlines the evolution of publication activity, algorithmic strategies, and global research distribution, illustrating how the field has progressed from early predictive modelling to emerging paradigms of generative and autonomous molecular design.As shown in Fig. 2, research activity in this area has grown exponentially over the past decade. Fewer than ten studies per year appeared before 2019, but this number exceeded fifty by 2024. Two major growth phases can be identified. The first, emerging around 2018–2019, coincided with the widespread adoption of supervised learning algorithms and graph-based molecular representations,53–55 which substantially improved predictive accuracy and data efficiency. The second, during 2023–2024, was driven by advances in transfer learning and active learning frameworks,56–61 along with the introduction of large foundation models capable of cross-domain generalisation. Although generative design currently accounts for a smaller proportion of publications, its steady growth since 2020 marks a conceptual turning point, from passive property prediction to inverse design and closed-loop discovery. This shift reflects a broader transition in the field, from descriptive modelling toward proactive and autonomous exploration of chemical space.
A central driver of this transition is the significant reduction in computational cost achieved by ML surrogates relative to first-principles calculations. For example, in the transfer-learning D-MPNN framework reported by Nie et al.,56 prediction of HOMO and LUMO energy levels for candidate OPV molecules requires approximately 1–1.2 seconds per molecule, whereas the corresponding DFT calculations require between 2 and 4 days of wall-clock time per molecule, depending on molecular size and functionalisation. This represents an acceleration of approximately five orders of magnitude on a per-molecule basis. Crucially, this speedup applies to routine model inference rather than to active-learning cycles. Once trained, the model enables near-instantaneous screening of thousands of molecules that would otherwise require months of cumulative quantum-chemical computation. Comparable reductions in computational cost are reported across ML-driven MSC studies, where trained models replace explicit quantum-chemical evaluations during large-scale virtual screening and allow chemical spaces comprising 103–106 candidates to be explored at negligible marginal computational cost.62–64
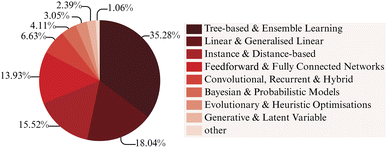
Complementing this temporal expansion, Fig. 3 summarises the algorithmic diversity across the reviewed literature. As can be seen, tree-based methods and ensemble learners constitute the dominant class of algorithms applied in MSC research, representing approximately 35% of the surveyed studies. Their prevalence underscores the enduring reliability of ensemble techniques such as random forests,65–67 gradient boosting models,68–70 bagging60,71,72 and decision tree models,61,73,74 which combine strong predictive performance with transparent feature importance analysis. These methods have proven particularly effective for modestly sized datasets that characterise much of the available experimental and computational literature on MSCs. Linear and generalised linear models, including linear regression,75–77 lasso,78–80 ridge regression69,81,82 elastic net regression,77,79,83 and orthogonal matching pursuit60,61,80 form the second largest category, accounting for around 18% of the total. Their simplicity, interpretability, and computational efficiency make them valuable as baseline predictors and as tools for mechanistic insight. Feedforward and fully connected networks make up roughly 14% reported applications. This group spans traditional multilayer perceptrons,55,84,85 general neural networks,86–88 deep learning,89–91 and feedforward neural networks.92,93 Instance- and distance-based algorithms, including support vector machines,94–96k-nearest neighbour,97–99 and kernel ridge regression,100–102 comprise about 9% of the literature. Although their relative prominence has declined in recent years, they remain highly competitive in low-data regimes and continue to serve as strong benchmarks for molecular property prediction. About 7% of the literature utilises convolutional, recurrent or hybrid networks. These include (convolutional) recurrent neural networks,103–105 message passing neural networks,92,106,107 as well as more advanced architectures such as graph (convolutional) neural networks that directly encode molecular connectivity and electronic interactions, thereby capturing structure–property relationships in a physically meaningful way. Bayesian and probabilistic models account for approximately 4% of studies. These methods, which include Gaussian process regression,108–110 and Bayesian optimisation,60,111,112 are particularly valuable for uncertainty quantification and for guiding active learning workflows that iteratively refine training data through targeted experimentation or computation. Evolutionary and optimisation-based techniques contribute around 3%, typically in applications involving multi-objective molecular design or inverse optimisation of optoelectronic properties.113–115 Generative models, such as variational autoencoders,105,116,117 generative adversarial networks,118 and generative pretrained transformers,119,120 currently represent about 2% of the reviewed work, reflecting a rapidly expanding area of research focused on de novo molecular generation. The remaining fraction, about 1%, includes hybrid and miscellaneous algorithms that do not align with conventional classifications but often combine multiple paradigms within integrated discovery pipelines.
The overall distribution reveals a field that remains grounded in established supervised learning methods while increasingly incorporating probabilistic reasoning, generative modelling, and hybrid optimisation strategies. This diversification signifies a methodological transition from purely predictive analytics toward adaptive and exploratory frameworks capable of autonomous molecular discovery. The convergence of interpretable, data-efficient, and generative approaches is gradually redefining the computational landscape of MSC research, promoting workflows that are not only accurate and scalable but also transparent and physically grounded.
Fair and transparent performance evaluation remains crucial for comparing results across this heterogeneous literature. In predictive modelling, regression and classification tasks employ complementary metrics. Regression models are typically evaluated using the coefficient of determination (R2), root-mean-square error (RMSE), and mean absolute error (MAE), which collectively quantify accuracy and precision relative to the reference data.121 Classification models rely on accuracy, precision, recall, and the F1 score to evaluate categorical performance,122 while receiver operating characteristic curves and the area under the curve provide additional measures of discriminative power.123,124 In generative modelling, evaluation extends beyond numerical accuracy to assess the chemical and functional realism of generated molecules. Common metrics include molecular validity (the fraction of chemically plausible structures), uniqueness (non-duplicate outputs), and novelty (the proportion of molecules not present in the training data).125,126 Many recent studies further incorporate task-specific objectives, such as predicted property enhancement, synthetic accessibility, or thermodynamic stability, to ensure that generated candidates are both realistic and experimentally meaningful.92,127,128 These practices reflect the field's gradual movement toward quantitative, multi-objective benchmarks that facilitate reproducibility and cross-study comparison.
The geographical distribution of studies, shown in Fig. 4, underscores the international and interdisciplinary nature of AI-driven MSC research. China leads with 87 publications, followed by the United States (39), South Korea (28), Japan (27), and Saudi Arabia (27). Other major contributors include Germany, Pakistan, and the United Kingdom, each producing more than twenty studies. Additional contributions from Australia, Brazil, Canada, Egypt, France, India, Italy, Singapore, Spain, Switzerland, Taiwan, and Turkey further demonstrate the breadth of global engagement. At the same time, emerging outputs from Africa, South America, and Eastern Europe indicate an expanding international presence. The combination of increasing global participation, methodological diversification, and integration of AI across the molecular design pipeline highlights a field transitioning from exploratory adoption toward systematic, data-driven discovery frameworks.
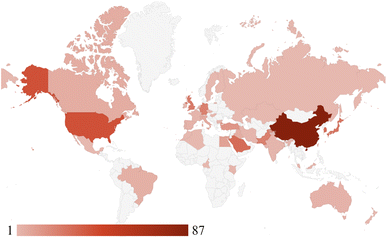 | ||
| Fig. 4 Geographical distribution of studies included in the systematic review. Darker shading corresponds to a higher number of publications originating from each country. | ||
Building on these observations, the following sections examine how AI has been applied across key facets of MSC research. These areas, spanning molecular electronic structure, photoactive and emissive materials, and charge-transport phenomena, capture the progression from molecular design to device-level function. The discussion focuses on how algorithmic strategies, data practices, and validation approaches have evolved within each domain, revealing common challenges and emerging opportunities that define the current trajectory of AI-driven discovery.
3.1 Electronic structure and spectroscopic properties
As shown in Fig. 5, the prediction and design of molecules with tailored electronic and spectroscopic characteristics constitute the most active area of AI application in MSC research. A total of 123 studies were identified in this category, including 87 focused on predictive modelling and 36 employing generative design strategies.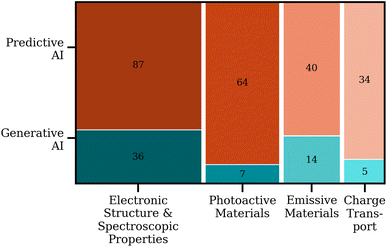 | ||
| Fig. 5 Distribution of predictive and generative AI applications across major research domains in MSCs studies. The size of each block corresponds to the number of publications within that domain. | ||
Electronic-structure parameters such as the frontier molecular orbital energies (HOMO and LUMO), bandgaps, optical absorption spectra, and exciton binding energies (Eb) govern essential photophysical processes, including charge separation and light absorption and emission.129,130 Traditionally, these quantities are evaluated using first-principles quantum-chemical methods, most notably DFT131–133 and many-body perturbation theory within the GW–BSE formalism.134 While these approaches offer high accuracy and physical interpretability, their computational cost limits their use in large-scale screening or high-throughput discovery.
ML provides a scalable alternative, capable of capturing complex, non-linear relationships between molecular structure and target properties with near-first-principles precision at far lower computational cost. Recent progress has expanded its scope from predictive modelling to generative molecular design, where algorithms autonomously propose new chemical structures optimised for desired optoelectronic performance. This shift marks a broader transition from data-driven screening of known molecules toward proactive exploration of chemical space and automated discovery of functional materials.
The following analysis focuses on two complementary paradigms within this domain: (1) predictive frameworks that estimate electronic and spectroscopic properties from molecular representations, and (2) generative approaches that create new molecules optimised for target functionalities. These directions illustrate how AI accelerates both the understanding and the design of high-performance MSCs.
Montavon et al. (2013)135 employed Coulomb matrix representations with deep neural networks to predict HOMO and LUMO energies for approximately 7200 molecules, achieving MAEs of 0.15 and 0.12 eV, respectively. Using alternative fixed descriptors, Pereira et al. (2017)136 demonstrated that neural networks trained on simple molecular features could reach comparable accuracy, with MAEs between 0.1 and 0.2 eV. Scaling descriptor-based learning to substantially larger datasets, Pyzer-Knapp et al. (2015)84 trained multilayer perceptrons on approximately 250![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 molecules represented by Morgan fingerprints,137 reaching MAEs of 0.028 eV for HOMO energies and 0.12 eV for LUMO energies. These studies showed that low-order descriptor encodings can support accurate electronic-property prediction when training and target chemical spaces are closely aligned.
000 molecules represented by Morgan fingerprints,137 reaching MAEs of 0.028 eV for HOMO energies and 0.12 eV for LUMO energies. These studies showed that low-order descriptor encodings can support accurate electronic-property prediction when training and target chemical spaces are closely aligned.
To address limitations in chemical diversity and transferability, subsequent work introduced representations that explicitly encode higher-order structural information or learn it directly from molecular geometry. Many-body tensor representations (MBTRs) capture element-resolved distributions of interatomic distances and angles, while graph neural networks learn local chemical environments through message passing, enabling more expressive and transferable descriptions of molecular structure. Stuke et al. (2019)101 applied kernel ridge regression (KRR)138 with many-body tensor representations to predict HOMO energies across datasets of increasing chemical complexity, including QM9,139 amino acids and dipeptides,140 and optoelectronic compounds from the Cambridge Structural Database (CSD).141 The resulting MAEs, 0.086 eV (QM9), 0.100 eV (amino acids), and 0.173 eV (CSD), highlighted both the improved expressiveness of higher-order descriptors and the persistent challenges of cross-domain generalisation. More recently, Gaul et al. (2024)142 employed a SchNet-based graph neural network143 with Set2Set aggregation, achieving RMSE errors of 0.063 eV for HOMO energies and 0.059 eV for LUMO energies, demonstrating the advantages of geometry-aware, learned representations.
Beyond ground-state properties, ML has also been applied to model complex excited-state phenomena that underpin optoelectronic behaviour. Schröder et al. (2019)144 developed a hybrid simulation combining ML and tensor-network methods to study singlet fission in a pentacene dimer, a process central to enhancing photovoltaic efficiency. By coupling time-dependent DFT (TD-DFT) with ML-based clustering of vibrational modes, they simulated non-Markovian quantum dynamics and identified specific vibrational groups that promote efficient fission. Similarly, Liu et al. (2022)145 employed the SISSO algorithm to derive interpretable models for singlet-fission thermodynamics, achieving RMSEs below 0.2 eV and identifying three new candidate crystals (BCPP, TBPT, DPNP). Gao et al. (2025)146 extended this strategy to model singlet and triplet excitation energies and exciton binding energies in polycyclic aromatic hydrocarbons, reaching MAEs around 0.2 eV. These studies highlight how physics-informed ML accelerates the identification of materials with targeted excited-state characteristics while retaining interpretability and physical grounding.
Recent research has increasingly prioritised model generalisation and data efficiency, two enduring challenges in AI-driven property prediction. Because high-quality training data remain limited, particularly for experimentally validated molecules, several strategies have emerged to leverage existing datasets more effectively. Transfer learning147 has proven particularly valuable, but reported performance gains depend strongly on data provenance, noise levels, and evaluation metrics. Jeong et al. (2022)148 pretrained a graph convolutional network on experimentally derived optical data and fine-tuned it using 3026 experimentally measured HOMO/LUMO values spanning diverse solvents and solid-state environments, achieving MAEs of 0.050–0.065 eV. Notably, these errors are comparable to or smaller than the reported experimental uncertainties themselves (0.089 eV for HOMO and 0.112 eV for LUMO), placing the model performance near the intrinsic noise floor of the measurements. In contrast, Peng et al. (2024)57 pretrained models on 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 626 DFT-computed frontier orbital energies and fine-tuned them on 1198 experimental measurements, reporting correlation coefficients of 0.75 (HOMO) and 0.84 (LUMO) alongside MAEs of 0.094 and 0.117 eV, respectively. The absolute errors remain substantially larger than those achieved by Jeong et al. (2022),148 reflecting both the smaller experimental fine-tuning set and the propagation of DFT-specific biases into the learned representation. Parallel advances in foundation and language-based models have introduced transferable chemical representations; for example, Xie et al. (2024)120 fine-tuned GPT-3 to classify molecules by frontier orbital energies, attaining accuracies above 90%, despite the model's origin in natural language processing. These developments signal a shift from task-specific featurisation toward generalisable molecular representations that can bridge computational and experimental data regimes.
626 DFT-computed frontier orbital energies and fine-tuned them on 1198 experimental measurements, reporting correlation coefficients of 0.75 (HOMO) and 0.84 (LUMO) alongside MAEs of 0.094 and 0.117 eV, respectively. The absolute errors remain substantially larger than those achieved by Jeong et al. (2022),148 reflecting both the smaller experimental fine-tuning set and the propagation of DFT-specific biases into the learned representation. Parallel advances in foundation and language-based models have introduced transferable chemical representations; for example, Xie et al. (2024)120 fine-tuned GPT-3 to classify molecules by frontier orbital energies, attaining accuracies above 90%, despite the model's origin in natural language processing. These developments signal a shift from task-specific featurisation toward generalisable molecular representations that can bridge computational and experimental data regimes.
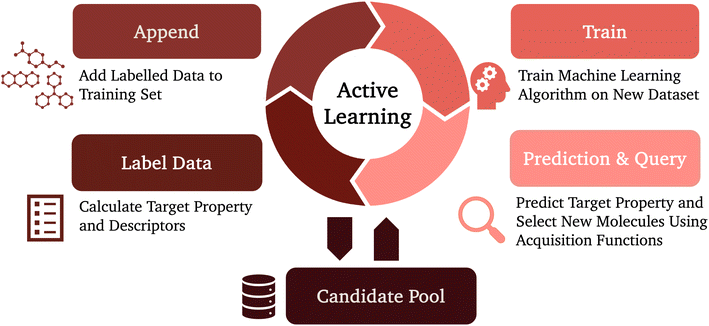
The cost of generating new training data nevertheless remains a major constraint, especially when exploring chemically diverse spaces. Active learning has emerged as a powerful approach to maximise predictive performance while minimising labelling effort. Instead of random sampling, the model iteratively selects the most informative molecules for evaluation, typically those associated with the highest uncertainty or potential improvement, thereby achieving high accuracy with fewer data points (Fig. 6). This approach is particularly well-suited to MSCs, where both DFT calculations and experimental synthesis are resource-intensive. Several recent studies have demonstrated the effectiveness of this approach. Butler et al. (2024)149 employed active learning to train machine-learned interatomic potentials for organic crystal polymorph prediction. By selectively sampling crystal configurations based on model uncertainty, their workflow achieved near-DFT accuracy (RMSE = 1.2 kJ mol−1) with significantly reduced computational effort. Saqib et al. (2024)150 combined active learning with a BRICS-based fragment recombination strategy to generate and screen low-bandgap molecules. Their hist-gradient boosting model achieved RMSE = 0.18 eV and R2 = 0.69 for bandgap prediction across thousands of candidates. The same workflow also predicted UV-vis absorption maxima with RMSE = 42 nm and R2 = 0.703. The model iteratively identified promising candidates while constraining the search to synthetically accessible chemistries, effectively narrowing a large combinatorial space into a tractable and meaningful design region.
These developments collectively signify a transition toward data-efficient, generalisable, and scalable predictive pipelines in AI research. By integrating strategies such as transfer learning, foundation models, and active learning, recent approaches have significantly reduced dependence on costly quantum-chemical and experimental labels. Beyond improving the accuracy and speed of property prediction, these frameworks are establishing the foundation for closed-loop discovery systems, in which molecular generation, screening, and validation occur autonomously within a continuous feedback cycle.
Recent work has demonstrated that these models can be conditioned on diverse quantum-chemical and device-relevant objectives, including frontier orbital energies,151–153 bandgaps,154–156 excited- and charge-transfer-state energies,54,118 and singlet–triplet gaps relevant to emissive materials.157 In several cases, auxiliary constraints such as oscillator strength, or synthetic accessibility are introduced to ensure that generated molecules are not only property-optimal but also physically meaningful and experimentally realisable.115,158 These studies signal a clear evolution from unguided chemical generation toward goal-oriented design strategies that link molecular architecture directly to targeted electronic and optical performance.
To achieve this, a variety of algorithmic frameworks have been employed, including deep generative networks,159 reinforcement learning,160 evolutionary algorithms,161 and inverse-design strategies.162 Many contemporary workflows integrate these approaches with active learning or Bayesian optimisation, forming closed feedback loops that iteratively refine the search process and prioritise molecules with the highest predicted potential. Such frameworks enable both local optimisation within learned chemical spaces and global exploration beyond them.
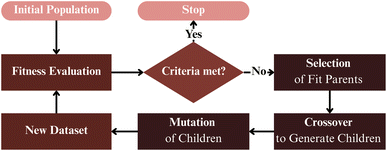
One of the earliest demonstrations of generative molecular design for MSCs was presented by Huwig et al. (2017).151 Using a population-based evolutionary algorithm (see Fig. 7), they optimised the initial population of benzene-core derivatives represented as six-site substitution patterns. Candidate fitness was evaluated using a suite of quantum-chemical descriptors, including the HOMO–LUMO gap, spatial orbital overlap, oscillator strength, and reorganisation energy. Through iterative cycles of selection, crossover, and random mutation, the population converged toward molecules exhibiting narrower bandgaps and enhanced oscillator strengths. Despite its simplicity, this approach demonstrated how evolutionary search can efficiently traverse combinatorial chemical spaces while preserving molecular validity and synthetic feasibility.
Building on this foundation, Kwon et al. (2021)153 developed a hybrid framework that combined a genetic algorithm with deep neural networks trained on a database of approximately 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 molecules with precomputed S1 excitation energies and frontier orbital levels. The optimisation objective, minimisation of the S1 energy, was achieved by combining multiple correlated descriptors within a unified scoring function. This integration of predictive modelling with evolutionary search improved optimisation efficiency and produced molecules with systematically reduced excitation energies, establishing a scalable approach for multi-objective design.
000 molecules with precomputed S1 excitation energies and frontier orbital levels. The optimisation objective, minimisation of the S1 energy, was achieved by combining multiple correlated descriptors within a unified scoring function. This integration of predictive modelling with evolutionary search improved optimisation efficiency and produced molecules with systematically reduced excitation energies, establishing a scalable approach for multi-objective design.
Subsequent advances incorporated property prediction directly into the generation process. Nigam et al. (2024)113 introduced the JANUS framework, which couples a genetic algorithm with neural-network classifiers to guide exploration through chemical space. Using the SELFIES molecular representation and the STONED mutation scheme, the workflow generated over 800![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 candidate molecules and identified more than 10
000 candidate molecules and identified more than 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 exhibiting inverted singlet–triplet (INVEST) gaps and strong oscillator strengths, essential for blue-emitting materials. Wavefunction-based excited-state calculations validated the top candidates, highlighting how classifier-guided evolutionary workflows can balance chemical diversity with targeted optoelectronic optimisation.
000 exhibiting inverted singlet–triplet (INVEST) gaps and strong oscillator strengths, essential for blue-emitting materials. Wavefunction-based excited-state calculations validated the top candidates, highlighting how classifier-guided evolutionary workflows can balance chemical diversity with targeted optoelectronic optimisation.
Alternative strategies have employed reinforcement learning to achieve property-driven generation. Li and Tabor (2023)163 implemented a recurrent neural network agent trained to generate SMILES sequences with reward functions derived from quantum chemical simulations. The agent autonomously designed molecules optimised for excited-state alignment relevant to singlet fission, producing both known and novel anthracene derivatives with favourable electronic configurations. This work exemplifies how physics-informed reinforcement learning can constrain generative exploration to synthetically accessible and functionally relevant molecular regions.
Diffusion-based methods have recently emerged as a robust alternative to traditional generative models. Weiss et al. (2023)156 introduced a guided diffusion framework that integrates gradients from property predictors into the generative trajectory, allowing molecules to be sampled directly along property-optimised directions. Unlike variational autoencoders or generative adversarial networks, diffusion models provide more stable training and broader chemical diversity. Their framework generated structurally novel aromatic compounds with targeted HOMO and LUMO energies, demonstrating the capacity of diffusion models to extrapolate beyond the distribution of training data.
Scalability and efficient chemical-space exploration have also become central concerns. Ohno et al. (2023)164 addressed this challenge with a graph-based molecular generator capable of producing over 4.8 million n-type MSC candidates, of which more than 740![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 exceeded an electron-affinity threshold of 3.0 eV. This large-scale enumeration was enabled by coupling the generator with a graph neural network surrogate model trained to rapidly predict electronic properties, exemplifying how generative and predictive approaches can operate synergistically to expand the accessible chemical landscape.
000 exceeded an electron-affinity threshold of 3.0 eV. This large-scale enumeration was enabled by coupling the generator with a graph neural network surrogate model trained to rapidly predict electronic properties, exemplifying how generative and predictive approaches can operate synergistically to expand the accessible chemical landscape.
Generative frameworks have further evolved to include supramolecular and morphological design, extending beyond molecular composition to structural organisation. Tom et al. (2023)165 used a property-based genetic algorithm to perform the inverse design of tetracene polymorphs optimised for singlet-fission performance. Rather than altering chemical substituents, their algorithm explored three-dimensional crystal packings using a multi-objective fitness function combining thermodynamic stability and theoretical fission rates. The model rediscovered known polymorphs and identified several new low-energy packings with enhanced performance. Fan et al. (2024)98 later introduced a theory-guided evolutionary framework for non-linear optical materials that couples a chemically interpretable group-contribution model with a multistage Bayesian neural network. Mutation operations on donor, acceptor, and bridge fragments were used to optimise first-order hyperpolarisability, and several high-performing candidates were validated using DFT. This hybrid methodology demonstrates how interpretable models and data-driven algorithms can be combined to balance accuracy, efficiency, and physical insight.
Across these studies, a unifying trend is evident such that the generative AI for AIs is becoming increasingly integrated with physics-based reasoning and heuristic search. Evolutionary algorithms remain attractive because they preserve molecular diversity and avoid premature convergence, while surrogate ML models accelerate property evaluation and guide exploration toward promising regions of chemical space. These hybrid frameworks achieve both the rediscovery of known high-performance molecules and the discovery of novel structures with optimised electronic and spectroscopic properties, reflecting the growing maturity of generative AI as a practical tool for AI design.
While these generative approaches demonstrate clear potential for accelerating electronic structure design, their practical translation to functional MSCs remains at an early stage. Among the 36 generative studies identified in this domain, only 4 (11%) reported experimental validation of computationally designed molecules,54,92,128,157 and just 3 (8%) included external validation using independent test sets beyond their training domains.117,166,167 This limited level of validation reflects the broader challenges associated with transferring data-driven predictions to experimentally realised materials, rather than deficiencies of individual methodologies. In addition, many studies rely on relatively small or chemically homogeneous starting datasets, often drawn from specific molecular families used as fragment sources for generation. Such constraints introduce inherent data bias and restrict exploration of genuinely novel regions of chemical space. The absence of reported negative experimental outcomes further suggests the presence of publication bias, which limits insight into failure modes and hampers systematic assessment of model robustness. These structural limitations are examined in greater detail in Section 3.5.
3.2 Photoactive materials
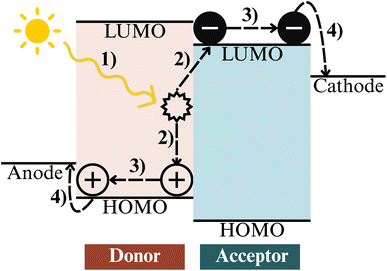
Building on advances in electronic-structure and spectroscopic modelling, recent research has increasingly extended AI methodologies to the design and optimisation of photoactive materials, which convert absorbed light into separated charge carriers. Among these systems, OPVs have emerged as the most intensively investigated platform and now serve as a benchmark for assessing the impact of data-driven discovery in AI research.Fig. 8 illustrates the fundamental photophysical processes governing OPV operation. Upon photoexcitation in the donor layer (1), an exciton is generated and subsequently migrates to the donor–acceptor interface (2), where charge separation occurs. The resulting free carriers, i.e., holes in the donor HOMO and electrons in the acceptor LUMO, are then transported through their respective energy levels (3) and finally collected at the electrodes (4). Each of these steps is influenced by the interplay between molecular electronic structure, interfacial alignment, and nanoscale morphology, which together determine the overall device efficiency.168–170
Despite significant progress, the power-conversion efficiencies of OPVs remain lower than those of inorganic technologies, limited by this intricate coupling between exciton dynamics, charge transport, and recombination.171 Traditional screening frameworks, such as the Scharber equation,172 offer fast, semi-empirical estimates of device performance but fail to capture the non-linear correlations that emerge across these multiple physical scales.114,173,174 The chemical diversity of modern donor–acceptor systems,175 together with the proliferation of non-fullerene acceptors (NFAs),176 has therefore motivated the adoption of AI as a scalable tool capable of learning and predicting the complex dependencies that dictate OPV behaviour.
By learning directly from experimental and computational datasets, AI-based approaches integrate information across molecular, morphological, and device scales. These models link chemical composition and structure to optoelectronic response and operational stability, offering a data-driven route to the rational optimisation of photovoltaic materials and device architectures.177 In doing so, they move beyond empirical design heuristics toward multiscale predictive frameworks that connect molecular chemistry with macroscopic performance.
Across the reviewed literature, 71 studies fall within this domain. The majority (64) employ predictive ML to estimate device-level metrics, most prominently the power-conversion efficiency (PCE), which quantifies the ratio of electrical output to incident solar energy. PCE is determined by three key parameters: the short-circuit current density (JSC), representing the current under zero bias; the open-circuit voltage (VOC), corresponding to the potential difference at zero current; and the fill factor (FF), which measures how closely the current–voltage curve approaches an ideal rectangular shape.178 A smaller subset (7 studies) explores generative strategies that search chemical and interfacial design spaces for new materials optimised for these performance targets.
A substantial body of work has focused on structure-derived descriptors as the primary representation of molecular candidates. Pyzer-Knapp et al. (2015)84 trained neural networks on Morgan fingerprints derived from the Harvard Clean Energy Project, achieving mean absolute errors of approximately 0.28% in power conversion efficiency (PCE) prediction and establishing a widely used benchmark for data-driven screening. Similarly, Sun et al. (2019)91 applied tree-based ensemble models to roughly 1700 donor molecules, demonstrating that molecular fingerprints outperform raw SMILES strings or image-based representations when ranking high-efficiency candidates. Morishita et al. (2024)179 extended this descriptor-based paradigm by combining principal component analysis, random forest feature selection, and support-vector regression to predict JSC across 47 donor–PCBM systems, achieving R2 = 0.64 and using genetic optimisation to propose 250 new donor candidates.
An alternative strategy replaces predefined descriptors with learned molecular representations. Zhang et al. (2025)180 employed graph neural networks operating directly on SMILES strings to screen over 45![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 donor–acceptor pairs, exemplifying an end-to-end representation learning approach that removes manual feature engineering while improving scalability across chemical space.
000 donor–acceptor pairs, exemplifying an end-to-end representation learning approach that removes manual feature engineering while improving scalability across chemical space.
In parallel, several studies have examined the contribution of quantum-chemical electronic descriptors, either alone or in combination with structural features. Sahu et al. (2018)181 augmented electronic descriptor sets with donor–acceptor energetic offsets, achieving R2 ≈ 0.8. Padula et al. (2019)102 demonstrated that integrating frontier orbital energies with structural descriptors within k-nearest-neighbour and kernel ridge regression frameworks enhances predictive accuracy across chemically heterogeneous datasets. However, the utility of electronic descriptors is not universal. Alwadai et al. (2022)71 and Janjua et al. (2022)182 independently showed that purely structural descriptors can outperform frontier orbital energies in PCE prediction, reaching R2 values up to 0.89. This discrepancy likely reflects differences in model expressivity: Padula et al. employed relatively simple algorithms such as k-nearest neighbour and kernel ridge regression, which may benefit from the inclusion of explicit electronic features, whereas the more flexible, nonlinear models used by Alwadai and Janjua (e.g. random forests) are better able to infer relevant electronic structure information implicitly from structural descriptors alone.
Beyond isolated molecular properties, several approaches incorporate descriptors that reflect interfacial, morphological, or processing effects. Yang et al. (2022)183 introduced a multi-fidelity framework that integrates morphology-derived latent variables with low-cost molecular descriptors, enabling consistent performance across distinct donor–acceptor classes. Lee et al. (2024)184 used gradient-boosted decision trees to model the fill factor (FF) of 180 donor–acceptor systems, revealing that high FF values correlate with small HOMO offsets (<0.3 eV) and balanced hole–electron mobilities (1.8 < µh/µe < 3.3). Complementary models of the open-circuit voltage further demonstrated the importance of dielectric and interfacial descriptors for capturing voltage losses.185 Liu et al. (2024)186 applied Gaussian process regression with spectral decomposition and mRMR feature selection to correlate processing parameters with operational lifetime, identifying the Huang–Rhys factor as a predictor of FF decay and the stabilising role of [70]PCBM on morphology and trap density. Vubangsi et al. (2024)187 similarly incorporated dielectric constants and VOC-loss descriptors into XGBoost regressors, improving voltage prediction and revealing dielectric mismatch as a dominant contributor to operational instability.
A subset of studies further integrates ML models with active-learning and automatic fabrication. Du et al. (2021)188 combined Gaussian process regression with a robotic fabrication platform to jointly optimise efficiency and photostability across more than 100 processing conditions within 70 hours. Their analysis identified spectral features such as absorption peak position, amplitude, and ordering as critical indicators of device degradation, while stable configurations favoured thinner active layers and moderate annealing. Almalki et al. (2024)94 employed active learning to optimise non-fullerene OPV fabrication under sparse data regimes, efficiently identifying solvent ratios, annealing temperatures, and film thicknesses that improved PCE. Such closed-loop strategies mark a departure from static prediction toward adaptive experimentation, effectively redefining ML from an analytical tool into a decision-making component of materials discovery.
Finally, ML frameworks have also been applied to stability-focused targets beyond device efficiency. Bornschlegl et al. (2025)109 used Gaussian process regression trained on structural fingerprints to predict UV-C photostability in hole-transport materials, identifying substructural motifs associated with photochemical resilience or degradation and extending data-driven design principles to operational robustness.
These studies collectively demonstrate a clear methodological evolution in ML-guided OPV research, progressing from simple structure-based descriptors toward increasingly expressive representations that integrate electronic, interfacial, and device-level information. Early fingerprint-based models established the feasibility of large-scale screening, while subsequent work showed that the choice of descriptors, whether structural or electronic, can strongly influence predictive performance depending on the application context. More recent approaches that incorporate morphology proxies, dielectric effects, and explicit donor–acceptor pairing have further improved physical interpretability and relevance to device operation. In parallel, advances in representation learning have reduced reliance on manual feature engineering, and closed-loop optimisation frameworks have begun to couple ML models directly with experimental control. These developments mark a transition from static property prediction toward adaptive, multiscale design strategies that more accurately reflect the coupled physical processes governing OPV performance and stability.
The first significant demonstration of generative molecular design for OPVs was reported by Khazaal et al. (2020),154 who introduced the PooMa (“Poor Man's Materials Optimization”) framework. This system combined a genetic algorithm with a density-functional tight-binding evaluation (DFTB) engine to perform computationally efficient exploration of enormous combinatorial design spaces. The algorithm targeted a composite performance index derived from a quantitative structure–property relationship model incorporating descriptors related to light-harvesting efficiency, oscillator strength, and electronic coupling. Within this framework, a tetrathiophene core was functionalised at seven substitution sites using 22 donor and acceptor groups, yielding over 2.5 billion possible molecular combinations. Through iterative selection and mutation cycles, PooMa identified 20 branched oligothiophenes predicted to display strong absorption and favourable HOMO–LUMO alignment. Subsequent DFT and TD-DFT calculations confirmed these predictions, establishing the feasibility of evolutionary optimisation guided by low-cost electronic-structure calculations. This study laid the conceptual groundwork for data-driven generative exploration of OPV-relevant chemical spaces.
Building upon this foundation, Greenstein et al. (2022)191 extended the evolutionary framework to design high-efficiency NFAs. Their hybrid pipeline coupled a genetic algorithm with TD-DFT-based fitness evaluation to recombine donor and acceptor building blocks into new NFAs, generating a library of 5426 unique compounds. The fitness function estimated PCE values using electronic and optical descriptors computed at the TD-DFT level, while the donor component remained fixed. Remarkably, 1087 generated molecules were predicted to exceed 18% PCE, and 159 surpassed 20%, demonstrating that automated generative strategies can recover and surpass the performance of known materials. Moreover, the terminal acceptor motifs repeatedly selected by the algorithm, such as indanone and rhodanine derivatives, mirrored those frequently used in experimental NFAs, providing data-driven validation of empirical design heuristics.
In a follow-up study, Greenstein et al. (2023)114 expanded this framework to model tandem OPV architectures, thereby incorporating multi-junction device simulation into the generative design process. Using a dataset of over 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 donor and acceptor structures, they employed fragment-based recombination and hierarchical optimisation to identify complementary pairs that optimised the absorption and voltage characteristics across both subcells. Analysis of the resulting high-performance NFAs revealed that molecules containing diphenylamine substituents and three-dimensional terminal groups exhibited superior optical coverage and reduced non-radiative voltage losses. These findings not only reinforced earlier empirical observations but also provided quantitative, design-level insight into how molecular geometry and functional group orientation influence tandem device behaviour. The study exemplifies the growing sophistication of generative frameworks capable of integrating molecular and device-level considerations within a unified optimisation loop.
000 donor and acceptor structures, they employed fragment-based recombination and hierarchical optimisation to identify complementary pairs that optimised the absorption and voltage characteristics across both subcells. Analysis of the resulting high-performance NFAs revealed that molecules containing diphenylamine substituents and three-dimensional terminal groups exhibited superior optical coverage and reduced non-radiative voltage losses. These findings not only reinforced earlier empirical observations but also provided quantitative, design-level insight into how molecular geometry and functional group orientation influence tandem device behaviour. The study exemplifies the growing sophistication of generative frameworks capable of integrating molecular and device-level considerations within a unified optimisation loop.
A more recent contribution by Morishita et al. (2024)179 further illustrated the synergistic potential of combining predictive and generative learning. Their study employed the alva-Builder genetic algorithm to design 250 new donor molecules for fullerene-based OPVs, specifically targeting improvements in JSC. A support-vector regression model trained on alvaDesc descriptors served as a surrogate fitness function, predicting the performance of newly generated candidates without expensive quantum calculations. Iterative optimisation revealed that molecules containing 4H-cyclopentadithiophene cores, fluorine-substituted aromatic rings, and carbonyl groups adjacent to thiophene units consistently achieved higher predicted JSC values. The combination of generative exploration and data-driven evaluation created a closed feedback loop that accelerated the discovery of promising donor motifs while providing interpretable correlations between substructural features and device performance.
Parallel to these molecular-level advances, emerging studies have begun extending generative methodologies to mesoscale and morphological optimisation.193 These frameworks aim to capture the influence of film structure, phase separation, and interfacial orientation on charge separation and transport. For instance, algorithmic searches guided by coarse-grained simulations or machine-learned morphology descriptors have been proposed to identify processing pathways that yield optimal percolation networks and minimal energetic disorder. Although still in the early stages of development, such models signal an important broadening of generative AI from single-molecule optimisation toward holistic design encompassing both chemical composition and supramolecular organisation.
Generative approaches applied to OPV discovery to date are dominated by evolutionary algorithms, with occasional use of rule-based molecular enumeration methods such as STONED.63 Neural generative models, including generative adversarial networks and diffusion models, have not yet seen substantive adoption for small-molecule OPV design. Evolutionary algorithms remain prevalent, because they operate naturally on discrete molecular building blocks, enforce chemical validity through explicit mutation and recombination rules, and readily incorporate quantum-chemical or device-level fitness functions.154,191 Their principal limitations are sample inefficiency and strong dependence on the fidelity of the fitness function.194,195 Rule-based approaches such as STONED enable rapid and chemically valid local exploration of molecular space, but do not learn underlying data distributions and therefore lack intrinsic global optimisation capability.63 As a result, STONED is well suited for local chemical-space traversal and hypothesis generation, but poorly matched to directed inverse design or multi-objective optimisation of OPV performance. The limited exploration of GANs and diffusion models likely reflects their reliance on large, well-curated datasets and the difficulty of enforcing chemical validity alongside multiple coupled physical constraints.
Across these developments, the trajectory of generative AI in OPV research reveals increasing integration between physical modelling, data-driven learning, and heuristic optimisation. Early studies relied primarily on rule-based genetic algorithms and surrogate quantum calculations, whereas more recent approaches incorporate predictive surrogate models, active learning strategies, and explicit multi-objective optimisation. This progression reflects a shift from heuristic enumeration toward closed-loop, physics-informed discovery workflows in which candidate generation, property evaluation, and model refinement proceed autonomously and iteratively.
Despite these advances, critical limitations continue to constrain the practical applicability of generative AI for photoactive material design. Among the generative OPV studies reviewed, only one reported experimental validation of computationally designed molecules and only one employed external validation using independent test sets.114,193 This near absence of real-world verification raises significant concerns regarding the transferability of computationally optimised photoactive molecules to functional devices. Moreover, most studies rely on small or chemically homogeneous datasets or narrowly defined molecular families as starting points, and none report negative experimental outcomes. As noted earlier, the lack of failure reporting limits insight into model robustness and prevents a systematic understanding of when and why generative approaches succeed or fail for photoactive materials. These challenges are discussed further in Section 3.5.
3.3 Emissive materials
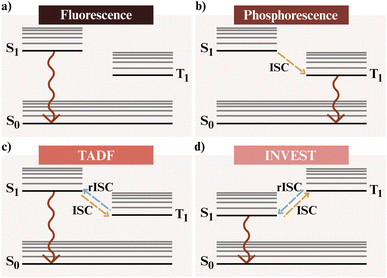
The application of AI to emissive AIs has expanded rapidly, driven by the need to improve the performance and stability of OLEDs. OLEDs are now the dominant technology in high-performance displays and emerging solid-state lighting, yet their efficiency and lifetime remain limited by the coupled excited-state processes governing light emission, colour purity, and degradation.196,197 These properties originate from a complex interplay between molecular electronic structure, spin-state dynamics, and solid-state organisation, making their simultaneous optimisation a persistent challenge for conventional design strategies.Fig. 9 summarises the principal excited-state mechanisms that underpin OLED operation. In fluorescent materials, radiative decay from the lowest singlet excited state (S1 → S0) restricts internal quantum efficiency to about 25%, as triplet excitons are non-emissive.198,199 Phosphorescent systems overcome this limit through spin–orbit coupling that enables emission from the triplet manifold (T1 → S0), achieving near-unity exciton utilisation.200,201 Thermally activated delayed fluorescence (TADF) emitters202,203 exploit reverse intersystem crossing to convert triplets into emissive singlets, while INVEST materials invert the energy ordering of the two states (ES1 < ET1), allowing ultrafast radiative decay without thermal activation.204,205
The evolution from fluorescence to phosphorescence, TADF, and INVEST emitters reflects a progressive refinement in exciton management and energy utilisation. However, high-performance OLED design remains a multidimensional optimisation problem involving the simultaneous control of charge injection, exciton generation and diffusion, intersystem conversion, and radiative decay.206 These parameters are inherently coupled to molecular conformation, packing geometry, and electronic coupling, creating a design space too complex for exhaustive computational or experimental exploration.
AI-based approaches address this complexity by learning non-linear relationships between molecular structure and emissive behaviour. Predictive models trained on experimental or theoretical datasets can estimate excited-state parameters such as singlet–triplet gaps, oscillator strengths, transition dipoles, or non-radiative decay rates with accuracy comparable to first-principles methods but at vastly reduced cost. Generative models extend this paradigm by enabling inverse design, autonomously proposing new emitters with tailored optical or stability characteristics.
These approaches, therefore, establish a data-driven framework that links molecular design, excited-state physics, and device-level performance within a unified modelling pipeline. Within the reviewed literature, 54 studies explore AI in emissive materials: 40 focus on predictive modelling of optical, electronic, or thermal properties, while 14 employ generative strategies for targeted molecular design. These developments mark a transition from empirical screening toward autonomous discovery of high-efficiency OLED emitters, where AI serves not only as a predictive surrogate but as a creative partner in molecular design.
ML approaches based on simple molecular descriptors have played a central role in establishing quantitative links between molecular structure and emissive performance in organic optoelectronic materials. Golin et al.85 trained neural networks and support vector machines on small molecular datasets described by 1688 physicochemical descriptors to model electroluminescence, identifying extended π-conjugation and charge delocalisation as the dominant factors governing emission intensity. More recently, Zhao et al.224 employed LightGBM regressors with molecular fingerprints to predict Stokes shifts across 6064 fluorescent compounds, achieving R2 = 0.86 and an RMSE of 19.16 nm. Guided by the model, the authors synthesised PXZ-F, whose experimentally measured Stokes shift (183 nm) closely matched the predicted value (153 nm), demonstrating the practical utility of fingerprint-based screening. In a complementary classification setting, Zhao et al.225 trained a LightGBM model on 3074 compounds to distinguish aggregation-induced emission-active molecules from aggregation-caused quenchers with 97.4% accuracy. Experimental validation confirmed the discovery of new aggregation-induced TADF emitters, illustrating how simple structural descriptors can bridge molecular photophysics and device-relevant behaviour.
Beyond purely structural representations, several studies have incorporated explicit electronic descriptors derived from quantum-chemical calculations to improve physical interpretability and predictive fidelity. Sato et al.77 developed a hierarchical ML pipeline that combines DFT-derived and empirical descriptors to design triazine-based electron-transport materials. Screening a virtual library of 3.67 million candidates led to the synthesis of nine compounds, with the top performer (T2-6970) exhibiting enhanced efficiency and operational lifetime. Shi et al.226 further demonstrated the value of electronic features by applying XGBoost models to predict transition-dipole orientations relevant to radiative efficiency, achieving R2 ≈ 0.8 and revealing that planar donor–acceptor geometries promote preferential horizontal dipole alignment.
An alternative modelling paradigm replaces manually engineered descriptors with representations learned directly from molecular graphs. Li et al.219 introduced the SOGCN architecture, a structure-aware graph neural network capable of simultaneously predicting singlet–triplet energy gaps (ΔEST) and emission bandwidths, attaining mean absolute errors of 0.037 eV and 10–12 nm, respectively. Barneschi et al.216 trained a three-dimensional graph neural network on more than 85![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 DFT-optimised and experimental structures, achieving mean errors of 0.02 eV in predicting inverted singlet–triplet gaps. Extending graph-based learning to higher levels of device complexity, Lee et al.227 incorporated crystal-structure information into graph neural networks to predict current efficiency in multilayer OLED stacks, achieving R2 = 0.83 and outperforming fully connected baselines. Nikhitha and Mondal212 further combined semi-empirical calculations with Δ-learning and SchNet-based architectures, obtaining an RMSE of 0.004 eV and R2 = 0.95 for ΔEST while successfully generalising to benchmark INVEST emitters.
000 DFT-optimised and experimental structures, achieving mean errors of 0.02 eV in predicting inverted singlet–triplet gaps. Extending graph-based learning to higher levels of device complexity, Lee et al.227 incorporated crystal-structure information into graph neural networks to predict current efficiency in multilayer OLED stacks, achieving R2 = 0.83 and outperforming fully connected baselines. Nikhitha and Mondal212 further combined semi-empirical calculations with Δ-learning and SchNet-based architectures, obtaining an RMSE of 0.004 eV and R2 = 0.95 for ΔEST while successfully generalising to benchmark INVEST emitters.
While most ML models focus on intrinsic molecular properties, some studies have directly incorporated device-level observables and architectures into the learning process. Lim et al.89 trained deep neural networks on time-resolved electroluminescence measurements to extract triplet–triplet annihilation kinetics with R2 = 0.99, eliminating the need for iterative kinetic fitting and demonstrating that transient device signals can be used directly as model inputs. Similarly, Kim et al. (2023)88 trained an artificial neural network on transient electroluminescence decay profiles to directly extract polaron recombination coefficients, achieving R2 values up to 0.949 and enabling quantitative reconstruction of polaron dynamics from device-level time-resolved measurements alone.
In line with other target domains such as OPV performance and electronic-structure prediction, these studies demonstrate that ML models for emissive organic semiconductors can be formulated across multiple representational levels, ranging from simple structural fingerprints and electronic descriptors to learned graph-based embeddings and device-resolved observables. While molecular-level descriptors enable efficient screening and retain a high degree of physical interpretability, representation-learning approaches offer greater expressivity and scalability by alleviating the need for manual feature engineering. The repeated experimental validation of models based on both structural and electronic descriptors underscores their practical reliability and establishes ML as a robust tool for connecting molecular photophysics to emissive performance.
Across the reviewed literature, generative deep-learning frameworks have been employed to design materials with optimised (i) singlet–triplet energy gaps (ΔEST),115,118,157,230 (ii) singlet and triplet excitation energies,118,128,153,231 (iii) photoluminescence quantum yield,117,166 (iv) glass-transition temperature,232 and (v) spectral efficiency and emission profiles.233
An early example of generative OLED design is the work by Kim et al. (2018),54 who employed an encoder–decoder architecture for the inverse design of OLED host materials. Their model significantly increased the proportion of molecules achieving target triplet energies, demonstrating the feasibility of AI-driven molecular generation for optoelectronic applications.
Despite these successes, a key challenge persists: molecules proposed by AI models must not only exhibit desirable photophysical properties but also be synthetically feasible and chemically stable. Without these constraints, generative algorithms may yield unrealistic or impractical structures. To address this limitation, researchers have incorporated synthetic accessibility, stability scoring, and multi-objective optimisation directly into the generative process. For example, Lim et al. (2018)234 introduced a conditional variational autoencoder that allows controlled molecular generation based on multiple target properties, including ease of synthesis. Although initially developed for pharmaceutical applications, this strategy has clear implications for optoelectronic materials, where balancing electronic performance with manufacturability is essential. Building on this concept, Kim et al. (2018)54 developed an inverse design framework integrating predictive property models with multi-objective optimisation to generate molecules that simultaneously satisfy thermal stability, optical gap, and synthetic accessibility criteria. Similarly, Kwak et al. (2022)232 implemented a goal-directed generative model combined with high-throughput molecular simulations, optimising singlet–triplet energy gaps, oscillator strengths, and molecular stability in parallel. These studies highlight the importance of embedding chemical realism into generative design pipelines.
More recent efforts have focused on improving the scalability and generalisability of these frameworks. Tan et al. (2022)118 combined autoencoders with deep property predictors and TD-DFT-based filtering to identify TADF emitters with favourable ΔEST and enhanced spin–orbit coupling, thereby bridging data-driven design with physics-based validation.
These advances illustrate a transition from purely data-driven molecular generation toward physically informed, multi-objective design of OLED materials. By integrating synthetic feasibility and quantum-chemical insight into generative workflows, AI models are becoming capable not only of proposing high-efficiency emitters but also of prioritising those that are experimentally realisable. Notwithstanding these computational achievements, significant limitations challenge the practical utility of generative AI approaches for emissive material design. Of the 14 generative studies reviewed in this domain, only three reported experimental validation of computationally designed molecules,54,128,157 and only two included external validation using independent test sets.117,166 This near absence of experimental verification represents a critical knowledge gap, as the field lacks empirical evidence demonstrating that these computationally optimized emissive molecules translate into materials with the desired photophysical properties, including emission wavelength, quantum yield, colour purity, and operational stability. The reliance on small, homogenous datasets or specific chromophore families as starting points for generation further limits these approaches, introducing substantial data bias and constraining exploration to incremental modifications of known structures rather than discovery of genuinely novel emissive scaffolds. The absence of any reported negative results compounds these concerns, preventing practitioners from learning about failure modes or calibrating expectations about the reliability of computational predictions. Systematic challenges including data bias, chemical validity, and publication bias are addressed in Section 3.5.
3.4 Charge transport
The preceding sections have examined AI applications in optoelectronic function, including absorption and excitation in photoactive materials, power conversion in OPVs, and emission in OLEDs. A complementary and equally important aspect of AIs performance is charge transport. Efficient transport governs current extraction in photovoltaics, charge injection and balance in OLEDs, and switching in thin-film transistors. Unlike spectral or radiative properties, charge transport arises from collective condensed-phase dynamics rather than any single molecular parameter, making it one of the most complex phenomena to model and design.235–237Charge transport in AIs differs fundamentally from that in crystalline inorganic materials. In these soft, π-conjugated lattices, charge carriers move in a regime where electronic and nuclear motions are strongly coupled. Consequently, neither a purely band-like description,238 which assumes long-range coherence, nor a purely hopping-based model,239 which treats carriers as localised, is sufficient. Thermal fluctuations in molecular geometry continuously modulate intermolecular electronic couplings, leading to time-dependent fluctuations collectively described as dynamic disorder.240 This coupling places charge transport in an intermediate regime between coherent band motion and incoherent hopping, often referred to as the transient localisation regime.241
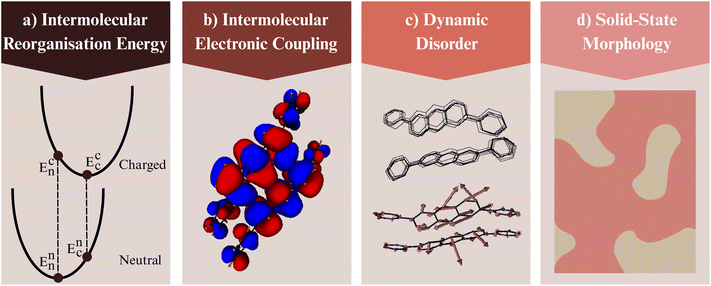
At the microscopic level, dynamic disorder arises from coupling between electronic and vibrational degrees of freedom. It can be expressed as σ = ∇J·Q, where ∇J is the gradient of the transfer integral J with respect to nuclear displacements, and Q represents vibrational normal-mode vectors.240,242 This relation shows how phonon-induced nuclear motions drive fluctuations in electronic coupling, linking charge mobility to the vibrational landscape of the solid state. The resulting mixed transport regime has motivated extensive theoretical and experimental efforts to connect molecular electronic structure with mesoscale structural dynamics.243,244
Within this framework, charge-carrier mobility is not an intrinsic molecular property but an emergent feature of the condensed phase. As illustrated schematically in Fig. 10, mobility reflects the combined influence of four interdependent factors: (i) the intramolecular reorganisation energy, which quantifies the structural relaxation cost during charge transfer; (ii) the intermolecular electronic coupling, which determines the efficiency of wavefunction overlap; (iii) the magnitude and timescale of dynamic disorder induced by thermal vibrations; and (iv) the solid-state morphology, including molecular packing, orientational order, and percolation pathways.245–248
Because these factors are strongly coupled and sensitive to morphology and temperature, accurate prediction or optimisation of charge transport requires models that capture multiple length and time scales, from local electronic structure to mesoscale organisation. Traditional simulations combining molecular dynamics, quantum-chemical calculations, and kinetic modelling are computationally intensive and difficult to generalise. ML now offers a scalable alternative. In this review, we identify 39 studies that apply AI to charge transport in AIs: 34 focus on predictive modelling and 5 on generative design. Predictive models estimate transport-relevant quantities from molecular or morphological descriptors, while generative approaches propose new molecular scaffolds or packing motifs with improved mobility under realistic synthetic and processing constraints. The following subsections provide a detailed discussion of these directions.
Beyond molecular descriptors, charge mobility is critically influenced by intermolecular electronic couplings (Fig. 10, panel b). These transfer integrals are traditionally computed using computationally demanding quantum-chemical methods.263 To accelerate such calculations, Wang et al. (2019)264 developed a KRR model to predict electronic couplings between ethylene dimers. The optimal model, based on Gaussian kernels and intermolecular descriptors, achieved a MAE of 3.5 meV and correctly predicted coupling signs in more than 98% of cases, while providing computational speed-ups of 104–1010 compared with ab initio calculations. Subsequent studies extended this approach to more complex systems. Wang et al. (2020)265 trained neural networks on naphthalene dimers extracted from molecular dynamics simulations, obtaining MAE of 6.5 meV while capturing orientation-dependent variations. Krämer et al. (2020)82 used KRR models trained on DFTB266 data to predict site energies and electronic couplings in anthracene crystals, reproducing hole mobilities within 8.5% of DFTB reference and 34% of experimental values using only 1000 samples. Bhat et al. (2024)267 introduced a three-dimensional message-passing neural network trained on 438![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 dimer configurations extracted from 25
000 dimer configurations extracted from 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 organic crystals. Their model predicted HOMO–HOMO and LUMO–LUMO couplings with MAEs of approximately 3 meV, enabling Marcus-theory-based screening of 60
000 organic crystals. Their model predicted HOMO–HOMO and LUMO–LUMO couplings with MAEs of approximately 3 meV, enabling Marcus-theory-based screening of 60![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 crystal structures within minutes. In a related effort, Nematiaram et al. (2025)37 utilised LightGBM classifiers to predict charge-transport two-dimensionality, an important indicator of mobility, achieving 95% accuracy with geometric and chemical descriptors. They identified crystal volume, molecular rigidity, and intermolecular distance as key features.
000 crystal structures within minutes. In a related effort, Nematiaram et al. (2025)37 utilised LightGBM classifiers to predict charge-transport two-dimensionality, an important indicator of mobility, achieving 95% accuracy with geometric and chemical descriptors. They identified crystal volume, molecular rigidity, and intermolecular distance as key features.
Thermal molecular motions further modulate electronic couplings, introducing dynamic disorder (Fig. 10, panel c).240,242,268 Reiser et al. (2021)86 showed that static Gaussian disorder models fail to capture the full complexity of charge-transfer fluctuations. Building on this, Wang et al. (2023)269 combined molecular dynamics with ML models, including KRR and neural networks, to evaluate time-resolved charge-transfer integrals in ethylene and naphthalene dimers. Their spectral density analysis revealed that low-frequency intermolecular motions, such as translations and rotations, dominate coupling fluctuations. The spectral density exhibited a sub-ohmic character with cut-off frequencies between 100 and 200 cm−1, consistent with inelastic neutron scattering measurements.270,271
Beyond molecular-scale parameters, charge transport is strongly governed by the solid-state morphology (Fig. 10, panel d). The relationship between morphology and charge-carrier mobility poses a fundamental multiscale challenge. Electronic couplings between neighbouring molecules depend sensitively on sub-angstrom variations in relative geometry, while macroscopic mobility emerges from percolation pathways that extend over micrometre length scales through structurally heterogeneous films comprising crystalline domains, grain boundaries, and amorphous regions. Traditional modelling approaches face an inherent trade-off. Quantum-chemical methods can accurately resolve intermolecular electronic couplings, but their computational cost prohibits application to the millions of molecular pairs present in realistic morphologies. Conversely, coarse-grained or mesoscale models efficiently capture large-scale structural organisation but lack the electronic resolution required to describe charge transfer processes. ML provides a viable route to bridge these length scales by learning surrogates of quantum-chemical calculations at greatly reduced computational cost, thereby enabling direct coupling between electronic-structure predictions and morphology-resolved simulations. Lederer et al. (2019)272 coupled KRR with molecular dynamics and kinetic Monte Carlo simulations to predict mobility in disordered pentacene, successfully reproducing mobility anisotropy while reducing computational cost. Tan and Wang (2023)36 extended this concept by training symmetry-adapted neural networks on transfer integrals computed for rubrene, pentacene, DNTT, and BTBT. The networks mapped molecular geometries directly to electronic couplings, which were then incorporated into kinetic Monte Carlo simulations. The resulting hole mobilities closely matched those from ab initio workflows while being several orders of magnitude faster. Tan and Wang (2024)273 further developed a multiscale framework for small-molecule thin films, combining molecular dynamics, ML, and kinetic Monte Carlo simulations. Their study on quadruple thiophene demonstrated how polymorphism, grain boundaries, and molecular orientation influence charge mobility. Neural networks pre-trained on crystalline dimers were fine-tuned on 68![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 844 dimers extracted from disordered film morphologies, achieving near-quantum accuracy for transfer integrals and enabling large-scale mobility predictions. These studies illustrate how ML models approximate morphology–mobility relationships not by explicitly resolving mesoscale morphology in full detail, but by learning effective mappings between local structural environments and transport-relevant electronic properties such as transfer integrals. Rather than treating morphology as a single global descriptor, contemporary approaches encode its influence through ensembles of local molecular arrangements sampled from molecular dynamics simulations or crystal databases. By embedding these local configurations into ML-predicted coupling distributions, kinetic Monte Carlo simulations can recover emergent transport behaviour arising from disorder, anisotropy, and polymorphism.
844 dimers extracted from disordered film morphologies, achieving near-quantum accuracy for transfer integrals and enabling large-scale mobility predictions. These studies illustrate how ML models approximate morphology–mobility relationships not by explicitly resolving mesoscale morphology in full detail, but by learning effective mappings between local structural environments and transport-relevant electronic properties such as transfer integrals. Rather than treating morphology as a single global descriptor, contemporary approaches encode its influence through ensembles of local molecular arrangements sampled from molecular dynamics simulations or crystal databases. By embedding these local configurations into ML-predicted coupling distributions, kinetic Monte Carlo simulations can recover emergent transport behaviour arising from disorder, anisotropy, and polymorphism.
At the device level, ML has been used to correlate molecular and interfacial properties with measured charge transport in OFETs. Lee et al. (2019)274 employed random forest and gradient boosting algorithms to predict electron mobilities in n-type OFETs using features such as HOMO/LUMO levels and electrode work functions. The models identified energy-level alignment and air stability as key determinants of device performance, guiding the optimisation of both materials and contacts.
Active learning has also been applied to improve data efficiency in charge-transport prediction. Antono et al. (2020)275 implemented a closed-loop platform using random forest surrogate models within the FUELS framework, which includes uncertainty estimation to guide sample selection. By combining expected-improvement and uncertainty-based acquisition strategies, their approach identified hole-transporting materials with mobilities 26% higher than the best in the initial dataset after only 165 evaluations. Kunkel et al. (2021)108 expanded this approach to explore an open-ended space of π-conjugated molecules using Gaussian process regression and chemically valid transformations such as ring fusion and side-chain modification. Iterative retraining guided the search toward high-mobility candidates, discovering previously unreported compounds within fifty iterations and outperforming brute-force screening.
Unsupervised learning and pattern-analysis approaches have also been applied to uncover structure–property relationships in existing datasets. Kunkel et al. (2019)276 employed network analysis on 350 π-conjugated molecules, identifying recurring structural motifs, referred to as “molecular LEGO bricks”, that frequently appear in high-mobility compounds. These motifs included specific fused rings, heterocycles, and side chains that can be recombined to form new candidates with enhanced transport properties. Tufail et al. (2024)60 applied a related fragment-based strategy to design small-molecule acceptors with low reorganisation energies, validating top candidates through quantum-chemical calculations. Such fragment-oriented methods provide chemically intuitive design rules and help constrain the search space for generative and active-learning-based exploration.
Across all four domains discussed above, electronic structure, photoactive materials, emissive materials, and charge transport, predictive ML has evolved from small datasets and empirical regressors to scalable, physics-informed neural architectures. These studies demonstrate that ML can now reproduce or even surpass traditional quantum-chemical accuracy for well-characterised targets while revealing transferable design principles across diverse materials classes. Table S1 summarises representative studies in predictive modelling, illustrating the progression from modest datasets and simple descriptors to big-data and deep-learning approaches. Predictive ML for AIs has matured to a point where several properties (e.g., DFT-level orbital energies, OPV VOC, photoluminescence quantum yield) can be estimated reliably without direct calculation. In more complex areas, particularly those entangled with device physics such as full device efficiency or operational stability, performance remains limited by sparse and noisy data. Nevertheless, as datasets expand and models become more physically grounded, ML predictions are approaching the accuracy required to guide experiments, dramatically reducing the search space for high-performance materials.
The first explicit demonstration of a transport-oriented generative framework was presented by Kunkel et al. (2021),108 who coupled molecular morphing with Bayesian active learning to explore π-conjugated chemical space. Their closed-loop workflow jointly optimised reorganisation energy, charge-injection barriers, and mobility-related proxies, successfully identifying previously unreported high-mobility candidates. Building on this foundation, Marques et al. (2021)105 implemented the REINVENT reinforcement-learning algorithm to design heteroacenes with reduced hole reorganisation energies, demonstrating that on-policy reinforcement learning can effectively guide molecular generation toward improved transport characteristics.
Subsequent studies have refined and benchmarked generative algorithms for transport-oriented optimisation. Staker et al. (2022)103 compared four de novo frameworks, MolDQN, GraphGA, GENTRL, and ChemTS, using a dataset of 250![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 DFT-calculated reorganisation energies. GraphGA achieved the best balance between chemical validity, novelty, and exploration efficiency, producing synthetically plausible low-λ molecules later confirmed through quantum-chemical validation. In a related effort, Kwak et al. (2022)232 combined a goal-directed recurrent neural network with deep reinforcement learning to design hole-transport materials. The model integrated high-throughput molecular simulations directly into the generation loop, yielding compounds with hole reorganisation energies below 0.2 eV and linking molecular design to predicted mobility. More recently, Kawagoe et al. (2024)251 coupled Bayesian optimisation with learned molecular embeddings to efficiently locate low-λ candidates while quantifying the role of descriptor selection and acquisition strategy on search performance.
000 DFT-calculated reorganisation energies. GraphGA achieved the best balance between chemical validity, novelty, and exploration efficiency, producing synthetically plausible low-λ molecules later confirmed through quantum-chemical validation. In a related effort, Kwak et al. (2022)232 combined a goal-directed recurrent neural network with deep reinforcement learning to design hole-transport materials. The model integrated high-throughput molecular simulations directly into the generation loop, yielding compounds with hole reorganisation energies below 0.2 eV and linking molecular design to predicted mobility. More recently, Kawagoe et al. (2024)251 coupled Bayesian optimisation with learned molecular embeddings to efficiently locate low-λ candidates while quantifying the role of descriptor selection and acquisition strategy on search performance.
As discussed across preceding domains, a persistent challenge in generative molecular design is achieving a balance between electronic performance, chemical realism, and experimental feasibility. Unconstrained optimisation often yields electronically ideal but synthetically inaccessible or thermally unstable structures. To address this, recent studies have incorporated physical constraints and empirical design rules directly into generative objectives, enabling models to account for both functionality and manufacturability. Multi-objective optimisation frameworks have become particularly effective, coupling charge-transport descriptors with stability-related metrics to ensure holistic performance. For instance, Kwak et al. (2022)232 employed a composite reward function integrating HOMO–LUMO alignment, hole reorganisation energy, and glass-transition temperature (Tg), a surrogate for morphological robustness, resulting in candidate materials that combined high mobility with enhanced thermal and structural stability. These developments mark a shift from single-property optimisation toward integrated, closed-loop frameworks that unify molecular generation, surrogate prediction, and physics-based validation. In such systems, generative models propose candidate structures, ML surrogates estimate charge-transport descriptors such as λ and electronic coupling, and quantum-chemical or kinetic simulations provide final verification. The convergence of data-driven design and physical modelling thus establishes a scalable pathway for the rational discovery of high-mobility MSCs.
Table S2 summarises representative generative and inverse-design studies across multiple domains, illustrating the growing methodological diversity and scope of AI-assisted molecular discovery. A decade ago, the design of new small-molecule semiconductors relied primarily on chemical intuition and analogue synthesis; today, AI-driven pipelines routinely propose novel scaffolds with superior predicted performance, accelerating the exploration of chemical space far beyond human intuition.92,166 Remaining challenges include enforcing synthetic accessibility, ensuring accurate surrogate predictions, and balancing conflicting objectives such as mobility, stability, and ease of processing, issues that define the next frontier of generative AI in AI design.
These generative strategies highlight the computational promise of inverse molecular design; however, their translation to practical charge-transport material discovery remains limited. Among the five generative studies identified in this domain, none reported experimental validation of computationally proposed molecules, nor did they include external validation using independent test sets. This gap is particularly relevant for charge-transport applications, where predicted mobilities must ultimately be corroborated through device fabrication and testing to assess energy-level alignment, thin-film morphology, and environmental stability. Broader challenges common to generative approaches, including dataset bias and limited reporting of negative outcomes, are discussed in Section 3.5.
3.5 Challenges and opportunities
The integration of AI into AI discovery has substantially advanced materials design, yet several persistent challenges constrain its full transformative potential. These include the scarcity, heterogeneity, and limited interoperability of available datasets; the restricted generalisability of models across distinct chemical families; and the reliance on incomplete or task-specific molecular representations that impede transferability and mechanistic insight. Further limitations arise from the insufficient treatment of predictive uncertainty, the gap between computational prediction and experimental realisation, and the absence of interpretable, physically grounded frameworks linking model outputs to chemical principles. Addressing these interdependent issues within a unified methodological framework is essential to enable robust, transferable, and experimentally meaningful AI-guided materials discovery. The following sections provide a detailed discussion of these challenges.To compensate for the scarcity of experimental data, most studies rely on computationally generated datasets derived from DFT, TD-DFT, or semi-empirical quantum-chemical calculations.88,89,219 These datasets offer scalability, internal consistency, and systematic exploration of chemical space, forming the foundation of current ML development. Large community initiatives such as the Harvard Clean Energy Project,26 the Harvard Organic Photovoltaic Dataset,279 and the PubChemQC Project99 have established valuable computational baselines for benchmarking and model training.
However, these resources remain inherently theoretical and therefore lack the experimental fidelity required for accurate prediction of device performance. Information essential to real-world functionality, such as morphology, molecular packing, processing conditions, and degradation pathways, is typically beyond the scope of first-principles datasets.280 Likewise, metadata central to experimental reproducibility, including fabrication parameters, measurement protocols, and environmental stability metrics, are intrinsically absent, limiting model transferability across data domains.
A further limitation lies in their narrow chemical diversity. Many available datasets are confined to specific donor–acceptor families or fused-ring systems,139 which limits model generalisability to new molecular scaffolds or device architectures.281 In addition, negative or low-performing examples, molecules that are synthetically feasible but fail to meet target thresholds, are rarely reported. Without such counterexamples, models tend to overfit to successful data and cannot learn to distinguish productive from unproductive designs. The deliberate inclusion of synthetic negative samples or controlled noise, as demonstrated in two-dimensional materials discovery,282 could enhance model discrimination and robustness by exposing algorithms to the boundaries of viable chemical space.
Progress in this area also depends on improving the quality, consistency, and accessibility of available data. The absence of standardised data formats, metadata conventions, and measurement protocols continues to limit reproducibility and comparability across studies. The adoption of FAIR data principles (Findable, Accessible, Interoperable, and Reusable)283 and the systematic reporting of low-performing materials284 would establish a more balanced and transparent data ecosystem. Open, version-controlled repositories that incorporate uncertainty estimates and standardised metadata would further enable reproducible benchmarking and closer collaboration between computational and experimental communities.
Because generating new experimental data remains slow and resource-intensive, data-efficient learning strategies have emerged as practical solutions for maximising the value of existing information. Active learning algorithms identify simulations or experiments expected to yield the greatest information gain,62,285 guiding exploration toward the most informative regions of chemical space. Complementary approaches, such as transfer learning, enable models trained on related datasets to improve predictions in data-scarce domains by reusing learned representations.58 These strategies have been successfully applied to predict key molecular properties including frontier orbital energies,56 reorganisation energies, and charge-carrier mobilities,58,273 demonstrating that meaningful generalisation can be achieved even under data constraints.
Beyond data availability, how molecular information is represented within ML algorithms constitutes a second major limitation. Molecular representations define how structural and electronic features are encoded, directly influencing model accuracy, interpretability, and transferability (Fig. 11). Early studies relied on handcrafted descriptors such as extended-connectivity fingerprints,137 topological indices,286 and quantum-chemical features.139 These representations are computationally efficient and interpretable but fail to capture critical aspects such as three-dimensional conformation, stereochemistry, and intermolecular interactions, which strongly affect charge transport and excited-state behaviour. The introduction of graph-based learning marked a turning point, with graph neural networks enabling data-driven feature extraction directly from molecular connectivity.287 Despite their success, many implementations remain restricted to two-dimensional molecular graphs and therefore neglect spatial information that is essential for accurately describing structure–property relationships.
Recent progress has focused on incorporating explicit three-dimensional information into molecular representations. Message-passing neural networks with geometric edge features,38 equivariant neural networks,288 and conformer-aware embeddings289 have all demonstrated improved performance for geometry-sensitive properties such as reorganisation energies257 and charge-transfer integrals.273 However, these models depend on accurate molecular geometries, which are computationally expensive to generate and sensitive to conformer selection, especially for flexible or disordered systems.
Capturing solid-state and mesoscale effects represents the next frontier in molecular representation for MSCs. Device performance depends not only on molecular structure but also on packing, morphology, and interfacial interactions.290–292 To bridge this molecular-to-device gap, new representations must integrate information across multiple length scales, combining molecular, crystallographic, and morphological descriptors. Promising directions include the use of packing efficiency metrics, crystal symmetry parameters, and intermolecular coupling features derived from molecular dynamics simulations or experimental characterisation.293 Physics-informed neural networks and differentiable molecular dynamics frameworks provide additional opportunities to embed physical constraints directly into model architectures, enabling the prediction of temperature-dependent behaviour, morphological evolution, and long-term stability.
Looking forward, advancing data and representation frameworks will be essential for achieving robust and interpretable AI-driven materials discovery. Future priorities include the integration of experimental and computational data within FAIR-compliant repositories, the use of active and transfer learning to enhance data efficiency, and the development of multiscale, physics-informed representations that capture both molecular structure and device-level phenomena. These advances will enable reliable, transparent, and experimentally relevant ML models capable of accelerating the rational design of next-generation MSCs.
This fragility often stems from the chemical homogeneity of available datasets. Models trained primarily on fused heterocycles,295 donor–acceptor cores,296 or specific π-conjugated frameworks69 learn narrow structure–property heuristics297 that fail when encountering unfamiliar bonding topologies or heteroatom arrangements.75 Even minor structural variations, such as introducing electron-withdrawing substituents or modifying π-bridge positions, can substantially alter optoelectronic behaviour and invalidate extrapolations. Expanding chemical diversity and adopting scaffold-aware validation protocols are therefore essential for improving model robustness. Complementary strategies, including transfer learning and domain-adaptation techniques,56,57,104,298 can help extend model applicability across related datasets. In parallel, geometric and equivariant neural networks offer the potential to learn more universal structure-based features that enhance generalisation across chemical families.57,148,299
Looking forward, improving model generalisation will require community-wide adoption of scaffold-based validation standards, greater chemical diversity in benchmark datasets, and systematic integration of domain-adaptation and uncertainty-analysis methods. Such practices will enable models that not only interpolate within known systems but also extrapolate reliably to new chemical spaces.
Several complementary approaches exist for quantifying predictive uncertainty. Model ensembling estimates confidence from the variance across independently trained models,301 while Bayesian methods, including Gaussian processes262 and Bayesian neural networks,302 offer principled probabilistic predictions with credible intervals. Monte Carlo dropout provides a computationally tractable approximation of Bayesian inference by applying dropout at inference time to produce distributions from a single network.303–305 Each method involves trade-offs: ensembles improve robustness but are computationally demanding, whereas dropout-based methods are efficient but can underestimate uncertainty in extrapolative regions.
Applicability-domain (AD) analysis complements UQ by identifying whether a molecule lies within the scope of the training distribution. AD frameworks employ distance-based similarity measures,100,306 density estimation, or statistical outlier detection algorithms307 to flag predictions that are likely unreliable. Integrating UQ and AD analyses into ML pipelines enables early detection of low-confidence predictions and more efficient allocation of experimental resources. Despite these advantages, uncertainty-aware practices remain under-represented in MSC research.108,308
Looking forward, embedding UQ and AD analysis directly into model development and evaluation will be crucial for establishing trust and reproducibility. Future benchmarks should assess both predictive accuracy and calibration quality, while active-learning workflows should incorporate uncertainty-based acquisition functions to prioritise the most informative experiments.
Where experimental validation is reported, it is typically limited to a small number of top-ranked candidates. None of the reviewed studies provides systematic statistics on unsuccessful syntheses or failed experimental outcomes. The absence of negative-result reporting prevents quantitative assessment of success rates for AI-predicted candidates and likely reflects a combination of publication bias and practical constraints associated with experimental follow-up. As a result, objective evaluation of model reliability and generalisability remains limited.
Post hoc screening techniques such as the synthetic accessibility score (SAscore),312 synthetic complexity score (SCScore),313 and retrosynthetic planning tools including ASKCOS314 and AiZynthFinder315 are often used to filter unrealistic candidates. While these methods help eliminate impractical designs, they are inherently reactive and inefficient because they discard large regions of generated chemical space after screening. A more effective approach incorporates feasibility directly into the generative process. Fragment-based assembly rules such as BRICS316 and RECAP317 can constrain molecular construction to synthetically plausible motifs. Surrogate descriptors such as log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P, Hansen solubility parameters, bond-dissociation energies, and predicted glass-transition temperatures318–320 can also be included as optimisation objectives to balance processability and stability alongside electronic performance. Integrating retrosynthetic analysis within closed-loop optimisation workflows further ensures that synthetic accessibility influences candidate prioritisation from the outset.
P, Hansen solubility parameters, bond-dissociation energies, and predicted glass-transition temperatures318–320 can also be included as optimisation objectives to balance processability and stability alongside electronic performance. Integrating retrosynthetic analysis within closed-loop optimisation workflows further ensures that synthetic accessibility influences candidate prioritisation from the outset.
Since high-performing MSCs must satisfy multiple, often competing requirements, including electronic performance, stability, and manufacturability, multi-objective optimisation frameworks are indispensable. Techniques such as evolutionary algorithms, reinforcement learning, and Bayesian optimisation have been used to identify Pareto-optimal solutions that balance trade-offs across diverse property spaces.321–323 Adaptive weighting schemes that incorporate experimental feedback represent a promising avenue toward more realistic optimisation of material performance.
Looking forward, embedding synthetic feasibility, processability, and stability constraints directly within generative and optimisation models will be essential for translating computational predictions into experimentally viable materials. Equally important will be improved reporting practices, including transparent disclosure of unsuccessful predictions and failed experimental validations, to enable rigorous benchmarking and fair assessment of AI-driven design strategies. The development of benchmark datasets that include experimentally measured stability and manufacturability metrics will further enable AI systems to propose MSCs that are both high-performing and practical to realise in the laboratory.
The opacity of many high-performing ML models also complicates error analysis and limits the ability to derive transferable design principles. Without insight into which features drive a prediction, it becomes difficult to identify biases, detect failure modes, or translate computational results into physical intuition.112 To address this challenge, recent work has focused on developing interpretability frameworks that explain how input features influence model outputs. Among the most widely used approaches are SHAP (SHapley Additive exPlanations),327 LIME (Local Interpretable Model-Agnostic Explanations),328 and gradient-based feature attribution methods.329 These techniques provide quantitative or visual insight into the relationship between molecular descriptors and predicted properties.
SHAP has become one of the most effective and widely adopted methods for model interpretation. It assigns each input feature a contribution value indicating how much that feature increases or decreases the predicted outcome. Fig. 12 illustrates a representative SHAP summary plot, where each point corresponds to a molecule. The colour indicates the magnitude of a specific descriptor, while the horizontal position shows whether that descriptor increases or decreases the target property. The overall spread of points reflects the global importance of that feature. For example, a cluster of red points with positive SHAP values for electron affinity indicates that higher electron affinity tends to enhance predicted device efficiency. By transforming high-dimensional numerical outputs into chemically meaningful patterns, SHAP enables us to connect data-driven predictions with mechanistic insight.
This approach has been successfully applied in several recent studies. Das et al. (2024)330 used SHAP analysis to identify the molecular descriptors most strongly affecting PCE, VOC, JSC, and FF in OPVs. Their analysis revealed that the acceptor oscillator strength, electron affinity, and the Gibbs free energy of charge transfer were the dominant contributors to device performance. Similarly, Abadi et al. (2022)331 demonstrated that reorganisation energy and heteroatom count were key determinants of PCE. These studies illustrate how SHAP facilitates a direct link between model predictions and chemically interpretable variables.
Beyond feature-level analysis, interpretability can also be achieved through methods embedded directly within model architectures. Graph-based attention mechanisms332 and gradient-based saliency maps329 have been applied to graph neural networks to visualise which atoms or structural motifs most strongly influence a prediction. Fig. 13 shows a schematic example in which atomic regions with greater influence are highlighted in red and less influential regions appear in blue. These maps are particularly valuable in MSCs research, where small structural modifications, such as adjusting π-bridge length or substituent position, can significantly alter optoelectronic properties.
Another promising direction is the development of hybrid or physics-informed models that combine interpretable physical descriptors with data-driven architectures.333–335 These models integrate features such as HOMO and LUMO energies, reorganisation energies, dipole moments, and singlet–triplet energy gaps within interpretable frameworks such as tree-based algorithms or symbolic regressors. While they may sacrifice some predictive accuracy compared to deep neural networks, they provide clearer physical insight and better alignment with established theory.
In some cases, symbolic regression and equation-discovery techniques have been employed to derive explicit, human-readable relationships that capture structure–property trends.218,336,337 This integration of mechanistic understanding with statistical learning represents a key step toward transparent and scientifically grounded AI models.
Latent-space visualisation provides another valuable interpretability tool.338 When applied to embeddings produced by autoencoders or graph neural networks, low-dimensional maps can reveal clusters of molecules with shared features, uncover hidden structure–property relationships, and identify outliers that deviate from expected design patterns.55,339 When coloured or labelled by experimental performance, such maps offer an intuitive overview of how the model organises chemical space, providing a bridge between prediction and intuition.
Looking forward, improving interpretability will be crucial for building trust and extracting mechanistic understanding from AI-guided molecular design. Future progress will require frameworks that integrate physically meaningful descriptors with explainable architectures, combine uncertainty estimation with interpretability analysis, and establish community benchmarks for transparent and reproducible model evaluation. Embedding interpretability as a core principle, rather than a post hoc addition, will enable ML models that not only predict performance but also reveal the physical and chemical logic that governs it.
The challenges discussed above are deeply interlinked. Limited and biased data restrict model generalisation, weak generalisation amplifies uncertainty, ignoring synthetic constraints undermines experimental translation, and limited interpretability obscures mechanistic insight. Addressing these issues requires unified, physics-aware frameworks that jointly advance data infrastructure, domain-aware validation, uncertainty quantification, and interpretability tools. Achieving this level of integration will transform AI from a primarily predictive instrument into a reliable and explanatory partner in the rational design of next-generation MSCs.
One tangible response to these interlinked challenges is the emergence of closed-loop, AI-driven experimental workflows that embed ML models directly within iterative design–make–test–analyse cycles. Such frameworks provide a unifying mechanism through which restricted generalisation, uncertainty amplification, limited interpretability, and weak experimental translation can be addressed simultaneously. By coupling predictive, generative, and active-learning strategies with automated synthesis and experimental feedback, closed-loop systems enable continuous model refinement under physically and synthetically realistic constraints. In this setting, interpretability and uncertainty estimation become integral to experimental decision-making rather than retrospective analytical tools.
Importantly, recent methodological advances demonstrate that this level of integration is no longer purely conceptual but increasingly experimentally operational.340,341 Within MSC research, however, such paradigms remain the exception rather than the norm. As shown across the preceding sections of this review, the majority of AI-driven studies continue to rely on in silico screening or limited experimental validation applied after model development, with only a small subset embedding ML models within genuinely iterative experimental workflows.
Progress in laboratory automation has nevertheless enabled the first realisations of fully closed-loop, “self-driving” laboratories for organic electronic materials. To date, the most advanced demonstrations are found in small-molecule organic semiconductor lasers, where autonomous platforms couple ML-guided molecular selection with automated cross-coupling synthesis and in-line or quasi-in-line optical characterisation.342,343 In these systems, candidate molecules are proposed algorithmically, synthesised through automated workflows, and evaluated using rapid optical measurements, with experimental outputs returned directly to the learning model to guide subsequent iterations. The optimisation and coordination of these workflows are typically achieved using Bayesian optimisation frameworks, including ChemOS344 and Phoenics,345 as well as more recent efforts aimed at improving the accessibility and integration of Bayesian optimisation tools within experimental environments.346
While these demonstrations validate the technical feasibility of AI-driven experimental autonomy, they also highlight the constraints that currently limit broader translation to MSCs. In particular, reliable automation of synthesis, materials handling, and performance-relevant characterisation remains a dominant bottleneck, especially for solid-state assembly and device-level evaluation. Consequently, and consistent with the validation gaps identified across predictive and generative studies in this review, existing self-driving platforms are largely restricted to solution-phase measurements or proxy performance metrics.347 Nevertheless, these early successes establish a credible pathway toward future autonomous discovery frameworks that integrate molecular design with materials processing and device-level performance assessment, positioning AI as an experimentally grounded, decision-making component of MSC research rather than solely a predictive surrogate.
4 Conclusion and outlook
AI is redefining the discovery and design of MSCs by connecting chemical structure to electronic and photonic function through data-driven understanding. This systematic review analysed 237 studies published between 2010 and 2025 to provide a comprehensive assessment of how AI has advanced the field from conventional computational modelling to predictive, generative, and physically informed discovery. The collective evidence demonstrates that AI has become an essential tool for accelerating innovation and deepening mechanistic insight in molecular materials research.Across the four principal research domains, i.e., electronic structure, photoactive materials, emissive systems, and charge transport, AI has achieved significant progress. In electronic structure prediction, ML models have reached near-quantum accuracy while reducing computational cost by orders of magnitude. In photoactive and emissive materials, predictive and generative frameworks have enabled the rational design of high-performance donors, acceptors, and emitters by directly linking molecular architecture to optoelectronic behaviour. In charge transport, hybrid and physics-informed models have begun to clarify the combined effects of morphology, disorder, and electronic coupling on carrier mobility. These advances demonstrate that AI now functions not only as a predictive tool but also as a powerful framework for uncovering the physical principles that underpin material performance.
The methodological foundations of this research have expanded in both depth and diversity. Tree-based and linear models remain valuable due to their interpretability, robustness, and reliability, while graph-based and message-passing neural networks have become essential for encoding molecular topology and electronic interactions. Generative and diffusion-based models further extend these capabilities by enabling inverse design, allowing algorithms to propose and optimise molecular structures that satisfy defined physical or functional objectives. These advances reflect a shift from purely data-driven prediction toward intelligent exploration, in which AI functions as an integrated scientific framework for guiding molecular design.
Despite these advances, several key challenges persist. Data availability and quality remain uneven, which restricts reproducibility and transferability across studies. Only 110 of the 237 studies reviewed provide openly accessible datasets, and just 58 release executable code, limiting independent benchmarking, methodological reuse, and long-term reproducibility. Benchmarking standards also vary widely across the literature, constraining cross-comparability and obscuring true model performance. In addition, many high-capacity neural models continue to function as opaque systems that deliver accurate predictions but offer limited chemical insight. Experimental validation remains the exception rather than the norm, with only 38 studies reporting experimental confirmation of AI-predicted MSCs. This pattern highlights a persistent disconnect between computational discovery and laboratory realisation. Addressing these challenges will require coordinated, community-wide efforts to establish open and standardised datasets, transparent validation protocols, and learning architectures that incorporate physical constraints and uncertainty quantification. Building these foundations will help ensure that AI-driven discovery remains credible, interpretable, and scientifically meaningful.
AI now occupies a central position in MSC research, providing a unifying framework that links theory, computation, and experiment within increasingly adaptive discovery pipelines. While predictive and generative models have already transformed how chemical space is explored, their long-term impact will depend on the maturation of data infrastructure, transparent validation practices, and deeper integration with experimental workflows. The convergence of AI with automation, robotics, and high-throughput experimentation offers a credible route toward self-driving discovery systems, but realising this vision will require community-wide commitments to open data, reproducible benchmarks, and experimentally grounded feedback loops. As these foundations are established, AI will evolve from a powerful accelerant of screening into a reliable, decision-making partner in materials design, enabling MSCs to be developed not by intuition alone, but through systematic, evidence-driven optimisation for future electronic and photonic technologies.
Conflicts of interest
There are no conflicts to declare.Data availability
All data supporting this review are publicly available in the supplementary information (SI). Supplementary information: a PDF document containing the PRISMA 2020 checklist, the PRISMA 2020 flow diagram, and the full Boolean search strings used for literature retrieval, as well as a CSV file providing a structured database of all studies included in the review. The CSV dataset contains extracted metadata for each study, including publication details, application domain (e.g. electronic structure, photoactive, emissive, or charge-transport materials), AI and ML methodologies employed, target properties, data provenance (experimental, computational, or hybrid), validation strategy or performance metrics, and the presence or absence of experimental verification, open datasets or open code. See DOI: https://doi.org/10.1039/d5dd00552c.Notes and references
- B. Kippelen and J.-L. Brédas, Energy Environ. Sci., 2009, 2, 251–261 RSC.
- R. Gómez-Bombarelli, J. Aguilera-Iparraguirre, T. D. Hirzel, D. Duvenaud, D. Maclaurin, M. A. Blood-Forsythe, H. S. Chae, M. Einzinger, D.-G. Ha and T. Wu, et al. , Nat. Mater., 2016, 15, 1120–1127 CrossRef PubMed.
- T. Q. Trung, N. T. Tien, Y. G. Seol and N.-E. Lee, Org. Electron., 2012, 13, 533–540 CrossRef CAS.
- S. R. Forrest, Nature, 2004, 428, 911–918 CrossRef CAS PubMed.
- H. Sirringhaus, Adv. Mater., 2005, 17, 2411–2425 CrossRef CAS.
- Y. Sun, G. C. Welch, W. L. Leong, C. J. Takacs, G. C. Bazan and A. J. Heeger, Nat. Mater., 2012, 11, 44–48 CrossRef CAS PubMed.
- C. J. Brabec, M. Heeney, I. McCulloch and J. Nelson, Chem. Soc. Rev., 2011, 40, 1185–1199 RSC.
- T. Nemati Aram, P. Anghel-Vasilescu, A. Asgari, M. Ernzerhof and D. Mayou, J. Chem. Phys., 2016, 145, 124116 CrossRef PubMed.
- M. Suzuki, K. Suzuki, T. Won and H. Yamada, J. Mater. Chem. C, 2022, 10, 1162–1195 RSC.
- J. Guo, C. Shi, Y. Zhen and W. Hu, Acc. Mater. Res., 2024, 5, 907–919 CrossRef CAS.
- L. R. Blair and T. Nematiaram, J. Mater. Chem. C, 2025, 13, 17769–17779 RSC.
- A. Pulido, L. Chen, T. Kaczorowski, D. Holden, M. A. Little, S. Y. Chong, B. J. Slater, D. P. McMahon, B. Bonillo and C. J. Stackhouse, et al. , Nature, 2017, 543, 657–664 CrossRef CAS PubMed.
- W. Shi, T. Deng, Z. M. Wong, G. Wu and S.-W. Yang, npj Comput. Mater., 2021, 7, 107 CrossRef CAS.
- P. Yu, Y. Zhen, H. Dong and W. Hu, Chem, 2019, 5, 2814–2853 CAS.
- C. K. Trinh and N. I. Abdo, J. Mol. Struct., 2022, 1269, 133764 CrossRef CAS.
- M. A. Erickson, M. T. Beels and I. Biaggio, J. Opt. Soc. Am. B, 2016, 33, E130–E142 CrossRef CAS.
- D. Cappello, F. L. Buguis and J. B. Gilroy, ACS Omega, 2022, 7, 32727–32739 CrossRef CAS PubMed.
- Y. Cui, Y. Xu, H. Yao, P. Bi, L. Hong, J. Zhang, Y. Zu, T. Zhang, J. Qin and J. Ren, et al. , Adv. Mater., 2021, 33, 2102420 CrossRef CAS PubMed.
- H. Sirringhaus, Adv. Mater., 2014, 26, 1319–1335 CrossRef CAS PubMed.
- L. Ruddigkeit, R. Van Deursen, L. C. Blum and J.-L. Reymond, J. Chem. Inf. Model., 2012, 52, 2864–2875 CrossRef CAS PubMed.
- D. S. Sholl and J. A. Steckel, Density Functional Theory: A Practical Introduction, John Wiley & Sons, 2022 Search PubMed.
- R. J. Bartlett and J. F. Stanton, Reviews in Computational Chemistry, 1994, pp. 65–169 Search PubMed.
- C. Sutton, J. S. Sears, V. Coropceanu and J.-L. Bredas, J. Phys. Chem. Lett., 2013, 4, 919–924 CrossRef CAS PubMed.
- F. Coppola, P. Cimino, F. Perrella, L. Crisci, A. Petrone and N. Rega, J. Phys. Chem. A, 2022, 126, 7179–7192 CrossRef CAS PubMed.
- J. M. Herbert, Phys. Chem. Chem. Phys., 2024, 26, 3755–3794 RSC.
- J. Hachmann, R. Olivares-Amaya, S. Atahan-Evrenk, C. Amador-Bedolla, R. S. Sánchez-Carrera, A. Gold-Parker, L. Vogt, A. M. Brockway and A. Aspuru-Guzik, J. Phys. Chem. Lett., 2011, 2, 2241–2251 CrossRef CAS.
- Ö. H. Omar, M. Del Cueto, T. Nematiaram and A. Troisi, J. Mater. Chem. C, 2021, 9, 13557–13583 RSC.
- M. Bursch, J.-M. Mewes, A. Hansen and S. Grimme, Angew. Chem., 2022, 134, e202205735 CrossRef.
- A. Tkatchenko, Nat. Commun., 2020, 11, 4125 CrossRef CAS PubMed.
- C. P. Gomes, B. Selman and J. M. Gregoire, MRS Bull., 2019, 44, 538–544 CrossRef.
- A. K. Cheetham and R. Seshadri, Chem. Mater., 2024, 36, 3490–3495 CrossRef CAS PubMed.
- D. E. Rumelhart, G. E. Hinton and R. J. Williams, nature, 1986, 323, 533–536 CrossRef.
- J. H. Friedman, Ann. Stat., 2001, 1189–1232 Search PubMed.
- C. Cortes and V. Vapnik, Mach. Learn., 1995, 20, 273–297 CrossRef.
- B. Mazouin, A. A. Schöpfer and O. A. von Lilienfeld, Mater. Adv., 2022, 3, 8306–8316 RSC.
- T. Tan and D. Wang, J. Chem. Phys., 2023, 158, 094102 CrossRef CAS PubMed.
- T. Nematiaram, Z. Lamprou and Y. Moshfeghi, Chem. Commun., 2025, 61, 3676–3679 RSC.
- J. Gilmer, S. S. Schoenholz, P. F. Riley, O. Vinyals and G. E. Dahl, International Conference on Machine Learning, 2017, pp. 1263–1272 Search PubMed.
- P. Reiser, M. Neubert, A. Eberhard, L. Torresi, C. Zhou, C. Shao, H. Metni, C. van Hoesel, H. Schopmans and T. Sommer, et al. , Commun. Mater., 2022, 3, 93 CrossRef CAS PubMed.
- C. Bilodeau, W. Jin, T. Jaakkola, R. Barzilay and K. F. Jensen, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2022, 12, e1608 Search PubMed.
- L. Schneider, M. Schwarting, J. Mysona, H. Liang, M. Han, P. M. Rauscher, J. M. Ting, S. Venkatram, R. B. Ross and K. Schmidt, et al. , Mol. Syst. Des. Eng., 2022, 7, 1611–1621 RSC.
- F. Strieth-Kalthoff, H. Hao, V. Rathore, J. Derasp, T. Gaudin, N. H. Angello, M. Seifrid, E. Trushina, M. Guy and J. Liu, et al. , Science, 2024, 384, eadk9227 CrossRef CAS PubMed.
- Z. Wu, B. Ramsundar, E. N. Feinberg, J. Gomes, C. Geniesse, A. S. Pappu, K. Leswing and V. Pande, Chem. Sci., 2018, 9, 513–530 RSC.
- B. Dou, Z. Zhu, E. Merkurjev, L. Ke, L. Chen, J. Jiang, Y. Zhu, J. Liu, B. Zhang and G.-W. Wei, Chem. Rev., 2023, 123, 8736–8780 CrossRef CAS PubMed.
- R. A. Mata and M. A. Suhm, Angew. Chem., 2017, 56, 11011 CrossRef CAS PubMed.
- E. Bhardwaj, H. Gujral, S. Wu, C. Zogheib, T. Maharaj and C. Becker, Proceedings of the 2024 ACM Conference on Fairness, Accountability, and Transparency, 2024, pp. 1055–1067 Search PubMed.
- X. Zhong, B. Gallagher, S. Liu, B. Kailkhura, A. Hiszpanski and T. Y.-J. Han, npj Comput. Mater., 2022, 8, 204 CrossRef.
- F. Oviedo, J. L. Ferres, T. Buonassisi and K. T. Butler, Acc. Mater. Res., 2022, 3, 597–607 CrossRef CAS.
- H. Choubisa, P. Todorović, J. M. Pina, D. H. Parmar, Z. Li, O. Voznyy, I. Tamblyn and E. H. Sargent, npj Comput. Mater., 2023, 9, 117 CrossRef.
- A. Kovacs, J. Fischbacher, H. Oezelt, A. Kornell, Q. Ali, M. Gusenbauer, M. Yano, N. Sakuma, A. Kinoshita and T. Shoji, et al. , Front. Mater., 2023, 9, 1094055 CrossRef.
- A. H. Cheng, C. T. Ser, M. Skreta, A. Guzmán-Cordero, L. Thiede, A. Burger, A. Aldossary, S. X. Leong, S. Pablo-Garcia and F. Strieth-Kalthoff, et al. , Faraday Discuss., 2025, 256, 10–60 RSC.
- M. J. Page, J. E. McKenzie, P. M. Bossuyt, I. Boutron, T. C. Hoffmann, C. D. Mulrow, L. Shamseer, J. M. Tetzlaff, E. A. Akl and S. E. Brennan, et al. , BMJ, 2021, 372, n71 CrossRef PubMed.
- S. Atahan-Evrenk, RSC Adv., 2018, 8, 40330–40337 RSC.
- K. Kim, S. Kang, J. Yoo, Y. Kwon, Y. Nam, D. Lee, I. Kim, Y.-S. Choi, Y. Jung and S. Kim, et al. , npj Comput. Mater., 2018, 4, 67 CrossRef.
- R. Gómez-Bombarelli, J. N. Wei, D. Duvenaud, J. M. Hernández-Lobato, B. Sánchez-Lengeling, D. Sheberla, J. Aguilera-Iparraguirre, T. D. Hirzel, R. P. Adams and A. Aspuru-Guzik, ACS Cent. Sci., 2018, 4, 268–276 CrossRef PubMed.
- C. Nie, K. Wang, H. Zhou, J. Deng, Z. Chen, K. Zhang, L. Chen, D. Huang, J. Liang and L. Zhao, ACS Appl. Mater. Interfaces, 2024, 16, 66316–66326 CrossRef CAS PubMed.
- X. Peng, J. Liang, K. Wang, X. Zhao, Z. Peng, Z. Li, J. Zeng, Z. Lan, M. Lei and D. Huang, npj Comput. Mater., 2024, 10, 213 CrossRef CAS.
- X. Zhang, G. Ye, C. Wen and Z. Bi, Comput. Mater. Sci., 2023, 228, 112361 CrossRef CAS.
- M. H. Tahir, N. Sultan, Z. Shafiq, I. M. Moussa, S. Sridhara and M. R. S. A. Janjua, Opt. Quant. Electron., 2024, 56, 1324 CrossRef.
- M. K. Tufail, S. S. A. Shah, S. Khan, F. Ahmad, L. W. Kiruri, M. S. Abbasi and A. Ahmad, Chem. Phys. Lett., 2024, 834, 140974 CrossRef CAS.
- M. Saqib, M. Rani, T. Mubashir, M. H. Tahir, M. Maryam, A. Mushtaq, R. Razzaq, M. A. El-Sheikh and H. O. Elansary, Opt. Mater., 2024, 150, 115295 CrossRef CAS.
- H. Abroshan, H. S. Kwak, Y. An, C. Brown, A. Chandrasekaran, P. Winget and M. D. Halls, Front. Chem., 2022, 9, 800371 CrossRef PubMed.
- A. Nigam, R. Pollice, M. Krenn, G. dos Passos Gomes and A. Aspuru-Guzik, Chem. Sci., 2021, 12, 7079–7090 RSC.
- C. Wechwithayakhlung, G. R. Weal, Y. Kaneko, P. A. Hume, J. M. Hodgkiss and D. M. Packwood, J. Chem. Phys., 2023, 158, 204106 CrossRef CAS PubMed.
- K. M. Katubi, S. Naeem, M. Y. Mehboob, Z. Alrowaili and M. Al-Buriahi, Chem. Phys. Lett., 2023, 813, 140326 CrossRef.
- C. Güleryüz, A. U. Hassan, H. Güleryüz, H. A. Kyhoiesh and M. H. Mahmoud, Mater. Sci. Eng., B, 2025, 317, 118212 CrossRef.
- M.-H. Lee, Adv. Intell. Syst., 2020, 2, 1900108 CrossRef.
- T. Mubashir, M. H. Tahir, M. Mahmoud, Z. Shafiq, M. Ashraf, I. H. El Azab, Z. M. El-Bahy and M. R. S. A. Janjua, J. Photochem. Photobiol., A, 2023, 444, 114977 CrossRef CAS.
- S. K. Pandey and K. Roy, Mater. Today Commun., 2024, 41, 110430 CrossRef CAS.
- M. A. B. Janai, K. L. Woon and C. S. Chan, Org. Electron., 2018, 63, 257–266 CrossRef CAS.
- N. Alwadai, S. U.-D. Khan, Z. M. Elqahtani and S. Ud-Din Khan, Molecules, 2022, 27, 5905 CrossRef CAS PubMed.
- F. Ahmad, A. Mahmood, I. H. El Azab, N. Ahmad, M. Mahmoud and Z. M. El-Bahy, J. Photochem. Photobiol., A, 2024, 453, 115670 CrossRef CAS.
- K. M. Katubi, M. Saqib, M. Sulaman, Z. Alrowaili and M. Al-Buriahi, Chem. Phys., 2024, 582, 112295 CrossRef.
- N. Alfryyan, M. Saqib, S. Ali, T. Mubashir, M. H. Tahir, Z. Alrowaili and M. Al-Buriahi, Mater. Today Commun., 2023, 36, 106556 CrossRef CAS.
- N. Alwadai, Z. M. Elqahtani, S. U.-D. Khan, A. M. Pembere, A. Badshah, M. Y. Mehboob and M. F. Nazar, J. Phys. Org. Chem., 2022, 35, e4388 CrossRef CAS.
- A. Irfan, M. Hussien, M. Y. Mehboob, A. Ahmad and M. R. S. A. Janjua, Energy Technol., 2022, 10, 2101096 CrossRef CAS.
- K. Sato, K. Hattori, F. Uehara, T. Kitaguni, T. Nishiura, T. Yamagata, K. Nomura, N. Matsumoto, T. Tanaka and H. Aihara, Sci. Rep., 2024, 14, 4336 CrossRef CAS PubMed.
- S. Zhong, W. Hsu, H. Chen, T. Yang, J. Yi, C. Zhu, S. Yin, Z. Li, L. Gao and J. Lin, et al. , Sol. RRL, 2024, 8, 2400288 CrossRef CAS.
- D. Brian and X. Sun, J. Phys. Chem. B, 2021, 125, 13267–13278 CrossRef CAS PubMed.
- B. Siddique, T. S. Alomar, M. H. Tahir, N. AlMasoud and Z. M. El-Bahy, J. Photochem. Photobiol., A, 2025, 459, 116026 CrossRef CAS.
- Y. Zhao, C. Fu, L. Fu, Y. Liu, Z. Lu and X. Pu, Mater. Today Chem., 2021, 22, 100625 CrossRef CAS.
- M. Krämer, P. M. Dohmen, W. Xie, D. Holub, A. S. Christensen and M. Elstner, J. Chem. Theor. Comput., 2020, 16, 4061–4070 CrossRef PubMed.
- J. W. Shin, S. Song, Y. Ko, M. K. Choi, C. Y. Go and K. C. Kim, Ind. Eng. Chem. Res., 2024, 63, 16651–16661 CrossRef CAS.
- E. O. Pyzer-Knapp, K. Li and A. Aspuru-Guzik, Adv. Funct. Mater., 2015, 25, 6495–6502 CrossRef CAS.
- A. F. Golin and R. Stefani, J. Supercond. Nov. Magnetism, 2013, 26, 2533–2536 CrossRef CAS.
- P. Reiser, M. Konrad, A. Fediai, S. Léon, W. Wenzel and P. Friederich, J. Chem. Theor. Comput., 2021, 17, 3750–3759 CrossRef CAS PubMed.
- B. Li, H. Sun, H. Shu and X. Wang, ACS Omega, 2021, 7, 168–175 CrossRef PubMed.
- J.-M. Kim, K. H. Lee and J. Y. Lee, Adv. Mater., 2023, 35, 2209953 CrossRef CAS PubMed.
- J. Lim, J.-M. Kim and J. Y. Lee, Adv. Mater., 2024, 36, 2312774 CrossRef CAS PubMed.
- L.-F. Lv, C.-R. Zhang, R. Cao, X.-M. Liu, M.-L. Zhang, J.-J. Gong, Z.-J. Liu, Y.-Z. Wu and H.-S. Chen, J. Mater. Chem. A, 2024, 12, 23859–23871 RSC.
- W. Sun, Y. Zheng, K. Yang, Q. Zhang, A. A. Shah, Z. Wu, Y. Sun, L. Feng, D. Chen and Z. Xiao, et al. , Sci. Adv., 2019, 5, eaay4275 CrossRef CAS PubMed.
- J. D. Tan, B. Ramalingam, V. Chellappan, N. K. Gupta, L. Dillard, S. A. Khan, C. Galvin and K. Hippalgaonkar, ACS Energy Lett., 2024, 9, 5240–5250 CrossRef CAS.
- M. Elkabous, A. Karzazi and Y. Karzazi, Comput. Mater. Sci., 2024, 243, 113146 CrossRef CAS.
- M. Almalki, Y. A. Chapuis and N. Lachiche, Data Science for Photonics and Biophotonics, 2024, pp. 36–50 Search PubMed.
- H. Li, Y. Cui, Y. Liu, W. Li, Y. Shi, C. Fang, H. Li, T. Gao, L. Hu and Y. Lu, IEEE Access, 2018, 6, 34118–34126 Search PubMed.
- M. Rinderle, W. Kaiser, A. Mattoni and A. Gagliardi, J. Phys. Chem. C, 2020, 124, 17733–17743 CrossRef CAS.
- T. Hao, S. Leng, Y. Yang, W. Zhong, M. Zhang, L. Zhu, J. Song, J. Xu, G. Zhou and Y. Zou, et al. , Patterns, 2021, 2, 100333 CrossRef CAS PubMed.
- J. Fan, B. Yuan, C. Qian and S. Zhou, Precis. Chem., 2024, 2, 263–272 CrossRef CAS PubMed.
- M. Nakata and T. Shimazaki, J. Chem. Inf. Model., 2017, 57, 1300–1308 CrossRef CAS PubMed.
- M. Seifrid, S. Lo, D. G. Choi, G. Tom, M. L. Le, K. Li, R. Sankar, H.-T. Vuong, H. Wakidi and A. Yi, et al. , J. Mater. Chem. A, 2024, 12, 14540–14558 RSC.
- A. Stuke, M. Todorović, M. Rupp, C. Kunkel, K. Ghosh, L. Himanen and P. Rinke, J. Chem. Phys., 2019, 150, 204121 CrossRef PubMed.
- D. Padula, J. D. Simpson and A. Troisi, Mater. Horiz., 2019, 6, 343–349 RSC.
- J. Staker, K. Marshall, K. Leswing, T. Robertson, M. D. Halls, A. Goldberg, T. Morisato, H. Maeshima, T. Ando and H. Arai, et al. , J. Phys. Chem. A, 2022, 126, 5837–5852 CrossRef CAS PubMed.
- Y. Li, Y. Xu and Y. Yu, Molecules, 2021, 26, 7257 CrossRef CAS PubMed.
- G. Marques, K. Leswing, T. Robertson, D. Giesen, M. D. Halls, A. Goldberg, K. Marshall, J. Staker, T. Morisato and H. Maeshima, et al. , J. Phys. Chem. A, 2021, 125, 7331–7343 CrossRef CAS PubMed.
- R. J. Richards and A. Paul, Sol. Energy, 2021, 224, 43–50 CrossRef CAS.
- C. Lu, Q. Liu, Q. Sun, C.-Y. Hsieh, S. Zhang, L. Shi and C.-K. Lee, J. Phys. Chem. C, 2020, 124, 7048–7060 CrossRef CAS.
- C. Kunkel, J. T. Margraf, K. Chen, H. Oberhofer and K. Reuter, Nat. Commun., 2021, 12, 2422 CrossRef CAS PubMed.
- A. J. Bornschlegl, P. Duchstein, J. Wu, J. S. Rocha-Ortiz, M. Caicedo-Reina, A. Ortiz, B. Insuasty, D. Zahn, L. Luer and C. J. Brabec, J. Am. Chem. Soc., 2025, 147, 1957–1967 CrossRef CAS PubMed.
- T. Won, N. Aizawa, Y. Harabuchi, R. Kurihara, M. Suzuki, S. Maeda, Y.-J. Pu and K.-i. Nakayama, Chem. Sci., 2025, 16, 9303–9310 RSC.
- D. M. Packwood, Y. Kaneko, D. Ikeda and M. Ohno, Adv. Theory Simul., 2023, 6, 2300159 CrossRef CAS.
- X. Rodríguez-Martínez, E. Pascual-San-José, Z. Fei, M. Heeney, R. Guimerà and M. Campoy-Quiles, Energy Environ. Sci., 2021, 14, 986–994 RSC.
- A. Nigam, R. Pollice, P. Friederich and A. Aspuru-Guzik, Chem. Sci., 2024, 15, 2618–2639 RSC.
- B. L. Greenstein and G. R. Hutchison, J. Phys. Chem. C, 2023, 127, 6179–6191 CrossRef CAS.
- X. Wang, S. Wang, J. Wang and S. Yin, J. Phys. Chem. A, 2023, 127, 5930–5941 CrossRef CAS PubMed.
- A. Yakubovich, A. Odinokov, S. Nikolenko, Y. Jung and H. Choi, Front. Chem., 2021, 9, 800133 CrossRef CAS PubMed.
- Y. Shi, H. Shi, H. Wang, C.-J. Chen, Y. Li, B. Qiao, Z. Liang, S. Zhao, D. Hang and Z. Xu, et al. , Chem. Eng. J., 2024, 500, 157082 CrossRef CAS.
- Z. Tan, Y. Li, Z. Zhang, X. Wu, T. Penfold, W. Shi and S. Yang, ACS Omega, 2022, 7, 18179–18188 CrossRef CAS PubMed.
- W. Xu, H. Chen, R. He, X. Song, L. Ma and J. Song, SID Symposium Digest of Technical Papers, 2024, pp. 2163–2166 Search PubMed.
- Z. Xie, X. Evangelopoulos, Ö. H. Omar, A. Troisi, A. I. Cooper and L. Chen, Chem. Sci., 2024, 15, 500–510 RSC.
- D. Chicco, M. J. Warrens and G. Jurman, PeerJ Comput. Sci., 2021, 7, e623 CrossRef PubMed.
- G. M. Foody, PLoS One, 2023, 18, e0291908 CrossRef CAS PubMed.
- D. J. Hand and R. J. Till, Mach. Learn., 2001, 45, 171–186 CrossRef.
- J. Eng, Acad. Radiol., 2005, 12, 909–916 CrossRef PubMed.
- Z. Zhang, Q. Liu, C.-K. Lee, C.-Y. Hsieh and E. Chen, Chem. Sci., 2023, 14, 8380–8392 RSC.
- T. Khater, S. A. Alkhatib, A. AlShehhi, C. Pitsalidis, A. M. Pappa, S. T. Ngo, V. Chan and V. K. Truong, J. Cheminf., 2025, 17, 116 Search PubMed.
- S.-Y. Lu, S. Mukhopadhyay, R. Froese and P. M. Zimmerman, J. Chem. Inf. Model., 2018, 58, 2440–2449 CrossRef CAS PubMed.
- S. An, Y. H. Jung, G. Nam, E. Jeon, J. H. Ham, S. C. Cha, M. Y. Chae, J. H. Kwon and Y. Jung, Chem. Eng. J., 2025, 505, 159697 CrossRef CAS.
- J.-L. Brédas, J. E. Norton, J. Cornil and V. Coropceanu, Accounts Chem. Res., 2009, 42, 1691–1699 CrossRef PubMed.
- S. R. Forrest, Philos. Trans. R. Soc. A Math. Phys. Eng. Sci., 2015, 373, 20140320 CrossRef PubMed.
- P. I. Djurovich, E. I. Mayo, S. R. Forrest and M. E. Thompson, Org. Electron., 2009, 10, 515–520 CrossRef CAS.
- S. J. Akram, N. Hadia, A. M. Shawky, J. Iqbal, M. I. Khan, N. S. Alatawi, M. A. Ibrahim, M. Ans and R. A. Khera, ACS Omega, 2023, 8, 11118–11137 CrossRef CAS PubMed.
- X. Wu, X. Xie and A. Troisi, J. Mater. Chem. C, 2024, 12, 18886–18892 RSC.
- G. Onida, L. Reining and A. Rubio, Rev. Mod. Phys., 2002, 74, 601 CrossRef CAS.
- G. Montavon, M. Rupp, V. Gobre, A. Vazquez-Mayagoitia, K. Hansen, A. Tkatchenko, K.-R. Müller and O. A. Von Lilienfeld, New J. Phys., 2013, 15, 095003 CrossRef CAS.
- F. Pereira, K. Xiao, D. A. Latino, C. Wu, Q. Zhang and J. Aires-de Sousa, J. Chem. Inf. Model., 2017, 57, 11–21 CrossRef CAS PubMed.
- D. Rogers and M. Hahn, J. Chem. Inf. Model., 2010, 50, 742–754 CrossRef CAS PubMed.
- V. Vovk, Empirical Inference: Festschrift in Honor of Vladimir N. Vapnik, Springer, 2013, pp. 105–116 Search PubMed.
- R. Ramakrishnan, P. O. Dral, M. Rupp and O. A. Von Lilienfeld, Sci. Data, 2014, 1, 1–7 Search PubMed.
- M. Ropo, M. Schneider, C. Baldauf and V. Blum, Sci. Data, 2016, 3, 1–13 Search PubMed.
- C. R. Groom, I. J. Bruno, M. P. Lightfoot and S. C. Ward, Struct. Sci., 2016, 72, 171–179 CAS.
- C. Gaul and S. Cuesta-Lopez, Phys. Status Solidi B, 2024, 261, 2200553 CrossRef CAS.
- K. Schütt, P.-J. Kindermans, H. E. Sauceda Felix, S. Chmiela, A. Tkatchenko and K.-R. Müller, Adv. Neural Inf. Process. Syst., 2017, 30, 992–1002 Search PubMed.
- F. A. Schröder, D. H. Turban, A. J. Musser, N. D. Hine and A. W. Chin, Nat. Commun., 2019, 10, 1062 CrossRef PubMed.
- X. Liu, X. Wang, S. Gao, V. Chang, R. Tom, M. Yu, L. M. Ghiringhelli and N. Marom, npj Comput. Mater., 2022, 8, 70 CrossRef.
- S. Gao, Y. Luo, X. Liu and N. Marom, Digital Discovery, 2025, 305, 121128 Search PubMed.
- L. Torrey and J. Shavlik, Handbook of Research on Machine Learning Applications and Trends: Algorithms, Methods, and Techniques, IGI Global Scientific Publishing, 2010, pp. 242–264 Search PubMed.
- M. Jeong, J. F. Joung, J. Hwang, M. Han, C. W. Koh, D. H. Choi and S. Park, npj Comput. Mater., 2022, 8, 147 CrossRef CAS.
- P. W. Butler, R. Hafizi and G. M. Day, J. Phys. Chem. A, 2024, 128, 945–957 CrossRef CAS PubMed.
- M. Saqib, M. Sagir, M. L. Joshi, S. Bashir, M. I. Halawa, S. Ali, H. O. Elansary and G. M. Kamal, Mater. Today Commun., 2024, 38, 108062 CrossRef CAS.
- K. Huwig, C. Fan and M. Springborg, J. Chem. Phys., 2017, 147, 234105 CrossRef PubMed.
- P. M. Tagade, S. P. Adiga, S. Pandian, M. S. Park, K. S. Hariharan and S. M. Kolake, npj Comput. Mater., 2019, 5, 127 CrossRef.
- Y. Kwon, S. Kang, Y.-S. Choi and I. Kim, Sci. Rep., 2021, 11, 17304 CrossRef CAS PubMed.
- A. S. Khazaal, M. Sprinborg, C. Fan and K. Huwig, Comput. Condens. Matter, 2020, 25, e00503 CrossRef.
- Q. Yuan, A. Santana-Bonilla, M. A. Zwijnenburg and K. E. Jelfs, Nanoscale, 2020, 12, 6744–6758 RSC.
- T. Weiss, E. Mayo Yanes, S. Chakraborty, L. Cosmo, A. M. Bronstein and R. Gershoni-Poranne, Nat. Comput. Sci., 2023, 3, 873–882 CrossRef PubMed.
- S. Xu, J. Li, P. Cai, X. Liu, B. Liu and X. Wang, J. Am. Chem. Soc., 2021, 143, 19769–19777 CrossRef CAS PubMed.
- A. S. Khazaal, M. Springborg, C. Fan and K. Huwig, J. Mol. Graph. Model., 2020, 100, 107654 CrossRef CAS PubMed.
- D. P. Kingma, M. Welling, et al., arXiv, 2013, preprint, arXiv:1312.6114, DOI:10.48550/arXiv.1312.6114.
- R. S. Sutton, A. G. Barto, et al., Reinforcement Learning: An Introduction, MIT press, Cambridge, 1998, vol. 1 Search PubMed.
- J. H. Holland, Adaptation in Natural and Artificial Systems: An Introductory Analysis with Applications to Biology, Control, and Artificial Intelligence, MIT press, 1992 Search PubMed.
- B. Sanchez-Lengeling and A. Aspuru-Guzik, Science, 2018, 361, 360–365 CrossRef CAS PubMed.
- C.-H. Li and D. P. Tabor, Chem. Sci., 2023, 14, 11045–11055 RSC.
- A. Ohno, J.-i. Hanna, H. Iino, K. Nakago, T. Yamaguchi, M. Abe, H. Akita and M. Takemoto, Chem.–Asian J., 2023, 18, e202300029 CrossRef CAS PubMed.
- R. Tom, S. Gao, Y. Yang, K. Zhao, I. Bier, E. A. Buchanan, A. Zaykov, Z. Havlas, J. Michl and N. Marom, Chem. Mater., 2023, 35, 1373–1386 CrossRef CAS PubMed.
- M. Han, J. F. Joung, M. Jeong, D. H. Choi and S. Park, ACS Cent. Sci., 2024, 11, 219–227 CrossRef PubMed.
- J. Westermayr, J. Gilkes, R. Barrett and R. J. Maurer, Nat. Comput. Sci., 2023, 3, 139–148 CrossRef CAS PubMed.
- M. Einax and A. Nitzan, J. Phys. Chem. C, 2014, 118, 27226–27234 CrossRef CAS.
- M. Ernzerhof, M.-A. Bélanger, D. Mayou and T. Nemati Aram, J. Chem. Phys., 2016, 144, 134102 CrossRef PubMed.
- T. N. Aram, A. Asgari, M. Ernzerhof, P. Quémerais and D. Mayou, EPJ Photovoltaics, 2017, 8, 85503 CrossRef.
- E. K. Solak and E. Irmak, RSC Adv., 2023, 13, 12244–12269 RSC.
- M. C. Scharber, D. Mühlbacher, M. Koppe, P. Denk, C. Waldauf, A. J. Heeger and C. J. Brabec, Adv. Mater., 2006, 18, 789–794 CrossRef CAS.
- T. N. Aram, M. Ernzerhof, A. Asgari and D. Mayou, J. Chem. Phys., 2018, 149, 064102 CrossRef PubMed.
- T. N. Aram, A. Asgari and D. Mayou, Europhys. Lett., 2016, 115, 18003 CrossRef.
- Y. Takeda, Acc. Chem. Res., 2024, 57, 2219–2232 CrossRef CAS PubMed.
- Z.-W. Zhao, O. H. Omar, D. Padula, Y. Geng and A. Troisi, J. Phys. Chem. Lett., 2021, 12, 5009–5015 CrossRef CAS PubMed.
- A. Mahmood and J.-L. Wang, Energy Environ. Sci., 2021, 14, 90–105 RSC.
- Y. Lin, Y. Li and X. Zhan, Chem. Soc. Rev., 2012, 41, 4245–4272 RSC.
- Y. Morishita, M. Yarimizu, M. Kaneko and A. Muraoka, Chem. Phys. Lett., 2024, 857, 141719 CrossRef CAS.
- S. Zhang, S. Li, S. Song, Y. Zhao, L. Gao, H. Chen, H. Li and J. Lin, Adv. Mater., 2025, 37, 2407613 CrossRef PubMed.
- H. Sahu, W. Rao, A. Troisi and H. Ma, Adv. Energy Mater., 2018, 8, 1801032 CrossRef.
- M. R. S. A. Janjua, A. Irfan, M. Hussien, M. Ali, M. Saqib and M. Sulaman, Energy Technol., 2022, 10, 2200019 CrossRef CAS.
- C.-H. Yang, B. S. S. Pokuri, X. Y. Lee, S. Balakrishnan, C. Hegde, S. Sarkar and B. Ganapathysubramanian, Comput. Mater. Sci., 2022, 213, 111599 CrossRef.
- M.-H. Lee, Sol. Energy, 2024, 267, 112191 CrossRef CAS.
- B. Yang, C.-R. Zhang, Y. Wang, M. Zhao, H.-Y. Yu, Z.-J. Liu, X.-M. Liu, Y.-H. Chen, Y.-Z. Wu and H.-S. Chen, Int. J. Quantum Chem., 2023, 123, e27039 CrossRef CAS.
- C. Liu, L. Lüer, V. M. L. Corre, K. Forberich, P. Weitz, T. Heumüller, X. Du, J. Wortmann, J. Zhang and J. Wagner, et al. , Adv. Mater., 2024, 36, 2300259 CrossRef CAS PubMed.
- M. Vubangsi, A. S. Mubarak and F. Al-Turjman, Energy Rep., 2024, 11, 3824–3835 CrossRef.
- X. Du, L. Lüer, T. Heumueller, J. Wagner, C. Berger, T. Osterrieder, J. Wortmann, S. Langner, U. Vongsaysy and M. Bertrand, et al. , Joule, 2021, 5, 495–506 CrossRef CAS.
- L. Ju, M. Li, L. Tian, P. Xu and W. Lu, Mater. Today Commun., 2020, 25, 101604 CrossRef CAS.
- W. Sun, Y. Zheng, Q. Zhang, K. Yang, H. Chen, Y. Cho, J. Fu, O. Odunmbaku, A. A. Shah and Z. Xiao, et al. , J. Phys. Chem. Lett., 2021, 12, 8847–8854 CrossRef CAS PubMed.
- B. L. Greenstein, D. C. Hiener and G. R. Hutchison, J. Chem. Phys., 2022, 156, 174107 CrossRef CAS PubMed.
- T. W. David and J. Kettle, J. Phys. Chem. C, 2022, 126, 4774–4784 CrossRef CAS.
- M. Hosseinnezhad, M. R. Saeb, S. Garshasbi and Y. Mohammadi, Sol. Energy, 2017, 149, 314–322 CrossRef CAS.
- L. Keller, D. Tanneberg, S. Stark and J. Peters, 2020 IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS), 2020, pp. 9675–9680 Search PubMed.
- P. Li, J. Hao, H. Tang, X. Fu, Y. Zhen, K. Tang, arXiv, 2024, preprint, arXiv:2401.11963, DOI:10.48550/arXiv.2401.11963.
- D. P.-K. Tsang, T. Matsushima and C. Adachi, Sci. Rep., 2016, 6, 22463 Search PubMed.
- I. Siddiqui, S. Kumar, Y.-F. Tsai, P. Gautam, Shahnawaz, K. Kesavan, J.-T. Lin, L. Khai, K.-H. Chou and A. Choudhury, et al. , Nanomaterials, 2023, 13, 2521 CrossRef CAS PubMed.
- L. Zhao, J. Li, L. Li and W. Hu, J. Mater. Chem. C, 2024, 12, 13745–13761 RSC.
- A. Brown, K. Pichler, N. Greenham, D. Bradley, R. H. Friend and A. Holmes, Chem. Phys. Lett., 1993, 210, 61–66 Search PubMed.
- S. Kappaun, C. Slugovc and E. J. List, Int. J. Mol. Sci., 2008, 9, 1527–1547 Search PubMed.
- M. A. Baldo, D. F. O'Brien, Y. You, A. Shoustikov, S. Sibley, M. E. Thompson and S. R. Forrest, Electrophosphorescent Materials and Devices, Jenny Stanford Publishing, 2023, pp. 1–11 Search PubMed.
- J. M. Dos Santos, D. Hall, B. Basumatary, M. Bryden, D. Chen, P. Choudhary, T. Comerford, E. Crovini, A. Danos and J. De, et al. , Chem. Rev., 2024, 124, 13736–14110 CrossRef CAS PubMed.
- H. Uoyama, K. Goushi, K. Shizu, H. Nomura and C. Adachi, Nature, 2012, 492, 234–238 CrossRef CAS PubMed.
- N. Aizawa, Y.-J. Pu, Y. Harabuchi, A. Nihonyanagi, R. Ibuka, H. Inuzuka, B. Dhara, Y. Koyama, K.-i. Nakayama and S. Maeda, et al. , Nature, 2022, 609, 502–506 CrossRef CAS PubMed.
- R. Pollice, P. Friederich, C. Lavigne, G. dos Passos Gomes and A. Aspuru-Guzik, Matter, 2021, 4, 1654–1682 CrossRef CAS.
- Y. Bu and Q. Peng, J. Phys. Chem. C, 2023, 127, 23845–23851 CrossRef CAS.
- J. F. Joung, M. Han, M. Jeong and S. Park, J. Chem. Inf. Model., 2022, 62, 2933–2942 CrossRef CAS PubMed.
- J. F. Joung, M. Han, J. Hwang, M. Jeong, D. H. Choi and S. Park, JACS Au, 2021, 1, 427–438 CrossRef CAS PubMed.
- Z. He, H. Bi, B. Liang, Z. Li, H. Zhang and Y. Wang, Light: Sci. Appl., 2025, 14, 75 CrossRef CAS PubMed.
- R. C. Souza, J. C. Duarte, R. R. Goldschmidt and I. Borges Jr, J. Chem. Inf. Model., 2025, 65, 3270–3281 CrossRef CAS PubMed.
- J. H. Kim, H. Kim and W. Y. Kim, Bull. Korean Chem. Soc., 2022, 43, 645–649 CrossRef CAS.
- R. Nikhitha and A. Mondal, J. Chem. Phys., 2025, 162, 144103 CrossRef PubMed.
- H. S. Kwak, D. J. Giesen, T. F. Hughes, A. Goldberg, Y. Cao, J. Gavartin, S. Dixon and M. D. Halls, Organic Light Emitting Materials and Devices XX, 2016, pp. 121–129 Search PubMed.
- D. D. Tarakanovskaya and E. A. Mostovich, J. Phys. Chem. A, 2025, 129, 4458–4470 CrossRef CAS PubMed.
- K. L. Woon, Z. X. Chong, A. Ariffin and C. S. Chan, J. Mol. Graph. Model., 2021, 105, 107891 CrossRef CAS PubMed.
- L. Barneschi, L. Rotondi and D. Padula, J. Phys. Chem. A, 2024, 128, 2417–2426 CrossRef CAS PubMed.
- A. E. Sifain, L. Lystrom, R. A. Messerly, J. S. Smith, B. Nebgen, K. Barros, S. Tretiak, N. Lubbers and B. J. Gifford, Chem. Sci., 2021, 12, 10207–10217 RSC.
- C. K. Borislavova, S. B. Djumayska, Y. D. Zagranyarski and A. N. Ivanova, Theor. Chem. Acc., 2024, 143, 71 Search PubMed.
- Y. Li, B. Zhang, A. Ren, D. Wang, J. Zhang, C. Nie, Z. Su and L. Zou, Chem. Eng. J., 2024, 501, 157676 CrossRef CAS.
- M.-H. Lee, Phys. Chem. Chem. Phys., 2020, 22, 16378–16386 RSC.
- H. Shi, W. Jing, W. Liu, Y. Li, Z. Li, B. Qiao, S. Zhao, Z. Xu and D. Song, ACS Omega, 2022, 7, 7893–7900 CrossRef CAS PubMed.
- W. Lu, L. Zhiyang, L. Shuyao, W. Zhipeng, S. Entao, L. Song and G. Wenzheng, SID Symposium Digest of Technical Papers, 2024, pp. 53–55 Search PubMed.
- H. Guo, G. Jiang, B. Diao, J. Du, W. Sun, J. Fan and X. Peng, J. Mater. Chem. C, 2024, 12, 14515–14522 RSC.
- Y. Zhao, K. Chen, L. Zhu and Q. Huang, Dyes Pigm., 2023, 220, 111670 CrossRef CAS.
- Y. Zhao, K. Chen, B. Yu, Q. Wan, Y. Wang, F. Tang and X. Li, J. Mol. Struct., 2024, 1317, 139126 CrossRef CAS.
- Y. Shi, H. Shi, Y. Zhang, X. Zang, Z. Zhao, S. Zhao, B. Qiao, Z. Liang, Z. Xu and L. Wang, et al. , Adv. Opt. Mater., 2024, 12, 2301768 CrossRef CAS.
- T. Lee, J. Choi, I. Na, I. Yoo, S. Woo, K. J. Kim, M. Park, J. Yang, J. Min and S. Lee, et al. , Adv. Intell. Syst., 2024, 2400598 Search PubMed.
- P. Li, Z. Wang, W. Li, J. Yuan and R. Chen, J. Phys. Chem. Lett., 2022, 13, 9910–9918 CrossRef CAS PubMed.
- X. Niu, Z. Su, L. Wang, W. Shi, H. Zhang, Y. Dang, Y. Yuan, Y. Sun and W. Hu, J. Mater. Inf., 2025, 5, 45 CrossRef.
- C. Tu, W. Huang, S. Liang, K. Wang, Q. Tian and W. Yan, RSC Adv., 2022, 12, 30962–30975 RSC.
- W. Xu, J. Shen, H. Chen, R. He, X. Song, Z. Xia, L. Ma and J. Song, SID Symposium Digest of Technical Papers, 2023, pp. 1571–1574 Search PubMed.
- H. S. Kwak, Y. An, D. J. Giesen, T. F. Hughes, C. T. Brown, K. Leswing, H. Abroshan and M. D. Halls, Front. Chem., 2022, 9, 800370 CrossRef PubMed.
- S. Kim, J. M. Shin, J. Lee, C. Park, S. Lee, J. Park, D. Seo, S. Park, C. Y. Park and M. S. Jang, Nanophotonics, 2021, 10, 4533–4541 Search PubMed.
- J. Lim, S. Ryu, J. W. Kim and W. Y. Kim, J. Cheminf., 2018, 10, 1–9 Search PubMed.
- V. Coropceanu, J. Cornil, D. A. da Silva Filho, Y. Olivier, R. Silbey and J.-L. Brédas, Chem. Rev., 2007, 107, 926–952 Search PubMed.
- A. Troisi, Chem. Soc. Rev., 2011, 40, 2347–2358 Search PubMed.
- G. Schweicher, Y. Olivier, V. Lemaur and Y. H. Geerts, Isr. J. Chem., 2014, 54, 595–620 Search PubMed.
- S. H. Glarum, J. Phys. Chem. Solids, 1963, 24, 1577–1583 CrossRef CAS.
- F. W. Schmidlin, Philos. Mag. B, 1980, 41, 535–570 CAS.
- A. Troisi, J. Chem. Phys., 2011, 134, 034702 CrossRef PubMed.
- S. Fratini, D. Mayou and S. Ciuchi, Adv. Funct. Mater., 2016, 26, 2292–2315 CrossRef CAS.
- T. Nematiaram, S. Ciuchi, X. Xie, S. Fratini and A. Troisi, J. Phys. Chem. C, 2019, 123, 6989–6997 CrossRef CAS.
- H. Oberhofer, K. Reuter and J. Blumberger, Chem. Rev., 2017, 117, 10319–10357 CrossRef CAS PubMed.
- T. Nematiaram and A. Troisi, J. Chem. Phys., 2020, 152, 190902 CrossRef CAS PubMed.
- C. Liu, K. Huang, W.-T. Park, M. Li, T. Yang, X. Liu, L. Liang, T. Minari and Y.-Y. Noh, Mater. Horiz., 2017, 4, 608–618 RSC.
- S. Giannini and J. Blumberger, Acc. Chem. Res., 2022, 55, 819–830 CrossRef CAS PubMed.
- J. Ostmeyer, T. Nematiaram, A. Troisi and P. Buividovich, Phys. Rev. Appl., 2024, 22, L031004 CrossRef CAS.
- T. Nematiaram, D. Padula, A. Landi and A. Troisi, Adv. Funct. Mater., 2020, 30, 2001906 CrossRef CAS.
- S. Yang, M. Sun, C. Shi, Y. Liu, Y. Guo, Y. Liu, Z. Lu, Y. Huang and X. Pu, J. Chem. Theory Comput., 2024, 20, 10633–10648 CrossRef CAS PubMed.
- T. Ando, N. Shimizu, N. Yamamoto, N. N. Matsuzawa, H. Maeshima and H. Kaneko, J. Phys. Chem. A, 2022, 126, 6336–6347 CrossRef CAS PubMed.
- R. Kawagoe, T. Ando, N. N. Matsuzawa, H. Maeshima and H. Kaneko, ACS Omega, 2024, 9, 48844–48854 CrossRef CAS PubMed.
- X. Niu, Y. Dang, Y. Sun and W. Hu, J. Energy Chem., 2023, 81, 143–148 CrossRef CAS.
- K. M. Katubi, M. Saqib, M. Maryam, T. Mubashir, M. H. Tahir, M. Sulaman, Z. Alrowaili and M. Al-Buriahi, Inorg. Chem. Commun., 2023, 151, 110610 CrossRef CAS.
- I. F. Graña, S. Varsamopoulos, T. Ando, H. Maeshima and N. N. Matsuzawa, arXiv, 2025, preprint, arXiv:2503.09517, DOI:10.48550/arXiv.2503.09517.
- S. Atahan-Evrenk and F. B. Atalay, J. Phys. Chem. A, 2019, 123, 7855–7863 CrossRef CAS PubMed.
- K. Chen, C. Kunkel, K. Reuter and J. T. Margraf, Digital Discovery, 2022, 1, 147–157 RSC.
- C.-H. Li and D. P. Tabor, J. Phys. Chem. A, 2023, 127, 3484–3489 CrossRef CAS PubMed.
- A. E. Hoerl and R. W. Kennard, Technometrics, 1970, 12, 55–67 CrossRef.
- C. Saunders, A. Gammerman and V. Vovk, Proceedings of the 15th International Conference on Machine Learning, ICML '98, 1998, pp. 515-521 Search PubMed.
- Y. LeCun, Y. Bengio and G. Hinton, Nature, 2015, 521, 436–444 CrossRef CAS PubMed.
- T. N. Kipf and M. Welling, arXiv, 2016, preprint, arXiv:1609.02907, DOI:10.48550/arXiv.1609.02907.
- M. Seeger, Int. J. Neural Syst., 2004, 14, 69–106 CrossRef PubMed.
- A. Troisi and G. Orlandi, Chem. Phys. Lett., 2001, 344, 509–518 CrossRef CAS.
- C.-I. Wang, M. K. E. Braza, G. C. Claudio, R. B. Nellas and C.-P. Hsu, J. Phys. Chem. A, 2019, 123, 7792–7802 CrossRef CAS PubMed.
- C.-I. Wang, I. Joanito, C.-F. Lan and C.-P. Hsu, J. Chem. Phys., 2020, 153, 214113 CrossRef CAS PubMed.
- M. Elstner, D. Porezag, G. Jungnickel, J. Elsner, M. Haugk, T. Frauenheim, S. Suhai and G. Seifert, Phys. Rev. B: Condens. Matter Mater. Phys., 1998, 58, 7260 CrossRef CAS.
- V. Bhat, B. Ganapathysubramanian and C. Risko, J. Phys. Chem. Lett., 2024, 15, 7206–7213 CrossRef CAS PubMed.
- T. Nematiaram and A. Troisi, Mater. Horiz., 2020, 7, 2922–2928 RSC.
- Y.-S. Wang, C.-I. Wang, C.-H. Yang and C.-P. Hsu, J. Chem. Phys., 2023, 159, 034103 CrossRef CAS PubMed.
- V. Dantanarayana, T. Nematiaram, D. Vong, J. E. Anthony, A. Troisi, K. Nguyen Cong, N. Goldman, R. Faller and A. J. Moulé, J. Chem. Theor. Comput., 2020, 16, 3494–3503 CrossRef CAS PubMed.
- D. Vong, T. Nematiaram, M. A. Dettmann, T. L. Murrey, L. S. Cavalcante, S. M. Gurses, D. Radhakrishnan, L. L. Daemen, J. E. Anthony and K. J. Koski, et al. , J. Phys. Chem. Lett., 2022, 13, 5530–5537 CrossRef CAS PubMed.
- J. Lederer, W. Kaiser, A. Mattoni and A. Gagliardi, Adv. Theory Simul., 2019, 2, 1800136 CrossRef.
- T. Tan, L. Duan and D. Wang, Adv. Funct. Mater., 2024, 34, 2313085 CrossRef CAS.
- M.-H. Lee, Adv. Electron. Mater., 2019, 5, 1900573 CrossRef CAS.
- E. Antono, N. N. Matsuzawa, J. Ling, J. E. Saal, H. Arai, M. Sasago and E. Fujii, J. Phys. Chem. A, 2020, 124, 8330–8340 CrossRef CAS PubMed.
- C. Kunkel, C. Schober, J. T. Margraf, K. Reuter and H. Oberhofer, Chem. Mater., 2019, 31, 969–978 CrossRef CAS.
- Ö. H. Omar, T. Nematiaram, A. Troisi and D. Padula, Sci. Data, 2022, 9, 54 CrossRef PubMed.
- J. F. Joung, M. Han, M. Jeong and S. Park, Sci. Data, 2020, 7, 295 CrossRef CAS PubMed.
- S. A. Lopez, E. O. Pyzer-Knapp, G. N. Simm, T. Lutzow, K. Li, L. R. Seress, J. Hachmann and A. Aspuru-Guzik, Sci. Data, 2016, 3, 1–7 Search PubMed.
- D. Jha, K. Choudhary, F. Tavazza, W.-k. Liao, A. Choudhary, C. Campbell and A. Agrawal, Nat. Commun., 2019, 10, 5316 CrossRef CAS PubMed.
- Z.-W. Zhao, M. Del Cueto and A. Troisi, Digital Discovery, 2022, 1, 266–276 RSC.
- J. Hu, S. Stefanov, Y. Song, S. S. Omee, S.-Y. Louis, E. M. Siriwardane, Y. Zhao and L. Wei, npj Comput. Mater., 2022, 8, 65 CrossRef.
- M. D. Wilkinson, M. Dumontier, I. J. Aalbersberg, G. Appleton, M. Axton, A. Baak, N. Blomberg, J.-W. Boiten, L. B. da Silva Santos and P. E. Bourne, et al. , Sci. Data, 2016, 3, 1–9 CrossRef PubMed.
- P. Raccuglia, K. C. Elbert, P. D. Adler, C. Falk, M. B. Wenny, A. Mollo, M. Zeller, S. A. Friedler, J. Schrier and A. J. Norquist, Nature, 2016, 533, 73–76 CrossRef CAS PubMed.
- H. Abroshan, H. S. Kwak, A. Chandrasekaran, A. K. Chew, A. Fonari and M. D. Halls, Chem. Mater., 2023, 35, 5059–5070 CrossRef CAS.
- M. Randić and J. Zupan, J. Chem. Inf. Comput. Sci., 2001, 41, 550–560 CrossRef PubMed.
- Y. Kim, Y. Jeong, J. Kim, E. K. Lee, W. J. Kim and I. S. Choi, Chem.–Asian J., 2022, 17, e202200269 CrossRef CAS PubMed.
- S. Batzner, A. Musaelian, L. Sun, M. Geiger, J. P. Mailoa, M. Kornbluth, N. Molinari, T. E. Smidt and B. Kozinsky, Nat. Commun., 2022, 13, 2453 CrossRef CAS PubMed.
- S. Axelrod and R. Gomez-Bombarelli, Mach. Learn.: Sci. Technol., 2023, 4, 035025 Search PubMed.
- S. Varghese and S. Das, J. Phys. Chem. Lett., 2011, 2, 863–873 CrossRef CAS PubMed.
- T. Nematiaram, D. Padula and A. Troisi, Chem. Mater., 2021, 33, 3368–3378 CrossRef CAS PubMed.
- T. Nematiaram and A. Troisi, Chem. Mater., 2022, 34, 4050–4061 CrossRef CAS PubMed.
- H. M. Johnson, F. Gusev, J. T. Dull, Y. Seo, R. D. Priestley, O. Isayev and B. P. Rand, J. Am. Chem. Soc., 2024, 146, 21583–21590 CrossRef PubMed.
- R. P. Sheridan, J. Chem. Inf. Model., 2013, 53, 783–790 CrossRef CAS PubMed.
- Q.-Y. Meng, R. Wang, H.-Y. Shao, Y.-L. Wang, X.-L. Wen, C.-Y. Yao and J. Qiao, J. Phys. Chem. Lett., 2024, 15, 4422–4429 CrossRef CAS PubMed.
- Y. Wu, J. Guo, R. Sun and J. Min, npj Comput. Mater., 2020, 6, 120 CrossRef CAS.
- A. S. Anker, A. Aspuru-Guzik, C. B. Mahmoud, S. Bennett, K. R. Briling, A. Changiarath, S. Chong, C. M. Collins, A. I. Cooper and D. Crusius, et al. , Faraday Discuss., 2025, 256, 373–412 RSC.
- J. M. Shin, S. Kim, S. G. Menabde, S. Park, I.-G. Lee, I. Kim and M. S. Jang, Nanophotonics, 2025, 14, 1091–1099 CrossRef CAS PubMed.
- E. Knapp, M. Battaglia, T. Stadelmann, S. Jenatsch and B. Ruhstaller, 2021 8th Swiss Conference on Data Science (SDS), 2021, pp. 46–51 Search PubMed.
- K. T. Butler, D. W. Davies, H. Cartwright, O. Isayev and A. Walsh, Nature, 2018, 559, 547–555 CrossRef CAS PubMed.
- L. Breiman, Mach. Learn., 1996, 24, 123–140 CrossRef.
- I. Kononenko, Biol. Cybern., 1989, 61, 361–370 CrossRef.
- D. Milanés-Hermosilla, R. Trujillo Codorniú, R. López-Baracaldo, R. Sagaró-Zamora, D. Delisle-Rodriguez, J. J. Villarejo-Mayor and J. R. Núñez-Álvarez, Sensors, 2021, 21, 7241 CrossRef PubMed.
- A. Lemay, K. Hoebel, C. P. Bridge, B. Befano, S. De Sanjosé, D. Egemen, A. C. Rodriguez, M. Schiffman, J. P. Campbell and J. Kalpathy-Cramer, npj Digit. Med., 2022, 5, 174 CrossRef PubMed.
- J. Padarian, B. Minasny and A. McBratney, Geoderma, 2022, 425, 116063 CrossRef.
- Z.-W. Zhao, M. del Cueto, Y. Geng and A. Troisi, Chem. Mater., 2020, 32, 7777–7787 CrossRef CAS.
- C.-T. Lu, D. Chen and Y. Kou, Third IEEE International Conference on Data Mining, 2003, pp. 597–600 Search PubMed.
- A. R. Tan, S. Urata, S. Goldman, J. C. Dietschreit and R. Gómez-Bombarelli, npj Comput. Mater., 2023, 9, 225 CrossRef CAS.
- G. Gryn’ova, K.-H. Lin and C. Corminboeuf, J. Am. Chem. Soc., 2018, 140, 16370–16386 Search PubMed.
- J. Zeng, T. J. Giese, S. Ekesan and D. M. York, J. Chem. Theor. Comput., 2021, 17, 6993–7009 CrossRef CAS PubMed.
- W. Gao and C. W. Coley, J. Chem. Inf. Model., 2020, 60, 5714–5723 Search PubMed.
- P. Ertl and A. Schuffenhauer, J. Cheminf., 2009, 1, 8 Search PubMed.
- C. W. Coley, W. Jin, L. Rogers, T. F. Jamison, T. S. Jaakkola, W. H. Green, R. Barzilay and K. F. Jensen, Chem. Sci., 2019, 10, 370–377 RSC.
- C. W. Coley, W. H. Green and K. F. Jensen, J. Chem. Inf. Model., 2019, 59, 2529–2537 CrossRef CAS PubMed.
- S. Genheden, A. Thakkar, V. Chadimová, J.-L. Reymond, O. Engkvist and E. Bjerrum, J. Cheminf., 2020, 12, 70 Search PubMed.
- J. Degen, C. Wegscheid-Gerlach, A. Zaliani and M. Rarey, ChemMedChem, 2008, 3, 1503 Search PubMed.
- X. Q. Lewell, D. B. Judd, S. P. Watson and M. M. Hann, J. Chem. Inf. Comput. Sci., 1998, 38, 511–522 CrossRef CAS PubMed.
- C. M. Hansen, Hansen Solubility Parameters: A User's Handbook, CRC press, 2007 Search PubMed.
- G. Bao, R. Y. Abe and Y. Akutsu, J. Therm. Anal. Calorim., 2021, 143, 3439–3445 CrossRef CAS.
- M. J. Uddin and J. Fan, Polymers, 2024, 16, 1049 CrossRef CAS PubMed.
- R. Schiano Lo Moriello, D. Ruggiero, L. Angrisani, E. Caputo, F. de Pandi and G. de Alteriis, Electronics, 2021, 10, 939 Search PubMed.
- M. Deutel, G. Kontes, C. Mutschler and J. Teich, ACM Transactions on Evolutionary Learning, 2025, 5(3), 17 Search PubMed.
- P. Xu, Y. Ma, W. Lu, M. Li, W. Zhao and Z. Dai, J. Mater. Inf., 2025, 5, N–A Search PubMed.
- V. Sze, Y.-H. Chen, T.-J. Yang and J. S. Emer, Proc. IEEE, 2017, 105, 2295–2329 Search PubMed.
- M. Edwards and X. Xie, arXiv, 2016, preprint, arXiv:1609.08965, DOI:10.48550/arXiv.1609.08965.
- A. Aspuru-Guzik, T. Bechtel, V. Bernales, P. C. Biggin, F. Bigi, I. Borges, K. R. Briling, J. Cheung, C. M. Collins and K. K. Darmawan, et al. , Faraday Discuss., 2025, 256, 177–220 RSC.
- S. M. Lundberg and S.-I. Lee, Adv. Neural Inf. Process. Syst., 2017, 30, 1–10 Search PubMed.
- M. T. Ribeiro, S. Singh and C. Guestrin, Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 2016, pp. 1135–1144 Search PubMed.
- K. Simonyan, A. Vedaldi and A. Zisserman, arXiv, 2013, preprint, arXiv:1312.6034, DOI:10.48550/arXiv.1312.6034.
- B. Das and A. Mondal, ACS Appl. Energy Mater., 2024, 7, 9349–9363 CrossRef CAS.
- E. A. J. Abadi, H. Sahu, S. M. Javadpour and M. Goharimanesh, Mater. Today Energy, 2022, 25, 100969 CrossRef.
- E. Choi, M. T. Bahadori, L. Song, W. F. Stewart and J. Sun, Proceedings of the 23rd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 2017, pp. 787–795 Search PubMed.
- T. Wang and Q. Lin, J. Mach. Learn. Res., 2021, 22, 1–38 Search PubMed.
- C.-W. Ju, E. J. French, N. Geva, A. W. Kohn and Z. Lin, J. Phys. Chem. Lett., 2021, 12, 9516–9524 CrossRef CAS PubMed.
- M.-H. Lee, Adv. Energy Mater., 2019, 9, 1900891 CrossRef.
- C. Kunkel, C. Schober, H. Oberhofer and K. Reuter, J. Mol. Model., 2019, 25, 87 CrossRef PubMed.
- H. Sahu, F. Yang, X. Ye, J. Ma, W. Fang and H. Ma, J. Mater. Chem. A, 2019, 7, 17480–17488 RSC.
- L. v. d. Maaten and G. Hinton, J. Mach. Learn. Res., 2008, 9, 2579–2605 Search PubMed.
- E. J. Bjerrum and B. Sattarov, Biomolecules, 2018, 8, 131 CrossRef PubMed.
- Z. Zhou, S. Veeramani, F. Munguia-Galeano, H. Fakhruldeen and A. I. Cooper, Commun. Chem., 2025, 8, 384 CrossRef PubMed.
- G. Tom, S. P. Schmid, S. G. Baird, Y. Cao, K. Darvish, H. Hao, S. Lo, S. Pablo-García, E. M. Rajaonson and M. Skreta, et al. , Chem. Rev., 2024, 124, 9633–9732 CrossRef CAS PubMed.
- B. P. MacLeod, F. G. Parlane, T. D. Morrissey, F. Häse, L. M. Roch, K. E. Dettelbach, R. Moreira, L. P. Yunker, M. B. Rooney and J. R. Deeth, et al. , Sci. Adv., 2020, 6, eaaz8867 CrossRef CAS PubMed.
- T. C. Wu, A. Aguilar-Granda, K. Hotta, S. A. Yazdani, R. Pollice, J. Vestfrid, H. Hao, C. Lavigne, M. Seifrid and N. Angello, et al. , Adv. Mater., 2023, 35, 2207070 CrossRef CAS PubMed.
- L. M. Roch, F. Häse, C. Kreisbeck, T. Tamayo-Mendoza, L. P. Yunker, J. E. Hein and A. Aspuru-Guzik, Sci. Robot., 2018, 3, eaat5559 CrossRef PubMed.
- F. Hase, L. M. Roch, C. Kreisbeck and A. Aspuru-Guzik, ACS Cent. Sci., 2018, 4, 1134–1145 CrossRef CAS PubMed.
- A. M. Mroz, P. N. Toka, E. A. del Río Chanona and K. E. Jelfs, Faraday Discuss., 2025, 256, 221–234 RSC.
- M. Seifrid, R. Pollice, A. Aguilar-Granda, Z. Morgan Chan, K. Hotta, C. T. Ser, J. Vestfrid, T. C. Wu and A. Aspuru-Guzik, Acc. Chem. Res., 2022, 55, 2454–2466 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |