 Open Access Article
Open Access ArticleMapping sleep-promoting volatiles in aromatic plants with machine learning: a comprehensive survey of 2300 molecules
Peiqin
Shi†
ad,
Xing
Huang†
a,
Qinfei
Ke
a,
Xingran
Kou
*a and
Dachuan
Zhang
 *bc
*bc
aCollaborative Innovation Center of Fragrance Flavour and Cosmetics, Faculty of Flavour Fragrance and Cosmetics, Shanghai Institute of Technology, Shanghai 201418, China. E-mail: kouxr@sit.edu.cn
bDepartment of Food Science and Technology, Faculty of Science, National University of Singapore, 117542, Singapore. E-mail: dachuan.zhang@nus.edu.sg
cNational University of Singapore (Suzhou) Research Institute, 377 Lin Quan Street, Suzhou Industrial Park, Jiangsu 215123, China
dSchool of Food Science and Technology, Jiangnan University, Wuxi, Jiangsu 214122, China
First published on 10th February 2026
Abstract
Sleep disturbances affect up to one-third of the global population, yet current pharmacological therapies based on insomnia medications carry notable risks and side effects. Aromatic plants have long been valued for their capacity to ease stress and promote sleep; however, bioactive volatiles driving these benefits remain poorly understood. This study presents a comprehensive survey of 2391 volatiles across 991 aromatic plants, integrated with an ensemble machine-learning approach to identify their potential sleep-promoting activity. To evaluate the predictive accuracy of our approach, five candidate volatiles were computationally prioritized for in vivo testing. Four of these (an 80% success rate) robustly induced sleep-promoting effects, as evidenced by electroencephalogram analysis and modulation of γ-aminobutyric acid (GABA) receptor expression. In parallel, this work identified plant families such as Asteraceae, Lamiaceae, and Lauraceae as particularly enriched in high-potential volatiles and highlighted individual species—including Lavandula angustifolia and Perilla frutescens—as promising candidates for further pharmacological investigation. By combining large-scale data mining, computational prediction, and in vivo experimentation, this work first provides a comprehensive understanding of the landscape of sleep-promoting volatiles and aromatic plants and offers a reusable workflow to accelerate the discovery of bioactive compounds with potential applications in medicine, functional foods, and natural therapeutics.
Introduction
Sleep disorders affect nearly one-third of the global population, posing significant health risks such as cognitive impairment, cardiovascular diseases, and metabolic disorders.1,2 Current pharmacological treatments, including benzodiazepines and non-benzodiazepine sedatives, provide symptomatic relief but are often associated with adverse effects such as dependency and cognitive decline.3 These limitations have motivated the search for alternative strategies to modulate sleep, including approaches based on natural products.4 Among these, aroma plants, rich in bioactive compounds, have been traditionally used for their sedative properties.5,6 For instance, ajowan and bay leaves exhibit central nervous system inhibition properties, prolonging pentobarbital-induced sleep durations.7 Similarly, Hibiscus syriacus Linnaeus has been traditionally utilized to treat sleep disorders by enhancing rapid eye movement (REM) sleep and upregulating γ-aminobutyric acid type A (GABA_A) receptor mRNA levels.8 While their empirical use is well-documented, the specific chemical constituents responsible for sleep-promoting effects remain poorly understood.9Aromatic plants contain a wide array of bioactive compounds, many of which have potential neuropharmacological effects.10 For instance, compounds such as 3,5-dimethoxytoluene exhibit central nervous system modulation, influencing neurotransmitter pathways.11–13 Traditional methods for identifying bioactive molecules rely on bioassay-guided fractionation and in vivo studies, which are labor-intensive, time-consuming, and restricted by the availability of chemical standards. Recent advances in machine learning (ML) have enabled the prediction of molecular bioactivity using computational models, facilitating high-throughput screening of natural products.14–16 For instance, Erlina et al. and Wang et al. utilized ML to identify phytochemicals with inhibitory activity against SARS-COV-2 from a wide range of plant-derived chemicals.17,18 Likewise, Srisongkram et al. applied ML to screen for potential α-amylase and α-glucosidase inhibitors in indigenous Thai plants.19 Brown et al. successfully employed ML strategies to discover anti-inflammatory compounds from hops.20 Additionally, Zhang et al. employed ML to predict the biological activity of genes and enzymes for food and agricultural applications.21,22 However, existing studies predominantly focus on non-volatile compounds, leaving a substantial gap in the systematic identification of bioactive molecules that exert their effects through olfactory stimulation.16,22–29 The structure–activity relationships governing the sleep-promoting effects of volatile organic compounds (VOCs) remain poorly understood. Furthermore, unlike non-volatile bioactives, which interact with the body through digestion and systemic circulation, VOCs primarily act via olfactory receptors and the olfactory bulb, directly influencing brain activity and neurotransmitter release. Despite increasing evidence supporting the neuroactive properties of VOCs, the molecular determinants of their activity, such as functional groups, element compositions, and lipophilicity, are still not well characterized. Moreover, previous studies have largely concentrated on a narrow subset of molecules, neglecting to explore the bioactivity of structurally complex compounds present in plant extracts and essential oils.30
Despite growing interest in alternative sleep aids, identifying sleep-promoting VOCs in aromatic plants remains challenging due to the chemical complexity of plant extracts and the low efficiency of conventional bioassay-guided fractionation and in vivo methods. To address this limitation, this work presents a data-driven approach combining ML with big data on aromatic plant composition (Fig. 1). It systematically reveals the sleep-promoting potential of over 2300 VOCs and approximately 1000 types of aromatic plants. In addition to evaluating individual compounds, we further integrated predicted sleep-promoting potential scores of VOCs with occurrence and abundance data across plant species to prioritize botanical sources with the highest sleep-promoting potential. This prioritization helps to identify promising leads for further pharmacological studies and supports the selection of candidate plants for the development of natural sleep aids. The implications of this research extend beyond sleep disorders, contributing to the broader fields of natural product-based drug discovery and the development of functional perfumes, cosmetics, and foods.
Results
Collection and analysis of plant-derived VOCs
We manually reviewed over 970 relevant publications (spanning from 1965 to 2022) to curate plant-derived VOCs. This thorough screening process yielded a curated library of 2391 VOCs from 991 unique plant species across 136 families. For each VOC, we recorded comprehensive metadata, including its plant source (species, family, or cultivar), reported content levels, the specific plant parts or tissues from which it was extracted (e.g., leaves, flowers, or seeds), analytical methods (e.g., GC-MS or LC-MS), and validated molecular structures (see Supporting Dataset 1). These VOCs served as a basis for characterizing the diversity and chemical profile of sleep-promoting aromatic plants.To examine their sleep-promoting activity, we compiled a dataset for ML modeling consisting of 244 sleep-promoting compounds (positive samples, e.g., GABA agonists) and 244 non-sleep-promoting compounds (negative samples, e.g., GABA inhibitors) from the literature, public databases, and patents (Fig. 2a and b; see Supporting Dataset 2). A molecular scaffold analysis was conducted to explore the chemical diversity within the dataset. The positive compounds exhibited 78 distinct molecular scaffolds (Fig. 2c), while the negative compounds displayed 108 unique scaffolds (Fig. 2d). Positive compounds tend to share common structural characteristics, likely because they are designed for specific targets, such as GABA_A, or optimized for improved pharmacophore features, resulting in a higher degree of structural similarity. Further analysis using t-distributed stochastic neighbor embedding mapping showed that positive and negative samples are separated into distinct groups in most cases, indicating distinct structural characteristics. A similar trend is observed in their physicochemical properties: negative samples generally have higher molecular weight, more rotatable bonds, a larger topological polar surface area, more rings, and more hydrogen bond acceptors, whereas positive samples tend to have a lower number of hydrogen bond donors (Fig. S1). These findings collectively highlight distinct structural characteristics differentiating sleep-promoting compounds from other compounds, underscoring the feasibility of developing a classification model capable of effectively identifying novel sleep-promoting VOCs (Fig. 2e).
Proposed approach surpasses individual machine learning models
We developed 19 classification models by combining nine ML algorithms with different molecular representations. The area under the receiver operating characteristic curve (AUC-ROC), accuracy, precision, and recall scores for all models are summarized in Table S1. Most models demonstrated good generalization capability, with the test set yielding approximately equal prediction outcomes compared to the validation set. Deep learning models, namely message-passing neural networks31 and Attentive FP,32 showed lower performance on the independent test sets compared to ML models, with AUC-ROC values of 0.876 ± 0.025 and 0.893 ± 0.033, respectively. This relatively lower performance is likely due to the limited size of the training dataset, which may be insufficient for training a fully optimized deep-learning model. In contrast, in this task, classical ML models such as the random forest (RF) model, gradient boosting decision trees (GBDT), support vector machines (SVM), and K-nearest neighbor (KNN), when combined with molecular fingerprints and descriptors, outperformed deep learning models.Among all the models, the RF model built on RDKit descriptors (RF-RDKit) exhibited strong predictive power, achieving an AUC-ROC of 0.957 ± 0.02 on the independent test sets. To further improve predictive accuracy, we developed a stacking ensemble model by combining the four best-performing classifiers: RF model built on Molecular ACCess System keys (RF-MACCS), RF-RDKit, XGBoost-MACCS, and SVM-MACCS (Fig. S2). The stacking model significantly surpassed the individual models, achieving an AUC-ROC of 0.994 ± 0.08, an accuracy of 0.961 ± 0.024, a precision of 0.957 ± 0.033, and a recall of 0.967 ± 0.024 (Table S1). Its false positive and false negative rates are 4.4% and 3.2%, respectively (Fig. 3a). These results highlight the effectiveness of model stacking, which leverages the strengths of multiple classifiers to improve overall predictive performance.
Another strategy to mitigate the challenge of limited training data is few-shot learning with pre-trained models. To compare this strategy with the model stacking approach, we tested the performance of two pre-trained models, namely CHEM-BERT33 and knowledge graph-enhanced molecular contrastive learning with functional prompts (KANO)34 fine-tuned on the dataset. CHEM-BERT leverages self-supervised learning on large-scale molecular datasets to capture chemical semantics. KANO incorporates domain-specific molecular graphs and functional group information to improve molecular representation learning. Both methods utilized pre-trained models and chemical knowledge to enhance the model's predictive ability on small datasets. However, we found that both models demonstrated lower performance than the stacking model in this task, with AUC-ROC values of 0.482 ± 0.045 and 0.901 ± 0.037, respectively. The lower performance of CHEM-BERT might be attributed to the substantial distribution shift between its pre-training datasets and our specific dataset. Although it has captured extensive chemical semantics on large molecular datasets, its representations appear to lack adequate specificity to accurately distinguish bioactive sleep-promoting volatiles, resulting in near-random predictive performance. Although KANO, which incorporates domain-specific molecular graphs and functional prompts, achieved better results than CHEM-BERT, it still underperformed when compared to the simple stacking ensemble approach. One possible reason for this limited performance is that, despite leveraging domain knowledge, the representational power of graph-based pre-training may still fail to adequately capture subtle pharmacological characteristics critical for predicting biological activities in highly specialized datasets. Additionally, fine-tuning on limited-instance data could lead these relatively complex pre-trained models to overfit due to the large number of parameters being adjusted, resulting in instability and reduced generalizability on our dataset. Collectively, these results underscore that, in this scenario of limited bioactive data, the simple and straightforward ensemble stacking strategy can be a better solution to address the data limitation compared to the pre-train/fine-tune strategy.
C–N bonds and five-membered rings strongly impact sleep-promoting activity
Shapley Additive ExPlanations (SHAP) analysis was employed to assess the influence of individual molecular features on the model's predictions, capturing both positive and negative contributions.35Fig. 3b–e and S3 show the key structural elements that significantly affect model performance. For the RF-MACCS, XGBoost-MACCS, and SVM-MACCS models, the most important descriptors were the C–N bond (a carbon atom directly bonded to a nitrogen atom) and the 5-membered ring (the presence of a five-membered ring).When a molecule contains at least one C–N bond or features a five-membered ring, the corresponding SHAP values are negative (highlighted in red), indicating a negative impact on the model's prediction of GABA activity (Fig. 3b, d, and e). Conversely, when these features are absent, the SHAP values are positive (highlighted in blue), suggesting a positive effect on the model's classification. In the RF-RDKit model, the most influential descriptor was SlogP_VSA8, which indicates the logarithm of the hydrophobic contribution surface area, ranging from 12.0 to 13.5 Å2. A higher SlogP_VSA8 value correlated with a decreased probability of sleep-promoting activity (Fig. 3c).
For the decision tree-based models (e.g., RF-MACCS and XGBoost-MACCS), we analyzed SHAP interaction values to explore how feature interactions influence model predictions (Fig. S4). Strong interaction effects were observed involving the C–N bond and the absence of a five-membered ring, indicating that both their individual contributions and interactions with other features play a significant role in the model's output. In addition, the interaction between the N–H group and aromatic rings > 1 consistently showed a negative effect for certain samples, suggesting that the co-occurrence of N–H groups and multiple aromatic rings reduces the sleep-promoting activity of VOCs.
These insights enhance interpretability by illuminating the “black box” of ML models, providing valuable guidance for identifying new sleep-promoting molecules and understanding their mechanisms.
Computationally predicting and experimentally validating sleep-promoting VOCs
Among 2391 VOCs from aromatic plants, 2373 (99.25%) were identified within the model's applicability domain using the Euclidean distance-based method,24 indicating strong applicability. These compounds were screened using the stacking model to predict their potential sleep-promoting activity. Finally, based on a high predicted sleep-promoting probability score from our ensemble ML model, we randomly selected five VOCs that are commercially available. These VOCs included linalool, carvacrol, safranal, vanillin, and methyl eugenol (Table S2).During sleep, electroencephalogram (EEG) signals can be classified into distinct stages, including non-rapid eye movement (NREM) sleep, REM sleep, and wakefulness (Fig. 4a). NREM sleep is marked by higher EEG amplitude, indicating synchronized brain activity, and lower electromyography (EMG) amplitude, reflecting muscle relaxation. In contrast, REM sleep exhibits lower EEG amplitude, accompanied by continued low EMG activity, indicating a state of reduced brain activity and muscle atonia. Wakefulness, however, is associated with low EEG amplitude and increased EMG activity, signifying an alert and active state of both the brain and muscles. According to EEG analysis, mice exposed to carvacrol, safranal, vanillin, and methyl eugenol exhibited a significant reduction in wakefulness duration compared to the normal control group (NOR) (Fig. 4b; p < 0.01, p < 0.05, p < 0.01, and p < 0.05, respectively). Additionally, total sleep duration was significantly extended in these groups (Fig. 4e; p < 0.01, p < 0.05, p < 0.01, and p < 0.05, respectively). The observed increase in total sleep duration was primarily driven by an increase in NREM sleep (Fig. 4c; p < 0.01, p < 0.01, p < 0.05, and p < 0.05, respectively), rather than REM sleep. These findings suggest that carvacrol, safranal, vanillin, and methyl eugenol effectively prolong NREM sleep duration and enhance overall sleep time in mice.
To investigate the sleep-promoting mechanisms of aroma compounds, we analyzed their effects on GABA_A receptor protein expression using Western blot analysis. The results indicated that carvacrol, safranal, vanillin, and methyl eugenol increased the expression of one or more GABA_A subunits (Fig. 5). Compared to the NOR group, the expression of GABA_Aα1 was significantly elevated in the vanillin and methyl eugenol groups, increasing by 1.55-fold and 1.63-fold, respectively (Fig. 5a; p < 0.05 and p < 0.05, respectively). Similarly, GABA_Aβ2 expression was significantly upregulated in the carvacrol, vanillin, and methyl eugenol groups, with increases of 1.91-fold, 1.93-fold, and 1.75-fold, respectively (Fig. 5b; p < 0.05, p < 0.05, and p < 0.05, respectively). Additionally, methyl eugenol significantly increased the expression of GABA_Aγ2 by 1.69-fold (Fig. 5c; p < 0.05). These findings suggest that these VOCs enhance GABAergic signaling by upregulating GABA_A receptor subunits, further supporting their potential role in promoting sleep.
We applied SHAP analysis to four validated sleep-promoting VOCs—carvacrol, safranal, vanillin, and methyl eugenol—using our trained models. Across all compounds, the models consistently highlighted structural features that contribute positively to the predicted sleep-promoting activity. These include the presence of no more than one aromatic ring, absence of heterocycles, lack of C–N bonds, absence of five-membered rings, and no nitrogen atom bonded to two aromatic atoms. Additionally, all compounds exhibited SlogP_VSA8 = 0, indicating no atomic hydrophobic contributions within the 12.0–13.5 Å2 surface area range. These findings reinforce our previous SHAP-based conclusions and demonstrate that the machine-learning models have captured meaningful and relevant patterns linked to sleep-promoting properties (Fig. S5–S8).
Prioritizing aromatic plants with high sleep-promoting potential
To evaluate the sleep-promoting potential of aromatic plants, we combined a predictive scoring approach with comprehensive data on the content of VOCs for each species. This integrated strategy allowed us to identify potentially sleep-promoting VOCs and gauge their abundance and diversity in different plant families.Through this process, 991 unique species with recorded aromatic properties were identified, and they were classified into three major plant classes: Magnoliopsida (dicotyledons), Pinopsida (conifers), and Equisetopsida (ferns). Among these, the Magnoliopsida class emerged as the most abundant and extensively studied, comprising 128 families and 914 species. In contrast, Pinopsida included seven families and 76 species, and Equisetopsida only comprised one family and one species (see Supporting Dataset 3).
Fig. 6 illustrates the number of VOCs with high sleep-promoting potential (prediction scores exceeding 0.95) detected in plant extracts from various families, while Fig. S9 displays the combined score of the number of sleep-promoting VOCs and their content. Within Magnoliopsida, three families stood out for their high diversity of sleep-promoting VOCs: Asteraceae, Lamiaceae, and Lauraceae (Fig. 6 and S10). Aromatic plants from Asteraceae, Lamiaceae, and Lauraceae are well-documented for their calming, sedative, anxiolytic, and central nervous system-suppressing properties in traditional medicine, which align with our data-driven findings.36–40 When examining specific species, we identified several candidates that not only contain a broad spectrum of putative sleep-promoting VOCs but also exhibit relatively high concentrations of these compounds. For example, Silybum marianum, Sphagneticola trilobata, and Petasites japonicus from the Asteraceae family, Lavandula angustifolia, Ocimum basilicum, Perilla frutescens, and Vitex negundo from the Lamiaceae family, and Litsea monopetala and Litsea cubeba from the Lauraceae family (Fig. S11). These findings suggest that these particular species may be valuable for further research and potential therapeutic applications related to sleep disorders. The synergy of multiple VOCs, including terpenes and phenolic compounds, may be contributing to the overall sleep-promoting effect—an area that warrants detailed pharmacological and mechanistic studies.
Beyond Magnoliopsida, notable sleep-promoting potential was also observed in Pinopsida (Fig. 6, S12, and S13). Although the families in the Pinopsida class exhibited relatively lower scores overall, two of them, particularly Pinus sylvestris (Pinaceae) and Taxodium distichum (Cupressaceae), demonstrated considerable potential. This suggests that Pinopsida plants may serve as an alternative direction and provide promising leads for further biochemical investigations. In particular, Cupressaceae and Pinaceae demonstrated elevated levels of terpenes (e.g., cedrol and alpha-fenchone), which are linked to sedative or sleep-regulating pathways. A summary of high-potential aromatic plants is presented in Fig. S14 and S15. In addition, among the lower-scoring families, some species exhibit high bioactive potential. For instance, Aconitum carmichaelii in the Ranunculaceae family, Artocarpus heterophyllus in the Moraceae family, and Linum usitatissimum in the Linaceae family (see SI) showcase concentrations of potent VOCs that exceed expectations based on a family-wide assessment.
Experimental
VOC data collection and processing
Using the keywords “aroma plants,” “essential oil,” and “volatile compounds,” we systematically searched the Google Scholar and Web of Science databases for publications focused on plant-derived VOCs (accessed October 2023). Each report was manually curated to capture detailed information on VOCs, including their plant sources, concentration levels, plant extraction sites, analytical methodologies, and molecular structures.To ensure data consistency and reliability, we performed rigorous data cleaning steps: (1) we restricted the dataset to include only compounds that fell within the model's applicability domain, ensuring valid concentration values and clearly defined plant sources. (2) To prevent inflated estimations of compound frequency and concentration due to multiple extracts from different plant parts (e.g., roots and leaves), we retained only unique instances of compounds, removing duplicate entries from various plant tissues. For the same compound in different parts, we only retain the concentration values of each compound when it first appeared in different plants. (3) Given the variation in compound naming conventions across different studies, we standardized all compound names according to the PubChem41 database to facilitate accurate comparisons. (4) Compounds for which the literature described presence qualitatively (e.g., “detected,” “major constituent,” or “trace amount”) without numerical values were excluded from these analyses.
Training data collection and visualization
To develop a predictive model, we compiled a dataset that includes both sleep-promoting compounds (positive samples) and non-sleep-promoting compounds (negative samples) (Fig. 2a). The dataset was curated from various sources, including the literature, patents, and public chemical databases such as ChEMBL,42 PubChem,41 and DrugBank43 (accessed October 2023). Specifically, compounds verified to have known sleep-promoting activities or effects on GABA_A receptor activation were selected as positive samples.44 Additionally, non-sleep-promoting compounds, such as GABA_A receptor inhibitors, were compiled from the aforementioned sources as negative samples (Fig. 2b). To ensure data quality and reliability, all molecules underwent rigorous preprocessing based on the following criteria: (1) molecules without available structural representations were excluded. (2) Molecules with a molecular weight below 30 Da or above 1000 Da were removed to avoid extremely small or large compounds that may be pharmacologically irrelevant. (3) Redundant entries and conflicting records were eliminated based on compound names and their simplified molecular input line entry system representations.To visualize the diversity of the dataset, we used MoleculeCloud,45 which generates molecular scaffolds by representing the core molecular structure with all non-carbon atoms replaced by carbon, thereby showcasing the structural variability within the dataset. Additionally, we applied t-distributed stochastic neighbor embedding to map the chemical space of the dataset and evaluate the clustering patterns between positive and negative samples using three types of molecular descriptors, including Extended Connectivity Fingerprints with a bond diameter of 4 (ECFP4), MACCS, and RDKit fingerprints.
Machine learning model development, evaluation, and application
Four different molecular representations were used to construct the prediction models, each capturing distinct structural and chemical properties, including molecular graphs, ECFP4, MACCS, and RDKit fingerprints. Molecular graphs encode the topological structure of molecules, representing atoms as nodes and chemical bonds as edges, thereby preserving complex connectivity patterns. The ECFP4 fingerprint encodes atomic neighborhoods based on two bond lengths, offering a substructure-based molecular representation.46 The MACCS fingerprint consists of 166 predefined structural fragments that encode specific substructural patterns relevant to molecular activity prediction.47 The RDKit fingerprint is a 208-bit descriptor that captures various physicochemical, topological, and connectivity-based properties of chemical compounds. The molecular representation was generated using the RDKit package (version 2023.3.1, https://www.rdkit.org/) and DeepChem (version 2.7.1, https://deepchem.io/). Before training, features with a correlation greater than 0.98 were removed to reduce redundancy and enhance model generalizability.We developed multiple models using conventional ML algorithms, including RF, KNN, SVM, XGBoost, and GBDT,48 as well as deep learning frameworks, including message-passing neural networks,31 Attentive FP,32 CHEM-BERT,33 and KANO.34 These models were trained using various molecular graph representations mentioned earlier. The conventional ML algorithms were implemented with scikit-learn (version 1.0.2) and XGBoost (version 1.7.6) in Python (version 3.8.16).
To optimize the ML models' performance, hyperparameter tuning was conducted using a grid search approach combined with five-fold cross-validation. The hyperparameters used are presented in the SI. To further enhance predictive accuracy, we employed a stacking ensemble strategy49 combining multiple ML models to capture different aspects of the data. Specifically, we selected the four best-performing individual models and integrated them into a stacking framework to leverage their complementary strengths (Fig. S2).
The models were evaluated using AUC-ROC, accuracy, precision, and recall (eqn (1)–(3)).
 | (1) |
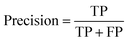 | (2) |
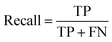 | (3) |
To ensure reliable predictions, we employed a previously reported method that quantifies the model's applicability domain using Euclidean distance and KNN.24 First, the threshold T for determining whether a compound falls within the applicability domain was calculated from the training set using the following equation:
| T = Zσ + Y | (4) |
We employed the SHAP algorithm to interpret the ML model's predictions. SHAP calculates feature importance by assigning Shapley values derived from cooperative game theory to quantify the individual contribution of each feature to the model's predictions.
Validation assays: materials and experimental animals
Linalool (Adamas, 83484A), carvacrol (MERYER, M17729-25G), safranal (Adamas, 15800B), methyl eugenol (Yuanye, S25645), vanillin (Beida Zhengyuan, SF020), fragrance lamp (MUJI, 4549337287815), Zoletil 50 (Virbac, 06516 CARROS), Veet hair removal cream (Reckitt Benckiser China Co., Ltd, G20161336), methyl alcohol (Thermo Fisher, A452-4), acetonitrile (Thermo Fisher, 036423.AP), GABA (Solarbio, SA8240), RIPA buffer (Solarbio, R0020), BCA protein assay kit (Solarbio, PC0020), sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) color preparation kit (Sangon Biotech, C671103), SDS-PAGE sample loading buffer (Biosharp, BL502A), M5 prestained protein ladder (Mei5bio, MF2190-plus-01), Goat anti-Rabbit IgG (H + L) HRP (Affinity, S0001), Goat Anti-Mouse IgG (H + L) HRP (Affinity, S0002), PVDF transfer membrane (Millipore, IPVH00010), enhanced chemiluminescence (ECL) western blotting substrate (Solarbio, PE0010), ice-free bath rapid film transfer solution (Servicebio, G2028-1 L), SWE fast high-resolution electrophoresis buffer (Servicebio, G2081-1 L), Tris buffered saline (Servicebio, G0001-2 L), protein-free rapid blocking buffer (Servicebio, G2052-500 ML), GABRA1 polyclonal antibody (Proteintech, 12410-1-AP), GABRG2 polyclonal antibody (Proteintech, 14104-1-AP), and anti-GABRB2 rabbit polyclonal antibody (HUABIO, HA500289).Male C57BL/6 mice (20–25 g, aged 4–8 weeks) were sourced from Shanghai SLAC Laboratory Animal Co., Ltd. The animals were acclimated for a minimum of one week under controlled conditions: temperature (20 °C ± 3 °C), humidity (50% ± 5%), and a 12-hour light/dark cycle, with ad libitum access to food and water. All animal experiments received approval from the Experimental Animal Ethics Committee of Wuchuang Biotechnology (Shanghai) Co., Ltd (approval no. WTPZ20230812001, Shanghai, China). All procedures were conducted in accordance with the ARRIVE 2.0 guidelines for reporting animal research.
Preparation for the EEG experiment
Based on model predictions and the availability of compound standards, five VOCs with potential sleep-promoting activity were selected for experimental validation. Mice were anesthetized with Zoletil® 50 (30 mg kg−1) before surgery. For EEG recording, electrodes were surgically implanted in the skull, with two EEG electrodes positioned above the hippocampus and ground and reference electrodes placed above the cerebellum. For EMG recording, two electrodes were sutured into the dorsal nuchal muscles. All electrode wires were routed to the center of the skull and secured with dental cement. After a one-week recovery period, the mice were randomly assigned to six groups (n = 6 per group). To ensure consistent exposure to the fragrance environment, a customized glass chamber was designed to accommodate three mice at a time, with each mouse housed in a separate cage (30 × 20 × 13.5 cm). The chamber featured ventilation holes and electrode access points on the top. A perforated plate was placed inside the chamber, beneath which a fragrance lamp released VOCs. A squirrel cage fan was situated above the plate to ensure even diffusion of the VOCs.The mice were divided into experimental and control groups based on whether they were exposed to the aroma or not. In the experimental group, five different VOCs were tested, with six mice assigned to each compound. The sleep conditions of the mice were monitored to evaluate the effects of each VOC. In the control group, six mice were used to assess sleep conditions without the presence of aromatic plant VOCs. For aroma exposure, 600 grams of pure water were added to the fragrance lamp, followed by 0.20 to 0.25 grams of the VOCs. The mice were exposed to the aroma from 8:00 a.m. to 4:00 p.m. to investigate its effect on sleep.
EEG/EMG acquisition and analysis
EEG and EMG data were recorded using the NeuroKey-16 device (Nanjing Greathink Medical Technology Co., Ltd) to extract raw signals. The signals underwent EEG-specific filtering with a bandpass filter operating in a frequency range of 0–30 Hz. Sleep analysis was conducted using Simplescore-v1.5 software (Nanjing Greathink Medical Technology Co., Ltd) to examine both EEG and EMG signals. NREM sleep was characterized by large, slow brain waves with delta activity below 4 Hz in the EEG. During the transition from NREM to REM sleep, a shift from low-frequency delta activity to a rapid, low-voltage EEG within the theta range (6–10 Hz) was observed. Wakefulness was identified by low- to moderate-voltage brain waves in the EEG, accompanied by high EMG activity, distinguishing it from sleep states. The Simplescore software was used with default parameters only. The operation manual, a detailed description of the default analysis workflow, and the immediate outputs generated by this workflow are publicly available in the Zenodo archive.Western blot analysis for receptor protein expression in mice
After 8 hours of aroma exposure, the mice were sacrificed, and their brains were extracted for protein analysis. Brain protein extracts were prepared using RIPA buffer, and the protein concentration was determined using the Bradford colorimetric method. The samples were analyzed via polyacrylamide gel electrophoresis and then transferred onto PVDF membranes. The membranes were blocked with a protein-free rapid-blocking buffer for 15 minutes at room temperature. For protein detection, anti-GABA_Aβ2, anti-GABA_Aα1, and anti-GABA_Aγ2 were used as primary antibodies, and the membranes were incubated with the antibodies at 4 °C overnight. After three washes with TBS-Tween buffer, secondary antibodies were added and incubated at room temperature for 2 hours. The membranes were then rinsed with TBS-Tween buffer, and protein signals were detected using the ECL luminescent solution.Prioritizing aromatic plants with high sleep-promoting potential
To identify aromatic plant species enriched in sleep-promoting volatiles, we integrated compound-level predictive scores with plant-level occurrence and abundance data. This analysis aimed to prioritize plant taxa based on both the number and relative concentration of volatiles predicted to exhibit sleep-promoting bioactivity. Each VOC was assigned a sleep-promoting probability score based on predictions from the ensemble ML model described above. VOCs with a model prediction probability exceeding 0.95 were considered high-confidence candidates for sleep-promoting activity. To assess the sleep-promoting potential of each plant species, we implemented two complementary scoring approaches: (1) diversity score: the number of high-confidence sleep-promoting VOCs (prediction score > 0.95) detected in each species. (2) Content-weighted score: a weighted sum of prediction scores multiplied by the relative abundance (normalized content) of each VOC in the plant, when such data were available. Both metrics were computed for all 991 aromatic plant species in the dataset. Species were then ranked within their respective families and across all plant taxa. Using the scientific names of aromatic plants, we employed Python to retrieve their complete taxonomic classifications (including kingdom, phylum, class, order, family, genus, and species) from “iPlant” (https://www.iplant.cn/). Then, with pycirclize (https://moshi4.github.io/pyCirclize/phylogenetic_tree/), we built circular phylogenetic trees to visualize the diversity and distribution of sleep-promoting VOCs across major plant groups. The trees were annotated with heatmaps indicating the number of high-confidence sleep-promoting VOCs per plant family.Conclusions
This study introduces an efficient approach to address the longstanding gap in our understanding of which specific bioactive VOCs in aromatic plants contribute to their reputed sleep-promoting effects. By adopting an ensemble ML approach, we implemented a comprehensive survey of the sleep-promoting activity of 2391 volatiles across 991 aromatic plants and found over 1000 of these have strong potential (predicted score > 0.95). Another noteworthy aspect of our study is the use of a SHAP-based interpretability framework to highlight key molecular substructures that informed the ML model's predictions. This transparency enhances confidence in the computational results and creates opportunities for rational molecular design. Understanding how specific functional groups affect sleep-related bioactivity can speed up the targeted development of novel therapeutic agents derived from plant VOCs.Additionally, we validated our approach with EEG measurements and GABA_A receptor expression analysis in mouse brain tissue and found that four out of five (an 80% success rate) predicted sleep-promoting VOCs—carvacrol, safranal, vanillin, and methyl eugenol—demonstrated the expected sleep-promoting activity. Our findings confirmed that the abovementioned VOCs significantly prolonged the duration of NREM sleep, indicating their potential to enhance sleep quality. The observation that these VOCs effectively modulate NREM sleep aligns with previous evidence highlighting the importance of NREM in physical restoration and energy replenishment. Detailed EEG data revealed that the increase in NREM was accompanied by heightened theta-wave activity, a hallmark of deeper, restorative sleep stages. These results support the hypothesis that specific VOCs can exert sedative or sleep-regulating effects by influencing particular pathways.
Interestingly, although linalool was predicted to have sleep-promoting properties, our experiments did not reveal significant increases in NREM sleep or total sleep time for this VOC, nor did it influence GABA_A receptor levels. These findings suggest that linalool may exert weaker or context-dependent effects on sleep modulation. First, the concentration of linalool used in the in vivo tests may not have been optimal; either too low to exert a measurable effect or too high, potentially leading to paradoxical excitation or adverse effects. Second, rapid metabolism or clearance of linalool in vivo may limit its bioavailability and central nervous system penetration, thereby attenuating its hypnotic effects. Third, species-specific differences in receptor binding or physiological response could also contribute, as computational predictions are often based on data from human or model organism targets that may not fully translate to the experimental model used. Indeed, linalool may have more subtle or context-specific modulatory effects on neural pathways, necessitating further research under various conditions to fully elucidate its role in sleep regulation.
In our comprehensive survey, we identified plant families with high potential for sleep-promoting activity, as well as individual species with strong bioactive potential even within lower-scoring families. This discrepancy highlights the chemotypic variability of plant secondary metabolite production: even families with minimal representation of sleep-promoting chemicals at the aggregate level can harbor “outliers” that produce specific compounds in higher concentrations or with greater synergy. Several factors may drive this phenomenon. First, plants often evolve specialized metabolic pathways in response to localized ecological pressures—such as herbivory, pathogen defense, or pollinator attraction—which can lead to the accumulation of distinct, potent VOCs in a handful of species. Consequently, while an entire family might be characterized by a moderate or minimal presence of sleep-promoting molecules, a few species could stand out due to their unique evolutionary trajectories. Second, these species may have been traditionally valued for medicinal properties unrelated to sleep (e.g., analgesia, cardioprotection, or antimicrobial activity), yet some of the same compounds responsible for these effects might also induce sedation or modulate sleep pathways. As research techniques advance and more sophisticated analytical tools become commonplace, it is increasingly clear that pharmacologically relevant bioactivity can stem from a broad range of overlapping or multifunctional plant metabolites.
Several aspects of this study could be expanded further. First, while our in vivo assays provide compelling evidence for the sleep-promoting effects of certain VOCs, long-term safety and efficacy studies are still required before these molecules can be considered for clinical use. Second, our focus on GABA_A receptor mechanisms does not rule out the possibility that other neurotransmitter systems (e.g., serotonin and melatonin) may also contribute to the observed sleep-promoting effects. Third, synergy among multiple VOCs within a single plant—and how these interactions might enhance or diminish sedative outcomes—is still poorly understood, highlighting the need for more sophisticated analytical and modeling approaches.52,53 Fourth, it is possible that yet-to-be-identified biochemical pathways54 in these species contribute to sleep-promoting activity. For example, some VOCs in lower-scoring families might be incompletely characterized, either because the plants are less commonly studied or because advanced analytical techniques (e.g., metabolomics, multi-omics approaches)55 have not been applied extensively to these taxa. Looking ahead, incorporating additional omics data (e.g., proteomics, metabolomics, and genomics) and conducting synergistic studies that evaluate multiple receptor pathways could further enhance our understanding of how these plant-derived VOCs influence sleep and other physiological processes. Finally, it is important to recognize that not all plant-derived essential oils or VOCs are universally safe. For instance, lavender oil (derived from Lavandula angustifolia) is usually regarded as safe for most adults when used appropriately for aromatherapy, topical application, or as a food flavoring. Nevertheless, concerns have been raised about its use in young boys before puberty due to potential hormonal disturbances.56 Such considerations highlight the need for safety assessments alongside efficacy studies in future research to ensure that sleep-promoting plant-derived VOCs can be applied both effectively and responsibly.
Nevertheless, we are confident that the current study provides an effective approach for estimating the sleep-promoting activity of plant-derived VOCs and prioritizes key plant families and species with high sleep-promoting potential for further investigation. Moreover, when high-quality training data are available, the general framework proposed in this study can be readily applied to prediction tasks other than sleep-promoting VOCs, such as the prediction of natural antioxidants, antidiabetic agents, or other health-promoting molecules.
Author contributions
D. Z., X. K., P. S., and X. H. designed the research. P. S. developed and evaluated the machine learning models. P. S. and X. H. implemented the experiments and organized the datasets. P. S., X. H., and D. Z. wrote the paper with input from X. K. and Q. K. All authors approved the final paper.Conflicts of interest
There are no conflicts to declare.Data availability
Data and code for model training and computational screening are available in a Zenodo repository (https://doi.org/10.5281/zenodo.18012782). Supporting Datasets 1–3 are also available in the same Zenodo repository (https://doi.org/10.5281/zenodo.18012782).Supplementary information (SI): detailed descriptions of machine learning models and hyperparameters, additional performance evaluations, extended figures and tables. See DOI: https://doi.org/10.1039/d5dd00173k.
Acknowledgements
This project was supported by the Collaborative Innovation Center of Fragrance Flavour and Cosmetics, Ministry of Education, Singapore, under the Academic Research Fund Tier 1 (A-8003718-00-00) and NUS IT (NUSREC-HPC-00001).Notes and references
- V. K. Chattu, M. D. Manzar, S. Kumary, D. Burman, D. W. Spence and S. R. Pandi-Perumal, Healthcare, 2019, 7, 1 CrossRef.
- J. M. Siegel, Lancet Neurol., 2022, 21, 937–946 Search PubMed.
- A. Reimers, P. Odin and H. Ljung, Drug Saf., 2025, 48, 339–361 CrossRef CAS.
- Z. Z. Hu, S. Oh, T. W. Ha, J. T. Hong and K. W. Oh, Biomol. Ther., 2018, 26, 343–349 CrossRef CAS.
- E. Christaki, E. Bonos, I. Giannenas and P. Florou-Paneri, Agriculture, 2012, 2, 228–243 CrossRef.
- M. Han, D. Zhang, S. Ding, Y. Tian, X. Cheng, L. Yuan, D. Sun, D. Liu, L. Gong, C. Jia, P. Cai, W. Tu, J. Chen and Q.-N. Hu, Bioinformatics, 2021, 37, 4275–4276 CrossRef CAS.
- T. Rahman, K. A. Rahman, S. Rajia, M. Alamgir, M. T. H. Khan and M. S. K. Choudhuri, Orient. Pharm. Exp. Med., 2010, 10, 86–89 CrossRef.
- Y. R. Kim, S. Y. Lee, S. M. Lee, I. Shim and M. Y. Lee, Biomed. Pharmacother., 2022, 146, 112301 CrossRef CAS.
- J. H. Liu, L. Ghastine, P. Um, E. Rovit and T. N. Wu, Environ. Res., 2021, 196, 110406 CrossRef CAS.
- X. Zeng, P. Zhang, Y. Wang, C. Qin, S. Chen, W. He, L. Tao, Y. Tan, D. Gao, B. Wang, Z. Chen, W. Chen, Y. Y. Jiang and Y. Z. Chen, Nucleic Acids Res., 2019, 47, D1118–D1127 CrossRef PubMed.
- M. Zotti, M. Colaianna, M. G. Morgese, P. Tucci, S. Schiavone, P. Avato and L. Trabace, Molecules, 2013, 18, 6161–6172 CrossRef CAS.
- B. K. Lee, A. N. Jung and Y. S. Jung, Biomol. Ther., 2018, 26, 368–373 Search PubMed.
- J. M. Tankam and M. Ito, Biol. Pharm. Bull., 2013, 36, 1608–1614 Search PubMed.
- D. Zhang, M. Liu, Z. Yu, H. Xu, S. Pfister, G. Menichetti, X. Kou, J. Zhu, D. Fan and P. Rao, Trends Food Sci. Technol., 2025, 164, 105272 Search PubMed.
- D. Zhang, D. Liu, J. Jing, B. Jia, Y. Tian, Y. Le, Y. Yu and Q.-N. Hu, Trends Food Sci. Technol., 2024, 148, 104513 CrossRef CAS.
- X. Kou, P. Shi, C. Gao, P. Ma, H. Xing, Q. Ke and D. Zhang, J. Agric. Food Chem., 2023, 71, 6789–6802 Search PubMed.
- L. Erlina, R. I. Paramita, W. A. Kusuma, F. Fadilah, A. Tedjo, I. P. Pratomo, N. S. Ramadhanti, A. K. Nasution, F. K. Surado, A. Fitriawan, K. A. Istiadi and A. Yanuar, BMC Complementary Med. Ther., 2022, 22, 207 CrossRef CAS.
- Z. Wang, T. Belecciu, J. Eaves, M. Reimers, M. H. Bachmann and D. Woldring, J. Biomol. Struct. Dyn., 2023, 41, 6643–6663 CrossRef CAS PubMed.
- T. Srisongkram, S. Waithong, T. Thitimetharoch and N. Weerapreeyakul, Nutrients, 2022, 14, 267 CrossRef CAS.
- K. S. Brown, P. Jamieson, W. Wu, A. Vaswani, A. Alcazar Magana, J. Choi, L. M. Mattio, P. H.-Y. Cheong, D. Nelson, P. N. Reardon, C. L. Miranda, C. S. Maier and J. F. Stevens, Antioxidants, 2022, 11, 1400 Search PubMed.
- Y. Tian, D. Zhang, H. Xing, M. Tang, C. Zhao, W. He, H. Lin, W. Yan, Q.-N. Hu and A. Wu, J. Adv. Res., 2025 DOI:10.1016/j.jare.2025.08.045.
- D. Zhang, H. Xing, D. Liu, M. Han, P. Cai, H. Lin, Y. Tian, Y. Guo, B. Sun, Y. Le, Y. Tian, A. Wu and Q.-N. Hu, ACS Catal., 2024, 14, 3336–3348 CrossRef CAS.
- M. Han, S. Liu, D. Zhang, R. Zhang, D. Liu, H. Xing, D. Sun, L. Gong, P. Cai, W. Tu, J. Chen and Q.-N. Hu, Molecules, 2022, 27(12), 3931 CrossRef CAS.
- D. Zhang, Z. Wang, C. Oberschelp, E. Bradford and S. Hellweg, ACS Sustain. Chem. Eng., 2024, 12, 2700–2708 CrossRef CAS PubMed.
- D. Zhao, Y. Zhang, Y. Chen, B. Li, W. Zhou and L. Wang, J. Chem. Inf. Model., 2024, 64, 9098–9110 CrossRef CAS.
- M. Njirjak, L. Žužić, M. Babić, P. Janković, E. Otović, D. Kalafatovic and G. Mauša, Nat. Mach. Intell., 2024, 6, 1487–1500 CrossRef.
- D. Zhang, C. Jia, D. Sun, C. Gao, D. Fu, P. Cai and Q.-N. Hu, J. Agric. Food Chem., 2023, 71, 8488–8496 Search PubMed.
- V. Smer-Barreto, A. Quintanilla, R. J. R. Elliott, J. C. Dawson, J. Sun, V. M. Campa, A. Lorente-Macias, A. Unciti-Broceta, N. O. Carragher, J. C. Acosta and D. A. Oyarzun, Nat. Commun., 2023, 14, 3445 CrossRef CAS.
- J. Qian, X. Wang, F. Song, Y. Liang, Y. Zhu, Y. Fang, W. Zeng, D. Zhang and J. Dong, Food Chem., 2025, 463, 141362 CrossRef PubMed.
- D. Zhang, Food Chem., 2026, 499, 147281 CrossRef.
- E. Heid, K. P. Greenman, Y. S. Chung, S. C. Li, D. E. Graff, F. H. Vermeire, H. Y. Wu, W. H. Green and C. J. Mcgill, J. Chem. Inf. Model., 2023, 64, 9–17 CrossRef PubMed.
- Z. P. Xiong, D. Y. Wang, X. H. Liu, F. S. Zhong, X. Z. Wan, X. T. Li, Z. J. Li, X. M. Luo, K. X. Chen, H. L. Jiang and M. Y. Zheng, J. Med. Chem., 2020, 63, 8749–8760 CrossRef.
- H. Kim, J. Lee, S. Ahn and J. R. Lee, Sci. Rep., 2021, 11, 11028 CrossRef PubMed.
- Y. Fang, Q. Zhang, N. Zhang, Z. Chen, X. Zhuang, X. Shao, X. Fan and H. Chen, Nat. Mach. Intell., 2023, 5, 542–553 CrossRef.
- S. M. Lundberg and S.-I. Lee, arXiv, 2017, preprint, arXiv:1705.07874, DOI:10.48550/arXiv.1705.07874.
- A. J. Alonso-Castro, J. R. Zapata-Morales, C. Solorio-Alvarado, A. Hernández-Santiago, L. A. Espinoza-Ramírez, C. Carranza-Álvarez and V. Ramadoss, Inflammopharmacology, 2020, 28, 749–757 CrossRef.
- G. J. M. Ketcha Wanda, S. Djiogue, F. Z. Gamo, S. G. Ngitedem and D. Njamen, J. Ethnopharmacol., 2015, 176, 494–498 CrossRef.
- G. Sotoing Taïwe, E. Ngo Bum, E. Talla, A. Dawe, F. C. Okomolo Moto, G. Temkou Ngoupaye, N. Sidiki, B. Dabole, P. D. Djomeni Dzeufiet, T. Dimo and M. De Waard, J. Ethnopharmacol., 2012, 143, 213–220 CrossRef.
- R. Estrada-Reyes, M. Martínez-Vázquez, A. Gallegos-Solís, G. Heinze and J. Moreno, J. Ethnopharmacol., 2010, 130, 1–8 CrossRef.
- F. C. F. de Sousa, B. A. Pereira, V. T. M. Lima, C. D. G. Lacerda, C. T. V. Melo, J. M. Barbosa-Filho, S. M. M. Vasconcelos and G. S. B. Viana, Phytother. Res., 2005, 19, 282–286 CrossRef PubMed.
- S. Kim, J. Chen, T. Cheng, A. Gindulyte, J. He, S. He, Q. Li, B. A. Shoemaker, P. A. Thiessen, B. Yu, L. Zaslavsky, J. Zhang and E. E. Bolton, Nucleic Acids Res., 2020, 49, D1388–D1395 CrossRef PubMed.
- D. Mendez, A. Gaulton, A. P. Bento, J. Chambers, M. De Veij, E. Félix, M. P. Magariños, J. F. Mosquera, P. Mutowo, M. Nowotka, M. Gordillo-Marañón, F. Hunter, L. Junco, G. Mugumbate, M. Rodriguez-Lopez, F. Atkinson, N. Bosc, C. J. Radoux, A. Segura-Cabrera, A. Hersey and A. R. Leach, Nucleic Acids Res., 2018, 47, D930–D940 CrossRef PubMed.
- D. S. Wishart, Y. D. Feunang, A. C. Guo, E. J. Lo, A. Marcu, J. R. Grant, T. Sajed, D. Johnson, C. Li, Z. Sayeeda, N. Assempour, I. Iynkkaran, Y. Liu, A. Maciejewski, N. Gale, A. Wilson, L. Chin, R. Cummings, D. Le, A. Pon, C. Knox and M. Wilson, Nucleic Acids Res., 2017, 46, D1074–D1082 CrossRef PubMed.
- C. Gottesmann, Neuroscience, 2002, 111, 231–239 CrossRef PubMed.
- P. Ertl and B. Rohde, J. Cheminf., 2012, 4, 12 Search PubMed.
- D. Rogers and M. Hahn, J. Chem. Inf. Model., 2010, 50, 742–754 CrossRef PubMed.
- J. L. Durant, B. A. Leland, D. R. Henry and J. G. Nourse, J. Chem. Inf. Comput. Sci., 2002, 42, 1273–1280 CrossRef.
- Z. Wen, J. Shi, B. He, J. Chen, K. Ramamohanarao and Q. Li, IEEE Trans. Parallel Distr. Syst., 2019, 30, 2706–2717 Search PubMed.
- M. Hajihosseinlou, A. Maghsoudi and R. Ghezelbash, Expert Syst. Appl., 2024, 237, 121668 CrossRef.
- S. Zhang, A. Golbraikh, S. Oloff, H. Kohn and A. Tropsha, J. Chem. Inf. Model., 2006, 46, 1984–1995 CrossRef.
- K. Mansouri, C. M. Grulke, R. S. Judson and A. J. Williams, J. Cheminf., 2018, 10, 10 Search PubMed.
- J. Zhang, H. Xing, A. Di Pizio, Q. Ke, X. Kou and D. Zhang, bioRxiv, 2026 DOI:10.64898/2026.01.21.700072.
- Q. Ke, J. Zhang, X. Huang, X. Kou and D. Zhang, Machine learning unveils three layers of food complexity, npj Science of Food, 2026 DOI:10.1038/s41538-026-00730-w.
- S. Ding, D. Liu, Y. Tian, D. Zhang, H. Xing, J. Chen, Z. Liu and Q. N. Hu, Synth. Syst. Biotechnol., 2025, 10, 1038–1049 CrossRef PubMed.
- P. Shi, S. Liu, J. Mao, X. Liu, R. Tu, H. Qin, A. Sun, D. Zhang and J. Mao, Trends Food Sci. Technol., 2026, 167, 105450 CrossRef CAS.
- D.V. Henley, N. Lipson, K.S. Korach and C.A. Bloch, N. Engl. j. Med., 2007, 356, 479–485 CrossRef CAS.
Footnote |
| † Co-first authors. |
| This journal is © The Royal Society of Chemistry 2026 |






