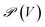Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Higher order structures in chemistry: hypergraphs reshape the molecule and the reaction†
Guillermo
Restrepo
 *
*
Max Planck Institute for Mathematics in the Sciences, Inselstraße 22, 04103 Leipzig, Germany. E-mail: restrepo@mis.mpg.de; Fax: +49 (0) 341 9959 658; Tel: +49 (0) 341 9959 601
First published on 15th January 2026
Abstract
Chemical systems contain higher-order relationships that exceed the binary constraints of traditional graph-based models. Although graph theory has long supported the digital representation of molecules and reactions, many fundamental chemical phenomena—such as multi-centre bonding, aromaticity, cooperative interactions, and the inherently set-theoretical nature of reactions—escape pairwise encodings. This work introduces hypergraphs and an extended “zoo” of higher-order mathematical structures as a unified framework for modelling both molecular structures and reaction networks. Molecular hypergraphs naturally capture multi-atomic interactions, while directed hypergraphs offer a mathematically faithful representation of reactions as transformations between arbitrary sets of substances. More sophisticated variants—including ordered, directed, binary, and directed-ordered hypergraphs—enable the incorporation of additional chemical information, such as atomic ordering, ligand–pocket affinities, and cavity organisation in porous materials at the substance level, as well as toxicity and economic constraints at the reaction level. Recent advances in hypergraph spectral theory, random models, and higher-order network statistics have opened new chemical, mathematical, and computational avenues. These developments coincide with emerging machine-learning evidence showing that hypergraph-based representations of molecules can outperform graph-based and even 3D-coordinate models. By outlining both the capabilities and current limitations of hypergraph approaches, this work argues that higher-order mathematical structures will be central to the next generation of digital discovery, enabling more faithful representations of chemical complexity and deeper integration across chemistry, mathematics, and computer science.
1 Introduction
Chemistry is characterised by its concern with the organisation, behaviour, and transformation of matter at multiple scales. Throughout its history, chemists have relied on high levels of abstraction to interpret, organise, and systematise the practice of their discipline.1 From molecular models, encoding atomic relationships, to the periodic system, including sets and order relationships,2 abstract frameworks have long been essential in advancing chemical knowledge.Building on this historical interplay between chemistry and mathematical abstraction, the present Perspective introduces hypergraphs as a mathematical structure of particular relevance for addressing chemical questions at both the molecular and reaction levels. In doing so, it aims to broaden the ongoing dialogue between chemistry and mathematics and to point towards new directions for chemical theory, computation, and practice.
2 Higher order structures
Gerhardt, the visionary 19th-century chemist,3–5 produced scientific results that addressed two foundational pillars of chemistry. Through his type representation of substances, he helped lay the groundwork for what later became molecular structural theory,4 a conceptual framework that remains central to the language of chemistry and to the way we encode chemical knowledge for computational analysis.3 Gerhardt also contributed to the modern representation of chemical reactions in the now-standard form A + B → C + D.‡ Viewed from a contemporary mathematical perspective, these contributions highlight the fundamental role of relationships in chemistry, from the atomic to the molecular level, that is form substances to reactions.The second half of the 19th century further strengthened the connection between chemistry and the mathematics of relations. Sylvester, a leading 19th-century mathematician, recognised a connection between the molecular structures of his contemporary chemists and what mathematicians would later call graphs.7,8 In this connection, atoms correspond to vertices and chemical bonds to edges (pairs of vertices), a correspondence Sylvester himself termed “chemicographs.”8 This insight established a lasting bridge between chemical structural theory and mathematical graph theory,9,10 which has generated a large amount of research and has led to applications ranging from structural prediction to QSAR (Quantitative Structure-Activity Relationships) modelling.1,11–13
Similar ideas emerged when chemists began to use graphs to model the reactivity of substances, ultimately leading in the 20th century to the study of reaction networks, in which substances are represented as vertices and their reactivity relations as edges.14–17 This mathematical framework is at the basis of the multiple studies on metabolic networks and on the gigantic network spanning the chemical space.14,15,18
Despite these successes, graph-based models face intrinsic limitations. As discussed in the next section, not all molecules can be reduced to binary atomic relations. Aromatic compounds, multicentre bonds, organometallic species, and numerous other systems require the interaction of sets of atoms rather than simple pairs. Whenever two sets of atoms interact—and especially when at least one of those sets contains multiple atoms—graphs, in the strict sense, are insufficient.
Chemical reactions exhibit a parallel limitation: they are fundamentally set-theoretical entities. A reaction relates a set of educts to a set of products, and each of these sets frequently contains more than one substance. Representing such transformations through binary edges inevitably leads to the loss or distortion of essential chemical information.
Before turning to the next section, it is therefore important to emphasise that chemistry is fundamentally shaped by higher-order relationships. The discipline extends far beyond binary interactions: both molecular structure and chemical reactivity routinely depend on the collective behaviour of atom sets and substance sets. Higher-order relations are not rare exceptions but a natural feature of both molecular structure and chemical reactivity, and recognising them is essential for developing mathematical frameworks that truly reflect chemical reality.
2.1 Graphs, hypergraphs and the molecular structure
This section outlines the achievements and constraints of chemical graph theory before turning to hypergraphs as a natural generalisation capable of encoding higher-order atomic relations. | ||
| Fig. 1 Graph and hypergraph molecular models. (a) Methane, (b) benzene, (c) diborane, (d) ferrocene, (e) non-covalent interactions. In (a–c) hyperedges correspond to chemical bonds (shown as coloured regions), with shared electrons (dots). In (b) equivalent aromatic carbons are represented by a grey hyperedge. In (d) each cyclopentadienyl ring forms a hyperedge describing its collective interaction with the Fe centre. In (e) the ligand–protein pocket interaction is represented by hyperedges gathering the highly interacting protein regions (red) together with the corresponding ligand fragments (figure reproduced from ,19 published under a CC BY 2.0 license. Reproduced with permission under the Creative Commons Attribution 2.0 International License). Structures in (b) (left) as depicted by Kekulé (reproduced from 20 with permission from John Wiley and Sons, copyright 2026). Structure in (c) (left) as published by Longuet-Higgins and Bell and reproduced from 21 with permission from The Royal Society of Chemistry, copyright 2026. Structures in (d) (left) as depicted by Wilkinson, Rosenblum, Whiting, and Woodward (reproduced from 22 with permission from the American Chemical Society, copyright 2026). | ||
A key strength of molecular graph theory, key for computational encoding and applications, is that graphs can be algebraised. Once encoded as matrices—typically adjacency or incidence matrices—graphs become amenable to linear algebra.§ This enables the derivation of eigenvalues, eigenvectors, spectra, path counts, cycle detection, and many other mathematically defined quantities with chemical interpretation.23–25 From these representations emerged the vast field of molecular descriptors, now numbering in the thousands,26 forming the backbone of QSAR modelling.1,11–13
Despite these successes, a fundamental limitation remains: not all molecules can be adequately represented using only binary atomic relations.27,28 The inadequacy was already evident in the 19th century for benzene and other aromatic systems,20,29 and re-emerged in the mid-20th century for diborane and ferrocene.21,22 In modern contexts such as receptor–ligand complexes, binary-only representations are even more restrictive.30
In all these cases—aromaticity, multi-centre bonding, organometallic bonding, and protein–ligand recognition—the essential feature is their set-theoretical nature: benzene involves six atoms interacting collectively; the bridging bonds in diborane involve three atoms; each cyclopentadienyl ring in ferrocene interacts as a five-atom unit with Fe; protein pockets interact with a set of ligand atoms (Fig. 1). These interactions cannot be reduced without loss to pairs of atoms.
The recognition that molecular structure routinely depends on higher-order atomic relationships lies at the heart of the hypergraph approach. Hypergraphs emerged in the chemical literature in the 1990s,11,28,31–33 and have recently been rediscovered—both as a faithful mathematical model of multi-atom interactions and as a powerful computational framework.11,28 One goal of this perspective is to encourage chemists, mathematicians, and computer scientists to revisit hypergraphs as a natural representation of molecular structure.
Formally, a hypergraph is defined as H = (V, E), where V is the set of vertices and E the set of hyperedges. A hyperedge may be any subset of V. For instance, a hypergraph model of diborane B2H6 can be written as V = {{B1, B2, H1, H2, H3, H4, H5, H6}} and E = {{B1,H1}, {B1,H2}, {B1,H3,B2}, {B1,H4,B2}, {B2,H5},{B2,H6}}, where the two three-atom hyperedges {B1, H3, B2} and {B1, H4, B2} encode the two bridging 3-centre–2-electron bonds (Fig. 1c, where atoms are not labelled for simplicity). Note how these hyperedges cannot be represented in a graph setting, since they contain more than two vertices. Parallel representations for methane, benzene, and ferrocene are shown in Fig. 1.
As with molecular graphs, the usefulness of hypergraphs becomes most apparent when they are algebraically encoded so that computations can be performed. In analogy with graphs, hypergraphs can be represented by adjacency matrices, where each entry records whether the i-th and j-th atoms participate together in at least one hyperedge. Here, “participation in a relation” generalises the notion of a chemical bond: for diborane, for example, the adjacency matrix records the three-atom relations of the bridging {B, H, B} bonds, the cyclic six-atom relation in benzene, and the five-atom cyclopentadienyl rings of ferrocene. Hypergraphs also admit incidence matrices, recording vertex membership in each hyperedge.
Based on these matrix representations, spectral hypergraph theory has begun to emerge, encompassing Laplacians, eigenvalues, eigenvectors, and associated invariants.40–46 Remarkably, the connection to chemistry was already recognised in the 1990s, when the first hypergraph-based molecular descriptors were proposed.31–33 These developments parallel the trajectory of molecular graph theory, where algebraic descriptors became central to QSAR research.
Recent machine learning work further strengthens the case for hypergraph formulations. Hypergraph neural networks have shown clear advantages over graph-based models—and even over some 3D-coordinate–based representations—in predicting molecular properties and reactivity.28 These methods leverage the ability of hypergraphs to encode multi-atom interactions natively rather than inferring them indirectly from pairwise data.
The trajectory that once took graph theory from a mathematical curiosity to a central tool of theoretical and computational chemistry now appears to be unfolding for molecular hypergraphs. Their capacity to encode higher-order atomic interactions makes them a natural extension of chemical modelling. With the rapid development of hypergraph mathematics and computation, this framework offers a fertile arena for future collaboration between chemists, mathematicians, and computer scientists.
Yet chemistry encompasses both substances and their transformations. Just as hypergraphs enrich the modelling of molecular structure, they also provide expressive representations of chemical reactions and reaction networks. This is the focus of the next section.
2.2 Graphs, hypergraphs and chemical reactions
The algebraic representation of chemical reactions, developed since Gerhardt's times,3 is now commonly interpreted through graph-theoretical lenses. Graphs have enabled large-scale analyses of reactivity patterns, retrosynthetic pathways, and chemical networks.14–17 Yet, as with molecular structure, the binary constraints of graphs impose limitations on the faithful encoding of chemical transformations. This section outlines the possibilities and shortcomings of graph-based reaction models and shows how higher-order structures—specifically directed hypergraphs—provide the appropriate mathematical generalisation.One of the simplest and most widely used approaches is the educt–product model.|| In this representation, a transformation in which substance B is formed from A is encoded as A → B.27 Formally, such a relation is represented as an arc, or directed edge, (A and B). This construction has been adopted extensively in studies of metabolic, synthetic, and general chemical reaction networks.14–18
However, the model suffers from a fundamental limitation inherent to its binary nature.27,47–49 Consider the reaction A + B → C + D. The educt–product model represents this transformation as the four arcs A → C, A → D, B → C, B → D, yielding the graph G = (V, E) with V = {A, B, C, D} and E = {(A, C), (A, D), (B, C), (B, D)} (see reaction r1 in Fig. 2).
 | ||
| Fig. 2 Chemical reaction models. Directed graph and directed hypergraph representations of the three reactions shown at the top. In the graph-based model, educts and products are connected by directed edges (arrows), each encoding a binary relation. In contrast, the hypergraph model relates sets of educts to sets of products via directed hyperedges, thereby capturing the set-theoretical structure of reactions. The arrow-based directed hypergraph notation follows Fig. 1 in.50 | ||
As discussed in,27 recovering reactions from this graph leads to multiple spurious interpretations, all of which are consistent with the graph but inconsistent with chemistry. From G, one may infer not only the true reaction A + B → C + D, but also unimolecular reactions A → C, A → D, B → C, and B → D, as well as the artificial bimolecular reactions A + B → C and A + B → D. These arise because the model decomposes each true reaction into pairwise relations between individual substances, ignoring the essential fact that educts act jointly.
The consequences become even more problematic in network-level analyses. In the network shown in Fig. 2, the graph model incorrectly predicts that substances D and G remain reachable even when A is absent, owing to paths such as B → D, B → C → G. Yet the underlying chemistry—embodied in the reactions at the top of the figure—shows clearly that none of these products can form without A. The root of the problem is conceptual: a chemical reaction does not relate individual substances but rather sets of substances.
Formally, a reaction maps a set of educts to a set of products. The educt–product model only faithfully represents rearrangements, where both sets have cardinality 1. But empirical analysis of the chemical literature reveals that chemists overwhelmingly report reactions involving one to three educts and one or two products.51 This diversity of set sizes cannot be encoded in the binary framework of graphs.
A mathematically faithful representation of chemical reactions requires a framework that accommodates relations between sets of arbitrary size. This is precisely what directed hypergraphs provide, as discussed in the next subsection.
Formally, a directed hypergraph H = (V, E) consists of a set of vertices V and a set of hyperedges E. Each hyperedge is an ordered pair of vertex-sets (ei, ej), where ei is the tail (or source) and ej the head (or target). In chemical terms, ei represents the set of educts and ej the set of products. The sets ei and ej are often called hypervertices or hypernodes to emphasise that they generalise the notion of a vertex by potentially containing several vertices.46
For example, the reaction A + B → C + D is represented by the directed hypergraph H = (V, E) with V = {A, B, C, D} and E = ({A, B}, {C, D}). This contrasts sharply with its graph representation (Section 2.2.1), which requires four arcs: A → C, A → D, B → C, and B → D. Likewise, the three reactions shown in Fig. 2 are captured by only three directed hyperedges rather than 10 arcs—illustrating the parsimony and clarity of the hypergraph representation.
These examples show that directed hypergraphs encode the set-theoretical essence of chemical reactions: both the educt set and the product set may contain any number of substances. Importantly, the chemical consequences of the absence of a substance become immediately transparent. As discussed in Section 2.2.1, removing A from the system makes the reaction r1 impossible, which in turn prevents the formation of C and thus blocks reaction r3 and the production of G. Such dependencies are obscured in the educt–product model.
As with molecular and reaction graphs, directed hypergraphs admit algebraic representations. Adjacency matrices of dimension m × m (with m the number of hypervertices) encode whether a hypervertex ei leads to another hypervertex ej. Incidence matrices of dimension m × n (with n the number of reactions) indicate whether a hypervertex acts as educt (entry +1) or product (entry −1) in a given reaction; a zero indicates non-participation. These matricial forms enable the study of the structural and dynamical properties of reaction networks.
Recent developments have extended classical network statistics to directed hypergraphs, including clustering coefficients, spectral measures, curvature, shortest paths, communicability, and random models.46,49,52–62 Several have already been applied to large biochemical networks. For instance, clustering coefficients reveal that metabolic networks are far less clustered than human-made systems such as email networks.** Spectral centralities have likewise been used to study biochemical networks, urban transit systems, and propositional-logic databases.53
Thermodynamic constraints have recently been incorporated into directed hypergraph reaction networks to identify pathways composed exclusively of energetically favourable reactions.63 Despite these advances, further work is needed to unify approaches, identify redundancies, and determine whether chemical networks possess structural signatures distinguishing them from other directed hypergraphs.
A further active avenue of research concerns random models for reaction hypergraphs, analogous to random graph models in classical network theory.64,65 An Erdős–Rényi–type model has been proposed for chemical hypergraphs,46 enabling comparisons between empirical networks and suitable null models.†† Yet many well-known random-graph models—including Watts–Strogatz small-world66 and Barabási–Albert preferential-attachment models67—remain to be generalised to the hypergraph setting. A key variable unique to hypergraphs, and absent from graph-based studies, is hypervertex size. This additional degree of freedom is essential and must be incorporated into any statistical or generative model of reaction networks.
In summary, directed hypergraphs provide a mathematically expressive and chemically faithful framework for encoding reactions and reaction networks. Their recent mathematical development opens many avenues for future research at the interplay of chemistry, mathematics, and computer sciences, offering fresh perspectives on the structure and organisation of large chemical networks—including those spanning the full chemical space. The following section extends the discussion to further hypergraph structures of chemical relevance.
2.3 A zoo of hypergraph structures of chemical application
In this section, I introduce a family of mathematical structures obtained by generalising different aspects of graphs, and I outline their potential—and in some cases already demonstrated—applications in chemistry.Beyond graphs, hypergraphs, and binary hypergraphs, richer structures emerge once order relations are introduced. A familiar case is the directed graph, obtained by endowing each two-element edge with a direction, thereby producing arcs (Fig. 3). If sets of arbitrary size are allowed while retaining an internal order structure, ordered hypergraphs are obtained.§§ Ordered hypergraphs thus constitute collections of partially ordered sets (posets)71 over V. If, in addition, order relations are permitted between pairs of these posets, the result is a directed ordered hypergraph (Fig. 3). Removing the internal order within hypervertices yields a directed hypergraph,57,72 while removing direction but retaining internal ordering leads to an ordered binary hypergraph. Adding direction to the latter reconstructs the directed ordered hypergraph. These interrelations are examples of morphisms between higher-order structures, some of which are illustrated in Fig. 3.
Although often not explicitly recognised as such, many chemical systems already exhibit the defining features of these higher-order structures. In the remainder of this section, I discuss several examples and highlight how exploring the mathematics of these structures may deepen our understanding of chemicals and reactivity.
 | ||
| Fig. 4 Graphs and hypergraphs as molecular structure models. Directed graphs representing (a) inductive effects as depicted by Ingold (reproduced from ref. 74 with permission from the American Chemical Society, copyright 2026) and (b) hydrogen bonds (adapted from Fig. 7 in ref. 73 with permission from the Royal Society of Chemistry, copyright 2026). (c) Ordered hypergraphs encoding electronegativity differences within bonds (left), and encoding the poset structure (white arrows) of specific regions (white dots) inside protein pockets (white curves) (right). (d) Directed ordered hypergraph on a 2D layer of MOF-5 (adapted from Fig. 1 in75 with permission from the Royal Society of Chemistry, copyright 2026). Cavities (hypervertices) are ordered according to host–guest affinity (orange arrows), and internal hypervertex order is shown at top right, which illustratively indicates the exposition of some atoms to the cavity. (e) Directed hypergraphs encoding σ-donation and π-backdonation between B(SiH3)3 and N2 (ref. 76) (left), and the side–on interaction between O2 and Ni in [NiIII(13-TMC)(O2)]+ (right), with 13-TMC as the N-tetramethylated cyclam chelate with 13 atoms of ring size.77 (f) Binary hypergraphs depicting bond–bond interactions in an ester (left) and Se–P spin–spin coupling in 1-Ph2P(C10H6)-8-P(:Se)Ph2 (right).78 | ||
Hypergraphs offer a natural way to reduce and navigate chemical complexity. The chemical space contains millions of known substances;51 classification is therefore essential.50,79 Classes of compounds or classes of reactions correspond to hyperedges in a hypergraph of the chemical space. Alkali metals, halogens, endocrine disruptors, and metal–organic frameworks (MOFs) are examples of hyperedges. Likewise, reaction classes—such as amide formation, Diels–Alder cycloadditions, Buchwald–Hartwig couplings—form another family of hyperedges. These sets often intersect, as molecules or reactions commonly belong to multiple classes, yielding a hypergraph with non-disjoint hyperedges.
At the molecular scale, hypergraphs arise naturally when atomic equivalence classes or multi-centre bonding motifs are used to define hyperedges. Any molecule can thus be modelled as a hypergraph whose hyperedges encode general m-centre–n-electron bonding patterns (Fig. 1).
Ordered hypergraphs are equally widespread. Ordering chemical elements by atomic radius or electronegativity yields ordered hypervertices within the hypergraph of the periodic system. In this setting, hypervertices correspond to families of chemical elements. When several parameters are used simultaneously to characterise chemical elements, posets arise, reflecting cases where no total order exists.2¶¶ Toxicity rankings, ligand–pocket affinities, electrochemical series, or spectrochemical series all constitute ordered hypergraphs. At the molecular level, bonds may be ordered by electronegativity difference or by polarisability (Fig. 4c). Protein pockets are also amenable of being modelled using ordered hypergraphs, where products contain regions ordered by their interaction strength with specific ligands (Fig. 4c). MOFs accept a similar treatment, where cavities may be ordered by their host–guest affinity.80,81
The periodic system, besides being the icon of chemistry, is a chemical object with a rich mathematical structure.2,82 Here, the corresponding hypergraph structure is the directed ordered hypergraph (Fig. 3), where hypervertices are element families endowed with internal order (for instance, by atomic size or electronegativity), while hypervertices are themselves ordered, as clearly recognised in group trends such as alkali metals being more electropositive than halogens. A further case of a chemical directed ordered hypergraph is the resulting from ordering chemicals by their degree of substitution.83,84 In this case, hypervertices are chemicals with the same degree of substitution.||||.
Directed ordered hypergraphs also naturally arise in catalyst selection. Suppose catalysts are classified into Pd-based and Ni-based families (hypervertices), with overlapping members. Additional parameters—cost, toxicity, availability—induce internal poset structures. Directed ordered relations between hypervertices enable the selection of the most promising catalyst class, after which internal ordering identifies the optimal candidate. Therefore, exploring the mathematical properties of this model may improve the AI-driven studies on synthesis planning under realistic chemical and external constraints.
The same framework supports retrosynthetic analysis and decision-making in self-driving laboratories.85 Substances form posets based on several criteria (toxicity, solubility, cost), reactions form hypervertices, and directed order between hypervertices identifies preferable synthetic routes.
Directed ordered hypergraphs also model the ordering of cavities within MOFs (Fig. 4d). Cavities (hypervertices) are ordered by their adsorption capacity or accessibility, while their internal order reflects steric and electronic factors arising from secondary building units.
Back to Fig. 4, a further hypergraph structure is the directed hypergraph, which, as shown earlier, provides a rigorous framework for modelling chemical reactions.46,51 They also offer molecular-level insight, for example by encoding directed interactions between atom sets, as in σ-donation and π-backdonation in metal complexes or the side-on binding of O2 to transition metals (Fig. 4e).76,77,86
Binary hypergraphs, in turn, capture interactions between sets of atoms and are particularly relevant for bond–bond couplings. Therefore, these hypergraph structures constitute suitable molecular models of application in different spectroscopies. For instance, vibrational spectroscopies such as IR and Raman, which reveal mode couplings—for example, between C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O and C–O stretches in esters (Fig. 4f)—could find a natural framework for interpretation with binary hypergraphs, where interacting bonds become the centre of attention. Likewise, the long-range spin–spin couplings (Fig. 4f), detected with NMR spectroscopy find in binary hypergraphs a suitable model.
O and C–O stretches in esters (Fig. 4f)—could find a natural framework for interpretation with binary hypergraphs, where interacting bonds become the centre of attention. Likewise, the long-range spin–spin couplings (Fig. 4f), detected with NMR spectroscopy find in binary hypergraphs a suitable model.
Chemical reaction networks can also be modelled as binary hypergraphs when the goal is to study global connectivity rather than the educt–product distinction. This binary hypergraph model was used recently for developing a random model for the chemical space.46
Finally, when atoms or groups of atoms are characterised by multiple parameters—connectivity, conformational state, vibrational signatures, mechanistic role, stereochemistry, electronic structure—they form posets. Interactions between such posets are central in diverse chemical phenomena: coupling between conformational substates during protein folding;87 vibrational poset interactions through anharmonic couplings;88 mechanistic posets constraining multistep pathways;89 stereochemical posets governing diastereomer stability;90 and electronic-structure posets whose interactions give rise to conjugation or aromaticity.91 In all these cases, chemical behaviour emerges from the interplay of multiple interacting posets, each encoding a distinct facet of molecular organisation. These cases are suitably modelled by ordered binary hypergraphs (Fig. 4).
3 Conclusions
The modelling of chemical systems at the molecular and reaction levels has traditionally relied on graphs. While these structures have proved extremely fruitful, they also exhibit clear limitations. Graphs are intrinsically binary, and therefore struggle to capture the multi-centric, higher-order, and context-dependent interactions characteristic of both molecular structure and chemical reactions. Hypergraphs, by contrast, naturally encode relations among sets of arbitrary size and therefore provide a far richer mathematical language for representing chemical complexity.In addition to presenting the uses of some hypergraph structures in chemistry, namely molecular hypergraphs and directed hypergraphs as a model for chemical reactions, a zoo of other hypergraph structures is presented, from ordered hypergraphs to directed ordered hypergraphs and ordered binary hypergraphs, which nuance the description of molecules and chemical reactions.
Despite their advantages, hypergraph-based models must be used with appropriate caution. Like graphs, these higher-order structures capture selected aspects of chemical systems—specifically their relational or topological organisation—but they do not, by themselves, encode all chemically relevant information. Hypergraphs significantly enrich the representation of multi-atomic and multi-molecular relationships, yet whenever three-dimensional geometric details are essential—such as quantifying cavity sizes in MOFs, characterising protein pockets, or distinguishing molecular conformations—additional structural information must be incorporated. This need mirrors the well-known limitations of graph-based molecular representations: embedding topological structures into metric space requires complementing the (hyper)graph with weighted vertices and edges and, crucially, with explicit coordinate systems. In such settings, hypergraphs serve as a powerful relational scaffold, but geometry must be supplied through appropriate metric or spatial augmentations.
A further element of caution when using hypergraphs in chemistry is related to the relative novelty of the hypergraph literature. Hypergraph theory is still an emerging and rapidly developing field, and its nomenclature has not yet reached the level of standardisation enjoyed by graph theory. As a consequence, the same mathematical object may carry different names in different subfields, while distinct objects may be referred to by similar terminology. For this reason, reading the hypergraph literature requires close attention to definitions rather than reliance on terminology alone. This conceptual heterogeneity is a challenge, but also a reflection of the vibrant and expanding nature of the field.
From a computational perspective, the situation is equally nuanced. Several hypergraph statistics can be mapped into statistics on bipartite graphs, which sometimes permits the transfer of efficient graph-theoretical algorithms. Yet this reduction is not universally applicable.47 In fact, many core tasks in hypergraph analysis remain computationally demanding. For example, computing shortest paths in weighted hypergraphs is NP-hard.47 Further examples of hypergraph computational complexity are discussed in.92–94 These challenges have direct implications for chemistry, where algorithmic efficiency is crucial. A case in point is the determination of whether a molecular fragment occurs within a larger structure—a task that reduces to subgraph matching in graph theory but becomes considerably more complex in hypergraphs.95,96 Understanding and addressing these complexities is essential for the practical use of hypergraphs in chemistry.
The mathematical and computational study of hypergraphs is, as shown throughout this perspective, an active and rapidly evolving domain. When combined with the richer and more sophisticated hypergraph structures introduced here—structures that extend well beyond classical hypergraphs into what is dubbed as a “zoo” of higher-order relational frameworks—the challenges are amplified. There remains a substantial amount of theoretical and algorithmic work to be done. Addressing these issues is not only a promising direction for mathematics and computer sciences, but also one from which chemistry stands to benefit enormously.
Another frontier concerns machine learning. Recent studies have already shown that hypergraph representations of molecular systems outperform both traditional graph-based models and models incorporating full three-dimensional information.28 These advances derive from the extension of graph neural networks into hypergraph neural networks,28 which can exploit higher-order structural information unavailable to pairwise models. It is natural to ask what would happen if the novel hypergraph structures introduced in this paper—ordered hypergraphs, directed hypergraphs, and other enriched forms—were incorporated into new machine learning architectures. Such developments could radically expand the representational and predictive power of models for chemical discovery.
In summary, hypergraphs and their mathematical extensions offer a powerful framework for modelling the richness and complexity of chemical systems. Their integration with modern computational approaches, including algorithmic advances and machine learning, opens a path toward a new generation of tools for digital discovery. By embracing these higher-order structures, chemistry gains access to a deeper and more expressive mathematical language—one capable of capturing complexity that lies beyond the reach of traditional, pairwise models.
Author contributions
G. R. was responsible for conceptualisation, formal analysis, investigation, methodology development, visualisation, and manuscript writing.Conflicts of interest
There are no conflicts to declare.Data availability
There is no original data associated to this document.Acknowledgements
G. R. gratefully acknowledges many years of discussions on topics related to those presented here with Rainer Brüggemann, Douglas Klein, Jürgen Jost, Peter F. Stadler, Angel Garcia-Chung, Marisol Bermúdez-Montaña, Wilmer Leal, and numerous other colleagues and collaborators. This paper was motivated by a lecture given at the Workshop on Random Simplicial Complexes and Hypergraphs, Nottingham, UK, September 1-2, 2025, organised by Jason P. Smith and Karim Bibars. Open Access funding provided by the Max Planck Society.Notes and references
- G. Restrepo, Essays in the Philosophy of Chemistry, Oxford University Press, 2016 Search PubMed.
- W. Leal and G. Restrepo, Proc. R. Soc. A, 2019, 475, 20180581 CrossRef PubMed.
- C. Gerhardt, Traité de chimie organique, F. Didot frères, 1860.
- H. Chang, Is Water H2O?: Evidence, Realism and Pluralism, Springer, Netherlands, 2012 Search PubMed.
- A. J. Rocke, Chemical Atomism in the Nineteenth Century, Ohio State University Press, 1984 Search PubMed.
- C. Gillispie, The Edge of Objectivity: An Essay in the History of Scientific Ideas, Princeton University Press, 1973 Search PubMed.
- J. J. Sylvester, Am. J. Math., 1878, 1, 64–104 CrossRef.
- J. J. Sylvester, Nature, 1878, 17, 284 Search PubMed.
- G. Restrepo, Found. Chem., 2013, 15, 185–197 CrossRef CAS.
- D. J. Klein, Hyle, 2013, 19, 35–85 CAS.
- L. S. G. Leite, S. Banerjee, Y. Wei, J. Elowitt and A. E. Clark, WIREs Comput Mol Sci, 2024, 14, e1729 CrossRef CAS.
- P.-Y. Cao, Y. He, M.-Y. Cui, X.-M. Zhang, Q. Zhang and H.-Y. Zhang, J. Cheminf., 2024, 16, 133 Search PubMed.
- H. Qin, A. F. Hashem, M. F. Hanif and O. A. Fiidow, Sci. Rep., 2025, 15, 28025 CrossRef CAS PubMed.
- M. Fialkowski, K. J. M. Bishop, V. A. Chubukov, C. J. Campbell and B. A. Grzybowski, Angew. Chem., Int. Ed., 2005, 44, 7263–7269 CrossRef CAS PubMed.
- K. J. M. Bishop, R. Klajn and B. A. Grzybowski, Angew. Chem., Int. Ed., 2006, 45, 5348–5354 CrossRef CAS PubMed.
- S. Fujita, Computer-Oriented Representation of Organic Reactions, Yoshioka Shoten Publishing Company, 2001.
- S. Rao, A. van der Schaft and B. Jayawardhana, J. Math. Chem., 2013, 51, 2401–2422 CrossRef CAS.
- H. Jeong, B. Tombor, R. Albert, Z. N. Oltvai and A.-L. Barabási, Nature, 2000, 407, 651–654 CrossRef CAS PubMed.
- V. Bianchi, P. F. Gherardini, M. Helmer-Citterich and G. Ausiello, BMC Bioinf., 2012, 13, S17 CrossRef CAS PubMed.
- A. Kekulé, Justus Liebigs Ann. Chem., 1872, 162, 77–124 CrossRef.
- H. C. Longuet-Higgins and R. P. Bell, J. Chem. Soc., 1943, 250–255 RSC.
- G. Wilkinson, M. Rosenblum, M. C. Whiting and R. B. Woodward, J. Am. Chem. Soc., 1952, 74, 2125–2126 CrossRef CAS.
- N. Trinajstić, Chemical Graph Theory, CRC Press, 2018 Search PubMed.
- N. Biggs, Algebraic Graph Theory, Cambridge University Press, 1993 Search PubMed.
- X. Li, Y. Shi and I. Gutman, Graph Energy, Springer, 2012 Search PubMed.
- R. Todeschini, V. Consonni, R. Mannhold, H. Kubinyi and G. Folkers, Molecular Descriptors for Chemoinformatics: Volume I: Alphabetical Listing/Volume II: Appendices, References, Wiley, 2009 Search PubMed.
- G. Restrepo, Digital Discovery, 2022, 1, 568–585 RSC.
- J. Chen and P. Schwaller, J. Chem. Phys., 2024, 160, 144307 CrossRef CAS PubMed.
- A. J. Rocke, Angew. Chem., Int. Ed., 2015, 54, 46–50 CrossRef CAS PubMed.
- K. A. Murgas, E. Saucan and R. Sandhu, Sci. Rep., 2022, 12, 20879 CrossRef CAS PubMed.
- E. V. Konstantinova and V. A. Skoroboratov, J. Struct. Chem., 1998, 39, 958–966 CrossRef CAS.
- X. Liu, L. Wang and X. Li, J. Comb. Optim., 2020, 39, 351–364 Search PubMed.
- E. V. Konstantinova and V. A. Skoroboratov, J. Chem. Inf. Comput. Sci., 1995, 35, 472–478 Search PubMed.
- P. Corsini and V. Leoreanu, Applications of Hyperstructure Theory, Springer US, 2003 Search PubMed.
- C. Berge, Graphs and Hypergraphs, North-Holland Publishing Company, 1973 Search PubMed.
- J. Bilsky and A. E. Clark, J. Chem. Phys., 2025, 163, 090901 CrossRef CAS PubMed.
- K. Xia, X. Feng, Y. Tong and G. W. Wei, J. Comput. Chem., 2015, 36, 408–422 CrossRef CAS PubMed.
- K. Xia and G.-W. Wei, J. Comput. Chem., 2015, 36, 1502–1520 Search PubMed.
- A. R. Benson, D. F. Gleich and D. J. Higham, Higher-order Network Analysis Takes Off, Fueled by Classical Ideas and New Data, arXiv, 2021, DOI:10.48550/arXiv.2103.05031.
- R. Mulas, C. Kuehn, T. Böhle and J. Jost, Discret. Appl. Math., 2022, 317, 26–41 CrossRef.
- R. Mulas, D. Horak and J. Jost, in Graphs, Simplicial Complexes and Hypergraphs: Spectral Theory and Topology, ed. F. Battiston and G. Petri, Springer International Publishing, Cham, 2022, pp. 1–58 Search PubMed.
- J. Jost, R. Mulas and D. Zhang, Vietnam J. Math., 2022, 50, 323–358 CrossRef.
- R. Mulas, C. Kuehn and J. Jost, Phys. Rev. E, 2020, 101, 062313 Search PubMed.
- E. Andreotti and R. Mulas, Electron. J. Graph Theory Appl., 2022, 10, 485–500 CrossRef.
- R. Mulas and J. Nie, Acta Math. Hung., 2023, 170, 168–182 CrossRef.
- A. Garcia-Chung, M. Bermúdez-Montaña, P. F. Stadler, J. Jost and G. Restrepo, J. Math. Chem., 2024, 62, 1357–1383 Search PubMed.
- S. Klamt, U.-U. Haus and F. Theis, PLoS Comput. Biol., 2009, 5, 1–6 CrossRef PubMed.
- M. Wen, E. W. C. Spotte-Smith, S. M. Blau, M. J. McDermott, A. S. Krishnapriyan and K. A. Persson, Nat. Comput. Sci., 2023, 3, 12–24 CrossRef PubMed.
- C. Flamm, B. M. R. Stadler and P. F. Stadler, Generalized Topologies: Hypergraphs, Chemical Reactions, and Biological Evolution, in Advances in mathematical chemistry and applications, ed. S. C. Basak, G. Restrepo and J. L.Villaveces, Bentham-Elsevier, 2015, ch. 2, pp. 300–328 Search PubMed.
- J. Schummer, Hyle, 1998, 4, 129–162 CAS.
- E. J. Llanos, W. Leal, D. H. Luu, J. Jost, P. F. Stadler and G. Restrepo, Proc. Natl. Acad. Sci. U. S. A., 2019, 116, 12660–12665 Search PubMed.
- G.-G. Ha, I. Neri and A. Annibale, J. Nonlinear Sci., 2024, 34, 043102 Search PubMed.
- G. Contreras-Aso, R. Criado and M. Romance, J. Complex Netw., 2024, 12, cnae037 CrossRef.
- W. Leal, G. Restrepo, P. F. Stadler and J. Jost, Adv. Complex Syst., 2021, 24, 2150003 Search PubMed.
- M. Eidi and J. Jost, Sci. Rep., 2019, 10, 12466 Search PubMed.
- S. Kim, M. Choe, J. Yoo and K. Shin, Data Min. Knowl. Discov., 2023, 37, 2330–2388 Search PubMed.
- G. Ausiello and L. Laura, Theor. Comput. Sci., 2017, 658, 293–306 CrossRef.
- S. S. Shetty and K. A. Bhat, Arab J. Basic Appl. Sci., 2025, 32, 347–360 Search PubMed.
- R. Fagerberg, C. Flamm, R. Kianian, D. Merkle and P. F. Stadler, J. Cheminf., 2018, 10, 19 Search PubMed.
- P. Traversa, G. Ferraz de Arruda, A. Vazquez and Y. Moreno, Entropy, 2023, 25 Search PubMed.
- S. Krieger and J. Kececioglu, J. Comput. Biol., 2023, 30, 1198–1225 CrossRef CAS PubMed.
- S. G. Marehalli Srinivas, M. Esposito and N. Freitas, J. Chem. Phys., 2025, 163, 194112 CrossRef CAS PubMed.
- A. Pal, R. Fagerberg, J. L. Andersen, C. Flamm, P. Dittrich and D. Merkle, J. Chem. Inf. Model., 2025, 65, 6772–6787 CrossRef CAS PubMed.
- P. Erdős and A. Rényi, Publ. Math. Debr., 1959, 6, 290 CrossRef.
- M. Newman, Networks, Oxford University Press, 2018 Search PubMed.
- D. J. Watts and S. H. Strogatz, Nature, 1998, 393, 440–442 CrossRef CAS PubMed.
- A.-L. Barabási and R. Albert, Science, 1999, 286, 509–512 CrossRef PubMed.
- J. Balogh, B. Bollobás and R. Morris, Eur. J. Commun., 2006, 27, 1263–1281 CrossRef.
- C. Eslahchi and A. M. Rahimi, Mat. Vesn., 2007, 59, 9–13 Search PubMed.
- F. Feng, X. He, Y. Liu, L. Nie and T.-S. Chua, Proceedings of the 2018 World Wide Web conference, Republic and Canton of Geneva, Switzerland, 2018, pp. 1523–1532 Search PubMed.
- W. Trotter, Combinatorics and Partially Ordered Sets: Dimension Theory, Johns Hopkins University Press, 2002 Search PubMed.
- M. Thakur and R. Tripathi, Theor. Comput. Sci., 2009, 410, 2592–2618 CrossRef.
- S. Bougueroua, A. A. Kolganov, C. Helain, C. Zens, D. Barth, E. A. Pidko and M.-P. Gaigeot, Phys. Chem. Chem. Phys., 2025, 27, 1298–1309 RSC.
- C. K. Ingold, Chem. Rev., 1934, 15, 225–274 CrossRef CAS.
- P. Cubillas, K. Etherington, M. W. Anderson and M. P. Attfield, CrystEngComm, 2014, 16, 9834–9841 RSC.
- T. Brinck, F. Sagan and M. Mitoraj, Eur. J. Inorg. Chem., 2025, 28, e202400845 CrossRef CAS.
- J. Cho, H. Y. Kang, L. V. Liu, R. Sarangi, E. I. Solomon and W. Nam, Chem. Sci., 2013, 4, 1502–1508 RSC.
- A. Karaçar, M. Freytag, H. Thönnessen, J. Omelanczuk, P. G. Jones, R. Bartsch and R. Schmutzler, Z. Anorg. Allg. Chem., 2000, 626, 2361–2372 CrossRef.
- A. Bernal, E. Llanos, W. Leal and G. Restrepo, Advances in Mathematical Chemistry and Applications, Bentham Science Publishers, 2015, pp. 24–54 Search PubMed.
- H. Li, M. Eddaoudi, M. O'Keeffe and O. M. Yaghi, Nature, 1999, 402, 276–279 Search PubMed.
- K. Tan, S. Zuluaga, E. Fuentes, E. C. Mattson, J.-F. Veyan, H. Wang, J. Li, T. Thonhauser and Y. J. Chabal, Nat. Commun., 2016, 7, 13871 Search PubMed.
- D. Rouvray and R. King, The Mathematics of the Periodic Table, Nova Science Publishers, 2006 Search PubMed.
- G. Restrepo and D. J. Klein, J. Math. Chem., 2011, 49, 1311–1321 Search PubMed.
- D. J. Klein and L. Bytautas, Commun. Math. Comput. Chem., 2000, 42, 261–290 CAS.
- G. Restrepo and P. F. Stadler, ACS Sustain. Chem. Eng., 2016, 4, 2191–2199 CrossRef CAS.
- B. A. Smith and K. D. Vogiatzis, J. Phys. Chem. A, 2021, 125, 7956–7966 CrossRef CAS PubMed.
- H. Ishikawa, K. Kwak, J. K. Chung, S. Kim and M. D. Fayer, Proc. Natl. Acad. Sci. U. S. A., 2008, 105, 8619–8624 CrossRef CAS PubMed.
- K. K. Liang and A. A. Villaeys, Phys. Rev. A, 2010, 82, 013423 CrossRef.
- J. L. Andersen, C. Flamm, D. Merkle, P. F. Stadler, Chemical Transformation Motifs - Modelling Pathways as Integer Hyperflows, arXiv, 2017, DOI:10.48550/arXiv.1712.02594.
- S. Datta and T. Limpanuparb, RSC Adv., 2021, 11, 20691–20700 RSC.
- R. Grande-Aztazi, E. Matito, J. M. Ugalde and J. M. Mercero, ChemPhysChem, 2025, 26, e202400713 CrossRef CAS PubMed.
- M. Thakur, R. Tripathi and FSTTCS, Foundations of Software Technology and Theoretical Computer Science, Berlin, Heidelberg, 2004, vol. 2005, pp. 481–493 Search PubMed.
- M. Bressan, J. Brinkmann, H. Dell, M. Roth and P. Wellnitz, The Complexity of Counting Small Sub-Hypergraphs, arXiv, 2025, DOI:10.48550/arXiv.2506.14081.
- G. Csáji, Theor. Comput. Sci., 2022, 931, 1–16 CrossRef.
- Z. Yang, W. Zhang, X. Lin, Y. Zhang and S. Li, HGMatch: A Match-by-Hyperedge Approach for Subgraph Matching on Hypergraphs, arXiv, 2023, DOI:10.48550/arXiv.2302.06119.
- Y. Su, Y. Gu, Z. Wang, Y. Zhang, J. Qin and G. Yu, IEEE Trans. Knowl. Data Eng., 2023, 35, 5808–5822 Search PubMed.
Footnotes |
| † To the memory of Rainer Brüggemann, pioneer on the theory and application of partially ordered sets to chemistry and environmental sciences. |
| ‡ A notation whose roots, as some historians have argued, can be traced back to Lavoisier at the end of the 18th century.6 |
| § The adjacency matrix is an m × m array indicating whether atom i of the m atoms in the molecule is directly bonded to atom j. A value of 1 in the (i, j) entry denotes the presence of a bond, and 0 its absence. Alternative schemes also exist in which bond orders, rather than simple presence/absence, are recorded. The incidence matrix, by contrast, is an m × n array encoding the relationship between atoms and bonds: an entry of 1 in row i, column j indicates that atom i is incident to (that is, participates in) bond j; otherwise the entry is 0. Further details on these matrix representations of molecular graphs may be found in ref. 23. |
| ¶ Other higher-order frameworks include simplicial complexes, which have also found chemical applications through topological data analysis.11,36–38 Further structures include tensors and higher-order Markov chains.39. |
| || In ref. 47 this model is referred to as the “substrate graph.” |
| ** In an email hypergraph, vertices correspond to users and a directed hyperedge links the sender to the set of recipients; the sending hypervertex has size 1, whereas the receiving hypervertex may have arbitrary size.52 |
| †† In this model the distinction between educt and product sets is suppressed, enabling analysis of the undirected connectivity backbone before reintroducing reaction directionality. |
| ‡‡ As shown in Fig. 3, binary hypergraphs can be obtained from directed hypergraphs by disregarding the order between hypervertices. Binary hypergraphs preserve the binary relations between hypervertices and may also be viewed as hypergraphs whose hyperedges are partitioned into two subsets of vertices. In this sense, they could be treated as bipartite hypergraphs. |
| §§ Ordered hypergraphs are often defined as hypergraphs endowed with a total order on their vertices.68,69 Here, following the spirit of,70 I consider hypervertices as partially ordered sets. |
| ¶¶ In formal terms a poset is defined as a set V endowed with an order relation ⪯. This later is a binary relation satisfying that for all x, y, z ∈ V: (1) x ⇒ x ⪯ x (reflexivity), (2) x ⪯ y and y ⪯ x ⇒ x = y (antisymmetry), and (3) x ⪯ y and y ⪯ z ⇒ x ⪯ z (transitivity).71 |
| |||| Interestingly, since the order relation between chemicals is given by the embedding of subgraphs of one chemical structure within another, vertices in hypervertices are not comparable, which also equates this chemical system with a directed hypergraph. See 83 for further details. |
| This journal is © The Royal Society of Chemistry 2026 |