Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Automated and robotic sample delivery systems for mass spectrometry and ion-mobility spectrometry
Chikondi
Shaba
,
Decibel P.
Elpa
and
Pawel L.
Urban
 *
*
Department of Chemistry, National Tsing Hua University, 101 Section 2, Kuang-Fu Rd, Hsinchu, 300044, Taiwan. E-mail: urban@mx.nthu.edu.tw
First published on 20th February 2026
Abstract
Mass spectrometry (MS) and ion-mobility spectrometry (IMS) are two complementary tools enabling atomic and molecular analysis. While MS provides mass-to-charge ratio values and fragmentation patterns facilitating molecular identification, IMS enables rapid separation of chemical species by size. The two techniques are often combined to benefit from the advantages offered by both. To match the needs of contemporary research and industrial activities, these techniques have been upgraded by integrating them with automated and robotic sample delivery systems. Automation eliminates or decreases human labor involved in sample handling. It also allows for high-throughput analysis, thus increasing productivity. This is especially important considering the capital cost of MS. The common approaches to automation involve the use of autosamplers, flow-injection analysis systems, microfluidics, and robotics. In this perspective, we highlight single-cell analysis as a prominent application area of automated MS methods.
Introduction
Automation has become a buzzword across STEM fields in the 21st century, reflecting both widespread interest and diverse interpretations. It is also a notable trend in modern chemical science and engineering. Interestingly, the word “automat” and its derivatives have different meanings around the world. In English, it signifies a type of self-service restaurants, which used to be popular in major US cities in the beginning of the 20th century,1 while in some other languages related terms refer, more generally, to other automatic devices such as vending machines, gambling machines, or even, colloquially, automatic transmissions or ATMs. On the other hand, the word “robot” is derived from the Czech noun “robota” (“forced labor”), and it first appeared in the play titled “Rossum's Universal Robots” by Karel Čapek from 1920.2 Later, in the 1939 New York World's Fair, the Westinghouse Electric Corporation presented a humanoid robot called “Elektro”, which could walk, talk, and even smoke cigarettes.3 Invented by George Devol, Unimate holds the distinction of being the earliest industrial robot ever created.4 It functioned as a hydraulic manipulator arm engineered to carry out repetitive manufacturing tasks. Automakers employed it to streamline operations like metalworking and welding. The International Union of Pure and Applied Chemistry defines automation as “mechanization with process control, where process means a sequence of manipulations”.5 In practice, however, the term is applied much more broadly in common usage. Further discussion provides an overview of efforts to integrate broadly defined automated and robotized systems with mainstream analytical platforms such as mass spectrometry (MS) and ion-mobility spectrometry (IMS). The examples and references highlighted here are intended to illustrate key developments, rather than to exhaustively cover the field.Mass spectrometry
MS is a key tool used for identification and quantification of elements and compounds.6–9 It enables ultrasensitive analysis of organic molecules.10 Mass spectrometers consist of three main blocks: ion source, mass analyzer, and detector. Considering a variety of potential applications, the ion source is the component that determines a mass spectrometer's flexibility and usability. Frequently used ion sources include: electron ionization, chemical ionization, atmospheric pressure chemical ionization, electrospray ionization (ESI), and matrix-assisted laser desorption/ionization (MALDI).7 Efficient formation of gaseous ions and their transmission to the mass analyzer is essential for achieving high sensitivity. For example, ESI enables transferring analytes from the liquid phase to the gas phase, generating charged species.11,12 Ion sources are often interfaced with other analytical devices to expedite the analysis of many samples and enable analysis of matrix-rich samples. Some MS interfaces are simplistic (e.g., a tubing connecting two components), while others are complex, and incorporate many elements (e.g., transmission lines, transducers, or elements of robotics).Ion-mobility spectrometry
Ion-mobility measurements were pioneered by John Zeleny in the end of the 19th century.13 He demonstrated that the velocity of atomic ions traveling through a gas-filled tube (utilizing air, oxygen, nitrogen, and carbon dioxide) under the influence of a weak electric field varied depending on both the type of gas and the ion.14 He noted that the observed velocity difference in two ions can be due to an inequality in the size of the two ions.13 Currently, there exist several types of IMS techniques: drift tube IMS (DT-IMS), travelling wave IMS (TWIMS), trapped IMS (TIMS), field asymmetric IMS (FAIMS), and differential mobility analyzer (DMA) IMS. DT-IMS—often regarded as the classic IMS model—is valued for its simplicity, straightforward operation, and capacity to directly measure ion mobility and calculate collision cross-section (CCS).15Although standalone IMS systems are very convenient and downscalable, combining IMS with MS has significantly expanded the resolution of chemical analysis.15 The complementary separation processes in both the ion mobility and mass dimensions offer remarkable improvements in selectivity and sensitivity.15 In the early days, ion mobility-mass spectrometry (IM-MS) instruments were primarily custom-built in academic laboratories. Wider adoption as a routine analytical tool began in 2006 with the launch of the Waters Synapt HDMS, the first widely available commercial IM-MS platform. Following the success of this commercial implementation, other instrument manufacturers quickly developed their own IM-MS systems, incorporating different ion mobility separation techniques to enhance the selectivity of MS, especially for analyzing complex mixtures.15 Most recently, two high-performance IMS platforms have been developed: cyclic IMS16 and structures for lossless ion manipulations (SLIM).17 These techniques are based on the general concept of TWIMS. They achieve high-resolution separations due to the very long ion migration pathlengths, which are in the order of tens of meters.18 Because some IMS separations can be accomplished on a millisecond timescale, they can also be easily integrated into traditional gas chromatography-mass spectrometry (GC-MS) and liquid chromatography-mass spectrometry (LC-MS) workflows.15 Ion mobility introduces an additional layer of separation to LC-MS-based untargeted metabolomics.19 The developments in IM-MS have enhanced metabolite annotation by providing an additional conditional molecular descriptor: CCS, thereby increasing confidence in identification.20 An ion-mobility CCS atlas has been created for metabolite annotation.21
Despite its potential, the complexity of ion-mobility-resolved metabolomics data poses significant challenges for processing, thus limiting its broader adoption. Advanced computational tools now facilitate the distinction of co-eluted metabolite isomers that exhibit subtle variations in chromatographic and ion-mobility dimensions.19 It is appealing to further develop coupling of IMS with sample preparation and separation systems, to enhance metabolomic analyses.
Our perspective
In this perspective, we discuss approaches to automation, highlighting single-cell analysis as a key area that increasingly benefits from automated workflows. Furthermore, we categorize two key areas that define how robotics are currently applied in modern online MS and IM-MS workflows: sample preparation and sample delivery. Here, robotic sample preparation systems refer to automated platforms that use robotics in sample preparation and subsequent delivery to MS or IM-MS, while robotic sample delivery systems refer to automated platforms that utilize robotics for sample handling and direct delivery to MS or IM-MS without multistep sample preparation. We highlight and discuss robotic systems for sample preparation and delivery, providing representative examples of their application in MS and IM-MS analytical workflows. Advancements in technologies such as artificial intelligence (AI) continue to drive the evolution of analytical methods. Although still in their early stages, other robotic applications—including those integrating AI—are also discussed later on.Automation
Automated flow injection-based techniques
Conventional direct infusion ESI-MS measurements are conducted by filling a glass syringe with the sample, fixing the syringe in a syringe pump holder, attaching sample flow line to the syringe tip, and starting the pump. However, this mode of operation requires many manual operations, is time consuming, and does not enable high-throughput sampling. Therefore, other ways of sample introduction are used to increase sample throughput, and—in some cases—collect analytes from surfaces. Flow-based analytical techniques have been pivotal in streamlining sample handling and delivery in MS workflows. The concept originated from flow injection analysis (FIA), which was introduced in the 1970s by Růžička and Hansen.22,23 FIA enabled the generation of reproducible transient signals by precise and controlled injection of a defined volume of the sample into a flowing carrier stream.22–24 While conceptually distinct from segmented flow analysis introduced by Skeggs in the 1950s—which used discrete sample zones for clinical assays—FIA laid the foundation for contemporary flow-based methods.25–27 Over time, its development can be described in three generations, reflecting the evolution from classical FIA to sequential injection analysis (SIA) and, subsequently, to bead injection-lab-on-valve (LOV) systems.24,28,29Flow techniques, such as FIA, SIA, or multisyringe flow injection analysis (MSFIA), permit controlled and reproducible handling of liquid samples.26,30 The flow techniques use valves, tubing, columns, phase separators, or even multiple syringes30 to manipulate the flow of liquids, merging segments of liquids,30 and extraction processes.26 They have been used as a strategy to automate microextraction techniques.26 SIA was also used for dispersive liquid–liquid microextraction (DLLME) on a commercial FIAlab 3500 system connected to a fiber-optic charge-coupled device for detection.31 Other modifications incorporated a syringe as a DLLME extraction vessel (in-syringe DLLME).32–35 Direct infusion sample delivery to ESI-MS can also be “automated” allowing for the control of sample concentration,36 optimization of sample flow rate,37,38 or switching modifier vapors in the ionization region.39
It has been argued, however, that the use of flow systems does not necessarily imply automation.30 Rather, automation is characterized by the utilization of a computerized control system that incorporates a feedback loop.30 Therefore, while the attempts discussed above may have addressed several analytical challenges, it is important to note that they may not be strictly classified as “automated” because they were either not computerized or lacked a feedback loop.30 Although classical FIA is not inherently automated, as it can be performed manually using simple pumps, modern implementations often utilize LC autosamplers and computer-controlled pumps to deliver sample plugs directly to the mass spectrometer. In these setups, FIA effectively becomes automated, enabling precise, reproducible, and high-throughput operation. This approach is commonly applied on existing LC-MS systems by bypassing the chromatographic column, allowing rapid sample delivery into the ion source.40–43 Coupling FIA to a drift tube IM-MS (DTIM-MS) has also been demonstrated using this approach.44 While it does not fully replace conventional LC-MS, FIA—combined with tandem MS—provides a practical, high-throughput alternative for screening applications.
Microfluidic systems for automated sample delivery
Microfluidics uses micrometer-scale channels to manipulate below nanoliter volumes of fluid.45,46 The fluid flow, whether pressure- or electrically driven, is highly controlled and typically characterized by laminar behavior.45,46 Microfluidics streamlines complex workflows by supporting multiplexed, automated control and sample processing within a single integrated system.47–49 The automated control is mostly achieved through the use of electronic circuits, particularly, transistor-based analog and digital circuit designs.50 The sample processing steps include droplet sorting and isolation, cell lysis, protein digestion, sample cleanup, and sample deposition. Microfluidic approaches, such as continuous-flow systems,51 microchip electrophoresis,52–54 centrifugal,51 digital,51,55,56 droplet,57,58 and paper59,60 microfluidics, have been coupled with MS. Direct coupling can be challenging due to differences in geometry, pressure, phase, wettability, and electric current.55,61 This was particularly evident in attempts to integrate continuous flow systems with MALDI and matrix-free laser desorption/ionization,62 as well as multi-phase or discrete flow systems with spray ionization techniques.55 Developments in droplet microfluidics have progressed significantly. Key focus areas include droplet generation and manipulation, integration into chip-based systems, improved resolution and sensitivity, applications in microscale and molecular technologies, and interfacing with MS.53–55,61Consequently, efforts have focused on developing interfaces that enable online analysis. The most common type is the continuous flow microfluidics system, which is pressure-driven,51 whereas microchip electrophoresis is also a continuous flow system, but the flow is driven by an electric field.51 Microchip electrophoresis evolved from the miniaturization of capillary electrophoresis (CE).52 Centrifugal microfluidics is primarily used for automated, controlled sample preparation by using centrifugal forces to move fluids radially outward from the center of a disc.51 These continuous flow regime microfluidics have been employed in microchip electrophoresis-IMS,63 chip-electrochromatography-IMS,64 and in centrifugal microfluidics integrated with probe ESI-MS to enable automated, time-controlled sample processing.65
Continuous-flow systems are commonly used in perfusion or separation-based experiments,65 as well as in sample preparation workflows66–68 and lab-on-chip devices.69,70 They facilitate the delivery of minute sample volumes to microarrays or MALDI plates, thereby enabling automated wofklows.71,72 Automation permitted tuning of culture conditions for real-time measurements by incorporating a culture chamber/chip, multiplexed chips, and a control device equipped with valves and software.48,49 Furthermore, chip-based and capillary-based microfluidics have been integrated to simplify the setup of nanospray desorption electrospray ionization (DESI) mass spectrometry imaging (MSI),73,74 and to streamline the interfacing and assembly of chip-based supercritical fluid chromatography (SFC) with MS75,76 and IMS.77
Compartmentalization involves physically isolating samples into discrete volumes, reducing interference and cross-contamination.46,78,79 This enables spatially confined reactions79 and parallelized assays to improve throughput.79 Compartmentalization is achieved through several methods, such as by closing channels with valves, typically implemented using multilayer elastic chips with control channels;79,80 by isolating samples in micro/nanowells;58,80 or by generating water-in-oil droplets.45,46,79–82 The Fluidigm C1 system is a microfluidic chip-based automated commercial system that uses a compartmentalized microfluidic approach. In the C1 system, single cells are captured and isolated into discrete reaction chambers within integrated fluidic circuits (IFCs).83 It is an electrically and pneumatically operated desktop instrument with built-in vacuum pump to position IFCs.83 While not interfaced with MS, it has been used to automate entire molecular biology workflows (e.g., cell capture, lysis) for single cell genomics.80,84,85
Microfluidic platforms interfaced with MS
Interfacing droplet microfluidics with MS was considered challenging because the sub-microliter monodisperse bubbles or droplets of a dispersed gas or liquid are created in a continuous flow of immiscible carrier/mobile liquid,45,82,86 on- or off-chip.72,82,86 The carrier/mobile liquid of the microdroplet-encapsulated analyte generally contains halogens and surfactants for droplet stability but these may cause instability of the Taylor cone and contamination of the mass spectrometer.82,86–88 Coupling strategies have involved isolating the droplets from the carrier flow prior to ionization and diverting the carrier flow to waste.86 For example, Fidalgo et al. used fluorescence screening to trigger application of voltage that merges the droplet with a sheath liquid, thereby isolating the droplet.87 However, this process causes dispersion of the droplet contents.87 Furthermore, Küster et al. automated the deposition of aqueous droplets onto microarrays of hydrophilic spots within hydrophobic coating by using an optical detection system for real-time differentiation of droplet deposition.57 In another variation, Gasilova et al. isolated individual droplets from segmented flow via an on-chip spyhole, applying high-voltage pulses beneath to enable electrostatic-spray ionization of aqueous droplets.89 This enabled direct coupling to MS with minimized sample dilution and improved analytical sensitivity.89 Alternatively, the segmented flow is simply directly infused into the MS,81,86,90 but with limitations of spray voltage and accumulation of carrier liquid at the spray tip.86 Both limitations can be addressed as explored by Belder's group91 and Kennedy's group.92–95 In another instance, the flexibility of a droplet microfluidic chip interface using a stainless steel ESI capillary was demonstrated by coupling to three commercially available mass spectrometers—Orbitrap, DTIM quadrupole time-of-flight (Q-TOF), and TWIMS Q-TOF mass spectrometers.88Unlike conventional droplet microfluidics, the so-called digital microfluidics (DMF) controls discrete droplets that are not confined in fluidic channels, using an array of patterned electrodes, and performs a finite set of operations.51,55,56 These operations include droplet generation, merging, reagent addition, and splitting, at ambient pressure,51,55,56 as well as coupling solid-phase microextraction (SPME) with high-performance liquid chromatography (HPLC)-MS.96 Similar to droplet microfluidics, there were challenges in online coupling of DMF with MS, particularly in the efficient transfer of droplets to the ESI source. This is because droplets on a DMF device are unconfined and at ambient pressure, requiring pressure-assisted delivery and, at times, additional sample processing before MS analysis.55,61 With advances over the years, DMF systems have been directly coupled to MS in various ways, such as DMF-MS interfaced by Venturi easy ambient sonic-spray ionization,97 DMF with MALDI-MS for multiplexed sample preparation,47 with ESI-MS interfaced by a microfluidic eductor,55 with HPLC-MS directly autosampling from a 3D-printed manifold,98 with nanoESI-MS interfaced by a specially fitted nanoESI emitter,99 and a folded polyimide nanoESI emitter.100 For example, Shih et al. demonstrated automated dried blood spot analysis using DMF with nanoESI-MS (Fig. 1I), employing a feedback-controlled system (Fig. 1II).99 The system enabled precise droplet handling and on-chip processing entirely without manual intervention, significantly advancing rapid and reliable bioanalytical workflows.99 A more recent approach for coupling DMF with MS eliminates the use of transfer capillaries and rather employs a microspray hole in the chip top plate.61
Automated platforms for single-cell analysis
The ability to analyze individual cells has long been regarded as a Holy Grail of analytical science. Single-cell omics investigates cellular heterogeneity from structural, functional, environmental, or genetic variations,101–107 and requires high sensitivity, high throughput, and a broad coverage of analytes.103,108 In this context, automation reduces operator variability, increases throughput and reproducibility, improves cell sorting and targeting, and addresses several limitations, thereby broadening the scope of analysis in single-cell workflows. These limitations include cell clustering and random positioning of cells in a flow, resulting in an overlap of output (peaks),53,108 preservation of the cells' native metabolic state,108 post-lysis degradation of analytes,108 and sample injection volume scaling.108,109High-throughput single-cell analysis
High-throughput approaches, such as MALDI and laser-ablation ESI (LAESI), as well as flow cytometry using droplet extraction combined with pulsed direct current ESI (pulsed-DC-ESI) have been adapted for single-cell analysis, enabling automated workflows.110–115 MALDI faces limitations due to specialized sample preparation requirements, including cell extraction and mixing the extract with the matrix, as well as its operation under vacuum conditions, both of which hinder its applicability for in situ studies.106,108 In contrast, LAESI allows for ambient ionization.105 However, early LAESI applications had limited spatial resolution108 and manual cell targeting,113 which have undergone significant improvements over the years. Earlier attempts to adapt MALDI, LAESI, and pulsed-DC-ESI for automated single-cell analysis included, respectively, the microarrays for mass spectrometry (MAMS) platform,102,116–119 a fiber-based LAESI ion mobility-mass spectrometry (f-LAESI-IM-MS) integrated with a feedback-controlled autofocus system,105 and an MS system integrated with real-time visual feedback and robotic micromanipulation.115 Application of MAMS involves aliquoting of cells into hydrophilic spots surrounded by a hydrophobic coating, which is followed by MALDI matrix application, and MALDI-MS scan. Automation of the f-LAESI-IM-MS enabled minimal sample preparation, preserved spatial information for metabolite mapping, supported robust statistical analysis of cellular heterogeneity, and increased throughput by approximately 13-fold compared to earlier semi-automated approaches.105 The automation by MAMS and f-LAESI-IM-MS requires specialized instrumentation and expertise that are financially and technically demanding, limiting accessibility.Challenges in achieving true single-cell resolution
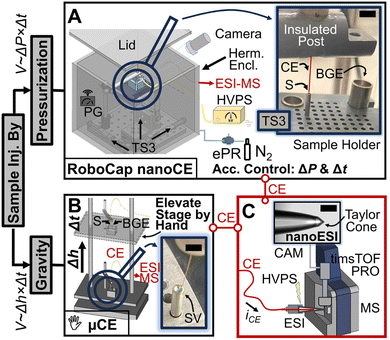
Despite the progress, the reported approaches challenge the fidelity of single-cell analysis, as defined by the term “single-cell”. One, multiple, or no cells may be sampled or deposited, yet the signals obtained are regarded as equivalent to single-cell data.102 Some approaches also fall short in terms of capability for live-cell sampling, time-lapse measurements, adequate sample volume, and repeated spot or cell sampling,105,113,120 resulting in difficulty determining repeatability and measurement certainty. Automated systems that incorporate such functions enable comparable metabolite coverage,105,115,116,121,122 supporting complex experimental designs and scalable analysis. These are capabilities which manual methods cannot reliably or practically achieve. However, despite these advancements—including the recent developments mentioned below and the microfluidic systems described earlier—such platforms still offer limited flexibility in the culture formats that can be sampled.The following recent developments represent a critical technological leap for single-cell MS and IMS analyses, enabling true single-cell sampling and precise, high-throughput, and physiologically relevant investigations of cellular heterogeneity. Recent years have seen significant advancements in automated, nanoliter volume, and scalable sample handling and delivery platforms specifically designed for single-cell MS and IMS. One such system is the robotic capillary (RoboCap) employed for automated nanoflow capillary electrophoresis electrospray ionization mass spectrometry (nanoCE-ESI-MS) (Fig. 2).109 RoboCap represents a pivotal development in single-cell analysis by automating CE-MS, particularly for very small sample volumes.109 This robotic platform eliminates manual handling bottlenecks that once limited throughput, enabling consistent and reproducible analysis with minimal human intervention.109 Additionally, automation enhances sample use efficiency compared to manual conventional µCE.109 Therefore, RoboCap paves the way for high-throughput single-cell analysis, addressing major challenges related to throughput, scalability, sample utilization, and robustness.109 To remotely control the system, RoboCap uses a custom-written software virtual instrument.109 In a recent improvement of MALDI-MSI, a Python program called microMS was used to automate coordinate transformation and laser targeting during fluorescence-guided sequential single-cell MS for cell-type-specific lipid classification and true single-cell analysis.112 In this automated workflow, microMS converts fluorescence microscopy pixel coordinates to physical stage coordinates of the MALDI system and—in the same action—removes cell clusters.112 This multimodal strategy, integrating lipidomics and proteomics on the same cells, together with precise automated targeting, significantly advances both throughput and resolution in single-cell MS.
Commercial systems have also been introduced such as the Single Cellome System 2000, which includes incubation controls that maintain optimal conditions during extraction,123 time-lapse analysis,124 and single-cell cloning, ensuring cell and culture integrity.125 Another commercial platform is the modular single-cell microfluidics sampling platform from iotaSciences126,127 which can sort and verify single cells,126 and automates the formation of liquid chambers, single-cell dispensing and visualization, and transferring selected cells into vials for LC-MS analysis.127
Automated chemical mapping of non-flat surfaces
Chemical mapping of real specimens with non-flat surfaces requires the use of adaptable sampling systems that can control the vertical position of the sampling probe establishing contact with the specimen surface. Therefore, the automation of surface sampling has focused on sensing height differences, especially for sampling uneven surfaces, to derive temporal information and spatial information for topographic molecular mapping, without causing sample damage or loss of microjunction. An earlier attempt made use of image-guided feedback control to automate the formation of a microjunction in surface sampling.128 However, this method was not capable of spot sampling, thereby limiting its application. The use of a robotic arm integrated with a distance sensor was one improved approach, it included a camera and a height calibrated laser pointer, then coupled to a water-assisted laser desorption ionization technology for MSI.129 More recently, an electrical conductance feedback system was integrated with a modified 3D printer, whereby the sampling probe was mounted on the printer head and its movements were controlled by a custom Python code.130 Repurposing a 3D printer is a cost-effective approach, and integrating it with a feedback loop is a significant step toward creating accessible, automated, and versatile analytical workflows.Automation hardware and software ecosystem
The Opentrons Python protocol API—an open-source framework—enables users to create or customize protocols for the Opentrons robots, automating liquid handling and precise coordination of steps.146 In a recent study, an automated sample preparation platform for analyzing proteins and protein modifications, called “AUTO-SP”, was written in Python and used to automate the key sample preparation procedures in the 2018 Clinical Proteomic Tumor Analysis Consortium protocol.147 AUTO-SP was used with the Opentrons OT-2 and Opentrons Flex robots for automation of liquid handling prior to MS analysis.147 Additional examples of the use of electronic modules are discussed in other sections, such as those covering digital microfluidics, while further examples related to robotics are provided in the subsequent Robotics section. Adoption of some of these approaches may not be immediate as they require varying levels of programming and hardware integration skills.
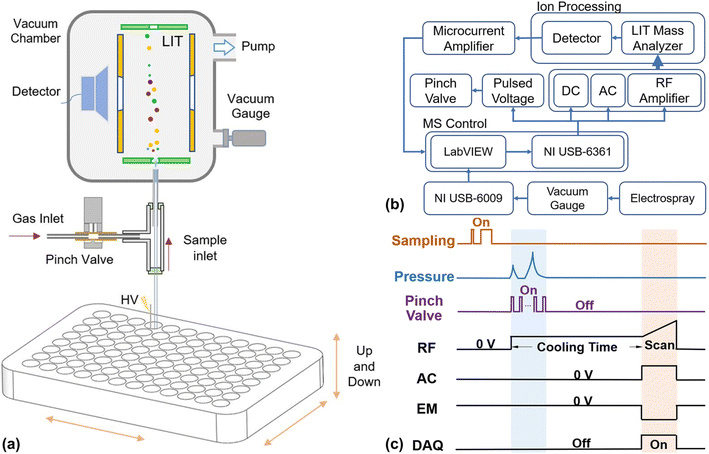
In contrast to open-source software, vendor-specific software is usually associated with licensing costs, as well as restricted accessibility and customizability. These limitations can hinder adaptation to changing methodological requirements. Integration across software platforms is a critical enabler of automation and high-throughput workflows. Several software platforms were incorporated using Analytical Studio,151 which automates data processing through a graphical interface rather than manual scripting.152 Together, these platforms created an integrated laboratory informatics and data analysis ecosystem. This ecosystem managed entire workflows, including robotic liquid handling using Tecan robots with RP-HPLC-MS and SFC-MS, to support automated analysis in drug discovery.151 Open Platform Communications Unified Architecture (OPC-UA) compliant software was integrated with LC instrumentation in an automated online sampling workflow for monoclonal antibody (mAb) bioreactors.153 OPC-UA facilitates reliable machine-to-machine communication.154,155 The workflow automated adjustment of process parameters, sampling, and measurement based on real-time data transmission and feedback control across multiple bioreactors.153 A custom software was developed from proprietary hardware and software modules for an induced self-aspiration vacuum electrospray ionization source (ISA-VESI) coupled to miniature ion trap MS (Fig. 4).156 These modules precisely synchronize vertical motion stage actuation with MS data acquisition, enabling pulsed sample injection through intermittent capillary–sample contact.156
LabVIEW is a system-design software and graphical programming environment that integrates with various hardware and software platforms.157–159 It supports a wide range of devices with real-time data acquisition, automated control, and seamless instrument communication network.157,160 LabVIEW offers significant advantages for controlling automated platforms (e.g., RoboCap109). Its graphical programming interface158,159 is user-friendly and less intimidating for beginners, making prototyping more accessible compared to text-based languages.158 It enhances workflow efficiency by reducing manual intervention, improving data quality, and ensuring compliance with regulatory standards.158 However, using LabVIEW requires software license, which incurs associated costs, but the costs can be minimized by controlling a DAQ device with a Python script. LabVIEW projects can also become complex at scale, requiring careful code organization for maintainability.158
Commercial autosampler systems are typically operated through vendor-specific software, often equipped with multiple application-specific, customizable configurations and preprogrammed workflows. The autosamplers are standalone, or coupled to an MS directly, and vary in functionality.161 The functionality features range from simple solution transfer to sample heating, agitation, and vial transportation, all of which streamline sample preparation and cleanup workflows. These automating systems can also be interfaced to MS through other sampling systems, e.g., droplet-liquid microjunction-surface sampling probe,162 laser ablation/liquid phase collection surface sampling,163 and sample delivery platforms such as TriVersa NanoMate,164–166 PAL 3,167 and SampleStream.168–170 Several ambient ionization sources, including direct analysis in real time,171–173 DESI,174 and paper spray,175 are now commercially available. These systems enable rapid and direct MS analysis of samples under ambient conditions with minimal or no sample preparation. Many commercial configurations are integrated with autosamplers or robotic stages, allowing automated, sequential analysis of multiple samples. Their software-controlled operation—managing parameters such as voltage, gas flow, and probe positioning—further enhances analytical reproducibility and throughput.172,176 Automation of paper spray ionization, which was first introduced in 2010 by Cooks, Ouyang and team,177–179 included the Prosolia paper spray disposable cartridges and autosampler,180–183 and then VeriSpray PaperSpray which can be coupled to Thermo Fisher Scientific triple quadrupole MS systems.184–189 The VeriSpray PaperSpray, like most other commercial autosamplers discussed in the Robotics section, uses vendor-specific software which may limit its extendability and customizability.
Robotics
The increasing demand for sensitive, precise, reproducible, and high-throughput sample analysis has driven significant advancements in MS190–192 and IM-MS.193–195 MS has become a widely adopted analytical tool for both routine analysis and advanced scientific research.190 Similarly, recent advancements have transformed IMS from a tool for detecting chemical warfare agents and explosives to a versatile instrument for analytical and bioanalytical applications.196 To keep pace with the evolving analytical requirements, robotic systems are increasingly integrated in online analytical workflows. Robotics improves both operational efficiency and reliability of analytical methods by minimizing manual intervention, reducing variability, and enabling continuous operation.Robotic sample preparation systems
Robotics was first introduced into the analytical laboratory settings in the 1980s, where it substantially contributed to the advancement of analytical chemistry.197–199 Its primary application at the time was the automation of sample preparation processes to improve throughput and overall analytical performance.197 Even today, the majority of robotic applications in analytical laboratories remain focused on sample preparation, as it continues to be a critical step in the analytical workflow. The sample preparation process involves procedures conducted on samples before detection by an analytical instrument.200,201 Generally, these laboratory procedures are repetitive and routine in nature, which make them particularly well-suited for automating analytical workflows across various analytical instruments. For instance, solid-phase extraction (SPE) is a widely-used technique for isolating target analytes from complex sample matrices. The steps involved in SPE are typically performed manually and are well-established in many laboratories, making the automation of these routine processes both practical and desirable. For example, Fleischer et al. demonstrated the automation of the entire SPE-GC-MS analysis of benzoic acids in water using a dual-arm robotic system.202 The transition from manual to automated SPE was made possible by the robotic system's ability to perform complex movements. In their work, each arm of the dual-arm robot featured seven servo-controlled joints, facilitating human-like motions to perform extraction, sample handling, and sample delivery to the GC-MS instrument.202Liquid-handling platforms for sample preparation
Cartesian robots are commonly integrated with liquid-handling platforms and are also extensively used to automate extraction techniques.150,203–207 Cartesian robotic arms operate along linear X, Y, and Z axes. This robotic arm configuration makes them ideal for automating repetitive procedures such as pipetting, liquid transfers, reagent addition, and cleanup.208 Given that chromatographic and spectroscopic techniques are readily interfaced with MS, Cartesian robotic systems—which are commonly used to automate online analysis for these techniques—are consequently widely employed in MS and IM-MS workflows.209–214The RapidFire is an SPE-based sample preparation robotic system,166,215 equipped with multiple pumps synchronized in time and function.216 It samples from a well-plate,166,215 and is directly coupled with MS and IM-MS.209,215 Although the RapidFire system has drawbacks such as fluidic line clogging and sample carryover from complex matrices, it remains a reliable platform widely used for high-throughput screening applications.166 More recently, a software platform called AutonoMS was developed for automated end-to-end IM-MS metabolomic fingerprinting.217 Although currently only integrated with RapidFire IM-MS, AutonoMS integrates software control layers of multiple specialized instruments, coordinating entire workflows from sample injection to data transfer and analysis with an automated trigger. Another SPE-based system—called Prospekt 2—integrates various modules, such as an automated cartridge exchange, for automated, online SPE coupled to other analytical instruments such as LC-MS.218 In addition to SPE, SPME is a well-established sample preparation technique known for its simplicity, solvent-free operation, and ability to preconcentrate analytes from various matrices.219,220 Since its development in the early 1990s,221,222 commercial autosamplers for SPME have become available.167,223,224 Although automation improves reproducibility, SPME is not inherently a high-throughput technique. Recent studies by Pawliszyn's research group have addressed this limitation by implementing automated multiextractions using different SPME geometries, such as fibers and blades, enabling high-throughput analysis in a 96-well plate format.225–229
Evosep Eno is a technology that streamlines LC workflows by integrating sample handling steps.230,231 Its key feature, the Evotip, simplifies sample preparation by integrating desalting with LC-MS sample introduction. Evotip also functions as a temporary storage device by immobilizing analytes, such as peptides, which can be stored and recovered later without loss.232 Sample loading on Evotips can be automated using platforms such as the cellenONE instrument,233,234 Agilent Assaymap Bravo,234 Opentrons OT-2,234,235 or Biomek I-series.234 The cell processing, peptide digestion, and other sample preparation steps are performed on a chip of the cellenONE,233,236 equipped with environmental controls.236 Samples are then either transferred to Evotips for cleanup233,236,237 or pooled by directly connecting the chip with the LC autosampler238–240 followed by MS analysis.233,236,237,239 Liquid extraction surface analysis (LESA)—developed by van Berkel and group241—conventionally uses the commercially available TriVersa NanoMate robotic system242–244 that is capable of X, Y, and Z positional movements. LESA has been directly coupled to IMS,245,246 MS,241,247–249 and IM-MS.250
Prototype sample preparation robotic systems
In addition to existing commercial robotic systems, prototyped robotic systems for sample preparation are emerging.198,251–254 Santos-Neto's research group, for instance, created an open-source, multipurpose Cartesian robotic platform designed for automated sample preparation and online coupling with LC or LC-MS.251,252 This prototype was developed to perform diverse liquid- and solid-phase microextraction techniques such as single-drop microextraction (SDME),252 hollow-fiber liquid phase microextraction,251 and microextraction by packed sorbent.253 A significant feature of their developed open-source robotic prototype is the integration of SDME with LC or LC-MS. This broadened the range of target compounds to include thermally labile analytes that are better suited for LC-MS. Unlike typical automated SDME setups that rely on expensive commercial robotic autosamplers designed primarily for GC or GC-MS, this prototype employed a lab-made Cartesian robot controlled by open-source electronics (e.g., Arduino). All steps of the SDME process—including syringe rinsing, filling, droplet exposure and withdrawal, and injection into the LC system—were fully automated by the lab-made robot.252 This system was subsequently improved with a multisyringe configuration for microextraction by packed sorbent.253 The setup utilized an Arduino microcontroller board to automate six parallel microextractions.253In robotic configurations where multiple samples are processed simultaneously, sample preparation throughput is significantly increased. However, although extractions or multistep sample preparations are performed simultaneously for multiple samples, MS detection is still performed sequentially, resulting in a bottleneck. This stage of the analytical workflow continues to present limitations, emphasizing the need for technical improvements to bridge the gap between high-throughput sample preparation with downstream MS analytical detection. To fully realize the benefits of simultaneous multi-sample processing—whether using commercial or prototype robotic systems—improvements in both extraction and detection are necessary. Integrating multiplexed injection or ionization systems, utilizing fast MS acquisition modes or automating MS acquisition are some approaches that may further reduce the impact of simultaneous sample preparation with sequential instrument detection.
Robotic sample delivery systems
Robotic systems provide precision and flexibility that are challenging to achieve manually. By minimizing operator-dependent variability, these platforms ensure consistent performance across various applications and facilitate the extension of analytical workflows beyond the laboratory. Although robotic arms are less commonly used for routine sample preparation due to their complexity and cost, they are particularly valuable in sample delivery applications where precise positioning, transfer, and reproducibility are crucial.198 Industrial robotic arms are more complex than automated liquid handlers, pipetting robots, or syringe-based systems, and they also require programming expertise.255 Laboratories often prefer straightforward, dedicated devices such as Cartesian robotic systems for high-throughput routine sample preparation.198,206,256 However, robotic arms excel in sample delivery when precise positioning, sample transfer, or integration with multiple instruments is necessary. Their articulated joints and enhanced freedom of motion enable them to handle various sample types, including solids and irregularly shaped materials, with high accuracy.256,257 Robotic arms can integrate seamlessly with analytical instruments such as mass spectrometers, facilitating consistent and automated sample transfer. This versatility enables specialized robotic approaches for liquid handling, solid materials, volatile compounds, and portable, in-field applications, as will be discussed further in this section.Robotic liquid handling workstations
Currently, the most common robotic systems found in analytical laboratories are liquid handling workstations. Examples of commercially available robotic platforms include Eppendorf epMotion,258 Opentrons OT-2 robot,259 CTC Analytics HTS PAL Autosampler,260 Hamilton Microlab Vantage robot,261 Waters Andrew+ pipetting robot,262 and Beckman Biomek workstations,263 among others. These systems are primarily employed to automate routine liquid-handling tasks such as pipetting, serial dilutions, reagent addition, and plate filling. Although setting up these systems requires a considerable investment, they provide significant value in high-throughput analyses, particularly in fields such as pharmaceutical development, clinical diagnostics, and large-scale biochemical studies, where precision, reproducibility, and efficiency are essential.210,264–268 Beyond routine liquid handling, these platforms can be integrated with other automated modules to enable fully automated analytical workflows. For example, liquid handling platforms can be combined with high-throughput analytical technologies such as acoustic droplet ejection (ADE). ADE uses focused acoustic energy to transfer nanoliter-scale droplets from a source plate without physical contact, providing precise, reproducible sample delivery while minimizing cross-contamination.269–271First demonstrated in the early 2000s by Ellson and coworkers for “moving liquids with sound” and then later commercialized, ADE established the foundation for automated, contactless liquid handling.269,270,272,273 Coupled with an open port interface (OPI), which continuously directs droplets into a mass spectrometer, this technology forms the Echo MS system, a commercially available platform for ADE–OPI–MS.267,274–276 Winter et al. demonstrated its first application for high-throughput drug discovery, screening over one million compounds.274 They highlighted that a critical factor to consider in fully automated ADE–OPI–MS workflows is the risk of clogging in the transfer capillary connecting the OPI to the mass spectrometer, which can occur due to salt deposition after thousands of analyses.274,277 To mitigate this, hardware modifications were implemented, including in-line capillary cleaning, which allows seamless switching between standard carrier liquid and washing liquid within the automated ADE–OPI–MS workflow.274,277
Although ADE–OPI–MS dramatically increases throughput for rapid, label-free analysis of complex biological samples, sample preparation remains a bottleneck, as manual pipetting, reagent addition, and enrichment are slow and variable. Given the high sampling rate of ADE–OPI–MS, an equally high-throughput sample preparation and liquid handling platform is required. Van Puyvelde et al. addressed this bottleneck by integrating a robotic liquid handling platform with ADE–OPI–MS for protein biomarker quantification.267 Because ADE–OPI–MS does not include sample cleanup or separation, the authors evaluated the feasibility of an automated immunocapture protocol. Using a Biomek i7 robotic liquid handler, they automated the immunocapture protocol to improve selectivity even without LC separation, achieving a 15-fold speed improvement over LC-MS across 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 peptide measurements.267
000 peptide measurements.267
Multi-jointed robotic arms
In addition to robotic liquid handling workstations that operate along three linear axes, multi-jointed robotic arms with higher degrees of freedom are increasingly integrated into analytical MS workflows. Cartesian, or gantry-style, robotic systems provide highly specialized precision liquid handling within a defined, rectilinear area.208 In contrast, articulated robotic arms offer more flexibility and a broader range of motion.208,278 Hence, the articulated robotic arms are integrated into the analytical workflow to perform more human-like repetitive tasks such as moving plates, transferring bulk liquids, or loading and unloading instruments—effectively complementing existing liquid handling platforms.208,278–280 Achieving full automation of an analytical workflow typically requires the coordinated integration of multiple robotic systems. Collaborative robots, or ‘cobots’, are industrial robotic arms designed to safely perform automated tasks with human operators in the same workspace.281Similar to ADE–OPI–MS, another technique that enables rapid MS analysis is infrared matrix-assisted laser desorption electrospray ionization mass spectrometry (IR-MALDESI-MS).279,282 Ultrahigh-speed IR-MALDESI-MS measurements with sampling frequencies of up to ∼22 Hz have been previously demonstrated.282 As discussed earlier, with high-throughput techniques such as ADE–OPI–MS and IR-MALDESI-MS, advancing sample handling capabilities is essential to optimize overall throughput and avoid bottlenecks in the analytical workflow. To address this challenge, Shanley et al. developed a plate transfer system using UR5e cobots from Universal Robots to support high-throughput IR-MALDESI-MS analysis.279 This system was estimated to have the capacity to screen approximately one million compounds in 6–7 working days.279
The plate transfer system (Fig. 5) featured a cobot arm mounted on a three-meter seventh-axis linear rail and equipped with a dual-actuator gripper fitted with custom fingers.279 Additional components included a high-capacity assay plate loading window, a regrip and barcode scanning station, the loading nest of the IR-MALDESI-MS, and a plate conveyor integrated with a plate management system. With this setup, the only human intervention required is the initial loading of assay plates into a carrier and entering plate and assay information into the software. All subsequent steps—including plate handling, motion scripting, and MS control—are carried out automatically by the system. Because plate retrieval is faster than plate scanning, the IR-MALDESI-MS spends nearly all of its operating time on data acquisition rather than waiting for plates, with the only downtime being the brief interval required for plate exchange. For a system of this scale, it is essential to characterize the accuracy and precision of movement, handling, and positioning—not only of the robotic arm but of all integrated modules—to ensure reliable sample transfer and minimize the risk of errors or misalignment.279
Fleischer's group also employed a cobot to handle and transport samples, demonstrating how cobots can be incorporated into high-throughput workflows.280 In their work, the UR5 cobot from Universal Robots was used to continuously load samples to the autosampler of an inductively coupled mass spectrometry (ICP-MS) instrument. Fleischer's group had earlier shown that sample preparation steps, including microwave digestion, could be automated for elemental analysis with ICP-MS.283,284 To bridge the gap between automated sample preparation and ICP-MS analysis, the authors have integrated an automated sample transportation and handling cobot to achieve full automation of the entire ICP-MS analytical workflow. Specifically, the cobot was programmed to transport screw-cap vessels to the capping station, open the sample vessel, and load it into the ICP-MS autosampler in a predefined sequence.280 Analyzing trace metals via a fully automated workflow using hardware constructed from metals could be a potential source of contamination. The authors performed a contamination test of their automated system and reported a minor contamination of iron, copper, and zinc in the low ppb-range.280
In addition to high-throughput screening, robotic arms have been directly integrated with ionization sources and mass spectrometers to facilitate the analysis of complex, non-planar, or in vivo surfaces.129,285–287 Some approaches employ fully automated operation of a sampling probe (i.e., needle285,286 or laser probe129) mounted on a robotic arm. The robotic system automatically maneuvers the probe to perform surface sampling and subsequent delivery to the MS inlet for ionization. These systems are applied in MSI and are beginning to find potential use in clinical settings. By contrast, in one application, robotic arms rely on partial automation combined with surgical or operator control. For example, the da Vinci Xi surgical system, a minimally invasive robotic surgery platform, has been integrated with the MasSpec Pen for in vivo molecular analysis of porcine tissues.287 The MasSpec Pen operates by dispensing a small, controlled droplet of solvent (commonly water) onto the tissue surface to extract biomolecules through solid–liquid extraction.288 After a few seconds of contact, the droplet containing extracted biomolecules is aspirated and transported via a transfer tube to the inlet of a mass spectrometer.287,288 To accommodate integration with the da Vinci Xi system, the design and dimensions of the original handheld MasSpec Pen were modified to function as a laparoscopic device attached to one of the robotic arms. The surgeon controlled the MasSpec Pen remotely through the robotic system's interface, allowing precise positioning and operation during in vivo tissue analysis and enabling seamless, real-time chemical profiling in a robotic-assisted porcine surgery model.
Robotic arms in prototype sample delivery systems
Prototype robotic systems offer flexible and adaptable alternatives for automating MS analytical workflows at a lower cost than industrial robotic systems. Our research group has prototyped robotic analytical systems for MS using low-cost robotic arms and open-source electronic modules. For instance, we have demonstrated a reaction-based MS assay without human intervention.289,290 The entire workflow—including sample recognition, aliquoting, incubation, ion source delivery, and initiation of data acquisition—was fully automated. The robotic-assisted MS analysis was initially developed using a single robotic arm and later improved with a second robotic arm to enable simultaneous processing of multiple samples.289,290 We have also recently developed robotized systems for sampling VOCs emanating from solid surfaces.291,292 A pen-shaped probe was attached to the robotic arm for contactless VOC sampling at multiple defined positions on a solid surface.291 The robotic sampling system was prototyped using inexpensive electronic modules to enable automated VOC sampling and subsequent delivery to the MS ion source. Specifically, the Arduino Uno R3 controlled the relays for switching the pump on and off, controlling the solenoid valve for nitrogen gas outflow, and triggering the mass spectrometer. This prototyped robotic sampling system—which uses a pen probe to aspirate VOCs—offers versatility that enables adaptation to various detection techniques. For instance, building on this robotic pen-probe sampling, our group has developed a computer vision-assisted variant.292 In this robotic VOC sampling system, we have incorporated modifications and improvements to automatically sample VOCs from a solid surface and subsequently transfer them to a tritium-based ion source of a DT-IMS.292 While the precision and accuracy of low-cost robotic arms are limited compared to industrial-grade robotic systems, their use provides a practical and accessible approach to automating MS workflows—particularly when the aim is not high throughput but streamlining the workflow through automation. Moreover, these systems serve as valuable platforms for training researchers, fostering the development of practical automation skills.Robotic systems for on-site sampling and analysis
Another important application of robotic systems is sampling and analysis in environments that are hazardous for human operators. Robots play a crucial role by enabling sample collection under conditions that pose significant risks, such as radioactive sites or environments with potentially dangerous and toxic VOC emissions that are inaccessible to humans. In such cases, deploying robotic systems for on-site sampling and analysis is particularly advantageous. For example, Hu's research group developed a ‘lab-on-robot’ portable MS system to facilitate on-site MS analysis of hazardous and toxic compounds.293,294 Their on-site MS system incorporated a switchable in-house developed robotic arm sampler, a quadruped robot, and a miniature mass spectrometer (Fig. 6). An earlier design primarily focused on VOC sampling using a needle trap device mounted on a robotic arm, with subsequent analysis using a portable GC-MS instrument (Fig. 6I). Their on-site robotic system was later improved with the addition of a switchable robotic arm sampler and a dual, switchable ionization source (Fig. 6II). This enhanced versatility enabled the analysis of diverse sample types—gases and aerosols delivered to the nESI ionization region, liquids pumped to the paper-capillary spray ionization emitter, and bulk ores processed into powders and introduced for paper-capillary spray ionization—all conducted unmanned and on-site. These advances demonstrate how integrating robotics with MS further extends the analytical capabilities of MS, highlighting the importance of combining creativity and automation with analytical skills.Artificial intelligence and its integration to automated and robotic systems
Automation may require extensive programming, and AI can assist by generating coding scripts. In some cases, the corresponding source code files are freely accessible. Moreover, a free and open-source set of software libraries and tools such as the Robot Operating System is available for developing a control system software.315,316 Automated intelligent robotic sample treatment platforms are also available, and AI has been used to customize functions and improve instrument performance. Recent advances in deep learning-based computer vision models, particularly the You Only Look Once (YOLO) series of real-time object detection models, have enabled vision-guided automation in laboratory systems. For instance, YOLOv8 was integrated with the Opentrons OT-2 for real-time feedback on errors and precise recognition of pipette tips and liquid volumes without manual intervention.317 Moreover, Zheng et al. developed an automated, intelligent microfluidic platform for microalgae species detection.300 YOLOv5 was used for data analysis, while a Raspberry Pi microcomputer controlled the user interface, integrating a low-cost portable USB microscope and a mini-motorized stage.300 While YOLOv5 is renowned for its speed and ease of use, YOLOv8 offers improved accuracy, multi-task capabilities, and broader flexibility, making it ideal for applications demanding high precision and advanced vision tasks.318
While augmented reality (AR) and virtual reality (VR) have been widely adopted in other disciplines, they are still considered emerging technologies in laboratory automation and analytical sciences. AR and VR are expected to further enhance laboratory automation and robotics by expanding their accessibility and functionality. These technologies are already being applied in laboratory education and training, where immersive simulations allow users to learn and interact with instruments in a virtual-assisted environment.337–340 For instance, the Hilton research group developed a VR digital twin of an HPLC system339 and, subsequently, a full digital twin laboratory,340 offering realistic and interactive experiences for both educational training and research. Beyond virtual environments, AR has been successfully applied in robotic-assisted surgery, improving precision and spatial awareness during complex procedures.341,342 Building on such advances, it is conceivable that AR and VR will soon play a greater role in analytical laboratories, enabling remote operation, immersive interaction, and improved safety. In the longer term, collaborative robots and humanoid systems are increasingly becoming integral to self-driving laboratories, guided by scientists from remote locations—a development that is gradually advancing toward practical realization and, although not yet realized, has the potential to become a routine part of scientific research.
Concluding remarks
Automation of analytical systems involving MS and IMS is not just trendy; it is a must, especially in applications that require high throughput and handling small samples, such as single-cell analysis. Simple “automation” can be achieved by taking advantage of the FIA system. This can be done by adapting a commercial HPLC system with an autosampler. Sample plugs can be introduced to the flow line—following a preset program—without further intervention of the analyst. Miniaturization has been instrumental in automating MS workflows. However, implementing microfluidic systems for sample handling is generally more cumbersome than conventional FIA, as it requires access to microfabrication infrastructure. Although some commercial microchips and fabrication services are available, these are generally expensive, thus beyond the reach of most analytical labs. Coupling multi-axis robotic systems with MS and IMS provides the highest degree of flexibility enabling direct interfacing of raw sample collection with sample preparation stages. However, such systems are highly sophisticated, and require much expertise during their setup and operation. The proliferation of AI platforms, as well as 3D printing, may likely lower the entry barrier for the prospective users of such systems. Drone technology has also skyrocketed in recent years, partly due to its extensive military uses.343 Attempts have already been made to implement IMS systems on lightweight drones.344,345 It is imaginable that unmanned vehicles—whether aerial, aquatic, or terrestrial—will soon carry miniaturized IMS or MS instruments to fulfill chemical surveillance tasks. AI tools will likely be used for data treatment in automated MS and IMS analyses as well as for the design and operation of the newly established automated systems.Author contributions
All the authors participated in the writing and editing process.Conflicts of interest
P. L. U. is co-inventor of the patented MAMS technology.Data availability
No primary research results, software, or code have been included, and no new data were generated or analyzed as part of this perspective article.Acknowledgements
We acknowledge the National Science and Technology Council, Taiwan (grant numbers 114-2113-M-007-019-MY3 and 114-2811-M-007-041).References
- C. Kim, A dining experience to remember: a brief history of the automat – History Associates Incorporated, https://www.historyassociates.com/automat/, accessed 19 May 2025 Search PubMed.
- Who did invent the word ‘robot’ and what does it mean? – Adelaide Robotics Academy, https://www.roboticsacademy.com.au/who-invented-the-word-robot-and-what-does-it-mean/, accessed 19 May 2025 Search PubMed.
- Jenny Norman & co, Elektro, The most famous robot of the 1930s: History of Information, https://historyofinformation.com/detail.php?entryid=3570, accessed 19 May 2025 Search PubMed.
- ROBOTS, https://robotsguide.com/robots/unimate, accessed 19 May 2025 Search PubMed.
- Automation, https://goldbook.iupac.org/terms/view/A00527, accessed 5 September 2025 Search PubMed.
- G. L. Glish and R. W. Vachet, The basics of mass spectrometry in the twenty-first century, Nat. Rev. Drug Discovery, 2003, 2, 140–150 CrossRef CAS PubMed.
- P. L. Urban, Y.-C. Chen and Y.-S. Wang, Time-resolved mass spectrometry, John Wiley & Sons, Inc., 2016 Search PubMed.
- P. L. Urban, Quantitative mass spectrometry: an overview, Philos. Trans. R. Soc. Math. Phys. Eng. Sci., 2016, 374, 20150382 Search PubMed.
- T. Van Acker, S. Theiner, E. Bolea-Fernandez, F. Vanhaecke and G. Koellensperger, Inductively coupled plasma mass spectrometry, Nat. Rev. Methods Primer, 2023, 3, 52 CrossRef CAS.
- M. Kandiah and P. L. Urban, Advances in ultrasensitive mass spectrometry of organic molecules, Chem. Soc. Rev., 2013, 42, 5299–5322 RSC.
- J. B. Fenn, M. Mann, C. K. Meng, S. F. Wong and C. M. Whitehouse, Electrospray ionization for mass spectrometry of large biomolecules, Science, 1989, 246, 64–71 CrossRef CAS PubMed.
- G. R. D. Prabhu, E. R. Williams, M. Wilm and P. L. Urban, Mass spectrometry using electrospray ionization, Nat. Rev. Methods Primer, 2023, 3, 23 CrossRef CAS.
- E. Christofi and P. Barran, Ion mobility mass spectrometry (IM-MS) for structural biology: insights gained by measuring mass, charge, and collision cross section, Chem. Rev., 2023, 123, 2902–2949 CrossRef CAS PubMed.
- J. Zeleny, VI. On the ratio of the velocities of the two ions produced in gases by Röntgen radiation; and on some related phenomena, Lond. Edinb. Dublin Philos. Mag. J. Sci., 1898,(46), 120–154 CrossRef.
- J. N. Dodds and E. S. Baker, Ion mobility spectrometry: fundamental concepts, instrumentation, applications, and the road ahead, J. Am. Soc. Mass Spectrom., 2019, 30, 2185–2195 CrossRef CAS PubMed.
- K. Giles, J. Ujma, J. Wildgoose, S. Pringle, K. Richardson, D. Langridge and M. Green, A cyclic ion mobility-mass spectrometry system, Anal. Chem., 2019, 91, 8564–8573 CrossRef CAS PubMed.
- R. Wojcik, G. Nagy, I. K. Attah, I. K. Webb, S. V. B. Garimella, K. K. Weitz, A. Hollerbach, M. E. Monroe, M. R. Ligare, F. F. Nielson, R. V. Norheim, R. S. Renslow, T. O. Metz, Y. M. Ibrahim and R. D. Smith, SLIM ultrahigh resolution ion mobility spectrometry separations of isotopologues and isotopomers reveal mobility shifts due to mass distribution changes, Anal. Chem., 2019, 91, 11952–11962 CrossRef CAS PubMed.
- C. N. Naylor and G. Nagy, Recent advances in high-resolution traveling wave-based ion mobility separations coupled to mass spectrometry, Mass Spectrom. Rev., 2025, 44, 581–598 CrossRef CAS PubMed.
- M. Luo, Y. Yin, Z. Zhou, H. Zhang, X. Chen, H. Wang and Z.-J. Zhu, A mass spectrum-oriented computational method for ion mobility-resolved untargeted metabolomics, Nat. Commun., 2023, 14, 1813 CrossRef CAS PubMed.
- T. Mairinger, T. J. Causon and S. Hann, The potential of ion mobility-mass spectrometry for non-targeted metabolomics, Curr. Opin. Chem. Biol., 2018, 42, 9–15 CrossRef CAS PubMed.
- Z. Zhou, M. Luo, X. Chen, Y. Yin, X. Xiong, R. Wang and Z.-J. Zhu, Ion mobility collision cross-section atlas for known and unknown metabolite annotation in untargeted metabolomics, Nat. Commun., 2020, 11, 4334 CrossRef CAS PubMed.
- J. Ṙu
![[z with combining breve]](https://www.rsc.org/images/entities/char_007a_0306.gif) i
i![[c with combining breve]](https://www.rsc.org/images/entities/char_0063_0306.gif) ka and E. H. Hansen, Flow injection analyses: part I. A new concept of fast continuous flow analysis, Anal. Chim. Acta, 1975, 78, 145–157 CrossRef.
ka and E. H. Hansen, Flow injection analyses: part I. A new concept of fast continuous flow analysis, Anal. Chim. Acta, 1975, 78, 145–157 CrossRef. - B. Horstkotte, M. Miró and P. Solich, Where are modern flow techniques heading to?, Anal. Bioanal. Chem., 2018, 410, 6361–6370 CrossRef CAS PubMed.
- E. H. Hansen and J. Wang, The three generations of flow injection analysis, Anal. Lett., 2004, 37, 345–359 CrossRef CAS.
- L. T. Skeggs Jr., An automatic method for colorimetric analysis, Am. J. Clin. Pathol., 1957, 28, 311–322 CrossRef CAS PubMed.
- V. Cerdà, J. Avivar and A. Cerdà, Laboratory automation based on flow techniques, Pure Appl. Chem., 2012, 84, 1983–1998 CrossRef.
- M. Trojanowicz and K. Kołacińska, Recent advances in flow injection analysis, Analyst, 2016, 141, 2085–2139 RSC.
- J. Ruzicka, Lab-on-valve: universal microflow analyzer based on sequential and bead injection, Analyst, 2000, 125, 1053–1060 RSC.
- J. Wang and E. H. Hansen, Sequential injection lab-on-valve: the third generation of flow injection analysis, TrAC Trends Anal. Chem., 2003, 22, 225–231 CrossRef CAS.
- M. Trojanowicz and M. Pyszynska, Flow-injection methods in water analysis—recent developments, Molecules, 2022, 27, 1410 CrossRef CAS PubMed.
- V. Andruch, C. C. Acebal, J. Škrlíková, H. Sklenářová, P. Solich, I. S. Balogh, F. Billes and L. Kocúrová, Automated on-line dispersive liquid-liquid microextraction based on a sequential injection system, Microchem. J., 2012, 100, 77–82 CrossRef CAS.
- M. B. Melwanki and M. R. Fuh, Dispersive liquid-liquid microextraction combined with semi-automated in-syringe back extraction as a new approach for the sample preparation of ionizable organic compounds prior to liquid chromatography, J. Chromatogr. A, 2008, 1198, 1–6 CrossRef PubMed.
- F. Maya, B. Horstkotte, J. M. Estela and V. Cerdà, Lab in a syringe: fully automated dispersive liquid-liquid microextraction with integrated spectrophotometric detection, Anal. Bioanal. Chem., 2012, 404, 909–917 CrossRef CAS PubMed.
- B. Horstkotte, M. Alexovič, F. Maya, C. M. Duarte, V. Andruch and V. Cerdá, Automatic determination of copper by in-syringe dispersive liquid-liquid microextraction of its bathocuproine-complex using long path-length spectrophotometric detection, Talanta, 2012, 99, 349–356 CrossRef CAS PubMed.
- M. A. Vargas-Muñoz, V. Cerdà, L. S. Cadavid-Rodríguez and E. Palacio, Automated method for volatile fatty acids determination in anaerobic processes using in-syringe magnetic stirring assisted dispersive liquid-liquid microextraction and gas chromatography with flame ionization detector, J. Chromatogr. A, 2021, 1643, 462034 CrossRef PubMed.
- P.-H. Liu and P. L. Urban, Plug-volume-modulated dilution generator for flask-free chemistry, Anal. Chem., 2016, 88, 11663–11669 CrossRef CAS PubMed.
- G. R. D. Prabhu, H. A. Witek and P. L. Urban, Programmable flow rate scanner for evaluating detector sensitivity regime, Sens. Actuators, B, 2019, 282, 992–998 CrossRef CAS.
- G. R. D. Prabhu, V. K. Ponnusamy, H. A. Witek and P. L. Urban, Sample flow rate scan in electrospray ionization mass spectrometry reveals alterations in protein charge state distribution, Anal. Chem., 2020, 92, 13042–13049 CrossRef CAS PubMed.
- Y.-R. Hwang, D. P. Elpa and P. L. Urban, FlexESI: an automated vapor-switching interface for enhanced flexibility and sensitivity in electrospray ionization, ACS Meas. Sci. Au, 2025, 5(6), 814–822 CrossRef CAS PubMed.
- S. C. Nanita and L. G. Kaldon, Emerging flow injection mass spectrometry methods for high-throughput quantitative analysis, Anal. Bioanal. Chem., 2016, 408, 23–33 CrossRef CAS PubMed.
- S. A. Berger, C. Grimm, J. Nyenhuis, S. E. Payer, I. Oroz-Guinea, J. H. Schrittwieser and W. Kroutil, Rapid, label-free screening of diverse biotransformations by flow-injection mass spectrometry, ChemBioChem, 2023, 24, e202300170 CrossRef CAS PubMed.
- X. Shi, J. Zhang, Z. Sun and J. You, The determination of mycotoxins using isotope-coded labeling and dispersive liquid–liquid microextraction by high throughput flow injection analysis coupled with tandem mass spectrometry, Microchem. J., 2024, 201, 110550 CrossRef CAS.
- P.-R. Bai, N. An, Y.-Z. Wang, Y.-Y. Chen, Q.-F. Zhu and Y.-Q. Feng, iTASO: a novel photosensitive probe for high-throughput and selective submetabolomic analysis via flow injection-mass spectrometry, Anal. Chem., 2025, 97, 1495–1499 CrossRef CAS PubMed.
- C. M. Nichols, J. C. May, S. D. Sherrod and J. A. McLean, Automated flow injection method for the high precision determination of drift tube ion mobility collision cross sections, Analyst, 2018, 143, 1556–1559 RSC.
- G. M. Whitesides, The origins and the future of microfluidics, Nature, 2006, 442, 368–373 CrossRef CAS PubMed.
- D. T. Chiu, A. J. DeMello, D. Di Carlo, P. S. Doyle, C. Hansen, R. M. Maceiczyk and R. C. R. Wootton, Small but perfectly formed? Successes, challenges, and opportunities for microfluidics in the chemical and biological sciences, Chem, 2017, 2, 201–223 CAS.
- H. Moon, A. R. Wheeler, R. L. Garrell, J. A. Loo and C. J. Kim, An integrated digital microfluidic chip for multiplexed proteomic sample preparation and analysis by MALDI-MS, Lab Chip, 2006, 6, 1213–1219 RSC.
- B. Schuster, M. Junkin, S. S. Kashaf, I. Romero-Calvo, K. Kirby, J. Matthews, C. R. Weber, A. Rzhetsky, K. P. White and S. Tay, Automated microfluidic platform for dynamic and combinatorial drug screening of tumor organoids, Nat. Commun., 2020, 11, 5271 CrossRef CAS PubMed.
- S. T. Seiler, G. L. Mantalas, J. Selberg, S. Cordero, S. Torres-Montoya, P. V. Baudin, V. T. Ly, F. Amend, L. Tran, R. N. Hoffman, M. Rolandi, R. E. Green, D. Haussler, S. R. Salama and M. Teodorescu, Modular automated microfluidic cell culture platform reduces glycolytic stress in cerebral cortex organoids, Sci. Rep., 2022, 12, 20173 CrossRef CAS PubMed.
- K. A. Gopinathan, A. Mishra, B. R. Mutlu, J. F. Edd and M. Toner, A microfluidic transistor for automatic control of liquids, Nature, 2023, 622, 735–741 CrossRef CAS PubMed.
- N. S. Ha, M. De Raad, L. Z. Han, A. Golini, C. J. Petzold and T. R. Northen, Faster, better, and cheaper: harnessing microfluidics and mass spectrometry for biotechnology, RSC Chem. Biol., 2021, 2, 1331–1351 RSC.
- A. Manz, D. J. Harrison, E. M. J. Verpoorte, J. C. Fettinger, A. Paulus, H. Lüdi and H. M. Widmer, Planar chips technology for miniaturization and integration of separation techniques into monitoring systems: Capillary electrophoresis on a chip, J. Chromatogr. A, 1992, 593, 253–258 CrossRef CAS.
- X. Li, S. Zhao, H. Hu and Y. M. Liu, A microchip electrophoresis-mass spectrometric platform with double cell lysis nano-electrodes for automated single cell analysis, J. Chromatogr. A, 2016, 1451, 156–163 CrossRef CAS PubMed.
- L. Yang, S. Wang, X. Li, W. Wang, F. Xu and C.-F. Ding, Microchip capillary electrophoresis-mass spectrometry for high-throughput simultaneous analysis of B-complex vitamins, J. Chromatogr. A, 2025, 1740, 465589 CrossRef CAS PubMed.
- C. A. Baker and M. G. Roper, Online coupling of digital microfluidic devices with mass spectrometry detection using an eductor with electrospray ionization, Anal. Chem., 2012, 84, 2955–2960 CrossRef CAS PubMed.
- N. Sathishkumar, J. G. Camacho Valenzuela, N. H. Le, A. K. C. Yong, M. A. Rossotti, J. Dahmer, A. A. Sklavounos, M. Plante, D. Brassard, L. Malic, A. N. Moraitis, R. Biga, I. El Idrissi, J. Tanha, J. Labrecque, T. Veres and A. R. Wheeler, A combined digital microfluidic test for assessing infection and immunity status for viral disease in saliva, Lab Chip, 2025, 25, 3197–3207 RSC.
- S. K. Küster, S. R. Fagerer, P. E. Verboket, K. Eyer, K. Jefimovs, R. Zenobi and P. S. Dittrich, Interfacing droplet microfluidics with matrix-assisted laser desorption/ionization mass spectrometry: label-free content analysis of single droplets, Anal. Chem., 2013, 85, 1285–1289 Search PubMed.
- J. C. Villarreal, K. Kow, B. Pham, A. Egatz-Gomez, T. R. Sandrin, P. D. Coleman and A. Ros, Intracellular amyloid-β detection from human brain sections using a microfluidic immunoassay in tandem with MALDI-MS, Anal. Chem., 2023, 95, 5522–5531 CrossRef CAS PubMed.
- S. Nishat, A. T. Jafry, A. W. Martinez and F. R. Awan, Paper-based microfluidics: simplified fabrication and assay methods, Sens. Actuators, B, 2021, 336, 129681 CrossRef CAS.
- S. Jackson, S. Lee and A. K. Badu-Tawiah, Automated immunoassay performed on a 3D microfluidic paper-based device for malaria detection by ambient mass spectrometry, Anal. Chem., 2022, 94, 5132–5139 Search PubMed.
- A. Das, C. Weise, M. Polack, R. D. Urban, B. Krafft, S. Hasan, H. Westphal, R. Warias, S. Schmidt, T. Gulder and D. Belder, On-the-fly mass spectrometry in digital microfluidics enabled by a microspray hole: toward multidimensional reaction monitoring in automated synthesis platforms, J. Am. Chem. Soc., 2022, 144, 10353–10360 CrossRef CAS PubMed.
- P. L. Urban, A. Amantonico and R. Zenobi, Lab-on-a-plate: Extending the functionality of MALDI-MS and LDI-MS targets, Mass Spectrom. Rev., 2011, 30, 435–478 CrossRef CAS PubMed.
- M. Masár, J. Hradski, M. Nováková, R. Szucs, M. Sabo and Š. Matejčík, Online coupling of microchip electrophoresis with ion mobility spectrometry for direct analysis of complex liquid samples, Sens. Actuators, B, 2020, 302, 127183 CrossRef.
- N. T. Hartner, C. R. Raddatz, C. Thoben, S. K. Piendl, S. Zimmermann and D. Belder, On-line coupling of chip-electrochromatography and ion mobility spectrometry, Anal. Chem., 2020, 92, 15129–15136 Search PubMed.
- M. Futami, H. Naito, S. Ninomiya, L. C. Chen, T. Iwano, K. Yoshimura and Y. Ukita, Automated sample preparation for electrospray ionization mass spectrometry based on CLOCK-controlled autonomous centrifugal microfluidics, Biomed. Microdevices, 2024, 26, 22 Search PubMed.
- C. Furlan, R. A. M. Dirks, P. C. Thomas, R. C. Jones, J. Wang, M. Lynch, H. Marks and M. Vermeulen, Miniaturised interaction proteomics on a microfluidic platform with ultra-low input requirements, Nat. Commun., 2019, 10, 1525 Search PubMed.
- S. T. Gebreyesus, A. A. Siyal, R. B. Kitata, E. S.-W. Chen, B. Enkhbayar, T. Angata, K.-I. Lin, Y.-J. Chen and H.-L. Tu, Streamlined single-cell proteomics by an integrated microfluidic chip and data-independent acquisition mass spectrometry, Nat. Commun., 2022, 13, 37 CrossRef CAS PubMed.
- S. Y. Han, B. J. Treves Brown, M. A. Higginson, P. Kaye, C. A. Sharrad and S. L. Heath, Development of an automated microfluidic system for actinide separation and analysis, J. Chromatogr. A, 2025, 1742, 465646 CrossRef CAS PubMed.
- A. Oedit, P. Vulto, R. Ramautar, P. W. Lindenburg and T. Hankemeier, Lab-on-a-Chip hyphenation with mass spectrometry: strategies for bioanalytical applications, Curr. Opin. Biotechnol., 2015, 31, 79–85 CrossRef CAS PubMed.
- Q. Xue, F. Foret, Y. M. Dunayevskiy, P. M. Zavracky, N. E. McGruer and B. L. Karger, Multichannel microchip electrospray mass spectrometry, Anal. Chem., 1997, 69, 426–430 CrossRef CAS PubMed.
- J. Lee, S. A. Soper and K. K. Murray, Development of an efficient on-chip digestion system for protein analysis using MALDI-TOF MS, Analyst, 2009, 134, 2426–2433 RSC.
- J. P. Smithers and M. A. Hayes, Interfacing microfluidics with information-rich detection systems for cells, bioparticles, and molecules, Anal. Bioanal. Chem., 2022, 414, 4575–4589 CrossRef CAS PubMed.
- L.-X. Jiang, M. Polack, X. Li, M. Yang, D. Belder and J. Laskin, A monolithic microfluidic probe for ambient mass spectrometry imaging of biological tissues, Lab Chip, 2023, 23, 4664–4673 RSC.
- L.-X. Jiang, X. Li, M. Polack, D. Belder and J. Laskin, High-spatial-resolution mass spectrometry imaging of biological tissues using a microfluidic probe, Nat. Protoc., 2026, 21, 18–36 CrossRef CAS PubMed.
- J. Schwieger, C. Weise and D. Belder, Novel pinhole emitter chip for micro supercritical fluid chromatography-mass spectrometry with integrated dilution-free fluidic back-pressure regulation, Anal. Chem., 2024, 96, 20107–20114 CrossRef CAS PubMed.
- C. Weise, M. Schirmer, M. Polack, A. Korell, H. Westphal, J. Schwieger, R. Warias, S. Zimmermann and D. Belder, Modular Chip-Based nanoSFC-MS for Ultrafast Separations, Anal. Chem., 2024, 96, 13888–13896 CAS.
- J. Schwieger, K. Welters, C. Thoben, A. Nitschke, S. Zimmermann and D. Belder, Miniaturized supercritical fluid chromatography coupled with ion mobility spectrometry: a chip-based platform for rapid chiral and complex mixture analysis, Anal. Chem., 2025, 97, 7954–7960 CrossRef CAS PubMed.
- T. A. Hakala, F. Bialas, Z. Toprakcioglu, B. Bräuer, K. N. Baumann, A. Levin, G. J. L. Bernardes, C. F. W. Becker and T. P. J. Knowles, Continuous flow reactors from microfluidic compartmentalization of enzymes within inorganic microparticles, ACS Appl. Mater. Interfaces, 2020, 12, 32951–32960 Search PubMed.
- K. F. Sonnen and C. A. Merten, Microfluidics as an emerging precision tool in developmental biology, Dev. Cell, 2019, 48, 293–311 CrossRef CAS PubMed.
- X. Luo, J.-Y. Chen, M. Ataei and A. Lee, Microfluidic compartmentalization platforms for single cell analysis, Biosensors, 2022, 12, 58 CrossRef CAS PubMed.
- M. Volný, J. Rolfs, B. Hakimi, P. Fryčák, T. Schneider, D. Liu, G. Yen, D. T. Chiu and F. Tureček, Nanoliter segmented-flow sampling mass spectrometry with online compartmentalization, Anal. Chem., 2014, 86, 3647–3652 CrossRef PubMed.
- T. Moragues, D. Arguijo, T. Beneyton, C. Modavi, K. Simutis, A. R. Abate, J. C. Baret, A. J. DeMello, D. Densmore and A. D. Griffiths, Droplet-based microfluidics, Nat. Rev. Methods Primer, 2023, 3, 32 CrossRef CAS.
- C1 system, https://fluidigm.my.salesforce.com/sfc/p/#700000009DAw/a/4u0000019jok/DX5TU5FkEvr4knHoRR5gcd1nnv0lukP5C94aI_vHKW0, accessed 22 October 2025 Search PubMed.
- D. M. DeLaughter, The use of the Fluidigm C1 for RNA expression analyses of single cells, Curr. Protoc. Mol. Biol., 2018, 122, e55 CrossRef PubMed.
- C. L. G. Lambert, G. van Mierlo, J. J. Bues, O. J. Guillaume-Gentil and B. Deplancke, Engineering next-generation microfluidic technologies for single-cell phenomics, Nat. Genet., 2025, 57, 1344–1356 CrossRef CAS PubMed.
- B. E. Murray, L. I. Penabad and R. T. Kennedy, Advances in coupling droplet microfluidics to mass spectrometry, Curr. Opin. Biotechnol., 2023, 82, 102962 CrossRef CAS PubMed.
- L. M. Fidalgo, G. Whyte, B. T. Ruotolo, J. L. P. Benesch, F. Stengel, C. Abell, C. V. Robinson and W. T. S. Huck, Coupling microdroplet microreactors with mass spectrometry: reading the contents of single droplets online, Angew. Chem., Int. Ed., 2009, 48, 3665–3668 CrossRef CAS PubMed.
- E. E. Kempa, C. A. Smith, X. Li, B. Bellina, K. Richardson, S. Pringle, J. L. Galman, N. J. Turner and P. E. Barran, Coupling droplet microfluidics with mass spectrometry for ultrahigh-throughput analysis of complex mixtures up to and above 30 Hz, Anal. Chem., 2020, 92, 12605–12612 Search PubMed.
- N. Gasilova, Q. Yu, L. Qiao and H. H. Girault, On-Chip spyhole mass spectrometry for droplet-based microfluidics, Angew. Chem., Int. Ed., 2014, 53, 4408–4412 CrossRef CAS PubMed.
- D. Liu, B. Hakimi, M. Volny, J. Rolfs, X. Chen, F. Turecek and D. T. Chiu, Controlled generation of double emulsions in air, Anal. Chem., 2013, 85, 6190–6194 Search PubMed.
- R. J. Beulig, R. Warias, J. J. Heiland, S. Ohla, K. Zeitler and D. Belder, A droplet-chip/mass spectrometry approach to study organic synthesis at nanoliter scale, Lab Chip, 2017, 17, 1996–2002 Search PubMed.
- D. J. Steyer and R. T. Kennedy, High-throughput nanoelectrospray ionization-mass spectrometry analysis of microfluidic droplet samples, Anal. Chem., 2019, 91, 6645–6651 Search PubMed.
- C. I. D'Amico, D. A. Polasky, D. J. Steyer, B. T. Ruotolo and R. T. Kennedy, Ion mobility-mass spectrometry coupled to droplet microfluidics for rapid protein structure analysis and drug discovery, Anal. Chem., 2022, 94, 13084–13091 Search PubMed.
- C. I. D'Amico, G. Robbins, I. Po, Z. Fang, T. R. Slaney, G. Tremml, L. Tao, B. T. Ruotolo and R. T. Kennedy, Screening clones for monoclonal antibody production using droplet microfluidics interfaced to electrospray ionization mass spectrometry, J. Am. Soc. Mass Spectrom., 2023, 34, 1117–1124 Search PubMed.
- S. S. Wells, I. J. Bain, A. C. Valenta, A. E. Lenhart, D. J. Steyer and R. T. Kennedy, Microdialysis coupled with droplet microfluidics and mass spectrometry for determination of neurotransmitters in vivo with high temporal resolution, Analyst, 2024, 149, 2328–2337 Search PubMed.
- K. Choi, E. Boyaci, J. Kim, B. Seale, L. Barrera-Arbelaez, J. Pawliszyn and A. R. Wheeler, A digital microfluidic interface between solid-phase microextraction and liquid chromatography-mass spectrometry, J. Chromatogr. A, 2016, 1444, 1–7 Search PubMed.
- J.-B. Hu, T. R. Chen, C.-H. Chang, J.-Y. Cheng, Y.-C. Chen and P. L. Urban, A compact 3D-printed interface for coupling open digital microchips with Venturi easy ambient sonic-spray ionization mass spectrometry, Analyst, 2015, 140, 1495–1501 Search PubMed.
- C. Liu, K. Choi, Y. Kang, J. Kim, C. Fobel, B. Seale, J. L. Campbell, T. R. Covey and A. R. Wheeler, Direct interface between digital microfluidics and high performance liquid chromatography-mass spectrometry, Anal. Chem., 2015, 87, 11967–11972 Search PubMed.
- S. C. C. Shih, H. Yang, M. J. Jebrail, R. Fobel, N. McIntosh, O. Y. Al-Dirbashi, P. Chakraborty and A. R. Wheeler, Dried blood spot analysis by digital microfluidics coupled to nanoelectrospray ionization mass spectrometry, Anal. Chem., 2012, 84, 3731–3738 Search PubMed.
- A. E. Kirby and A. R. Wheeler, Microfluidic origami: a new device format for in-line reaction monitoring by nanoelectrospray ionization mass spectrometry, Lab Chip, 2013, 13, 2533–2540 RSC.
- S. S. Rubakhin, E. V. Romanova, P. Nemes and J. V. Sweedler, Profiling metabolites and peptides in single cells, Nat. Methods, 2011, 8, S20–S29 CrossRef CAS PubMed.
- A. J. Ibáñez, S. R. Fagerer, A. M. Schmidt, P. L. Urban, K. Jefimovs, P. Geiger, R. Dechant, M. Heinemann and R. Zenobi, Mass spectrometry-based metabolomics of single yeast cells, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 8790–8794 CrossRef PubMed.
- B. B. Misra, S. M. Assmann and S. Chen, Plant single-cell and single-cell-type metabolomics, Trends Plant Sci., 2014, 19, 637–646 Search PubMed.
- A. Ali, S. Davidson, E. Fraenkel, I. Gilmore, T. Hankemeier, J. A. Kirwan, A. N. Lane, I. Lanekoff, M. Larion, L. I. McCall, M. Murphy, J. V. Sweedler and C. Zhu, Single cell metabolism: current and future trends, Metabolomics, 2022, 18, 77 Search PubMed.
- M. Dolatmoradi, S. A. Stopka, C. Corning, G. Stacey and A. Vertes, High-throughput f-LAESI-IMS-MS for mapping biological nitrogen fixation one cell at a time, Anal. Chem., 2023, 95, 17741–17749 CrossRef CAS PubMed.
- Q. Liu, S. Martínez-Jarquín and R. Zenobi, Recent advances in single-cell metabolomics based on mass spectrometry, CCS Chem., 2023, 5, 310–324 CrossRef CAS.
- R. Zenobi, Single-cell metabolomics: analytical and biological perspectives, Science, 2013, 342, 1243259 CrossRef CAS PubMed.
- A. Amantonico, P. L. Urban and R. Zenobi, Analytical techniques for single-cell metabolomics: State of the art and trends, Anal. Bioanal. Chem., 2010, 398, 2493–2504 CrossRef CAS PubMed.
- D. Jia and P. Nemes, Development and validation of RoboCap, a robotic capillary platform to automate capillary electrophoresis mass spectrometry en route to high-throughput single-cell proteomics, Anal. Chem., 2024, 96, 16985–16993 CrossRef CAS PubMed.
- J. C. McKinnon, R. Balez, R. S. E. Young, M. L. Brown, J. S. Lum, L. Robinson, M. E. Belov, L. Ooi, S. Tortorella, T. W. Mitchell and S. R. Ellis, MALDI-2-enabled oversampling for the mass spectrometry imaging of metabolites at single-cell resolution, J. Am. Soc. Mass Spectrom., 2024, 35, 2729–2742 CrossRef CAS PubMed.
- K. K. Krestensen, T. F. E. Hendriks, A. Grgic, M. Derweduwe, F. De Smet, R. M. A. Heeren and E. Cuypers, Molecular profiling of glioblastoma patient-derived single cells using combined MALDI-MSI and MALDI-IHC, Anal. Chem., 2025, 97, 3846–3854 CrossRef CAS PubMed.
- M. Asadian, S. W. Croslow, T. J. Trinklein, S. S. Rubakhin, F. Lam and J. V. Sweedler, High-throughput fluorescence-guided sequential single-cell MALDI-ICC mass spectrometry, Anal. Chem., 2025, 97, 15864–15872 CrossRef CAS PubMed.
- L. Z. Samarah, R. Khattar, T. H. Tran, S. A. Stopka, C. A. Brantner, P. Parlanti, D. Veličković, J. B. Shaw, B. J. Agtuca, G. Stacey, L. Paša-Tolić, N. Tolić, C. R. Anderton and A. Vertes, Single-cell metabolic profiling: metabolite formulas from isotopic fine structures in heterogeneous plant cell populations, Anal. Chem., 2020, 92, 7289–7298 CrossRef CAS PubMed.
- M. J. Taylor, A. Liyu, A. Vertes and C. R. Anderton, Ambient single-cell analysis and native tissue imaging using laser-ablation electrospray ionization mass spectrometry with increased spatial resolution, J. Am. Soc. Mass Spectrom., 2021, 32, 2490–2494 CrossRef CAS PubMed.
- A. Chen, M. Yan, J. Feng, L. Bi, L. Chen, S. Hu, H. Hong, L. Shi, G. Li, B. Jin, X. Zhang and L. Wen, Single cell mass spectrometry with a robotic micromanipulation system for cell metabolite analysis, IEEE Trans. Biomed. Eng., 2022, 69, 325–333 Search PubMed.
- P. L. Urban, K. Jefimovs, A. Amantonico, S. R. Fagerer, T. Schmid, S. Mädler, J. Puigmarti-Luis, N. Goedecke and R. Zenobi, High-density micro-arrays for mass spectrometry, Lab Chip, 2010, 10, 3206–3209 RSC.
- Micro-array for mass spectrometry (MAMS) for single cell analysis, https://www.sigmaaldrich.com/TW/zh/product/sial/50757?srsltid=AfmBOoqXpgs5MnvaOFRkHHPL7O2eiip72E7uZFFz9xIUHAU2PyYWB-Mw, accessed 3 October 2025 Search PubMed.
- P. L. Urban, A. M. Schmidt, S. R. Fagerer, A. Amantonico, A. Ibañez, K. Jefimovs, M. Heinemann and R. Zenobi, Carbon-13 labelling strategy for studying the ATP metabolism in individual yeast cells by micro-arrays for mass spectrometry, Mol. Biosyst., 2011, 7, 2837–2840 CrossRef CAS PubMed.
- S. R. Fagerer, T. Schmid, A. J. Ibáñez, M. Pabst, R. Steinhoff, K. Jefimovs, P. L. Urban and R. Zenobi, Analysis of single algal cells by combining mass spectrometry with raman and fluorescence mapping, Analyst, 2013, 138, 6732–6736 RSC.
- L. Zhang, D. P. Foreman, P. A. Grant, B. Shrestha, S. A. Moody, F. Villiers, J. M. Kwak and A. Vertes, In Situ metabolic analysis of single plant cells by capillary microsampling and electrospray ionization mass spectrometry with ion mobility separation, Analyst, 2014, 139, 5079–5085 RSC.
- D. Zhang and L. Qiao, Microfluidics coupled mass spectrometry for single cell multi-omics, Small Methods, 2024, 8, e2301179 CrossRef PubMed.
- G. Abdi, M. Jain, R. Bhardwaj, K. Thakkar, I. Dewangan, R. Dhariwal and M. A. Tarighat, in Advances in metabolomics, ed. V. Singh, Springer Nature, Singapore, 2024, pp. 49–89 Search PubMed.
- A. Kontiza, J. von Gerichten, M. Spick, E. Fraser, C. Costa, K. D. G. Saunders, A. D. Whetton, C. F. Newman and M. J. Bailey, Single-cell lipidomics: protocol development for reliable cellular profiling using capillary sampling, Analyst, 2025, 150, 1261–1270 RSC.
- J. von Gerichten, K. D. G. Saunders, A. Kontiza, C. F. Newman, G. Mayson, D. J. V. Beste, E. Velliou, A. D. Whetton and M. J. Bailey, Single-Cell untargeted lipidomics using liquid chromatography and data-dependent acquisition after live cell selection, Anal. Chem., 2024, 96, 6922–6929 CrossRef CAS PubMed.
- Single Cellome System, 2000, https://www.yokogawa.com/eu/solutions/products-and-services/life-science/single-cellome/ss2000/#Overview, accessed 7 August 2025 Search PubMed.
- Iota Sciences, Gentle single-cell picking simplified, https://iotasciences.com/picking-platform/, accessed 11 September 2025 Search PubMed.
- A. Kontiza, J. von Gerichten, K. D. G. Saunders, M. Spick, A. D. Whetton, C. F. Newman and M. J. Bailey, Single-cell lipidomics: an automated and accessible microfluidic workflow validated by capillary sampling, Anal. Chem., 2024, 96, 17594–17601 CrossRef CAS PubMed.
- V. Kertesz, M. J. Ford and G. J. Van Berkel, Automation of a surface sampling probe/electrospray mass spectrometry system, Anal. Chem., 2005, 77, 7183–7189 CrossRef CAS PubMed.
- N. Ogrinc, A. Kruszewski, P. Chaillou, P. Saudemont, C. Lagadec, M. Salzet, C. Duriez and I. Fournier, Robot-assisted SpiderMass for in vivo real-time topography mass spectrometry imaging, Anal. Chem., 2021, 93, 14383–14391 CrossRef CAS PubMed.
- M. Hermann, H. Metwally, J. Yu, R. Smith, H. Tomm, M. Kaufmann, K. Y. M. Ren, C. Liu, Y. LeBlanc, T. R. Covey, A. C. Ross and R. D. Oleschuk, 3D printer platform and conductance feedback loop for automated imaging of uneven surfaces by liquid microjunction-surface sampling probe mass spectrometry, Rapid Commun. Mass Spectrom., 2025, 39, e9492 CrossRef CAS PubMed.
- P. L. Urban, Open-source electronics as a technological aid in chemical education, J. Chem. Educ., 2014, 91, 751–752 CrossRef CAS.
- P. L. Urban, Universal electronics for miniature and automated chemical assays, Analyst, 2015, 140, 963–975 RSC.
- P. Urban, Self-built labware stimulates creativity, Nature, 2016, 532, 313 CrossRef CAS PubMed.
- G. R. D. Prabhu and P. L. Urban, The dawn of unmanned analytical laboratories, TrAC Trends Anal. Chem., 2017, 88, 41–52 CrossRef CAS.
- P. L. Urban, Prototyping instruments for the chemical laboratory using inexpensive electronic modules, Angew. Chem., Int. Ed., 2018, 57, 11074–11077 CrossRef CAS PubMed.
- G. R. D. Prabhu, T.-H. Yang, C. Y. Hsu, C.-P. Shih, C.-M. Chang, P.-H. Liao, H.-T. Ni and P. L. Urban, Facilitating chemical and biochemical experiments with electronic microcontrollers and single-board computers, Nat. Protoc., 2020, 15, 925–990 CrossRef CAS PubMed.
- G. R. D. Prabhu and P. L. Urban, Elevating chemistry research with a modern electronics toolkit, Chem. Rev., 2020, 120, 9482–9553 CrossRef CAS PubMed.
- P. L. Urban, Practical electronics and robotics for chemists, Nat. Chem., 2025, 17, 1 CrossRef CAS PubMed.
- J.-B. Hu, S.-Y. Chen, J.-T. Wu, Y.-C. Chen and P. L. Urban, Automated system for extraction and instantaneous analysis of millimeter-sized samples, RSC Adv., 2014, 4, 10693–10701 RSC.
- H.-C. Yang, C.-M. Chang and P. L. Urban, Automation of fizzy extraction enabled by inexpensive open-source modules, Heliyon, 2019, 5, e01639 CrossRef PubMed.
- C.-H. Chang and P. L. Urban, Automated dual-chamber sampling system to follow dynamics of volatile organic compounds emitted by biological specimens, Anal. Chem., 2018, 90, 13848–13854 CrossRef CAS PubMed.
- K.-T. Hsieh, P.-H. Liu and P. L. Urban, Automated on-line liquid–liquid extraction system for temporal mass spectrometric analysis of dynamic samples, Anal. Chim. Acta, 2015, 894, 35–43 Search PubMed.
- N. S. García-Rojas, H. Guillén-Alonso, S. MacKay, C. Torres-Calzada, L. D. Soto-Rodriguez, R. Winkler and D. S. Wishart, The open SprayBot: a high-throughput paper spray mass spectrometry platform for disease screening, HardwareX, 2024, 19, e00551 CrossRef PubMed.
- H. Ting, J.-B. Hu, K.-T. Hsieh and P. L. Urban, A pinch-valve interface for automated sampling and monitoring of dynamic processes by gas chromatography-mass spectrometry, Anal. Methods, 2014, 6, 4652–4660 RSC.
- C.-H. Chen, G. R. D. Prabhu, K.-C. Yu, D. P. Elpa and P. L. Urban, Portable fizzy extraction ion-mobility spectrometry system, Anal. Chim. Acta, 2022, 1204, 339699 CrossRef CAS PubMed.
- Python Protocol API v2, https://docs.opentrons.com/v2/index.html, accessed 21 October 2025 Search PubMed.
- T. M. Lih, L. Jiao, L. Chen, J. Woo, Y. Wang and H. Zhang, AUTO-SP: automated sample preparation for analyzing proteins and protein modifications, Anal. Chem., 2025, 97(31), 16751–16758 CrossRef CAS PubMed.
- HighRes Biosolutions, Advanced lab automation solutions, https://highresbio.com, accessed 11 September 2025 Search PubMed.
- H. E. Waldenmaier, E. Gorre, M. L. Poltash, H. P. Gunawardena, X. A. Zhai, J. Li, B. Zhai, E. J. Beil, J. C. Terzo, R. Lawler, A. M. English, M. Bern, A. D. Mahan, E. Carlson and H. Nanda, “Lab of the future” ─ today: fully automated system for high-throughput mass spectrometry analysis of biotherapeutics, J. Am. Soc. Mass Spectrom., 2023, 34, 1073–1085 Search PubMed.
- M. Wu, Q. Ma, M. Li, Z. Zhou, J. Xu, G. I. N. Waterhouse, N. Song, W.-W. Zhao and G. Chen, Online sequential analysis of volatile and semivolatile organic compounds in water matrices by double robotic sample preparations and dual-channel mono and comprehensive two-dimensional gas chromatography coupled with time-of-flight mass spectrometry system, J. Chromatogr. A, 2024, 1726, 464963 Search PubMed.
- K. Chen, J. L. Dores-Sousa, A. Fontana, C. Grosanu, H. M. McAllister, G. Bai, K. Bartkowiak, S. Cañellas, D. Corens, A. De Groot, J. Nevarez, M. Serrano, K. Raeymaekers, R. Rodriguez, L. Van Eynde, R. Zhou and Z. Shi, Automated high-throughput RP-HPLC-MS and SFC-MS analytical and purification platforms to support drug discovery, J. Chromatogr. A, 2025, 1742, 465648 Search PubMed.
- Analytical Studio – Virscidian, https://www.virscidian.com/analytical-studio, accessed 22 October 2025 Search PubMed.
- Z. D. Dunn, P. Bohman, A. Quinteros, B. Sauerborn, F. Milman, M. Patel, R. Kargupta, S. Wu, M. Hornshaw, R. Barrientos, J. Bones, V. S. Tayi, N. Abaroa, B. Patel, E. Appiah-Amponsah and E. L. Regalado, Automated online-sampling multidimensional liquid chromatography with feedback-control capability as a framework for real-time monitoring of mAb critical quality attributes in multiple bioreactors, Anal. Chem., 2023, 95, 18130–18138 Search PubMed.
- M. Krohn, What is OPC UA?, https://www.opc-router.com/what-is-opc-ua/, accessed 16 September 2025 Search PubMed.
- OPC UA explained, https://maplesystems.com/industrial-internet-of-things-iiot/opcua/, accessed 16 September 2025 Search PubMed.
- Y. Zhu, Q. Zhang, J. Lu, K. Wang, R. Zhang and Q. Yu, High-throughput screening using a synchronized pulsed self-aspiration vacuum electrospray ionization miniature mass spectrometer, Anal. Chem., 2022, 94, 7417–7424 Search PubMed.
- What is NI LabVIEW?, https://www.ni.com/en/shop/labview.html, accessed 6 November 2025 Search PubMed.
- LabVIEW in the life science industry, https://www.aristo-group.com/en/guide/labview-in-the-life-science-industry/, accessed 21 October 2025 Search PubMed.
- Exploring the advantages of LabVIEW, https://www.linkedin.com/pulse/exploring-advantages-labview-versatile-platform, accessed 21 October 2025 Search PubMed.
- NI LabVIEW empowers scientific research at CERN's antimatter lab, https://www.ni.com/en/solutions/academic-research/case-studies/labview-automation-empowers-cerns-antimatter-research.html, accessed 6 November 2025 Search PubMed.
- M. Alexovič, P. L. Urban, H. Tabani and J. Sabo, Recent advances in robotic protein sample preparation for clinical analysis and other biomedical applications, Clin. Chim. Acta, 2020, 507, 104–116 CrossRef PubMed.
- V. P. Sica, H. A. Raja, T. El-Elimat, V. Kertesz, G. J. Van Berkel, C. J. Pearce and N. H. Oberlies, Dereplicating and spatial mapping of secondary metabolites from fungal cultures in situ, J. Nat. Prod., 2015, 78, 1926–1936 Search PubMed.
- O. S. Ovchinnikova, V. Kertesz and G. J. Van Berkel, Combining laser ablation/liquid phase collection surface sampling and high-performance liquid chromatography-electrospray ionization-mass spectrometry, Anal. Chem., 2011, 83, 1874–1878 Search PubMed.
- C. D. Wenger, M. T. Boyne, J. T. Ferguson, D. E. Robinson and N. L. Kelleher, Versatile online-offline engine for automated acquisition of high-resolution tandem mass spectra, Anal. Chem., 2008, 80, 8055–8063 Search PubMed.
- R. K. Scheerle, J. Graßmann and T. Letzel, Enzymatic conversion continuously monitored with a robotic nanoESI-MS tool: experimental status, Anal. Methods, 2011, 3, 822–830 RSC.
- E. E. Kempa, K. A. Hollywood, C. A. Smith and P. E. Barran, High throughput screening of complex biological samples with mass spectrometry – from bulk measurements to single cell analysis, Analyst, 2019, 144, 872–891 RSC.
- PAL autosampler systems, https://www.agilent.com/en/product/gas-chromatography/gc-sample-preparation-introduction/pal-auto-sampler-systems, accessed 25 August 2025 Search PubMed.
- Integrated protein technologies, The SampleStream™ platform is here, https://www.iptinc.com/our-technology/#techoverview, accessed 25 August 2025 Search PubMed.
- H. D. Santos Seckler, H. M. Park, C. M. Lloyd-Jones, R. D. Melani, J. M. Camarillo, J. T. Wilkins, P. D. Compton and N. L. Kelleher, New interface for faster proteoform analysis: immunoprecipitation coupled with SampleStream-mass spectrometry, J. Am. Soc. Mass Spectrom., 2021, 32, 1659–1670 Search PubMed.
- B. J. Des Soye, J. P. McGee, M. A. R. Hollas, E. Forte, R. T. Fellers, R. D. Melani, J. T. Wilkins, P. D. Compton, J. O. Kafader and N. L. Kelleher, Automated immunoprecipitation, sample preparation, and individual ion mass spectrometry platform for proteoforms, Anal. Chem., 2024, 96, 13879–13887 Search PubMed.
- B. Musselman, J. Tice and E. Crawford, in Ambient ionization mass spectrometry, ed. M. Domin and R. Cody, The Royal Society of Chemistry, 2014, vol. 18, pp. 445–461 Search PubMed.
- Direct analysis in real time mass spectrometry, https://www.bruker.com/en/products-and-solutions/mass-spectrometry/dart-ms.html, accessed 3 November 2025 Search PubMed.
- Direct analysis in real time, https://www.jeolusa.com/PRODUCTS/Mass-Spectrometers/AccuTOF-DART, accessed 3 November 2025 Search PubMed.
- DESI XS, https://www.waters.com/nextgen/us/en/products/mass-spectrometry/mass-spectrometry-ion-sources/desi-xs.html, accessed 3 November 2025.
- VeriSpray PaperSpray ion source for mass spectrometry, https://www.thermofisher.com/tw/zt/home/industrial/mass-spectrometry/liquid-chromatography-mass-spectrometry-lc-ms/lc-ms-accessories/verispray-paperspray-ion-source-mass-spectrometry.html, accessed 3 November 2025 Search PubMed.
- DESI XS – designed to deliver, https://www.waters.com/nextgen/tw/zh/library/application-notes/2020/desi-xs-designed-to-deliver.html, accessed 3 November 2025 Search PubMed.
- H. Wang, J. Liu, R. G. Cooks and Z. Ouyang, Paper spray for direct analysis of complex mixtures using mass spectrometry, Angew. Chem., Int. Ed., 2010, 49, 877–880 CrossRef CAS PubMed.
- Therapeutic drug monitoring research, https://www.chem.purdue.edu/media/news/2010/therapeutic_drug_research.html, accessed 28 September 2025 Search PubMed.
- Purdue research park, AMI – pre-clinical PaperSpray ion source for rapid analysis, https://www.youtube.com/watch?v=Zlq1sRND3A8, accessed 27 September 2025 Search PubMed.
- R. D. Espy, N. E. Manicke, Z. Ouyang and R. G. Cooks, Rapid analysis of whole blood by paper spray mass spectrometry for point-of-care therapeutic drug monitoring, Analyst, 2012, 137, 2344–2349 RSC.
- R. D. Espy, S. F. Teunissen, N. E. Manicke, Y. Ren, Z. Ouyang, A. van Asten and R. G. Cooks, Paper spray and extraction spray mass spectrometry for the direct and simultaneous quantification of eight drugs of abuse in whole blood, Anal. Chem., 2014, 86, 7712–7718 CrossRef CAS PubMed.
- R.-Z. Shi, E. T. M. El Gierari, N. E. Manicke and J. D. Faix, Rapid measurement of tacrolimus in whole blood by paper spray-tandem mass spectrometry (PS-MS/MS, Clin. Chim. Acta, 2015, 441, 99–104 CrossRef CAS PubMed.
- R.-Z. Shi, E. T. M. El Gierari, J. D. Faix and N. E. Manicke, Rapid measurement of cyclosporine and sirolimus in whole blood by paper spray–tandem mass spectrometry, Clin. Chem., 2016, 62, 295–297 CrossRef CAS PubMed.
- C. Görgens, K. Walker, C. Boeser, N. Wijeratne, C. Martins, S. Guddat and M. Thevis, Paper spray mass spectrometry – a potential complementary technique for the detection of polar compounds in sports drug testing, Drug Test. Anal., 2020, 12, 1658–1665 CrossRef PubMed.
- C. Skaggs, H. Zimmerman, N. Manicke and L. Kirkpatrick, Development and validation of a paper spray mass spectrometry method for the rapid quantitation of remdesivir and its active metabolite, GS-441524, in human plasma, J. Mass Spectrom. Adv. Clin. Lab, 2022, 25, 27–35 CrossRef CAS PubMed.
- G. Ren, N. Manicke, C. Boeser and N. R. Wijeratne, Detection of controlled substances in blood samples using the VeriSpray ion source with TSQ altis MS for clinical research and forensic toxicology, Braz. J. Anal. Chem., 2022, 9, 256–261 CAS.
- A. Saatchi, T. M. Zarkovic, S. A. Borden, J. Palaty and C. G. Gill, Therapeutic drug monitoring of clozapine in human serum by high-throughput paper spray mass spectrometry, J. Mass Spectrom. Adv. Clin. Lab, 2024, 32, 41–46 CrossRef CAS PubMed.
- Paper spray ionization: The faster path to mass spec analysis, https://documents.thermofisher.com/TFS-Assets/CMD/brochures/eb-73396-ms-paper-spray-ionization-eb73396-en.pdf, accessed 28 September 2025 Search PubMed.
- R. G. Lahr, M. Meyer, L. Nelson, L. A. Kottschade, P. J. Jannetto and Y. K. Yang, Performance comparison of liquid chromatography and paper spray ionization with mass spectrometry for measuring kinase inhibitors in human plasma, ACS Pharmacol. Transl. Sci., 2025, 8, 557–565 CrossRef CAS PubMed.
- D. P. Elpa, G. R. D. Prabhu, S.-P. Wu, K. S. Tay and P. L. Urban, Automation of mass spectrometric detection of analytes and related workflows: a review, Talanta, 2020, 208, 120304 Search PubMed.
- C. Li, S. Chu, S. Tan, X. Yin, Y. Jiang, X. Dai, X. Gong, X. Fang and D. Tian, Towards higher sensitivity of mass spectrometry: a perspective from the mass analyzers, Front. Chem., 2021, 9, 813359 CrossRef CAS PubMed.
- E. González-Infante, A. S. Román, J. F. Ayala-Cabrera, N. Etxebarria, B. González-Gaya, N. Lopez-Herguedas, M. Musatadi, M. Olivares, A. Prieto and O. Zuloaga, Mass spectrometry-based high-throughput sample treatment methods for analysis of xenobiotics in human biofluids, Adv. Sample Prep., 2025, 14, 100183 CrossRef.
- C. J. Gray, B. Thomas, R. Upton, L. G. Migas, C. E. Eyers, P. E. Barran and S. L. Flitsch, Applications of ion mobility mass spectrometry for high throughput, high resolution glycan analysis, Biochim. Biophys. Acta BBA – Gen. Subj., 2016, 1860, 1688–1709 CrossRef CAS PubMed.
- T. O. Metz, E. S. Baker, E. L. Schymanski, R. S. Renslow, D. G. Thomas, T. J. Causon, I. K. Webb, S. Hann, R. D. Smith and J. G. Teeguarden, Integrating ion mobility spectrometry into mass spectrometry-based exposome measurements: what can it add and how far can it go?, Bioanalysis, 2017, 9, 81–98 CrossRef CAS PubMed.
- D. H. Ross, A. Bilbao, R. D. Smith and X. Zheng, in High-throughput mass spectrometry in drug discovery, John Wiley & Sons, Ltd, 2023, pp. 183–213 Search PubMed.
- A. T. Kirk, A. Bohnhorst, C. R. Raddatz, M. Allers and S. Zimmermann, Ultra-high-resolution ion mobility spectrometry—current instrumentation, limitations, and future developments, Anal. Bioanal. Chem., 2019, 411, 6229–6246 CrossRef CAS PubMed.
- R. Dessy, Robots in the laboratory: part II, Anal. Chem., 1983, 55, 1232A–1264A CrossRef.
- D. A. V. Medina, E. V. S. Maciel and F. M. Lanças, Modern automated sample preparation for the determination of organic compounds: a review on robotic and on-flow systems, TrAC Trends Anal. Chem., 2023, 166, 117171 CrossRef.
- J. N. Little, The Zymate laboratory automation systems, J. Liq. Chromatogr., 1986, 9, 3197–3201 CrossRef.
- T. Martinoví, M. Martina, M. Gajdošik, G. Gajdošik and D. Josí, Sample preparation in foodomic analyses, Electrophoresis, 2018, 39, 1527–1542 CrossRef PubMed.
- D. P. Elpa and P. L. Urban, Bubble-assisted sample preparation techniques for mass spectrometry, Mass Spectrom. Rev., 2026, 45, 167–192 CrossRef CAS PubMed.
- H. Fleischer, L. A. Kroos, S. Joshi, T. Roddelkopf, R. Stoll and K. Thurow, Dual-arm robotic compound-oriented measurement system: integration of a positive pressure solid phase extraction unit, Institute of Electrical and Electronics Engineers Inc., Glasgow, United Kingdom, 2021, pp. 1–6 Search PubMed.
- L. Löfgren, G.-B. Forsberg and M. Ståhlman, The BUME method: a new rapid and simple chloroform-free method for total lipid extraction of animal tissue, Sci. Rep., 2016, 6, 27688 CrossRef PubMed.
- L. Löfgren, M. Ståhlman, G.-B. Forsberg, S. Saarinen, R. Nilsson and G. I. Hansson, The BUME method: a novel automated chloroform-free 96-well total lipid extraction method for blood plasma, J. Lipid Res., 2012, 53, 1690–1700 CrossRef PubMed.
- E. Carasek, L. Morés and J. Merib, Basic principles, recent trends and future directions of microextraction techniques for the analysis of aqueous environmental samples, Trends Environ. Anal. Chem., 2018, 19, e00060 CrossRef CAS.
- M. Alexovič, Y. Dotsikas, P. Bober and J. Sabo, Achievements in robotic automation of solvent extraction and related approaches for bioanalysis of pharmaceuticals, J. Chromatogr. B, 2018, 1092, 402–421 CrossRef PubMed.
- S. Dugheri, N. Mucci, G. Cappelli, L. Trevisani, A. Bonari, E. Bucaletti, D. Squillaci and G. Arcangeli, Advanced solid-phase microextraction techniques and related automation: a review of commercially available technologies, J. Anal. Methods Chem., 2022, 2022, 8690569 CrossRef PubMed.
- F. Kong, L. Yuan, Y. F. Zheng and W. Chen, Automatic liquid handling for life science: a critical review of the current state of the art, SLAS Technol., 2012, 17, 169–185 CrossRef CAS PubMed.
- X. Zhang, M. Romm, X. Zheng, E. M. Zink, Y.-M. Kim, K. E. Burnum-Johnson, D. J. Orton, A. Apffel, Y. M. Ibrahim, M. E. Monroe, R. J. Moore, J. N. Smith, J. Ma, R. S. Renslow, D. G. Thomas, A. E. Blackwell, G. Swinford, J. Sausen, R. T. Kurulugama, N. Eno, E. Darland, G. Stafford, J. Fjeldsted, T. O. Metz, J. G. Teeguarden, R. D. Smith and E. S. Baker, SPE-IMS-MS: an automated platform for sub-sixty second surveillance of endogenous metabolites and xenobiotics in biofluids, Clin. Mass Spectrom., 2016, 2, 1–10 CrossRef PubMed.
- A. Gaun, K. N. Lewis Hardell, N. Olsson, J. J. O’brien, S. Gollapudi, M. Smith, G. Mcalister, R. Huguet, R. Keyser, R. Buffenstein and F. E. Mcallister, Automated 16-plex plasma proteomics with real-time search and ion mobility mass spectrometry enables large-scale profiling in naked mole-rats and mice, J. Proteome Res., 2021, 20, 1280–1295 CrossRef CAS PubMed.
- J. Zhang, Y. Yang, M. Ashraf, C. N. Cruz, S. Lee and P. J. Faustino, An advanced automation platform coupled with mass spectrometry for investigating in vitro human skin permeation of UV filters and excipients in sunscreen products, Rapid Commun. Mass Spectrom., 2022, 36, e9273 CrossRef CAS PubMed.
- K. E. Butler and E. S. Baker, A high-throughput ion mobility spectrometry-mass spectrometry screening method for opioid profiling, J. Am. Soc. Mass Spectrom., 2022, 33, 1904–1913 CrossRef CAS PubMed.
- S. Zhang, N. Song, Z. He, M. Zeng and J. Chen, Multi-pesticide residue analysis method designed for the robot experimenters, J. Agric. Food Chem., 2022, 70, 16424–16434 CrossRef CAS PubMed.
- A. Schürmann, C. Crüzer, V. Duss, R. Kämpf, T. Preiswerk and H. J. Huebschmann, Automated micro-solid-phase extraction clean-up and gas chromatography-tandem mass spectrometry analysis of pesticides in foods extracted with ethyl acetate, Anal. Bioanal. Chem., 2024, 416, 689–700 CrossRef PubMed.
- B. R. Juliano, J. W. Keating, H. W. Li, A. G. Anders, Z. Xie and B. T. Ruotolo, Development of an automated, high-throughput methodology for native mass spectrometry and collision-induced unfolding, Anal. Chem., 2023, 95, 16717–16724 CrossRef CAS PubMed.
- B. T. Veach, T. K. Mudalige and P. Rye, RapidFire mass spectrometry with enhanced throughput as an alternative to liquid-liquid salt assisted extraction and LC/MS analysis for sulfonamides in honey, Anal. Chem., 2017, 89, 3256–3260 CrossRef CAS PubMed.
- G. K. Reder, E. Y. Bjurström, D. Brunnsåker, F. Kronström, P. Lasin, I. Tiukova, O. I. Savolainen, J. N. Dodds, J. C. May, J. P. Wikswo, J. A. McLean and R. D. King, AutonoMS: automated ion mobility metabolomic fingerprinting, J. Am. Soc. Mass Spectrom., 2024, 35, 542–550 CrossRef CAS PubMed.
- E. Bourgogne, C. Grivet, E. Varesio and G. Hopfgartner, Generic on-line solid phase extraction sample preparation strategies for the analysis of drugs in biological matrices by LC-MS/MS, J. Pharm. Biomed. Anal., 2015, 102, 290–298 Search PubMed.
- A. Naccarato, R. Elliani and A. Tagarelli, Microextraction and eco-friendly techniques applied to solid matrices followed by chromatographic analysis, Separations, 2025, 12, 124 CrossRef CAS.
- A. Das, M. Thiruvenkatakumar, H. Poojary, K. Selvamuthukumar, S. Koo and C. Ghosh, Current advances in solid-phase microextraction techniques for the estimation of pesticides from fruits and vegetables, Microchem. J., 2025, 215, 114412 CrossRef CAS.
- R. P. Belardi and J. B. Pawliszyn, The application of chemically modified fused silica fibers in the extraction of organics from water matrix samples and their rapid transfer to capillary columns, Water Qual. Res. J., 1989, 24, 179–191 CrossRef CAS.
- C. L. Arthur and J. Pawliszyn, Solid phase microextraction with thermal desorption using fused silica optical fibers, Anal. Chem., 1990, 62, 2145–2148 CrossRef CAS.
- Solid phase microextraction (SPME), https://www.gerstelus.com/products-solid-phase-microextraction/, accessed 3 November 2025 Search PubMed.
- Autosamplers | gas chromatography (GC), https://www.thermofisher.com/tw/zt/home/industrial/chromatography/gas-chromatography-gc/gc-autosamplers.html, accessed 3 November 2025 Search PubMed.
- J. P. Hutchinson, L. Setkova and J. Pawliszyn, Automation of solid-phase microextraction on a 96-well plate format, J. Chromatogr. A, 2007, 1149, 127–137 Search PubMed.
- D. Vuckovic, E. Cudjoe, D. Hein and J. Pawliszyn, Automation of solid-phase microextraction in high-throughput format and applications to drug analysis, Anal. Chem., 2008, 80, 6870–6880 CrossRef CAS PubMed.
- W. Zhou, E. Nazdrajić and J. Pawliszyn, Rapid screening and quantitation of drugs of abuse by both positive and negative modes via coated blade spray–mass spectrometry, J. Am. Soc. Mass Spectrom., 2022, 33, 1187–1193 CrossRef CAS PubMed.
- W. Zhou, E. Nazdrajić and J. Pawliszyn, High-throughput and rapid screening of drugs of abuse in saliva by multi-segment injection using solid-phase microextraction-automated microfluidic open interface-mass spectrometry, Anal. Chem., 2023, 95, 6367–6373 CrossRef CAS PubMed.
- W. Zhou, M. N. Wieczorek and J. Pawliszyn, High throughput and automated solid-phase microextraction and determination by liquid chromatography-mass spectrometry for the analysis of mycotoxins in beer, Food Chem., 2023, 426, 136557 CrossRef CAS PubMed.
- Evosep, BR-003G: Evosep One, a standardized separation tool for proteomics, https://www.evosep.com/wp-content/uploads/2024/03/BR-003G-EVOSEP-brochure.pdf, accessed 2 September 2025 Search PubMed.
- Evosep, Evotip pure: A new and improved generation of Evotip, https://www.evosep.com/evotip/, accessed 2 September 2025 Search PubMed.
- Evosep, AN-022D: evotip pure simplifies workflows with excellent reproducibility, storage and recovery, https://www.evosep.com/wp-content/uploads/2024/03/AN-022D_Evotip-Pure-simplifies-workflows-with-excellent-reproducibility-storage-and-recovery-WEB.pdf, accessed 2 September 2025 Search PubMed.
- C. Krisp, M. Lubeck, G. Kruppa, A. Almeida, J. Sandow, D. Hartlmayr and A. Seth, AN-021A: Pushing the boundaries for robust and high-throughput single cell analysis, Evosep, 2022, https://www.evosep.com/wp-content/uploads/2022/12/AN-021A-Pushing-the-boundaries-for-robust-and-high-throughput-single-cell-proteomics.pdf Search PubMed.
- Evosep, Automation, https://www.evosep.com/support/automation/, accessed 3 September 2025 Search PubMed.
- A. H. Kverneland, F. Harking, J. M. Vej-Nielsen, M. Huusfeldt, D. B. Bekker-Jensen, I. M. Svane, N. Bache and J. V. Olsen, Fully automated workflow for integrated sample digestion and evotip loading enabling high-throughput clinical proteomics, Mol. Cell. Proteomics, 2024, 23, 100790 CrossRef CAS PubMed.
- Z. Ye, P. Sabatier, L. van der Hoeven, M. Y. Lechner, T. Phlairaharn, U. H. Guzman, Z. Liu, H. Huang, M. Huang, X. Li, D. Hartlmayr, F. Izaguirre, A. Seth, H. J. Joshi, S. Rodin, K. H. Grinnemo, O. B. Hørning, D. B. Bekker-Jensen, N. Bache and J. V. Olsen, Enhanced sensitivity and scalability with a chip-tip workflow enables deep single-cell proteomics, Nat. Methods, 2025, 22, 499–509 CrossRef CAS PubMed.
- C. Ctortecka, N. M. Clark, B. W. Boyle, A. Seth, D. R. Mani, N. D. Udeshi and S. A. Carr, Automated single-cell proteomics providing sufficient proteome depth to study complex biology beyond cell type classifications, Nat. Commun., 2024, 15, 5707 CrossRef CAS PubMed.
- C. Ctortecka, D. Hartlmayr, A. Seth, M. Ali, G. Tourniaire and C. Sasu, ProteoCHIP & cellenONE taking multiplexed single cell proteomics sample preparation to the next level, Cellenion, 2021 Search PubMed , https://www.scienion.com/wp-content/uploads/2025/12/AppNote-20-IMP-Vienna-Multiplexed-single-cell-proteomics-sample-preparation-using-the-proteoCHIP.pdf, accessed on 2 September 2025.
- C. Ctortecka, D. Hartlmayr, A. Seth, S. Mendjan, G. Tourniaire, N. D. Udeshi, S. A. Carr and K. Mechtler, An automated nanowell-array workflow for quantitative multiplexed single-cell proteomics sample preparation at high sensitivity, Mol. Cell. Proteomics, 2023, 22, 100665 CrossRef CAS PubMed.
- A. Leduc, L. Khoury, J. Cantlon, S. Khan and N. Slavov, Massively parallel sample preparation for multiplexed single-cell proteomics using nPOP, Nat. Protoc., 2024, 19, 3750–3776 CrossRef CAS PubMed.
- V. Kertesz and G. J. Van Berkel, Fully automated liquid extraction-based surface sampling and ionization using a chip-based robotic nanoelectrospray platform, J. Mass Spectrom., 2010, 45, 252–260 CrossRef CAS PubMed.
- D. Eikel and J. Henion, Liquid extraction surface analysis (LESA) of food surfaces employing chip-based nano-electrospray mass spectrometry, Rapid Commun. Mass Spectrom., 2011, 25, 2345–2354 CrossRef CAS PubMed.
- S. Das and R. Bhatia, Liquid extraction surface analysis-mass spectrometry: an advanced and environment-friendly analytical tool in modern analysis, J. Sep. Sci., 2022, 45, 2746–2765 CrossRef CAS PubMed.
- TriVersa NanoMate® – Advion Interchim Scientific, https://www.advion.com/products/triversa-nanomate/, accessed 24 August 2025 Search PubMed.
- R. L. Griffiths, A. L. Simmonds, J. G. Swales, R. J. A. Goodwin and H. J. Cooper, LESA MS imaging of heat-preserved and frozen tissue: benefits of multistep static FAIMS, Anal. Chem., 2018, 90, 13306–13314 CrossRef CAS PubMed.
- R. L. Griffiths, J. W. Hughes, S. E. Abbatiello, M. W. Belford, I. B. Styles and H. J. Cooper, Comprehensive LESA mass spectrometry imaging of intact proteins by integration of cylindrical FAIMS, Anal. Chem., 2020, 92, 2885–2890 CrossRef CAS PubMed.
- J. Havlikova, R. C. May, I. B. Styles and H. J. Cooper, Liquid extraction surface analysis mass spectrometry of ESKAPE pathogens, J. Am. Soc. Mass Spectrom., 2021, 32, 1345–1351 CrossRef CAS PubMed.
- S. Zhou, Z. Fatma, P. Xue, S. Mishra, M. Cao, H. Zhao and J. V. Sweedler, Mass spectrometry-based high-throughput quantification of bioproducts in liquid culture, Anal. Chem., 2023, 95, 4067–4076 CrossRef CAS PubMed.
- J. Lim, S. Zhou, J. Baek, A. Y. Kim, E. Valera, J. Sweedler and R. Bashir, A blood drying process for DNA amplification, Small, 2024, 20, 2307959 CrossRef CAS PubMed.
- L. Lamont, M. Baumert, N. Ogrinc Potočnik, M. Allen, R. Vreeken, R. M. A. Heeren and T. Porta, Integration of ion mobility MSE after fully automated, online, high-resolution liquid extraction surface analysis micro-liquid chromatography, Anal. Chem., 2017, 89, 11143–11150 CrossRef CAS PubMed.
- D. A. V. Medina, L. F. Rodriguez Cabal, F. M. Lanças and Á. J. Santos-Neto, Sample treatment platform for automated integration of microextraction techniques and liquid chromatography analysis, HardwareX, 2019, 5, e00056 CrossRef.
- D. A. V. Medina, L. F. Rodríguez Cabal, G. M. Titato, F. M. Lanças and Á. J. Santos-Neto, Automated online coupling of robot-assisted single drop microextraction and liquid chromatography, J. Chromatogr. A, 2019, 1595, 66–72 CrossRef CAS PubMed.
- M. D. Bocelli, D. A. V. Medina, F. M. Lanças and Á. J. dos Santos-Neto, Automated microextraction by packed sorbent of endocrine disruptors in wastewater using a high-throughput robotic platform followed by liquid chromatography-tandem mass spectrometry, Anal. Bioanal. Chem., 2023, 415, 6165–6176 CrossRef CAS PubMed.
- G. Peris-Pastor, J. Grau, L. Arruza, J. L. Benedé and A. Chisvert, Lab-made open-source controlled robotic workstation for sorbent-based dispersive microextraction of low-volume samples, Anal. Chim. Acta, 2025, 1356, 344033 CrossRef CAS PubMed.
- S. Joshi, X. Chu, H. Fleischer, T. Roddelkopf, M. Klos and K. Thurow, Analysis of measurement process design for a dual-arm robot using graphical user interface, Institute of Electrical and Electronics Engineers Inc., Auckland, New Zealand, 2019, pp. 1–6 Search PubMed.
- H. Fleischer, S. Joshi, T. Roddelkopf, M. Klos and K. Thurow, Automated analytical measurement processes using a dual-arm robotic system, SLAS Technol., 2019, 24, 354–356 CrossRef PubMed.
- H. Fleischer, D. Baumann, S. Joshi, X. Chu, T. Roddelkopf, M. Klos and K. Thurow, Analytical measurements and efficient process generation using a dual–arm robot equipped with electronic pipettes, Energies, 2018, 11, 2567 Search PubMed.
- Eppendorf epMotion | your solution for reproducible pipetting, https://www.eppendorf.com/us-en/Products/Liquid-Handling/Automated-Pipetting-c-WebPSub-H-540035, accessed 3 November 2025 Search PubMed.
- Robots, https://opentrons.com/products/categories/robots, accessed 3 November 2025 Search PubMed.
- Automated sample preparation solution to save time and money, https://www.palsystem.com/en/, accessed 3 November 2025 Search PubMed.
- Automated liquid handling | Hamilton liquid handling platforms, https://www.hamiltoncompany.com/automated-liquid-handling, accessed 3 November 2025 Search PubMed.
- Andrew+ pipetting robot | Andrew alliance, https://www.andrewalliance.com/pipetting-robot/, accessed 3 November 2025 Search PubMed.
- Biomek, https://www.beckman.tw/landing/ppc/liquid-handlers/biomek, accessed 3 November 2025 Search PubMed.
- R. Lerner, D. Baker, C. Schwitter, S. Neuhaus, T. Hauptmann, J. M. Post, S. Kramer and L. Bindila, Four-dimensional trapped ion mobility spectrometry lipidomics for high throughput clinical profiling of human blood samples, Nat. Commun., 2023, 14, 937 CrossRef CAS PubMed.
- J. F. Cahill, V. Kertesz, P. Saint-Vincent, H. Valentino, E. Drufva, N. Thiele and J. K. Michener, High-throughput characterization and optimization of polyamide hydrolase activity using open port sampling interface mass spectrometry, J. Am. Soc. Mass Spectrom., 2023, 34, 1383–1391 CrossRef CAS PubMed.
- O. H. L. Williams, O. Rusli, L. Ezzedinloo, T. M. Dodgen, J. K. Clegg and N. J. Rijs, Automated structural activity screening of β-diketonate assemblies with high-throughput ion mobility-mass spectrometry, Angew. Chem., Int. Ed., 2024, 63, e202313892 Search PubMed.
- B. Van Puyvelde, C. L. Hunter, M. Zhgamadze, S. Savant, Y. O. Wang, E. Hoedt, K. Raedschelders, M. Pope, C. A. Huynh, V. K. Ramanujan, W. Tourtellotte, M. Razavi, N. L. Anderson, G. Martens, D. Deforce, Q. Fu, M. Dhaenens and J. E. Van Eyk, Acoustic ejection mass spectrometry empowers ultra-fast protein biomarker quantification, Nat. Commun., 2024, 15, 5114 CrossRef CAS PubMed.
- T. Kipura, M. Hotze, A. Hofer, A.-S. Egger, L. E. Timpen, C. A. Opitz, P. A. Townsend, L. A. Gethings, K. Thedieck and M. Kwiatkowski, Automated liquid handling extraction and rapid quantification of underivatized amino acids and tryptophan metabolites from human serum and plasma using dual-column U(H)PLC-MRM-MS and its application to prostate cancer study, Metabolites, 2024, 14, 370 CrossRef CAS PubMed.
- R. Ellson, Picoliter: enabling precise transfer of nanoliter and picoliter volumes, Drug Discov. Today, 2002, 7, S32–S34 CrossRef.
- R. Ellson, M. Mutz, B. Browning, L. Lee, M. F. Miller and R. Papen, Transfer of low nanoliter volumes between microplates using focused acoustics—automation considerations, SLAS Technol., 2003, 8, 29–34 Search PubMed.
- N. M. Nebane, T. Coric, S. McKellip, L. Woods, M. Sosa, L. Rasmussen, M.-A. Bjornsti and E. L. White, Acoustic droplet ejection technology and its application in high-throughput RNA interference screening, SLAS Technol., 2016, 21, 198–203 Search PubMed.
- B. Hadimioglu, R. Stearns and R. Ellson, Moving liquids with sound: the physics of acoustic droplet ejection for robust laboratory automation in life sciences, SLAS Technol., 2016, 21, 4–18 CrossRef PubMed.
- Echo Acoustic Liquid Handlers, https://www.beckman.com/liquid-handlers/echo-acoustic, accessed 30 September 2025 Search PubMed.
- M. Winter, R. P. Simon, T. T. Häbe, R. Ries, Y. Wang, D. Kvaskoff, A. Fernández-Montalván, A. H. Luippold, F. H. Büttner and W. Reindl, Label-free high-throughput screening via acoustic ejection mass spectrometry put into practice, SLAS Discov., 2023, 28, 240–246 CrossRef CAS PubMed.
- Echo® MS+ system, https://sciex.com/products/integrated-solutions/echo-ms-plus, accessed 30 September 2025 Search PubMed.
- H. Zhang, C. Liu, W. Hua, L. P. Ghislain, J. Liu, L. Aschenbrenner, S. Noell, K. J. Dirico, L. F. Lanyon, C. M. Steppan, M. West, D. W. Arnold, T. R. Covey, S. S. Datwani and M. D. Troutman, Acoustic ejection mass spectrometry for high-throughput analysis, Anal. Chem., 2021, 93, 10850–10861 CrossRef CAS PubMed.
- R. P. Simon, T. T. Häbe, R. Ries, M. Winter, Y. Wang, A. Fernández-Montalván, D. Bischoff, F. Runge, W. Reindl, A. H. Luippold and F. H. Büttner, Acoustic ejection mass spectrometry: a fully automatable technology for high-throughput screening in drug discovery, SLAS Discov., 2021, 26, 961–973 CrossRef CAS PubMed.
- H. Fleischer, D. Baumann, X. Chu, T. Roddelkopf, M. Klos and K. Thurow, Integration of electronic pipettes into a dual-arm robotic system for automated analytical measurement processes behaviors, Institute of Electrical and Electronics Engineers Inc., Munich, Germany, 2018, pp. 22–27 Search PubMed.
- J. Shanley, F. Pu, J. D. Williams, N. L. Elsen, S. M. Gopalakrishnan, J. Y. Pan and A. J. Radosevich, Collaborative robotics to enable ultra-high-throughput IR-MALDESI, SLAS Technol., 2024, 29, 100163 CrossRef CAS PubMed.
- H. Fleischer, M. A. Tlili, T. Roddelkopf and K. Thurow, Automated sample transportation and handling for determination of trace metals in cell growth media using ICP-MS, Institute of Electrical and Electronics Engineers Inc., Bari, Italy, 2024, pp. 3300–3305 Search PubMed.
- V. Gusan, A. M. Titu, A. Bogorin-Predescu, P. Tertereanu, D. Bâlc and A. B. Pop, in From industrial robots to collaborative robots: exploring the transformation in the collaborative work environment, ed. I. Karabegovic, A. Kovačević and S. Mandzuka, New Technologies, Development and Application VII, NT 2024, Lecture Notes in Networks and Systems, Springer, Cham, 2024, vol. 1069, pp. 32–41 Search PubMed.
- A. J. Radosevich, F. Pu, D. Chang-Yen, J. W. Sawicki, N. N. Talaty, N. L. Elsen, J. D. Williams and J. Y. Pan, Ultra-high-throughput ambient MS: direct analysis at 22 samples per second by infrared matrix-assisted laser desorption electrospray ionization mass spectrometry, Anal. Chem., 2022, 94, 4913–4918 CrossRef CAS PubMed.
- E. Vorberg, H. Fleischer, S. Junginger, N. Stoll and K. Thurow, Automated sample preparation for mercury analysis in wood materials, IET Sci. Meas. Technol., 2016, 10, 398–404 CrossRef.
- H. Fleischer, C. Lutter, A. Büttner, W. Mittelmeier and K. Thurow, Semi-automated determination of heavy metals in autopsy tissue using robot-assisted sample preparation and ICP-MS, Molecules, 2021, 26, 3820 CrossRef CAS PubMed.
- R. V. Bennett, E. M. Morzan, J. O. Huckaby, M. E. Monge, H. I. Christensen and F. M. Fernández, Robotic plasma probe ionization mass spectrometry (RoPPI-MS) of non-planar surfaces, Analyst, 2014, 139, 2658–2662 Search PubMed.
- A. Li, M. R. L. Paine, S. Zambrzycki, R. B. Stryffeler, J. Wu, M. Bouza, J. Huckaby, C.-Y. Chang, M. Kumar, P. Mukhija and F. M. Fernández, Robotic surface analysis mass spectrometry (RoSA-MS) of three-dimensional objects, Anal. Chem., 2018, 90, 3981–3986 Search PubMed.
- M. F. Keating, J. Zhang, C. L. Feider, S. Retailleau, R. Reid, A. Antaris, B. Hart, G. Tan, T. E. Milner, K. Miller and L. S. Eberlin, Integrating the MasSpec pen to the da vinci surgical system for in vivo tissue analysis during a robotic assisted porcine surgery, Anal. Chem., 2020, 92, 11535–11542 Search PubMed.
- J. Zhang, J. Rector, J. Q. Lin, J. H. Young, M. Sans, N. Katta, N. Giese, W. Yu, C. Nagi, J. Suliburk, J. Liu, A. Bensussan, R. J. DeHoog, K. Y. Garza, B. Ludolph, A. G. Sorace, A. Syed, A. Zahedivash, T. E. Milner and L. S. Eberlin, Nondestructive tissue analysis for ex vivo and in vivo cancer diagnosis using a handheld mass spectrometry system, Sci. Transl. Med., 2017, 9, eaan3968 CrossRef PubMed.
- S.-H. Chiu and P. L. Urban, Robotics-assisted mass spectrometry assay platform enabled by open-source electronics, Biosens. Bioelectron., 2015, 64, 260–268 Search PubMed.
- C.-L. Chen, T.-R. Chen, S.-H. Chiu and P. L. Urban, Dual robotic arm “production line” mass spectrometry assay guided by multiple Arduino-type microcontrollers, Sens. Actuators, B, 2017, 239, 608–616 CrossRef CAS.
- N. H. Abu Bakar, K.-C. Yu and P. L. Urban, Robotized noncontact open-space mapping of volatile organic compounds emanating from solid specimens, Anal. Chem., 2021, 93, 6889–6894 Search PubMed.
- C.-C. Chan, N. H. Abu Bakar, C. M. Raju and P. L. Urban, Computer vision-assisted robotized sampling of volatile organic compounds, Anal. Chem., 2024, 96, 16307–16314 CrossRef CAS PubMed.
- X. Liu, Q. Huang, J. Deng, X. Liu and B. Hu, Portable mass spectrometry for on-site detection of hazardous volatile organic compounds via robotic extractive sampling, Anal. Chem., 2024, 96, 9325–9331 Search PubMed.
- X. Liu, X. Liu, B. Li, X. Zhang and B. Hu, Lab-on-robot: unmanned mass spectrometry robot for direct sample analysis in hazardous and radioactive environments, Anal. Chem., 2025, 97, 9126–9130 CrossRef CAS PubMed.
- P. H. Winston, Artificial intelligence, Addison-Wesley Pub. Co., Reading, Massachusetts, 3rd edn, 1993. Search PubMed.
- K. Guo, Z. Yang, C.-H. Yu and M. J. Buehler, Artificial intelligence and machine learning in design of mechanical materials, Mater. Horiz., 2021, 8, 1153–1172 RSC.
- Q. Salamat, Z. P. Gumus and M. Soylak, Recent developments and applications of artificial intelligence in solid/liquid extraction studies, TrAC Trends Anal. Chem., 2025, 182, 118057 Search PubMed.
- Y. Mahdi and K. Daoud, Microdroplet size prediction in microfluidic systems via artificial neural network modeling for water-in-oil emulsion formulation, J. Dispers. Sci. Technol., 2017, 38, 1501–1508 CrossRef CAS.
- E. A. Galan, H. Zhao, X. Wang, Q. Dai, W. T. S. Huck and S. Ma, Intelligent microfluidics: the convergence of machine learning and microfluidics in materials science and biomedicine, Matter, 2020, 3, 1893–1922 CrossRef.
- J. Zheng, T. Cole, Y. Zhang, N. Bayinqiaoge, D. Yuan and S.-Y. Tang, An automated and intelligent microfluidic platform for microalgae detection and monitoring, Lab Chip, 2024, 24, 244–253 Search PubMed.
- J. Cheng, R. Anne and Y.-C. Chen, Transforming microfluidics for single-cell analysis with robotics and artificial intelligence, Lab Chip, 2025, 25, 6100–6125 RSC.
- O. J. Dressler, P. D. Howes, J. Choo and A. J. Demello, Reinforcement learning for dynamic microfluidic control, ACS Omega, 2018, 3, 10084–10091 CrossRef CAS PubMed.
- H. Parastar and P. Weller, Benchtop volatilomics supercharged: How machine learning based design of experiment helps optimizing untargeted GC-IMS gas phase metabolomics, Talanta, 2024, 272, 125788 Search PubMed.
- W. Shigeeda, R. Yosihimura, Y. Fujita, H. Saiki, H. Deguchi, M. Tomoyasu, S. Kudo, Y. Kaneko, H. Kanno, Y. Inoue and H. Saito, Utility of mass spectrometry and artificial intelligence for differentiating primary lung adenocarcinoma and colorectal metastatic pulmonary tumor, Thorac. Cancer, 2021, 13, 202 Search PubMed.
- O. E. Olajide, M. Zirpoli, K. Y. Kartowikromo, J. Zheng and A. M. Hamid, Discrimination of common E. Coli strains in urine by liquid chromatography-ion mobility-tandem mass spectrometry and machine learning, J. Am. Soc. Mass Spectrom., 2024, 35, 2706–2713 Search PubMed.
- B. Shen, F. Zhou and P. Nemes, Real-time eco–AI, electrophoresis-correlative data-dependent acquisition with AI-based data processing broadens access to single-cell mass spectrometry proteomics, Angew. Chem., Int. Ed., 2025, 64, e202510692 CrossRef CAS PubMed.
- S. Tiwary, R. Levy, P. Gutenbrunner, F. Salinas Soto, K. K. Palaniappan, L. Deming, M. Berndl, A. Brant, P. Cimermancic and J. Cox, High-quality MS/MS spectrum prediction for data-dependent and data-independent acquisition data analysis, Nat. Methods, 2019, 16, 519–525 Search PubMed.
- K. Y. Kartowikromo, O. E. Olajide and A. M. Hamid, Collision cross section measurement and prediction methods in omics, J. Mass Spectrom., 2023, 58, e4973 Search PubMed.
- J. Xue, B. Wang, H. Ji and W. Li, RT-Transformer: retention time prediction for metabolite annotation to assist in metabolite identification, Bioinformatics, 2024, 40, btae084 CrossRef CAS PubMed.
- A. G. Beck, M. Muhoberac, C. E. Randolph, C. H. Beveridge, P. R. Wijewardhane, H. I. Kenttämaa and G. Chopra, Recent developments in machine learning for mass spectrometry, ACS Meas. Sci. Au, 2024, 4, 233–246 CrossRef CAS PubMed.
- J. Liu, C. Bao, J. Zhang, Z. Han, H. Fang and H. Lu, Artificial intelligence with mass spectrometry-based multimodal molecular profiling methods for advancing therapeutic discovery of infectious diseases, Pharmacol. Ther., 2024, 263, 108712 Search PubMed.
- D. H. Ross, H. Bhotika, X. Zheng, R. D. Smith, K. E. Burnum-Johnson and A. Bilbao, Computational tools and algorithms for ion mobility spectrometry-mass spectrometry, Proteomics, 2024, 24, 2200436 CrossRef PubMed.
- A. Bilbao, The future of a myriad of accelerated biodiscoveries lies in AI-powered mass spectrometry and multiomics integration, J. Mass Spectrom., 2025, 60, e5157 Search PubMed.
- K. Y. Kartowikromo, J. S. Pizzo, I. Jerin and A. M. Hamid, Advancements in ion mobility-based diagnostics for infectious diseases, Proteomics, 2025, 25, e13976 Search PubMed.
- S. Macenski, T. Foote, B. Gerkey, C. Lalancette and W. Woodall, Robot operating system 2: design, architecture, and uses in the wild, Sci. Robot., 2022, 7, eabm6074 CrossRef PubMed.
- ROS – robot operating system, https://www.ros.org/, accessed 3 November 2025 Search PubMed.
- S. U. Khan, V. K. Møller, R. J. N. Frandsen and M. Mansourvar, Real-time AI-driven quality control for laboratory automation: a novel computer vision solution for the opentrons OT-2 liquid handling robot, Appl. Intell., 2025, 55, 524 Search PubMed.
- YOLOv5 vs. YOLOv8, Ultralytics, https://docs.ultralytics.com/compare/yolov5-vs-yolov8, accessed 21 October 2025 Search PubMed.
- H.-F. Tsai, S. Podder and P.-Y. Chen, Microsystem advances through integration with artificial intelligence, Micromachines, 2023, 14, 826 Search PubMed.
- M. Abolhasani and E. Kumacheva, The rise of self-driving labs in chemical and materials sciences, Nat. Synth., 2023, 2, 483–492 Search PubMed.
- H. G. Martin, T. Radivojevic, J. Zucker, K. Bouchard, J. Sustarich, S. Peisert, D. Arnold, N. Hillson, G. Babnigg, J. M. Marti, C. J. Mungall, G. T. Beckham, L. Waldburger, J. Carothers, S. S. Sundaram, D. Agarwal, B. A. Simmons, T. Backman, D. Banerjee, D. Tanjore, L. Ramakrishnan and A. Singh, Perspectives for self-driving labs in synthetic biology, Curr. Opin. Biotechnol., 2023, 79, 102881 Search PubMed.
- J. Li, C. Ding, D. Liu, L. Chen and J. Jiang, Autonomous laboratories in China: an embodied intelligence-driven platform to accelerate chemical discovery, Digit. Discov., 2025, 4, 1672–1684 Search PubMed.
- S. Asche, G. J. T. Cooper, G. Keenan, C. Mathis and L. Cronin, A robotic prebiotic chemist probes long term reactions of complexifying mixtures, Nat. Commun., 2021, 12, 3547 Search PubMed.
- T. Dai, S. Vijayakrishnan, F. T. Szczypiński, J.-F. Ayme, E. Simaei, T. Fellowes, R. Clowes, L. Kotopanov, C. E. Shields, Z. Zhou, J. W. Ward and A. I. Cooper, Autonomous mobile robots for exploratory synthetic chemistry, Nature, 2024, 635, 890–897 Search PubMed.
- Q. Zhu, F. Zhang, Y. Huang, H. Xiao, L. Zhao, X. Zhang, T. Song, X. Tang, X. Li, G. He, B. Chong, J. Zhou, Y. Zhang, B. Zhang, J. Cao, M. Luo, S. Wang, G. Ye, W. Zhang, X. Chen, S. Cong, D. Zhou, H. Li, J. Li, G. Zou, W. Shang, J. Jiang and Y. Luo, An all-round AI-chemist with a scientific mind, Natl. Sci. Rev., 2022, 9, nwac190 Search PubMed.
- B. Burger, P. M. Maffettone, V. V. Gusev, C. M. Aitchison, Y. Bai, X. Wang, X. Li, B. M. Alston, B. Li, R. Clowes, N. Rankin, B. Harris, R. S. Sprick and A. I. Cooper, A mobile robotic chemist, Nature, 2020, 583, 237–241 Search PubMed.
- V. Dragone, V. Sans, A. B. Henson, J. M. Granda and L. Cronin, An autonomous organic reaction search engine for chemical reactivity, Nat. Commun., 2017, 8, 15733 Search PubMed.
- J. M. Granda, L. Donina, V. Dragone, D.-L. Long and L. Cronin, Controlling an organic synthesis robot with machine learning to search for new reactivity, Nature, 2018, 559, 377–381 CrossRef CAS PubMed.
- D. Salley, J. S. Manzano, P. J. Kitson and L. Cronin, Robotic modules for the programmable chemputation of molecules and materials, ACS Cent. Sci., 2023, 9, 1525–1537 Search PubMed.
- A. C. Vaucher, F. Zipoli, J. Geluykens, V. H. Nair, P. Schwaller and T. Laino, Automated extraction of chemical synthesis actions from experimental procedures, Nat. Commun., 2020, 11, 3601 CrossRef CAS PubMed.
- C. Bettenhausen, IBM debuts chemical synthesis robot, CEN Glob. Enterp., 2020, 98, 10 Search PubMed.
- IBM RXN for chemistry, https://rxn.res.ibm.com/rxn/robo-rxn/welcome, accessed 3 November 2025 Search PubMed.
- T. Song, M. Luo, X. Zhang, L. Chen, Y. Huang, J. Cao, Q. Zhu, D. Liu, B. Zhang, G. Zou, G. Zhang, F. Zhang, W. Shang, Y. Fu, J. Jiang and Y. Luo, A multiagent-driven robotic AI chemist enabling autonomous chemical research on demand, J. Am. Chem. Soc., 2025, 147, 12534–12545 Search PubMed.
- Mars science laboratory, https://science.nasa.gov/mission/msl-curiosity/, accessed 3 November 2025 Search PubMed.
- Curiosity science instruments, https://science.nasa.gov/mission/msl-curiosity/science-instruments/, accessed 3 November 2025 Search PubMed.
- J. P. Grotzinger, J. Crisp, A. R. Vasavada, R. C. Anderson, C. J. Baker, R. Barry, D. F. Blake, P. Conrad, K. S. Edgett, B. Ferdowski, R. Gellert, J. B. Gilbert, M. Golombek, J. Gómez-Elvira, D. M. Hassler, L. Jandura, M. Litvak, P. Mahaffy, J. Maki, M. Meyer, M. C. Malin, I. Mitrofanov, J. J. Simmonds, D. Vaniman, R. V. Welch and R. C. Wiens, Mars science laboratory mission and science investigation, Space Sci. Rev., 2012, 170, 5–56 Search PubMed.
- M.-P. Chen and B.-C. Liao, Augmented reality laboratory for high school electrochemistry course, Institute of Electrical and Electronics Engineers Inc., Hualien, Taiwan, 2015, pp. 132–136 Search PubMed.
- S. D. Rihm, J. Bai, A. Kondinski, S. Mosbach, J. Akroyd and M. Kraft, Transforming research laboratories with connected digital twins, Nexus, 2024, 1, 100004 Search PubMed.
- M. Taylor, N. B. Abdullah, A. Al-Dargazelli, M. B. Montaner, F. Kareem, A. Locks, Z. Cao, B. Bowles, J. C. Sarraf, T. Fajinmi, Z. Muwaffak, C. Beckwith, G. N. Parkinson, Z. A. E. Waller, B. R. Szulc and S. T. Hilton, Breaking the access to education barrier: enhancing HPLC learning with virtual reality, J. Chem. Educ., 2024, 101, 4093–4101 Search PubMed.
- M. V. Taylor, Z. Muwaffak, M. R. Penny, B. R. Szulc, S. Brown, A. Merritt and S. T. Hilton, Optimising digital twin laboratories with conversational AIs: enhancing immersive training and simulation through virtual reality, Digit. Discov., 2025, 4, 1134–1141 Search PubMed.
- R. Schiavina, L. Bianchi, F. Chessa, U. Barbaresi, L. Cercenelli, S. Lodi, C. Gaudiano, B. Bortolani, A. Angiolini, F. M. Bianchi, A. Ercolino, C. Casablanca, E. Molinaroli, A. Porreca, R. Golfieri, S. Diciotti, E. Marcelli and E. Brunocilla, Augmented reality to guide selective clamping and tumor dissection during robot-assisted partial nephrectomy: a preliminary experience, Clin. Genitourin. Cancer, 2021, 19, e149–e155 Search PubMed.
- S.-Y. Hong and B.-L. Qin, Recent advances in robotic surgery for urologic tumors, Medicina, 2024, 60, 1573 Search PubMed.
- J. Glynn, Russian force generation & technological adaptations update, 2025, https://understandingwar.org/research/russia-ukraine/russian-force-generation-technological-adaptations-update-october-9-2025/, accessed 6 November 2025 Search PubMed.
- Y. Kostyukevich, D. Efremov, V. Ionov, E. Kukaev and E. Nikolaev, Remote detection of explosives using field asymmetric ion mobility spectrometer installed on multicopter, J. Mass Spectrom., 2017, 52, 777–782 CrossRef CAS PubMed.
- M. Allers, A. Ahrens, M. Hitzemann, H. Bock, T. Wolf, J. Radunz, F. Meyer, F. Wilsenack, S. Zimmermann and A. Ficks, Real-time remote detection of airborne chemical hazards—an unmanned aerial vehicle (UAV) carrying an ion mobility spectrometer, IEEE Sens. J., 2023, 23, 16562–16570 Search PubMed.
| This journal is © The Royal Society of Chemistry 2026 |