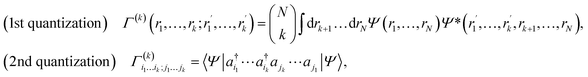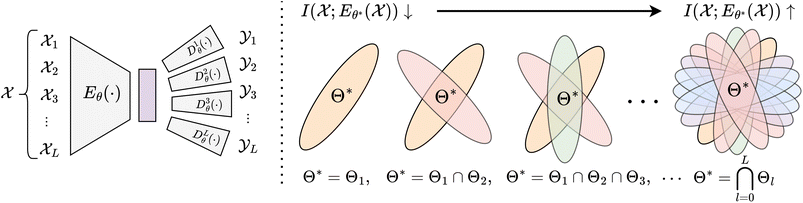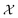Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Connecting the concepts of quantum state tomography and molecular representations for machine learning
Raul
Ortega-Ochoa†
 *ab,
Luis Mantilla
Calderón†
*cd,
Juan Bernardo
Perez Sanchez
*ab,
Luis Mantilla
Calderón†
*cd,
Juan Bernardo
Perez Sanchez
 cd,
Mohsen
Bagherimehrab
fc,
Abdulrahman
Aldossary
cd,
Tejs
Vegge
cd,
Mohsen
Bagherimehrab
fc,
Abdulrahman
Aldossary
cd,
Tejs
Vegge
 ab,
Tonio
Buonassisi
e and
Alán
Aspuru-Guzik
ab,
Tonio
Buonassisi
e and
Alán
Aspuru-Guzik
 *cdfghij
*cdfghij
aDepartment of Energy Conversion and Storage, Technical University of Denmark, Kongens Lyngby 2800, Denmark. E-mail: rauoc@dtu.dk
bCAPeX Pioneer Center for Accelerating P2X Materials Discovery, DK 2800 Kgs. Lyngby, Denmark
cDepartment of Computer Science, University of Toronto, 40 St George St., Toronto, ON M5S 2E4, Canada. E-mail: luis@cs.toronto.edu; alan@aspuru.com
dVector Institute for Artificial Intelligence, Schwartz Reisman Innovation Campus, W1140-108 College St., Toronto, ON M5G 0C6, Canada
eDepartment of Mechanical Engineering, Massachusetts Institute of Technology, Cambridge, MA 02139, USA
fDepartment of Chemistry, University of Toronto, 80 St. George St., Toronto, ON M5S 3H6, Canada
gDepartment of Chemical Engineering & Applied Chemistry, University of Toronto, 200 College St., Toronto, ON M5S 3E5, Canada
hDepartment of Materials Science & Engineering, University of Toronto, 184 College St., Toronto, ON M5S 3E4, Canada
iAcceleration Consortium, 700 University Ave., Toronto, ON M7A 2S4, Canada
jNVIDIA, 431 King St. W #6th, Toronto, ON M5V 1K4, Canada
First published on 19th February 2026
Abstract
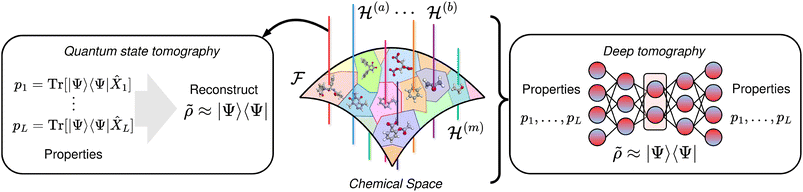
Quantum state tomography has been widely used to reconstruct the quantum state of a system from a set of informationally-complete measurements. Obtaining enough information about, e.g., the wavefunction of a molecule allows its complete characterization. On the other hand, deep learning models have proven useful to perform molecular property prediction (forward design) and inverse design subject to property constraints within the approximate bounds of the data manifold, suggesting that their learned representations are reliable within the region of chemical compound space spanned by their training data. In this work, from the tomographic perspective, we argue that enforcing faithful prediction of an increasing number of diverse molecular descriptors from a shared learned representation progressively constrains the space of admissible internal explanations, driving the inter-alignment of models as they converge towards representation that can explain all observed properties. In the limit where the set of descriptors approaches information-completeness, this alignment drives the learned representations to states that can act, locally, as informationally-equivalent to the molecule's reduced quantum density matrix – a deep tomography. Under this lens, the generalization capabilities of a deep learning model, and the alignment among successful models, arise from unphysical or shortcut solutions becoming progressively incompatible as supervision approaches informational completeness.
1 Introduction
Information can be quantified through the outcomes of questions posed about a system.1,2 Let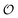 be a random variable taking values in a set of possible objects according to some probability distribution. Any question is a measurable function q whose outcome induces a conditional distribution over
be a random variable taking values in a set of possible objects according to some probability distribution. Any question is a measurable function q whose outcome induces a conditional distribution over 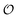 , leaving a remaining uncertainty quantified by
, leaving a remaining uncertainty quantified by 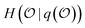 , which satisfies
, which satisfies 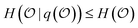 . Its answer may be discrete, in which case we can quantify its uncertainty using Shannon entropy, or continuous, in which case we use the differential entropy.3 A set of questions
. Its answer may be discrete, in which case we can quantify its uncertainty using Shannon entropy, or continuous, in which case we use the differential entropy.3 A set of questions 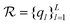 is sufficient for
is sufficient for 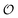 if
if 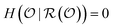 , meaning the answers completely determine
, meaning the answers completely determine 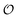 , regardless of whether the set is overcomplete or contains redundant questions. Any such sufficient representation
, regardless of whether the set is overcomplete or contains redundant questions. Any such sufficient representation 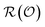 is a random variable satisfying
is a random variable satisfying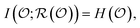 | (1) |
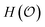 quantifies the uncertainty of
quantifies the uncertainty of 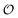 , and
, and 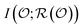 is the mutual information between the object and its representation. In this way,
is the mutual information between the object and its representation. In this way, 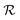 serves as the encoding of the object into a representation space.
serves as the encoding of the object into a representation space.
In quantum chemistry, the main objects of interest are molecules, which are arrangements of interacting electrons and nuclei whose quantum mechanical state is described by a density matrix ρ. Such a density matrix contains all the information of the system.4 It can be either a pure state (e.g., the ground state), a Gibbs state representing a thermal state (i.e., the equilibrium density operator of a system at temperature T for a given Hamiltonian), or any other state that quantifies the status of the molecule. Measuring observables on such quantum states can be thought of as asking a question about the molecule, and the resulting measurement outcomes as the answers to those questions.5 Here, we refer to these answers as molecular descriptors, which contain information about structure, energy, electron density, spin state, oxidation state, or any other property of the system.
By measuring a sufficiently large set of properties of a molecule, known as an informationally-complete positive operator-valued measure (IC-POVM),6–8 we can uniquely determine the density matrix via quantum tomography.9–11 Because this set is informationally complete, any other molecular property can be reconstructed from it. We call the reconstruction from properties to properties a quantum property map (QPM)
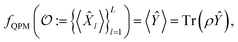 | (2) |
![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) l and Ŷ are quantum observables, and
l and Ŷ are quantum observables, and 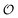 is the set of properties that can be encoded in the molecule's density matrix ρ. Learning QPMs has been the target of deep learning models for molecular science, enabling molecular property prediction (forward design)12–20 and inverse design subject to property constraints21–23 within the bounds of the chemical space spanned by the training set, without explicit reference to the density matrix. In self-supervised learning, QPMs map inputs to an intermediate representation
is the set of properties that can be encoded in the molecule's density matrix ρ. Learning QPMs has been the target of deep learning models for molecular science, enabling molecular property prediction (forward design)12–20 and inverse design subject to property constraints21–23 within the bounds of the chemical space spanned by the training set, without explicit reference to the density matrix. In self-supervised learning, QPMs map inputs to an intermediate representation 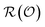 from which the original data can be recovered. Autoencoders fθ = Dθ × Eθ are the clearest example, with
from which the original data can be recovered. Autoencoders fθ = Dθ × Eθ are the clearest example, with 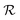 produced by a learned encoder Eθ and inverted by a learned decoder Dθ. Here, each latent coordinate
produced by a learned encoder Eθ and inverted by a learned decoder Dθ. Here, each latent coordinate 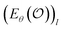 is part of the model's information bottleneck24–26 and can be interpreted as the answer to an unknown continuous-valued question about the input data
is part of the model's information bottleneck24–26 and can be interpreted as the answer to an unknown continuous-valued question about the input data 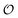 . Training such a model is thus equivalent to learning an interrogation protocol that extracts the essential information of the object,27 and determining the physical meaning of this interrogation process would enable meaningful AI explainability.28–30 A QPM is learnable only when a well-defined input–output mapping exists. Otherwise, models can only resort to memorization (not interpolatable) or trivial statistical predictors (e.g., mean regression under an MSE loss). For example, in a minimal quantum system such as a single spin-
. Training such a model is thus equivalent to learning an interrogation protocol that extracts the essential information of the object,27 and determining the physical meaning of this interrogation process would enable meaningful AI explainability.28–30 A QPM is learnable only when a well-defined input–output mapping exists. Otherwise, models can only resort to memorization (not interpolatable) or trivial statistical predictors (e.g., mean regression under an MSE loss). For example, in a minimal quantum system such as a single spin-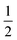 degree of freedom, a single descriptor given by 〈Sz〉 is insufficient to predict 〈Sy〉 since no functional map exists between them, in this case ML can only memorize training samples or perform trivial regression.
degree of freedom, a single descriptor given by 〈Sz〉 is insufficient to predict 〈Sy〉 since no functional map exists between them, in this case ML can only memorize training samples or perform trivial regression.
In a general setting, after successful training, these models learn structured representations that are semantically coherent: similar objects tend to map to nearby points in the learned representation space. This property enables interpolation and generative capabilities. Although a model must arrange objects in a semantically meaningful way in latent space to enable generative behavior,31 there is no unique way to achieve this arrangement—because what counts as “semantic” is itself context-dependent. For example, benzene and pyridine are structurally very similar when viewed as molecular graphs, yet they differ greatly in their dipole moments. An even more striking case is thalidomide,32 its two enantiomers are mirror images of each other, nearly identical structurally, yet their biological activities are drastically different.33 Different contexts imply different semantic arrangements, and learning many competing contexts can force convergence towards representations that preserve physically meaningful information, assuming that the model is not exploiting spurious correlations as documented in the “Clever Hans” effect, where it was shown that some vision models had learned to correctly classify images by using the image's watermarks.34,35
Crucially, molecular descriptors are not independent labels: they are induced by a shared physical state. Here, we argue that if the data is obtained from informationally-complete observables, ML models learning QPMs converge towards an informationally-complete latent space that is semantically well-defined and behaves as a compressed representation of the system's quantum state, analogous to a state tomography reconstruction (cf.Fig. 1). Such models would effectively become surrogates for quantum mechanics, though likely in an approximate form, capturing only mean-field or low-order reduced density matrix information in the subset of the chemical space accessible through the dataset. We call this internal representation a deep tomography representation.
2 Information in one molecule
Molecules are composed of electrons and nuclei, which interact with each other through electromagnetic forces, and are represented by a wave function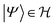 or density matrix
or density matrix 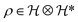 , where
, where 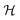 is the system's Hilbert space. This defines the molecular Hamiltonian, the operator that dictates the energy, dynamics, and symmetries of the molecule:
is the system's Hilbert space. This defines the molecular Hamiltonian, the operator that dictates the energy, dynamics, and symmetries of the molecule:| Ĥ = Tn + Te + Vnn + Vne + Vee | (3) |
However, this many-body problem is intractable for large molecules since the dimension of the Hilbert space grows exponentially with the number of electrons and nuclei.36,37 Even if we had a quantum computer that could store all the degrees of freedom of the full wave function of a molecule, solving for the ground or Gibbs state of a general molecule is a QMA-complete problem.38–40 Thus, heuristic methods that make use of a compressed representation of the wavefunction, like Hartree–Fock,41 DFT,42 coupled cluster,43 VQE,44 or others, are needed and used to approximate the eigenstates.
When calculating molecular properties using these methods, it is common to express the Hamiltonian in second quantization, where operators are written in terms of creation and annihilation operators over a set of basis functions {ϕµ}Kµ=1 that span the relevant Hilbert space. In the standard Born–Oppenheimer approach, this basis is electronic only; however, the same formalism applies if the basis functions span the joint electron-nuclear space. On one hand, this formalism uses fewer degrees of freedom to approximate the molecular wave function compared to the first quantization representation. On the other hand, some properties, such as the fermionic and bosonic symmetries of the wave function, are easily encoded in the algebra of these creation and annihilation operators. The molecular Hamiltonian36 can thus be written as
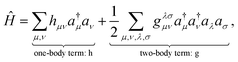 | (4) |
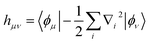 are one-body integrals,
are one-body integrals, 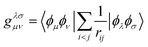 are two-body integrals, and a†µ and aµ are the creation and annihilation operator for the basis function ϕµ. Since Ĥ only depends on one- and two-body terms, calculating expectation values requires reduced information about the wave function, captured in the one- and two-body reduced density matrices.
are two-body integrals, and a†µ and aµ are the creation and annihilation operator for the basis function ϕµ. Since Ĥ only depends on one- and two-body terms, calculating expectation values requires reduced information about the wave function, captured in the one- and two-body reduced density matrices.
Formally, a k-reduced density matrix (k-RDM) is a partial trace of the full density matrix ρ = |ψ〉〈 ψ| over N – k particles. The k-RDM is defined by
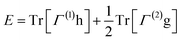 . This raises the question: is the information in k-RDMs enough to reconstruct the full N-body wave function? This is the k-body N-representability problem.45–49 Solving it for arbitrary k and N is NP-hard on classical computers50 and QMA-hard for quantum computers,49,51,52 but multiple heuristic methods are used to circumvent this issue.53–55
. This raises the question: is the information in k-RDMs enough to reconstruct the full N-body wave function? This is the k-body N-representability problem.45–49 Solving it for arbitrary k and N is NP-hard on classical computers50 and QMA-hard for quantum computers,49,51,52 but multiple heuristic methods are used to circumvent this issue.53–55
3 Information in many molecules
While the discussion so far has focused on a fixed quantum system with a defined number of particles, a foundation model that generalizes across chemistry must learn not only a representation of one system (as quantum tomography does), but also the shared representation across systems. This calls for a generalization of quantum tomography over Fock space, the space of quantum states across all molecular systems,56,57 and the corresponding space of observables across molecules. Such a formulation may provide a natural framework for designing foundation models that aim to learn transferable QPMs across chemistry. By newly establishing a connection with quantum tomography, t hese ideas extend the formalization of chemical compound space (CCS) introduced by von Lilienfeld et al..58–60 The underlying complexity scales exponentially with Hilbert space dimension, although this challenge is partially mitigated by restricting attention to chemically relevant states.61Fock space 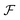 is defined as the direct sum of Hilbert spaces over varying particle number
is defined as the direct sum of Hilbert spaces over varying particle number 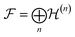 , where n enumerates “sectors” associated to a set of electrons and nuclei (cf.Fig. 1). Similarly, observables across molecules correspond to expectation values of operators acting on this Fock space. This unifying picture can guide architectural decisions in model design, helping ensure that learned representations respect physical principles already encoded in quantum theory. For instance, the indistinguishability of particles imply the foundation model should be permutationally invariant, and respect other symmetries depending on the systems being studied.
, where n enumerates “sectors” associated to a set of electrons and nuclei (cf.Fig. 1). Similarly, observables across molecules correspond to expectation values of operators acting on this Fock space. This unifying picture can guide architectural decisions in model design, helping ensure that learned representations respect physical principles already encoded in quantum theory. For instance, the indistinguishability of particles imply the foundation model should be permutationally invariant, and respect other symmetries depending on the systems being studied.
Another key aspect of this formulation is the notion of generalization across systems. All possible states of a molecule can be viewed as belonging to a sector. A foundation model should not only learn QPMs within single sectors (cf.eqn (2)), it should also be capable of learning the relationships between observables across different sectors,58,62
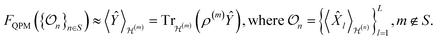 | (5) |
Such cross-sector generalization is particularly relevant for frameworks that seek to infer properties of large systems by leveraging knowledge about smaller “motifs”,63–71 where the target quantum state in sector m, encoded in ρ(m), can be inferred from an informationally rich set of observables measured in other sectors, 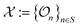 . This reflects the intuitive observation that motifs make similar contributions across different molecular systems,72 while corrections emerge from the interactions among these substructures.73 This is very well understood in quantum chemistry due to the locality of the coulomb operator, and for gapped systems the effects are restricted by such limits.74,75
. This reflects the intuitive observation that motifs make similar contributions across different molecular systems,72 while corrections emerge from the interactions among these substructures.73 This is very well understood in quantum chemistry due to the locality of the coulomb operator, and for gapped systems the effects are restricted by such limits.74,75
Graph neural networks leverage this modular perspective by representing molecules as atomistic graphs, where message passing architectures learn to aggregate and propagate local information across systems.12,76–79 Through iterative nonlinear transformations, these models aim to capture both local atomic environments and the aforementioned nonlocal correlations between molecular substructures. We formalize this idea with the deep tomography hypothesis discussed below.
4 The deep tomography hypothesis
Across vision, language, and materials, independently trained models on similar data tend to learn internal representations that are equivalent up to a simple transformation. This “alignment” has been observed empirically,80–85 allowing, for instance, re-using layers from one network into another with minimal adaptation,86,87 or translation between models' embedding spaces.88,89 Different lines of work frame this in various ways: from practical tools like model stitching to more abstract proposals like the Platonic representation hypothesis,90,91 its “strong” variant,89 or the tomographic interpretation.27 A recurring metaphor for this phenomenon is the Anna Karenina scenario: “all happy families are alike; each unhappy family is unhappy in its own way.”92 Well-generalizing models tend to converge on similar information-sufficient representations, while models that overfit, like “Clever Hans”,34 each fail in their own unique way. For a single descriptor, many distinct internal explanations, including unphysical shortcuts remain viable, as they can achieve low error on a single task.The unifying insight across these perspectives is that well-trained models tend to arrive at one of a family of approximate information-sufficient representations.25,93 This convergence arises from two complementary and interrelated mechanisms, which operate when supervision is mediated by a shared representation bottleneck: a common network trunk producing a single latent representation from which multiple targets are decoded. If targets are instead handled through separate disjoint networks the constraints do not accumulate on a common representation, and the convergence argument no longer applies. Moreover, the enforced targets must be sufficiently diverse: redundant or highly correlated descriptors do not impose additional constraints and thus do not meaningfully restrict the space of admissible representations.
4.1 Preservation of information
Integrating multiple views,90,94–102 greatly constrains the solution space. Every property![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) l encoded in the shared representation space restricts solutions θ* ∈ Θl to those minimizing the reconstruction error d(·, ·) for that property. Thus, in well-trained models every property
l encoded in the shared representation space restricts solutions θ* ∈ Θl to those minimizing the reconstruction error d(·, ·) for that property. Thus, in well-trained models every property ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) l must be approximately recoverable (cf.eqn (6) and Fig. 2):
l must be approximately recoverable (cf.eqn (6) and Fig. 2):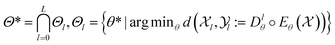 | (6) |
4.2 Preservation of structure
In practice, the reconstruction error for![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) l is minimized via a loss
l is minimized via a loss 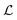 (e.g., mean squared error) over M molecular states:
(e.g., mean squared error) over M molecular states: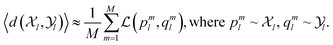 | (7) |
With limited data, an over-parameterized model can solve this by memorization (i.e., a lookup table), failing to generalize effectively. However, given sufficient samples and model capacity, it must instead learn to minimize the reconstruction error across the full random variable ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) l, enabling interpolation. This is only possible if the representation
l, enabling interpolation. This is only possible if the representation 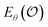 encodes information in a structured way, as discussed in the contrastive learning literature.94,103–105 Specifically, for a valid similarity metric Siml(·, ·), any ordering Siml(pal, pbl) ≤ Siml(pal, pcl) on realizations
encodes information in a structured way, as discussed in the contrastive learning literature.94,103–105 Specifically, for a valid similarity metric Siml(·, ·), any ordering Siml(pal, pbl) ≤ Siml(pal, pcl) on realizations 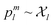 must be preserved in latent space by a corresponding metric
must be preserved in latent space by a corresponding metric 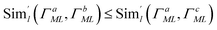 with
with 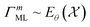 .
.
When the model integrates another property ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) l+1, generalization demands that the latent space also preserve its similarity ordering Siml+1, which may conflict with that of
l+1, generalization demands that the latent space also preserve its similarity ordering Siml+1, which may conflict with that of ![[X with combining circumflex]](https://www.rsc.org/images/entities/i_char_0058_0302.gif) l. This structural encoding of potentially competing contexts imposes additional constraints, further narrowing the viable solution space.
l. This structural encoding of potentially competing contexts imposes additional constraints, further narrowing the viable solution space.
These constraints define an equivalence class of representations under invertible transformations that aim to approximately preserve both mutual information and local structure. The joint requirement to maintain semantic organization while satisfying multiple, potentially competing objectives sharply reduces the space of admissible solutions, providing a natural explanation for the convergence observed across independently trained models and sets the basis of transfer learning,106–108 zero-shot learning88 and the justification of foundation models.101,109 According to the first postulate of Quantum Mechanics, the state of a physical system is completely described by its wavefunction. The k-RDM Γ(k) serves as the canonical object containing all k-body information about a quantum system.110 We therefore propose the Deep Tomography Hypothesis: large ML models trained on molecular properties derived from the k-RDM will, in the limit of sufficient capacity and data, converge to latent representations that are informationally complete, and thus informationally equivalent to Γ(k) on the subset of the chemical space they are trained on. Importantly, even when an informationally-complete representation may not be accessible, e.g., due to computational limits, or is not contained in a model's hypothesis space Θ, each model is biased towards the closest admissible approximation. Below, we formalize Hypothesis 1 and discuss its implications.
This hypothesis suggests that foundation models trained across many molecules and properties are, in effect, implementing a novel kind of cross-sector tomography. The model learns not only to interpolate within a sector of the Hilbert space, but also to extrapolate across different sectors of Fock space—a capacity that traditional quantum tomography does not possess, yet which is indispensable for scalable chemical prediction. Such models, when sufficiently expressive and trained on diverse molecular data, effectively extend the informational reach of quantum measurements: they enable predictions of quantum observables without direct quantum access to the system of interest, using knowledge transferred from related systems. This hypothesis should be understood as a statement on the limit behaviour, as in practice limitations regarding informational-completeness set of descriptors, model capacity and learning efficiency arise. In such non-ideal scenarios, the learned representations do not uniquely determine a sufficient statistics of the underlying quantum system consistent with all observed properties, but rather concentrate the probability mass over a subset of states consistent with the observations.
The conditions for the deep tomography hypothesis to hold rely on overcoming two bottlenecks: a data and a capacity bottleneck. The former assumes that we can reliably sample the massively large Fock space of chemicals. The latter assumes the model is sufficiently wide and deep. Since Hilbert spaces grow exponentially in the number of particles, a loose restriction on the underlying network is that it also be exponentially wide and deep. However, since these two limits are not feasible in practice, we believe the following:
• Width limit: since the dimension of the k-RDM is O(N2k), i.e., exponential in k and polynomial in the size of the underlying molecule, fixing a value for k will permit a reliable encoding of the k-RDM in a polynomial (in N) amount of neurons. The dimension of Γ(k) is the width of the model in this case, fixed for maximum size of the molecule considered.
• Depth limit: recovering the k-RDM from an informationally-complete set of properties is as expensive as a matrix inversion problem.111 However, recovering ρ = Γ(N) from Γ(k) is QMA hard (cf. Section 2). This means that in the worst case, a network would need exponentially many layers to recover ρ but polynomially many layers to recover Γ(k).
This means that, for a fixed k, neither bottleneck is expected to be exponentially hard—making it feasible for the deep tomography hypothesis to be tested. We expect that this convergence in representation will be physically meaningful, allowing polynomially-large models to achieve transfer-learning capabilities.112–114
5 Outlook
The bitter lesson115 has taught us that general ML models will get better as we scale compute and data.116,117 In chemistry, larger and diverse datasets will give rise to the next generation of ML models, those capable of combining all types of information about molecules, crystals, proteins, materials, and other forms of matter—including structure, numerical properties, or other descriptors.101 We believe these models will learn representations increasingly closer to informational-completeness, which we call deep tomographies, and these should be informationally equivalent to k-RDMs on the subset of the chemical space for which they are trained. Under this hypothesis, insights from quantum tomography, such as the role of information completeness, the structure of operator algebras, and the interpretability of reduced descriptions, can guide the design of better learning models. In particular, they motivate a move away from arbitrary descriptors toward physically grounded representations that preserve the relational structure among quantum observables.112,118,119Viewed through this lens, symmetries and physical laws can be understood as additional physics constraints imposed on the model either through its architecture or the objective function. The difference lies not in the solution itself, which under the perspective adopted here would ultimately exhibit the required symmetries if enough and diverse data is provided, but in the optimization dynamics. While architectural enforcement of physics constraints can be beneficial in data-scarce regimes by ruling out unphysical solutions when those are viable,120,121 such hard constraints may also restrict the optimization dynamics by disallowing temporary violations that could facilitate convergence toward a physically valid solution.122,123 In data-rich regimes, this raises the question whether softer, penalty-based enforcement of constraints may allow more flexibility during optimization while leveraging the guidance they provide. Understanding this trade-off remains an open problem.
This work is conceptually related to PAC learning (Valiant's model),124,125 while it addresses a different question. PAC learning assumes an unknown distribution 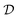 over a set of objects
over a set of objects 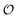 and a target function
and a target function 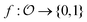 . The central concern is whether access to labelled sample pairs (o, f(o)) drawn from
. The central concern is whether access to labelled sample pairs (o, f(o)) drawn from 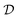 , a learning algorithm can output a hypothesis
, a learning algorithm can output a hypothesis 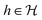 that with high probability achieves small generalization error, with emphasis on sample and computationally efficiency. In contrast, we argue that in molecular science target functions are not arbitrary elements but induced by a shared physical variable, the molecule's quantum state. Enforcing faithful prediction over an increasing number of diverse molecular descriptors through a shared learned representation object, progressively restricts the space of admissible hypothesis. Unphysical or shortcut solutions become incompatible with all tasks, driving convergence toward representations that act like sufficient statistics, locally, of the underlying physics.
that with high probability achieves small generalization error, with emphasis on sample and computationally efficiency. In contrast, we argue that in molecular science target functions are not arbitrary elements but induced by a shared physical variable, the molecule's quantum state. Enforcing faithful prediction over an increasing number of diverse molecular descriptors through a shared learned representation object, progressively restricts the space of admissible hypothesis. Unphysical or shortcut solutions become incompatible with all tasks, driving convergence toward representations that act like sufficient statistics, locally, of the underlying physics.
For quantum chemistry, this perspective hints at a path to foundation models that have tomographically-meaningful latent spaces, from which all other physical properties are recoverable. It suggests a principled and alternate benchmark procedure for representational completeness of foundation models beyond task-specific accuracy: freeze the encoder and train decoders to predict molecular properties not included during training. Models whose frozen representations support accurate reconstruction for a wider range of unseen observables can be regarded as more informationally-complete.
Such benchmarks can potentially bridge the gap between ML embeddings of molecules and quantum-mechanical state reconstructions—laying the groundwork for AI systems that operate as compressed yet faithful surrogates of the underlying physics. As the field advances, progress in foundation models for quantum chemistry coming from richer training data will enable the convergence towards deep tomographic representations. For example, the development of quantum computing will allow efficient computation of k-RDM properties of molecules126–130 providing training data that surpasses the accuracy achievable by any classical simulation131—relevant for highly-correlated systems like FeMoco.132 In such case, the ML architecture remains the same, but the dataset creation within a time-window is of higher quality. Another opportunity enabled by quantum processor would be access to learning algorithms with latent representations that are quantum in nature26 and emerging devices like the 25–50 logical qubit machine “MAGNE”133 would facilitate deep tomography representations learned from low-order measurements (1/2-RDMs) to serve as generalizable property-to-property maps, compressing complex many-body physics into experimentally grounded representations. In this second scenario, the ML model would be implemented as a quantum algorithm, and while we believe that the deep tomography hypothesis still hold—small changes on what is considered a decoder must be made. This would open a new practical route – or paradigm – to chemically reliable surrogates for strongly correlated motifs, supporting scalable prediction and validation of spin states, redox chemistry, and catalytic pathways in Fe–S type systems.134 In summary, larger models trained on such data or algorithms, encompassing diverse molecular properties, will be better positioned to converge towards platonic representations capturing the essential structure of quantum-mechanical information.
Conflicts of interest
There are no conflicts of interest to declare.Data availability
No new data was generated or analyzed in support of this research.Acknowledgements
The authors would like to acknowledge valuable discussions with Anatole von Lilienfeld. L. M. C. is supported by the Novo Nordisk Foundation, Grant number NNF22SA0081175, NNF Quantum Computing Programme. J. B. P. acknowledge funding of this project by the National Sciences and Engineering Research Council of Canada (NSERC) Alliance Grant #ALLRP587593-23. M. B. and A. A.-G. acknowledge funding of this project by the National Sciences and Engineering Research Council of Canada (NSERC) Alliance Consortia Quantum Grants #ALLRP587590-23. A. A. gratefully acknowledges King Abdullah University of Science and Technology (KAUST) for the KAUST Ibn Rushd Postdoctoral Fellowship. R. O. O. and T. V. acknowledge financial support from the Technical University of Denmark (DTU) through the Alliance PhD Scholarship, the Pioneer Center for Accelerating P2X Materials Discovery (CAPeX), DNRF Grant P3, and Novo Nordisk Foundation grants no. NNF24OC0089800 and NNF23SA0087929 (NitroScale), and the MIT-Danish Fellowship, administered by the Danish Ministry of Higher Education and Science to promote research collaboration between Denmark and the Massachusetts Institute of Technology (MIT). A. A.-G. and T. B. are CIFAR Fellows in the Accelerated Decarbonization Program; this research is based in part on work supported by CIFAR through a catalyst award. A. A.-G. thanks Anders G. Frøseth for his generous support. A. A.-G. also acknowledges the generous support of Natural Resources Canada and the Canada 150 Research Chairs program. This research is part of the University of Toronto's Acceleration Consortium, which receives funding from the CFREF-2022-00042 Canada First Research Excellence Fund.References
- E. S. Claude, A mathematical theory of communication, Bell Syst. Tech. J., 1948, 27(3), 379–423, DOI:10.1002/j.1538-7305.1948.tb00917.x.
- D. J. C. MacKay, Information Theory, Inference and Learning Algorithms, Cambridge University Press, 2003, ISBN 978-0521642989 Search PubMed.
- T. M. Cover, Elements Of Information Theory, John Wiley & Sons, 1999, ISBN 978-0471241959 Search PubMed.
- D. J. Griffiths and D. F. Schroeter, Introduction To Quantum Mechanics, Cambridge University Press, 2018, DOI:10.1017/9781316995433.
- J. Archibald Wheeler, Information, physics, quantum: The search for links, Feynman And Computation, 2018, pp. 309–336, DOI:10.1201/9780429500459-19.
- J. M. Renes, R. Blume-Kohout, A. J. Scott and C. M. Caves, Symmetric informationally complete quantum measurements, J. Math. Phys., 2004, 45(6), 2171–2180, DOI:10.1063/1.1737053.
- G. Mauro D'Ariano and P. Perinotti, Optimal data processing for quantum measurements, Phys. Rev. Lett., 2007, 98(2), 020403, DOI:10.1103/physrevlett.98.020403.
- J. Malmi, K. Korhonen, D. Cavalcanti and G. García-Pérez, Enhanced observable estimation through classical optimization of informationally overcomplete measurement data: Beyond classical shadows, Phys. Rev. A, 2024, 109(6), 062412, DOI:10.1103/physreva.109.062412.
- M. Paris and J. Rehacek, Quantum State Estimation, Springer Science & Business Media, 2004, 649, DOI:10.1007/b98673.
- J. Yuen-Zhou, J. K. Jacob, M. Mohseni and A. Aspuru-Guzik, Quantum state and process tomography of energy transfer systems via ultrafast spectroscopy, Proc. Natl. Acad. Sci. U. S. A., 2011, 108(43), 17615–17620, DOI:10.1073/pnas.1110642108.
- H.-Y. Huang, R. Kueng and J. Preskill, Predicting many properties of a quantum system from very few measurements, Nat. Phys., 2020, 16(10), 1050–1057, DOI:10.1038/s41567-020-0932-7.
- J. Gilmer, S. S. Schoenholz, P. F. Riley, O. Vinyals and G. E. Dahl, NeuralMessage Passing For Quantum Chemistry, arXiv, 2017, preprint, arXiv:1704.01212, DOI:10.48550/arXiv.1704.01212, https://arxiv.org/abs/1704.01212.
- D. K. Duvenaud, D. Maclaurin, J. Iparraguirre, R. Bombarell, T. Hirzel, A. Aspuru-Guzik, and R. P. Adams, Convolutional networks on graphs for learning molecular fingerprints, Advances In Neural Information Processing Systems, ed. C. Cortes, N. Lawrence, D. Lee, M. Sugiyama, and R. Garnett, Curran Associates, Inc., 2015, vol. 28, https://proceedings.neurips.cc/paper_files/paper/2015/file/f9be311e65d81a9ad8150a60844bb94c-Paper.pdf Search PubMed.
- Y. Li, D. Tarlow, M. Brockschmidt, and R. Zemel, Gated Graph Sequence Neural Networks, arXiv, 2017, preprint, arXiv:1511.05493, DOI:10.48550/arXiv.1511.05493, https://arxiv.org/abs/1511.05493.
- S. Kearnes, K. McCloskey, M. Berndl, V. Pande and P. Riley, Molecular graph convolutions: moving beyond fingerprints, J. Comput. Aided Mol. Des., 2016, 30(8), 595–608, DOI:10.1007/s10822-016-9938-8.
- K. T. Schütt, F. Arbabzadah, S. Chmiela, K. R. Müller and A. Tkatchenko, Quantum-chemical insights from deep tensor neural networks, Nat. Commun., 2017, 8(1), 13890, DOI:10.1038/ncomms13890.
- K. Schütt, P.- Jan Kindermans, H. E. S. Felix, S. Chmiela, A. Tkatchenko, and K.-R. Müller, Schnet: A continuous-filter convolutional neural network for modeling quantum interactions, Advances In Neural Information Processing Systems, I. Guyon, U. Von Luxburg, S. Bengio, H. Wallach, R. Fergus, S. Vishwanathan, and R. Garnett, Curran Associates, Inc., 2017, vol. 30, https://proceedings.neurips.cc/paper_files/paper/2017/file/303ed4c69846ab36c2904d3ba8573050-Paper.pdf Search PubMed.
- S. Ahmed, C. Sanchez Munoz, F. Nori and A. F. Kockum, Quantum state tomography with conditional generative adversarial networks, Phys. Rev. Lett., 2021, 127(14), 140502, DOI:10.1103/PhysRevLett.127.140502.
- P. Reiser, M. Neubert, A. Eberhard, L. Torresi, C. Zhou, C. Shao, H. Metni, C. van Hoesel, H. Schopmans and T. Sommer, et al., Graph neural networks for materials science and chemistry, Commun. Mater., 2022, 3(1), 93, DOI:10.1038/s43246-022-00315-6.
- R. Ashtari Mahini, G. Casanola-Martin, S. A. Ludwig and B. R. . Mixturemetrics, A comprehensive package to develop additive numerical features to describe complex materials for machine learning modeling, SoftwareX, 2024, 28, 101911, DOI:10.1016/j.softx.2024.101911 . URL https://www.sciencedirect.com/science/article/pii/S2352711024002814.
- A. Zunger, Inverse design in search of materials with target functionalities, Nat. Rev. Chem., 2018, 2(4), 0121, DOI:10.1038/s41570-018-0121.
- B. Sanchez-Lengeling and A. Aspuru-Guzik, Inverse molecular design using machine learning: Generative models for matter engineering, Science, 2018, 361(6400), 360–365, DOI:10.1126/science.aat2663.
- R. Gómez-Bombarelli, J. N. Wei, D. Duvenaud, J. M. Hernández-Lobato, B. Sánchez-Lengeling, D. Sheberla, J. Aguilera-Iparraguirre, T. D. Hirzel, R. P. Adams and A. Aspuru-Guzik, Automatic chemical design using a data-driven continuous representation of molecules, ACS Cent. Sci., 2018, 4(2), 268–276, DOI:10.1021/acscentsci.7b00572.
- N. Tishby, F. C. Pereira, and W. Bialek, The information bottleneck method, arXiv, 2000, preprint physics/0004057, DOI:10.48550/arXiv.physics/0004057.
- N. Tishby and N. Zaslavsky, Deep learning and the information bottleneck principle, In 2015 IEEE Information Theory Workshop (ITW), 2015, pp. 1–5, DOI:10.1109/ITW.2015.7133169.
- J. Romero, J. P. Olson and A. Aspuru-Guzik, Quantum autoencoders for efficient compression of quantum data, Quantum Sci. Technol., 2017, 2(4), 045001, DOI:10.1088/2058-9565/aa8072.
- R. Ortega-Ochoa, A. Aspuru-Guzik, T. Vegge, and T. Buonassisi, A tomographic interpretation of structure-property relations for materials discovery, arXiv, 2025, preprint, arXiv:2501.18163, DOI:10.48550/arXiv.2501.18163, https://arxiv.org/abs/2501.18163.
- M. Tulio Ribeiro, S. Singh, and C. Guestrin, why should i trust you?” explaining the predictions of any classifier, Proceedings Of The 22nd Acm Sigkdd International Conference On Knowledge Discovery And Data Mining, 2016, pp. 1135–1144, DOI:10.1145/2939672.2939778.
- S. M. Lundberg and S.-I. Lee, A unified approach to interpreting model predictions, Adv. Neural Inf. Process. Syst., 2017, 30, 4768–4777 Search PubMed.
- M. Esders, T. Schnake, J. Lederer, A. Kabylda, G. Montavon, A. Tkatchenko and K.-R. Müller, Analyzing atomic interactions in molecules as learned by neural networks, J. Chem. Theory Comput., 2025, 21(2), 714–729, DOI:10.1021/acs.jctc.4c01424.
- L. Tětková, T. Brüsch, T. Dorszewski, F. Martin Mager, R. Ø. Aagaard, J. Foldager, T. S. Alstrøm and L. K. Hansen, On convex decision regions in deep network representations, Nat. Commun., 2025, 16(1), 5419, DOI:10.1038/s41467-025-60809-y.
- W. D. Figg, E. Reed, S. Green, and J. M. Pluda. Thalidomide, Humana Press, Totowa, NJ, 1999, pp. 407–422, DOI:10.1007/978-1-59259-453-5_24.
- J. H. Kim and A. R. Scialli, Thalidomide: The tragedy of birth defects and the effective treatment of disease, Toxicol. Sci., 04 2011, 122(1), 1–6, DOI:10.1093/toxsci/kfr088.
- K. Jacob, J. Dippel, L. Ruff, W. Samek, K.-R. Müller and G. Montavon, Explainable ai reveals clever hans effects in unsupervised learning models, Nat. Mach. Intell., 2025, 7(3), 412–422, DOI:10.1038/s42256-025-01000-2.
- J. K. Winkler, C. Fink, F. Toberer, A. Enk, T. Deinlein, R. Hofmann-Wellenhof, L. Thomas, A. Lallas, A. Blum and S. Wilhelm, et al., Association between surgical skin markings in dermoscopic images and diagnostic performance of a deep learning convolutional neural network for melanoma recognition, JAMA Dermatol., 2019, 155(10), 1135–1141, DOI:10.1001/jamadermatol.2019.1735.
- T. Helgaker, P. Jorgensen, and J. Olsen, Molecular Electronic-Structure Theory, John Wiley & Sons, 2013, DOI:10.1002/9781119019572.
- C. J. Cramer, Essentials Of Computational Chemistry: Theories And Models, John Wiley & Sons, 2013. ISBN 978-0-470-09182-1 Search PubMed.
- J. Kempe, A. Kitaev and O. Regev, The complexity of the local hamiltonian problem, SIAM J. Comput., 2006, 35(5), 1070–1097, DOI:10.1007/978-3-540-30538-5_31.
- D. Aharonov, I. Arad and T. Vidick, Guest column: the quantum pcp conjecture, ACM SIGART Newsl., 2013, 44(2), 47–79, DOI:10.1145/2491533.249154.
- J. Watrous, Quantum computational complexity, Computational Complexity, Springer, 2012, pp. 2361–2387, DOI:10.1007/978-1-4614-1800-9_147.
- R. H. Douglas, The wave mechanics of an atom with a non-coulomb central field. part i. theory and methods, Mathematical Proceedings Of The Cambridge Philosophical Society, Cambridge University Press, 1928, vol. 24, pp. 89–110, DOI:10.1017/s0305004100011919.
- P. Hohenberg and W. Kohn, Inhomogeneous electron gas, Phys. Rev., 1964, 136(3B), B864, DOI:10.1103/physrev.136.b864.
- J. Čížek, On the correlation problem in atomic and molecular systems. calculation of wavefunction components in ursell-type expansion using quantum-field theoretical methods, J. Chem. Phys., 1966, 45(11), 4256–4266, DOI:10.1063/1.1727484.
- A. Peruzzo, J. McClean, P. Shadbolt, M.-H. Yung, X.-Qi Zhou, P. J. Love, A. Aspuru-Guzik and J. L. O’brien, A variational eigenvalue solver on a photonic quantum processor, Nat. Commun., 2014, 5(1), 4213, DOI:10.1038/ncomms5213.
- A. Klyachko, Quantum marginal problem and representations of the symmetric group, arXiv, 2004, preprint quant-ph/0409113, DOI:10.48550/arXiv.quant-ph/0409113.
- A. A. Klyachko, Quantum marginal problem and n-representability, Journal Of Physics: Conference Series, IOP Publishing, 2006, 36, p. 72, DOI:10.1088/1742-6596/36/1/014.
- C. A. Coulson, Present state of molecular structure calculations, Rev. Mod. Phys., 1960, 32(2), 170, DOI:10.1103/revmodphys.32.170.
- A. J. Coleman, Structure of fermion density matrices, Rev. Mod. Phys., 1963, 35(3), 668, DOI:10.1103/revmodphys.35.668.
- W. Tzu-Chieh, M. Mosca and A. Nayak, Interacting boson problems can be qma hard, Phys. Rev. Lett., 2010, 104(4), 040501, DOI:10.1103/physrevlett.104.040501.
- D. A. Mazziotti, Structure of fermionic density matrices: Complete n-representability conditions, Phys. Rev. Lett., 2012, 108(26), 263002, DOI:10.1103/PhysRevLett.108.263002.
- Yi-K. Liu, Consistency of local density matrices is qma-complete, in Approximation, Randomization, and Combinatorial Optimization. Algorithms and Techniques: 9th International Workshop on Approximation Algorithms for Combinatorial Optimization Problems, Approx 2006 and 10th International Workshop on Randomization and Computation, Random 2006, Barcelona, Spain, August 28-30 2006 Proceedings, Springer, 2006, pp. 438–449, DOI:10.1007/11830924_40.
- Yi-K. Liu, M. Christandl and F. Verstraete, Quantum computational complexity of the n-representability problem: Qma complete, Phys. Rev. Lett., 2007, 98(11), 110503, DOI:10.1103/PhysRevLett.98.110503.
- T. L. Gilbert, Hohenberg-kohn theorem for nonlocal external potentials, Phys. Rev. B, 1975, 12(6), 2111, DOI:10.1103/physrevb.12.2111.
- M. Nakata, H. Nakatsuji, M. Ehara, M. Fukuda, K. Nakata and K. Fujisawa, Variational calculations of fermion second-order reduced density matrices by semidefinite programming algorithm, J. Chem. Phys., 2001, 114(19), 8282–8292, DOI:10.1063/1.1360199.
- D. A. Mazziotti, Variational minimization of atomic and molecular ground-state energies via the two-particle reduced density matrix, Phys. Rev. A, 2002, 65(6), 062511, DOI:10.1103/physreva.65.062511.
- Al. Ivanov, The structure of chemical particles, J. Math. Chem., 2007, 42(2), 141–152, DOI:10.1007/s10910-005-9044-y.
- V. J. Härkönen, Quantum field theory of electrons and nuclei, J. Phys. A: Math. Theor., 2024, 57(46), 465402, DOI:10.1088/1751-8121/ad8a2c.
- O. Anatole von Lilienfeld, First principles view on chemical compound space: Gaining rigorous atomistic control of molecular properties, Int. J. Quantum Chem., 2013, 113(12), 1676–1689, DOI:10.1002/qua.24375.
- R. Ramakrishnan and O. Anatole von Lilienfeld, Machine Learning, Quantum Chemistry, And Chemical Space, John Wiley & Sons, Ltd, 2017, ch 5, pp. 225–256, ISBN 9781119356059 Search PubMed.
- B. Huang and O. Anatole von Lilienfeld, Ab initio machine learning in chemical compound space, Chem. Rev., 2021, 121(16), 10001–10036, DOI:10.1021/acs.chemrev.0c01303.
- A. Steffens, C. A. Riofrío, R. Hübener and J. Eisert, Quantum field tomography, New J. Phys., 2014, 16(12), 123010, DOI:10.1088/1367-2630/16/12/123010.
- N. Segal, A. Netanyahu, K. P. Greenman, P. Agrawal, and R. Gomez-Bombarelli, Known unknowns: Out-of-distribution property prediction in materials and molecules, arXiv, 2025, preprint arXiv:2502.05970, DOI:10.48550/arXiv.2502.05970.
- S. A. Wildman and G. M. Crippen, Prediction of physicochemical parameters by atomic contributions, J. Chem. Inf. Comput. Sci., 1999, 39, 868–873, DOI:10.1021/ci990307l.
- P. Ertl and A. Schuffenhauer, Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions, J. Cheminf., 2009, 1(1), 8, DOI:10.1186/1758-2946-1-8.
- M. Gastegger, C. Kauffmann, J. Behler and P. Marquetand, Comparing the accuracy of high-dimensional neural network potentials and the systematic molecular fragmentation method: A benchmark study for all-trans alkanes, J. Chem. Phys., 2016, 144(19), 194110, DOI:10.1063/1.4950815.
- R. Zubatyuk, J. S. Smith, J. Leszczynski and O. Isayev, Accurate and transferable multitask prediction of chemical properties with an atoms-in-molecules neural network, Sci. Adv., 2019, 5(8), eaav6490, DOI:10.1126/sciadv.aav6490.
- A. S. Christensen, L. A. Bratholm, F. A. Faber and O. Anatole von Lilienfeld, Fchl revisited: Faster and more accurate quantum machine learning, J. Chem. Phys., 2020, 152(4), 044107, DOI:10.1063/1.5126701.
- O. T. Unke, S. Chmiela, M. Gastegger, K. T. Schütt, H. E. Sauceda and K.-R. M. Spookynet, Learning force fields with electronic degrees of freedom and nonlocal effects, Nat. Commun., 2021, 12(1), 7273, DOI:10.1038/s41467-021-27504-0.
- P. Gao, J. Zhang, H. Qiu and S. Zhao, A general qspr protocol for the prediction of atomic/inter-atomic properties: a fragment based graph convolutional neural network (f-gcn), Phys. Chem. Chem. Phys., 2021, 23, 13242–13249, 10.1039/D1CP00677K.
- K.-D. Luong and A. Singh, Fragment-based pretraining and finetuning on molecular graphs, arXiv, 2023, preprint, arXiv:2310.03274, DOI:10.48550/arXiv.2310.03274, https://arxiv.org/abs/2310.03274.
- R. Ortega-Ochoa, T. Vegge, and J. Frellsen, Molminer: Towards controllable, 3d-aware, fragment-based molecular design, arXiv, 2025, preprint, arXiv:2411.06608, DOI:10.48550/arXiv.2411.06608, https://arxiv.org/abs/2411.06608.
- C. Borgelt and M. R. Berthold, Mining molecular fragments: finding relevant substructures of molecules, 2002 IEEE International Conference on Data Mining, 2002. Proceedings, 2002, pp. 51–58, DOI:10.1109/ICDM.2002.1183885.
- Z. Wu, J. Wang, H. Du, D. Jiang, Yu Kang, D. Li, P. Pan, Y. Deng, D. Cao, C.-Yu Hsieh and T. Hou, Chemistry-intuitive explanation of graph neural networks for molecular property prediction with substructure masking, Nat. Commun., 2023, 14(1), 2585, DOI:10.1038/s41467-023-38192-3.
- S. Ismail-Beigi and T. A. Arias, Locality of the density matrix in metals, semiconductors, and insulators, Phys. Rev. Lett., 1999, 82(10), 2127, DOI:10.1103/physrevlett.82.2127.
- J. Eisert, M. Cramer and M. B. Plenio, Colloquium: Area laws for the entanglement entropy, Rev. Mod. Phys., 2010, 82(1), 277–306, DOI:10.1103/revmodphys.82.277.
- J. Gilmer, S. S. Schoenholz, P. F. Riley, O. Vinyals, and G. E. Dahl, Message Passing Neural Networks, Springer International Publishing, Cham, 2020, pp. 199–214, ISBN 978-3-030-40245-7 Search PubMed.
- M. Tang, B. Li and H. Chen, Application of message passing neural networks for molecular property prediction, Curr. Opin. Struct. Biol., 2023, 81, 102616, DOI:10.1016/j.sbi.2023.102616 . https://www.sciencedirect.com/science/article/pii/S0959440X23000908.
- K. T. Schütt, H. E. Sauceda, P.-J. Kindermans, A. Tkatchenko and K.-R. Müller, Schnet – a deep learning architecture for molecules and materials, J. Chem. Phys., 2018, 148(24), 241722, DOI:10.1063/1.5019779.
- D. K. Duvenaud, D. Maclaurin, J. Iparraguirre, R. Bombarell, T. Hirzel, A. Aspuru-Guzik and R. P. Adams, Convolutional networks on graphs for learning molecular fingerprints, Adv. Neural Inf. Process. Syst., 2015, 28, 2224–2232 Search PubMed.
- C. Olah, Visualizing representations: Deep learning and human beings, 2015, http://colah.github.io/posts/2015-01-Visualizing-Representations/, Accessed: 2025-07-26.
- G. Roeder, L. Metz, and D. Kingma, On linear identifiability of learned representations, Proceedings Of The 38th International Conference On Machine Learning, Volume 139 Of Proceedings Of Machine Learning Research, M. Meila and T. Zhang, PMLR, 2021, pp. 9030–9039, https://proceedings.mlr.press/v139/roeder21a.html Search PubMed.
- Y. Li, J. Yosinski, J. Clune, H. Lipson, and J. Hopcroft, Convergent learning: Do different neural networks learn the same representations?, arXiv, 2016, preprint, arXiv:1511.07543, DOI:10.48550/arXiv.1511.07543, https://arxiv.org/abs/1511.07543.
- A. Dravid, Y. Gandelsman, A. A. Efros, and A. Shocher, Rosetta neurons: Mining the common units in a model zoo, arXiv, 2023, preprint, arXiv:2306.09346, DOI:10.48550/arXiv.2306.09346, https://arxiv.org/abs/2306.09346.
- A. Morcos, M. Raghu, and S. Bengio. Insights on representational similarity in neural networks with canonical correlation, Advances In Neural Information Processing Systems, S. Bengio, H. Wallach, H. Larochelle, K. Grauman, N. Cesa-Bianchi, and R. Garnett, Curran Associates, Inc., 2018, vol. 31, https://proceedings.neurips.cc/paper_files/paper/2018/file/a7a3d70c6d17a73140918996d03c014f-Paper.pdf Search PubMed.
- S. Yue, T. A. Keller, N. Sebe, and M. Welling. Topographic variational autoencoders, Structured Representation Learning: From Homomorphisms and Disentanglement to Equivariance and Topography, Springer, 2025, pp. 21–48, DOI:10.1007/978-3-031-88111-4_3.
- K. Lenc and A. Vedaldi, Understanding image representations by measuring their equivariance and equivalence, arXiv, 2015, preprint, arXiv:1411.5908, DOI:10.48550/arXiv.1411.5908, https://arxiv.org/abs/1411.5908.
- Y. Bansal, P. Nakkiran, and B. Barak, Revisiting model stitching to compare neural representations, Advances in Neural Information Processing Systems, M. Ranzato, A. Beygelzimer, Y. Dauphin, P. S. Liang, and J. Wortman Vaughan, Curran Associates, Inc., 2021, vol. 34, pp. 225–236, https://proceedings.neurips.cc/paper_files/paper/2021/file/01ded4259d101feb739b06c399e9cd9c-Paper.pdf Search PubMed.
- L. Moschella, V. Maiorca, M. Fumero, A. Norelli, F. Locatello, and E. Rodolà, Relative representations enable zero-shot latent space communication, International Conference on Learning Representations, 2023, https://openreview.net/forum?id=SrC-nwieGJ Search PubMed.
- R. Jha, C. Zhang, V. Shmatikov, and J. X. Morris, Harnessing the universal geometry of embeddings, arXiv, 2025, preprint, arXiv:2505.12540, DOI:10.48550/arXiv.2505.12540, https://arxiv.org/abs/2505.12540.
- M. Huh, B. Cheung, T. Wang, and P. Isola, The platonic representation hypothesis, arXiv, 2024, preprint, arXiv:2405.07987, DOI:10.48550/arXiv.2405.07987, https://arxiv.org/abs/2405.07987.
- Z. Liu and I. Chuang, Proof of a perfect platonic representation hypothesis, arXiv, preprint arXiv:2507.01098, 2025, DOI:10.48550/arXiv.2507.01098.
- Leo Tolstoy, Anna Karenina, 1878, DOI:10.7326/0003-4819-128-8-199804150-00008, First published in Russian; various English translations available.
- Ravid Shwartz-Ziv and Naftali Tishby, Opening the black box of deep neural networks via information, arXiv, 2017, preprint, arXiv:1703.00810, DOI:10.48550/arXiv.1703.00810, https://arxiv.org/abs/1703.00810.
- P. Bachman, R. Devon Hjelm, and W. Buchwalter. Learning representations by maximizing mutual information across views, Advances in Neural Information Processing Systems, H. Wallach, H. Larochelle, A. Beygelzimer, F. d’ Alché-Buc, E. Fox, and R. Garnett, Curran Associates, Inc., 2019, vol. 32, https://proceedings.neurips.cc/paper_files/paper/2019/file/ddf354219aac374f1d40b7e760ee5bb7-Paper.pdf Search PubMed.
- C. Xu, D. Tao, and C. Xu, A survey on multi-view learning, arXiv, 2013, preprint, arXiv:1304.5634, DOI:10.48550/arXiv.1304.5634, https://arxiv.org/abs/1304.5634.
- Y. Song, T. Anderson Keller, N. Sebe, and M. Welling, Topographic Variational Autoencoders, Springer, Nature Switzerland, Cham, 2026, pp. 21–48, ISBN 978-3-031-88111-4, DOI:10.1007/978-3-031-88111-4_3.
- Z. Yu, Z. Dong, C. Yu, K. Yang, Z. Fan and C. L. Philip Chen, A review on multi-view learning, Front. Comput. Sci., 2024, 19(7), 197334, DOI:10.1007/s11704-024-40004-w.
- T. Jin, V. Singla, H.-H. Hsu and B. M. Savoie, Large property models: a new generative machine-learning formulation for molecules, Faraday Discuss., 2025, 256, 104–119, 10.1039/D4FD00113C.
- H. Abu Alhaija, J. Alvarez, M. Bala, T. Cai, T. Cao, L. Cha, J. Chen, M. Chen, F. Ferroni, S. Fidler, D. Fox, Y. Ge, J. Gu, H. Ali, M. Isaev, P. Jannaty, S. Lan, T. Lasser, H. Ling, M.-Yu Liu, X. Liu, Y. Lu, A. Luo, Q. Ma, H. Mao, F. Ramos, X. Ren, T. Shen, X. Sun, S. Tang, T.-C. Wang, J. Wu, J. Xu, S. Xu, K. Xie, Y. Ye, X. Yang, X. Zeng, and Yu Zeng, Cosmos-transfer1: Conditional world generation with adaptive multimodal control, arXiv, 2025, preprint, arXiv:2503.14492, DOI:10.48550/arXiv.2503.14492, https://arxiv.org/abs/2503.14492.
- R. Chang, Yu-X. Wang and E. Ertekin, Towards overcoming data scarcity in materials science: unifying models and datasets with a mixture of experts framework, npj Comput. Mater., 2022, 8(1), 242, DOI:10.1038/s41524-022-00929-x.
- V. Moro, C. Loh, R. Dangovski, G. Ali, A. Ma, Z. Chen, S. Kim, P. Y. Lu, T. Christensen and M. Soljačić, Multimodal foundation models for material property prediction and discovery, Newton, 2025, 1(1), 100016, DOI:10.1016/j.newton.2025.100016.
- P. P. De Breuck, G. Hautier and G. M. Rignanese, Materials property prediction for limited datasets enabled by feature selection and joint learning with modnet, npj Comput. Mater., 2021, 7(1), 83, DOI:10.1038/s41524-021-00552-2.
- R. Hadsell, S. Chopra, and Y. LeCun, Dimensionality reduction by learning an invariant mapping, 2006 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR’06), 2006, vol. 2, pp. 1735–1742, DOI:10.1109/CVPR.2006.100.
- Aaron van den Oord, Y. Li, and O. Vinyals, Representation learning with contrastive predictive coding, arXiv, 2019, preprint, arXiv:1807.03748, DOI:10.48550/arXiv.1807.03748, https://arxiv.org/abs/1807.03748.
- A. Frome, G. S. Corrado, J. Shlens, S. Bengio, J. Dean, M. ’ A. Ranzato, and T. Mikolov, Devise: A deep visual-semantic embedding model, Advances in Neural Information Processing Systems, C. J. Burges, L. Bottou, M. Welling, Z. Ghahramani, and K. Q. Weinberger, Curran Associates, Inc., 2013, vol. 26, https://proceedings.neurips.cc/paper_files/paper/2013/file/7cce53cf90577442771720a370c3c723-Paper.pdf Search PubMed.
- L. Y. Pratt, Discriminability-based transfer between neural networks, Advances in Neural Information Processing Systems, S. Hanson, J. Cowan, and C. Giles, Morgan-Kaufmann, 1992, vol. 5, https://proceedings.neurips.cc/paper_files/paper/1992/file/67e103b0761e60683e83c559be18d40c-Paper.pdf Search PubMed.
- S. Jialin Pan and Q. Yang, A survey on transfer learning, IEEE Trans. Knowl. Data Eng., 2010, 22(10), 1345–1359, DOI:10.1109/TKDE.2009.191.
- E. O. Pyzer-Knapp, M. Manica, P. Staar, L. Morin, P. Ruch, T. Laino, J. R. Smith and A. Curioni, Foundation models for materials discovery – current state and future directions, npj Comput. Mater., 2025, 11(1), 61, DOI:10.1038/s41524-025-01538-0.
- I. Batatia, P. Benner, Y. Chiang, A. M. Elena, D. . P. Kovács, J. Riebesell, X. R. Advincula, M. Asta, M. Avaylon, W. J. Baldwin, F. Berger, N. Bernstein, A. Bhowmik, S. M. Blau, V. Cărare, J. P. Darby, S. De, F. Della Pia, V. L. Deringer, R. Elijošius, Z. El-Machachi, F. Falcioni, E. Fako, A. C. Ferrari, A. Genreith-Schriever, J. George, R. E. A. Goodall, C. P. Grey, P. Grigorev, S. Han, W. Handley, H. H. Heenen, K. Hermansson, C. Holm, J. Jaafar, S. Hofmann, K. S. Jakob, H. Jung, V. Kapil, A. D. Kaplan, N. Karimitari, J. R. Kermode, N. Kroupa, J. Kullgren, M. C. Kuner, D. Kuryla, G. Liepuoniute, J. T. Margraf, I.-B. Magdău, A. Michaelides, J. Harry Moore, A. A. Naik, S. P. Niblett, S. W. Norwood, N. O'Neill, C. Ortner, K. A. Persson, K. Reuter, A. S. Rosen, L. L. Schaaf, C. Schran, B. X. Shi, E. Sivonxay, T. K. Stenczel, V. Svahn, C. Sutton, T. D. Swinburne, J. Tilly, C. van der Oord, E. Varga-Umbrich, T. Vegge, M. Vondrák, Y. Wang, W. C. Witt, F. Zills, and G. . Csányi, A foundation model for atomistic materials chemistry, arXiv, 2024, preprint, arXiv:2401.00096, DOI:10.48550/arXiv.2401.00096, https://arxiv.org/abs/2401.00096.
- J. Wetherell, A. Costamagna, M. Gatti and L. Reining, Insights into one-body density matrices using deep learning, Faraday Discuss., 2020, 224, 265–291, 10.1039/d0fd00061b.
- Z. Hradil, Quantum-state estimation, Phys. Rev. A, 1997, 55(3), R1561, DOI:10.1103/physreva.55.r1561.
- M. Tsubaki and T. Mizoguchi, Quantum deep field: data-driven wave function, electron density generation, and atomization energy prediction and extrapolation with machine learning, Phys. Rev. Lett., 2020, 125(20), 206401, DOI:10.1103/physrevlett.125.206401.
- M. Tsubaki and T. Mizoguchi, Quantum deep descriptor: Physically informed transfer learning from small molecules to polymers, J. Chem. Theory Comput., 2021, 17(12), 7814–7821, DOI:10.1021/acs.jctc.1c00568.
- N. C. Frey, R. Soklaski, S. Axelrod, S. Samsi, R. Gomez-Bombarelli, C. W. Coley and V. Gadepally, Neural scaling of deep chemical models, Nat. Mach. Intell., 2023, 5(11), 1297–1305, DOI:10.26434/chemrxiv-2022-3s512.
- R. Sutton, The bitter lesson, Incomplete Ideas (blog), 2019, vol. 13, iss. 1, p. 38 Search PubMed.
- J. Kaplan, S. McCandlish, T. Henighan, T. B. Brown, B. Chess, R. Child, S. Gray, A. Radford, J. Wu, and D. Amodei. Scaling laws for neural language models. arXiv, 2020, preprint arXiv:2001.08361, DOI:10.48550/arXiv.2001.08361.
- A. H. Cheng, C. T. Ser, M. Skreta, A. Guzmán-Cordero, L. Thiede, A. Burger, A. Aldossary, S. X. Leong, S. Pablo-Garcia and F. Strieth-Kalthoff, et al., Spiers memorial lecture: How to do impactful research in artificial intelligence for chemistry and materials science, Faraday Discuss., 2025, 256, 10–60, 10.1039/d4fd00153b.
- K. T. Schütt, M. Gastegger, A. Tkatchenko, K.-R. Müller and R. J. Maurer, Unifying machine learning and quantum chemistry with a deep neural network for molecular wavefunctions, Nat. Commun., 2019, 10(1), 5024, DOI:10.1038/s41467-019-12875-2.
- W. Yu, E. Abdelaleem, I. Nemenman and J. C. Burton, Physics-tailored machine learning reveals unexpected physics in dusty plasmas, Proc. Natl. Acad. Sci. U. S. A., 2025, 122(31), e2505725122, DOI:10.1073/pnas.2505725122.
- T. Cohen and M. Welling, Group equivariant convolutional networks, Proceedings of The 33rd International Conference on Machine Learning, volume 48 of Proceedings of Machine Learning Research, M. F. Balcan and K. Q. Weinberger, PMLR, New York, New York, USA, 2016, pp. 2990–2999, https://proceedings.mlr.press/v48/cohenc16.html Search PubMed.
- B. Elesedy and S. Zaidi, Provably strict generalisation benefit for equivariant models, Proceedings of the 38th International Conference on Machine Learning, volume 139 of Proceedings of Machine Learning Research, M. Meila and T. Zhang, PMLR, 2021, pp. 2959–2969, https://proceedings.mlr.press/v139/elesedy21a.html Search PubMed.
- S. Pertigkiozoglou, E. Chatzipantazis, S. Trivedi and K. Daniilidis, Improving equivariant model training via constraint relaxation, Adv. Neural Inf. Process. Syst., 2024, 37, 83497–83520 Search PubMed.
- A. A. A. Elhag, T. Konstantin Rusch, F. Di Giovanni, and M. M. Bronstein, Relaxed equivariance via multitask learning, In ICLR 2025 Workshop on Machine Learning for Genomics Explorations, 2025, https://openreview.net/forum?id=8kZSO4WbTh Search PubMed.
- L. G. Valiant, A theory of the learnable, Commun. ACM, 1984, 27(11), 1134–1142, DOI:10.1145/1968.1972.
- A. S. Rocco, The probably approximately correct learning model in computational learning theory, arXiv, 2025, preprint, arXiv:2511.08791, DOI:10.48550/arXiv.2511.08791, https://arxiv.org/abs/2511.08791.
- X. Bonet-Monroig, R. Babbush and T. E. O'Brien, Nearly optimal measurement scheduling for partial tomography of quantum states, Phys. Rev. X, 2020, 10, 031064, DOI:10.1103/PhysRevX.10.031064 . https://link.aps.org/doi/10.1103/PhysRevX.10.031064.
- A. Zhao, N. C. Rubin and A. Miyake, Fermionic partial tomography via classical shadows, Phys. Rev. Lett., 2021, 127(11), 110504, DOI:10.1103/PhysRevLett.127.110504.
- E. Knill, G. Ortiz and R. D. Somma, Optimal quantum measurements of expectation values of observables, Phys. Rev. A, 2007, 75, 012328, DOI:10.1103/PhysRevA.75.012328 , https://link.aps.org/doi/10.1103/PhysRevA.75.012328.
- A. Alase, R. R. Nerem, M. Bagherimehrab, P. Høyer and B. C. Sanders, Tight bound for estimating expectation values from a system of linear equations, Phys. Rev. Res., 2022, 4, 023237, DOI:10.1103/PhysRevResearch.4.023237 ., https://link.aps.org/doi/10.1103/PhysRevResearch.4.023237.
- W. J. Huggins, K. Wan, J. McClean, T. E. O'Brien, N. Wiebe and R. Babbush, Nearly optimal quantum algorithm for estimating multiple expectation values, Phys. Rev. Lett., 2022, 129(24), 240501, DOI:10.1103/physrevlett.129.240501.
- H.-Y. Huang, R. Kueng, G. Torlai, V. V. Albert and J. Preskill, Provably efficient machine learning for quantum many-body problems, Science, 2022, 377(6613), eabk3333, DOI:10.1126/science.abk3333.
- M. Reiher, N. Wiebe, K. M. Svore, D. Wecker and M. Troyer, Elucidating reaction mechanisms on quantum computers, Proc. Natl. Acad. Sci., 2017, 114(29), 7555–7560, DOI:10.1073/pnas.1619152114.
- 2025, https://novonordiskfonden.dk/en/news/eifo-and-the-novo-nordisk-foundation-acquire-the-worlds-most-powerful-quantum-computer/.
- Y. Alexeev, V. S. Batista, N. Bauman, L. Bertels, D. Claudino, R. Dutta, L. Gagliardi, S. Godwin, N. Govind, M. Head-Gordon, M. Hermes, K. Kowalski, Li Ang, C. Liu, J. Liu, P. Liu, J. M. Garcia-Lustra, D. Mejia-Rodriguez, K. Mueller, M. Otten, Bo Peng, M. Raugus, M. Reiher, R. Paul, W. Shaw, M. van Schilfgaarde, T. Vegge, Yu Zhang, M. Zheng, and L. Zhu, A perspective on quantum computing applications in quantum chemistry using 25–100 logical qubits, arXiv, 2025, preprint, arXiv:2506.19337, DOI:10.48550/arXiv.2506.19337, https://arxiv.org/abs/2506.19337.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |