 Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Assessing the performance of quantum-mechanical descriptors in physicochemical and biological property prediction
Alejandra
Hinostroza Caldas
 a,
Artem
Kokorin
a,
Artem
Kokorin
 b,
Alexandre
Tkatchenko
b,
Alexandre
Tkatchenko
 *b and
Leonardo
Medrano Sandonas†
*b and
Leonardo
Medrano Sandonas†
 *b
*b
aUniversidad Nacional de Ingeniería, Av. Túpac Amaru 210, Rímac, Lima 15333, Peru
bDepartment of Physics and Materials Science, University of Luxembourg, L-1511 Luxembourg City, Luxembourg. E-mail: alexandre.tkatchenko@uni.lu; leonardo.medrano@tu-dresden.de
First published on 19th January 2026
Abstract
Machine learning (ML) approaches have drastically advanced the exploration of structure–property and property–property relationships in computer-aided drug discovery. A central challenge in this field is the identification of molecular descriptors that can effectively capture both geometric- and electronic structure-derived features, enabling the development of reliable and interpretable predictive models. While numerous descriptors focusing solely on structural characteristics have been recently proposed, improvements in model accuracy often come at the cost of increased computational demands, thereby restricting their practical applicability. To address this challenge, we introduce the “QUantum Electronic Descriptor” (QUED) framework, which integrates both structural and electronic data of molecules to develop ML regression models for property prediction. In doing so, a quantum-mechanical (QM) descriptor is derived from molecular and atomic properties computed using the semi-empirical density functional tight-binding (DFTB) method, which allows for efficient modelling of both small and large drug-like molecules. This descriptor is combined with inexpensive geometric descriptors—capturing two-body and three-body interatomic interactions—to form comprehensive molecular representations used to train Kernel Ridge Regression and XGBoost models. As a proof of concept, we validate QUED using the QM7-X dataset, which comprises equilibrium and non-equilibrium conformations of small drug-like molecules, demonstrating that incorporating electronic structure data notably enhances the accuracy of ML models for predicting physicochemical properties. For biological endpoints, we find that QM properties provide some predictive value for toxicity and lipophilicity prediction, as assessed using the TDCommons-LD50 and the MoleculeNet benchmark datasets. Moreover, a SHapley Additive exPlanations (SHAP) analysis of the toxicity and lipophilicity predictive models reveals that molecular orbital energies and DFTB energy components are among the most influential electronic features. Hence, our work underscores the importance of incorporating QM descriptors to enhance both the accuracy and interpretability of ML models for predicting multiple properties relevant to pharmaceutical and biological applications.
1 Introduction
Quantum-mechanical (QM) descriptors have emerged as powerful tools in molecular property prediction, effectively bridging theoretical physics and chemistry with practical applications in drug discovery1–4 and material science.5–8 These descriptors are derived from the electronic structure of molecules, which is obtained by solving the Schrödinger equation. While this process can be computationally demanding—particularly for large and flexible drug-like molecules—advancements in computational chemistry have made it more tractable through approximate methods such as density functional theory (DFT)9,10 and semi-empirical (SE) methods.11,12 A critical input for these QM methods is the 3D spatial arrangement of atoms forming a molecular system, which is often overlooked in the development of predictive models within computer-aided drug discovery.13,14 Traditional descriptors used in this context are typically based on easily computable molecular and atomic features, such as SMILES (Simplified Molecular Input Line Entry System) strings,15 molecular weight,16 Morgan fingerprints,17 and Fukui functions.18 Despite their practicality, these descriptors lack the mechanistic insight into short- and long-range molecular interactions that electronic structure-based descriptors can provide. As a result, QM descriptors are attracting increasing attention for predicting physicochemical properties,8,19 environmental-related properties,20,21 and ADMET (Absorption, Distribution, Metabolism, Excretion, and Toxicity) endpoints.22–25Among the ADMET endpoints, toxicity stands out as a critical factor due to its direct implications for drug safety, spanning from mild adverse effects to severe, life-threatening outcomes. Indeed, pre-clinical and clinical (animal and human) toxicity issues account for over 30% of drug attrition,26 highlighting the urgent need for reliable early-stage prediction methods. Late-stage failures not only entail significant losses in time and resources but also often require revisiting earlier development phases, even when compounds have shown favorable pharmacokinetic properties.27,28 Consequently, computer-aided screening is essential for identifying potential toxicity risk early in the drug discovery pipeline, thereby supporting more efficient compound prioritization and structural optimization. Recent studies have emphasized the predictive value of descriptors that encode electronic properties, such as molecular orbital energies, polarization, reactivity, and total energy, in modelling toxicity across various biological datasets.29–34 These features are particularly relevant for capturing complex covalent interactions, which are central to many toxicological endpoints. Moreover, the increasing availability of large and curated toxicity datasets (e.g., ToxBenchmark,35 MoleculeNet36) has spurred the development of numerous machine learning (ML) frameworks for toxicity prediction.15,37,38
A critical challenge in developing ML models for property prediction that incorporate conformational sampling lies in the structural representation of conformers. Specifically, one must define a multidimensional function that transforms discrete atomic information—such as Cartesian coordinates and nuclear charges—into a task-appropriate structural representation, commonly referred to as a geometric descriptor39,40 These transformations must satisfy several essential criteria: they should preserve fundamental physical symmetries (i.e., be invariant under translations, rotations, and permutations of identical atoms), ensure smoothness (so that small changes in atomic positions lead to small changes in the descriptor), and be complete (ensuring that distinct molecular configurations are not mapped to the same representation).41,42 Early examples of such descriptors include the Coulomb Matrix (CM)43 and the Bag-of-Bonds (BOB),44 which achieve rotational and translational invariance by encoding two-body coulombic interactions. Building on these foundations, more expressive and permutationally invariant many-body descriptors45—such as the Spectrum of London and Axilrod–Teller–Muto (SLATM) potentials46 and the Faber–Christensen–Huang–Lilienfeld (FCHL) representation—have demonstrated higher accuracy in predicting both extensive and intensive physicochemical properties of small drug-like molecules. More sophisticated descriptors have since been introduced,39,47 offering even greater predictive performance. However, these improvements often come at the expense of increased computational cost, particularly when applied to large and flexible molecules common in pharmaceutical and biological contexts. To address this challenge in the development of ML models for biomedical endpoint prediction, one promising strategy is to augment inexpensive geometric descriptors with QM-derived features. This hybrid approach can enhance both accuracy and generalizability while maintaining computational efficiency. Furthermore, due to their inherent simplicity and informative nature, such combined descriptors may also facilitate the development of more interpretable ML models for toxicity prediction.48
In this work, we introduce the “QUantum Electronic Descriptor” (QUED) framework, which integrates structural features and electronic structure data of molecules to develop ML regression models for property prediction (see Fig. 1). Our primary goal is to evaluate the performance and interpretability of these hybrid descriptors in predicting physicochemical properties and biological responses of drug-like molecules. To this end, we employ inexpensive structural descriptors such as BOB and SLATM to capture essential geometrical information. These are complemented by an electronic descriptor (DQM), composed of molecular and atomic properties computed using the semi-empirical QM method density functional tight-binding (DFTB).49 Both types of descriptors are used in combination with kernel ridge regression (KRR) and XGBoost algorithms to investigate different strategies for enhancing predictive reliability. As a proof-of-concept, we first assess QUED ability to predict extensive and intensive physicochemical properties of both equilibrium and non-equilibrium small molecules from the QM7-X dataset.50 The gained insights are then leveraged to evaluate the effectiveness of hybrid descriptors for toxicity prediction in large and flexible drug-like molecules from the LD50 dataset.51,52 We further explore QUED potential to predict lipophilicity using data from the MoleculeNet benchmark.36 To interpret model predictions and identify key electronic features relevant to target properties, we employ SHapley Additive exPlanations (SHAP).53 Our results demonstrate that DFTB-derived electronic descriptors capture subtle molecular interactions that purely geometrical representations often miss, thereby enhancing the performance of specific ML regression models. In particular, molecular orbital energies and DFTB energy components are key contributors to this improvement. Overall, QUED offers a robust and interpretable framework for developing predictive models of physicochemical properties and biological responses.
2 Methods
2.1 The QUED framework
2.1.1.1 Kernel ridge regression (KRR). As part of the QUED framework, we have developed the
 toolbox that can be used to train ML models for property prediction using the KRR method (also known as the ‘kernel trick’).54 Various features, including kernel functions, molecular descriptors, and metrics, have been implemented to capture the unknown structure-to-property relationships in complex molecular systems.
toolbox that can be used to train ML models for property prediction using the KRR method (also known as the ‘kernel trick’).54 Various features, including kernel functions, molecular descriptors, and metrics, have been implemented to capture the unknown structure-to-property relationships in complex molecular systems.  toolbox also considers a quasi-Newton algorithm such as the limited-memory Broyden–Fletcher–Goldfarb–Shanno (BFGS) for hyperparameter optimization. Within
toolbox also considers a quasi-Newton algorithm such as the limited-memory Broyden–Fletcher–Goldfarb–Shanno (BFGS) for hyperparameter optimization. Within  , the target property array ŷ is given as
, the target property array ŷ is given as | (1) |
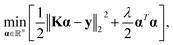 | (2) |
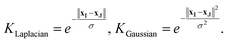 | (3) |
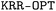 algorithm. The determination of the optimal set of hyperparameters relies on the simultaneous optimization of the hyperparameters, training set, and validation set. To carry out this process,
algorithm. The determination of the optimal set of hyperparameters relies on the simultaneous optimization of the hyperparameters, training set, and validation set. To carry out this process, 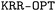 considers a given number of randomly distributed molecular sets. To train the KRR models, each benchmark dataset was randomly split into training, validation, and test sets. The number of samples allocated to the training and validation sets is reported in Table S2 of the SI, while the remaining samples were used for testing. The
considers a given number of randomly distributed molecular sets. To train the KRR models, each benchmark dataset was randomly split into training, validation, and test sets. The number of samples allocated to the training and validation sets is reported in Table S2 of the SI, while the remaining samples were used for testing. The 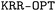 toolbox can be accessed in the QUED Github repository.
toolbox can be accessed in the QUED Github repository.
2.1.1.2 eXtreme gradient boosting (XGBoost). As a member of the gradient boosting category, at each boosting iteration, the XGBoost algorithm55 augments the model with a new tree ft(x) by minimizing a regularized objective function given by
 | (4) |
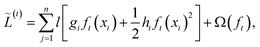 | (5) |
 | (6) |
 and
and  being the sum of gradients and Hessians of all samples in leaf j, λ and γ acting as regularization parameters, and T denoting the total number of leaves. Optimizing eqn (6) with respect to the leaf weight wj by setting its derivative to zero yields the optimal weight
being the sum of gradients and Hessians of all samples in leaf j, λ and γ acting as regularization parameters, and T denoting the total number of leaves. Optimizing eqn (6) with respect to the leaf weight wj by setting its derivative to zero yields the optimal weight | (7) |
Hyperparameter tuning was executed via a Bayesian optimization framework implemented in the Optuna56 package. In each iteration, a five-fold cross-validation was employed, with the objective of maximizing the negative root mean square error. Furthermore, each predictive model underwent 100 iterations of this optimization process. For all benchmark datasets, the training set sizes matched those used to develop the KRR models. Training samples were here selected using the farthest point sampling (FPS) technique,57 and the remaining samples constituted the test set. More details about the hyperparameter optimization can be found in Table S3 of SI. The module performing XGBoost calculations will be integrated into the 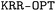 toolbox in a later version.
toolbox in a later version.
 | ||
| Fig. 2 Scheme of the QUED framework used to predict physicochemical and biological properties from structural and electronic descriptors. QUED integrates geometric representations, such as Bag-of-Bonds (BOB) and the Spectrum of London and Axilrod–Teller–Muto potential (SLATM), with a quantum mechanical (QM) descriptor (DQM) computed at the DFTB3 + MBD level. Predictive models are trained using Kernel Ridge Regression (KRR) and eXtreme Gradient Boosting (XGBoost) methods on two datasets, including QM7-X50 for physicochemical properties of small organic molecules and TDCommons-LD50 (ref. 51) for experimental acute toxicity values of drug-like compounds. Molecular size distributions and elemental compositions are shown for these datasets. We have also considered the MoleculeNet-Lipophilicity benchmark dataset,36 plots are provided in Fig. S6 of SI. | ||
2.1.2.1 Geometric representations. The two-body molecular descriptor Bag-of-Bonds (BOB), as well as the two and three-body descriptor SLATM, were used in the present work. BOB, a vectorized molecular representation, was introduced as a slightly more complex and improved version of the Coulomb Matrix (CM) representation,43 inspired by an ML approach in text processing bag-of-words.44 This descriptor is obtained by sorting into ‘bags’ (i.e., individual vectors) the types of bonds between pairs of atoms, which comprise a CM element. Each bag contains a single type of bond, and the bags are concatenated; the end of the vector is padded with zeros, so as to obtain the same vector length independent of the size of the molecules in a given dataset. However, the BOB representation is solely based on the atomic numbers and interatomic distances and, therefore, still lacks more precise spatial information about the molecule.
The SLATM descriptor involves a two-body term, which is a function of the coordinates and atomic numbers of the constituent atoms, and a three-body term, which includes a van der Waals potential contribution based on the Axilrod–Teller–Muto three-body potential.46 The presence of a three-body term and thus the inclusion of van-der-Waals interactions in the molecular descriptor indicate a much more elaborate picture of the impact of the surrounding environment on each atom. Accordingly, SLATM has proven to be a more complex molecular representation that yields better performance in ML models, albeit with increased computational costs for its generation and higher running costs due to larger sizes compared to two-body descriptors like BOB. However, SLATM still only considers neighboring atoms and is therefore not prohibitively expensive, making it suitable for extensive benchmark studies with multiple components where varying parameters and running calculations are required. Although this work primarily focuses on BOB and SLATM as baseline geometric representations, we additionally benchmark selected performance metrics against those obtained using the Smooth Overlap of Atomic Positions (SOAP) descriptor.58
2.1.2.2 Electronic representation. The second type of descriptor focuses on the electronic structure features of a given molecular system. Here, our main purpose is to define a reliable and efficient electronic descriptor that does not required large computational overheads rising from highly accurate QM methods (e.g., DFT, coupled-cluster) for which single-point calculations of big conformational datasets of large drug-like molecules are not sustainable for high-throughput studies. Accordingly, the QM properties of molecular conformations are calculated using the semi-empirical third-order DFTB method (DFTB3)59,60 supplemented with a treatment of many-body dispersion (MBD) for van der Waals interactions,61,62 as it is implemented in the
 code.63 The versatile performance of DFTB method has already been demonstrated in several works.50,64–69 For instance, DFTB simulations were used to gain a molecular understanding of the temperature-gradient degradation of polyethylene and polypropylene and to evaluate the subsequent oxidative upcycling reactions.64 DFTB + MBD was also recently used to investigate the relative stability of native states of several proteins in explicit solvation.65 Similarly, DFTB has been successfully applied to examine the electrostatic interactions and charge transfer in artificial molecular devices66,67 This method has also been extended to investigate excited-state properties, computing electronic transition dipole moments for organic chromophores.68
code.63 The versatile performance of DFTB method has already been demonstrated in several works.50,64–69 For instance, DFTB simulations were used to gain a molecular understanding of the temperature-gradient degradation of polyethylene and polypropylene and to evaluate the subsequent oxidative upcycling reactions.64 DFTB + MBD was also recently used to investigate the relative stability of native states of several proteins in explicit solvation.65 Similarly, DFTB has been successfully applied to examine the electrostatic interactions and charge transfer in artificial molecular devices66,67 This method has also been extended to investigate excited-state properties, computing electronic transition dipole moments for organic chromophores.68
Single-point DFTB3 + MBD calculations were carried out for all molecular systems studied in this work by considering hydrogen correction and the electronic Hamiltonian described by the 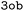 parameters set.70 The QM properties extracted from the DFTB output files are listed in Table 1, and have been divided into global properties (Dglob), molecular orbital energies (DeMO), and atomic properties (Datom). To create a standardized representation across the dataset, Mulliken charges arrays (whose dimension depends on the number of atoms in the molecule) are zero-padded to match the array size corresponding to the largest molecular structure in the dataset, so that all properties are included in a fixed-size array. This ensures consistency in the descriptor representation, allowing for effective input to ML models.
parameters set.70 The QM properties extracted from the DFTB output files are listed in Table 1, and have been divided into global properties (Dglob), molecular orbital energies (DeMO), and atomic properties (Datom). To create a standardized representation across the dataset, Mulliken charges arrays (whose dimension depends on the number of atoms in the molecule) are zero-padded to match the array size corresponding to the largest molecular structure in the dataset, so that all properties are included in a fixed-size array. This ensures consistency in the descriptor representation, allowing for effective input to ML models.
| Subset | Label | Property name | Dim |
|---|---|---|---|
| Global (Dglob) | E Fermi | Fermi energy | 1 |
| E band | Band energy | 1 | |
| NE | Number of electrons | 1 | |
| E H0 | Reference density energy | 1 | |
| E scc | Self-consistent charge energy | 1 | |
| E 3rd | Third-order correction energy | 1 | |
| E rep | Repulsion energy | 1 | |
| E mbd | Many-body interaction energy | 1 | |
| ‖µTB‖ | Scalar dipole moment | 1 | |
| E TBgap | HOMO–LUMO energy gap | 1 | |
| MO energies (DeMO) | ε | Molecular orbital energies | 8 |
| Atomic (Datom) | Q | Atomic Mulliken charges | N |
2.2 Benchmark datasets
To understand the effect of considering both geometric and electronic descriptors on the performance of ML regression models, we have first investigated the accuracy in predicting highly accurate physicochemical properties of equilibrium and non-equilibrium conformations of small drug-like molecules contained in QM7-X dataset.50 Later, we generated QM structural and property data of molecular conformations associated with SMILES representation of large drug-like molecules extracted from LD50 toxicity51,52 and lipophilicity datasets.36Additionally, we investigated the prediction of lipophilicity in drug-like molecules. This property describes the tendency of a compound to partition into a non-polar lipid matrix rather than an aqueous matrix.73 Lipophilicity is strongly correlated with key physicochemical and biochemical properties such as permeability and solubility, which in turn influence drug potency, distribution, and elimination. Consequently, it is frequently measured in QSAR studies to better characterize the pharmacological profiles of drug-like compounds.74 The lipophilicity dataset from the MoleculeNet benchmark36 contains experimental measurements of the octanol/water distribution coefficient (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D at pH 7.4) for 4200 compounds. These compounds contain heavy atom counts (B, C, N, O, F, Cl, P, S, Se, Si, Br, and I) ranging from 7 to 100 and log
D at pH 7.4) for 4200 compounds. These compounds contain heavy atom counts (B, C, N, O, F, Cl, P, S, Se, Si, Br, and I) ranging from 7 to 100 and log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D values spanning −1.5 to 4.5. Eleven compounds were discarded because they contained Si (similar to TDCommons dataset) or B, P, or Se, which were underrepresented in the dataset (see summary of discarded molecules in Table S1 of the SI).
D values spanning −1.5 to 4.5. Eleven compounds were discarded because they contained Si (similar to TDCommons dataset) or B, P, or Se, which were underrepresented in the dataset (see summary of discarded molecules in Table S1 of the SI).
To achieve a more precise modelling and representation of these drug-like molecules for predicting toxicity and lipophilicity, we generated their 3D molecular structures. The initial 3D structures were created using the 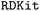 package, which constructs the graph of heavy atoms based on the connectivity information encoded in SMILES strings, adds hydrogen atoms, and optimizes the resulting geometry using the Merck Molecular Force Field (MMFF). For the next step, we considered only the molecules that were successfully generated and optimized using MMFF. While calculating ground-state energies and geometries via ab initio methods would offer greater accuracy, the computational demands are prohibitive. As a practical alternative, we employed the Conformer–Rotamer Ensemble Sampling Tool (CREST),75,76 which performs a comprehensive conformational search. CREST plays a crucial role in QUED by efficiently sampling both low- and high-energy conformers with a level of complexity beyond that of well-known conformational search methods based on classical force fields.77 This advantage arises from its more accurate treatment of long-range interactions (electrostatics and dispersion) and its ability to incorporate solvent effects. Indeed, CREST integrates the semi-empirical extended tight-binding method GFN2-xTB78 with a metadynamics-based search algorithm. We have applied energy (12.0 kcal mol−1) and root-mean-square deviation (RMSD) (0.1 Å) thresholds relative to the input structure to determine which geometries are included in the final Conformer–Rotamer Ensemble (CRE). All geometry optimization and conformational search calculations were performed using the GBSA implicit solvent model for water. As a result, 98.5% (7273 unique molecules) of the toxicity dataset and 97.0% (4073 unique molecules) of the lipophilicity dataset were successfully retrieved, yielding approximately 3.6 M and 1.8 M conformers, respectively. We then applied an RMSD-based hierarchical clustering method to refine these extensive sets, selecting ≈ 1.8 M and 618k representative conformers. This clustering approach ensures that the chosen conformers effectively represent the diversity of the explored conformational space. For each target property, the ML regression models were trained using only the conformer with the lowest DFTB3+MBD energy for each unique molecule. The computational costs associated with dataset generation are summarized in Fig. S5 of the SI.
package, which constructs the graph of heavy atoms based on the connectivity information encoded in SMILES strings, adds hydrogen atoms, and optimizes the resulting geometry using the Merck Molecular Force Field (MMFF). For the next step, we considered only the molecules that were successfully generated and optimized using MMFF. While calculating ground-state energies and geometries via ab initio methods would offer greater accuracy, the computational demands are prohibitive. As a practical alternative, we employed the Conformer–Rotamer Ensemble Sampling Tool (CREST),75,76 which performs a comprehensive conformational search. CREST plays a crucial role in QUED by efficiently sampling both low- and high-energy conformers with a level of complexity beyond that of well-known conformational search methods based on classical force fields.77 This advantage arises from its more accurate treatment of long-range interactions (electrostatics and dispersion) and its ability to incorporate solvent effects. Indeed, CREST integrates the semi-empirical extended tight-binding method GFN2-xTB78 with a metadynamics-based search algorithm. We have applied energy (12.0 kcal mol−1) and root-mean-square deviation (RMSD) (0.1 Å) thresholds relative to the input structure to determine which geometries are included in the final Conformer–Rotamer Ensemble (CRE). All geometry optimization and conformational search calculations were performed using the GBSA implicit solvent model for water. As a result, 98.5% (7273 unique molecules) of the toxicity dataset and 97.0% (4073 unique molecules) of the lipophilicity dataset were successfully retrieved, yielding approximately 3.6 M and 1.8 M conformers, respectively. We then applied an RMSD-based hierarchical clustering method to refine these extensive sets, selecting ≈ 1.8 M and 618k representative conformers. This clustering approach ensures that the chosen conformers effectively represent the diversity of the explored conformational space. For each target property, the ML regression models were trained using only the conformer with the lowest DFTB3+MBD energy for each unique molecule. The computational costs associated with dataset generation are summarized in Fig. S5 of the SI.
3 Results and discussion
3.1 Assessing QM descriptors for small molecules
We first explore the impact of combining geometric and electronic representations on the performance of ML regression models in predicting both extensive (atomization energy, EAT, and molecular polarizability, α) and intensive (dipole moment, µ, and HOMO–LUMO gap, Egap) physicochemical properties of small drug-like molecules from the QM7-X dataset (see Fig. 3 and S1 of SI). Two molecular subsets were analyzed: one consisting of equilibrium conformations (EQ) and the other of highly distorted non-equilibrium conformations (NEQ), each containing approximately 42k structures. Fig. 3A, D and S1A, D show the distributions of the target properties across both subsets. Our results indicate that combining baseline representations (BOB or SLATM) with electronic descriptors (DQM), i.e., BOB ⊕DQM and SLATM ⊕DQM, consistently improves prediction accuracy across all regression tasks when using KRR method. Among all models, SLATM ⊕DQM yields the highest accuracy. The benefit of including electronic descriptors is particularly significant for the NEQ subset, where the mean absolute error (MAE) is reduced by approximately 60% on average for all properties except α. This underscores the difficulty of capturing the relationship between strongly distorted molecular geometries and their properties using geometric features alone. In these cases, electronic descriptors derived from DFTB calculations enhance the connectivity between data points, leading to a more meaningful mapping between the high-dimensional feature space and the target properties. In contrast, for the EQ subset, incorporating electronic descriptors yields a more pronounced improvement for intensive properties than for extensive ones. Overall, the results obtained using the XGBoost method follow similar trends to those from KRR. However, while KRR slightly outperforms XGBoost in predicting extensive properties, XGBoost performs better for intensive properties (see dotted and dashed horizontal lines in Fig. 3 and S1 of SI). Table 2 presents the top results for predicting dipole moments and polarizabilities.| Target | Dataset | Regression method | Descriptor | MAE | R 2 |
|---|---|---|---|---|---|
| Dipole moment | EQ | KRR | SLATM ⊕DQM | 0.018 | 0.987 |
| D QM | 0.024 | 0.979 | |||
| XGBoost | SLATM ⊕DQM | 0.017 | 0.988 | ||
| D QM | 0.023 | 0.982 | |||
| NEQ | KRR | SLATM ⊕DQM | 0.059 | 0.924 | |
| D QM | 0.065 | 0.911 | |||
| XGBoost | SLATM ⊕DQM | 0.056 | 0.933 | ||
| D QM | 0.062 | 0.919 | |||
| Polarizability | EQ | KRR | SLATM ⊕DQM | 0.178 | 0.999 |
| D QM | 0.426 | 0.994 | |||
| XGBoost | BOB ⊕DQM | 0.377 | 0.995 | ||
| D QM | 0.552 | 0.988 | |||
| NEQ | KRR | SLATM ⊕DQM | 0.814 | 0.988 | |
| D QM | 1.657 | 0.958 | |||
| XGBoost | SLATM ⊕DQM | 1.030 | 0.984 | ||
| D QM | 1.620 | 0.957 |
We now turn to the prediction of µ, which serves as a clear example of how QM-derived features can enhance molecular representations for predicting intensive properties (results for Egap can be found in Tables S4, S5 and Fig. S1 of SI). As shown in Fig. 3B and C, DQM significantly outperforms the purely geometric descriptors BOB and SLATM, achieving MAE values of 0.024 and 0.064 eÅ for EQ and NEQ subsets, respectively, when using KRR models trained on 25k samples. Combining geometric and electronic descriptors further improves the prediction accuracy, yielding MAE values of 0.018 and 0.059 eÅ with the SLATM ⊕DQM. Similarly, XGBoost models confirm the advantage of incorporating QM-derived information: while DQM alone achieves an MAE of 0.023 eÅ, adding two-body interactions reduces this to 0.018 eÅ, and including three-body features further improves it to 0.017 eÅ, demonstrating consistently improved performance for µ prediction of EQ subset (similar trend is also found for NEQ subset). In contrast, for extensive properties, purely geometric descriptors generally outperform DQM across learning curves. When combining both types of descriptors, MAE values either remain unchanged or improve only marginally relative to the geometric descriptor alone. Notable improvements are mainly observed at larger training set sizes (16k and 25k), particularly for the SLATM representation. For instance, in predicting α for EQ subset (see Fig. 3E), SLATM shows a consistent improvement of approximately 15% when combined with DQM, achieving an MAE value of 0.18 a03. For NEQ subset (see Fig. 3F), both BOB and SLATM representations show enhanced performance when combined with DQM, with MAEs decreasing from 1.08 to 0.99 a03 and from 0.92 to 0.81 a03, respectively. Unlike µ prediction, XGBoost models generally yield higher errors than KRR models for extensive properties. Interestingly, for EQ subset, the best performance is achieved by the combination BOB ⊕DQM (MAE = 0.377 a03), diverging from the trend observed in other tasks, where SLATM ⊕DQM typically performs better. A similar behavior is seen in the prediction of EAT (see Tables S4, S5 and Fig. S1 of SI). Although the addition of DQM does not consistently outperform geometric descriptors for EQ subset, its value becomes more evident for NEQ subset. These findings are further supported by KRR models employing delta learning (see Table S6 of SI) and the XGBoost models using SOAP descriptor (see Table S10 of SI), which show consistent improvements in predictive accuracy when incorporating DQM.
3.1.1.1 Descriptor components. To determine which components of the DQM descriptor most significantly enhance model performance, we partitioned it into three subsets: global (Dglob), MO energies (DeMO), and atomic (Datom) properties. A detailed list of these properties is provided in Table 1. We assessed the performance of the geometric descriptor SLATM both “pure” and in combination with each property subset, i.e., SLATM ⊕Dglob, SLATM ⊕DeMO, and SLATM ⊕Datom, as well as with the full QM descriptor (SLATM ⊕DQM). The results from the KRR models trained on 16k samples are shown in Fig. 4 and 5 for the prediction of µ and α, respectively.
Optimal performance in physicochemical property prediction was generally achieved by combining the SLATM representation with the full electronic descriptor. However, for EQ subset, more accurate estimates of µ and α—20.5 × 10−3 eÅ and 0.18 a03, respectively—were obtained using SLATM ⊕Dglob, compared to 20.9 × 10−3 eÅ and 0.21 a03 for SLATM ⊕DQM. As shown in the boxplots of Fig. 4A and 5A, both models produce comparable error ranges, with SLATM ⊕Dglob exhibiting a slightly narrower error distribution. For NEQ subset, the global feature set also leads to substantial performance improvements when combined with SLATM, ranking second only to SLATM ⊕DQM. We attribute this improvement to the inclusion of the TB dipole moment and many-body dispersion energy in Dglob, which show strong correlations with the reference DFT-PBE0 µ (ρ(µ, µTB) = 0.94) and α (ρ(α, Embd) = −0.61), respectively (see other ρ values in Fig. S3 of SI). Interestingly, even though Dglob includes the HOMO–LUMO energy gap at the DFTB3 level (ETBgap)–a property that shows only moderate correlation with the DFT-PBE0 energy gap–SLATM ⊕DeMO outperforms all other models for predicting Egap. On the other hand, incorporating Mulliken charges (Q) generally degrades performance and appears to introduce noise, even though the target properties are related to the spatial distribution of charge. In most tasks, the SLATM ⊕Datom model performs worse than SLATM alone, likely due to the weak correlation between Q values and the target properties (see Fig. S3 of SI). A similar trend is observed for DeMO in the prediction of µ and α, which may also be correlated to the overall weak correlation of these properties with MO energies. However, SLATM ⊕DeMO model does outperform SLATM for predicting EAT and Egap, being the effect more remarkable for NEQ subset (see Fig. S2 of SI). These findings suggest that, while strong physical correlations with the target properties are important, model performance also depends critically on how electronic structure information is represented and integrated with geometric descriptors.
3.1.1.2 SHapley additive exPlanations (SHAP). Tree-based predictive ML models, such as XGBoost, greatly benefit from being coupled with interpretability tools like the SHAP method.53 Rooted in cooperative game theory, SHAP leverages Shapley values to quantify the contribution of each input feature to an individual prediction. In the context of our feature-based ML model, SHAP values estimate how each feature influences the deviation of a specific prediction from the expected output of the model. This enables a transparent interpretation of the learned relationships, revealing both the relative importance of features and how they interact to shape predicted outcomes. Fig. 4 and 5 present beeswarm plots that summarize the distribution of SHAP values for the most influential features in each property prediction task, based solely on DQM. In these plots, features are ordered by importance (top to bottom), and their SHAP values are shown along the x-axis. A positive SHAP value indicates that the feature pushes the prediction higher, while a negative value suggests it drives the prediction lower. The color gradient encodes the feature values: red denotes high values, and blue denotes low values.
Fig. 4C and D show that µTB is the most influential feature in predicting the DFT-PBE0 dipole moment (µ). This result is expected, as both quantities represent the same physical observable, albeit computed using different QM methods. In the EQ subset, we observe that higher values of the DFTB repulsion energy (Erep) and the third-order correction energy (E3rd) are associated with larger dipole moments. In contrast, higher values of the self-consistent charge energy (Escc) and many-body dispersion energy (Embd) tend to reduce the predicted µ. Notably, the TB-derived MO energies also rank among the top ten relevant features. A similar pattern is observed for the NEQ subset, where Mulliken charges (Q) also emerge as important contributors to the predictive performance. SHAP analysis for α predictions reveals a slightly different hierarchy of feature importance between EQ and NEQ subsets (see Fig. 5C and D). Overall, low values of Mulliken charges tend to negatively impact the predicted polarizability, with the notable exception of Q7, which deviates from this trend. We identify Embd as a key feature in this regression task: low values are linked to smaller polarizabilities, while higher values correlate with increased polarizability. This is consistent with the moderate negative correlation between Embd and α (ρ(Embd, α) = −0.61). Interestingly, the reference DFTB energy (EH0) also plays a significant role, likely due to its influence on the electron density distribution and its response to perturbations. In NEQ subset, the number of electrons contributes positively to the prediction, whereas it does not rank among the top 11 features for the EQ subset. Surprisingly, for EQ subset, the sixth MO energy ε6 gains more relevance, indicating that different structural regimes may be governed by distinct sets of driving QM features.
3.2 Predicting biological responses of large molecules
We now examine QUED performance to predict biological endpoints: toxicity and lipophilicity. To this end, we develop ML regression models using chemically diverse sets of large drug-like molecules from the TDCommons-LD50 dataset and the MoleculeNet-Lipophilicity dataset. These models were trained on the lowest-energy geometries (as determined by DFTB3+MBD) for each unique molecule.The distribution of LD50 values in this subset is shown in Fig. 6A. Fig. 6B and C present the learning curves for DQM and BOB ⊕DQM and SLATM ⊕DQM, using the KRR and XGBoost methods, respectively. In this task, XGBoost models consistently outperformed their KRR counterparts. This performance difference between XGBoost and KRR models is further illustrated in Fig. 6D and E, which display box plots of residuals (predicted minus true toxicity values) for different descriptor combinations. The KRR residuals show a broader spread, indicating higher prediction variance and reduced precision, whereas the XGBoost residuals are more tightly clustered around zero. Under KRR, the inclusion of geometric information improves the performance of the electronic descriptor. For instance, DQM alone yields an MAE of 0.539, which decreases to 0.469 and 0.445 when combined with BOB and SLATM, respectively. Interestingly, the combination SLATM ⊕DQM slightly underperforms pure SLATM, which achieves an MAE of 0.433—suggesting that in this case, the addition of DQM may not be beneficial. In contrast, under XGBoost, DQM achieves an MAE of 0.473. BOB shows improved performance when combined with DQM, achieving the best overall result with an MAE of 0.400—an improvement over the 0.451 obtained with pure BOB. In contrast, SLATM does not benefit from this addition: pure SLATM reaches an MAE of 0.403, slightly better than SLATM ⊕DQM, which yields 0.413.
 | ||
| Fig. 6 Prediction of acute toxicity (LD50) of large drug-like molecules from TDCommons-LD50 dataset.72 Panel (A) shows the LD50 distribution, colored by the number of non-hydrogen atoms. Panels (B) and (C) present learning curves for KRR and XGBoost models, respectively, using DQM, BOB ⊕DQM, and SLATM ⊕DQM. Shaded areas highlight improvements from adding DQM to geometric descriptors. Panels (D) and (E) display the corresponding residual distributions (prediction – true). In this task, adding DQM improves the performance of BOB but not SLATM, highlighting that the benefit of electronic information is not only task-specific but also descriptor-dependent. For these calculations, we used only the lowest-energy conformation of each unique molecule in the TDCommons-LD50 dataset. | ||
On the other side, geometric descriptors show no relevant improvement when combined with DQM for lipophilicity prediction (see Fig. S4 of SI). Alike toxicity results, XGBoost consistently outperforms KRR, e.g., pure DQM yields MAEs of 0.784 and 0.617 with KRR and XGBoost, respectively. For KRR, the best performance is achieved with SLATM ⊕DQM (MAE = 0.476), followed closely by pure SLATM (MAE = 0.480). For XGBoost, the best model overall is pure SLATM (MAE = 0.418), with performance slightly reduced upon inclusion of DQM (MAE = 0.432).
Table 3 summarizes the best results for predicting toxicity and lipophilicity. Our XGBoost model using BOB ⊕DQM achieves an MAE of 0.400 on the TDCommons-LD50 dataset, surpassing previous state-of-the-art approaches. These include 2D graph neural networks,79 which achieved an MAE of 0.45 when considering only the top 5% most confident predictions; equivariant transformers38 with an MAE of 0.653; and fingerprint-based surrogate models,80,81 reporting MAEs of 0.497 and an RMSE of 0.697, respectively. Furthermore, combining pure SLATM with XGBoost yields the best predictive accuracy for lipophilicity, with an RMSE of 0.567. This result surpasses the benchmark set by MoleculeNet using Extended-Connectivity Fingerprints with XGBoost (0.799) and is comparable to their results using graph-convolutional methods (0.655).36 While more complex state-of-the-art architectures reach comparable errors, such as convolutional neural networks trained on augmented SMILES representations82 (RMSE = 0.593), graph neural networks and multitask learning83 (RMSE = 0.537), 3D molecular representation learning framework Uni-Mol84 (RMSE = 0.603), and nested connected hierarchical GNN DenseNGN85 (MAE = 0.351), our approach remains competitive due to its simplicity and computational efficiency. Notice that regression models trained with the SOAP descriptor using XGBoost (see Table S10 in the SI) and with the state-of-the-art equivariant neural network MACE86 (see Table S11 in the SI) exhibited lower performance compared to the top-performing models summarized in Table 3.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) D) prediction using Kernel Ridge Regression (KRR) and XGBoost methods with different molecular descriptors
D) prediction using Kernel Ridge Regression (KRR) and XGBoost methods with different molecular descriptors
| Target | Regression method | Descriptor | MAE | RMSE | R 2 |
|---|---|---|---|---|---|
| Toxicity | KRR | SLATM | 0.433 | 0.606 | 0.595 |
| D QM | 0.539 | 0.719 | 0.430 | ||
| XGBoost | BOB ⊕DQM | 0.400 | 0.571 | 0.661 | |
| D QM | 0.473 | 0.637 | 0.572 | ||
| Lipophilicity | KRR | SLATM ⊕DQM | 0.476 | 0.661 | 0.643 |
| D QM | 0.784 | 0.991 | 0.339 | ||
| XGBoost | SLATM | 0.418 | 0.567 | 0.752 | |
| D QM | 0.617 | 0.813 | 0.519 |
The correlation between lipophilicity and the properties in the electronic descriptor is even weaker compared to toxicity (see ρ values in Fig. S3 of SI). Consequently, adding DeMO or Datom to the pure BOB descriptor results in larger MAEs, i.e., 0.585 and 0.593, respectively (see Table S8 of SI). In contrast, SLATM ⊕DeMO and SLATM ⊕Datom achieve slightly better performances (0.478 and 0.479) than pure SLATM. For both geometric descriptors, the inclusion of Dglob has no significant impact on performance. These results indicate that the benefit of incorporating QM features depends on the base geometric descriptor and target biological response; hence, the integration should be adapted to their specific characteristics.
4 Conclusions
In this work, we introduced the “QUantum Electronic Descriptor” (QUED) framework, which integrates both structural and electronic molecular information to develop ML regression models for physicochemical and biological property prediction. Central to QUED was the definition of a QM descriptor derived from molecular and atomic properties computed using the semi-empirical DFTB3 method supplemented with a many-body dispersion (MBD) treatment for van der Waals interactions. Indeed, to form comprehensive molecular representations, we combined this QM descriptor with computationally inexpensive geometric descriptors that capture two-body and three-body interatomic interactions, such as BOB and SLATM. As a proof of concept, we validated QUED performance by using two molecular subsets of the QM7-X dataset, which includes both equilibrium and non-equilibrium conformations of small drug-like molecules. The results demonstrated that incorporating electronic structure information significantly improves the accuracy of ML models in predicting physicochemical properties compared to only considering geometric features. In particular, combining SLATM with DQM led to a notable accuracy improvement, especially for highly distorted molecular structures. For QM7-X molecules, XGBoost models followed similar trends to those obtained by KRR models trained using toolbox. However, while KRR slightly outperforms XGBoost in predicting extensive properties, XGBoost performs better for intensive properties. Moreover, a detailed analysis combining property subsets with SHAP method revealed that certain electronic features are more relevant for specific target physicochemical properties, e.g., global properties play a more crucial role than MO energies or atomic charges in predicting µ, whereas atomic charges and DFTB energy components are more important for predicting α.
toolbox. However, while KRR slightly outperforms XGBoost in predicting extensive properties, XGBoost performs better for intensive properties. Moreover, a detailed analysis combining property subsets with SHAP method revealed that certain electronic features are more relevant for specific target physicochemical properties, e.g., global properties play a more crucial role than MO energies or atomic charges in predicting µ, whereas atomic charges and DFTB energy components are more important for predicting α.
QUED framework was also evaluated on the TDCommons-LD50 and MoleculeNet-Lipophilicity datasets to predict the toxicity levels and lipophilicity of larger and more chemically diverse drug-like molecules, respectively. Here, the benefits and insights of using QM descriptors for biological property prediction were more nuanced. SHAP analysis also confirmed that DFTB properties, such as MO energies and energy components, play a central role in these performance gains, with BOB ⊕DQM combined with XGBoost yielding the best performance for toxicity prediction. Overall, our findings highlight the importance of incorporating electronic structure data into ML workflows to enhance the reliability and interpretability of the predictive models. While geometric descriptors capture spatial patterns effectively, they may miss subtle electronic effects that are critical for accurately modelling complex molecular properties. That said, the computational demands associated with QM descriptor generation—even when using semi-empirical methods—can be a bottleneck for high-throughput workflows. To address this trade-off between descriptor complexity and computational efficiency, future research should explore strategies to optimize both aspects. One promising direction is the integration of ML-accelerated electronic structure methods,91–93 which can significantly reduce the time required to compute QM descriptors. Additionally, ML-enhanced DFTB approaches offer a way to improve the accuracy of QM properties without significantly increasing computational cost.94,95 Within the QUED framework, an analysis of computational time for each step revealed that conformational sampling with CREST represents the most time-consuming component (see Fig. S5 in the SI). In this context, generative AI models could aid conformational exploration, thereby increasing both the diversity and quality of biological datasets.96–98 Hence, we expect that the QUED framework can be extended to predict a wide range of biological endpoints, such as ADMET properties (beyond toxicity and lipophilicity), and protein–ligand interactions, further demonstrating the versatility and impact of integrating electronic structure information into molecular ML approaches.
Author contributions
The work was initially conceived by AH and LMS, and designed with contributions from AT. AK developed the KRR–OPT toolbox. AH and LMS generated the quantum-mechanical datasets, trained the regression models, and analyzed their performance. AT and LMS supervised and revised all stages of the work. AH and LMS drafted the original manuscript. All authors discussed the results and contributed to the final manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
All datasets, source code, running examples, and trained ML models presented in this work are available in the QUED GitHub repository (https://github.com/lmedranos/QUED) and are also publicly available on Zenodo under the DOI https://doi.org/10.5281/zenodo.17106019.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00411j.
Acknowledgements
A. H. C. and L. M. S. are grateful to the Research Experience for Peruvian Undergraduates (REPU) program for its organizational support. We acknowledge financial support from the Luxembourg National Research Fund via FNR “MBD-in-BMD C23/MS/18093472” project as well as from the European Research Council (ERC Advanced Grant 101054629 – “FITMOL”). The results presented in this publication have been obtained using computational resources provided by the Center for Information Services and High-Performance Computing (ZIH) at TU Dresden. This research also used the HPC facilities of the University of Luxembourg. The authors would like to express their gratitude to Li Chen for his guidance in the implementation of XGBoost models.Notes and references
- Z. Qiao, M. Welborn, A. Anandkumar, F. R. Manby and T. F. Miller, J. Chem. Phys., 2020, 153, 12 CrossRef PubMed.
- O. D. Abarbanel and G. R. Hutchison, J. Chem. Theory Comput., 2024, 20, 6946–6956 CrossRef CAS PubMed.
- S.-C. Li, H. Wu, A. Menon, K. A. Spiekermann, Y.-P. Li and W. H. Green, J. Am. Chem. Soc., 2024, 146, 23103–23120 CrossRef CAS PubMed.
- L. Medrano Sandonas, J. Hoja, B. G. Ernst, A. Vazquez-Mayagoitia, R. A. DiStasio and A. Tkatchenko, Chem. Sci., 2023, 14, 10702 RSC.
- Y. Zhang and X. Xu, Polym. Chem., 2021, 12, 843–851 RSC.
- Y. Guan, C. W. Coley, H. Wu, D. Ranasinghe, E. Heid, T. J. Struble, L. Pattanaik, W. H. Green and K. F. Jensen, Chem. Sci., 2021, 12, 2198–2208 Search PubMed.
- H. Shimakawa, A. Kumada and M. Sato, npj Comput. Mater., 2024, 10, 11 CrossRef.
- H.-C. Chang, M.-H. Tsai and Y.-P. Li, J. Chem. Inf. Model., 2025, 65, 1367–1377 Search PubMed.
- A. D. Becke, J. Chem. Phys., 2014, 140, 18A301 CrossRef PubMed.
- R. O. Jones, Rev. Mod. Phys., 2015, 87, 897–923 CrossRef.
- W. Thiel, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2014, 4, 145–157 CAS.
- A. V. Akimov and O. V. Prezhdo, Chem. Rev., 2015, 115, 5797–5890 CrossRef CAS PubMed.
- G. Maggiora, M. Vogt, D. Stumpfe and J. Bajorath, J. Med. Chem., 2014, 57, 3186–3204 CrossRef CAS PubMed.
- K. V. Chuang, L. M. Gunsalus and M. J. Keiser, J. Med. Chem., 2020, 63, 8705–8722 CrossRef CAS PubMed.
- J. Born, G. Markert, N. Janakarajan, T. B. Kimber, A. Volkamer, M. R. Martínez and M. Manica, Digital Discovery, 2023, 2, 674–691 RSC.
- O. Daoui, S. Elkhattabi, S. Chtita, R. Elkhalabi, H. Zgou and A. T. Benjelloun, Heliyon, 2021, 7, year CrossRef PubMed.
- E. Lounkine, M. J. Keiser, S. Whitebread, D. Mikhailov, J. Hamon, J. L. Jenkins, P. Lavan, E. Weber, A. K. Doak and S. Côté, et al. , Nature, 2012, 486, 361–367 CrossRef CAS PubMed.
- T. Stuyver and C. W. Coley, J. Chem. Phys., 2022, 156, 084104 CrossRef CAS PubMed.
- O. A. von Lilienfeld, K.-R. Müller and A. Tkatchenko, Nat. Rev. Chem., 2020, 4, 347–358 CrossRef PubMed.
- B. Muthiah, S.-C. Li and Y.-P. Li, J. Taiwan Inst. Chem. Eng., 2023, 151, 105123 CrossRef CAS.
- M.-H. Tsai, Y.-H. Lin and Y.-P. Li, ChemRxiv, 2025, preprint, ChemRxiv:chemrxiv-2025-hw9ff, DOI:10.26434/chemrxiv-2025-hw9ff.
- B. Tang, S. T. Kramer, M. Fang, Y. Qiu, Z. Wu and D. Xu, J. Cheminform., 2020, 12, 1–9 Search PubMed.
- C. Isert, J. C. Kromann, N. Stiefl, G. Schneider and R. A. Lewis, ACS Omega, 2023, 8, 2046–2056 CrossRef CAS PubMed.
- C. W. Coley, R. Barzilay, W. H. Green, T. S. Jaakkola and K. F. Jensen, J. Chem. Inf. Model., 2017, 57, 1757–1772 CrossRef CAS PubMed.
- M. A. Ghanavati, S. Ahmadi and S. Rohani, Digital Discovery, 2024, 3, 2085–2104 RSC.
- I. Kola and J. Landis, Nat. Rev. Drug Discovery, 2004, 3, 711–716 Search PubMed.
- F. P. Guengerich, Drug Metab. Pharmacokinet., 2011, 26, 3–14 CrossRef CAS PubMed.
- L. Di and E. H. Kerns, Drug-Like Properties: Concepts, Structure Design and Methods from ADME to Toxicity Optimization, Elsevier, 2016 Search PubMed.
- V. Reenu, J. Mol. Graphics Modell., 2015, 61, 89–101 CrossRef CAS PubMed.
- E. P. Sifonte, F. A. Castro-Smirnov, A. A. S. Jimenez, H. R. G. Diez and F. G. Martínez, J. Nanopart. Res., 2021, 23, 8 CrossRef.
- A. Singh, S. Kumar, A. Kapoor, P. Kumar and A. Kumar, Toxicol. Mech. Methods, 2023, 33, 222–232 Search PubMed.
- J. Kostal, Chem. Res. Toxicol., 2023, 36, 1444–1450 Search PubMed.
- D. Guan, R. Lui and S. T. Matthews, Curr. Res. Toxicol., 2024, 7, 100183 Search PubMed.
- J. Kostal, A. Voutchkova-Kostal, J. P. Bercu, J. C. Graham, J. Hillegass, M. Masuda-Herrera, A. Trejo-Martin and J. Gould, Chem. Res. Toxicol., 2024, 37, 1404–1414 Search PubMed.
- K. Hansen, S. Mika, T. Schroeter, A. Sutter, A. ter Laak, T. Steger-Hartmann, N. Heinrich and K.-R. Müller, J. Chem. Inf. Model., 2009, 49, 2077–2081 CrossRef CAS PubMed.
- Z. Wu, B. Ramsundar, E. Feinberg, J. Gomes, C. Geniesse, A. S. Pappu, K. Leswing and V. Pande, Chem. Sci., 2018, 9, 513–530 RSC.
- F. Mastrolorito, N. Gambacorta, F. Ciriaco, F. Cutropia, M. V. Togo, V. Belgiovine, A. R. Tondo, D. Trisciuzzi, A. Monaco, R. Bellotti, C. D. Altomare, O. Nicolotti and N. Amoroso, J. Chem. Inf. Model., 2025, 65, 1850–1861 CrossRef CAS PubMed.
- J. Cremer, L. Medrano Sandonas, A. Tkatchenko, D.-A. Clevert and G. De Fabritiis, Chem. Res. Toxicol., 2023, 36, 1561–1573 Search PubMed.
- J. A. Keith, V. Vassilev-Galindo, B. Cheng, S. Chmiela, M. Gastegger, K.-R. Muller and A. Tkatchenko, Chem. Rev., 2021, 121, 9816–9872 CrossRef CAS PubMed.
- B. Huang and O. A. Von Lilienfeld, Chem. Rev., 2021, 121, 10001–10036 CrossRef CAS PubMed.
- F. Musil, A. Grisafi, A. P. Bartók, C. Ortner, G. Csányi and M. Ceriotti, Chem. Rev., 2021, 121, 9759–9815 CrossRef CAS PubMed.
- S. N. Pozdnyakov, M. J. Willatt, A. P. Bartók, C. Ortner, G. Csányi and M. Ceriotti, Phys. Rev. Lett., 2020, 125, 166001 CrossRef CAS PubMed.
- A. Akbarzadeh, S. Prosandeev, E. J. Walter, A. Al-Barakaty and L. Bellaiche, Phys. Rev. Lett., 2012, 108, 257601 CrossRef CAS PubMed.
- K. Hansen, F. Biegler, R. Ramakrishnan, W. Pronobis, O. A. Von Lilienfeld, K.-R. Muller and A. Tkatchenko, J. Phys. Chem. Lett., 2015, 6, 2326–2331 CrossRef CAS PubMed.
- W. Pronobis, A. Tkatchenko and K.-R. Müller, J. Chem. Theory Comput., 2018, 14, 2991–3003 CrossRef CAS PubMed.
- F. A. Faber, A. S. Christensen, B. Huang and O. A. Von Lilienfeld, J. Chem. Phys., 2018, 148, 24 CrossRef PubMed.
- D. Khan, S. Heinen and O. A. von Lilienfeld, J. Chem. Phys., 2023, 159, 034106 CrossRef CAS PubMed.
- L. C. Gallegos, G. Luchini, P. C. St. John, S. Kim and R. S. Paton, Acc. Chem. Res., 2021, 54, 827–836 CrossRef CAS PubMed.
- F. Spiegelman, N. Tarrat, J. Cuny, L. Dontot, E. Posenitskiy, C. Martí, A. Simon and M. Rapacioli, Adv. Phys.: X, 2020, 5, 1710252 CAS.
- J. Hoja, L. Medrano Sandonas, B. G. Ernst, A. Vazquez-Mayagoitia, R. A. DiStasio Jr and A. Tkatchenko, Sci. Data, 2021, 8, 43 CrossRef CAS PubMed.
- D. Gadaleta, K. Vuković, C. Toma, G. J. Lavado, A. L. Karmaus, K. Mansouri, N. C. Kleinstreuer, E. Benfenati and A. Roncaglioni, J. Cheminform., 2019, 11, 1–16 Search PubMed.
- H. Zhu, T. M. Martin, L. Ye, A. Sedykh, D. M. Young and A. Tropsha, Chem. Res. Toxicol., 2009, 22, 1913–1921 Search PubMed.
- S. M. Lundberg and S.-I. Lee, in Advances in Neural Information Processing Systems 30, I. Guyon, U. V. Luxburg, S. Bengio, H. Wallach, R. Fergus, S. Vishwanathan and R. Garnett, Curran Associates, Inc., 2017, pp. 4765–4774 Search PubMed.
- K. T. Schütt, S. Chmiela, O. A. von Lilienfeld, A. Tkatchenko, K. Tsuda and K.-R. Müller, Machine learning meets quantum physics, Springer Nature, Cham, Switzerland, 2020 Search PubMed.
- T. Chen and C. Guestrin, Proceedings of the 22nd acm sigkdd international conference on knowledge discovery and data mining, 2016, pp. 785–794 Search PubMed.
- T. Akiba, S. Sano, T. Yanase, T. Ohta and M. Koyama, Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, 2019 Search PubMed.
- Y. Liu, L. Wang, W. Zhu and X. Yu, J. Inf. Rec. Mater., 2025, 5, 39 Search PubMed.
- S. De, A. P. Bartók, G. Csányi and M. Ceriotti, Phys. Chem. Chem. Phys., 2016, 18, 13754–13769 RSC.
- G. Seifert, D. Porezag and T. Frauenheim, Int. J. Quantum Chem., 1996, 58, 185–192 Search PubMed.
- M. Gaus, Q. Cui and M. Elstner, J. Chem. Theory Comput., 2011, 7, 931–948 CrossRef CAS PubMed.
- A. Tkatchenko, R. A. DiStasio, R. Car and M. Scheffler, Phys. Rev. Lett., 2012, 108, 236402 Search PubMed.
- A. Ambrosetti, A. M. Reilly, J. DiStasio, A. Robert and A. Tkatchenko, J. Chem. Phys., 2014, 140, 18A508 CrossRef PubMed.
- B. Hourahine, B. Aradi, V. Blum, F. Bonafe, A. Buccheri, C. Camacho, C. Cevallos, M. Deshaye, T. Dumitrică and A. Dominguez, et al. , J. Chem. Phys., 2020, 152, 12 Search PubMed.
- Z. Xu, N. E. Munyaneza, Q. Zhang, M. Sun, C. Posada, P. Venturo, N. A. Rorrer, J. Miscall, B. G. Sumpter and G. Liu, Science, 2023, 381, 666–671 CrossRef CAS PubMed.
- M. Stöhr and A. Tkatchenko, Sci. Adv., 2019, 5, eaax0024 Search PubMed.
- A. Santana Bonilla, R. Gutierrez, L. Medrano Sandonas, D. Nozaki, A. P. Bramanti and G. Cuniberti, Phys. Chem. Chem. Phys., 2014, 16, 17777–17785 RSC.
- A. Santana-Bonilla, L. Medrano Sandonas, R. Gutierrez and G. Cuniberti, J. Phys.: Condens. Matter, 2019, 31, 405502 CrossRef CAS PubMed.
- M. Y. Deshaye, A. T. Wrede and T. Kowalczyk, J. Chem. Phys., 2023, 158, 134104 Search PubMed.
- V. S. Naumov, A. S. Loginova, A. A. Avdoshin, S. K. Ignatov, A. V. Mayorov, B. Aradi and T. Frauenheim, Int. J. Quantum Chem., 2021, 121, e26427 Search PubMed.
- M. Gaus, A. Goez and M. Elstner, J. Chem. Theory Comput., 2013, 9, 338–354 CrossRef CAS PubMed.
- P. Ruiz, G. Begluitti, T. Tincher, J. Wheeler and M. Mumtaz, Molecules, 2012, 17, 8982–9001 Search PubMed.
- K. Huang, T. Fu, W. Gao, Y. Zhao, Y. Roohani, J. Leskovec, C. W. Coley, C. Xiao, J. Sun and M. Zitnik, arXiv, 2021, preprint, arXiv:2102.09548, DOI:10.48550/arXiv.2102.09548.
- L. Di and E. H. Kerns, Drug-like properties: concepts, structure design and methods from ADME to toxicity optimization, Academic press, 2015 Search PubMed.
- T. Ginex, J. Vazquez, E. Gilbert, E. Herrero and F. J. Luque, Future Med. Chem., 2019, 11, 1177–1193 Search PubMed.
- P. Pracht, F. Bohle and S. Grimme, Phys. Chem. Chem. Phys., 2020, 22, 7169–7192 RSC.
- P. Pracht, S. Grimme, C. Bannwarth, F. Bohle, S. Ehlert, G. Feldmann, J. Gorges, M. Müller, T. Neudecker and C. Plett, et al. , J. Chem. Phys., 2024, 160, 11 Search PubMed.
- L. Medrano Sandonas, D. Van Rompaey, A. Fallani, M. Hilfiker, D. Hahn, L. Perez-Benito, J. Verhoeven, G. Tresadern, J. Kurt Wegner, H. Ceulemans and A. Tkatchenko, Sci. Data, 2024, 11, 742 CrossRef CAS PubMed.
- C. Bannwarth, S. Ehlert and S. Grimme, J. Chem. Theory Comput., 2019, 15, 1652–1671 Search PubMed.
- A. P. Soleimany, A. Amini, S. Goldman, D. Rus, S. N. Bhatia and C. W. Coley, ACS Cent. Sci., 2021, 7, 1356–1367 CrossRef CAS PubMed.
- H. Y. I. Lam, R. Pincket, H. Han, X. E. Ong, Z. Wang, J. Hinks, Y. Wei, W. Li, L. Zheng and Y. Mu, Nat. Mach. Intell., 2023, 5, 754–764 CrossRef.
- S. Teng, C. Yin, Y. Wang, X. Chen, Z. Yan, L. Cui and L. Wei, Comput. Biol. Med., 2023, 164, 106904 CrossRef CAS PubMed.
- T. B. Kimber, M. Gagnebin and A. Volkamer, Artif. Intell. Life Sci., 2021, 1, 100014 CAS.
- N. Lukashina, A. Alenicheva, E. Vlasova, A. Kondiukov, A. Khakimova, E. Magerramov, N. Churikov and A. Shpilman, arXiv, 2020, preprint, arXiv:2011.12117, DOI:10.48550/arXiv.2011.12117.
- G. Zhou, Z. Gao, Q. Ding, H. Zheng, H. Xu, Z. Wei, L. Zhang and G. Ke, In The Eleventh International Conference on Learning Representations, 2023, https://openreview.net/forum?id=6K2RM6wVqKu Search PubMed.
- H. Du, J. Wang, J. Hui, L. Zhang and H. Wang, npj Comput. Mater., 2024, 10, 292 CrossRef CAS.
- D. P. Kovács, J. H. Moore, N. J. Browning, I. Batatia, J. T. Horton, Y. Pu, V. Kapil, W. C. Witt, I.-B. Magdău, D. J. Cole and G. Csányi, J. Am. Chem. Soc., 2025, 147, 17598–17611 Search PubMed.
- J. Seward, M. Cronin and T. Schultz, SAR QSAR Environ. Res., 2002, 13, 325–340 CrossRef CAS PubMed.
- S. Long, Y. Onitsuka, S. Nagao and M. Takahashi, Molecules, 2025, 30, 2947 CrossRef CAS PubMed.
- F. Reymond, P.-A. Carrupt, B. Testa and H. H. Girault, Chem.–Eur. J., 1999, 5, 39–47 CrossRef CAS.
- U. Argikar, M. Blatter, D. Bednarczyk, Z. Chen, Y. S. Cho, M. Dore, J. L. Dumouchel, S. Ho, K. Hoegenauer and T. Kawanami, et al. , J. Med. Chem., 2022, 65, 12386–12402 Search PubMed.
- W. Li, H. Ma, S. Li and J. Ma, Chem. Sci., 2021, 12, 14987–15006 RSC.
- G. Zhou, N. Lubbers, K. Barros, S. Tretiak and B. Nebgen, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2120333119 CrossRef CAS PubMed.
- B. Huang, G. F. von Rudorff and O. A. von Lilienfeld, Science, 2023, 381, 170–175 CrossRef CAS PubMed.
- L. Medrano Sandonas, M. Puleva, Z. Erarslan, R. Parra Payano, M. Stöhr, G. Cuniberti and A. Tkatchenko, Phys. Chem. Chem. Phys., 2026 10.1039/D6CP00038J.
- A. McSloy, G. Fan, W. Sun, C. Hölzer, M. Friede, S. Ehlert, N.-E. Schütte, S. Grimme, T. Frauenheim and B. Aradi, J. Chem. Phys., 2023, 158, 034801 CrossRef CAS PubMed.
- A. Fallani, L. Medrano Sandonas and A. Tkatchenko, Nat. Commun., 2024, 15, 6061 Search PubMed.
- C. Xu, X. Deng, Y. Lu and P. Yu, Digital Discovery, 2025, 4, 161–171 RSC.
- A. Volokhova, M. Koziarski, A. Hernández-García, C.-H. Liu, S. Miret, P. Lemos, L. Thiede, Z. Yan, A. Aspuru-Guzik and Y. Bengio, Digital Discovery, 2024, 3, 1038–1047 RSC.
Footnote |
| † Present address: Institute for Materials Science and Max Bergmann Center of Biomaterials, TUD Dresden University of Technology, 01062 Dresden, Germany. |
| This journal is © The Royal Society of Chemistry 2026 |





