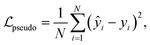Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Joint embedding predictive architecture for self-supervised pretraining on polymer molecular graphs
Francesco
Piccoli†‡
,
Gabriel
Vogel†
 and
Jana M.
Weber
and
Jana M.
Weber
 *
*
Department of Intelligent Systems, Delft University of Technology, Delft, 2629 HZ, The Netherlands. E-mail: j.m.weber@tudelft.nl; g.vogel@tudelft.nl
First published on 23rd January 2026
Abstract
Recent advances in machine learning (ML) have shown promise in accelerating the discovery of polymers with desired properties by aiding in tasks such as virtual screening via property prediction. However, progress in polymer ML is hampered by the scarcity of high-quality labeled datasets, which are necessary for training supervised ML models. In this work, we study the use of the very recent ‘Joint Embedding Predictive Architecture’ (JEPA), a type of architecture developed for self-supervised learning (SSL), on polymer molecular graphs to understand whether pretraining with the proposed SSL strategy improves downstream performance when labeled data is scarce. We first pretrain our polymer-JEPA model on a large dataset of conjugated copolymer photocatalysts. The pretrained model is then fine-tuned on two distinct downstream tasks: predicting electron affinity in the same chemical space and classifying phase behavior in diblock copolymers, a different chemical space. Our results indicate that JEPA-based self-supervised pretraining enhances downstream performance, particularly when labeled data is very scarce, achieving improvements across both tested datasets. The method provides performance gains in cross-domain fine-tuning, highlighting its potential to extract general knowledge across different classes of polymers. By leveraging large amounts of unlabeled polymer structures for pretraining, the proposed strategy can further reduce the dependence on extensive labeled datasets.
1 Introduction
Synthetic polymers are one of the most widespread classes of materials, constituting an essential component of numerous commodities in everyday life and industry.1–4 The large diversity of the chemical space of polymers provides an opportunity to design polymers whose properties match the demands of the application.3,5 However, the high number of combinations of monomer compositions, higher-order topologies, and processing methods brings challenges in the effective navigation of this large search space.3,4In recent years, machine learning (ML) has shown potential in the discovery of new materials, including polymers.6,7 ML methods are increasingly applied in polymer science, particularly in two key areas: virtual screening of predefined candidate structures to predict properties and inverse polymer design to generate novel structures with desired properties.5,8,9
However, the application of ML in polymer science is still in its infancy, primarily due to the scarcity of high-quality, large, publicly available labeled datasets. This limitation arises from the time- and cost-intensive procedure of generating labeled polymer data (via experimental synthesis and testing or accurate molecular simulations).4,6,10 To overcome the problem of limited labeled data, several common strategies have been applied in the polymer domain. Transfer learning involves pretraining models on polymer properties with abundant labeled data and fine-tuning them for properties with limited data.11,12 Multitask learning is an effective approach to train predictive models on multiple properties with varying levels of labeled data, leveraging interdependencies between these properties.2,13,14 Lastly, self-supervised learning (SSL) makes it possible to pretrain models on large volumes of unlabeled data through tasks defined directly on the input data. The learned representations can then be fine-tuned on smaller labeled datasets.3,12,15–17
Among these strategies, SSL has been particularly transformative across different data structures such as images,18–21 natural language,22,23 and graphs.24–26 In the molecular domain, graph-based SSL has shown considerable success with small molecules.27–29
In the context of polymers, the focus has largely been on text-based SSL, learning representations through tasks derived from the textual pSMILES representation,3,12,16,17 with limited exploration of graph-based SSL approaches. Polymer graphs beyond the polymer repeat unit graph, including weighted edges that describe monomer ensembles, their topology and their stochasticity, as proposed in ref. 1, present unique structural characteristics that distinguish them from small molecular graphs, posing challenges for directly applying SSL techniques developed for small molecular graphs. A recent study proposed a self-supervised graph neural network for such polymer graphs.15 The authors employ two SSL tasks: one at the node/edge level, masking nodes and edges and learning to predict them, and the other at the graph level, predicting a pseudolabel corresponding to the molecular weight of the polymer, derived from the monomers' weights. They test both tasks separately and together, and they discover that pretraining via both tasks proves to be the most effective. This result aligns with findings in the literature30 that SSL on graphs works better when using both node-level and graph-level tasks together.
In this work, we study a new architecture family, called Joint Embedding Predictive Architecture (JEPA),31 which was developed for self-supervised representation learning of images. Unlike traditional graph-based SSL methods, such as node or edge masking, which focus on reconstructing masked features directly in the input graph space, JEPAs operate in an embedding space. Predicting in the embedding space facilitates the learning of semantically-rich representations, avoiding the need to predict and reconstruct every (potentially noisy and hard to predict) detail of the input space, that in high-dimensional domains often leads to overfitting.31,32 The way JEPAs learn, is by predicting the embedding of a “target view” of the graph based on the embedding of a “context view”, typically by employing two encoders. We apply this architecture for self-supervised pretraining on stochastic polymer graphs to improve the accuracy in downstream tasks (e.g. property prediction) in label-scarce data scenarios.
We first use the proposed method for pretraining on a larger unlabeled corpus of data, and then finetune the model on available labeled data in a supervised fashion. While some aspects of our analysis apply broadly to JEPAs across various types of graphs (i.e. in different domains) and extend the study of JEPAs for graphs initiated in ref. 32, other results and experiments are specific to JEPAs in the molecular graph domain, specifically for stochastic polymer graphs.
2 Methods
2.1 Polymer representation and datasets
We represent polymers as stochastic graphs, as proposed in ref. 1. The graph representation, visualized in Fig. 1, connects monomer graphs through weighted edges indicating the probabilities of connections, representing the polymer chain architecture. We use the dataset of conjugated copolymer photocatalysts for hydrogen production,1 that is built upon the polymer space defined in ref. 33. As shown in Fig. 2a, the dataset contains 42![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 966 polymers, composed of nine A monomers and 862 B monomers. The polymers are created by combining each A monomer with each B monomer in three distinct chain architectures: alternating, random, and block. Additionally, they are further distinguished by three stoichiometry ratios: 1
966 polymers, composed of nine A monomers and 862 B monomers. The polymers are created by combining each A monomer with each B monomer in three distinct chain architectures: alternating, random, and block. Additionally, they are further distinguished by three stoichiometry ratios: 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1, 1
1, 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3, and 3
3, and 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. While this is a useful and extensive dataset, the variety of it is somewhat limited by the combinatorial approach to build the polymers. Aldeghi and Coley1 further performed oligomer DFT calculations to provide estimations for two properties, the electron affinity (EA) and ionization potential (IP), for each of the copolymers.1 While this provides a valuable resource for benchmarking polymer informatics, it potentially inherits biases from DFT approximations. We use this dataset for pretraining and keep aside a part of the dataset for finetuning.
1. While this is a useful and extensive dataset, the variety of it is somewhat limited by the combinatorial approach to build the polymers. Aldeghi and Coley1 further performed oligomer DFT calculations to provide estimations for two properties, the electron affinity (EA) and ionization potential (IP), for each of the copolymers.1 While this provides a valuable resource for benchmarking polymer informatics, it potentially inherits biases from DFT approximations. We use this dataset for pretraining and keep aside a part of the dataset for finetuning.
 | ||
| Fig. 1 Polymer graph representation as introduced in ref. 1. The representation uses stochastic edges (dashed) reflecting the connection probabilities between monomers, i.e. reflecting the stochastic nature and chain architecture. | ||
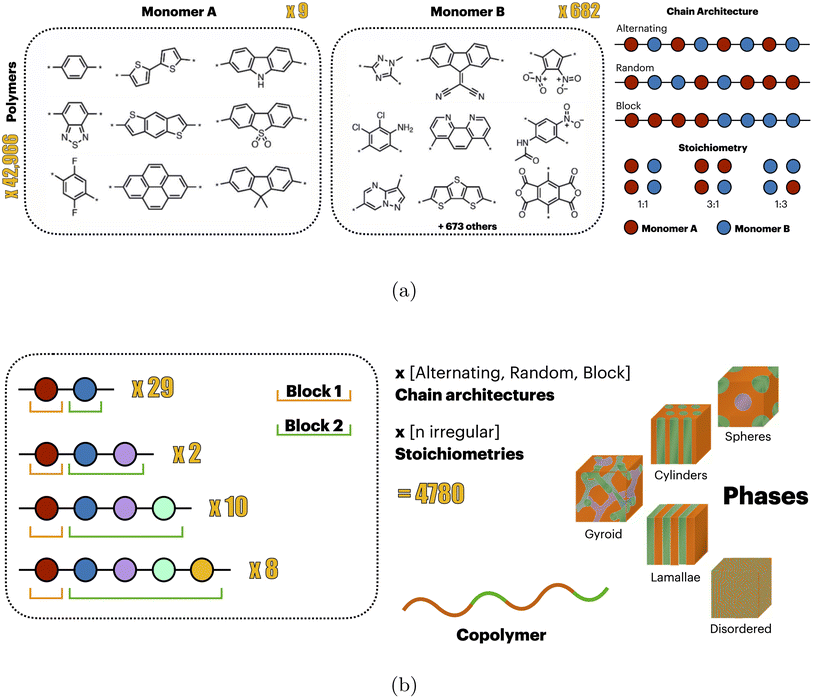 | ||
| Fig. 2 (a) Conjugated copolymer photocatalyst dataset1 with different stoichiometries and chain architectures. (b) Diblock copolymer dataset compiled by Arora et al.,34 reporting experimentally observed phase behavior (e.g., lamellae, cylinders, gyroid) for 50 diblock copolymers across varying stoichiometries and chain architectures. | ||
We test our approach on two distinct downstream tasks, both starting from a model pretrained on the aforementioned dataset. Firstly, we finetune on data from the same dataset (but different split). Secondly, we finetune on a different downstream task using a dataset of diblock copolymers.34 The dataset provides labels of the phase behavior (lamellae, hexagonal-packed cylinders, body-centred cubic spheres, a cubic gyroid, or disordered) of 49 diblock copolymers across various relative volume fractions, totaling 4780 labeled polymer samples (see Fig. 2b). Both datasets were downloaded from the open-source repository https://github.com/coleygroup/polymer-chemprop-data/, accessed on Jan. 30th, 2024.
2.2 Model
The idea of JEPAs for graphs is to first partition the graphs into patches (i.e. subgraphs) and define larger context subgraph x and smaller target subgraphs y. Secondly, the goal is to predict (reconstruct) the embedding sy of a target subgraph, from the embedding sx of the context subgraph, operating in the embedding space (see Fig. 3).• Motif-based subgraphing is a domain-specific approach to generate chemically meaningful subgraphs, such as functional groups or molecular subunits. We build on the BRICS (Breaking of Retrosynthetically Interesting Chemical Substructures) algorithm,36 specifically using the implementation in ref. 37. The motif-based subgraphing method ensures meaningful subgraphs but potentially limits the variability of subgraphs due to its deterministic nature.
• METIS35 is a popular subgraphing algorithm, using a clustering-based method, that partitions graphs into clusters while minimizing edge cuts and maximizing within-cluster links. Its widespread use is due to its low computational cost and the quality of the produced subgraphs. Despite being computationally efficient, METIS-based subgraphs lack chemical meaning compared to motif-based subgraphing.
• Random-walk subgraphing uses a stochastic approach to generate diverse subgraphs. It ensures greater flexibility and control over subgraph sizes while producing varying subgraphs at each training iteration. This method aligns well with the requirements for JEPA, particularly the need for dynamic changes to prevent overfitting.
The next step focuses on the prediction of the target subgraph embeddings through a multi-layer perceptron (MLP) from the context embedding and positional information of the target. We employed positional encoding (PE) via random-walk structural encoding (RWSE)38,39 at two levels: at the node level and at the subgraph (patch) level. The node-level PE allows us to maintain node positional information when working with subgraphs. The subgraph (patch)-level positional encoding contains information about the connectivity of two subgraphs, here the context and target subgraph. Given the output of the context encoder, sx, we wish to predict the m target subgraphs representations sy(1), …, sy(m). To that end, for a given target subgraph embedding sy(i), the predictor hϕ takes as input sx summed with the linearly transformed target subgraph positional token ![[p with combining tilde]](https://www.rsc.org/images/entities/b_char_0070_0303.gif) i:
i:
ŝy(i) = hϕ(sx + ![[p with combining tilde]](https://www.rsc.org/images/entities/b_char_0070_0303.gif) i i![[T with combining tilde]](https://www.rsc.org/images/entities/b_char_0054_0303.gif) ) ) | (1) |
![[T with combining tilde]](https://www.rsc.org/images/entities/b_char_0054_0303.gif) ∈
∈ ![[Doublestruck R]](https://www.rsc.org/images/entities/char_e175.gif)
![[k with combining tilde]](https://www.rsc.org/images/entities/i_char_006b_0303.gif) ×d, where
×d, where ![[k with combining tilde]](https://www.rsc.org/images/entities/i_char_006b_0303.gif) is the dimension of the positional encoding token and d is the embedding dimension. The predictor outputs the predicted target embedding ŝy(i). Since we wish to make predictions for m target blocks, we apply our predictor m times, obtaining predictions ŝy(1), …, ŝy(m). In practice, the predictor hϕ is implemented via a MLP. For each data point, the loss is the average L2 distance (Mean Square Error (MSE)) between the m predicted target subgraph representations and the m true target subgraph representations. The MSE loss in the embedding space can lead to fluctuating or increasing loss values, e.g., with changing magnitudes of embedding vectors over epochs. In Appendix E, we investigate the effect of layer normalization on training stability and convergence of JEPA pretraining.
is the dimension of the positional encoding token and d is the embedding dimension. The predictor outputs the predicted target embedding ŝy(i). Since we wish to make predictions for m target blocks, we apply our predictor m times, obtaining predictions ŝy(1), …, ŝy(m). In practice, the predictor hϕ is implemented via a MLP. For each data point, the loss is the average L2 distance (Mean Square Error (MSE)) between the m predicted target subgraph representations and the m true target subgraph representations. The MSE loss in the embedding space can lead to fluctuating or increasing loss values, e.g., with changing magnitudes of embedding vectors over epochs. In Appendix E, we investigate the effect of layer normalization on training stability and convergence of JEPA pretraining.
2.3 Training procedure
We split the full conjugated copolymer dataset1 in three parts: 40% for pretraining, 40% for the different finetuning scenarios, and the remaining 20% for testing the property prediction performance. We use random splits with a five-fold cross validation and three repetitions for different finetuning scenarios. Moreover, we provide an additional analysis on a scaffold-based data split scenario in Appendix F, performing a nine-fold cross validation where always one A-monomer is held out as a test set.The pretraining phase always entails training the JEPA architecture on 40% (17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 186 entries) of the conjugated copolymer dataset.1 After pretraining, only the trained target encoder is utilized in the finetuning step to obtain the polymer graph embedding. For the downstream task (e.g. polymer property prediction), an MLP is employed on top of the polymer graph embedding obtained from the target encoder. Finetuning is done end-to-end, wherein not only the MLP but also the target encoder weights are updated during the optimization process. This allows the polymer graph embeddings to also finetune to the specific downstream task. As mentioned in Section 2.1, we perform the finetuning on the two different datasets to investigate the difference between using the same chemical space for pretraining and finetuning in contrast to using two different polymer chemical spaces. The results for this study are presented in Sections 3.1 and 3.2.
186 entries) of the conjugated copolymer dataset.1 After pretraining, only the trained target encoder is utilized in the finetuning step to obtain the polymer graph embedding. For the downstream task (e.g. polymer property prediction), an MLP is employed on top of the polymer graph embedding obtained from the target encoder. Finetuning is done end-to-end, wherein not only the MLP but also the target encoder weights are updated during the optimization process. This allows the polymer graph embeddings to also finetune to the specific downstream task. As mentioned in Section 2.1, we perform the finetuning on the two different datasets to investigate the difference between using the same chemical space for pretraining and finetuning in contrast to using two different polymer chemical spaces. The results for this study are presented in Sections 3.1 and 3.2.
2.4 Additional pseudolabel objective
As mentioned in the introduction, a first study on self-supervised learning on the stochastic polymer graph representation1 used both node and edge masking and prediction of a pseudolabel as self-supervised tasks. The study found that transferring not only the pretrained weights of the wD-MPNN encoder (equivalent to our target encoder), but also the weights of the pseudolabel predictor (an MLP used to predict molecular weight during pretraining) improved downstream performance. These pseudolabel predictor weights of the MLP were reused for downstream tasks, such as predicting electron affinity (EA) in the conjugated copolymer dataset.1 Motivated by these results, we decided to adopt the pseudolabel objective also with our architecture. We do so by predicting the molecular weight from the polymer fingerprint learned through the target encoder, as visible in Fig. 4. Unlike the sequential training approach in ref. 15, which first trains on node-level tasks and then on the pseudolabel task, we jointly train the encoder with both the JEPA objective and the pseudolabel prediction task. The total training loss is calculated as the sum of the JEPA loss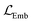 and the pseudolabel loss
and the pseudolabel loss 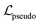 , aggregated with equal weights:
, aggregated with equal weights:The pseudolabel loss was computed as a mean squared error (MSE) between the predicted and target pseudolabels,
3 Results and discussion
In this section, we analyze the performance of our JEPA-based self-supervised pretraining strategy on polymer graphs. Section 3.1 evaluates the model's performance on downstream tasks under varying labeled data availability scenarios. As indicated in Section 2.1, we test the effectiveness of the self-supervised pretraining for predicting the electron affinity (EA) (regression) of the conjugated copolymer dataset (Section 3.1) and the phase behavior (classification) of diblock copolymers (Section 3.2). The latter task is based on a different polymer chemical space than the self-supervised pretraining, and thus evaluates the model's cross-domain generalization capability. Further, Section 3.3 compares our approach with an alternative self-supervised learning (SSL) method for polymer graphs, while Section 3.4 shows the performance against a simple baseline random forest model. Finally, we present an ablation study of subgraphing hyperparameters in Section 3.5, which provides general insights for JEPA-based graph machine learning.3.1 Downstream performance on data set of conjugated copolymers
For the task of predicting the electron affinity (EA) of conjugated copolymers, our pretraining approach improves the performance especially in low labeled data scenarios as shown in Fig. 5. We test from a data scenario of 0.4% (192 polymers) labeled data points to a scenario of 24% (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 311 polymers) labeled data points. In Fig. 5 we report results only up to the 8% scenario to better visualize the impact in low data regimes. The pretrained model especially demonstrates performance improvements in scenarios up to a data size of 4% (1728 polymers) labeled data points. However, beyond this threshold, the benefits of pretraining plateau. This suggest that the available labeled data is sufficient for supervised learning, rendering the transferred pretraining knowledge redundant. In practice, a small change in the R2 value (e.g. ±0.01) does not significantly impact molecular design task, however, in the low labeled data scenarios (i.e. 0.4% and 0.8%), pretraining leads to a significant improvement of the property prediction performance.
311 polymers) labeled data points. In Fig. 5 we report results only up to the 8% scenario to better visualize the impact in low data regimes. The pretrained model especially demonstrates performance improvements in scenarios up to a data size of 4% (1728 polymers) labeled data points. However, beyond this threshold, the benefits of pretraining plateau. This suggest that the available labeled data is sufficient for supervised learning, rendering the transferred pretraining knowledge redundant. In practice, a small change in the R2 value (e.g. ±0.01) does not significantly impact molecular design task, however, in the low labeled data scenarios (i.e. 0.4% and 0.8%), pretraining leads to a significant improvement of the property prediction performance.
 | ||
| Fig. 5 Effectiveness of our pretraining strategy for different finetune dataset sizes. The performance is evaluated on predicting the electron affinity of test data from the conjugated copolymer dataset1 with the performance measured as R2. | ||
3.2 Self-supervised pretraining and transfer across chemical spaces
For the task of predicting the diblock copolymer phase behavior, pretraining on the dataset of conjugated copolymers consistently improves the classification performance (between around 0.02 and 0.1 in AUPRC), even in higher labeled data scenarios, as illustrated in Fig. 6. We test from a training data scenario of 191 data points (4%) to 3824 data points (80%). The latter scenario corresponds to finetuning on the full dataset, retaining 20% for testing, as done in ref. 1. Since the pretraining dataset represents a different polymer chemical space compared to the finetuning dataset, we conclude that the knowledge acquired during pretraining is not overfitting or memorizing the training distribution (i.e. chemical space) but rather learning general chemical knowledge about polymers. This indicates the potential opportunity to utilise this strategy more broadly across different polymer data sets. | ||
| Fig. 6 Effectiveness of our pretraining strategy for transfer learning and different finetune dataset sizes. Self-supervised pretraining is performed on data from the conjugated copolymer dataset1 and finetuning is performed on test data from the diblock copolymer dataset.34 The classification performance for predicting the phase behavior is measured as AUPRC (area under the precision–recall curve). | ||
We see particular promise in scenarios, where a model pretrained on a large, unlabeled polymer space is fine-tuned across domains on a smaller, labeled dataset from another polymer space. We expect this to be especially advantageous for small datasets with complex structure–property relationships.
3.3 Comparison to input space SSL
We compare the effectiveness of our pretraining strategy to the SSL pretraining method by Gao et al.,15 which uses node and edge masking in the input spaces as described in the introduction. Due to the small finetune dataset size, we run both methods with a 5-fold cross validation and three repetitions with different splits to ensure a robust comparison. The size of the wD-MPNN is set to three layers a hidden dimension of 300 for both methods.Overall, the performance increase using input space self-supervised pretraining and embedding space self-supervised pretraining (JEPA, our method) is comparable. Fig. 7 reveals that our method is slightly better in the very low data scenarios, and Gao et al.'s method15 performs better with more available labelled data.
 | ||
| Fig. 7 Comparison between our pretraining strategy and the best performing SSL model from ref. 15. For different finetune dataset sizes we compare the R2 for prediction of EA of the conjugated copolymer test data.1 | ||
Using an additional pseudolabel objective as described in Section 2.4 leads to a consistent improvement in ref. 15. While we observed significant improvements in single scenarios, overall the performance improvements attributed to using an additional pseudolabel objective are smaller for our method than for the input-space SSL approach as shown in Fig. 8. This suggests that our strategy potentially already captures relevant information related to the polymer molecular weight pseudolabel.
 | ||
| Fig. 8 The effect of using an additional pseudolabel objective in input space SSL15 and embedding space SSL (ours). For different finetune dataset sizes we compare the R2 for prediction of EA of the conjugated copolymer test data.1 We include both the scenarios when only the wD-MPNN encoder weights are transferred (no PL), and the scenario when also the pseudolabel (molecular weight) predictor weights are transferred (PL). | ||
3.4 Comparison to tree-based baseline models
We also compare the performance of our model to a random forest and XGBoost model trained on fingerprints generated from polymer sequences as inputs. Typically, simpler ML models outperform complex ones in data-scarce scenarios. The wD-MPNN generates a polymer representation (fingerprint) through pooling node embeddings learned during training, which is then used as input for the MLP predictor. In contrast, the tree-based models are trained on handcrafted fingerprints as inputs. We follow the approach in ref. 1, utilizing count-vector Extended-Connectivity Fingerprints (ECFP)40 of size 2048 and radius 2, computed with RDKit.41 The polymer fingerprints are obtained by averaging across an ensemble of 32 oligomer sequence fingerprints.While the polymer-JEPA demonstrated improvements over our baseline model without pretraining in low-data regimes, we observed that the tree-based models outperform the pretrained wD-MPNN in the very low labeled data regimes. This trend is evident in Fig. 9a, where the random forest and XGBoost model exhibited an advantage over the pretrained wD-MPNN when using 0.4% (192 points) and 0.8% (384 points) of the dataset for finteuning. The advantage of the RF model vanishes as the dataset size increases to 1.6% and the advantage of the XGBoost model vanishes as the size increases to 4%. In practice, only for a small regime around 4.0% finetune data, we observe that the pretrained model outperforms both the random forest and the wD-MPNN without pretraining. There is no scenario in which the pretrained wD-MPNN model outperforms both the XGBoost model and the baseline wD-MPNN without pretraining.
 | ||
| Fig. 9 (a) Comparison between our pretraining-finetuning strategy and two tree-based baseline models (random forest and XGBoost) on the conjugated copolymer dataset,1 predicting the EA property, for different finetune dataset sizes. (b) Comparison of phase behavior classification performance between our pretraining–finetuning strategy and two tree-based baseline models (random forest and XGBoost) on the diblock copolymer dataset,34 for different finetune dataset sizes. | ||
For the smaller diblock dataset (Fig. 9b), we observed that the tree-based baseline models outperform our model throughout all data scenarios. This may be due to highly informative fingerprints, that correlate with the classification label. As Aldeghi and Coley1 point out, simply the mole fraction, correlated with the volume fractions of the two blocks is highly informative for determining the copolymer phase. They trained a RF model on mole fractions only which outperformed the wD-MPNN (no data scarce scenarios) without providing information about the chemistry. However, in many polymer systems the structure-to-property relationships are more complex and cannot be captured by a single, easily computed feature. While polymer fingerprinting combined with smaller (e.g. tree-based) models remains a strong baseline, several studies have demonstrated that learned representations and deep neural networks surpass classical models across a variety of molecular and polymer property prediction tasks.42–44 We anticipate that as polymer-specific representations mature – better encoding higher-order structural features such as branching, molecular-weight distributions, and sequence heterogeneity—representation learning approaches will deliver even greater performance gains.
Based on the results in our study, we advise to consider also the application of simpler models for small labeled datasets with relatively simple structure to property relationships. However, one key advantage of our proposed method lies in its ability to eliminate the reliance on handcrafted descriptors. By learning directly from the polymer graph structure, JEPA offers greater adaptability to new datasets without the need to tune fingerprints for specific polymer structures. Specifically, the used fingerprinting1 which involves the generation of oligomer ensembles and averaging their fingerprint, requires thoughtful engineering by experts and significant computation time.
Lastly, we hypothesize that more diverse pretraining datasets and more finetune data could further elevate the performance of our JEPA (wD-MPNN) pretrained model, beyond the two tasks covered in this study.
3.5 Evaluation of subgraphing strategies
This section presents an ablation study on subgraphing hyperparameters, including context and target subgraph size, the number of targets, and the subgraphing algorithm. These results explore the sensitivity of each parameter by varying them individually while keeping others fixed. Note, that we report suitable ranges and trends in this section. The ablation study itself is inherently stochastic as we utilise random initialization with stochastic subgraph generation and test the model under limited finetuning data availability. Further, pretraining effectively replaces random initialization of the wD-MPNN weights, while finetuning remains an end-to-end setup. Thus, the reported metrics reflect the integrated effect of pretraining and finetuning for this specific downstream task.Each experiment involved pretraining the model on 40% of the conjugated copolymer dataset (Section 2.3) and finetuning on 0.4% (192 data points) to simulate a label-scarce scenario. Overall, across ablation experiments, self-supervised pretraining with our model consistently improved downstream property prediction performance. For the ablation experiments related to context subgraph size, target subgraph size and number of targets, we used the random-walk algorithm for subgraph creation due to its direct control over subgraph size.
Within the searched subgraphing settings, we observe the highest performance increase using random walk subgraphing, a context size of 60% and one target with a subgraph size of 10% of the polymer graph. Yet, the model's sensitivity to subgraphing hyperparameters is comparatively low, indicating that (i) the resulting parameters should be interpreted as one out of many suitable configurations and that (ii), in the tested parameter space, JEPA pretraining provides performance improvements irrespective of the hyperparameter configuration.
| Context size | R 2 ↑ | RMSE ↓ |
|---|---|---|
| No pretraining | 0.46± 0.15 | 0.44± 0.06 |
| 20% | 0.56 ± 0.07 | 0.39 ± 0.03 |
| 40% | 0.60 ± 0.06 | 0.37 ± 0.03 |
| 60% | 0.65 ± 0.03 | 0.35 ± 0.02 |
| 80% | 0.62 ± 0.07 | 0.37 ± 0.03 |
| 95% | 0.61 ± 0.05 | 0.37 ± 0.02 |
| Target size | R 2 ↑ | RMSE ↓ |
|---|---|---|
| No pretraining | 0.46± 0.15 | 0.44± 0.06 |
| 5% | 0.61 ± 0.07 | 0.37 ± 0.03 |
| 10% | 0.66 ± 0.02 | 0.35 ± 0.01 |
| 15% | 0.65 ± 0.03 | 0.35 ± 0.02 |
| 20% | 0.63 ± 0.03 | 0.36 ± 0.02 |
| Number of targets | R 2 ↑ | RMSE ↓ |
|---|---|---|
| No pretraining | 0.46 ± 0.15 | 0.44 ± 0.06 |
| 1 | 0.67 ±0.01 | 0.34 ±0.01 |
| 2 | 0.64 ± 0.03 | 0.36 ± 0.01 |
| 3 | 0.66 ± 0.02 | 0.35 ± 0.01 |
| 4 | 0.65 ± 0.05 | 0.35 ± 0.02 |
| 5 | 0.61 ± 0.04 | 0.37 ± 0.02 |
| Subgraphing | R 2 ↑ | RMSE ↓ |
|---|---|---|
| No pretraining | 0.46 ± 0.15 | 0.44 ± 0.06 |
| Motif-based | 0.63 ± 0.05 | 0.36 ± 0.02 |
| Metis | 0.67 ± 0.04 | 0.34 ± 0.02 |
| Random walk (RW) | 0.67 ±0.01 | 0.34 ±0.01 |
Using a fixed context size of 60% and target size of 10% (single target), we compared the random-walk, motif-based, and METIS subgraphing. Interestingly, the motif-based method, which leverages domain knowledge to produce chemically meaningful subgraphs, exhibits slightly lower performance than the other two algorithms. While motif-based subgraphing generates chemically meaningful subgraphs, it tends to produce a relatively small number of subgraphs, in a deterministic fashion, potentially limiting model generalization by increasing the likelihood of encountering similar or identical subgraphs (both context and target ones) throughout training. On the other hand, the METIS algorithm, while also producing consistent subgraphs at each epoch, generates on average a higher number of subgraphs compared to the motif-based approach, introducing more variability across epochs. Finally, random-walk subgraphing generates different subgraphs at every epoch, thanks to the stochastic nature of the subgraphing process. Beyond the factor of variability, we investigated the adherence of these methods to the specified subgraph size. In Appendix D we provide a detailed analysis on the subgraph sizes for the different subgraphing types given the desired specification. While all methods systematically overshoot the specified subgraph size due to implementation constraints (chemical validity, connectivity preservation, and meaningfulness of partitions), the adherence to the specified size of the extracted subgraphs differs. The random-walk approach provided the best adherence to the desired context size (close to 60%). Conversely, METIS achieved the closest adherence to the specified target size (10%). In the previous sections, we identified a too small context subgraph size to be the main driver of performance drops. Since our implementation leads to larger subgraphs than specified for all methods, we naturally prevent too small context subgraphs.
While we observe the best performance for the random-walk approach, other subgraphing methods remain competitive and may favor slightly different optimal configurations (e.g., context size, target size, or number of targets). The analysis indicates that for our dataset the pretraining effect is consistently significant across most settings, whereas further hyperparameter tuning offers only minor improvements.
4 Conclusion
This study introduces a novel self-supervised pretraining strategy for the polymer domain, where labeled data is often scarce. The presented method operates on polymer molecular graphs, leveraging the concept of JEPAs. Our study stands as one of the initial efforts in exploring JEPAs for molecular graph-related tasks, thus enriching the understanding and analysis of this new architectural family in the molecular domain. We provide guidance for the exploration of hyperparameters for subgraphing; in particular on size and algorithm selection. While the exact optimal configuration likely depends on the considered dataset and fine-tune scenario, Section 3.5 reveals that, across all tested settings, pretraining consistently increased the performance in the tested downstream task, with limited sensitivity to subgraphing hyperparameter choices.Our experiments show that self-supervised pretraining on conjugated copolymer data consistently improves the downstream prediction accuracy for a dataset that describes a different polymer space (diblock copolymers), showcasing the ability of transferring general knowledge across polymer datasets of different applications. When pretraining and finetuning on the same polymer space of conjugated copolymers our method helps in label-scarce data scenarios up to up to around 8% (ca. 3440 polymers) of labeled data availability. The performance improvement (in R2) varies from 39.8% in the smallest labeled data scenario to 0.4% in the scenario with 8% labelled data.
Comparing our polymer-JEPA self-supervised model (embedding space) with node/edge-masking self-supervised learning (input space), we observe that we achieve comparable performance on the tested downstream task. Further, both methods benefit from including a pseudotask prediction (molecular weight), with the benefit being less pronounced in our embedding space self-supervised pretraining strategy. Additionally, we showed that simple tree-based baselines like a random forest or XGBoost model outperform our deep learning models in certain scenarios in the two downstream tasks considered, especially when labelled data is very scarce. However, these methods rely on expert-engineered polymer fingerprints, which is not necessary with our method that learns from the polymer graph directly.
Looking ahead, the embedding-based nature of JEPA offers promising opportunities for integrating multimodal data and utilizing a variety of different experimental and synthetic datasets for pretraining. Generally, we hypothesize that more diverse pretraining datasets could contribute to further increasing the performance of our JEPA (wD-MPNN) model compared to not using a pretrained model, also beyond the two tasks covered in this study.
Author contributions
FP: conceptualization, methodology, software (lead), formal analysis, writing – original draft. GV: supervision, methodology, software, formal analysis, writing – original draft. JW: conceptualization, supervision, writing – review and editing.Conflicts of interest
Not applicable.Data availability
The used data and code are provided in the Github repository https://github.com/Intelligent-molecular-systems/Polymer-JEPA, with the archived version at time of publication at https://doi.org/10.5281/zenodo.17640709. Additionally, data splits, preprocessed data to train the JEPA models, and polymer fingerprints to train tree-based models are provided on Zenodo: https://doi.org/10.5281/zenodo.17630815.Appendix A: Additional subgraphing requirements
The subgraphs used as inputs to our model comply with the following requirements:(1) In the case of a copolymer, the context subgraph should include elements from both monomers: predicting a part of monomer B, if monomer B is missing from the context is not possible.
(2) Every edge and every node should be in at least one subgraph32,39 to include full global information and full input representation.
(3) The context patch (subgraph) should be larger, hence more informative, than the targets patches (subgraphs) we are trying to predict from the context.45
(4) The target subgraphs should have minimal overlap with the context subgraph32,45 to make the prediction task less trivial.
(5) The context and targets subgraphs should change at every training loop to prevent overfitting.32,45
(6) For every directed edge evu in a subgraph, include also the edge euv, to comply with encoder architecture (wD-MPNN1).
Appendix B: Node-centred and edge-centred message passing with wD-MPNN
The choice of edge-centred convolutions differs from most common GNNs, which use node-centred message passing. The motivation behind this design, explained in ref. 43 is to prevent totters, that is, to avoid messages being passed along any path of the form v1v2…vn where vi = vi+2 for some i, which are thought to introduce noise into the graph representation by creating unnecessary loops in the message passing trajectory. However, we hypothesize that such loops are not less relevant for the dataset considered, as the polymer graphs utilized are unlikely to contain such small cyclic patterns. Working with edge-centred convolutions requires big and sparse adjacency matrices of shape Ae×e and Ae×n, which makes the training process more cumbersome, also due to the edge-centred convolution not being well supported in Pytorch Geometric. We compare the performance of the convolution methods predicting the EA and IP of the conjugated copolymer dataset in Tables 5 and 6 respectively. The results were obtained in the scenario where we train end-to-end on 80% of the data, and test on 20%. As visible in the tables, both methods achieve the same results, making the node-centred convolution a good solution, given the ease of implementation.| wD-MPNN | R 2 ↑ | RMSE ↓ |
|---|---|---|
| Edge-centred | 0.998 ± 0.0003 | 0.029 ± 0.002 |
| Node-centred | 0.998 ± 0.0002 | 0.027 ± 0.001 |
| wD-MPNN | R 2 ↑ | RMSE ↓ |
|---|---|---|
| Edge-centred | 0.998 ± 0.0007 | 0.022 ± 0.004 |
| Node-centred | 0.997 ± 0.0004 | 0.025 ± 0.003 |
Appendix C: Additional notes on model variants
In addition to the JEPA model variant presented in the main text (Model II, Fig. 3), we also experimented with an alternative architecture (Model I, Fig. 10). Model I is conceptually closer to Graph-JEPA,32 but was ultimately less effective in our setting. We include it here for the sake of completeness, along with a comparison between the two approaches.Architectural differences
Model I adopts a hybrid design, where an initial GNN encoder (wD-MPNN) processes all subgraphs obtained from the partitioning algorithm. The resulting subgraph embeddings are subsequently contextualized through a transformer-based target encoder Etarget, allowing global information exchange via self-attention. A symmetric context encoder Econtext is required when using momentum updates or weight sharing to prevent trivial information leakage.Model II (in the main manuscript), in contrast, simplifies the architecture by replacing the transformer encoders with wD-MPNNs for both context and target encoding, while removing the initial GNN stage. Here, Econtext operates on the context subgraph, and Etarget directly processes the full graph, from which target subgraph embeddings are pooled. This design yields a more parameter-efficient model, avoids intermediate subgraph embeddings, and naturally preserves positional information.
Performance comparison
Fig. 11 reports the predictive performance of both models under the configuration of 60% context size, 10% target size, one predicted target, and random-walk subgraphing.Model II clearly outperforms Model I, demonstrating better stability and accuracy, particularly in low-data regimes. Several factors may explain this: (i) Model I is more complex, combining wD-MPNN and transformer encoders, and likely requires more data to converge effectively. (ii) Model I contextualizes targets only through small subgraphs, which may discard positional cues critical for polymer graphs composed of multiple monomer units. In contrast, Model II trains the target encoder on the full graph, preserving this information. (iii) All hyperparameters (subgraph sizes, target number, and partitioning strategy) were tuned for Model II, potentially biasing results in its favor.
Regarding pretraining, the two variants behave differently. Model II benefits consistently from pretraining across all finetuning dataset sizes. In contrast, Model I shows improvement only in the most data-scarce scenarios; as the amount of finetuning data increases, the effect of pretraining becomes redundant or even slightly detrimental, suggesting that noise may dominate the learned representations in this configuration.
Appendix D: Subgraph size distribution analysis
We analyzed the distribution of context and target subgraph sizes across the three partitioning algorithms (motif-based, METIS, and random walk) under the parameter setting of 60% context size and 10% target size and one target subgraph. Table 7 reports the average number of nodes per subgraph, the resulting context–target ratios, and adherence to the specified parameters.| Method | Context | Target | Ratio | Adherence |
|---|---|---|---|---|
| Motif-based | 14.6 ± 2.8 (69.2%) | 4.8 ± 0.9 (23.2%) | 3.1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
Context 115%, target 232% |
| METIS | 14.6 ± 3.2 (68.5%) | 3.1 ± 0.7 (14.6%) | 4.7![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
Context 114%, target 146% |
| Random walk | 13.4 ± 2.6 (62.9%) | 3.8 ± 1.4 (18.6%) | 3.5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 1 |
Context 105%, target 186% |
We observe that all methods produce larger-than-expected targets due to implementation constraints (chemical validity, connectivity preservation, and meaningfulness of partitions). Nevertheless, the methods differ in how closely they approximate the intended proportions: random walk adheres most closely to the context size, METIS yields the most accurate sizes for targets, and motif-based partitions result in the largest context and targets compared to desired sizes.
Appendix E: Pretraining dynamics of JEPA
Fig. 12a and b show representative training and validation L2 (embedding) loss trajectories during JEPA pretraining, with and without a normalization layer after the graph pooling layer in context and target encoder.With the normalization layer, JEPA shows the expected smooth decline of both training and validation losses, indicating numerically stable optimization. In contrast, when the normalization layer is removed, the loss curves fluctuate and generally increase with increasing epochs. This behavior arises because the magnitude of latent embeddings can drift during training, leading to apparent divergence in the raw L2 loss scale.Despite this, we found the variant without normalization (closer to original JEPA implementation) to achieve superior downstream performance. We attribute this to increased representational flexibility: normalization constrains the embedding space and may reduce expressiveness. Moreover, in JEPA-style objectives, context–target pairs are resampled each epoch via random-walk subgraphing, so the prediction targets continually change, preventing the loss from flattening to a stationary minimum. Similar patterns have been reported in recent JEPA implementations (e.g., C-JEPA), and reflect the stochastic nature of the training dynamics rather than instability.
As a reference, we also provide the learning curves for self-supervised pretraining using node/edge masking (Fig. 13a) and pseudolabel prediction (Fig. 13b) of polymer graphs, as introduced by Gao et al.15
 | ||
| Fig. 13 MSE loss curves for node/edge masking and pseudolabel prediction, as introduced by Gao et al.15 (a) MSE loss curve for masked node/edge prediction. (b) MSE loss curve for pseudolabel prediction. | ||
Appendix F: Monomer-based data split
To evaluate out-of-distribution (OOD) generalization, we perform a monomer-based data split in which all polymers containing a specific A-monomer are excluded from training and used as test set. This setup imposes a stronger distribution shift than random splits, requiring models to generalize to unseen chemical scaffolds.We conduct a nine-fold cross validation, each fold holding out one A-monomer. The compared models include the JEPA-pretrained wD-MPNN, the same model fine-tuned only, the input-space SSL model of Gao et al.,15 and tree-based baselines random forest and XGBoost trained on oligomer fingerprints, as described in the main manuscript. Because the monomer-holdout setup induces substantial variability between folds, results are summarized as boxplots across all folds and random seeds in Fig. 14 and 15. Additionally, we provide per-monomer RMSE curves for the different finetune dataset size, shown in Fig. 16.
 | ||
| Fig. 14 R 2 performance under the monomer-holdout split for wD-MPNN without and with JEPA-based pretraining, tree-based baselines, and input-space SSL (Gao et al.15). Higher is better. | ||
 | ||
| Fig. 16 Monomer-specific RMSE curves for JEPA pretraining, input-space SSL (Gao et al.15), and the non-pretrained baseline, illustrating variability in generalization across monomer types. The titles of the subfigures are the respective monomer SMILES of the held out test A-monomer. | ||
Aggregated results
Embedding-space JEPA pretraining improves median performance and reduces variance across test monomer folds, though less strongly than input-space SSL. As shown in Fig. 14 and 15, JEPA pretraining generally increases the median R2 and lowers RMSE relative to training from scratch, while narrowing fold-to-fold variability. These gains are smaller than those obtained under random splits (Fig. 17). The input-space SSL approach (node/edge masking with a pseudolabel objective) yields larger median improvements, suggesting stronger OOD generalization than our embedding-based SSL strategy. | ||
| Fig. 17 RMSE performance under the random split for wD-MPNN without and with JEPA-based pretraining, tree-based baselines, and input-space SSL (Gao et al.15). Lower is better. | ||
The tree-based baselines random forest and XGBoost trained on handcrafted fingerprints are less competitive under the monomer-holdout split compared to the random split scenario. This indicates that fingerprint-based models tend to be more sensitive to distribution shifts than graph-based methods that learn directly from molecular structure.
Per-monomer analysis
Fig. 16 reveals strong monomer-specific variation in performance, reflected by differing RMSE scales on the y-axes. This aligns with the large variance in Fig. 15 compared to the more uniform results under random splits (Fig. 17).Interestingly, JEPA-based and input-space SSL models excel on different monomers. For instance, JEPA performs better for (*)c1ccc(*)cc1, whereas input-space SSL performs better for (*)c1cc2cc3sc(*)cc3cc2s1. For each held-out monomer, the plots also report the maximum Tanimoto similarity to any training-set monomer (upper-right corner). However, we observe no consistent relationship between similarity, RMSE magnitude, and the relative benefit of pretraining.
Appendix G: Additional plots
In Fig. 17, we show an additional boxplot for the RMSE in the random split scenario to illustrate the different fold-to-fold variances compared to the results in Appendix F.Acknowledgements
We thank Qinghe Gao for the discussions about SSL on polymer graphs. This work is supported by the AI4b.io program, a collaboration between TU Delft and dsm-firmenich, and is fully funded by dsm-firmenich and the RVO (Rijksdienst voor Ondernemend Nederland).References
- M. Aldeghi and C. W. Coley, A graph representation of molecular ensembles for polymer property prediction, Chem. Sci., 2022, 13(35), 10486–10498, 10.1039/D2SC02839E.
- C. Kuenneth, A. C. Rajan, H. Tran, L. Chen, C. Kim and R. Ramprasad, Polymer informatics with multi-task learning, Patterns, 2021, 2(4), 100238, DOI:10.1016/j.patter.2021.100238.
- C. Kuenneth and R. Ramprasad, polybert: a chemical language model to enable fully machine-driven ultrafast polymer informatics, Nat. Commun., 2023, 14(1), 4099, DOI:10.1038/s41467-023-39868-6.
- Y. Zhao, R. J. Mulder, S. Houshyar and T. C. Le, A review on the application of molecular descriptors and machine learning in polymer design, Polym. Chem., 2023, 14(29), 3325–3346, 10.1039/D3PY00395G.
- K. Sattari, Y. Xie and J. Lin, Data-driven algorithms for inverse design of polymers, Soft Matter, 2021, 17(33), 7607–7622, 10.1039/D1SM00725D.
- T. B. Martin and D. J. Audus, Emerging trends in machine learning: a polymer perspective, ACS Polym. Au, 2023, 3(3), 239–258, DOI:10.1021/acspolymersau.2c00053.
- C. Yan and G. Li, The rise of machine learning in polymer discovery, Advanced Intelligent Systems, 2023, 5(4), 2200243, DOI:10.1002/aisy.202200243.
- L. Chen, G. Pilania, R. Batra, T. D. Huan, C. Kim, C. Kuenneth and R. Ramprasad, Polymer informatics: current status and critical next steps, Mater. Sci. Eng., R, 2021, 144, 100595, DOI:10.1016/j.mser.2020.100595.
- P. Reiser, M. Neubert, A. Eberhard, L. Torresi, C. Zhou, C. Shao, H. Metni, C. van Hoesel, H. Schopmans, T. Sommer and P. Friederich, Graph neural networks for materials science and chemistry, Commun. Mater., 2022, 3(1), 1–18, DOI:10.1038/s43246-022-00315-6.
- R. A. Patel, C. H. Borca and M. A. Webb, Featurization strategies for polymer sequence or composition design by machine learning, Mol. Syst. Des. Eng., 2022, 7(6), 661–676, 10.1039/D1ME00160D.
- H. Yamada, C. Liu, S. Wu, Y. Koyama, S. Ju, J. Shiomi, J. Morikawa and R. Yoshida, Predicting materials properties with little data using shotgun transfer learning, ACS Cent. Sci., 2019, 5(10), 1717–1730, DOI:10.1021/acscentsci.9b00804.
- P. Zhang, L. Kearney, D. Bhowmik, Z. Fox, A. K. Naskar and J. Gounley, Transferring a Molecular Foundation Model for Polymer Property Predictions, J. Chem. Inf. Model., 2023, 63(24), 7689–7698 CrossRef CAS PubMed.
- R. Gurnani, C. Kuenneth, A. Toland and R. Ramprasad, Polymer informatics at scale with multitask graph neural networks, Chem. Mater., 2023, 35(4), 1560–1567, DOI:10.1021/acs.chemmater.2c02991.
- O. Queen, G. A. McCarver, S. Thatigotla, B. P. Abolins, C. L. Brown, V. Maroulas and K. D. Vogiatzis, Polymer graph neural networks for multitask property learning, npj Comput. Mater., 2023, 9(1), 1–10, DOI:10.1038/s41524-023-01034-3.
- Q. Gao, T. Dukker, A. M. Schweidtmann and J. M. Weber, Self-supervised graph neural networks for polymer property prediction, Mol. Syst. Des. Eng., 2024, 9(11), 1130–1143 RSC.
- C. Xu, Y. Wang and A. Barati Farimani, Transpolymer: a transformer-based language model for polymer property predictions, npj Comput. Mater., 2023, 9(1), 1–14, DOI:10.1038/s41524-023-01016-5.
- J. Zhou, Y. Yang, A. M. Mroz and K. E. Jelfs, Polycl: contrastive learning for polymer representation learning via explicit and implicit augmentations, Digital Discovery, 2025, 4(1), 149–160 Search PubMed.
- M. Caron, H. Touvron, I. Misra, H. Jégou, J. Mairal, P. Bojanowski and A. Joulin, Emerging properties in self-supervised vision transformers, in Proceedings of the IEEE/CVF International Conference on Computer Vision, 2021, pp. 9650–9660 Search PubMed.
- T. Chen, S. Kornblith, M. Norouzi and G. Hinton, A simple framework for contrastive learning of visual representations, in Proceedings of the 37th International Conference on Machine Learning, PMLR, Atlanta, Georgia, USA, 2020, pp. 1597–1607 Search PubMed.
- K. He, X. Chen, S. Xie, Y. Li, P. Dollár and R. Girshick, Masked autoencoders are scalable vision learners, in Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2022, pp. 16000–16009 Search PubMed.
- T. Uelwer, J. Robine, S. S. Wagner, M. Höftmann, E. Upschulte, S. Konietzny, M. Behrendt and S. Harmeling, A Survey on Self-Supervised Representation Learning, arXiv, 2023, preprint, arXiv:2308.11455, DOI:10.48550/arXiv.2308.11455.
- J. Devlin, M.-W. Chang, K. Lee and K. Toutanova, Bert: pre-training of deep bidirectional transformers for language understanding, in Proceedings of the 2019 conference of the North American chapter of the association for computational linguistics: human language technologies, 2019, vol. 1 (long and short papers), pp. 4171–4186 Search PubMed.
- A. Radford, K. Narasimhan, T. Salimans, I. Sutskever, et al., Improving language understanding by generative pre-training, 2018.
- W. Jin, T. Derr, H. Liu, Y. Wang, S. Wang, Z. Liu and J. Tang, Self-Supervised Learning on Graphs: Deep Insights and New Direction, arXiv, 2020, preprint, arXiv:2006.10141, DOI:10.48550/arXiv.2006.10141.
- Y. Liu, M. Jin, S. Pan, C. Zhou, Y. Zheng, F. Xia and P. S. Yu, Graph self-supervised learning: a survey, IEEE Transactions on Knowledge and Data Engineering, 2023, 35(6), 5879–5900, DOI:10.1109/TKDE.2022.3172903.
- Y. Xie, Z. Xu, J. Zhang, Z. Wang and S. Ji, Self-supervised learning of graph neural networks: a unified review, IEEE Trans. Pattern Anal. Mach. Intell., 2023, 45(2), 2412–2429, DOI:10.1109/TPAMI.2022.3170559.
- M. Sun, J. Xing, H. Wang, B. Chen and J. Zhou, Mocl: data-driven molecular fingerprint via knowledge-aware contrastive learning from molecular graph, in Proceedings of the 27th ACM SIGKDD Conference on Knowledge Discovery & Data Mining, ACM, Virtual Event Singapore, 2021, pp. 3585–3594, DOI:10.1145/3447548.3467186.
- Z. Zhang, Q. Liu, H. Wang, C. Lu and C.-K. Lee, Motif-based graph self-supervised learning for molecular property prediction, Advances in Neural Information Processing Systems, 2021, 34, 15870–15882 Search PubMed.
- Y. Rong, Y. Bian, T. Xu, W. Xie, Y. Wei, W. Huang and J. Huang, Self-supervised graph transformer on large-scale molecular data, Advances in Neural Information Processing Systems, 2020, 33, 12559–12571 Search PubMed.
- W. Hu, B. Liu, J. Gomes, M. Zitnik, P. Liang, V. Pande and J. Leskovec, Strategies for pre-training graph neural networks, arXiv, 2019, preprint, arXiv:1905.12265, DOI:10.48550/arXiv.1905.12265.
- Y. LeCun, A path towards autonomous machine intelligence version 0.9. 2, 2022-06-27, Open Rev., 2022, 62(1), 1–62 Search PubMed.
- G. Skenderi, H. Li, J. Tang and M. Cristani, Graph-Level Representation Learning with Joint-Embedding Predictive Architectures, arXiv, 2023, preprint, arXiv:2309.16014, DOI:10.48550/arXiv.2309.16014.
- Y. Bai, L. Wilbraham, B. J. Slater, M. A. Zwijnenburg, R. S. Sprick and A. I. Cooper, Accelerated discovery of organic polymer photocatalysts for hydrogen evolution from water through the integration of experiment and theory, J. Am. Chem. Soc., 2019, 141(22), 9063–9071 Search PubMed.
- A. Arora, T.-S. Lin, N. J. Rebello, S. H. Av-Ron, H. Mochigase and B. D. Olsen, Random forest predictor for diblock copolymer phase behavior, ACS Macro Lett., 2021, 10(11), 1339–1345 Search PubMed.
- G. Karypis and V. Kumar, A fast and high quality multilevel scheme for partitioning irregular graphs, SIAM Journal on Scientific Computing, 1998, 20(1), 359–392 Search PubMed.
- J. Degen, C. Wegscheid-Gerlach, A. Zaliani and M. Rarey, On the art of compiling and using ‘drug-like’ chemical fragment spaces, ChemMedChem, 2008, 3(10), 1503–1507, DOI:10.1002/cmdc.200800178.
- L. Zhang, V. Rao and W. Cornell, r-brics–a revised brics module that breaks ring structures and carbon chains, ChemMedChem, 2024, 202300202 Search PubMed.
- V. P. Dwivedi, A. T. Luu, T. Laurent, Y. Bengio and X. Bresson, Graph neural networks with learnable structural and positional representations, arXiv, 2021, preprint, arXiv:2110.07875, DOI:10.48550/arXiv.2110.07875.
- X. He, B. Hooi, T. Laurent, A. Perold, Y. Lecun and X. Bresson, A generalization of vit/mlp-mixer to graphs, in Proceedings of the 40th International Conference on Machine Learning, PMLR, Honolulu, Hawaii, USA, 2023, pp. 12724–12745 Search PubMed.
- D. Rogers and M. Hahn, Extended-connectivity fingerprints, J. Chem. Inf. Model., 2010, 50(5), 742–754 CrossRef CAS PubMed.
- G. Landrum, et al., RDKit: open-source cheminformatics, 2006 Search PubMed.
- E. R. Antoniuk, P. Li, B. Kailkhura and A. M. Hiszpanski, Representing polymers as periodic graphs with learned descriptors for accurate polymer property predictions, J. Chem. Inf. Model., 2022, 62(22), 5435–5445, DOI:10.1021/acs.jcim.2c00875.
- K. Yang, K. Swanson, W. Jin, C. Coley, P. Eiden, H. Gao, A. Guzman-Perez, T. Hopper, B. Kelley, M. Mathea, A. Palmer, V. Settels, T. Jaakkola, K. Jensen and R. Barzilay, Analyzing learned molecular representations for property prediction, J. Chem. Inf. Model., 2019, 59(8), 3370–3388, DOI:10.1021/acs.jcim.9b00237.
- E. I. Sanchez Medina, S. Kunchapu and K. Sundmacher, Gibbs–Helmholtz graph neural network for the prediction of activity coefficients of polymer solutions at infinite dilution, J. Phys. Chem. A, 2023, 127(46), 9863–9873 CrossRef CAS PubMed.
- M. Assran, Q. Duval, I. Misra, P. Bojanowski, P. Vincent, M. Rabbat, Y. LeCun and N. Ballas, Self-Supervised Learning from Images with a Joint-Embedding Predictive Architecture, in Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 2023, pp. 15619–15629 Search PubMed.
Footnotes |
| † These authors contributed equally to this work. |
| ‡ Francesco Piccoli completed this work before joining Amazon. |
| This journal is © The Royal Society of Chemistry 2026 |