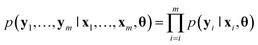Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
On-the-fly fine-tuning of foundational neural network potentials: a Bayesian neural network approach
Tim
Rensmeyer
 *,
Denis
Kramer
and
Oliver
Niggemann
*,
Denis
Kramer
and
Oliver
Niggemann
Helmut-Schmidt-University Hamburg, Hamburg, Germany. E-mail: rensmeyt@hsu.hamburg
First published on 7th April 2026
Abstract
Due to the computational complexity of evaluating interatomic forces from first principles, the creation of interatomic machine learning force fields has become a highly active field of research. However, the generation of training datasets of sufficient size and sample diversity itself comes with a computational burden that can make this approach impractical for modeling rare events or systems with a large configuration space. Fine-tuning foundation models that have been pre-trained on large-scale material or molecular databases offers a promising opportunity to reduce the amount of training data necessary to reach a desired level of accuracy. However, even if this approach requires less training data overall, creating a suitable training dataset can still be a very challenging problem, especially for systems with rare events and for end-users who don't have an extensive background in machine learning. In on-the-fly learning, the creation of a training dataset can be largely automated by using model uncertainty during the simulation to decide if the model is accurate enough or if a structure should be recalculated with quantum-chemical methods and used to update the model. A key challenge for applying this form of active learning to the fine-tuning of foundation models is how to assess the uncertainty of those models during the fine-tuning process, even though most foundation models lack any form of uncertainty quantification. In this paper, we overcome this challenge by introducing a fine-tuning approach based on Bayesian neural network methods and a subsequent on-the-fly workflow that automatically fine-tunes the model while maintaining a pre-specified accuracy and can detect rare events such as transition states and sample them at an increased rate relative to their occurrence.
1 Introduction
Ever since the discovery of the laws of quantum mechanics a century ago, the prediction of molecular and material properties such as stress–strain relationships or catalytic activity from first principles has, in theory, been possible.1,2 However, in practice, this task remains challenging even to this day.3,4 The major difficulty lies in the exponentially growing computational complexity of solving the underlying Schrödinger equation with an increasing number of electrons.1 As a consequence, several approximate methods for property prediction have been developed, which are computationally tractable for larger systems at the cost of varying degrees of accuracy.Density Functional Theory (DFT) in particular, has established itself as a valuable tool in computational chemistry that allows the computation of many material and molecular properties, such as electronic structure, binding energies and interatomic forces with a high degree of accuracy and a computational complexity that is feasible on a typical high-performance cluster for many tasks.4 While DFT has enabled the investigation of the properties of individual materials at quantum mechanical accuracy, high-throughput screening of materials or molecules for desired properties still remains very computationally demanding. Furthermore, Molecular Dynamics (MD) – the simulation of the time evolution of molecular systems and materials – remains challenging, as the forces on all atoms have to be calculated at each timestep. This severely limits the time horizon that can be achieved in a practical amount of time using DFT.2
Subsequently, the development of machine learning models that can predict interatomic forces has become an active field of research.5 Here, neural networks have become a promising approach. These models have made great strides in the past years and can achieve much higher accuracy than previous methods with just a few hundred well-sampled training configurations of a specific system, in some cases.6–11 Even more alluring is the prospect of not starting the training from scratch but instead using one of the models from the growing collection of publicly available foundation models.12–19 These models have been pre-trained on large databases of materials or molecules and can often model the overall dynamics of the system to a certain degree of accuracy, but frequently fail to capture subtle physical effects that might be of particular importance, as will be illustrated in Section 5. Moreover, the accuracy of a foundation model is inherently limited by the level of theory used for its pretraining dataset, which is often chosen on the basis of speed and stability rather than accuracy. Fine-tuning is, therefore, often necessary to model specific system details accurately and to bridge the gap between the level of theory used for pre-training and the level that is required for modeling specific properties accurately.20 Fine-tuning such pre-trained models can significantly reduce the amount of training data necessary to reach a desired accuracy when compared to training from scratch,20–25 making it an attractive approach.
However, a key challenge that remains even for fine-tuning is how to select data points for training datasets of a specific system of interest. While existing algorithms for fine-tuning foundation models perform well on benchmark datasets, those benchmark datasets are usually subsampled from very long MD-trajectories in DFT that cover the entire space of atomic arrangements that can occur. Simulating such an exhaustive trajectory to generate a training set in an applied context, where a certain system of interest is supposed to be investigated, can quickly become almost as computationally demanding as the simulation the neural network was supposed to replace. This is especially problematic for systems with high-dimensional configuration spaces with large degrees of freedom, for example, due to many rotational degrees of freedom of single σ-bonds or transition pathways between meta-stable configurations or phases of the system.
At first glance, computing the trajectory at a lower accuracy and computational complexity (e.g., with a minimal basis set) and then recalculating some subset of those configurations at the desired accuracy appears like a simple solution to increase sample diversity and reduce computational demand. However, this is often not feasible. For example, using minimal basis sets in DFT can alter the predicted equilibrium bond distances by a few percent when compared to large basis sets. In practice, this can lead to almost disjoint radial distributions, as shown for the nitrogen molecule in Fig. 1, even though many other physical properties, such as formation energy and vibrational frequency, can be similar to the higher accuracy method.
 | ||
| Fig. 1 Radial distribution functions of a nitrogen molecule at 500 kelvin computed with a minimal STO-3G basis set and an extensive triple-Zeta basis set. A B3LYP functional26–29 was used with both basis sets. Because the minimal basis set overestimates the equilibrium bond distance with a prediction of 1.134 Å, unlabeled data generated from a simulation at this temperature with this basis set will contain almost no molecular geometries where the bond distance is below the equilibrium bond distance of around 1.085 Å predicted by the more accurate triple-Zeta basis set. Therefore, labeling this data with the triple-zeta basis will lead to a biased training set that lacks information from the repulsive regime of this basis set, providing a significant risk for the model to degrade in accuracy for such states during a simulation. For reference, the experimental value for the bond distance is 1.098 Å which is significantly closer to the prediction with the triple-Zeta basis set. | ||
Using the foundational neural network model itself to generate the training data for fine-tuning is also often not that easy. For example, the model might predict the wrong phases at the target temperature, as will be illustrated in a later example. This is especially problematic in instances where the correct phase diagram of the material is not known beforehand.
In general, a problem with using a faster approximate method to generate unlabeled training data is that while trajectories generated by such models can often be accurate, it is difficult to assess if this is the case in any particular application. This is especially challenging because qualitatively inaccurate modeling of a system phenomenon by such a model might still appear plausible without extensive analysis a posteriori. If this model simulates a process of interest, such as a state transition or phase behaviour, qualitatively inaccurately, then the training data generated from such a simulation will lack explicit and accurate examples of this phenomenon of interest. Therefore, it is difficult to assess whether the subsequently finetuned model will be able to accurately simulate the processes of interest, since a lack of relevant training structures could potentially cause a significant degradation in its accuracy. This poses a significant obstacle to the trustworthiness of the fine-tuned model.
Even though researchers with sufficient experience in machine learning force fields often find ways to create training datasets for specific applications, the above discussion illustrates the significant challenges associated with and the work required in creating training datasets in general. The tedious and challenging burden of manually generating training datasets should, therefore, ideally be automated in order to allow computational material scientists and chemists to focus on the underlying physics and to lower the technical barrier of entry for fine-tuning models to a reliable, trustworthy and accurate state for less experienced users of machine learning force fields.
Moreover, for finetuning the foundational neural network potentials inside high-throughput materials discovery workflows, for example, for calculating thermodynamic properties of candidate materials, it would be very desirable if the finetuning process were automated completely without requiring human input for each material.
Uncertainty-based active learning methods have established themselves as a way to automate the selection of training data30–33 by sampling structures with high model uncertainty and potential energies that are low enough to be accessible at the target temperature. Uncertainty-quantification methods and uncertainty-based active learning are also attractive, because they provide an additional layer of trustworthiness and reliability by aiming to maintain high model confidence for all structures that have a high likelihood of occurring during simulations, even during rare events. Thus, the use of uncertainty-based active learning for fine-tuning foundation models appears very promising.
For example, on-the-fly learning is an elegant approach30,34 where the pre-trained model might be used to drive the dynamics (e.g., MD simulation, transition state optimization, etc.) until a configuration is encountered that exceeds a certain uncertainty threshold. This configuration is then recalculated with quantum-chemical simulation methods and used to update the model, which then resumes the task until the next configuration above the uncertainty threshold is encountered (Fig. 2). This approach has the additional benefit of being very easy to use for non-machine learning experts since essentially only the initial state of the system and the uncertainty threshold would have to be specified.
 | ||
| Fig. 2 An illustration of the on-the-fly learning approach based on Bayesian neural networks introduced in this work. | ||
Unfortunately, most foundational neural network potentials do not come with an uncertainty estimate for their predictions. Hence, the development of a framework to systematically update a foundational neural network potential on new data while also being able to assess its uncertainty would be very desirable.
Another challenge for fine-tuning foundational neural network potentials on-the-fly is that the fine-tuning algorithm has to be able to fit the training data and avoid overfitting neural networks with millions of trainable parameters while progressively building up the training dataset from a single initial sample to possibly hundreds of training samples, even though a validation dataset to detect overfitting is unavailable.
Moreover, a general challenge in uncertainty quantification with neural networks is that even if the predicted uncertainties are very well correlated with the error, this correspondence is often not one-to-one. As an example, while a higher standard deviation in the predictions of an ensemble of models might imply a larger error than a smaller one, a standard deviation of σpred = 1 in the predictions might correspond to an actual standard deviation in the error of σobserved = 1.5. If a validation set exists, the uncertainty estimates can be calibrated. For example, by changing the predicted standard deviation to σpred → ξ × σpred where the rescaling factor ξ is estimated on the validation set by matching the mean predicted variance to the observed mean squared error on the validation set. Unfortunately, in the case of on-the-fly learning, this is not an option, since no validation set exists. In general, only the predicted uncertainties and observed errors from the configurations that were recalculated in DFT due to a large uncertainty can be used as empirical data to estimate the miscalibration of the model.
A compounding challenge is that if at one point the recalibration factor is wrongfully estimated as too small, this miscalibration might not be correctable, since such a miscalibration inhibits new DFT calls that could be used to calibrate the model more accurately.
Bayesian neural networks have demonstrated a high quality of uncertainty quantification comparable to classical ensemble-based methods of uncertainty quantification for machine learning models of interatomic forces and potential energies.35,36 Further, they are inherently more robust to overfitting by weighing preexisting knowledge in the form of the Bayesian prior density and empirical evidence from training data via Bayes' theorem.37,38
Due to these inherent properties, the principal research question of this paper is whether it is possible to develop a framework for on-the-fly fine-tuning of foundation models based on the formalism of Bayesian neural networks that is capable of addressing the additional challenges mentioned above.
The main contributions of this work are the following:
• We develop a simple Bayesian framework for uncertainty-aware fine-tuning of foundation models, by harnessing a simple transfer learning prior as well as Monte Carlo Markov Chain (MCMC) sampling of an ensemble of models and assessing the uncertainty via the disagreement in the predictions of the models.
• We introduce a method for on-the-fly calibration of the uncertainties during the fine-tuning process.
• We demonstrate that the resulting on-the-fly learning workflow is capable of automatically fine-tuning foundation models to a pre-specified accuracy and biasing the training dataset towards rare events.
2 Interatomic force modeling using Bayesian neural networks
Throughout the rest of this paper, bold case symbols designate vectors and y|x denotes y conditioned on x.Machine-learned interatomic force fields for molecules aim to map an atomic configuration {(r1, z1), …, (rn, zn)}, composed of the nuclear coordinates  and nuclear charges zi, to the potential energy E and forces Fi acting on each nucleus i ∈ {1, …, n}. The predicted forces can then, for example, be used in combination with Newton's equations of motion to model the time evolution of the molecules. For materials, the input will usually also contain the lattice vectors, and the stress tensor will often be predicted as well. Because generating large amounts of high-quality training data is typically infeasible due to the high computational demand of quantum-chemical simulation methods, modern machine learning models have several forms of physical constraints built into them, in order to make them more data efficient,7–9,11,39,40 such as energy conservation by calculating the forces as the negative analytical gradient of the potential energy with respect to the atomic positions. Further, rotation invariance of the potential energy is explicitly built into modern neural network architectures.
and nuclear charges zi, to the potential energy E and forces Fi acting on each nucleus i ∈ {1, …, n}. The predicted forces can then, for example, be used in combination with Newton's equations of motion to model the time evolution of the molecules. For materials, the input will usually also contain the lattice vectors, and the stress tensor will often be predicted as well. Because generating large amounts of high-quality training data is typically infeasible due to the high computational demand of quantum-chemical simulation methods, modern machine learning models have several forms of physical constraints built into them, in order to make them more data efficient,7–9,11,39,40 such as energy conservation by calculating the forces as the negative analytical gradient of the potential energy with respect to the atomic positions. Further, rotation invariance of the potential energy is explicitly built into modern neural network architectures.
2.1 Bayesian neural networks
Bayesian neural networks have demonstrated promising results for modeling uncertainties in neural network predictions and, in particular, in machine learning force fields.35,36,41 The main difference between the Bayesian approach to neural networks and the regular approach is that the trainable parameters of the neural network, e.g., its weights and biases, are modeled probabilistically. For simplicity of notation, we denote by θ a vector containing all the trainable parameters of the neural network. For a given parameter vector θ and input sample x, the neural network predicts a probability density p(y|x, θ) over the target variable y. In the case of machine learning force fields for non-periodic systems x will be an atomic configuration {(r1, z1), …, (rn, zn)} and y will be the potential energy and atomic forces {E, F1, …, Fn}. For periodic systems, x will contain the lattice vectors as well, and y might additionally contain the stress tensor. The starting point of Bayesian methods is a prior density p(θ) over the parameters, which expresses a priori knowledge about which sets of parameters are likely to result in a good model of the underlying data distribution. Given some training dataset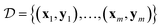 the prior density gets refined into the posterior density
the prior density gets refined into the posterior density 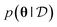 using Bayes' theorem:
using Bayes' theorem:With the mild assumption on conditional independence
this can be simplified to
where
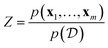 is a normalization constant. On a new input sample x, the probability distribution of the target variable y can then be calculated via
is a normalization constant. On a new input sample x, the probability distribution of the target variable y can then be calculated viaBecause this integral is almost never analytically tractable for neural networks, a Monte Carlo estimate is typically used:
where the parameter sets θi are sampled from the posterior density using either approximate inference42–44 or MCMC methods.45–47 MCMC methods, in particular, have displayed good results in uncertainty quantification48 due to their ability to sample different regions of the posterior. These methods work by simulating a stochastic process over the space of neural network parameters, which converges in distribution to the posterior.
2.2 Relation to other works
Even though Bayesian neural networks offer a very promising opportunity to systematically incorporate and update pre-existing knowledge via the prior density, we find that this approach is very underutilized in the literature. In fact, we could only find two instances in the literature where this was attempted.37,38 In the work by Chen et al.,38 transfer learning of simulated to experimental data was done via a Bayesian neural network prior. A simple isotropic Gaussian prior with a mean derived during pretraining was used in that work, which we will also employ here.A more sophisticated approach for constructing a transfer learning prior was introduced by Shwartz-Ziv et al.,37 where a rescaled local approximation of the posterior on the pretraining dataset was used as a prior. However, in the applications considered here, this approach is not practical since such a prior would have to be constructed during the pretraining of the foundation models, while we focus on fine-tuning foundation models that have already been pre-trained.
Transfer learning of a pre-trained model for interatomic force fields has been investigated by Kolluru et al.,21 Zaverkin et al.,22 Smith et al.23 and Falk et al.24 However, the modeling of uncertainty has not been under consideration in those works.
After presenting our initial investigation of the uncertainty-aware transfer learning approach in a non-archival peer-reviewed venue,49 there has very recently been one other work published that does uncertainty-aware fine-tuning.50 However, the integration into an active learning workflow, like on-the-fly learning, is not under investigation in that work. Further, they investigate instances where large training datasets of thousands of training samples are available and are more focused on fine-tuning for entire subclasses of materials. In contrast, we focus on fine-tuning for specific systems where the size of the training dataset varies from one to a few hundred training samples.
Finally, there are the seminal papers by Li et al.51 and Vandermause et al.30 for on-the-fly learning for molecular dynamics. However, they use Gaussian processes as their substitutional model, which are less data efficient than modern neural network architectures and furthermore require training from scratch instead of fine-tuning a pre-trained model.
3 Methodology
The fundamental prior assumption for fine-tuning neural network models to specific applications is that the weights of the pre-trained model are not quite right for the application and have to be adjusted by an unknown small change. The idea for the Bayesian fine-tuning approach is, that we model this uncertainty in the correct weights explicitly via a transfer learning prior that is a Gaussian distribution over the weights pTL ∼ N(θ0, σTL2I) which is centered around the weights of the pre-trained model θ0 and has a small standard deviation σTL. As we progressively build up our training dataset, we refine the initial uncertainty in the weights by applying Bayes' rule to calculate the posterior. From the posterior, we then generate several sets of neural network weights using Markov chain sampling, resulting in an ensemble of models. We can then assess the uncertainty in the prediction for new samples via the empirical variance of their predictions, which provides a Monte Carlo estimate of the posterior predictive uncertainty, while the average of their predictions can be used for the overall prediction.To calibrate the uncertainties on the fly, we use the following Bayesian procedure:
The uncalibrated Bayesian Neural Network (BNN) models the error eobs = Epred − Etrue as
To calibrate the model, we want to estimate a suitable value for λ in order to match the raw uncertainties produced by the model to actual error estimates. To do this, we use a Bayesian estimator for λ by introducing a prior in the form of a Gamma distribution over λ:
where Hyp2F1 denotes the hypergeometric function 2F1 and
 (see Appendix B.11 for the derivation of this result).
(see Appendix B.11 for the derivation of this result).
We use this result to decide if a DFT calculation should be done by specifying an error threshold K and calling a DFT calculation if the predicted probability of the error magnitude exceeding K is larger than five percent.If the probability is smaller than five percent, the model continues with the MD-simulation. Otherwise, a DFT reference calculation is done and added to the training dataset. Afterwards, we then use Markov chain sampling to generate an ensemble of eight neural networks from the updated posterior. To sample the posterior density, we use the AMSGrad version of the Stochastic Gradient Hamiltonian Monte Carlo (SGHMC) algorithm46 introduced by us in a previous work.36 We use the old Monte Carlo samples as a starting point for sampling the new ones and simulate short Markov Chains with those initial seeds and priority sampling of the newly added structure to achieve faster convergence (see Appendix B.5 for more details on the sampling procedure). Finally, the DFT forces and stresses are used to perform this MD step. The resulting workflow is illustrated in Fig. 2.
For our experiments, we use three different neural network potentials, NequIP,9 MACE12,13 and Equiformerv2.14 To apply the Bayesian neural network formalism to these models, we add a few layers to the models so that each model predicts a distribution instead of making point predictions. The details of these modifications are provided in Appendices B.1–B.5. Additional details about how predictions and uncertainty quantification are done with the BNN can be found in Appendix B.9.
We always label the initial atomic structure with DFT and include this first data point to sample the initial Monte Carlo samples. Because potential energies are only defined up to a constant and this constant might be different for the data the model was pre-trained on and the simulation software used for fine-tuning, we also use this initial labeled data point to construct an offset value that is added to all energies calculated by DFT during the on-the-fly simulation. This value is chosen so that the offset DFT calculated energy equals the potential energy predicted by the pre-trained model on the initial structure.
4 Empirical evaluation
The promise of transfer learning is to specialize a pre-trained model with limited computational effort to truthfully reflect subtle, but important aspects of a particular system of interest that are not adequately reproduced by the starting model. Hence, we have selected an initial illustrating example from a classical benchmark dataset of a surface-adsorbate system for neural network potentials, showcasing the improvement in data efficiency and the quality of the uncertainty quantification that results from the Bayesian neural network priors constructed from a pre-trained model. We then selected a number of “challenging” cases to demonstrate the power and general utility of the full on-the-fly finetuning workflow: we investigate a simple molecular system with several meta-stable conformers to highlight the ability to bias the training data towards transition states between meta-stable states, before turning to two solid state problems; first, we investigate a challenging phase transition due to subtle electronic effects in a highly relevant compound; finally, we address a dynamic problem where mobility of a migrating species is highly dependent on the dynamics of the host lattice.To illustrate the effect of training a Bayesian neural network model with the prior constructed from the weights of a pre-trained model, compared to training a model from scratch, we start out with a transfer learning scenario for potential energies of a surface adsorbate system. We use a publicly available EquiformerV2 model14 with 31 million trainable parameters, which was pre-trained on the OC20 dataset.52 For the target dataset, we choose the NbSiAs surface – COH adsorbate dataset from the OC20-Dense dataset.53 This system is not included in the pre-training data of this foundation model and both datasets are computed with identical DFT settings. Notably, this benchmark does not yet involve the on-the-fly learning workflow and is instead done on a publicly available benchmark dataset. We partition the dataset into training, validation, and test data and then generate Monte Carlo samples from the posterior resulting from the training dataset (see Appendix B.5 and C.4 for details). Here, we compare the model accuracy to a comparable BNN that is trained with an uninformative prior. Further, we illustrate the quality of the resulting uncertainty quantification by calculating the Area Under the Curve – Receiver Operating Characteristic (AUC-ROC) scores for determining whether a test sample has a prediction error that lies above or below 1 kcal mol−1 based on the standard deviation in the predictions of the model. Because recognizing samples with an error above a certain threshold becomes easier at higher accuracies, we plot the AUC-ROC scores over the Root Mean Squared Errors (RMSEs) on the test set for this metric. To assess the viability of this transfer learning approach to finetuning models, additional transfer learning ablation experiments were performed, which focused on transfer learning of molecular forces with a NequIP model in several settings, including a change of electronic structure theory from DFTB54 to DFT and DFT to CCSD(T).55 These experiments and their results are listed in Appendix A.
As the first benchmark for the proposed on-the-fly fine-tuning workflow, we investigate the on-the-fly fine-tuning of a NequIP9 model during a molecular dynamics simulation of an ethanol molecule at 300 kelvin. The NequIP model has 13.45 million parameters and was pre-trained by us on the SPICE dataset.56 Ethanol was not contained in the pre-training data and a different level of theory was used for fine-tuning (B3LYP26–29) and pre-training (ωB97M-D3(BJ)57–59). Despite its small size, ethanol is an interesting benchmark molecule because it has different meta-stable conformations – one anti and two symmetry-related gauche conformations (Fig. 4). Due to this, there are transition states between those conformations that are expected to occur during an MD-simulation at this temperature, which are relatively rare due to the higher potential energy when compared with the metastable conformations. In a good on-the-fly learning workflow, such transition states have to be identified by the uncertainty measure and added to the training dataset at a higher rate compared to the rate of their occurrence during the simulation to ensure the accuracy of the model during transitions. For this benchmark, we set the target accuracy as 0.5 kcal mol−1 and perform a 10 ps simulation.
For the second experiment with the proposed on-the-fly fine-tuning algorithm, we run a simulation of a 2 × 2 × 2 LaMnO3 supercell with the mace-mp0 medium model with 4.69 million trainable parameters12,13 pre-trained on the MPtrj dataset.15,60 This system is interesting because it undergoes a phase transition at around 750 kelvin (Fig. 5 a). LaMnO3 crystallizes in an orthorhombic phase with space group Pbnm below 750K due to the strong Jahn–Teller activity of manganese. The orthorhombic phase is an anti-ferromagnetic insulator in which an orbital ordering is established due to the cooperative Jahn–Teller effect breaking the degeneracy of the electronic configuration of Mn3+ (t2g3eg1).61 For this reason, we perform a 150 ps molecular dynamics simulation with two temperature jumps. For the first 25 ps, we run the simulation at 300 kelvin. At the 25 ps mark, we introduce a temperature jump to 800 kelvin. We keep this temperature for 100 ps, after which we introduce a second temperature jump back to 300 kelvin and continue the simulation for another 25 ps. Some geometries of this system were contained in the training data of the foundation model and we use the same level of theory for fine-tuning as was used for the pre-training (PBE + U62,63). However, as will be seen by the experiments, the non-fine-tuned model has insufficient accuracy to model this system physically accurately, making it a good benchmark for transfer learning experiments.
The final benchmark for the proposed on-the-fly fine-tuning algorithm is a proton diffusion simulation in a 160-atom CaZrS3 supercell at 1500 kelvin, again with the mace-mp0 medium model. This benchmark is challenging because proton mobility is intimately linked to the dynamics of the host lattice. Occasional close proximity of two sulfur atoms belonging to non-connected ZrS6 octahedra enables a direct jump between the two ZrS6 octahedra with virtually zero activation energy64 (Fig. 6a), resulting in unusually high proton mobility at low temperatures. The S–S distance is governed by a static lattice distortion due to size mismatches between Ca and Zr as well as the vibrational properties of the sulfur sublattice. Hence, this benchmark goes beyond the mere reflection of transition state energies. It also requires a correct representation of the dynamics of the host lattice. Accurately modeling the proton transport in this system, therefore, might require a very high accuracy of the model. Here, we evaluate the model with two different error thresholds, 5 kcal mol−1 and 15 kcal mol−1. We fine-tune both models on-the-fly during an initial 30 ps MD simulation and then use the fine-tuned models and the non-fine-tuned one to investigate the diffusion of protons in CaZrS3 with five additional 30 ps simulations, which was revealed by the initial training runs to be a well-suited time-horizon to estimate proton mobility. While some geometries of the host lattice were contained in the pre-training data of the foundation model, no such structures with interstitial protons were included. We use PBE62 level of theory for the fine-tuning run. We run the mace model and the DFT calculations without dispersion correction.
All experiments were done with 8 Monte Carlo samples, which we identified as a good tradeoff between computational complexity and quality of uncertainty quantification in previous benchmarks.36 Details of the sampling procedures can be found in Appendix B.5. More details for all the simulations can be found in Appendix B.6. We set σTL from the Bayesian neural network prior as 0.2 for all experiments involving the NequIP model, 0.5 for the MACE model and 0.02 for the EquiformerV2 model. For this method to be practically useful, it would be very desirable that the strength of the prior does not require precise fine-tuning to a value that is unique to each specific application. For this reason, we did not perform a deep hyperparameter optimization for each model and experiment. Instead, we started with a small value of σTL for each model and then incrementally relaxed it until it could fit the data from an initial task well and then used these values for all experiments with the corresponding model. For the MACE model, the calibration was the simulation of the LaMnO3 system, for the NequIP model, it was the paracetamol ablation experiment in Appendix A, and for the EquiformerV2 model, the NbSiAs surface – COH adsorbate dataset itself was used.
Based on the results of our transfer learning experiments in Appendix A, we chose a = 3 and b = 10 for the gamma-prior over the calibration parameter for the NequIP model. Because we did not have such results for the MACE model, we chose the more conservative parameters a = 1.5 and b = 10 for the corresponding experiments. The lengths of the training runs (or phases of training runs for LaMnO3) were determined adaptively by starting with a conservative time estimate for each system and extending it based on the frequency of DFT calls requested until DFT requests became infrequent. The most important factor influencing the difference in required training times is the size of the configuration space of each compound, with LaMnO3 requiring a significantly longer training time due to the occurring back and forth transitions between the high- and low-temperature configurations, which, due to finite size effects of the 2 × 2 × 2 supercell, still happen at 800K. Accuracy thresholds were chosen primarily by the size of the system and secondarily by the expected size of the configuration space for the specific simulations.
5 Results
5.1 Efficiency of the transfer learning approach on the NbSiAs surface – COH adsorbate dataset
Plotting the root mean square errors of the models for the potential energies on the test set, we see a considerable improvement in accuracy at a given size of the training dataset (Fig. 3 upper graph). Furthermore, the AUC-ROC scores for distinguishing low and high error samples from the test set based on the predicted uncertainty, which are shown in the lower graph of Fig. 3, demonstrate that the predicted uncertainty of the fine-tuned model is well suited for distinguishing low and high error samples. Also, we see the improvement in AUC-ROC scores on this benchmark with decreasing root mean square error on the test set, which was expected because samples with an error of more than 1 kcal mol−1 become more extreme outliers with increasing accuracy of the model. Note that an AUC-ROC score of 1 corresponds to perfect distinguishability and an AUC-ROC score of 0.5 corresponds to random guessing. All Monte Carlo samples were generated from a single very long Markov Chain, to make sure no pathological overfitting occurs (see Appendix B.5 for more details).5.2 Conformers and transition states of ethanol
On the ethanol benchmark, we find a very good consistency of the proposed on-the-fly fine-tuning algorithm with the specified error threshold of 0.5 kcal mol−1. Overall, we find that for the 65 DFT interventions, the model exceeded the error threshold 3 times, corresponding to 4.55 percent of interventions with an overall RMSE of 0.196 kcal mol−1 at the interventions. Only two DFT references were called after the first 5 ps of the simulation. We performed an additional analysis of 150 structures randomly sampled from the 10 ps trajectory, where the model exceeded the threshold only a single time, with an overall RMSE of 0.142 kcal mol−1, significantly outperforming the non-fine-tuned model, which has an RMSE of 0.958 kcal mol−1 on those structures.To evaluate this experiment qualitatively, we also analyzed the DFT interventions in relation to transition states between different meta-stable conformations. For this, we plotted the angle between the OH bond and the CC bond over time. By doing this, the anti-conformation can be identified by a small angle and the gauche conformation by a large angle. Further, transitions between anti and gauche conformations can be identified by large changes in the angle. Since gauche–gauche transitions are not very well visible via this angle, we highlighted them manually by labeling them by hand. The results are summarized in Fig. 4. As can be seen in this figure, after the initial learning phase, where the model has high uncertainty in general, the model is almost exclusively requesting DFT calls during the transitions between meta-stable conformations. Especially noteworthy is that in the initial three-ps timeframe, where almost all DFT calls happen, the gauche–gauche transitions make up only around five percent (0.155 ps) of the total time, but around 16 percent (10 out of 63) of the total DFT interventions of that timeframe happen during those transitions. Hence, gauche–gauche transition states were sampled at a rate more than three times higher relative to their rate of occurrence.5.3 Phase transition in LaMnO3
For the LaMnO3 system, we also find good consistency with the error threshold, with 4.5 percent of configurations at intervention time having an error above the threshold of 6 kcal mol−1. At the time of intervention, the on-the-fly learning model had an RMSE of 2.685 kcal mol−1 compared to an error of the non-fine-tuned model of 7.532 kcal mol−1 on the same structures. In contrast to the non-fine-tuned model, the fine-tuned model reproduces the correct phases at the two different temperatures, as can be seen by the three distinct MnO bond distances at lower temperatures in Fig. 5b. The observed bond distances of the fine-tuned model are in close agreement with the values of 2.235 Å, 2.002 Å and 1.947 Å that were computed by Gavin and Watson65 with DFT at the PBE + U level of theory, also used for this benchmark. To assess the quality of the uncertainty quantification, we recalculated 1800 structures from the simulation in DFT and evaluated the relationship between predicted (calibrated) standard deviations and error by scatter plotting the uncertainties over the error magnitude (Fig. 5c). Further, we estimated the RMSE at a given standard deviation by ordering the pairs of errors and predicted standard deviations by order of ascending standard deviations and then applying Gaussian smoothing (see Appendix B.10 for details). As can be seen in this figure, we have an overall good relationship between predicted standard deviation and error. However, it should be noted that there are instances where the error is quite large despite a low predicted standard deviation. Note that in order to better estimate the error at higher uncertainties, we did additional sampling of structures with a predicted standard deviation of around 2.1 kcal mol−1, 2.5 kcal mol−1 and 2.8 kcal mol−1, respectively, which causes the increase in the point density of the scatter plot at these values. Evaluating the accuracy for the sampled structures, we find that the model achieves an RMSE of 2.350 kcal mol−1 compared to 10.827 kcal mol−1 of the non-fine-tuned model, with 3.2 percent of samples being above the error threshold.Lastly, we also evaluated the number of DFT calls over time (Fig. 5d). We find that the model stops doing DFT reference calls very early during the initial low-temperature phase of the simulation. After the temperature jump, it starts doing DFT calls again, with the frequency decreasing over time. Further, the model stays well within the specified accuracy region with only four instances where the error is larger than the threshold at the time of intervention. For reference, we also plot the error of the non-fine-tuned model on the structures for which DFT calls were made, illustrating the improvement in accuracy.
5.4 Proton mobility in CaZrS3
Lastly, for the proton diffusion, we find that the models are outside the specified accuracy range of 5 and 15 kcal mol−1 at 2.5 and 3.2 percent of interventions, respectively. At interventions, the RMSEs of the models are 2.137 and 6.773 kcal mol−1 (0.57 and 1.78 meV atom−1). For comparison, the non-fine-tuned model has an RMSE of 10.99 kcal mol−1 (2.91 meV atom−1) on the intervention structures of the higher accuracy model. Towards the end of the training run, the disagreement thresholds for the models were 0.91 and 3.43 kcal mol−1. This highlights the necessity for model calibration because the RMSE is about 2 times larger than the model disagreement.After the initial training run, we use the fine-tuned models to analyse the square displacement of protons in CaZrS3. We perform five simulations with each model by first setting the target temperature with a Langevin thermostat during an initial one picosecond-long simulation. Afterwards, we perform a 30 ps simulation in the NVE ensemble. Because two protons were placed in the supercell of the simulation, this results in ten proton trajectories for each model.
The squared displacements for the different models are shown in Fig. 6c. We see a very large Mean Square Displacement (MSD) for the non-fine-tuned model, which is reduced significantly when fine-tuned with an error threshold of 15 kcal mol−1. Interestingly, fine-tuning to a lower error threshold of 5 kcal mol−1 leads to the MSD rising again slightly.
To explore the source of this phenomenon, we investigated the mechanism behind the proton jumps for the individual models. To do this, we analyzed the average sulfur–sulfur distance for the 50 fs leading up to proton jumps. For this, we have additional DFT reference data from a 30 ps DFT simulation from a previous research project in our group, where the average separation at the time of the jump was analyzed. The results are shown in Fig. 6b. For the low error threshold model, we find very good agreement with the DFT reference value. For the higher threshold model, the distance is slightly too large. This yields a possible explanation for the slightly lower proton mobility from this model, since the activation energy for jumps will be slightly larger at this separation. For the non-fine-tuned model, the average sulfur–sulfur distance at the time of the jump is much larger than the DFT reference. This demonstrates that the transition states underlying the proton jumps are not accurately modeled without finetuning, since the activation energy at such distances should be very high, resulting in low proton mobility instead of the erroneously high mobility that is predicted by the foundation model.
To further assess the accuracy of the models at test time, we sampled 100 random geometries from the first simulation for each of the already fine-tuned models. From the results shown in Fig. 6d, it can be seen that the models are within their target accuracy for those configurations. For reference, we again include an evaluation of the error of the non-fine-tuned model on the geometries sampled from the simulation of the high-accuracy model and find a much poorer accuracy.
Overall, it took 31 DFT calls to train the model to the higher threshold and 285 calls to train it to the lower error threshold.
We did an additional analysis to assess if, after the initial learning phase of 5000 simulation steps, there is a preference for DFT calls during transition states where the proton jumps between sulfur atoms. Interestingly, despite the large size of the system with 162 atoms, the overall uncertainty from the system did not completely drown out the uncertainty from the transition states in the low-threshold training run. In particular, we found a statistically highly significant increase in DFT calls (p = 0.0012) in 15 fs time windows around proton jumps, with an increase of 56 percent in the average number of DFT calls during time windows around proton jumps compared to time windows of equal size that do not contain proton jumps.
6 Discussion
As is illustrated on the NbSiAs surface – COH adsorbate benchmark dataset and in the appendix, the utilized transfer learning prior can improve the data efficiency substantially while enabling uncertainty-based detection of atomic structures that have an increased likelihood of a large prediction error. Furthermore, our full uncertainty-based on-the-fly fine-tuning workflow was able to maintain a specified accuracy and bias the training dataset toward rare events by selectively doing DFT reference calculations for samples with a high uncertainty. The workflow resulted in a significant improvement in the accuracy of the simulations over the non-fine-tuned models while only requiring a small amount of training data. The only simulation requiring more than 100 DFT interventions was for modeling the proton diffusion in CaZrS3 to an exceptionally high accuracy, achieving an RMSE of 0.57 meV atom−1 at time of intervention and at a temperature of 1500 kelvin. This increase in accuracy was particularly reflected in a much better modeling of the underlying physical processes and phenomena, such as the phase behaviour of LaMnO3 due to subtle effects in the electronic structure at different temperatures and the transport mechanism of protons in CaZrS3.Overall, the high data efficiency is primarily achieved via the transfer learning approach and not the uncertainty-guided on-the-fly learning, whose primary function is automation, ease of use and providing an additional layer of trustworthiness by maintaining high model confidence during the simulation. In particular, on-the-fly learning requires high intervention frequencies during the start of the simulation to maintain accuracy and to align the foundational neural network prior to the target system's specific potential energy surface. This might be suboptimal in terms of data efficiency since the initial structures will be correlated. Strategies that initially allow for a higher error threshold during a training run could be employed to reduce the initial sampling frequency, while risking that some of the structures sampled this way might retroactively turn out not to be relevant at the simulation temperature. This might, in practice, improve data efficiency further. However, error thresholds that are too large have the potential to lead to instabilities in the simulation since a certain level of catastrophic forgetting can't be ruled out even when using the transfer learning prior to counteract this effect. Due to the additional complications in terms of automation and ease of use that other active learning strategies can have, we chose to integrate the uncertainty-aware transfer learning approach into an on-the-fly learning workflow. However, due to its generality, the transfer learning approach introduced here can, in principle, be integrated into any uncertainty-guided active learning method.
Due to the simplicity of the uncertainty-aware transfer learning approach, it should further be straightforward to apply this approach to other neural network models not included in this work, since it worked out of the box for all neural network models investigated here after fitting only two hyperparameters for each model architecture individually, the strength of the BNN prior and the step size of the Markov chain. However, during our experiments, the same hyperparameters for each architecture gave good results across different simulations or datasets. This indicates that strong baseline values for those hyperparameters can be determined and given as default values to end-users. For large domain shifts, this value and the number of training steps at each update may need to be increased for the model to sufficiently adapt to the target simulation. However, post-hoc analysis indicates that the main driver behind variation in the scale of σTL across models seems to be largely driven by the scale of the weights of the respective foundation models, with the EquiformerV2 having substantially smaller weights on average, also needing a much smaller σTL. Therefore, even for large domain shifts, good values for σTL should still be of the same order of magnitude as the values determined here. Lastly, if the simulation is split into fine-tuning and production simulations, for example, to avoid violating energy conservation due to model updates, the same simulation-hyperparameters (i.e., temperature or pressure) can be used for both simulation types, which keeps the selection of those parameters for training runs simple.
While reliance on Bayesian neural network methods represents a likely source of methodological unfamiliarity for end-users compared with more classical optimization methods for neural networks, the effects that these hyperparameters have on the optimization process are conceptually intuitive to understand, with the strength of the prior controlling how far the fine-tuned model might deviate from the pretrained model in its predictions and the step size having an analogous role to step-sizes in regular optimizers. Combined with the existence of strong baseline values, the Bayesian nature should therefore not represent a high barrier of entry for computational material scientists and chemists.
Because we use the uncertainty in the energy predictions to decide if a DFT call should be done or not, the proposed fine-tuning workflow is geared towards applications where the accuracy in predicted potential energies is of primary interest, even though the DFT-computed forces and stresses are included as training labels during model optimization. An alternative approach would have been to use the uncertainty in the force predictions. We used the energy predictions because accuracy in the energies controls the accuracy of the thermodynamic ensembles, and accurate forces do not necessarily guarantee accurate potential energies since even small errors in the forces can add up to large errors in the potential energy over large changes in the geometry. However, an advantage of using the force uncertainties would be that forces are localized. This makes it possible to choose individual error thresholds for each atom, where some atoms or regions might be of particular interest, such as in the case of proton diffusion. For this reason, we plan to implement the option to include the force uncertainties as a decision criterion for DFT interventions in the future. For some physical properties that arise from higher-order derivatives of the potential energy surface, such as vibration frequencies which depend on the Hessian, it is challenging to ensure accuracy with any training methods, even on the training data itself, since the objective function used in the training of neural network potentials usually only involves the potential energy and its first-order derivatives. The main reason for this is that the first-order derivatives can be computed without significantly increasing the total computational complexity of the electronic structure calculation by leveraging the Hellmann–Feynman Theorem,66,67 while higher-order derivatives would be much more computationally demanding to compute. Therefore, it is currently unclear how accurately such properties are modeled within this workflow.
Regarding the speed up our fine-tuning approach can achieve for simulations, the exact results will be dependent on the chosen error threshold, hardware availability and the DFT software used. We give some illustrative examples and estimates for different hardware configurations for the CaZrS3 – proton system in Appendix D. It only took 2000 training steps at a batch size of 5 to update each Monte Carlo sample on a new DFT-labeled geometry. Additionally, updating the model is highly parallelizable because the Markov chain of each Monte Carlo sample can be run independently. Nonetheless, on limited hardware, it might be better to only sample 4 instead of 8 Monte Carlo samples. In previous works, we observed a decrease in log-likelihoods by a value comparable to the decrease observed by reducing the number of Monte Carlo samples from 16 to 836. However, it is not advisable to go below 4 Monte Carlo samples as the quality of uncertainty quantification starts to degrade more significantly, with a decrease of almost twice the value of going from 8 to 4 samples when reducing it further from 4 to 2. In cases of very constrained GPU resources, using more computationally efficient variational inference-based BNN methods instead of MCMC methods might therefore be a better option.
Overall, the quality of uncertainty quantification was sufficient to keep the model in the specified accuracy range, with the fraction of structures that at DFT intervention exceeding the error threshold consistently staying below the 5 percent target. Further, the uncertainty measure was able to bias the training set towards rare events for both the ethanol and the proton diffusion example. However, there were still instances where the error was large despite a small predicted standard deviation, which leaves room for improvement.
One possible cause for this might be that we sample the different sets of weights from a small region around the weights of the pre-trained model, because of the chosen prior over the weights. In this work, we use a Gaussian prior as a proof of concept to show that by harnessing the prior of a BNN for transfer learning, enhanced data efficiency and useful predictive uncertainty estimates can be achieved. However, there is likely room for improvement in the design of the prior by experimenting with different types of parametric distributions. Further, recently, an approach to sampling Bayesian neural network weights has been introduced, where the prior can be specified on the function space instead of the weight space.68 This is an attractive alternative because it would avoid biasing the weights of the model to stay close to the original ones and instead would only bias the weights to result in predictions that lie within a certain range of the original model. Attempting to adapt this formalism to construct a more sophisticated prior from the function space, based on the belief that the neural network might need to change its predictions by a certain amount to be accurate for the system of interest, might therefore be a promising approach to improve the quality of uncertainty quantification.
Other factors can additionally make uncertainty quantification challenging in an on-the-fly learning setting. There can be discontinuities in the electronic structure of molecules or materials, for example, when changes in the atomic geometry push previously occupied electronic states through the Fermi level. This can cause discontinuities in the curvature of the potential energy surface, which makes uncertainty quantification challenging because current machine learning models are unaware of the electronic structure at this detail. Because of this, the first occurrences of geometries with novel electronic structures in an on-the-fly learning scenario will likely always be predicted with some degree of overconfidence.
We currently make the assumption in our calibration of the uncertainty that the miscalibration of the uncalibrated model uncertainties remains approximately constant over the course of the simulation. This assumption can, of course, be violated, for example, during phase transitions, and hence it might be beneficial to adjust the calibration procedure for such cases. One approach could be to not calculate Mn from an average but instead from an exponential moving average that will weigh more recent examples more strongly compared to older ones. Another interesting future research direction is extending this fine-tuning approach from fine-tuning for specific systems to entire subclasses of systems. Currently, a model that is fine-tuned to a specific system with the proposed workflow here will likely diminish in accuracy for almost any other system. Further, the uncertainty is calibrated to a different system and will likely also not always be transferable to other systems. However, by automatically generating training data from system-specific fine-tuning runs for many individual systems from this subclass and then updating the foundation model based on the merged training data, it might be possible to fine-tune foundation models for entire subclasses of structures in an automated workflow.
7 Conclusion
In this work, we introduced an on-the-fly fine-tuning workflow for foundational neural network potentials that is capable of maintaining a user-specified accuracy by doing DFT reference calculations for high-uncertainty structures. As is illustrated in the paper and the appendix, the utilized transfer learning prior can improve the data efficiency substantially. Furthermore, the integration into our uncertainty-based on-the-fly fine-tuning workflow has demonstrated its ability for biasing the fine-tuning dataset towards rare events and maintaining a user-specified accuracy while making the fine-tuning process easy to use for end-users, especially those without an extensive technical background in neural network potentials. Moreover, it enables an automated fine-tuning of foundation models within high-throughput materials discovery workflows. Given the broad applicability of the uncertainty-aware transfer learning approach across neural network architectures and the strong performance of the on-the-fly learning workflow, we believe this method has the potential to become a standard for fine-tuning foundational neural network potentials in machine learning-accelerated molecular dynamics simulations for materials science.Author contributions
T. R., O. N. and D. K. conceptualized the project. T. R. and D. K. developed the experimental design. T. R. implemented the code and performed the evaluation. T. R. wrote the original draft and D. K. and O. N. helped with reviewing and editing.Conflicts of interest
There are no conflicts of interest to declare.Data availability
All code and data for reproducing the experiments in this paper are available.The code for running the on-the-fly learning simulations, including tutorials, is available at:
https://github.com/TimRensmeyer/OTFFineTune and has been archived on Zenodo (DOI: https://doi.org/10.5281/zenodo.18772098)
The ablation experiments for the transfer learning approach in isolation were done on public benchmark datasets. At the time of writing (25.02.2026), the datasets are available at the links below:
The stachyose dataset69 is available at:
https://www.quantum-machine.org/gdml/repo/datasets/md22_stachyose.npz.
The RMD17 dataset70 is available at:
https://figshare.com/articles/dataset/Revised_MD17_dataset_rMD17_/12672038.
The coupled cluster-level ethanol dataset71 is available at:
https://www.quantum-machine.org/gdml/data/npz/ethanol_ccsd_t.zip.
The OC20-Dense dataset53 is available at:
https://github.com/Open-Catalyst-Project/AdsorbML.
Only the NbSiAs surface – COH adsorbate data from this dataset was used in the ablation experiment.
Source code and in-depth illustrations on how the neural networks were trained and evaluated in those ablation experiments are available in the code appendix.
Supplementary information (SI): source code to perform the ablation experiments from the paper. See DOI: https://doi.org/10.1039/d5dd00392j.
Appendices
Appendix: A Additional transfer learning results
This section provides additional ablation experiments for the transfer learning approach to provide further evidence for the enhanced data efficiency and good quality of uncertainty quantification of this method beyond the NbSiAs benchmark. The experiments here go significantly beyond the evidence provided in the main manuscript by demonstrating these properties under different circumstances (quality of force predictions, transfer learning between different electronic structure methods). Thus, they additionally demonstrate the robustness of the proposed method. However, for brevity, they were omitted in the main manuscript.The second benchmark is a transfer learning scenario from DFTB level accuracy to DFT level accuracy. In particular, we generate a large dataset of different configurations of a stachyose molecule in DFTB for pre-training and then utilize the stachyose data from the MD22 dataset for the transfer learning task.
The third test scenario is a transfer learning task for reaching CC level accuracy on an ethanol molecule starting from a model pre-trained on the corresponding ethanol data from the MD17 dataset. The CC-level dataset used for this was introduced by Bogojeski et al.71
All experiments were done with 8 Monte Carlo samples generated from the same Markov chain, which has been identified as a good tradeoff between computational complexity and quality of uncertainty quantification in our previous work.74 Details of all the datasets can be found in Appendix C. We set σTL as 0.2 for all experiments.
Appendix: A.1.1 The evaluation metrics. On all tasks, we evaluate the model's overall accuracy in terms of the Root Mean Square Error (RMSE) of the force components in dependence on the size of the training dataset. We analyze the transfer learning models' accuracy and quality of uncertainty quantification in comparison to a model with a Gaussian mean field prior p(θ) ∼ N(0, I). For the evaluation of the uncertainties, we compare the Mean Log Likelihoods (MLLs) of the force components or energies as a function of the RMSE for both models. To smooth each predicted distribution of the 8 Monte Carlo samples on this metric, we fit a normal distribution to the means and variances of each predicted distribution and use these smoothed distributions instead. Further, since the main goal of the uncertainty measure is the identification of configurations with a large error in the prediction, we evaluate the models in the task of detecting force components with a large prediction error based on the predicted uncertainty. More specifically, we analyze the corresponding AUC-ROC scores for detecting large errors via the predicted variance and plotting them as a function of the RMSE. On the ethanol and paracetamol datasets, errors of more than 1 kcal mol−1 Å−1 were considered large, while on the more difficult stachyose dataset, the cutoff was set as 3 kcal mol−1 Å−1 because an error of 1 kcal mol−1 Å−1 could not be considered an outlier. Since ethanol is a very small molecule, it was computationally feasible to include a deep ensemble containing 8 models trained from scratch as a second baseline (see Appendix B.6 for the training details).
On the stachyose dataset, again, a clear improvement in accuracy at equal amounts of training samples is visible when compared to the baseline model (Fig. 7). However, both models have higher RMSEs than their counterparts on the paracetamol dataset at equal amounts of training configurations. The MLLs of the transfer learning model appear to be slightly lower than for a model trained from scratch when controlled for accuracy. The same is true only to a much smaller degree for the AUC-ROC scores. Further analysis revealed that the validation set was too small for the large configuration space of stachyose to properly recalibrate the uncertainties, which led to an overestimation of the errors on the test set for the transfer learned models but not for the baseline models. This also explains the absence of such reduced performance on the AUC-ROC scores, which are invariant under recalibration of uncertainties. Accounting for the slightly wrong calibration by recalibrating the uncertainties on the test set instead of the validation set confirmed miscalibration as the main source of the gap in MLLs. In particular, the gap between the MLLs of the transfer learning models that are closest in RMSE reduced from 0.191 to 0.086. Notably, we also attempted a transfer learning scenario on the stachyose dataset, where the model was only pre-trained on the small molecules of the MD17 dataset. However, this did not lead to any improvement in data efficiency. The most likely explanation for this is that the local atomic environments on the small molecule datasets, which the neural network uses to calculate potential energy contributions, are qualitatively very different from those of the stachyose molecule.
The biggest improvement in accuracy, when compared to the baseline model, was found on the ethanol dataset (Fig. 8), with an RMSE of less than 0.5 kcal mol−1 Å−1 with only 10 configurations. There appears to be no decrease in the quality of uncertainty quantification, both in terms of MLLs as well as AUC-ROCs on this benchmark, when compared to the baseline model. Comparing both the baseline model and the transfer learning model to the deep ensemble, almost identical performance can be seen on the outlier detection task, while the deep ensemble has slightly higher MLLs.
Additional analysis was performed by breaking down the MLLs into contributions from force components, whose prediction error falls into a certain interval, e.g. the contribution to the MLLs from samples where the prediction error is in the interval [0.1, 0.2) is given by the sum over the log-likelihoods of all force components in the test set where the prediction error is in the interval [0.1, 0.2) divided by the total number of force components in the test set. The results shown in Fig. 9 demonstrate that the total MLL score is dominated by samples whose prediction error is small. This offers an explanation for the slightly better MLLs without a similar performance gap on the outlier detection task on this benchmark: the deep ensembles achieve slightly better uncertainty quantification on configurations with a small prediction error but not on configurations with a large prediction error.
Notably, for all force transfer learning scenarios, the error of the pre-trained model was quite large with mean absolute errors of 2.31 kcal mol−1 Å−1 on the paracetamol validation set, 4.34 kcal mol−1 Å−1 on the stachyose validation set and 5.12 kcal mol−1 Å−1 on the ethanol validation set. Further, all pre-trained models achieved a validation loss smaller than 0.15 kcal mol−1 Å−1 on their pre-training datasets, strongly indicating that DFTB, DFT and CC methods disagree quite substantially in their force predictions for a given configuration. However, as was already alluded to in the introduction, this can potentially be traced back to simple biases in the simulation methods, such as slightly different equilibrium bond lengths. These small biases in different simulation methods can lead to qualitatively very similar force fields that may disagree substantially on the forces of a given configuration. This would also explain why transfer learning is very efficient in these cases, as the model mostly has to correct for those biases, such as equilibrium bond lengths. Importantly, those force fields will lead to similar predictions of physical and chemical properties despite their apparently large disagreement, while a machine-learned force field with a similar magnitude of error to one of those methods cannot, in general, be expected to yield those properties as well and hence needs to be trained to a much higher accuracy.
One additional result that stands out is the relatively high RMSE of both the transfer learning and the baseline model on the stachyose dataset when compared to the other two test scenarios. However, two factors make this dataset particularly challenging. First of all, stachyose is a larger molecule than paracetamol and ethanol, which, in addition, contains many single sigma bonds that allow for rotational degrees of freedom along the bond axis. This results in a very large configuration space for stachyose molecules, even relative to their size. The second factor that makes this benchmark more challenging for the transfer learning model is that, unlike in the ethanol case, the higher accuracy dataset was not composed of configurations generated from an MD trajectory of the lower accuracy method but instead from a trajectory at DFT-level accuracy. As a result, the distribution of configurations in the DFT dataset will be different from the one from the DFTB dataset.
Lastly, one important observation we made is that the transfer learning approach converges much faster than when training from scratch. While state-of-the-art models can take days to train from scratch, training and validation losses converged within minutes on the transfer learning tasks. The only reason we let the sampling algorithm run for as long as described in Appendix B is to make sure that no pathological overfitting takes place.
Appendix: B Methodological details
This section contains the methodological details that were omitted from the main manuscript to avoid breaking the flow of reading. . NequIP and MACE then calculate the forces acting on the atoms as the negative gradients of the potential energy
. NequIP and MACE then calculate the forces acting on the atoms as the negative gradients of the potential energy  via automatic differentiation libraries, while the Equiformerv2 model calculates the forces directly from a set of equivariant atomic feature vectors. To apply the Bayesian neural network framework to these models, the architectures were modified slightly by adding layers that compute standard deviations σ1, …, σn for the forces from the invariant features {v1, …, vn}. Further, an energy standard deviation σÊ is introduced and the distribution over the energy and forces is modeled as
via automatic differentiation libraries, while the Equiformerv2 model calculates the forces directly from a set of equivariant atomic feature vectors. To apply the Bayesian neural network framework to these models, the architectures were modified slightly by adding layers that compute standard deviations σ1, …, σn for the forces from the invariant features {v1, …, vn}. Further, an energy standard deviation σÊ is introduced and the distribution over the energy and forces is modeled aswhere N denotes a normal distribution and I is the identity matrix.
For the Equiformev2 and MACE models, the predictions are additionally conditioned on the lattice vectors L1, L2, L3. For the MACE model, the stress tensor is also predicted. For this, the predicted distribution is modified to
The standard deviations of the forces σi are predicted by a three-layer MLP with input dimension 64, latent dimensions 32 and 16 and output dimension 1.
SiLU activation functions are used for the latent layers and the output activation function is the exponential function.
A base with 5 interaction blocks was used.
Because there aren't a lot of labeled examples for the potential energies, a fixed energy standard deviation was used during the on-the-fly experiments, which was set to 10 percent of the target accuracy.
For the experiments in Appendix A, no energies were included in the training and train only the force labels were trained on.
The output of the mace-mp medium model contains feature vectors for each atom. 256 components of each feature vector are rotation invariant. From these features, the standard deviations of the forces σi are predicted by a three-layer MLP with input dimension 256, latent dimensions 64 and 16 and output dimension 1.
Again use a fixed energy standard deviation set to 10 percent of the target accuracy was used. For all experiments, a fixed stress standard deviation of 0.1/16 kcal mol−1 Å−3 was used.
Appendix: B.5.1 Sampling for the transfer learning experiments. For the ethanol and paracetamol test cases, the step size γ is exponentially decreased from 10−2 and 0.3·10−2 to 10−5 during the first 106 steps for the baseline model and transfer learning model, respectively. At the end of this phase, the first model is sampled. Afterward, a cyclical learning rate schedule is used:
with γ0 = 0.001 and cycle length K = 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 to sample the subsequent models from the same Markov chain at the end of each cycle.
000 to sample the subsequent models from the same Markov chain at the end of each cycle.
The same procedure is also utilized for the baseline model on the stachyose test case. However, the initial convergence phase is shortened to 0.5·106 steps for the transfer learning model, as the other two test cases had revealed a quicker convergence for the transfer learning models. For the paracetamol and ethanol cases, a batch size of 30 is used and for the stachyose case, it is set as 15. Analogous to the on-the-fly learning experiments, an energy offset was calculated for the training, validation and test data, of the equiformer model so that for the first sample in the training data, the energy equals the one predicted by the pre-trained model. For the surface-adsorbate transfer learning task, a batch size of 20 was used for the baseline model and for the transfer learning task, it is set as 15. Furthermore, the same sampling procedure is employed. For the baseline and transfer learning model, the step size γ is exponentially decreased from 10−4 and·10−5 to 10−7 and 10−8 respectively during the initial convergence phase. This phase was 0.5·106 steps long for the baseline model and 105 steps long for the transfer learning model, respectively. Then again, a cyclical sampling procedure is employed to generate the other samples with γ0 = 0.0001 and cycle length K = 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 After the first 90 percent of the initial convergence phase, the mass term is kept constant to ensure close convergence to the posterior.
000 After the first 90 percent of the initial convergence phase, the mass term is kept constant to ensure close convergence to the posterior.
Appendix: B.5.2 Sampling for the on-the-fly finetuning experiments. To sample the posterior for the on-the-fly fine-tuning experiments, 8 separate Markov Chains were run, one for each Monte Carlo sample. After each added training sample, the SGHMC algorithm with the adaptive mass term is run for 2000 steps to update the Monte Carlo samples. A batch size of 5 and a learning rate schedule
was used. For the MACE model, γ0 = 0.001 was used and for the NequIP model, it was set to γ0 = 0.00003. To improve the speed of convergence, priority sampling was used to increase the likelihood of sampling the newly added training sample at each iteration. The sampling probability was increased so that, on average, the newly added sample is contained once in each minibatch. Each sample in the minibatch estimator of the log-likelihood was weighted by the likelihood ratio of a uniform sampling procedure and the sampling probabilities used to ensure the minibatch estimator is unbiased.
Appendix: B.6.1 The ethanol on-the-fly simulation. A Langevin thermostat with 0.5 fs time steps and a friction term of 0.01 fs−1 of the Atomic Simulation Environment (ASE) library was used to drive the dynamics. The DFT calculations were done with the Vienna Ab initio Simulation Package (VASP) using a cubic 30 Å simulation cell. A plane wave energy cutoff of 800 eV with a convergence criterion of 1e–6 eV and the B3LYP exchange-correlation functional76–79 was used.
Appendix: B.6.2 The LaMnO3 on-the-fly simulation. The NPT thermostat of the Atomic Simulation Environment (ASE) library with a 0.5 fs time step, a time of 100 fs, a p-factor of 160 GPa·0.1·752 fs2 and an external pressure of 1 bar was used to drive the dynamics. A 2 × 2 × 2 supercell containing 40 atoms in total was simulated. The DFT calculations were done with the Vienna Ab initio Simulation Package (VASP). A plane wave energy cutoff of 500 eV with a convergence criterion of 1e–4 eV and the PBE exchange–correlation functional80 and a 4 × 4 × 4 gamma centered Monkhorst–pack grid was used. A Hubbard term of 3.9 was used for the Mn atoms.
Appendix: B.6.3 The proton diffusion on-the-fly simulations. The Langevin thermostat with a friction term of 0.5 of the Atomic Simulation Environment (ASE) library was used to drive the dynamics for 1 ps to set the temperature. Afterwards, The velocity Verlet integrator was used to continue the simulation for another 30 ps. One initial training run was done for each of the two fine-tuned models. Afterwards, 5 production runs were done to investigate the diffusivity of the protons. 1.5 fs time steps were used for all simulations. A CaZrS3 supercell containing 160 atoms and two additional protons was simulated. The DFT calculations were done with the Vienna Ab initio Simulation Package (VASP). A plane wave energy cutoff of 510 eV with a convergence criterion of 1e–4 eV and the PBE exchange–correlation functional80 and a 2 × 2 × 2 gamma centered Monkhorst–pack grid was used.
Appendix: B.7.1 Pretraining for the transfer learning experiments. To pre-train a model, it was converged to a local maximum of the log-posterior on the pre-training dataset with a Gaussian mean field prior p(θ)–N(0, I). Almost the same sampling algorithm and hyperparameters were used as in the sampling of the posterior of the corresponding baseline model. The only differences are that the injected noise is downscaled by a factor of 0.1 and only the first model is sampled. The injected noise was not set to zero, because we found that a small amount of injected noise actually speeds up convergence, especially at the beginning of the optimization.
Appendix: B.7.2 Pretraining the NequIP model on the spice dataset for the ethanol on-the-fly experiment. The NequIP model was trained at a batch size of 25 at a fixed learning rate of 3 × 10−5 with the Adam optimizer. The loss function L = (1/200)MSE(Energy) + MSE(Forces) was used, where the Mean Square Error (MSE) refers to the batch mean. We keep a validation set of 70 structures from the SPICE dataset, which are not included in the training dataset. Every 20
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 training steps, the energy MSE on the validation set were evaluated. The training was stopped after the energy validation loss hadn't improved for 3 epochs. The final model used is the one with the lowest validation loss during the training run.
000 training steps, the energy MSE on the validation set were evaluated. The training was stopped after the energy validation loss hadn't improved for 3 epochs. The final model used is the one with the lowest validation loss during the training run.
where j enumerates the predicted standard deviations of the individual Monte Carlo samples/ensemble models,
![[F with combining circumflex]](https://www.rsc.org/images/entities/i_char_0046_0302.gif) i,j is the predicted expectation value for the i-th force component of that particular model and
i,j is the predicted expectation value for the i-th force component of that particular model and  the corresponding standard deviation. The variance is calculated over the Monte Carlo samples/ensemble models. The mean of the predicted distribution was simply calculated as
the corresponding standard deviation. The variance is calculated over the Monte Carlo samples/ensemble models. The mean of the predicted distribution was simply calculated as 
The calculation of the final energy and stress distribution was done completely analogously.
where
 . By integrating this density from e* = −K to e* = K, the result:
. By integrating this density from e* = −K to e* = K, the result:can be derived. Further, with the identities
 and
and  for x → ∞ it can be verified, that for n → ∞ the predicted error distribution becomes
for x → ∞ it can be verified, that for n → ∞ the predicted error distribution becomesAppendix: C The datasets
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 configurations generated from a molecular dynamics trajectory calculated at DFT level accuracy. The training and test datasets of ethanol at CCSD(T) level accuracy introduced by Bogojeski et al.71 were used for the transfer learning task. The last 10 configurations of the training set were used as validation data. The actual training data consisted of the first
000 configurations generated from a molecular dynamics trajectory calculated at DFT level accuracy. The training and test datasets of ethanol at CCSD(T) level accuracy introduced by Bogojeski et al.71 were used for the transfer learning task. The last 10 configurations of the training set were used as validation data. The actual training data consisted of the first  configurations of the training dataset for varying values of m.
configurations of the training dataset for varying values of m.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 configurations from each MD17 dataset and all configurations from the MD22 datasets were used to form a pool of configurations from which 100
000 configurations from each MD17 dataset and all configurations from the MD22 datasets were used to form a pool of configurations from which 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 are randomly drawn as the pretraining dataset.
000 are randomly drawn as the pretraining dataset.
For the actual training set  , configurations are randomly sampled from the MD17 paracetamol dataset for varying values of m. 10 additional configurations are randomly sampled as a validation set. The rest of the 106
, configurations are randomly sampled from the MD17 paracetamol dataset for varying values of m. 10 additional configurations are randomly sampled as a validation set. The rest of the 106![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 490 configurations are used as a test set.
490 configurations are used as a test set.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 configurations. For both the geometry optimization as well as the MD simulation, a Hamiltonian with self-consistent charges83 and third-order corrections84 was used in correspondence with the 3ob-3-1 Slater Coster files.85 For all atoms, s- and p-orbitals were used in the Hamiltonian.
000 configurations. For both the geometry optimization as well as the MD simulation, a Hamiltonian with self-consistent charges83 and third-order corrections84 was used in correspondence with the 3ob-3-1 Slater Coster files.85 For all atoms, s- and p-orbitals were used in the Hamiltonian.
For the actual training set  , configurations are randomly sampled from the first 10
, configurations are randomly sampled from the first 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 configurations of the MD22 stachyose dataset for varying values of m. 10 additional configurations are randomly sampled from the configurations 10
000 configurations of the MD22 stachyose dataset for varying values of m. 10 additional configurations are randomly sampled from the configurations 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 to 10
100 to 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 900 as a validation set. Configurations 11
900 as a validation set. Configurations 11![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 up to 27
000 up to 27![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 are used as a test set.
000 are used as a test set.
8000 configurations were randomly sampled as possible training configurations. For a training dataset of size  , the first n configurations from that subset were used as a training set. Further, twenty randomly sampled configurations were used as a validation set to recalibrate the uncertainties. The rest of the configurations were used as the test set.
, the first n configurations from that subset were used as a training set. Further, twenty randomly sampled configurations were used as a validation set to recalibrate the uncertainties. The rest of the configurations were used as the test set.
Appendix: D Runtime estimates on different hardware configurations for the CaZrS3 – proton system
A single 30 ps proton diffusion simulation in CaZrS3 would take around 3 months on the two 36-core Intel Platinum 8360Y processors that were used to do the DFT interventions in VASP 5.4.4, while it took less than a day to finetune the 15 kcal mol−1 threshold model and around a week for the 5 kcal mol−1 threshold model during the initial 30 ps training runs. The time for the 5 production runs with each model was negligible. Increasing the CPU resources to eight processors will reduce the time of the simulation in VASP 5.4.4 to around one month and also reduce the time for the on-the-fly simulations to less than half of the previous values, since the DFT interventions were the computational bottleneck. Increasing CPU resources even more will start to result in diminishing returns, as the MPI communications overhead will start to become the limiting factor.GPU nodes containing 8 L40S GPUs were used for the experiments and updating all Monte Carlo samples on a new training data point takes only a few minutes on that hardware. To test the impact of constrained GPU resources, the experiment for the largest system, the 162-atom CaZrS3-proton system, was rerun using only two A100 GPUs. On that hardware, it takes around 20 minutes to update the model. However, it should be noted that DFT calculations for systems of that size will also be quite time-intensive and on the two Intel Platinum 8360Y processors that were used, they took around 24–25 minutes.
DFT calculations involving two 36-core Intel Platinum 8360Y processors were conducted on a single compute node with two CPU sockets (four NUMA domains, 36 logical cores and 64 GB RAM per domain). For calculations with eight 36-core Intel Platinum 8360Y processors, VASP was run on four compute nodes. VASP was run using the full nodes, with MPI ranks and OpenMP threads distributed across CPUs. No explicit NUMA binding or memory placement was applied. Memory allocation followed the system's default NUMA policy. VASP was compiled with Intel compilers (icx, icpx, ifx) and Intel MPI (mpiicx, mpiicpx, mpiifx). Linear algebra routines used the Intel MKL library. The VASP binary used was vasp_std (CPU variant). No custom compilation options beyond the default module build were applied.
Neural network optimization was performed with torch 2.6 and mace-torch 0.3.0 using CUDA 12.4.1 without cuEquivariance acceleration.
Acknowledgements
This research as part of the project CouplteIT! is funded by dtec.bw – Digitalization and Technology Research Center of the Bundeswehr which we gratefully acknowledge. dtec.bw is funded by the European Union – NextGenerationEU. Computational resources (HPC cluster HSUper) have been provided by the project hpc.bw, funded by dtec.bw.Notes and references
- P. W. Atkins and R. S. Friedman, Molecular Quantum Mechanics, OUP, Oxford, 2011, https://books.google.de/books?id=9k-cAQAAQBAJ Search PubMed.
- M. Gastegger and P. Marquetand, Molecular dynamics with neural network potentials, In Machine Learning Meets Quantum Physics, Springer, pp. 233–252, 2020 Search PubMed.
- K. Schütt, S. Chmiela, A. von Lilienfeld, A. Tkatchenko, K. Tsuda, and K.-R. Müller, Machine Learning Meets Quantum Physics, 2020, DOI:10.1007/978-3-030-40245-7.
- F. Giustino, Materials Modelling Using Density Functional Theory: Properties and Predictions, Oxford University Press, 2014, https://books.google.de/books?id=FzOTAwAAQBAJ Search PubMed.
- E. Kocer, T. W. Ko and J. Behler, Neural network potentials: A concise overview of methods, Annu. Rev. Phys. Chem., 2022, 73(1), 163–186, DOI:10.1146/annurev-physchem-082720-034254.
- J. Klicpera, F. Becker, and S. G. Gemnet, Universal directional graph neural networks for molecules, In Advances in Neural Information Processing Systems, 2021, https://openreview.net/forum?id=HS_sOaxS9K- Search PubMed.
- O. T. Unke, S. Chmiela, M. Gastegger, K. T. Schütt, H. E. Sauceda and K.-R. M. Spookynet, Learning force fields with electronic degrees of freedom and nonlocal effects, Nat. Commun., 2021, 12, 7273, DOI:10.1038/s41467-021-27504-0.
- K. T. Schütt, O. T. Unke, and M. Gastegger, Equivariant message passing for the prediction of tensorial properties and molecular spectra, In International Conference on Machine Learning, 2021 Search PubMed.
- S. Batzner, A. Musaelian, L. Sun, M. Geiger, J. P. Mailoa, M. Kornbluth, N. Molinari, T. E. Smidt and B. Kozinsky, E(3)-equivariant graph neural networks for data-efficient and accurate interatomic potentials, Nat. Commun., 2022, 13 DOI:10.1038/s41467-022-29939-5.
- M. Haghighatlari, J. Li, X. Guan, O. Zhang, A. Das, C. J. Stein, F. Heidar-Zadeh, M. Liu, M. Head-Gordon, L. Bertels, H. Hao, I. Leven and T. Head-Gordon, Newtonnet: a Newtonian message passing network for deep learning of interatomic potentials and forces, Digital Discovery, 2022, 1(3), 333–343, 10.1039/d2dd00008c.
- Z. Qiao, A. S. Christensen, M. Welborn, F. R. Manby, A. Anandkumar and T. F. Miller, Informing geometric deep learning with electronic interactions to accelerate quantum chemistry, Proc. Natl. Acad. Sci. U. S. A., 2022, 119(31), e2205221119, DOI:10.1073/pnas.2205221119.
- I. Batatia, D. P. Kovacs, G. Simm, C. Ortner, and G. Csanyi, Mace: Higher order equivariant message passing neural networks for fast and accurate force fields, In Advances in Neural Information Processing Systems, ed. S. Koyejo, S. Mohamed, A. Agarwal, D. Belgrave, K. Cho, and A. Oh, Curran Associates, Inc., 35, pp. 11423–11436, 2022, https://proceedings.neurips.cc/paper_files/paper/2022/file/4a36c3c51af11ed9f34615b81edb5bbc-Paper-Conference.pdf Search PubMed.
- D. . Péter Kovács, I. Batatia, E. S. Arany and G. Csányi, Evaluation of the mace force field architecture: From medicinal chemistry to materials science, J. Chem. Phys., 2023, 159(4), 044118, DOI:10.1063/5.0155322.
- Y.-L. Liao, B. M. Wood, A. Das, and T. Smidt, Equiformerv2: Improved equivariant transformer for scaling to higher-degree representations, In The Twelfth International Conference on Learning Representations, 2024, URL https://openreview.net/forum?id=mCOBKZmrzD Search PubMed.
- B. Deng, P. Zhong, K. J. Jun, J. Riebesell, K. Han, C. J. Bartel and G. Ceder, Chgnet as a pretrained universal neural network potential for charge-informed atomistic modelling, Nat. Mach. Intell., 2023, 5(9), 1031–1041, DOI:10.1038/s42256-023-00716-3.
- A. Musaelian, B. Simon, A. Johansson, L. Sun, C. J. Owen, M. Kornbluth and B. Kozinsky, Learning local equivariant representations for large-scale atomistic dynamics, Nat. Commun., 2023, 14(1), 579, DOI:10.1038/s41467-023-36329-y.
- F. Xiang, B. M. Wood, L. Barroso-Luque, D. S. Levine, M. Gao, M. Dzamba, and C. Lawrence Zitnick, Learning smooth and expressive interatomic potentials for physical property prediction, In Proceedings of the 42nd International Conference on Machine Learning, volume 267 of Proceedings of Machine Learning Research, ed. A. Singh, M. Fazel, D. Hsu, S. Lacoste-Julien, F. Berkenkamp, T. Maharaj, K. Wagstaff, and J. Zhu, PMLR, pp. 17875–17893, 2025, https://proceedings.mlr.press/v267/fu25h.html Search PubMed.
- B. M. Wood, M. Dzamba, F. Xiang, M. Gao, M. Shuaibi, L. Barroso-Luque, K. Abdelmaqsoud, V. Gharakhanyan, J. R. Kitchin, D. S. Levine, K. Michel, A. Sriram, T. Cohen, A. Das, S. J. Sahoo, A. Rizvi, Z. Ward Ulissi, and C. Lawrence Zitnick, UMA: A family of universal models for atoms, In The Thirty-ninth Annual Conference on Neural Information Processing Systems, 2025, https://openreview.net/forum?id=SvopaNxYWt Search PubMed.
- D. Péter Kovács, J. H. Moore, N. J. Browning, I. Batatia, J. T. Horton, Y. Pu, V. Kapil, W. C. Witt, I.-B. Magdău, D. J. Cole and G. Csányi, Mace-off: Short-range transferable machine learning force fields for organic molecules, J. Am. Chem. Soc., 2025, 147(21), 17598–17611, DOI:10.1021/jacs.4c07099.
- H. Kaur, F. D. Pia, I. Batatia, X. R. Advincula, B. X. Shi, J. Lan, G. Csányi, A. Michaelides and V. Kapil, Data-efficient fine-tuning of foundational models for first-principles quality sublimation enthalpies, Faraday Discuss., 2025, 256, 120–138, 10.1039/D4FD00107A.
- A. Kolluru, N. Shoghi, M. Shuaibi, S. Goyal, A. Das, C. Lawrence Zitnick and Z. Ulissi, Transfer learning using attentions across atomic systems with graph neural networks (TAAG), J. Chem. Phys., 2022, 156(18), 184702, DOI:10.1063/5.0088019.
- V. Zaverkin, D. Holzmüller, L. Bonfirraro and J. Kästner, Transfer learning for chemically accurate interatomic neural network potentials, Phys. Chem. Chem. Phys., 2023, 25, 5383–5396, 10.1039/D2CP05793J.
- J. S. Smith, B. Tyler Nebgen, R. I. Zubatyuk, N. Lubbers, C. Devereux, K. Barros, S. Tretiak, O. Isayev and A. E. Roitberg, Approaching coupled cluster accuracy with a general-purpose neural network potential through transfer learning, Nat. Commun., 2019, 10, 2903 CrossRef.
- J. Isak Texas Falk, L. Bonati, P. Novelli, M. Parrinello, and M. pontil, Transfer learning for atomistic simulations using GNNs and kernel mean embeddings, In Thirty-seventh Conference on Neural Information Processing Systems, 2023, https://openreview.net/forum?id=Enzew8XujO Search PubMed.
- A. Liebert, F. Dethof, S. Keßler, and O. Niggemann, Automated impact echo spectrum anomaly detection using u-net autoencoder, In 13th Conference on Prestigious Applications of Intelligent Systems, 2024, DOI:10.3233/FAIA241058.
- A. D. Becke, Density-functional thermochemistry. iii. the role of exact exchange, J. Chem. Phys., 1993, 98(7), 5648–5652, DOI:10.1063/1.464913.
- C. Lee, W. Yang and R. G. Parr, Development of the colle-salvetti correlation-energy formula into a functional of the electron density, Phys. Rev. B: Condens. Matter Mater. Phys., 1988, 37, 785–789, DOI:10.1103/PhysRevB.37.785.
- P. J. Stephens, F. J. Devlin, C. F. Chabalowski and M. J. Frisch, Ab initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields, J. Phys. Chem., 1994, 98(45), 11623–11627, DOI:10.1021/j100096a001.
- S. H. Vosko, L. Wilk and M. Nusair, Accurate spin-dependent electron liquid correlation energies for local spin density calculations: a critical analysis, Can. J. Phys., 1980, 58(8), 1200–1211, DOI:10.1139/p80-159.
- J. Vandermause, S. B. Torrisi, S. L. Batzner, Y. Xie, L. Sun, A. M. Kolpak and B. Kozinsky, On-the-fly active learning of interpretable Bayesian force fields for atomistic rare events, npj Comput. Mater., 2019, 6, 1–11 Search PubMed . URL https://api.semanticscholar.org/CorpusID:208635886.
- E. V. Podryabinkin, E. V. Tikhonov, A. V. Shapeev and A. R. Oganov, Accelerating crystal structure prediction by machine-learning interatomic potentials with active learning, Phys. Rev. B, 2019, 99, 064114, DOI:10.1103/PhysRevB.99.064114.
- E. V. Podryabinkin and A. V. Shapeev, Active learning of linearly parametrized interatomic potentials, Comput. Mater. Sci., 2017, 140, 171–180, DOI:10.1016/j.commatsci.2017.08.031 . URL https://www.sciencedirect.com/science/article/pii/S0927025617304536.
- K. Gubaev, E. V. Podryabinkin, G. L. W. Hart and A. V. Shapeev, Accelerating high-throughput searches for new alloys with active learning of interatomic potentials, Comput. Mater. Sci., 2019, 156, 148–156, DOI:10.1016/j.commatsci.2018.09.031 . URL https://www.sciencedirect.com/science/article/pii/S0927025618306372.
- R. Jinnouchi, K. Miwa, F. Karsai, G. Kresse and R. Asahi, On-the-fly active learning of interatomic potentials for large-scale atomistic simulations, J. Phys. Chem. Lett., 2020, 11, 6946–6955, DOI:10.1021/acs.jpclett.0c01061.
- L. Kahle and F. Zipoli, Quality of uncertainty estimates from neural network potential ensembles, Phys. Rev. E, 2022, 105, 015311, DOI:10.1103/PhysRevE.105.015311.
- T. Rensmeyer, B. Craig, D. Kramer and O. Niggemann, High accuracy uncertainty-aware interatomic force modeling with equivariant bayesian neural networks, Digital Discovery, 2024, 3, 2356–2366, 10.1039/D4DD00183D.
- R. Shwartz-Ziv, M. Goldblum, H. Souri, S. Kapoor, C. Zhu, Y. LeCun, and A. G. Wilson, Pre-train your loss: Easy Bayesian transfer learning with informative prior, In First Workshop on Pre-training: Perspectives, Pitfalls, Paths Forward at ICML, 2022, https://openreview.net/forum?id=ao30zaT3YL Search PubMed.
- C. H. Chen, P. Parashar, C. Akbar, S. M. Fu, M.-Y. Syu and A. Lin, Physics-prior Bayesian neural networks in semiconductor processing, IEEE Access, 2019, 7, 130168–130179, DOI:10.1109/ACCESS.2019.2940130.
- K. T. Schütt, H. E. Sauceda, P.-J. Kindermans, A. Tkatchenko and K.-R. Müller, Schnet – a deep learning architecture for molecules and materials, J. Chem. Phys., 2018, 148(24), 241722, DOI:10.1063/1.5019779.
- S. Chmiela, A. Tkatchenko, H. E. Sauceda, I. Poltavsky, K. T. Schütt and K.-R. Müller, Machine learning of accurate energy-conserving molecular force fields, Sci. Adv., 2017, 3(5), e1603015, DOI:10.1126/sciadv.1603015.
- Y. Kwon, J.-H. Won, B. J. Kim, and M. Cho Paik, Uncertainty quantification using Bayesian neural networks in classification: Application to ischemic stroke lesion segmentation, In Medical Imaging with Deep Learning, 2018, https://openreview.net/forum?id=Sk_P2Q9sG Search PubMed.
- W. J. Maddox, P. Izmailov, T. Garipov, D. P. Vetrov, and A. G. Wilson, A simple baseline for Bayesian uncertainty in deep learning, In Advances in Neural Information Processing Systems, 32, 2019, https://proceedings.neurips.cc/paper_files/paper/2019/file/118921efba23fc329e6560b27861f0c2-Paper.pdf Search PubMed.
- C. Blundell, J. Cornebise, K. Kavukcuoglu, and D. Wierstra, Weight uncertainty in neural network, In International Conference on Machine Learning, 2015 Search PubMed.
- Y. Gal and Z. Ghahramani, Dropout as a Bayesian approximation: Representing model uncertainty in deep learning, In International Conference on Machine Learning, pp. 1050–1059, 2016 Search PubMed.
- M. Welling and Y. Whye Teh, Bayesian learning via stochastic gradient Langevin dynamics, In International Conference on Machine Learning, 2011 Search PubMed.
- T. Chen, E. B. Fox, and C. Guestrin, Stochastic gradient Hamiltonian monte carlo. In International Conference on Machine Learning, pp. 1683–1691, 2014 Search PubMed.
- Y.-A. Ma, T. Chen, and E. Fox, A complete recipe for stochastic gradient mcmc, In Advances in Neural Information Processing Systems, 28, 2015 Search PubMed.
- J. Yao, W. Pan, S. Ghosh, and F. Doshi-Velez, Quality of uncertainty quantification for Bayesian neural network inference, In International Conference on Machine Learning: Workshop on Uncertainty & Robustness in Deep Learning (ICML), 2019 Search PubMed.
- T. Rensmeyer, W. Großmann, D. Kramer, and O. Niggemann, Bayesian transfer learning of neural network-based interatomic force models, In 38th Annual AAAI Conference on Artificial Intelligence | Workshop on AI to Accelerate Science and Engineering, 2023, https://ai-2-ase.github.io/papers/4 Search PubMed.
- J. A. Bilbrey, J. S. Firoz, M.-S. Lee and S. Choudhury, Uncertainty quantification for neural network potential foundation models, npj Comput. Mater., 11(1), 4–2025, DOI:10.1038/s41524-025-01572-y.
- Z. Li, J. R. Kermode and A. De Vita, Molecular dynamics with on-the-fly machine learning of quantum-mechanical forces, Phys. Rev. Lett., 2015, 114, 096405, DOI:10.1103/PhysRevLett.114.096405.
- L. Chanussot, A. Das, S. Goyal, T. Lavril, M. Shuaibi, M. Riviere, K. Tran, J. Heras-Domingo, C. Ho, W. Hu, A. Palizhati, A. Sriram, B. Wood, J. Yoon, P. Devi, C. Lawrence Zitnick and Z. Ulissi, Open Catalyst 2020 (OC20) Dataset and Community Challenges, ACS Catal., 2021, 11(10), 6059–6072, DOI:10.1021/acscatal.0c04525.
- J. Lan, A. Palizhati, M. Shuaibi, B. M. Wood, B. Wander, A. Das, M. Uyttendaele, C. Lawrence Zitnick, and Z. W. Ulissi, Adsorbml: Accelerating adsorption energy calculations with machine learning, 2022, preprint arXiv:2211.16486 Search PubMed.
- M. Elstner, D. Porezag, G. Jungnickel, J. Elsner, M. Haugk, T. Frauenheim, S. Suhai and G. Seifert, Self-consistent-charge density-functional tight-binding method for simulations of complex materials properties, Phys. Rev. B: Condens. Matter Mater. Phys., 1998, 58, 7260–7268, DOI:10.1103/PhysRevB.58.7260.
- K. Raghavachari, G. W. Trucks, J. A. Pople and M. Head-Gordon, A fifth-order perturbation comparison of electron correlation theories, Chem. Phys. Lett., 1989, 157(6), 479–483, DOI:10.1016/S0009-2614(89)87395-6 . URL https://www.sciencedirect.com/science/article/pii/S0009261489873956.
- P. Eastman, P. Kumar Behara, D. L. Dotson, R. Galvelis, J. E. Herr, J. T. Horton, Y. Mao, J. D. Chodera, B. P. Pritchard, Y. Wang, G. De Fabritiis and T. E. Markland, Spice, a dataset of drug-like molecules and peptides for training machine learning potentials, Sci. Data, 2023, 10(1), 11, DOI:10.1038/s41597-022-01882-6.
- N. Mardirossian and M. Head-Gordon, ωb97m-v: A combinatorially optimized, range-separated hybrid, meta-gga density functional with vv10 nonlocal correlation, J. Chem. Phys., 2016, 144(21), 214110, DOI:10.1063/1.4952647.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, A consistent and accurate ab initio parametrization of density functional dispersion correction (dft-d) for the 94 elements h-pu, J. Chem. Phys., 132(15), 154104, DOI:10.1063/1.3382344.
- S. Grimme, S. Ehrlich and L. Goerigk, Effect of the damping function in dispersion corrected density functional theory, J. Comput. Chem., 2011, 32(7), 1456–1465, DOI:10.1002/jcc.21759.
- A. Jain, S. P. Ong, G. Hautier, W. Chen, W. D. Richards, S. Dacek, S. Cholia, D. Gunter, D. Skinner, G. Ceder and K. A. Persson, Commentary: The materials project: A materials genome approach to accelerating materials innovation, APL Mater., 2013, 1(1), 011002, DOI:10.1063/1.4812323.
- J. Rodríguez-Carvajal, M. Hennion, F. Moussa, A. H. Moudden, L. Pinsard and A. Revcolevschi, Neutron-diffraction study of the jahn-teller transition in stoichiometric lamno3, Phys. Rev. B: Condens. Matter Mater. Phys., 1998, 57, R3189–R3192, DOI:10.1103/PhysRevB.57.R3189.
- J. P. Perdew, K. Burke and M. Ernzerhof, Generalized gradient approximation made simple, Phys. Rev. Lett., 1996, 77, 3865–3868, DOI:10.1103/PhysRevLett.77.3865.
- V. I. Anisimov, J. Zaanen and O. K. Andersen, Band theory and mott insulators: Hubbard u instead of stoner i, Phys. Rev. B: Condens. Matter Mater. Phys., 1991, 44, 943–954, DOI:10.1103/PhysRevB.44.943.
- S. Walder, A. Gueguen and D. Kramer, Proton diffusion in orthorhombic perovskite sulfides, Chem. Mater., 2025, 37(4), 1349–1357, DOI:10.1021/acs.chemmater.4c01841.
- A. L. Gavin and G. W. Watson, Modelling the electronic structure of orthorhombic lamno3, Solid State Ionics, 2017, 299, 13–17, DOI:10.1016/j.ssi.2016.10.007 . URL https://www.sciencedirect.com/science/article/pii/S0167273816303770.
- R. P. Feynman, Forces in molecules, Phys. Rev., 1939, 56, 340–343, DOI:10.1103/PhysRev.56.340.
- H. Hellmann, Einführung in die quantenchemie, In Hans Hellmann: Einführung in die Quantenchemie: Mit biografischen Notizen von Hans Hellmann jr., Springer, pp. 19–376, 1937 Search PubMed.
- M. Wu, J. Xuan and J. LuFunctional stochastic gradient MCMC for bayesian neural networks, Proceedings of The 28th International Conference on Artificial Intelligence and Statistics, PMLR 2025, vol. 258, pp. 2998–3006, https://openreview.net/forum?id=MSYG8XHh0U Search PubMed.
- S. Chmiela, V. Vassilev-Galindo, O. T. Unke, A. Kabylda, H. E. Sauceda, A. Tkatchenko and K.-R. Müller, Accurate global machine learning force fields for molecules with hundreds of atoms, Sci. Adv., 2023, 9(2), eadf0873, DOI:10.1126/sciadv.adf0873.
- A. S. Christensen and O. Anatole von Lilienfeld, On the role of gradients for machine learning of molecular energies and forces, Mach. Learn.: Sci. Technol., 2020, 1, 045018 Search PubMed.
- M. Bogojeski, L. Vogt-Maranto, M. E. Tuckerman, K.-R. Müller and K. Burke, Quantum chemical accuracy from density functional approximations via machine learning, Nat. Commun., 2019, 11, 5223 CrossRef PubMed.
- S. Chmiela, A. Tkatchenko, H. E. Sauceda, I. Poltavsky, K. T. Schütt and K.-R. Müller, Machine learning of accurate energy-conserving molecular force fields, Sci. Adv., 2017, 3(5), e1603015, DOI:10.1126/sciadv.1603015.
- S. Chmiela, V. Vassilev-Galindo, O. T. Unke, A. Kabylda, H. E. Sauceda, A. Tkatchenko and K.-R. Müller, Accurate global machine learning force fields for molecules with hundreds of atoms, Sci. Adv., 2023, 9(2), eadf0873, DOI:10.1126/sciadv.adf0873.
- T. Rensmeyer, B. Craig, D. Kramer and O. Niggemann, High accuracy uncertainty-aware interatomic force modeling with equivariant bayesian neural networks, Digital Discovery, 2024, 3, 2356–2366, 10.1039/D4DD00183D.
- T. Chen, E. B. Fox, and C. Guestrin. Stochastic gradient Hamiltonian monte carlo. in International Conference on Machine Learning, 2014, pp. 1683–1691 Search PubMed.
- A. D. Becke, Density-functional thermochemistry. iii. the role of exact exchange, J. Chem. Phys., 1993, 98(7), 5648–5652, DOI:10.1063/1.464913.
- C. Lee, W. Yang and R. G. Parr, Development of the colle-salvetti correlation-energy formula into a functional of the electron density, Phys. Rev. B, 1988, 37, 785–789, DOI:10.1103/PhysRevB.37.785.
- P. J. Stephens, F. J. Devlin, C. F. Chabalowski and M. J. Frisch, Ab initio calculation of vibrational absorption and circular dichroism spectra using density functional force fields, J. Phys. Chem., 1994, 98(45), 11623–11627, DOI:10.1021/j100096a001.
- S. H. Vosko, L. Wilk and M. Nusair, Accurate spin-dependent electron liquid correlation energies for local spin density calculations: a critical analysis, Can. J. Phys., 1980, 58(8), 1200–1211, DOI:10.1139/p80-159.
- J. P. Perdew, K. Burke and M. Ernzerhof, Generalized gradient approximation made simple, Phys. Rev. Lett., 1996, 77, 3865–3868, DOI:10.1103/PhysRevLett.77.3865.
- B. Hourahine, B. Aradi, V. Blum, F. Bonafé, A. Buccheri, C. Camacho, C. Cevallos, M. Y. Deshaye, T. Dumitrică, A. Dominguez, S. Ehlert, M. Elstner, T. van der Heide, J. Hermann, S. Irle, J. J. Kranz, C. Köhler, T. Kowalczyk, T. Kubař, I. S. Lee, V. Lutsker, R. J. Maurer, S. K. Min, I. Mitchell, C. Negre, T. A. Niehaus, A. M. N. Niklasson, A. J. Page, A. Pecchia, G. Penazzi, M. P. Persson, J. Řezáč, C. G. Sánchez, M. Sternberg, M. Stöhr, F. Stuckenberg, A. Tkatchenko, V. W.-z. Yu and T. Frauenheim, DFTB+, a software package for efficient approximate density functional theory based atomistic simulations, J. Chem. Phys., 2020, 152(12), 124101, DOI:10.1063/1.5143190.
- G. J. Martyna, M. E. Tuckerman, D. J. Tobias and M. L. Klein, Explicit reversible integrators for extended systems dynamics, Mol. Phys., 1996, 87(5), 1117–1157, DOI:10.1080/00268979600100761.
- M. Elstner, D. Porezag, G. Jungnickel, J. Elsner, M. Haugk, Th. Frauenheim, S. Suhai and G. Seifert, Self-consistent-charge density-functional tight-binding method for simulations of complex materials properties, Phys. Rev. B, 1998, 58, 7260–7268, DOI:10.1103/PhysRevB.58.7260.
- M. Gaus, Q. Cui and M. E. Dftb3, Extension of the self-consistent-charge density-functional tight-binding method (scc-dftb), J. Chem. Theory Comput., 2011, 7(4), 931–948, DOI:10.1021/ct100684s.
- M. Gaus, X. Lu, M. Elstner and Q. Cui, Parameterization of dftb3/3ob for sulfur and phosphorus for chemical and biological applications, J. Chem. Theory Comput., 2014, 10(4), 1518–1537, DOI:10.1021/ct401002w.
| This journal is © The Royal Society of Chemistry 2026 |