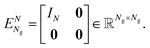Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
A symmetry-preserving and transferable representation for learning the Kohn–Sham density matrix
Liwei
Zhang
 *a,
Patrizia
Mazzeo
*a,
Patrizia
Mazzeo
 b,
Michele
Nottoli
b,
Michele
Nottoli
 c,
Edoardo
Cignoni
c,
Edoardo
Cignoni
 b,
Lorenzo
Cupellini
b,
Lorenzo
Cupellini
 b and
Benjamin
Stamm
b and
Benjamin
Stamm
 c
c
aInstitut für Geometrie und Praktische Mathematik, RWTH Aachen University, Templergraben 55, 52062 Aachen, Germany. E-mail: l.zhang@igpm.rwth-aachen.de
bDipartimento di Chimica e Chimica Industriale, Università di Pisa, Via G. Moruzzi 13, 56124 Pisa, Italy
cUniversität Stuttgart, Institute of Applied Analysis and Numerical Simulation, Pfaffenwaldring 57, 70569 Stuttgart, Germany
First published on 27th March 2026
Abstract
The Kohn–Sham (KS) density matrix is one of the most essential properties in KS density functional theory (DFT), from which many other physical properties of interest can be derived. In this work, we present a parameterized representation for learning the mapping from a molecular configuration to its corresponding density matrix using the Atomic Cluster Expansion (ACE) framework, which preserves the physical symmetries of the mapping, including isometric equivariance and Grassmannianity. Trained on several typical molecules, the proposed representation is shown to be systematically improvable with the increase of the model parameters and is transferable to molecules that are not part of and even more complex than those in the training set. The models generated by the proposed approach are illustrated as being able to generate reasonable predictions of the density matrix to either accelerate the DFT calculations or to provide approximations to some properties of the molecules.
1 Introduction
Computational chemistry often deals with many quantum mechanical calculations repeated on the same system or on similar systems. Examples are molecular dynamics (MD) simulations, repeated calculations on a statistical sampling, geometry optimizations, or even scans along some interesting coordinate. In all these cases, the results of already performed calculations can be used to fit a machine learning model able to predict energies and properties of subsequent calculations.1In the context of quantum chemistry, machine learning models have been used to fit properties, for example, the energy2–7 and atomic forces,8–10 or to predict more fundamental quantities like the Hamiltonian11–18 and the wavefunction.19,20 Machine learning models have also been used directly as interatomic potentials for molecular dynamics simulations of a variety of systems.21–25
Among the more fundamental quantities, various methods have been proposed to fit the electronic density matrix. These either target the electronic density in real space,26–37 or they target the corresponding electronic density matrix in a basis.3,38–44 Fitting the electronic density is a powerful strategy, as the density can then be directly used to compute different observables that arise from one-electron operators. Other strategies often need to train an ad hoc model for each property of interest, but multiple properties are often required for answering a scientific question (cf. ref. 45). Additionally, the electronic density matrix exhibits some extent of locality and sparsity,46,47 making it easier to derive a machine learning model based on local descriptors. By contrast, the Kohn–Sham Hamiltonian is less local. For polyalkenes, it has been shown to include long range charge–charge bielectronic interactions that create a systematic bias in the predictions.15
While being less general than fitting in real space, fitting the density on a suitable basis removes any projection error and removes the barrier between the predicted density and the quantum chemistry package of choice, which can be used to compute the properties of interest. Fitting the electronic density matrix provides two additional advantages. First, in the context of Hartree–Fock or density functional theory (DFT), an electronic density matrix can simply be used as an initial guess for the upcoming self-consistent field (SCF) calculation, instead of directly using it to access properties. This hybrid approach represents a middle ground between a full SCF calculation and directly using the density to access the properties: it retains the full accuracy of a normal SCF procedure, but at a reduced computational cost.42–44 The better the guess, the more efficient is the full accuracy model. Second, given a predicted electronic density matrix D, it is possible to assemble the corresponding Fock/Kohn–Sham matrix F = F(D), and the commutator FD − DF provides a measure of how accurate the prediction is, thus providing the opportunity to either discard low quality predictions or mark the data points with the worst predictions, which is useful in active learning strategies.48
However, the required mapping from the molecular configurations (coordinates and atomic numbers) to the corresponding density matrices is in general complicated and of high dimensionality, and therefore difficult to learn. The fitting problem becomes treatable by introducing appropriate molecular descriptors, which take into account physical knowledge such as invariance or equivariance of the target property. In this way, the required design space can be reduced. More specifically, the descriptors are functions of the molecular parameters satisfying a series of requisites: they are desired to be injective (exactly or approximately), economical to compute, and should capture the aforementioned symmetries of the target property. Depending on the order in which the various invariances are introduced, different classes of descriptors are obtained. A possible strategy is to compute translationally and rotationally invariant functions of the coordinates, and only then introducing the permutational invariance. Examples of descriptors of this kind are permutationally invariant polynomials (PIPs)49 and its variant atomic permutationally invariant polynomials (aPIPs)50 and the Moment Tensor Potentials (MTPs).51 Alternatively, it is possible to compute functions that are permutationally and translationally invariant, thereafter enforcing the rotational invariance. This is the strategy followed by the smooth overlap of atomic positions (SOAP),52 by the atomic cluster expansions (ACE)53 and by the Behler–Parrinello descriptors.54,55 The ACE descriptors are of particular interest as they include, in principle, many-body terms of arbitrarily high order, and are cheaper to compute than other alternatives.56 Notably, it has also been generalized to capture the equivariant properties.12,57
In this contribution, we propose a strategy that combines the strengths of the equivariant ACE descriptors with the flexibility of fitting the electronic density matrix in a basis, which respects the intrinsic properties of the density matrix. Specifically, the electronic density matrix is approximated with a linear regression in an ACE basis, similarly to the previous work on self-consistent Hamiltonians from one of the authors.12 This strategy for the density matrix differentiates from similar equivariant approaches43 as it is a linear model, for which the mathematical foundation is well established. Hence, the whole method becomes more tractable and interpretable. Similar differences are found between the work in ref. 12 and other equivariant approaches for the Hamiltonians.14,18
The strategy is used to train both specific models (that is, trained on a single molecule) and unified models (trained on multiple molecules). The resulting models are systematically improvable and, in the case of unified models, also transferrable to unseen molecules, provided that they share some chemical similarity with the training set. Both the specific and unified models can be used to reduce the number of SCF iterations or to directly predict the properties of interest.
2 Methods
Our approach targets the electronic density matrix D, which is the solution of the Kohn–Sham equation discretized with certain atomic orbital basis functions. In accordance with the atomic orbitals used, the density matrix can be split into blocks and sub-blocks, with each block corresponding to the interaction between one or two different atoms, and within which each sub-block corresponds to two orbital shells.Given a dataset containing the molecular coordinates {R(k)}k, the corresponding density matrices {D(k)}k and metadata such as the atomic orbitals adopted, the first step is to build suitable bases for representing the sub-blocks of the density matrix using the ACE descriptors. The ACE descriptors are particularly suitable as they provide a multi-set basis with the correct invariant and equivariant properties at low cost, allowing efficient description of molecular configurations with arbitrary numbers of particles. Such bases consist of functions of the positions and chemical elements of nearby atoms within some pre-defined local truncation cutoffs (Fig. 1(e and f)).
 | ||
Fig. 1 (a) A schematic of the process of learning the density matrix described in this paper. Here, the loss function 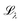 is defined as (7). The molecules for which the density matrices are predicted can be different from those in the training set. Finally, the predicted density matrix can be directly sent to quantum chemistry packages of choice for further operations. (b) Example block structure of the density matrix of C3H4O, where the atomic basis 6-31G(d) is used. (c) Block structure of the blocks DC, DO, DCC, DCO. (d) Block structure of the blocks DCH, DTHO. The block structure of the DHH block is omitted as it is a 2 by 2 matrix consisting of 4 ss-blocks. (e and f) Illustration of the local atomic environment for the construction of the onsite basis (e) and for the offsite basis (f). The red atoms are the centering atoms while the atoms in blue are the neighbors. The radii of the red and blue spheres indicate the onsite cutoff and the offsite cutoff, respectively. is defined as (7). The molecules for which the density matrices are predicted can be different from those in the training set. Finally, the predicted density matrix can be directly sent to quantum chemistry packages of choice for further operations. (b) Example block structure of the density matrix of C3H4O, where the atomic basis 6-31G(d) is used. (c) Block structure of the blocks DC, DO, DCC, DCO. (d) Block structure of the blocks DCH, DTHO. The block structure of the DHH block is omitted as it is a 2 by 2 matrix consisting of 4 ss-blocks. (e and f) Illustration of the local atomic environment for the construction of the onsite basis (e) and for the offsite basis (f). The red atoms are the centering atoms while the atoms in blue are the neighbors. The radii of the red and blue spheres indicate the onsite cutoff and the offsite cutoff, respectively. | ||
Next, a model is trained with a suitable training set to fit the density matrix. A small sub-model is required for each sub-block of the density matrix, that is, for each combination of elements and each combination of orbital shells assigned to the elements. Each sub-block of the density matrix is represented as a linear combination of the corresponding ACE basis, whose coefficients are determined through a standard least-squares minimization with a Tikhonov regularization to prevent overfitting.
The model can then be used to predict the density matrix for a given molecular geometry. The molecule can be part of the training set or beyond, provided that it contains elements found in the training set, and shares some chemical similarity with the training samples. Finally, the predicted density matrices are brought back to the manifold to which the real density matrices belong through a retraction step, which enforces other physical constraints on the predicted density matrices. Such predicted density matrices can be transmitted directly to certain Quantum Mechanical (QM) software packages to facilitate further validations or calculations.
A schematic representation of the entire workflow is given in Fig. 1(a).
Our approach of learning the density matrix was tested on various systems described with DFT ωB96X-D/6-31G(d). While the functional of choice ωB96X-D offers an accurate description of organic molecules in their different conformations, we present an implementation with a larger basis set (6-311G(d,p)) in Section S5 of the SI, demonstrating that the conclusions of this study are not affected by the choice of basis set. In total, we used datasets corresponding to 18 molecules within 3 chemical classes (aldehydes, aromatics, alcohols), 9 comprising 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 frames for training and 9 comprising 100 frames for testing. An overview of the datasets is given in Table S2 in the SI. Details on data generation can be found in Section 2.5. We designed tests of increasing complexity to validate the approach and assess its performance. The performance of the various models was assessed by computing the Root-Mean-Square-Error (RMSE) between the reference
000 frames for training and 9 comprising 100 frames for testing. An overview of the datasets is given in Table S2 in the SI. Details on data generation can be found in Section 2.5. We designed tests of increasing complexity to validate the approach and assess its performance. The performance of the various models was assessed by computing the Root-Mean-Square-Error (RMSE) between the reference 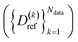 and predicted density matrices
and predicted density matrices 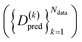 as
as
 | (1) |
The remaining part of this section provides detailed descriptions of each step of the workflow as well as additional information about the dataset preparation.
2.1 Density matrix
Let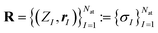 be a molecular configuration consisting of Nat atoms and N (valence) electron-pairs, with
be a molecular configuration consisting of Nat atoms and N (valence) electron-pairs, with 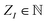 and
and 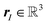 characterizing the atomic number and the position of the I-th atom, respectively. The union of all ZI characterizes the different elements in this given system, whose cardinality will be denoted by n. Other properties of the atoms can potentially also be included in σI but that would go beyond the scope of this paper. If an Ng(≥N) dimensional discretization space is adopted, in which the orbitals are approximated, then the corresponding KS equation will read as
characterizing the atomic number and the position of the I-th atom, respectively. The union of all ZI characterizes the different elements in this given system, whose cardinality will be denoted by n. Other properties of the atoms can potentially also be included in σI but that would go beyond the scope of this paper. If an Ng(≥N) dimensional discretization space is adopted, in which the orbitals are approximated, then the corresponding KS equation will read as| FR[DR]CR = SRCRER. |
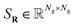 are the discretized KS operator (Hamiltonian) and the overlap matrix respectively,
are the discretized KS operator (Hamiltonian) and the overlap matrix respectively, 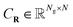 represents the coefficients of the orbitals in a given basis, DR = CRCTR is the density matrix, the main object of this paper, and ER is a diagonal matrix of order N, which contains the N corresponding eigenvalues of the system (sorted in ascending order). Without loss of generality, we assume that SR = INg, by adopting the Löwdin orthonormalization58 if necessary. Under this setting, the above eqn (1) can be rewritten as
represents the coefficients of the orbitals in a given basis, DR = CRCTR is the density matrix, the main object of this paper, and ER is a diagonal matrix of order N, which contains the N corresponding eigenvalues of the system (sorted in ascending order). Without loss of generality, we assume that SR = INg, by adopting the Löwdin orthonormalization58 if necessary. Under this setting, the above eqn (1) can be rewritten as| FR[DR]CR = CRER. | (2) |
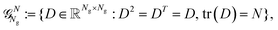 | (3) |
In the context of linear combinations of atomic orbitals (LCAO), the discretization space is spanned by a set of atomic orbitals 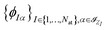 where
where 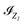 is the index set of the atomic orbitals centered at the I-th atom, depending only on the atomic number ZI. The density matrix, consequently, has elements that are invariant under translations of the system or permutations of the index of the atoms, and is equivariant under rotations and reflections of the whole system. It can be divided into several subblocks that have respective symmetries and can be learned independently. In Fig. 1(b–d), we take C3H4O as an example to illustrate the block structure of the density matrix. Note that a similar strategy is used in ref. 12 and is here extended to a case with multiple different elements. The detailed derivation of the block-wise equivariance of the density matrix can be found in the SI (Section S1). For simplicity, we omit the subscript R in DR hereafter when no ambiguity is introduced.
is the index set of the atomic orbitals centered at the I-th atom, depending only on the atomic number ZI. The density matrix, consequently, has elements that are invariant under translations of the system or permutations of the index of the atoms, and is equivariant under rotations and reflections of the whole system. It can be divided into several subblocks that have respective symmetries and can be learned independently. In Fig. 1(b–d), we take C3H4O as an example to illustrate the block structure of the density matrix. Note that a similar strategy is used in ref. 12 and is here extended to a case with multiple different elements. The detailed derivation of the block-wise equivariance of the density matrix can be found in the SI (Section S1). For simplicity, we omit the subscript R in DR hereafter when no ambiguity is introduced.
As can be seen from Fig. 1(b–d), there are two types of blocks appearing in the density matrix, the diagonal blocks and the off-diagonal ones. Depending on contexts, they are also called onsite and offsite, or homo- and hetero-orbital, respectively. We will use the terms onsite and offsite throughout this paper. For the targeted systems having n different elements, there exist n(n + 3)/2 matrix-valued functions which correspond to interactions of distinct elements (although some may be missing; for instance, when the system has only one oxygen atom, there is no O–O offsite interaction). As such, the block of the density matrix corresponding to the I-th and J-th atom is of the form
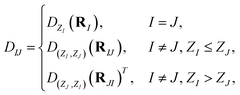 | (4) |
In addition, each of such matrix-valued functions can be further divided into completely independent sub-blocks corresponding to the atomic orbitals, as shown in Fig. 1(c) and (d), i.e.
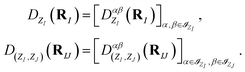 | (5) |
Our target then becomes the unified matrix-valued functionals 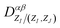 for various atomic numbers ZI, ZJ and the orbitals α, β assigned to the atoms, which have their distinct isometric equivariance and can be dealt with separately. Such structure of the density matrix forms the foundation of the transferability and parallelizability of the proposed method. We refer readers to Section S1 of the SI for a detailed discussion.
for various atomic numbers ZI, ZJ and the orbitals α, β assigned to the atoms, which have their distinct isometric equivariance and can be dealt with separately. Such structure of the density matrix forms the foundation of the transferability and parallelizability of the proposed method. We refer readers to Section S1 of the SI for a detailed discussion.
In practice, it is commonly believed that only atoms near the central atoms make substantial contributions to the corresponding part of the density matrix (also known as the nearsightedness of the object). As a result, certain cutoff strategies are often used when constructing the input atomic environment. We illustrate our truncation strategy in Fig. 1(e and f), where the particles in the red and blue spheres form the onsite and offsite environments RI and RIJ, respectively. The rigorous definitions of the truncated RI and RIJ are provided in the SI (Section S2).
Here, we assume that atoms of the same element are discretized by the same set of bases. However, the method proposed in this work can potentially be extended to the more general setting where atoms of the same element are assigned different atomic orbital basis functions, simply by artificially treating them as having different atom types.
2.2 Representation of the density matrix
One of the goals of this paper is to provide a faithful representation of the density matrix, respecting its inherent physical symmetries as much as possible to facilitate its learning. To this end, we adopt the equivariant ACE descriptors12,53 to approximate the functions D•, where the symbol • can be one of the indices appearing in the right hand side of (5).For each function D•, there exists a set of ACE bases 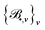 as functions of the local environments RI or RIJ, which has the same isometric equivariance as D• and asymptotically spans the function space to which D• belongs.59 The size of the basis is determined merely by two parameters: (i) the correlation order ν, which corresponds to the body order in physics (up to a constant, precisely, it is 1 and 2 less for onsite and offsite, respectively) and (ii) the maximum polynomial degree dmax, indicating the resolution of the one-particle basis. Additional details about these two parameters and the corresponding ACE basis, as well as the definition of the one-particle basis, can be found the SI (Section S2). Given the basis
as functions of the local environments RI or RIJ, which has the same isometric equivariance as D• and asymptotically spans the function space to which D• belongs.59 The size of the basis is determined merely by two parameters: (i) the correlation order ν, which corresponds to the body order in physics (up to a constant, precisely, it is 1 and 2 less for onsite and offsite, respectively) and (ii) the maximum polynomial degree dmax, indicating the resolution of the one-particle basis. Additional details about these two parameters and the corresponding ACE basis, as well as the definition of the one-particle basis, can be found the SI (Section S2). Given the basis 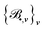 , we can approximate each function D• by a linear combination of
, we can approximate each function D• by a linear combination of 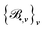 :
:
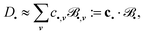 | (6) |
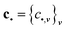 and
and  .
.
To predict the corresponding sub-block of the density matrix, the only thing left now is to estimate the coefficient c• for all possible indices •.
2.3 Parameter estimation
Suppose that a dataset is given of the form {(R(k), D(k))}k, where k is the index of the data point, R(k) is the k-th (global) molecular configuration and D(k) is the corresponding density matrix. The dataset can first be transformed into sets of local atomic clusters and their corresponding portions of the density matrix, according to the atomic number of each atom in R(k), as| {(R(k)IJ, D(k)•)}k,I,J, |
 | (7) |
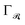 refers to some Tikhonov regularizer that can be customized with respect to the basis
refers to some Tikhonov regularizer that can be customized with respect to the basis 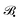 , and λ is a regularization parameter. Throughout our experiments, we use λ = 10−4 and choose
, and λ is a regularization parameter. Throughout our experiments, we use λ = 10−4 and choose 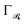 to be the smooth prior given in ref. 12. Once an (approximate) minimizer is found, it is possible to provide an approximation of the ground state density matrix through (6) for any given configuration R as long as its chemical composition of elements does not go beyond that of the training set.
to be the smooth prior given in ref. 12. Once an (approximate) minimizer is found, it is possible to provide an approximation of the ground state density matrix through (6) for any given configuration R as long as its chemical composition of elements does not go beyond that of the training set.
2.4 Retraction
The construction of the ACE basis as well as our representation (6) ensure that the predicted density matrix D has the desired isometric-equivariance. However, it does not guarantee that the prediction belongs to the Grassmann manifold (3), i.e. that it is a valid density matrix. To bring this restriction back, we introduce a retraction operator that maps D to the manifold.Since D is a real symmetric matrix of size Ng × Ng, its eigenvalue decomposition can be written as
| D = UDΣDUDT, |
 is unitary and ΣD is a diagonal matrix containing all the eigenvalues of D, sorted in descending order. The retraction is then defined as
is unitary and ΣD is a diagonal matrix containing all the eigenvalues of D, sorted in descending order. The retraction is then defined as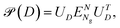 | (8) |
We mention that after applying the retraction, 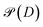 remains isometric equivariant and is the nearest element on the Grassmann manifold to D. A proof of this statement, as well as the well-definedness of
remains isometric equivariant and is the nearest element on the Grassmann manifold to D. A proof of this statement, as well as the well-definedness of 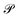 , is given in the SI (Section S3).
, is given in the SI (Section S3).
Fig. 1 summarizes our density matrix prediction scheme, from which we can see that the whole procedure, apart from the retraction step, is essentially parallelizable, including training and prediction.
It is worth noting that the proposed scheme does not require specific data origination, but just requires a consistent atomic basis discretization across the data set (or more abstractly, it requires only the equivariance of the data). The resulting density matrix can be used in many different application scenarios (cf. Subsections 3.4 and 3.5). We remark that a similar strategy is used in ref. 12 to fit the self-consistent Hamiltonian (i.e. Kohn–Sham) matrix for periodic crystal systems. We compared the aforementioned approach of learning the Hamiltonian matrix therein with the method proposed in this work, whose results can be found in the SI (Section S4).
2.5 Data preparation
To evaluate our density matrix learning strategy, we selected a set of neutral organic molecules, spanning a range of sizes (12 to 21 atoms) and functional groups, including carbonyls, alcohols, and substituted aromatic compounds. For alcohols, the test sets include both smaller and larger molecules, and different positioning of the hydroxyl group. For aromatic systems, we considered more challenging cases by combining functional groups present in the training data but exhibiting different behaviors as aromatic substituents. The molecules included in the training and test sets are listed in Table S3 of the SI.Each dataset was prepared with the same protocol, consisting of a sampling step and a QM calculation step. In the sampling step, each molecule was optimized with DFT B3LYP/6-31G in water, treated with IEFPCM,60 and solvated with an octahedral box of TIP3P waters,61 extending up to 35 Å from the molecule. The solvent was then minimized while keeping the molecule fixed. Thereafter, the whole system was heated from 0 K to 100 K in a 5 ps NVT simulation and from 100 K to 310 K in a 100 ps NPT simulation. The QM/MM production simulation was then run for 150 ps in the NVT ensemble, using the Langevin thermostat. The molecule was treated at the DFTB3 level of theory62 with 30b-3-1 parameters.63,64 Electrostatic interactions were treated with PME,65 using a 10 Å cutoff to divide the direct and reciprocal space. The first 50 ps of production trajectory were discarded. All simulations were performed with AMBER.66
In the second step, solvent molecules were stripped, and QM DFT calculations were run on equally spaced frames along the trajectory, using the ωB97X-D/6-31G(d) level of theory, and enforcing the use of spherical atomic basis functions. The training-and-testing dataset comprises nine organic molecules featuring different functional groups, whereas the test-only datasets comprise nine similar but distinct molecules. For training-and-testing datasets, the calculations were run on 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 frames, whereas for test only datasets, the calculations were run only on 100 frames. All the calculations for the datasets were performed using Gaussian 16.67
000 frames, whereas for test only datasets, the calculations were run only on 100 frames. All the calculations for the datasets were performed using Gaussian 16.67
The datasets were generated by storing the coordinates, the overlap matrices, the coefficient matrices, the Kohn–Sham matrices, as well as metadata such as the list of atoms and the calculation level in HDF5 binary files. An overview of the datasets is reported in Table S2 of the SI. All the datasets are available in the corresponding archive for the sake of reproducibility.68
3 Results
3.1 Specific models
To assess the method we proposed, we first show that it generates systematically improvable results, so that we can refrain from tuning the choice of model parameters (correlation orders ν and maximum polynomial degrees dmax, see Section S2 of the SI for more details). To this end, we show the results of molecule-specific models, each of which is trained with the geometries of only one molecule. For each molecule, the training set molecule contained less than 3000 frames in these tests. More details on dataset selection are given in Table S3 and Fig. S9 in the SI. The training set is then used to train the models with ν = 2, 3 and 4 ≤ dmax ≤ 8. For the local truncation, we tested cutoffs of 4.0 Å and 6.5 Å. For the sake of simplicity, we only show the results of aniline and propanol. We show in Fig. 2 the distribution of RMSE values as a function of the degree dmax used to generate the descriptors, for the two choices of the correlation order ν. | ||
| Fig. 2 The RMSEs (1) of the predicted density matrices for: (a) aniline with rcut = 4.0 Å, (b) aniline with rcut = 6.5 Å, (c) propanol with rcut = 4.0 Å and (d) propanol with rcut = 6.5 Å, obtained by the corresponding specific models with respect to different degrees dmax and for various orders ν. The solid and dashed lines refer to the average training and test set errors, and the shaded areas show the distribution of the errors for the corresponding models. | ||
As illustrated in Fig. 2, the training and test set RMSE distributions align with each other nicely, and are nearly normal, even with only 3000 samples used in the training phase. The similarity of the training/test-set errors suggests little overfitting in the training, validating the effects of the regularization we used (cf.eqn (7)). Comparing the two truncation parameters, we find that the models trained with larger cutoff radii can give better results, but the differences are not too substantial. The specific model reaches the smallest average RMSE at around 10−4 and 2 × 10−4 for aniline and propanol, respectively, which corresponds to a relative error in the whole density matrix of around 0.2% to 0.35% (note that ∥D∥2F = N the number of electrons, so the relative error is simply given by 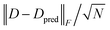 ). In all cases, the test set RMSE decreases monotonically with increasing ACE basis size, which implies that smaller errors can be expected simply by increasing either of the two model parameters. We also verified the robustness of our data preparation protocol by optimizing the geometries in vacuum at the reference level of theory and confirming that the RMSEs for these geometries remain lower than those reported for our test sets (see Table S8 in the SI).
). In all cases, the test set RMSE decreases monotonically with increasing ACE basis size, which implies that smaller errors can be expected simply by increasing either of the two model parameters. We also verified the robustness of our data preparation protocol by optimizing the geometries in vacuum at the reference level of theory and confirming that the RMSEs for these geometries remain lower than those reported for our test sets (see Table S8 in the SI).
3.2 Unified model
We then extend the training set to configurations from several distinct molecules. More specifically, the training set used in this subsection consists of a total of 2700 frames (300 frames evenly sampled from the first 9000 frames of each of the 9 training molecules). This extended training set is used to train the largest model mentioned above (ν = 3, dmax = 8) in order to obtain a unified model, which is then tested on the last 1000 configurations of the training molecules. Information on the molecules in the training set is given in Table 1. We compare the average test-set RMSE obtained by the unified model with those obtained by the corresponding specific models of the same size, whose results can be found in the first two columns of Table 1. To make the model more capable of capturing the similarity only of local chemical structures across different molecules, we choose rcut = 4.0 Å for the unified model. The results for the unified model with rcut = 6.5 Å are given in Table S3 in the SI, which indeed shows that a smaller cutoff is favorable in terms of the generalizability of the models.| Molecule | Specific model | Unified model | Unified Model-A |
|---|---|---|---|
| Acetaldehyde | 4.416 × 10−5 | 3.278 × 10−4 | 3.299 × 10−4 |
| Acrolein | 2.514 × 10−4 | 5.028 × 10−4 | 5.081 × 10−4 |
| Aniline | 4.300 × 10−4 | 4.868 × 10−4 | 4.876 × 10−4 |
| o-Toluidine | 5.430 × 10−4 | 5.962 × 10−4 | 5.940 × 10−4 |
| m-Toluidine | 5.384 × 10−4 | 5.824 × 10−4 | 5.822 × 10−4 |
| Benzene* | — | 4.058 × 10−4 | 3.646 × 10−4 |
| Toluene* | — | 6.369 × 10−4 | 5.980 × 10−4 |
| Phenol** | — | 4.809 × 10−3 | 6.770 × 10−4 |
| Benzaldehyde** | — | 4.129 × 10−3 | 1.201 × 10−3 |
| p-Toluidine** | — | 2.840 × 10−3 | 6.293 × 10−4 |
| 1-Propanol | 3.049 × 10−4 | 4.427 × 10−4 | 4.444 × 10−4 |
| 1-Butanol | 4.510 × 10−4 | 4.921 × 10−4 | 4.934 × 10−4 |
| 2-Butanol | 9.173 × 10−4 | 1.494 × 10−3 | 1.509 × 10−3 |
| 1-Hexanol | 1.031 × 10−3 | 5.324 × 10−4 | 5.314 × 10−4 |
| Ethanol* | — | 9.644 × 10−4 | 8.999 × 10−4 |
| 2-Propanol* | — | 7.701 × 10−4 | 7.480 × 10−4 |
| 2-Hexanol* | — | 7.384 × 10−4 | 7.353 × 10−4 |
| 1-Heptanol* | — | 8.896 × 10−4 | 8.980 × 10−4 |
As indicated in the first two columns of Table 1, the unified model overall achieves a performance comparable to the specific models trained on each molecule independently. This indicates that the model is able to gather information from distinct molecules, and offers the advantage of predicting the density matrices for multiple systems within a single model. Whereas the RMSE of the unified model for acetaldehyde is slightly higher, the unified model performs even better than the specific one for 1-hexanol. This demonstrates that the models generated by the proposed method are able to predict the density matrices of diverse molecular systems, despite their inherent structural dissimilarities, as long as a certain number of configurations is included in the training set. We remark that the training set of the unified model comprises a slightly smaller number of configurations than that of the specific models, and was sampled without particular strategies.
For reference, we also trained a unified model for the alcohols only, which is a simpler task compared to the unified one we introduce in this subsection. The results for the Unified Alcohols Model can be found in Table S3 in the SI.
3.3 Transfer to other molecules
As a more challenging task, we directly use the unified model obtained in Subsection 3.2 to predict the density matrices of some molecules beyond the training set, which may be larger or more complex. The corresponding average test set RMSEs as well as which molecules belong to the test set are reported in Table 1, from which we observe that the unified model can provide faithful predictions of the density matrices for benzene, toluene and all the alcohols, for which the obtained errors are similar to those within the training process. However, the unified model struggles with giving good predictions for phenol, benzaldehyde, and is less good at predicting the density matrices for p-toluidine. The poor prediction on these molecules can be attributed to the lack of information in the training set. Indeed, while the dataset includes both carbonyl compounds and alcohols, the chemical behaviour of these functional groups changes significantly when they are bonded to aromatic moieties. Additionally, when an aromatic molecule has two substituents, their effect depends on their relative positions. This explains why, despite the inclusion of o- and m-toluidine in the training set, the model struggles to accurately predict for p-toluidine.To test whether the weaker performance of the unified model on the three molecules is caused by the limitation of the method itself or just by the training set, we design an augmented training set, which consists of the data points of the previous unified training set, and 10 frames from each of the three molecules, evenly sampled from the first 90 frames. This augmented training set contains 2730 samples in total. We train the above (3,8)-model with the augmented training set, and obtain a new model Unified Model-A, with the suffix “A” indicating that it is trained with an augmented set. The augmented unified model is again used to predict the density matrices for all the involved molecules, and the corresponding test set RMSEs are listed in the third column of Table 1. The results show that the augmented unified model achieves a higher accuracy for the three molecules with previously critical accuracy, which is similar in magnitude to that of the training molecules, while maintaining a comparable effectiveness for the other systems involved. This indicates that the performance of the model is primarily limited by the variability of the training set rather than by its capability. In Fig. S3 and S4 of the SI, we applied a Uniform Manifold Approximation and Projection (UMAP)69 to our datasets, and show the first two components of each data point, to further justify this.
The RMSE results presented in this section suggest that the models generated by the proposed method can be uniformly refined simply by increasing the two model parameters. In addition, the proposed unified models can be transferred to the molecules that are not known at the training stage, provided some similarity in the chemical geometries. The performance of the generated models is mainly limited by the design of the training set, rather than the representation itself.
3.4 Accelerating the SCF iterations
A natural application of our model is to use the predictions as the initial guesses of the SCF procedure, in order to achieve SCF convergence faster. For each test geometry, we use the proposed models to predict the density matrix and provide it to Gaussian as an initial guess. For these calculations, we used the development version of the Gaussian suite of programs.70 Communication with Gaussian is possible thanks to the GauOpen open-source library.71 We compared the number of iterations required to achieve convergence with all our models and with the Harris guess.72 While other strategies exist for the generation of accurate densities,73,74 the Harris guess was chosen as a compromise between accuracy and simplicity, as it is already available in Gaussian. Table 2 reports the average of iterations obtained with the default guess, specific models with rcut = 4.0 Å and unified models for SCF convergence tolerance 10−6. The same results for the convergence levels 10−7 and 10−8 are reported in Table S6 in the SI. The value within parentheses indicates the percentage of reduction with respect to the default guess. The table shows that, as expected, the specific model achieves the highest reduction in the number of iterations for each molecule, with a maximum for acetaldehyde, where a 44% reduction is observed. We also compared the performance of the specific models with rcut = 4.0 Å and rcut = 6.5 Å, finding no significant differences (see Table S4 in the SI). On average, the specific models allow us to save three iterations (30% of reduction with respect to the default guess). Moving to the unified model, we observe that for the majority of the molecules in the training set, the predicted density is comparable to that obtained with the respective specific model, with a slightly greater loss of accuracy for the two carbonyl molecules. For what concerns the out-of-sample molecules, the model exhibits good transferability for alcohols, achieving comparable results for both known and unknown molecules. Conversely, the predictions for phenol, p-toluidine, and benzaldehyde were particularly poor, even falling below the accuracy of the default guess. However including just 10 frames in the training set for each of these three molecules (Unified Model-A) enhances the performance, as demonstrated by the RMSEs, and reduces the number of iterations by around two.| Molecule | Default guess | Specific model | Unified model | Unified Model-A |
|---|---|---|---|---|
| Acetaldehyde | 9.4 ± 0.1 | 5.2 ± 0.1 (∼44%) | 7.5 ± 0.1 (∼20%) | 7.5 ± 0.1 (∼20%) |
| Acrolein | 10.5 ± 0.1 | 7.5 ± 0.2 (∼29%) | 8.3 ± 0.2 (∼21%) | 8.3 ± 0.2 (∼21%) |
| Aniline | 9.9 ± 0.1 | 7.2 ± 0.1 (∼28%) | 7.5 ± 0.1 (∼24%) | 7.5 ± 0.1 (∼24%) |
| o-Toluidine | 10.0 ± 0.0 | 7.6 ± 0.1 (∼24%) | 7.8 ± 0.1 (∼22%) | 7.8 ± 0.1 (∼22%) |
| m-Toluidine | 10.0 ± 0.0 | 7.3 ± 0.1 (∼27%) | 7.6 ± 0.1 (∼24%) | 7.6 ± 0.1 (∼24%) |
| Benzene* | 9.0 ± 0.0 | — | 7.4 ± 0.1 (∼18%) | 7.4 ± 0.1 (∼18%) |
| Toluene* | 9.0 ± 0.0 | — | 8.0 ± 0.0 (∼11%) | 7.9 ± 0.1 (∼12%) |
| Phenol** | 9.9 ± 0.1 | — | 10.0 ± 0.1 (∼−1%) | 8.1 ± 0.1 (∼18%) |
| Benzaldehyde** | 10.7 ± 0.1 | — | 10.5 ± 0.1 (∼2%) | 9.0 ± 0.0 (∼16%) |
| p-Toluidine** | 10.0 ± 0.0 | — | 8.3 ± 0.1 (∼16%) | 7.8 ± 0.1 (∼21%) |
| 1-Propanol | 9.0 ± 0.0 | 6.0 ± 0.1 (∼33%) | 6.5 ± 0.1 (∼27%) | 6.5 ± 0.1 (∼28%) |
| 1-Butanol | 9.0 ± 0.0 | 6.4 ± 0.1 (∼29%) | 6.6 ± 0.1 (∼27%) | 6.5 ± 0.1 (∼27%) |
| 2-Butanol | 9.0 ± 0.0 | 7.2 ± 0.1 (∼20%) | 7.0 ± 0.1 (∼22%) | 7.1 ± 0.1 (∼22%) |
| 1-Hexanol | 9.0 ± 0.0 | 6.4 ± 0.1 (∼29%) | 6.5 ± 0.1 (∼28%) | 6.5 ± 0.1 (∼28%) |
| Ethanol* | 9.1 ± 0.0 | — | 7.2 ± 0.1 (∼21%) | 7.1 ± 0.1 (∼21%) |
| 2-Propanol* | 9.0 ± 0.0 | — | 7.1 ± 0.1 (∼21%) | 7.0 ± 0.0 (∼22%) |
| 2-Hexanol* | 9.0 ± 0.0 | — | 7.0 ± 0.1 (∼23%) | 7.0 ± 0.1 (∼22%) |
| 1-Heptanol* | 9.0 ± 0.0 | — | 6.6 ± 0.1 (∼27%) | 6.6 ± 0.1 (∼27%) |
It is worth mentioning that the computational time for predicting a density matrix for a given configuration using our model is almost negligible compared to a single SCF iteration. For example, it takes about 112 ms to obtain a predicted density matrix for a propanol molecule using the unified model in a single thread, whereas a single SCF iteration, even carried out on 6 threads, takes an average of 626 ms. Hence, the percentage of reduction is almost exactly the acceleration that we gain.
3.5 Predictions of physical properties
The predicted density matrices can also be directly used to derive physical properties of interest, obtaining satisfactory predictions. This was achieved by providing the predicted density matrix to Gaussian to perform the corresponding calculations directly. Fig. 3 illustrates the error in energy, Mulliken charges, dipole moment, and forces with respect to the results obtained from the corresponding converged density matrix. The plot compares the errors obtained using the density matrix predicted with Unified Model-A (pink) and the default guess available in Gaussian (blue). For the forces, in particular, the construction of the Fock matrix F = F(D) is required. Therefore, it is desirable to use the new density matrix obtained after a single SCF cycle to compute the forces for both the predicted density matrix and the Gaussian default guess, given the minimal additional cost. The same plots for specific models, unified alcohols model and unified model are reported in the SI (see Fig. S5–S7).Averaging over all molecules, we obtain a mean absolute error (MAE) of 2.7 kcal mol−1 for energy and 6.5 kcal mol−1 Å−1 for forces. Although they do not achieve chemical accuracy (1 kcal mol−1 for energies and 1 kcal mol−1 Å−1 for forces) except for the two aldehydes, the predictions can be considered as qualitatively correct results in most of the cases. Typically, the average errors of the properties derived from the predicted density matrix for all the involved molecules are 1 to 3 orders of magnitude smaller than those from the Gaussian default guesses. This trend holds consistently for both the aldehyde and aromatic families. Despite the existence of some outliers for the alcohols, especially 2-butanol, which also turned out to be the one having the largest test set RMSE within the training molecules (unified models), they are only rare occurrences, as indicated by the error distribution shown in the violin plots. We expect that this can be resolved by adjusting the training set to include the structures corresponding to the outliers. This result also suggests that one may need to give the alcohol family more weight in the training. It is therefore likely that a better selection of training points, obtained for example by active learning approaches (see e.g. ref. 48 and the references therein), will give more stable errors. In Subsection 3.6, we provide a potential way to determine whether a given prediction should be disregarded or whether the corresponding structure should be included in the training set to improve model performance.
3.6 Commutator and active learning
As the last application, we use the predicted density matrices to compute the corresponding KS matrix F = F(D), and check how well the commutator condition FD = DF is fulfilled. Indeed, when convergence is achieved, FD = DF must hold exactly. Therefore, the norm of FD − DF is a residue and can serve as a physical parameter to evaluate the accuracy of the prediction. In Fig. 4, we present the relationship between the commutator violation error, measured in the Frobenius norm, and the relative error in the predicted energy. It turns out that there is an empirical algebraic relation observed between the two errors. Similar plots for other properties are provided in the SI (Fig. S8), which also demonstrate positive correlations while the trend is less clear compared to that for the energies. Thus, one can use the commutator error to determine whether to disregard a prediction, without accessing the real physical properties of interest. From another perspective, one can also use the commutator error as an indicator of which geometries to include in the training process in an active learning framework, so as to avoid performing full SCF iterations for all the geometries in a relatively large training set.We use 1-propanol to illustrate a prototypical active learning loop. Given a training pool of molecular geometries (the first 5000 propanol frames in this example), we first train an ML model using the first 500 frames. The ML model is then used to compute the commutator errors for all the geometries in the pool, after which the 500 frames with the worst commutator errors are added to the training set. Another round of training is then carried out to produce a new ML model. This process is repeated until the frames to be added start to overlap with the existing training set. Note that during the active learning procedure, the full SCF calculations are only required, once each, for the geometries selected for the training set.
Fig. 5 shows a comparison of the average and maximum test set RMSEs in the test frames of the predicted density matrices obtained using (3,8)-models, which were trained using either the first few frames or the training sets generated by the active learning procedure described above. The last 5000 frames were used as the test set. As the RMSEs suggest, the training sets generated by the active learning strategy seem to capture quickly the structures with which the model is less familiar. Compared to the original strategy, the active learning strategy enables us to achieve a comparable average RMSE, yet a notably lower maximum RMSE, using significantly fewer training frames (only 1476 in the final training set). To rationalize the termination criterion, we performed an additional iteration, resulting in a new training set comprising 1737 samples. However, this had only a marginal impact on the outcome. The results demonstrate that the commutator-based active learning strategy is capable of efficiently generating reasonable training sets and has the potential to eliminate outliers.
As shown in the previous section, it is indeed the design of the training set that limits the accuracy and transferability of the proposed method. It is therefore one of our immediate future works to implement our commutator-based active learning strategy when dealing with multiple molecules.
4 Conclusions
We presented a simple yet powerful regression model for learning the ground-state density matrix of arbitrary molecules in an atom-centered basis set. Our model exploits the flexibility and the favorable symmetry properties of the equivariant ACE descriptors, which represent a natural set of features to represent the density matrix. The resulting model can be improved systematically by increasing the size of the ACE basis (order and degree) or by tuning the training samples. More importantly, our model can learn the relationship between molecular geometry and density matrix using information from multiple distinct molecules. This opens the possibility of building unified models for molecules with similar local structure, in contrast to other approaches.3,42 As a consequence, our model is transferable to unseen molecules, provided that they have a local chemical structure similar to the ones in the training set.A model generating fast predictions for the density matrices provides, first of all, a way to accelerate KS-DFT calculations by generating a better starting guess for self-consistent iterations. Besides the straightforward application to ab initio molecular dynamics or geometry optimizations, the possibility of extrapolating predictions to unseen molecules provides the opportunity to accelerate KS-DFT calculations on many different molecules without molecule-specific training. The guesses generated by our unified model allow saving about 20% of the SCF iterations compared to the default guess in most of the cases.
Secondly, the predicted density matrix can be used in a quantum chemistry code to directly compute multiple properties. We have tested how well this model predicts energy, Mulliken atomic charges, molecular dipole, and atomic forces. The density matrix predictions give significantly better estimates than the standard guess for all these properties, and especially for the energy, even for unseen molecules.
Lastly, the commutator error can be obtained from the predicted density matrix at a relatively low computational cost, which has been shown to be a reasonable indicator of prediction reliability. We have incorporated the commutator error into an active learning loop, which produced more robust results than the baseline sampling strategy while requiring a smaller training set, thereby demonstrating the potential of using the commutator error to assess whether a new geometry is already adequately represented.
Our model still presents some limitations. First, although the reduction in SCF iterations is significant, a substantial improvement would be necessary to consistently accelerate KS-DFT calculations by at least a factor of two. Second, the energies and forces predicted by our model still do not reach chemical accuracy, which would be needed for direct applications. Nonetheless, all models show significant room for improvement, both in the model flexibility and in the choice of the training set, making our strategy promising for both applications, especially thanks to the observed systematic improvability and rigorous methodology.
Overall, our results show that learning the density matrix from descriptors encoding the correct symmetry features represents a promising strategy towards more complete and transferable ML models. In particular, the density matrix represents the solution of the KS-DFT equations and thus gives direct access to numerous properties with one single ML model. Further, the model turns out to be transferable to unseen molecular structures, which is a central stepping stone towards this development.
Author contributions
L. Z., L. C. and B. S. conceived the study. All authors contributed to the methodology. L. Z. and M. N. wrote the code. P. M. and E. C. prepared the datasets. L. Z., P. M. and M. N. did the numerical investigation. L. Z., P. M., M. N., L. C. and B. S. wrote the manuscript. All authors read and approved the final manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
All datasets used in this contribution and the corresponding codes used to read the data, construct and train the models, and predict the density matrices are available at ref. 75 for reproducibility. The datasets are also available in identical form at ref. 68, and the corresponding codes are also hosted at https://github.com/ACEsuit/ACEdensitymatrix, along with a comprehensive script showing how the entire learning procedure is performed.Supplementary information (SI): equivariance of the density matrix, construction of the ACE basis, properties of the retraction, comparison of fitting the density matrix and the Kohn–Sham matrix, effect of the basis set, additional tables and additional plots. See DOI: https://doi.org/10.1039/d5dd00230c.
Acknowledgements
The authors thank Filippo Lipparini for performing the calculations with the Gaussian development version. M. N. and B. S. thank the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) for supporting this work by funding – EXC2075 – 390740016 under Germany's Excellence Strategy. We acknowledge the support by the Stuttgart Center for Simulation Science (SimTech). L. Z. and M. N. and B. S. acknowledge funding by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) – project number 442047500 through the Collaborative Research Center “Sparsity and Singular Structures” (SFB 1481).Notes and references
- N. Fedik, R. Zubatyuk, M. Kulichenko, N. Lubbers, J. S. Smith, B. Nebgen, R. Messerly, Y. W. Li, A. I. Boldyrev, K. Barros, O. Isayev and S. Tretiak, Nat. Rev. Chem., 2022, 6, 653–672 CrossRef CAS PubMed.
- Z. Qiao, M. Welborn, A. Anandkumar, F. R. Manby and T. F. Miller, J. Chem. Phys., 2020, 153, 124111 CrossRef CAS PubMed.
- X. Shao, L. Paetow, M. E. Tuckerman and M. Pavanello, Nat. Commun., 2023, 14, 6281 CrossRef CAS PubMed.
- Y. Chen, L. Zhang, H. Wang and W. E, J. Phys. Chem. A, 2020, 124, 7155–7165 CrossRef CAS PubMed.
- S. Dick and M. Fernandez-Serra, Nat. Commun., 2020, 11, 3509 CrossRef CAS.
- A. S. Christensen, S. K. Sirumalla, Z. Qiao, M. B. O'Connor, D. G. A. Smith, F. Ding, P. J. Bygrave, A. Anandkumar, M. Welborn, F. R. Manby and T. F. Miller, J. Chem. Phys., 2021, 155, 204103 CrossRef CAS PubMed.
- M. Welborn, L. Cheng and T. F. Miller, J. Chem. Theory Comput., 2018, 14, 4772–4779 CrossRef CAS PubMed.
- O. T. Unke, S. Chmiela, H. E. Sauceda, M. Gastegger, I. Poltavsky, K. T. Schütt, A. Tkatchenko and K. R. Müller, Chem. Rev., 2021, 121, 10142–10186 CrossRef CAS.
- M. Pinheiro, F. Ge, N. Ferré, P. O. Dral and M. Barbatti, Chem. Sci., 2021, 12, 14396–14413 RSC.
- O. T. Unke, S. Chmiela, M. Gastegger, K. T. Schütt, H. E. Sauceda and K. R. Müller, Nat. Commun., 2021, 12(1), 1–14 Search PubMed.
- K. T. Schütt, M. Gastegger, A. Tkatchenko, K.-R. Müller and R. J. Maurer, Nat. Commun., 2019, 10, 5024 CrossRef.
- L. Zhang, B. Onat, G. Dusson, A. McSloy, G. Anand, R. J. Maurer, C. Ortner and J. R. Kermode, npj Comput. Mater., 2022, 8, 158 CrossRef.
- J. Nigam, M. J. Willatt and M. Ceriotti, J. Chem. Phys., 2022, 156, 014115 CrossRef CAS PubMed.
- H. Li, Z. Wang, N. Zou, M. Ye, R. Xu, X. Gong, W. Duan and Y. Xu, Nat. Comput. Sci., 2022, 2, 367–377 CrossRef PubMed.
- E. Cignoni, D. Suman, J. Nigam, L. Cupellini, B. Mennucci and M. Ceriotti, ACS Cent. Sci., 2024, 10, 637–648 CrossRef CAS PubMed.
- M. Shakiba and A. V. Akimov, J. Chem. Theory Comput., 2024, 20, 2992–3007 CrossRef CAS PubMed.
- H. Tang, B. Xiao, W. He, P. Subasic, A. R. Harutyunyan, Y. Wang, F. Liu, H. Xu and J. Li, Nat. Comput. Sci., 2024, 5, 144–154 CrossRef PubMed.
- D. Suman, J. Nigam, S. Saade, P. Pegolo, H. Türk, X. Zhang, G. K.-L. Chan and M. Ceriotti, J. Chem. Theory Comput., 2025, 21, 6505–6516 CrossRef CAS.
- J. Hermann, Z. Schätzle and F. Noé, Nat. Chem., 2020, 12, 891–897 CrossRef CAS PubMed.
- X. Li, C. Fan, W. Ren and J. Chen, Phys. Rev. Res., 2022, 4, 013021 CrossRef CAS.
- S. Chmiela, H. E. Sauceda, I. Poltavsky, K. R. Müller and A. Tkatchenko, Comput. Phys. Commun., 2019, 240, 38–45 CrossRef CAS.
- K. Zinovjev, L. Hedges, R. M. Andreu, C. Woods, I. Tuñón and M. W. van der Kamp, J. Chem. Theory Comput., 2024, 20, 4514–4522 CrossRef CAS.
- R. Galvelis, A. Varela-Rial, S. Doerr, R. Fino, P. Eastman, T. E. Markland, J. D. Chodera and G. D. Fabritiis, J. Chem. Inf. Model., 2023, 63, 5701–5708 CrossRef CAS.
- A. Kabylda, J. T. Frank, S. S. Dou, A. Khabibrakhmanov, L. M. Sandonas, O. T. Unke, S. Chmiela, K.-R. Muller and A. Tkatchenko, Molecular Simulations with a Pretrained Neural Network and Universal Pairwise Force Fields, 2024, https://chemrxiv.org/engage/chemrxiv/article-details/6704263051558a15ef6478b6 Search PubMed.
- P. Mazzeo, E. Cignoni, A. Arcidiacono, L. Cupellini and B. Mennucci, Digital Discovery, 2024, 3, 2560–2571 RSC.
- F. Brockherde, L. Vogt, L. Li, M. E. Tuckerman, K. Burke and K.-R. Müller, Nat. Commun., 2017, 8, 872 CrossRef PubMed.
- J. M. Alred, K. V. Bets, Y. Xie and B. I. Yakobson, Compos. Sci. Technol., 2018, 166, 3–9 CrossRef CAS.
- A. Chandrasekaran, D. Kamal, R. Batra, C. Kim, L. Chen and R. Ramprasad, npj Comput. Mater., 2019, 5, 22 CrossRef.
- S. Gong, T. Xie, T. Zhu, S. Wang, E. R. Fadel, Y. Li and J. C. Grossman, Phys. Rev. B, 2019, 100, 184103 CrossRef CAS.
- A. Fabrizio, A. Grisafi, B. Meyer, M. Ceriotti and C. Corminboeuf, Chem. Sci., 2019, 10, 9424–9432 RSC.
- B. Cuevas-Zuviría and L. F. Pacios, J. Chem. Inf. Model., 2020, 60, 3831–3842 CrossRef PubMed.
- R. Meyer, M. Weichselbaum and A. W. Hauser, J. Chem. Theory Comput., 2020, 16, 5685–5694 CrossRef CAS PubMed.
- J. A. Ellis, L. Fiedler, G. A. Popoola, N. A. Modine, J. A. Stephens, A. P. Thompson, A. Cangi and S. Rajamanickam, Phys. Rev. B, 2021, 104, 035120 CrossRef CAS.
- B. Cuevas-Zuviría and L. F. Pacios, J. Chem. Inf. Model., 2021, 61, 2658–2666 CrossRef.
- P. B. Jørgensen and A. Bhowmik, npj Comput. Mater., 2022, 8, 183 CrossRef.
- B. Focassio, M. Domina, U. Patil, A. Fazzio and S. Sanvito, npj Comput. Mater., 2023, 9, 87 CrossRef.
- R.-G. Lee and Y.-H. Kim, npj Comput. Mater., 2024, 10, 248 CrossRef CAS.
- A. Grisafi, A. Fabrizio, B. Meyer, D. M. Wilkins, C. Corminboeuf and M. Ceriotti, ACS Cent. Sci., 2018, 5, 57–64 CrossRef PubMed.
- A. M. Lewis, A. Grisafi, M. Ceriotti and M. Rossi, J. Chem. Theory Comput., 2021, 17, 7203–7214 CrossRef CAS PubMed.
- K. R. Briling, A. Fabrizio and C. Corminboeuf, J. Chem. Phys., 2021, 155, 024107 CrossRef CAS PubMed.
- J. A. Rackers, L. Tecot, M. Geiger and T. E. Smidt, Mach. Learn.: Sci. Technol., 2023, 4, 015027 Search PubMed.
- S. Hazra, U. Patil and S. Sanvito, J. Chem. Theory Comput., 2024, 20, 4569–4578 CrossRef CAS PubMed.
- P. Febrer, P. B. Jørgensen, M. Pruneda, A. García, P. Ordejón and A. Bhowmik, Mach. Learn.: Sci. Technol., 2025, 6, 025013 Search PubMed.
- P. Stishenko, C. Qian, J. Westermayr, R. Maurer and A. Logsdail, Practical integration of machine learning into ab initio calculations and workflows: accelerating the SCF cycle via density matrix predictions, 2025, DOI:10.26434/chemrxiv-2025-xn2mp.
- M. Gastegger, J. Behler and P. Marquetand, Chem. Sci., 2017, 8, 6924–6935 RSC.
- P. E. Maslen, C. Ochsenfeld, C. A. White, M. S. Lee and M. Head-Gordon, J. Phys. Chem. A, 1998, 102, 2215–2222 CrossRef CAS.
- J. Kussmann, M. Beer and C. Ochsenfeld, Wiley Interdiscip. Rev. Comput. Mol. Sci., 2013, 3, 614–636 Search PubMed.
- C. van der Oord, M. Sachs, G. Csányi and C. Ortner, npj Comput. Mater., 2023, 9, 168 CrossRef CAS PubMed.
- Z. Xie and J. M. Bowman, J. Chem. Theory Comput., 2009, 6, 26–34 CrossRef PubMed.
- C. van der Oord, G. Dusson, G. Csányi and C. Ortner, Mach. Learn.: Sci. Technol., 2020, 1, 015004 Search PubMed.
- A. Shapeev, Multiscale Model. Simul., 2016, 14, 1153–1173 CrossRef.
- A. P. Bartók, R. Kondor and G. Csányi, Phys. Rev. B:Condens. Matter Mater. Phys., 2013, 87, 184115 CrossRef.
- R. Drautz, Phys. Rev. B, 2019, 99, 014104 CrossRef CAS.
- J. Behler, J. Chem. Phys., 2011, 134, 074106 CrossRef PubMed.
- J. Behler, Chem. Rev., 2021, 121, 10037–10072 CrossRef CAS PubMed.
- Y. Lysogorskiy, C. van der Oord, A. Bochkarev, S. Menon, M. Rinaldi, T. Hammerschmidt, M. Mrovec, A. Thompson, G. Csányi, C. Ortner and R. Drautz, npj Comput. Mater., 2021, 7, 97 CrossRef CAS.
- R. Drautz, Phys. Rev. B, 2020, 102, 024104 CrossRef CAS.
- P.-O. Löwdin, J. Chem. Phys., 1956, 18, 365–375 CrossRef.
- G. Dusson, M. Bachmayr, G. Csányi, R. Drautz, S. Etter, C. van der Oord and C. Ortner, J. Comput. Phys., 2022, 454, 110946 CrossRef CAS.
- E. Cancès, B. Mennucci and J. Tomasi, J. Chem. Phys., 1997, 107, 3032–3041 CrossRef.
- P. Mark and L. Nilsson, J. Phys. Chem. A, 2001, 105, 9954–9960 CrossRef CAS.
- M. Gaus, Q. Cui and M. Elstner, J. Chem. Theory Comput., 2011, 7, 931–948 CrossRef CAS PubMed.
- M. Gaus, X. Lu, M. Elstner and Q. Cui, J. Chem. Theory Comput., 2014, 10, 1518–1537 CrossRef CAS.
- X. Lu, M. Gaus, M. Elstner and Q. Cui, J. Phys. Chem. B, 2015, 119, 1062–1082 CrossRef CAS PubMed.
- T. Darden, D. York and L. Pedersen, J. Chem. Phys., 1993, 98, 10089–10092 CrossRef CAS.
- D. A. Case, H. M. Aktulga, K. Belfon, I. Y. Ben-Shalom, J. T. Berryman, S. R. Brozell, D. S. Cerutti, T. E. Cheatham III, G. A. Cisneros, V. W. D. Cruzeiro, T. A. Darden, R. E. Duke, G. Giambasu, M. K. Gilson, H. Gohlke, A. W. Goetz, R. Harris, S. Izadi, S. A. Izmailov, K. Kasavajhala, M. C. Kaymak, E. King, A. Kovalenko, T. Kurtzman, T. S. Lee, S. LeGrand, P. Li, C. Lin, J. Liu, T. Luchko, R. Luo, M. Machado, V. Man, M. Manathunga, K. M. Merz, Y. Miao, O. Mikhailovskii, G. Monard, H. Nguyen, K. A. O'Hearn, A. Onufriev, F. Pan, S. Pantano, R. Qi, A. Rahnamoun, D. R. Roe, A. Roitberg, C. Sagui, S. Schott-Verdugo, A. Shajan, J. Shen, C. L. Simmerling, N. R. Skrynnikov, J. Smith, J. Swails, R. C. Walker, J. Wang, J. Wang, H. Wei, R. M. Wolf, X. Wu, Y. Xiong, Y. Xue, D. M. York, S. Zhao and P. A. Kollman, AMBER 2022, University of California, San Francisco, 2022 Search PubMed.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, M. Caricato, A. V. Marenich, J. Bloino, B. G. Janesko, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. A. Montgomery Jr, J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian16 Revision C.01, Gaussian Inc., Wallingford CT, 2016 Search PubMed.
- L. Zhang, P. Mazzeo, M. Nottoli, E. Cignoni, L. Cupellini and B. Stamm, Replication Data for: a symmetry-preserving and transferable representation for learning the Kohn-Sham density matrix, 2025, DOI:10.18419/DARUS-4902.
- L. McInnes, J. Healy, N. Saul and L. Großberger, J. Open Source Softw., 2018, 3, 861 CrossRef.
- M. J. Frisch, G. W. Trucks, H. B. Schlegel, G. E. Scuseria, M. A. Robb, J. R. Cheeseman, G. Scalmani, V. Barone, G. A. Petersson, H. Nakatsuji, X. Li, A. V. Marenich, M. Caricato, J. Bloino, B. G. Janesko, J. Zheng, R. Gomperts, B. Mennucci, H. P. Hratchian, J. V. Ortiz, A. F. Izmaylov, J. L. Sonnenberg, D. Williams-Young, F. Ding, F. Lipparini, F. Egidi, J. Goings, B. Peng, A. Petrone, T. Henderson, D. Ranasinghe, V. G. Zakrzewski, J. Gao, N. Rega, G. Zheng, W. Liang, M. Hada, M. Ehara, K. Toyota, R. Fukuda, J. Hasegawa, M. Ishida, T. Nakajima, Y. Honda, O. Kitao, H. Nakai, T. Vreven, K. Throssell, J. J. A. Montgomery, J. E. Peralta, F. Ogliaro, M. J. Bearpark, J. J. Heyd, E. N. Brothers, K. N. Kudin, V. N. Staroverov, T. A. Keith, R. Kobayashi, J. Normand, K. Raghavachari, A. P. Rendell, J. C. Burant, S. S. Iyengar, J. Tomasi, M. Cossi, J. M. Millam, M. Klene, C. Adamo, R. Cammi, J. W. Ochterski, R. L. Martin, K. Morokuma, O. Farkas, J. B. Foresman and D. J. Fox, Gaussian Development Version, Revision J.19, 2020, Gaussian, Inc., Wallingford CT, 2020 Search PubMed.
- GauOpen, https://gaussian.com/interfacing/, accessed 20 Feb. 2025.
- J. Harris, Phys. Rev. B:Condens. Matter Mater. Phys., 1985, 31, 1770–1779 CrossRef CAS PubMed.
- S. Lehtola, J. Chem. Theory Comput., 2019, 15, 1593–1604 CrossRef CAS PubMed.
- S. Grimme, M. Müller and A. Hansen, J. Chem. Phys., 2023, 158, 124111 CrossRef CAS PubMed.
- L. Zhang, P. Mazzeo, M. Nottoli, E. Cignoni, L. Cupellini and B. Stamm, Replication Data for: A symmetry-preserving and transferable representation for learning the Kohn-Sham density matrix, 2026, DOI:10.5281/zenodo.19110103.
| This journal is © The Royal Society of Chemistry 2026 |