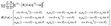Open Access Article
Open Access ArticleFiberForge: enabling high-throughput simulations of the mechanical properties of helical fibrils
Kieran
Nehil-Puleo
a and
Zhongyue John
Yang
 *bcdef
*bcdef
aInterdisciplinary Material Science Program, Vanderbilt University, 2301 Vanderbilt Pl, Nashville, TN 37235, USA
bDepartment of Chemistry, Vanderbilt University, Nashville, TN 37235-1826, USA
cDepartment of Chemical and Biomolecular Engineering, Vanderbilt University, 2301 Vanderbilt Place PMB 351826, Nashville, TN 37235-1826, USA
dCenter for Structural Biology, Vanderbilt University, Nashville, TN 37235, USA
eVanderbilt Institute of Chemical Biology, Vanderbilt University, Nashville, TN 37235, USA
fData Science Institute, Vanderbilt University, Nashville, TN 37235, USA
First published on 2nd February 2026
Abstract
The mechanical properties of amyloid-based materials are governed by fibril geometry, sequence, and polymorphism, yet systematic exploration of this vast design space has been limited by the lack of high-throughput modeling tools. Here we present FiberForge, an open-source workflow that automates construction of amyloid protofibrils, streamlines high-throughput simulations of amyloid deformation, and analyzes fibril trajectories to estimate mechanical properties and fracture mechanisms. Using 374 full-length amyloid crystal structures from the Protein Data Bank, FiberForge rebuilds fibrils with a mean per-chain RMSD of 1.7 Å (median 2.2 Å), demonstrating accurate structural recovery across wide sequence (18–420 aa) and symmetry ranges. Extensive SMD benchmarking on Aβ(1–42) (2MXU) yields a mean rupture force of 1.534 ± 0.164 nN from 232 replicas; bootstraping analysis shows that three replicas suffice for converged elastic-modulus and strength estimates. High-throughput screening of the amyloid fiber dataset produces elastic moduli of 0.2–20 GPa and ultimate tensile strengths of 0.1–1 GPa. Comparison with four AFM-characterized systems shows agreement within an order of magnitude, underscoring the method's predictive capability. FiberForge's screening results also enable larger-scale sequence–structure–property analysis, revealing that mechanical behavior is correlated with molecular assembly geometry, especially hydrogen-bond density. While earlier work suggested the relevance of these features for particular systems, our results demonstrate their importance across diverse fibril architectures. FiberForge thus provides an end-to-end platform for molecular modeling and design of amyloid materials, enabling physics-based identification of sequences and polymorphs with targeted mechanical behavior.
1 Introduction
The mechanical properties of amyloid fibers are remarkable.1 They exhibit mechanical properties comparable to other protein materials such as microtubules,2 actin filaments,3 and even spider silk.4 They possess substantial tensile strength, rivaling that of steel, due to hydrogen bonding between β-sheet structures along their length. Amyloid fibers are stiff with a tensile modulus ranging from 2 to 20 GPa.5 In addition to their impressive mechanical characteristics, amyloid fibers are typically more biocompatible due to their biosynthetic nature; this makes them particularly attractive for biomedical applications. Due to these extraordinary material properties, amyloid fibers have been the focus of numerous biomedical engineering efforts, including hydrogels6 for tissue engineering, carbon capture7 materials, drug delivery,8 and so on. When these fibers form larger networks, such as gels or plaques, the properties of the network emerge from the collective behavior of the individual fibers. For this reason, understanding the behavior of individual fibrils is of fundamental importance to understanding the properties of the collective material. Mechanical properties of amyloid fibrils, such as stiffness and elasticity, can be characterized experimentally using atomic force microscopy (AFM). Computationally, steered molecular dynamics (SMD) offers atomic-level insight into deformation mechanisms by simulating the effects of applied forces. Together, these methods reveal the molecular interactions that govern amyloid stability and structural organization.Despite the successful application of amyloids, the design of amyloid materials still faces several key challenges: the selection of the optimal fiber from the vast space of possible amyloid-forming proteins, prediction of the critical fiber length for the change of the fracture mechanism, differing properties exhibited by structural polymorphs, and lack of a curated dataset of amyloid structures relevant for mechanics investigation. On the issue of sequence selection, prior studies have shown that many proteins, even those not conventionally associated with amyloids, are capable of converting to amyloid fibers at high concentration under destabilizing conditions.9 The richness of the sequence space of possible amyloids necessitates a high-throughput (HTP), automated fiber construction, and simulation framework. In terms of structural polymorphisms, there can be a huge difference in properties between aggregate polymorphic structures for the same protein sequence.10 The sensitivity of mechanical properties to the underlying polymorphic structure highlights the importance of systematically sampling and characterizing different structural polymorphs. On the issue of fracture mechanism, insights from SMD modeling have revealed evidence of length-scale-dependent fracture mechanisms (stick-slip) that arise in both amyloids11 and silk crystals.12 As suggested by these studies, length-scale dependent mechanisms may be attributed to the fact that the fracture mechanism of a β-sheet-rich fibril is governed by the cooperative rupture of hydrogen bonds that stabilize the fibril structure. To investigate further cooperative rupture mechanisms arising in these materials, accurate construction of variable length amyloid structures is needed. Finally, we need a database that captures amyloid structural variations. Several amyloid databases have been curated, including AmyPro, AmyloGraph, and StAmP-DB.13–16 Although these databases are extremely helpful for the community interested in the pathological aspect of amyloids, they do not capture sufficient structural differences that are relevant for the design of amyloids for their mechanical applications.
One methodology to navigate the multi-factor design landscape of amyloids is high-throughput computational physics-guided protein design.17 High-throughput computational design of amyloid materials requires a framework for the autonomous building of fibril structures, the initialization of simulation systems, and the analysis of simulation results. Unfortunately, prior software focused on amyloid properties covers a broad range of applications not specifically focused on this challenge; these software include experimental image analysis of fibril microscopy,18de novo aggregate structure prediction,19,20 β-serpentine fibril structure prediction,21 and numerous examples of amylogenic region prediction.22–25
In this work, we seek to address the issues mentioned above by creating a software, FiberForge, that automates the construction of amyloid protofibrils, streamlines the deployment of high-throughput simulations of amyloid deformation, and automatically analyzes the trajectory of amyloid fibril deformation to estimate mechanical properties and fracture mechanisms. FibrilForge enables a deeper understanding of the structure–function relationships that dictate the mechanical properties of amyloid-based materials, providing critical insights for fundamental materials science. Moreover, it facilitates the in silico design and screening of amyloid mutants with tailored mechanical behaviors, accelerating the development of bioinspired materials for applications in nanotechnology, biomedical devices, and responsive materials.
1.1 Design and implementation
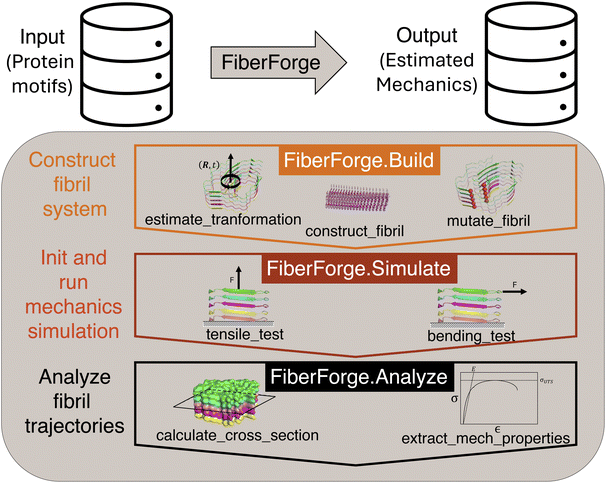
To address our goal of creating software to enable HTP computational design of amyloids, we need a data-structure that is as condensed as possible, but would still accurately describe the 3D geometry of amyloid protein assemblies. We created this data-structure by utilizing the underlying symmetry of amyloid structure: helical symmetry. This representation can describe amyloid fibrils using a 3-tuple: the growth axis vector, translation scalar, and rotation scalar. With this data structure, we were able to construct amyloid fibrils and molecular systems with high fidelity (See bottom left of Fig. 2 for depiction of data structure).After designing our foundational data-structure for the description of amyloids, we built accompanying modules and functions to utilize this structure to perform mechanical property modeling tasks. Specifically, we designed our software suite to have 3 modules: Build, Simulate, and Analyze (See Fig. 1 for depiction of FiberForge). The Build module consists of functions that enable the geometrical characterization of structures containing helical symmetry, the construction of assemblies based on their helical geometrical parameters and sequence mutation information, and finally the construction of a fully solvated amyloid fiber for simulation (see sample code in SI, Fig. S3). In our software, we are able to construct amyloids of different lengths by first learning the symmetry functions from multi-chain fibrils and then applying these symmetries to obtain the desired length fibril. The Simulate module consists of a tensile testing function and a bending testing function. These mechanics testing functions create an SMD protocol to conduct the application of an external force which depends on the testing conditions specified (see sample code in SI, Fig. S4). The Analyze module reads the output trajectories produced from the Simulate module and calculates mechanical properties of interest, such as elastic/bending modulus (see sample code in SI, Fig. S5). Together, these modules enable the construction of end-to-end workflows for fracture mechanics simulations of diverse amyloid structures.
2 Results and discussion
2.1 Amyloid dataset
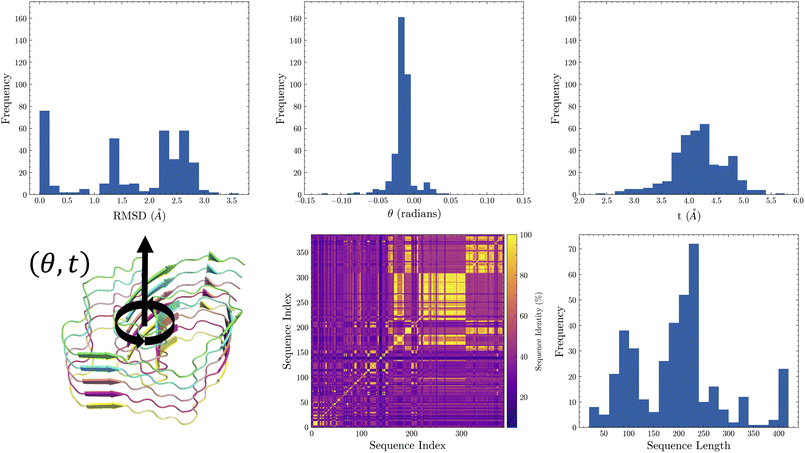
As a prerequisite for building and testing FiberForge, we collected and curated a dataset of experimental amyloid fiber structures. We searched the PDB for entries associated with the keyword “fiber” (performed on February 4th 2024) and then selected all entries that were full sequence fibers, according to our visual inspection of the 3D structure. The compiled dataset consists of 374 structures, with sequence lengths ranging from a minimum of 18 amino acids (aa) to a maximum of 420 aa. The mean sequence length is 197 aa, with a median of 207 aa and a standard deviation of 88 aa. The resolution of the structures has a mean of 3.1 Å, a median of 3.1 Å, and a standard deviation of 0.5 Å.2.2 Benchmarking of reconstruction from helical parameters
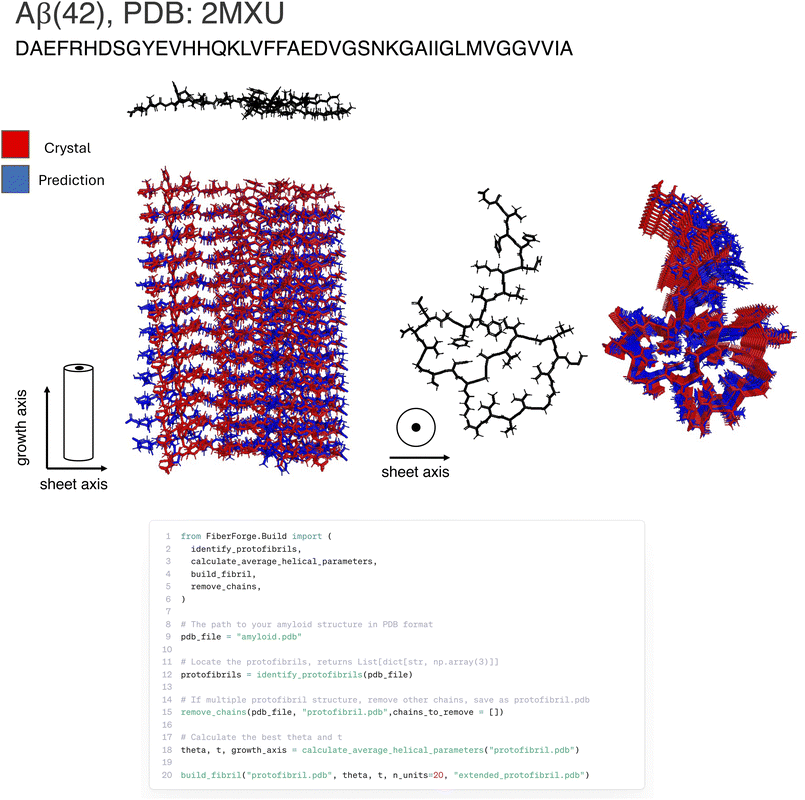
Accurate fibril construction affects the reliability of estimated mechanics from simulation. To validate our construction method, we applied our method to each structure in our curated dataset, rebuilding each structure using only a seed chain and the helical parameters. After fibril construction, we calculated the geometric deviation of our reconstructed fibril structures from their respective experimental structures (See Fig. 3, FiberForge API for this process, and summary reconstruction histogram for our curated dataset). Reconstruction statistics are as follows: mean RMSD per chain: 1.7 Å, median RMSD per chain 2.2 Å, standard deviation of RMSD per chain 1.4 Å. As such, the structural models produced by FiberForge.Build shows high consistency with experimental structures for amyloids of various lengths and across a wide range of geometric constructs. A representative example of this group is a hIAPP polymorph (7YL7) which has a RMSD of 0.07 Å (see Fig. S1 for a depiction of this fibril). This is believed to be attributed to intra-chain rotation occurring between stacking of successive chains in an amyloid, particularly for residues occurring along the outer region, which has not be accounted for in our current building model of the amyloid symmetry. For some fibril structures our method deviated slightly from experimental structures. The major outlier is 6NZN, which we do not show in Fig. 2. This amyloid has an anti-parallel stacking occurring between chains in a fibril. The current version of our helical symmetry representation does not account for this behavior, but could be extended to handle this type of structure in the next version. Excluding the outlier, the largest RMSD per chain is 3.6 Å with the structure 6RTB (See Fig. S2 for depiction of reconstruction).2.3 Benchmarking of steered molecular dynamics
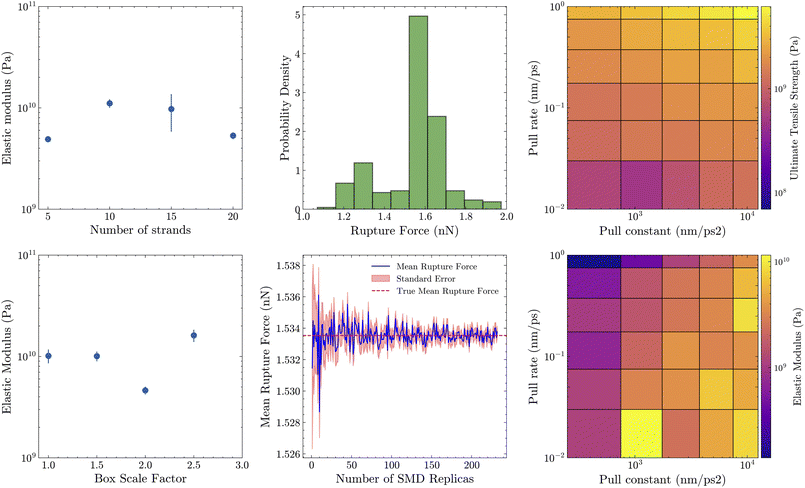
Essential to validating the reliability of mechanical property prediction is the expected variability of values predicted from SMD. Previous SMD studies of amyloid fracture involved very few replicates, as little as a single SMD simulation. This is likely because the primary focus was to qualitatively investigate fracture mechanism rather than predicting specific mechanical properties.11,26 Here, we assessed how much rupture-force modeling results fluctuate across SMD simulations to judge their accuracy and reproducibility (Fig. 4). Specifically, we carried out a case study on the Aβ(42) amyloid (PDB: 2MXU), the same structure used in our construction benchmark, to quantify this inherent variability. We took the same equilibrated system and the same pulling process under different random seeds (a total of 232 replicates). The results show that the mean rupture force is 1.530 nN, the standard deviation of the rupture force is 0.164 nN, and the standard error of the rupture force is 0.0108 nN. Across the entire replica dataset, the mean value fluctuates between 1.526 nN and 1.538 nN—a total spread of only 0.012 nN (12.0 pN), or one percent of the mean. This means that, even though the curve looks “noisy”, the absolute variation is small. As such, a handful of trajectories already gives a reasonable estimate: after just 1–3 replicas the mean typically sits within a few ×10−3 nN of the large-sampling-averaged mean (red dashed line). Thus, for qualitative comparisons or screening studies, running only a couple of replicas is usually sufficient. For applications that demand quantitative convergence to the second decimal place (±0.010 nN, i.e., ±5 pN), the shaded standard-error band shows that roughly 30–40 independent replicas are needed. Beyond this point the residual uncertainty falls below 0.005 nN, ensuring the mean is stable to 0.010 nN precision.Next, we quantified how simulation settings affect the mechanical constants extracted from SMD pulling (Fig. 4). Raising the spring constant from 5 × 102 → 1 × 104 nm ps−2 and the pulling rate from 0.01 → 1 nm ps−1 drives a clear, monotonic stiffening: the ultimate tensile strength climbs from around 5.3 × 109 Pa to 1.1 × 1010 Pa and the elastic modulus from around 3.0 × 109 Pa to 9.5 × 109 Pa, matching the behavior suggested by prior theory of SMD.27 In contrast, box size and number of strands can also perturb the calculated elastic modulus, but no significant change was observed on the scale of the estimated elastic modulus.
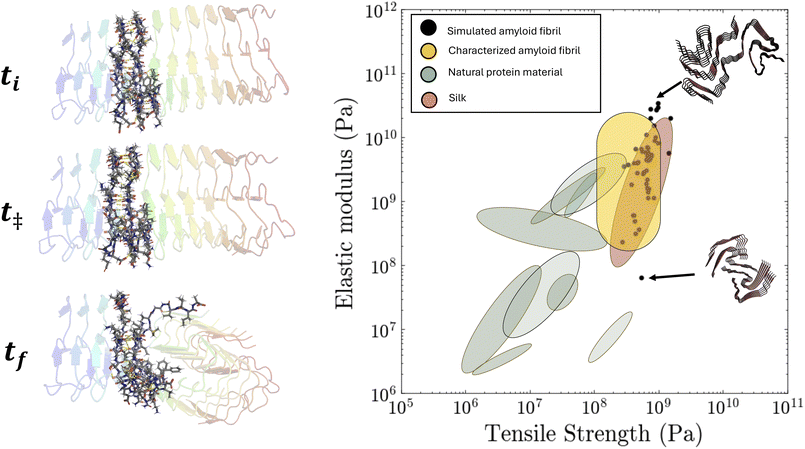
Lastly, we compared our estimated mechanical properties to those obtained from experimental methods (see Fig. 5 for results across the entire curated dataset). Our simulated values fall within the expected range for amyloid fibrils (roughly 109 to 1011 Pascals for elastic modulus and 108 to 109 Pascals for tensile strength).5 Furthermore, we compared our estimated properties with a single experimental study (see Table 1). Although our results showed reasonable agreement with experimental values, for example, for the insulin amyloid with PDB ID 8SBD, we obtained 4.7 GPa for the simulated elastic modulus compared to the reported experimental value of 3.2 GPa, our simulated estimates were consistently higher. This discrepancy is expected: theoretical predictions and atomistic simulations typically produce substantially larger elastic moduli than experiments because the deformation rates used in SMD are several orders of magnitude higher than those employed in AFM and related experimental techniques.28 Additionally, experimental measurements exhibit substantial variability across studies due to differences in fibril preparation, mechanical testing protocols, deformation rates, and solution conditions. For these reasons, and to mitigate the variability inherent in experimental datasets, comparison with experiment was limited to a single group. To test the generality of this approach, future work will focus on expanding comparisons across multiple experimental systems, despite the challenge posed by the relatively small number (on the order of 10) of available single-fibril mechanical measurements and their substantial inter-study heterogeneity.
| Common name | PDB entry | E exp ± SE | E sim ± SE |
|---|---|---|---|
| Insulin | 8SBD | 3.2 ± 0.6 | 4.7 ± 0.2 |
| τ-protein | 7YPG | 3.4 ± 0.7 | 11.3 ± 3 |
| β-lactoglobulin | 6GK3 | 3.7 ± 0.8 | 8.6 ± 1 |
| Aβ(1–42) | 2MXU | 3.2 ± 0.8 | 4.8 ± 0.9 |
In addition to the benchmarks, we should note the intrinsic stochasticity underlying the rupture force measurement from single-molecule AFM experiments. Early physical theories were developed to describe rupture force distributions in single hydrogen bonded systems29 (Bell–Evans model), these theories were later extended to protein unfolding, i.e. multi-hydrogen bonded system, then eventually the connection between AFM and SMD techniques.27,28 These theories were applied to describe amyloid fibril rupture.11 Essentially, amyloids are held together by a network of hydrogen bonds occurring between proteins which result in induced force being distributed across a network of bonds. The rupture of this network is a non-equilibrium, stochastic process; as such, the outcome of SMD mechanical property estimates inherently depends on many random factors, making statistical analysis essential.
2.4 Virtual screening results
We applied our FiberForge workflow to model the mechanical properties of 72 PDB-resolved amyloid fibrils spanning 18 peptide sequences to probe how sequence and geometry shape mechanical performance and to demonstrate the pipeline's throughput. For each structure, we prepared the workflow code to parse the helical rise and twist, build a 40 nm-long, 6–12 nm-wide atomistic model of about 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 atoms, equilibrate it, and ran a 10 ns constant-velocity pull at 0.1 nm ps−1, with post-processing handled automatically; the full set finished in 36 h on a single GPU node. The resulting Ashby plot (Fig. 5) contains 190 simulated data points: elastic moduli range from 7.0 × 107 to 1.1 × 1010 Pa (median 3.2 × 109 Pa) and ultimate tensile strengths from 9.0 × 107 to 1.0 × 1010 Pa (median 7.8 × 108 Pa). These values align with experimentally reported amyloid properties (the Young's modulus: 108–1010 Pa; tensile strength: 108–109 Pa) and sit well above those of typical structural proteins such as collagen or keratin. Wider multiprotofilament fibrils occupy the upper-right of the plot, while slimmer two-strand assemblies fall lower, reflecting the expected dependence on cross-section.5,30,31 The match to experiment and the ability to complete dozens of simulations overnight support both the reliability and scalability of our software for large-scale mapping of sequence–structure–property relationships in amyloids.
000 atoms, equilibrate it, and ran a 10 ns constant-velocity pull at 0.1 nm ps−1, with post-processing handled automatically; the full set finished in 36 h on a single GPU node. The resulting Ashby plot (Fig. 5) contains 190 simulated data points: elastic moduli range from 7.0 × 107 to 1.1 × 1010 Pa (median 3.2 × 109 Pa) and ultimate tensile strengths from 9.0 × 107 to 1.0 × 1010 Pa (median 7.8 × 108 Pa). These values align with experimentally reported amyloid properties (the Young's modulus: 108–1010 Pa; tensile strength: 108–109 Pa) and sit well above those of typical structural proteins such as collagen or keratin. Wider multiprotofilament fibrils occupy the upper-right of the plot, while slimmer two-strand assemblies fall lower, reflecting the expected dependence on cross-section.5,30,31 The match to experiment and the ability to complete dozens of simulations overnight support both the reliability and scalability of our software for large-scale mapping of sequence–structure–property relationships in amyloids.
2.5 Sequence–structure–property relations
To identify the key structural and sequence-derived determinants of amyloid mechanical performance, we examined correlations between the geometric descriptors extracted from PDB structures, sequence hydrophobicity metrics, and the mechanical properties computed via SMD (see SI, Fig. S6). Ultimate tensile strength (UTS) exhibited a moderate positive correlation (0.62) with elastic modulus, consistent with the expected coupling between stiffness and failure load in stiff materials. Elastic modulus showed moderate correlation with Hydrogen-bond density (HBD) and sequence hydrophobicity, (0.44 and 0.26, respectively), suggesting a dependence of geometric and sequence properties on elasticity. UTS showed a strong correlation with HBD (0.68) and a moderate negative correlation with the translation distance between chains (−0.23), suggesting the HBD greatly affects the maximum force that the amyloid can withstand before fracturing. Reducing the translation distance between chains also increase the UTS, which may be a causal effect or simply a result of stronger interactions between chains.Notably, HBD showed a strong positive correlation with both elastic modulus and UTS, reinforcing earlier theories that interchain HBD is the primary factor affecting deformation resistance. Though earlier studies demonstrated this behavior in specific amyloid systems,5,11,12 our results imply that it is a general design principle of planar amyloid systems. Helical twist and rise generally showed weak correlations with mechanical properties, suggesting that local geometric offsets alone do not strongly dictate mechanical response. These two properties showed moderately strong negative correlation with each other, suggesting that a greater degree of twist in an amyloid reduces the spacing between between chains. Sequence mean hydrophobicity displayed modest (0.26) but statistically significant correlations with elastic modulus, indicating that hydrophobic packing may stabilize inter-chain interfaces and thereby alter load-bearing capacity.
Having established key correlations among the structural and sequence descriptors, we next asked whether these features could predict mechanical behavior. To evaluate their predictive power, we performed an automated screening of machine-learning algorithms using AutoML,32 testing dozens of regression models across multiple random train–test splits. The random forest regressor consistently achieves good performance and demonstrates strong robustness to dataset resampling. Based on these results, we selected the random forest as the base model for all subsequent analyses, and assessed how different feature groups contribute to predictive accuracy for elastic modulus and UTS (see SI for the parity plots of cross-validation and hold-out test, Fig. S7).
For elastic modulus, feature sets composed solely of helical parameters (magnitude of translation and rotation), sequence–distance metrics (HBD, cross-sectional area, and H-bond count), hydrophobicity metrics (fraction of hydrophobic residues, mean residue hydrophobicity, standard deviation of residue hydrophobicity), or evolutionary-sequence metrics yield near zero or even negative R2, indicating that no single class of descriptors is sufficient to predict elasticity variations. Notably, inclusion of evolutionary sequence embeddings33 alone decreases predictive accuracy relative to the geometric features, suggesting that sequence evolutionary information does not meaningfully contribute to the elastic behavior of planar amyloids. The strongest predictive performance is achieved when geometric descriptors were combined with hydrophobicity features (R2 = 0.377 on the holdout set), highlighting the interplay between amyloid structure and amino acid properties on elasticity. For UTS, hydrophobicity or helical descriptors alone produced weak predictive performance, and ESM embeddings again performs poorly in isolation. The highest R2 values are achieved using the geometric, hydrophobicity, and ESM feature set (R2 = 0.516 on the holdout set). These results emphasize that amyloid UTS is influenced by the geometry of interchain organization, particularly features such as HBD, and evolutionary sequence information. However, the gap in R2 values between cross-validation and the hold-out test suggests that the model may be overfitting due to the limited size of the training dataset. We anticipate that this issue will diminish as additional molecular simulation data become available.
Overall, both properties are driven predominantly by geometric organization—most notably HBD, which emerges as the single most informative descriptor for elastic modulus and a significant predictor for UTS. Elastic modulus depends largely on HBD and hydrophobicity with only minor gains from any additional features and little to no improvement from evolutionary embeddings. By contrast, UTS also relied heavily on HBD but benefited substantially from the inclusion of evolutionary sequence information, indicating that tensile strength is more sensitive than elasticity to sequence-encoded nuances once the primary geometric scaffold (HBD) is accounted for. This establishes a path toward rapid, simulation-free prediction and optimization of amyloid-based materials, helping to define general design principles and highlighting the promise of integrating interpretable structural features with machine learning for next-generation biomolecular material design.
3 Conclusion
We have developed FiberForge, an end-to-end, high-throughput software suite that automates the construction, simulation, and analysis of amyloid protofibrils for mechanical characterization. By encoding fibril geometry with a helical-symmetry tuple, FiberForge.Build rebuilds experimental structures with good accuracy (median RMSD 2.2 Å across 374 PDB fibrils) and readily generates user-defined lengths and sequence variants. FiberForge.Simulate deploys tailored SMD tensile and bending protocols, while FiberForge.Analyze extracts mechanical observables—elastic modulus, rupture force, and ultimate tensile strength—directly from the resulting trajectories.Benchmarking shows that (i) reconstruction fidelity remains high across diverse polymorphs, (ii) rupture-force calculations converge within ±0.010 nN after 30–40 replicas, and (iii) the expected rate- and spring-constant-dependent stiffening follows classical non-equilibrium theory. Using FiberForge, we screened 72 experimentally resolved fibrils (18 sequences), completing 190 tensile tests in 36 h on a single GPU node. The simulated elastic moduli and tensile strengths overlap closely with single-fibril AFM measurements.
Our structure–property analyses reveal that amyloid mechanics arise from the coupled effects of molecular assembly geometry and residue-level interactions. Across fibrils, HBD emerges as the strongest determinant of elasticity and a strongly correlated with UTS. Evolutionary sequence information does not appear to aid in the prediction of elasticity but does appear to aid in the prediction of UTS. This establishes a route toward rapid, simulation-free property prediction and materials optimization.
While FiberForge provides a robust computational platform for mapping sequence–structure–property relationships in amyloids, several limitations remain. The current framework is primarily optimized for parallel, helical-symmetry protofibrils and does not yet capture anti-parallel, mixed-symmetry, or irregular architectures. Moreover, it is best suited for protofibril-scale fracture characterization, whereas the multi-stranded fibril bundles observed in mature fibers remain beyond its present modeling scope. From a simulation standpoint, the accessible timescales of classical molecular dynamics inherently limit the treatment of rate-dependent mechanical behavior.
Future developments will address these limitations by (i) extending FiberForge.Build to support anti-parallel, mixed-symmetry, and bundled fibril assemblies, (ii) coupling coarse-grained or multi-scale fracture models to capture longer timescales and larger systems, and (iii) integrating automated mutation, ranking, and machine-learning–based property prediction modules to enable high-throughput screening and accelerated design.
Together with our structure–property analysis—showing that mechanical behavior emerges from the interplay of supramolecular geometry and residue-level chemistry, and that features such as hydrogen-bond density, while important in specific amyloid systems, are not universally predictive across all fibrils, FiberForge provides a unified computational foundation for disentangling the multiscale determinants of amyloid mechanics.
FiberForge provides a computational platform for mapping sequence–structure–property relationships in amyloids and accelerates the in silico discovery of bio-inspired, high-performance protein materials. Combined with emerging single-fibril experimental techniques, FiberForge promises to bridge atomic-scale insight and macroscopic material design, paving the way for next-generation biomedical and sustainable engineering applications built upon amyloid architectures.
4 Methods
4.1 Molecular modeling protocol
The general procedure for the simulations involves the following steps: First, specify the PDB ID of the starting structure to ensure the correct protein is selected. For example, 2MXU of Aβ42 was used in our demo. Next, place the corresponding PDB file in the designated directory. The file is stripped of non-protein molecules. The (θ, t, a) parameters are then estimated (see parameter estimation section for details). After the parameters are estimated, the system is constructed (see system construction section for details).In SMD, an external force Fext(t) is applied to the system according to the equation:
| Fext(t) = −k(x(t) − x0(t)) |
4.2 Sequence–structure–property relations
To assess sequence contributions to mechanics, we calculated Kyte–Doolittle hydrophobicity38 values for each residue in the extracted sequence. Mean, standard deviation, and the fraction of residues with positive hydrophobicity score were used as sequence-level hydrophobicity features. To examine whether protein sequence information, beyond simple hydrophobicity metrics, improves prediction of fibril mechanical performance, we extracted primary sequences for each simulated amyloid structure and embedded them using a state-of-the-art protein language model. First, amino-acid sequences were parsed from the first chain of each PDB file using BioPython. For each valid fibril, we also retrieved precomputed structural descriptors including inter-chain hydrogen-bond count, cross-sectional area, helical rotation and translation offsets, elastic modulus, and ultimate tensile strength (UTS). To capture higher-order, nonlocal sequence information, we computed deep sequence embeddings using the ESM-2 protein transformer model (esm2_t33_650M_UR50D).33 Each sequence was batch-converted into token format, passed through the pretrained model, and the 33rd-layer per-residue representations were averaged to obtain a fixed-length embedding vector for each fibril sequence.
Finally, sequence embeddings were paired with inter-chain hydrogen-bond density, cross-sectional area, Kyte–Doolittle hydrophobicity statistics, as well as helical parameters, to construct an extended feature vector representing sequence, structure, and inter-chain interaction characteristics.
Before performing cross-validation analyses, we first identified an appropriate regression model using an automated model-selection procedure. We applied an AutoML workflow32 that screened a broad set of machine-learning algorithms across randomized train–test splits. We performed this operation several times on different train-test partitions to obtain model which we believed was robust to the train-test partitioning. This initial step enabled systematic comparison of dozens of candidate models under repeated resampling, providing a robust estimate of algorithm stability rather than performance on any single data partition. Across all tested regressors, the random forest consistently achieved strong and reproducible performance, demonstrating resilience to dataset reshuffling and minimal sensitivity to outlier splits. Based on these findings, the random forest model was selected as the base estimator for all subsequent analyses.
We trained random forest models under two complementary evaluation schemes. First, we performed 10-fold cross-validation on the full dataset to quantify model robustness and assess the relative predictive value of specific feature subsets. Feature groups included: (i) geometric offsets only, (ii) hydrogen-bonding and cross-sectional metrics, (iii) hydrophobicity features, and (iv) full sequence embeddings. Both features and targets were standardized within each training fold, and performance was assessed via the coefficient of determination R2. Second, to evaluate generalization, we performed a strict 80/20 train–holdout split, retrained models exclusively on the training portion (including normalization), and computed hold-out R2 on unseen fibrils. For both evaluation schemes, parity plots were generated for each target–feature-set combination to visualize predictive accuracy and systematic bias (See SI, Fig. S7).
4.3 Data structure
Successful methodologies41,42 to reduce the structural complexity of protein aggregates involve understanding and utilizing the symmetries that govern their structure. Prior work has focused on exhaustively characterizing the symmetries that can exist in amyloid structures.43 The symmetries that exist within a protofibril can be characterized by a translational (distance between proteins) and rotational (parallel vs. anti-parallel and equifacial vs. antifacial) symmetries which characterize the helical structure. In a structure with helical symmetry, a rotation around an axis followed by a translation along that axis leaves the structure invariant. If Rθ is a rotation by angle θ and Td is a translation by distance d along the same axis, the symmetry operation can be represented as a combined transformation RθTd. More formally, the helical group H is defined: | (1) |
The “classical” pathogenic amyloid structure is characterized by helical symmetry (See eqn (1) for formal description). The rotational symmetry about the growth axis can be represented by a single scalar  , the translational symmetry by a translation scalar
, the translational symmetry by a translation scalar  , and the axis about which these parameters are applied (the growth axis)
, and the axis about which these parameters are applied (the growth axis)  .
.
The tuple (θ, t, a) characterizes the symmetry of the protein sheets that make up the protein fibril. It should be noted that functional amyloids, amyloids characterized by solenoid fibrils such as CsgA, can also be described using helical symmetry. In this case θ is simply 1 i.e., only translational symmetry.
4.4 Core algorithms
We created the function identify_protofibrils to identify groups of protein chains forming protofibrils based on spatial proximity within a given PDB file. The PDB structure was first parsed using PDBParser from Biopython,35 and for each chain, the atomic coordinates were extracted. The center of mass for each chain was then computed by averaging the Cartesian coordinates of all its atoms.
To group chains into protofibrils, a clustering approach based on a predefined distance threshold was applied. Each chain's center of mass was compared against those in existing protofibril clusters. If the distance between a chain's center and any chain already assigned to a protofibril was below the threshold, the chain was incorporated into that protofibril. If no existing protofibril met the distance criterion, a new protofibril cluster was initiated.
This procedure resulted in a list of protofibrils, where each protofibril was represented as a dictionary containing chain identifiers and their corresponding centers of mass. This method facilitated the automated identification of spatially grouped chains for further structural analysis.
For a formal description, let the protein structure contain C chains, each with a center of mass  ,
,  as the set of protofibrils where each Pj is a subset of chains, dij = ‖ri − rj‖ as the Euclidean distance between chain centers ri and rj, and δ as the distance threshold for protofibril membership.
as the set of protofibrils where each Pj is a subset of chains, dij = ‖ri − rj‖ as the Euclidean distance between chain centers ri and rj, and δ as the distance threshold for protofibril membership.
The goal is to find a partitioning of chains into protofibrils that minimizes the maximum intra-protofibril distance:
where θ is the rotation angle (in radians), t is the translation along the symmetry axis, a is the axis of symmetry (as a unit vector), rij(1) is the coordinates of atom j in chain 1 of pair i, rij(2) are the coordinates of atom j in chain 2 of pair i, N is the number of chain pairs, Mi is the number of atoms in the i-th chain pair, a = [ax, ay, az], ‖a‖ = 1, and R(θ, a) is the rotation matrix about axis a by angle θ.
| rn = R(θ,a)rn−1 + ta | (2) |
The protofibril structure can be constructed using either the mBuild44 or Pymol45 software packages to enable direct conversion for molecular dynamics simulations.
A key challenge in SMD of amyloids is the large system size required, given the nanometer scale of fibrils. To automate system construction efficiently, we leveraged the amyloid's helical parameters to optimize its orientation in the simulation box. Specifically, we aligned the fibril along its identified growth axis—a 3D vector—placing it in a rectangular box with its longest dimension parallel to the growth direction. Once the fibril is correctly oriented we can place solvent molecules using PackMol.46
The FiberForge package offers two modes of deformation: tensile and shear. Tensile deformation refers to a pulling force applied parallel to the growth axis, while shear deformation involves a pulling force applied perpendicular to the growth axis (see Fig. 1 for a depiction of these mechanical tests).
To initialize these simulations without human input, we utilize pre-defined data structures. Specifically, we used the translational component t of the helical symmetry to orient the fibrils in the simulation box. For tensile tests, the fibrils are aligned parallel to the applied force, and for shear tests, they are aligned perpendicular to the applied force.
Springs are attached to the center of mass of the end chains of the amyloid for a tensile simulation. For a shear simulation restraints are placed on one end of the fibril and a spring is attached to the other end.
SMD simulations are performed using GROMACS.47 Systems are solvated using Packmol. After the fibril is placed in the correct orientation in the simulation box, energy minimization is performed. Following minimization, the system undergoes equilibration under an NVT ensemble for 50 ps, followed by further equilibration under an NPT ensemble for another 50 ps. Finally, SMD pulling is performed for 500 ps.
Parameters affecting SMD simulations, such as the pull rate and the force constant, are determined manually. The optimal pull rate is estimated to be 0.01 nm/ps and the optimal force constant is 1000 kJ (mol−1 nm−2). Temperature coupling is performed using the Nose–Hoover extended ensemble under standard conditions, and pressure coupling is performed using the Parrinello–Rahman extended ensemble.
The elastic modulus is a measure of a material's resistance to elastic deformation and is generally defined as
Similarly, the UTS is defined as
| UTS = max(σ) |
that is, the maximum stress a material can withstand before failure during a tensile test.
These properties are estimated by analyzing the geometry of the protofibril and the SMD trajectory. The Fext and the corresponding deformation distance produced from the SMD simulations are extracted from the output files. To reduce the noise we apply a moving average filter to the stress–strain curve. To further process the simulation data, automated routines were implemented to smooth and fit the stress–strain relationships using cubic spline interpolation. Next, we estimate the linear elastic region of the stress–strain curve by calculating the second derivative of the stress with respect to strain and then extracting the inflections points. The elastic modulus E is then obtained as the slope of the fitted curve within the linear elastic region, which is identified between the initial point and the first inflection point of the second derivative of the spline. This inflection point also defines the yield point, corresponding to the transition between elastic and plastic deformation. To estimate the tensile and shear strength, the maximum Fext obtained during the SMD simulation is used.
Author contributions
KNP and ZJY designed the study. KNP implemented the software, performed SMD simulations, analyzed and gathered computational data. KNP wrote the manuscript, and KNP and ZJY revised the manuscript.Conflicts of interest
The authors declare no competing financial interest.Data availability
Archived code for FiberForge can be found on GitHub https://github.com/ChemBioHTP/FiberForge/tree/v1.0.0 with DOI; https://doi.org/10.5281/zenodo.18202973. The most up to date version of FiberForge can be found at https://github.com/ChemBioHTP/FiberForge. All additional code and data used to produce figures can be found on Zenodo with DOI: https://doi.org/10.5281/zenodo.17782766.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5dd00307e.
Acknowledgements
This research was supported by the startup grant from Vanderbilt University. Z. J. Yang thanks the sponsorship from Rosetta Commons Seed Grant Award and the Dean's Faculty Fellowship in the College of Arts and Science at Vanderbilt. K. Nehil-Puleo thanks the NSF Graduate Research Fellowship Program for sponsorship.Notes and references
- M. Schleeger, T. Deckert-Gaudig, V. Deckert, K. P. Velikov, G. Koenderink and M. Bonn, et al. , Polymer, 2013, 54, 2473–2488 CrossRef CAS.
- M. Kikumoto, M. Kurachi, V. Tosa and H. Tashiro, Biophys. J., 2006, 90, 1687–1696 CrossRef CAS PubMed.
- J. M. Ferrer, H. Lee, J. Chen, B. Pelz, F. Nakamura, R. D. Kamm and M. J. Lang, Proceedings of the National Academy of Sciences, 2008, 105, 9221–9226 Search PubMed.
- F. Vollrath and D. P. Knight, Nature, 2001, 410, 541–548 Search PubMed.
- T. P. Knowles and M. J. Buehler, Nat. Nanotechnol., 2011, 6, 469–479 Search PubMed.
- G. Bhak, S. Lee, J. W. Park, S. Cho and S. R. Paik, Biomaterials, 2010, 31, 5986–5995 Search PubMed.
- D. Li, H. Furukawa, H. Deng, C. Liu, O. M. Yaghi and D. S. Eisenberg, Proceedings of the National Academy of Sciences, 2014, 111, 191–196 Search PubMed.
- S. K. Maji, D. Schubert, C. Rivier, S. Lee, J. E. Rivier and R. Riek, PLoS Biol., 2008, 6, e17 CrossRef PubMed.
- L. Goldschmidt, P. K. Teng, R. Riek and D. Eisenberg, Proceedings of the National Academy of Sciences, 2010, 107, 3487–3492 Search PubMed.
- G. Yoon, M. Lee, J. I. Kim, S. Na and K. Eom, PLoS One, 2014, 9, e88502 CrossRef PubMed.
- B. Choi, G. Yoon, S. W. Lee and K. Eom, Phys. Chem. Chem. Phys., 2015, 17, 1379–1389 Search PubMed.
- S. Keten, Z. Xu, B. Ihle and M. J. Buehler, Nat. Mater., 2010, 9, 359–367 CrossRef CAS PubMed.
- M. Varadi, G. De Baets, W. F. Vranken, P. Tompa and R. Pancsa, Nucleic Acids Res., 2018, 46, D387–D392 CrossRef CAS PubMed.
- N. Louros, K. Konstantoulea, M. De Vleeschouwer, M. Ramakers, J. Schymkowitz and F. Rousseau, Nucleic Acids Res., 2020, 48, D389–D393 Search PubMed.
- M. Burdukiewicz, D. Rafacz, A. Barbach, K. Hubicka, L. Bkakala, A. Lassota, J. Stecko, N. Szymanska, J. W. Wojciechowski and D. Kozakiewicz, et al. , Nucleic Acids Res., 2023, 51, D352–D357 Search PubMed.
- N. Louros, R. Van Der Kant, J. Schymkowitz and F. Rousseau, Bioinformatics, 2022, 38, 2636–2638 CrossRef CAS PubMed.
- Q. Shao, Y. Jiang and Z. J. Yang, J. Chem. Inf. Model., 2022, 62, 647–655 CrossRef CAS PubMed.
- I. Usov and R. Mezzenga, Macromolecules, 2015, 48, 1269–1280 Search PubMed.
- I. André, P. Bradley, C. Wang and D. Baker, Proceedings of the National Academy of Sciences, 2007, 104, 17656–17661 Search PubMed.
- C. W. Wood, M. Bruning, A. A. Ibarra, G. J. Bartlett, A. R. Thomson, R. B. Sessions, R. L. Brady and D. N. Woolfson, Bioinformatics, 2014, 30, 3029–3035 CrossRef CAS PubMed.
- S. A. Bondarev, O. V. Bondareva, G. A. Zhouravleva and A. V. Kajava, Bioinformatics, 2018, 34, 599–608 CrossRef CAS PubMed.
- S. Maurer-Stroh, M. Debulpaep, N. Kuemmerer, M. L. De La Paz, I. C. Martins, J. Reumers, K. L. Morris, A. Copland, L. Serpell and L. Serrano, et al. , Nat. Methods, 2010, 7, 237–242 CrossRef CAS PubMed.
- S. O. Garbuzynskiy, M. Y. Lobanov and O. V. Galzitskaya, Bioinformatics, 2010, 26, 326–332 CrossRef CAS PubMed.
- A. W. Bryan Jr, M. Menke, L. J. Cowen, S. L. Lindquist and B. Berger, PLoS Comput. Biol., 2009, 5, e1000333 CrossRef.
- C. Kim, J. Choi, S. J. Lee, W. J. Welsh and S. Yoon, Nucleic Acids Res., 2009, 37, W469–W473 CrossRef CAS PubMed.
- R. Paparcone and M. J. Buehler, Biomaterials, 2011, 32, 3367–3374 CrossRef CAS PubMed.
- H. Lu and K. Schulten, Proteins: Struct., Funct., Bioinf., 1999, 35, 453–463 CrossRef CAS.
- B. Isralewitz, M. Gao and K. Schulten, Curr. Opin. Struct. Biol., 2001, 11, 224–230 CrossRef CAS PubMed.
- E. Evans and K. Ritchie, Biophys. J., 1997, 72, 1541–1555 CrossRef CAS PubMed.
- U. G. Wegst and M. F. Ashby, Philos. Mag., 2004, 84, 2167–2186 CrossRef CAS.
- M. F. Ashby, L. Gibson, U. Wegst and R. Olive, Proceedings of the Royal Society of London. Series A: Mathematical and Physical Sciences, 1995, 450, 123–140 Search PubMed.
- S. Pandala, LazyPredict, GitHub repository, https://github.com/shankarpandala/lazypredict Search PubMed.
- Z. Lin, H. Akin, R. Rao, B. Hie, Z. Zhu, W. Lu, N. Smetanin, A. dos Santos Costa, M. Fazel-Zarandi, T. Sercu, S. Candidoet al., bioRxiv, 2022, preprint.
- J. Adamcik, C. Lara, I. Usov, J. S. Jeong, F. S. Ruggeri, G. Dietler, H. A. Lashuel, I. W. Hamley and R. Mezzenga, Nanoscale, 2012, 4, 4426–4429 RSC.
- P. J. Cock, T. Antao, J. T. Chang, B. A. Chapman, C. J. Cox, A. Dalke, I. Friedberg, T. Hamelryck, F. Kauff and B. Wilczynski, et al. , Bioinformatics, 2009, 25, 1422 CrossRef CAS PubMed.
- R. T. McGibbon, K. A. Beauchamp, M. P. Harrigan, C. Klein, J. M. Swails, C. X. Hernández, C. R. Schwantes, L.-P. Wang, T. J. Lane and V. S. Pande, Biophys. J., 2015, 109, 1528–1532 CrossRef CAS PubMed.
- A. Shrake and J. A. Rupley, J. Mol. Biol., 1973, 79, 351–371 CrossRef CAS PubMed.
- J. Kyte and R. F. Doolittle, J. Mol. Biol., 1982, 157, 105–132 CrossRef CAS PubMed.
- P. Virtanen, R. Gommers, T. E. Oliphant, M. Haberland, T. Reddy, D. Cournapeau, E. Burovski, P. Peterson, W. Weckesser and J. Bright, et al. , Nat. Methods, 2020, 17, 261–272 CrossRef CAS PubMed.
- M. L. Waskom, J. Open Source Softw., 2021, 6, 3021 CrossRef.
- F. DiMaio, A. Leaver-Fay, P. Bradley, D. Baker and I. André, PLoS One, 2011, 6, e20450 CrossRef CAS PubMed.
- J. L. Watson, D. Juergens, N. R. Bennett, B. L. Trippe, J. Yim, H. E. Eisenach, W. Ahern, A. J. Borst, R. J. Ragotte and L. F. Milles, et al. , Nature, 2023, 620, 1089–1100 CrossRef CAS PubMed.
- D. S. Eisenberg and M. R. Sawaya, Annu. Rev. Biochem., 2017, 86, 69–95 CrossRef CAS PubMed.
- C. Klein, J. Sallai, T. J. Jones, C. R. Iacovella, C. McCabe and P. T. Cummings, Foundations of molecular modeling and simulation, Select papers from FOMMS 2015, 2016, 79–92 Search PubMed.
- W. L. DeLano, et al. , CCP4 Newsl. Protein Crystallogr., 2002, 40, 82–92 Search PubMed.
- L. Martínez, R. Andrade, E. G. Birgin and J. M. Martínez, J. Comput. Chem., 2009, 30, 2157–2164 CrossRef PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1, 19–25 CrossRef.
| This journal is © The Royal Society of Chemistry 2026 |