Open Access Article
Open Access ArticleCreative Commons Attribution 3.0 Unported Licence
Catalytic strategies for post-translational modifications regulating protein higher-order structure and properties
Yuki
Yamanashi
 * and
Motomu
Kanai
* and
Motomu
Kanai
 *
*
Graduate School of Pharmaceutical Sciences, The University of Tokyo, Tokyo 113-0033, Japan. E-mail: yamanashi@mol.f.u-tokyo.ac.jp; kanai@mol.f.u-tokyo.ac.jp
First published on 30th March 2026
Abstract
Proteins, the major functional components of living organisms, undergo post-translational modifications (PTMs) that expand their structural and functional diversity. Recent advances in PTM profiling and functional analysis have revealed that many PTMs act as reversible modulators of protein behavior, operating with residue- and domain-level precision to reshape higher-order structures. Both biotic and abiotic catalyses are emerging means of deciphering and controlling PTMs. In this Tutorial Review, we outline how PTMs influence protein architecture across multiple structural scales and survey catalytic strategies that enable their analysis and manipulation.
Key learning points(1) Post-translational modifications (PTMs) as dynamic chemical switches that expand proteomic diversity.(2) How PTMs reshape protein higher-order structures. (3) Chemical catalysis to decipher or control PTMs. (4) How catalysis can help to decipher PTM-related biology. (5) How catalysis itself can be a drug via PTM regulation. |
1. Introduction
Catalysts accelerate chemical reactions without being consumed. In living organisms, enzymes function as highly evolved catalysts, operating with high precision to drive metabolism, reproduction, and countless cellular processes that sustain homeostasis. Because every biological function ultimately arises from networks of molecules and chemical reactions, catalysis constitutes a fundamental layer of control in cellular systems.Proteins, the primary executors of all molecular phenomena in living organisms, including these catalytic processes, are built from 20 canonical amino acids. The encoded polypeptide sequences (primary structure) form local folds, such as α-helices and β-sheets (secondary structure), which fold into three-dimensional architectures (tertiary structure) and assemble into multi-protein complexes (quaternary structure). These structures define proteins’ biological function and activity, which have been optimized through evolution over billions of years. Proteins continually remodel these higher-order structures, expanding functional diversity.
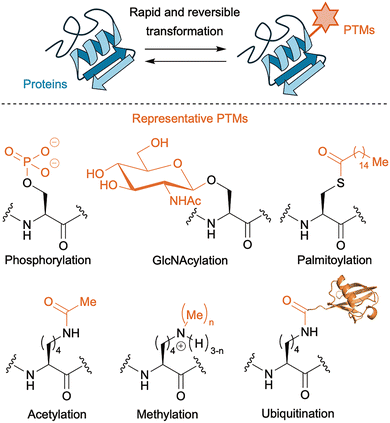
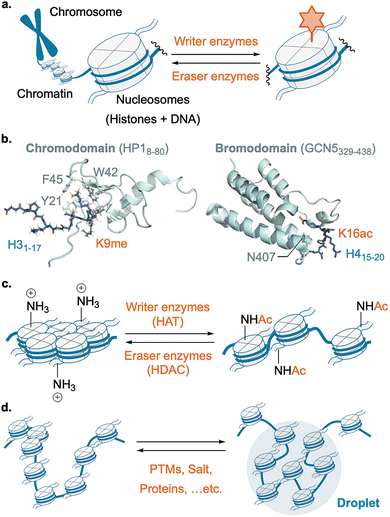
Post-translational modifications (PTMs), protein modifications performed after their production by the ribosome, further expand the diversity of protein structures beyond the 20 canonical amino acid residues by covalently modifying backbones and side chains.1 By modulating charge, hydrophobicity, sterics, and hydrogen-bonding capacity, PTMs can rapidly and reversibly reprogram protein behavior. In these regulatory systems, enzymes that install modifications are referred to as “writers,” those that remove them as “erasers,” and proteins that recognize modified residues and translate them into downstream biological responses as “readers.” Representative PTMs are phosphorylation on Ser/Thr/Tyr, O-GlcNAcylation on Ser/Thr, S-palmitoylation on Cys, acetylation on Lys/N-terminus, methylation on Lys/Arg, and ubiquitination/SUMOylation on Lys (Fig. 1). Irreversible PTMs, including proteolytic cleavage, also play important roles in protein maturation and disease-related protein processing.2 This Tutorial Review, however, primarily focuses on reversible PTMs, which are particularly attractive targets for manipulation using biocatalysis or abiotic catalysis.
Functionally, PTMs operate as chemical switches whose effects propagate from the nanometer to the cellular scale. They stabilize or destabilize local folds to tune the function of a specific protein. They can also remodel protein interfaces to alter interactions with other biomolecules. At larger scales, PTMs help encode the material state of macromolecular assemblies, including biomolecular condensates formed through liquid–liquid phase separation (LLPS).3,4 Through these mechanisms, PTMs regulate transcription by RNA polymerase II,5 cell-cycle progression,6 chromatin dynamics,7 and cytoskeletal organization.8 Dysregulation of PTM regulators is directly linked to aging and diseases such as cancer and neurodegeneration.9 The types and sites of identified PTMs are continually increasing due to the progress of proteomics,10,11 and over 600 types of PTMs have been categorized in UniProt Knowledgebase across all taxa so far.12
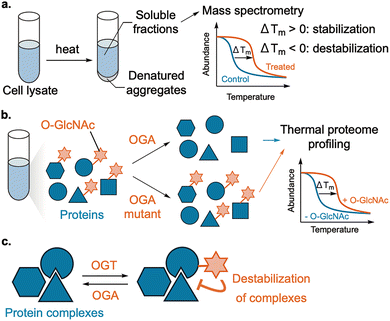
Despite their importance, the functions of PTMs in living systems remain difficult to dissect because they are often heterogeneous, substoichiometric, and transient. In addition, the functional consequences of PTMs are further complicated by the diversity of the proteome across tissues and cell types, as well as by differences in the expression levels of PTM writers, erasers, and readers. To address these challenges, elucidating the functions of PTMs in cells relies on two complementary strategies: profiling and perturbation. For profiling, mass spectrometry or antibody-based methods (immunoblotting, immunoprecipitation, and imaging) are effective. In particular, mass spectrometry-based proteomics is widely used to identify PTMs that correlate with specific phenotypes, diseases, or drug treatments (Fig. 2a).13,14 However, these approaches alone primarily report association rather than causation; temporal resolution can be limited by sample processing; site coverage depends on enrichment, ionization, and antibody specificity. Perturbation of PTMs is required to establish cause and effect. Small-molecule inhibitors of PTM writers/erasers remain the most accessible option and have yielded numerous biological insights (Fig. 2b). Starting from natural products, many studies have developed inhibitors with high selectivity, also motivated by therapeutic need.15,16 However, many inhibitors still act on multiple family members, complicating residue-level assignments of PTM roles. The promiscuous nature of writers/erasers is also a fundamental limitation in this method.
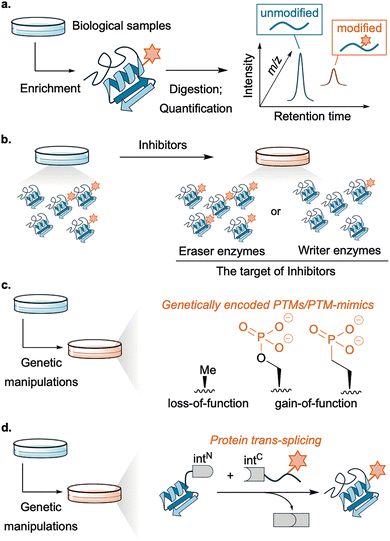 | ||
| Fig. 2 Methods for profiling (a) or perturbation ((b): inhibitor, (c): genetic code expansion, (d): protein trans-splicing) of PTMs. | ||
By contrast, biological approaches that leverage highly specific genetic manipulation can achieve higher precision (Fig. 2c and d). Site-directed mutagenesis creates PTM “mimics” (e.g. Ser to Ala for non-phosphorylated Ser, Ser to Asp for phosphorylated Ser, Lys to Gln for acetylated Lys), and is mainly used to probe the role of a specific PTM in test tubes.17 Genetic code expansion extends this strategy by installing PTMs or stable analogs in cells (Fig. 2c).18 Semi-synthetic strategies based on protein trans-splicing allow installation of defined PTMs onto cellular proteins with residue-level precision (Fig. 2d).19,20 In this method, a synthetic fragment of a split intein bearing the desired PTMs is delivered to cells expressing the target protein fused to the complementary intein fragment, thereby generating the protein with the desired modifications. Although these approaches have advantages in precision and generality, they typically act on exogenously expressed proteins and may not fully reproduce endogenous regulation. The one-shot nature of installation can also limit reversible spatiotemporal control of PTMs. Moreover, the reliance on genetic manipulation complicates therapeutic application. Overall, the lack of broadly applicable, residue- and protein-selective perturbation methods in living systems still constrains causal analysis of how PTMs influence biological phenomena, especially protein higher-order structures, where rapid and regioselective changes can be decisive.21,22
Catalysis offers a direct route to manipulate PTMs. Native or engineered enzymes, induced-proximity strategies, and abiotic chemical catalysts can directly manipulate PTMs with spatiotemporal precision (Fig. 3). These approaches enable cause-and-effect studies of PTMs, thereby bridging mechanistic biology and drug discovery. Moreover, this strategy is opening an emerging field in which catalysis itself functions as a therapeutic modality. The idea is to leverage the ability of catalysis that transforms diverse substrates into desired products, potentially in a manner beyond what nature does.
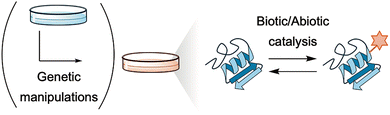 | ||
| Fig. 3 Catalysis for direct manipulation of PTMs. Notably, some methods, particularly those employing abiotic catalysis, enable PTM manipulation without the need for genetic manipulation. | ||
This Tutorial Review first outlines how PTMs shape protein higher-order structures (Section 2) and then surveys catalytic strategies to decipher (Sections 3 and 4) and control (Sections 5 and 6) them. Distinctively, we focus not only on how PTMs can be controlled but also on the consequences of such control for protein structure across scales, presenting an integrated, chemistry-based perspective aimed at an inclusive understanding of these phenomena. Although PTMs span multiple taxa, we focus primarily on human proteins, particularly in disease-related contexts.
2. PTMs and protein higher-order structures
2.1. Overview of PTM regulation
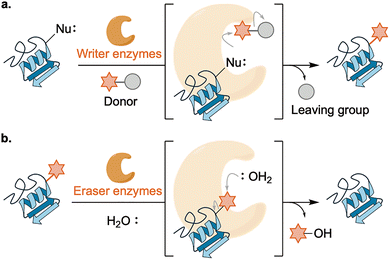
Regulation of most PTMs follows a unified scheme centered on writer and eraser enzymes equipped with catalytic and substrate-recognition motifs.23 Writers transfer a modification group from a donor molecule to a defined residue on a protein substrate, typically through nucleophilic attack by the target residue, although mechanistic variations exist depending on the PTM chemistry (Fig. 4a). Typical examples include the transfer of the phosphate group from ATP in phosphorylation, the acetyl group from acetyl-CoA in acetylation, and the methyl group from S-adenosyl-L-methionine (SAM) in methylation.Erasers employ several types of chemistries. The most common mode is hydrolysis, with water acting as the nucleophile (Fig. 4b); this includes many erasers for PTMs that have an electrophilic nature, like phosphatases, deacylases, and deubiquitinases. Cysteine- or serine-based hydrolases capture the PTM to generate a PTM–enzyme covalent intermediate that is subsequently hydrolyzed, whereas metallohydrolases activate water directly. A second mode couples co-substrates: NAD+-dependent sirtuin deacetylases consume NAD+ to cleave the amide bond of acetylated Lys, yielding nicotinamide and 2′-O-acyl-ADP-ribose.24 A third mode is an oxidation-dependent reaction: FAD-dependent demethylases oxidize the ε-N-methyl group to an iminium that is hydrolyzed to formaldehyde.25
In both directions, active sites preorganize donors and acceptors and tune their reactivity through catalytic residues, metal ions, and cofactors. Exceptions include enzyme-independent PTMs that arise from intrinsic chemical reactivity. Representative cases include oxidative modifications or acylations driven by reactive oxygen species or metabolites such as acyl-CoA thioesters and acyl phosphates. The roles of these nonenzymatic PTMs in disease and aging are being elucidated, and further studies are required.26,27
2.2. Classification of PTM-induced structural effects
Primary effects of PTMs can be classified into two types: intramolecular structural changes and intermolecular interaction changes.28 Intramolecular structural changes include alterations of local folding or covalent topology (e.g., crosslinking) (Fig. 5a). These can alter enzyme catalytic activity or propagate to new intermolecular interactions. Intermolecular structural changes include altered binding to other biomolecules (proteins, nucleic acids, lipids, or small molecules) (Fig. 5b).29 These lead to downstream consequences, including recognition by writer, eraser, or reader proteins,23 changes in localization,30 altered degradation rates,31 and remodeling of larger cellular assemblies such as chromatin,7 amyloid,32 cytoskeleton,8 and biomolecular condensates.33 Beltrao, Krogan, and colleagues reported that phosphorylation and acetylation sites located at protein–protein interfaces are more conserved between S. cerevisiae and humans than average PTM sites, indicating that interface-localized PTMs constitute “hot spots” under stronger functional constraint rather than neutral drift. This suggests that interface-localized PTMs are functionally constrained to preserve interaction regulation across evolution.34 Intermolecular interactions of writer or eraser proteins with PTMs further lead to crosstalk among PTMs, introducing or eliminating modifications proximal to one another, thereby forming a complex PTM network exemplified by the chromatin epigenetic code (epigenome) and the tubulin code.35,36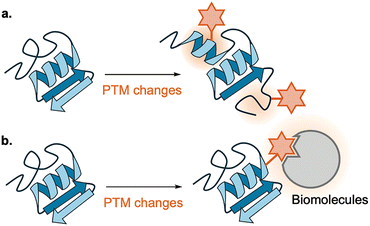 | ||
| Fig. 5 Primary effects of PTMs classified by structural perspective. (a) Intramolecular structural changes. (b) Intermolecular interaction changes. | ||
Stability regulation of biomolecules affects biological outcomes. PTMs critically control protein stability and are therefore major therapeutic targets, a theme that recurs throughout this review. Stability regulation by PTMs operates by either stabilization/destabilization of tertiary structures (intramolecular effects)37 or by installing degradation signals, called degrons, exemplified by polyubiquitin chains that target substrates for proteasomal degradation (intermolecular effects).31 In the following sections, we illustrate how each class of PTM effects, as classified above, manifests at the molecular and cellular scales.
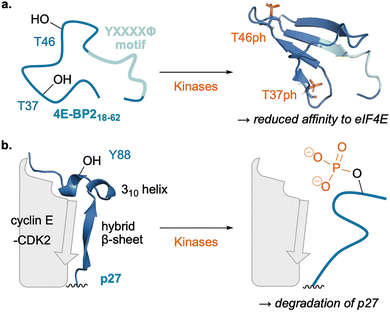
2.3. PTM-driven intramolecular structural changes
2.4. 14-3-3: PTM-driven intermolecular interaction changes
We next survey representative cases in which PTMs rewire intermolecular interactions. Once a PTM is installed or removed, it can alter a protein's binding ability to other proteins, nucleic acids, lipids, or small molecules. This section focuses on how PTMs change recognition by reader modules,23 stability,31 localization,30 and the assembly of larger complexes and condensates.7,8,32,33Writers and erasers install or remove PTMs, whereas reader proteins decode them. Reader domains typically recognize a local chemical epitope rather than a global protein fold, allowing a small repertoire of modules to interpret PTMs across diverse protein families.
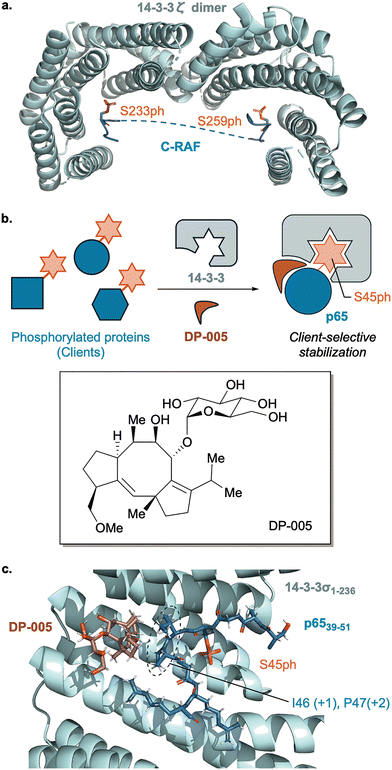
The 14-3-3 protein family exemplifies this logic: 14-3-3 dimers primarily recognize short phosphorylated Ser/Thr motifs rather than global folds, permitting interaction with hundreds of cellular clients across pathways (Fig. 7a).41,42 Human cells express seven 14-3-3 isoforms (β, γ, ε, ζ, η, σ, τ/θ), which form homo- or heterodimers. Recent thermodynamic profiling revealed isoform-specific hierarchies of phospho-target binding and distinct client selectivity, implying that isoform abundance and pairing stoichiometry help determine the phenotypic output of a given phosphorylation event.43 Functionally, 14-3-3 binding can mask localization or degron motifs, allosterically stabilize specific target conformations, or scaffold multi-protein assemblies. Because a limited set of 14-3-3 dimers can parse many phospho-signals, these adaptors operate as information hubs in phosphorylation networks, integrating diverse kinase inputs and distributing outputs to cell-cycle control, apoptosis, cytoskeletal remodeling, and metabolism.43–45
Given their hub role, reader proteins are often studied as therapeutic targets. For 14-3-3, its phospho-dependent protein–protein interactions (PPIs) have been defined structurally and mechanistically, and small molecule “molecular glues” have been developed to selectively stabilize reader-client interfaces.45 For example, Ottmann and co-workers reported a semisynthetic fusicoccane derivative (DP-005) that preferentially stabilized the 14-3-3/p65 (a subunit of NF-κB) complex with high cooperativity, yielding client-selective PPI enhancement (Fig. 7b).46 Fusicoccanes, a class of natural diterpenoid compounds, had long been known to stabilize a subset of 14-3-3 protein interactions,47 and subsequent synthetic studies generated derivatives with distinct client- and isoform-selectivity.48 However, the rational design principles governing selective molecular glue-mediated PPI stabilization remained largely underexplored.
In this study, DP-005 was identified as a potent stabilizer of the p65/14-3-3 interaction. Fluorescence anisotropy (FA) assays showed that the dissociation constant of the p65/14-3-3 complex decreased from 350 µM in the absence of compound to 2.8 µM in the presence of 100 µM DP-005, corresponding to more than a 100-fold stabilization. Structural and biophysical analyses showed that DP-005 occupied a hydrophobic pocket at the p65/14-3-3 interface, with its sugar moiety engaging the surrounding water shell (Fig. 7c). Because this pocket is largely shaped by the +1 and +2 residues adjacent to the phospho-Ser site (Ile and Pro for p65 S45ph), DP-005 exhibited weaker stabilization for other 14-3-3 client proteins with different +1/+2 residues. Quantitatively, FA measurements showed approximately 6-fold stabilization for CFTR, ∼2-fold for c-Raf, and little or no stabilization for p53. These trends were further supported by isothermal titration calorimetry (ITC) measurements. Notably, this selectivity did not correlate with the intrinsic affinity of DP-005 for individual client proteins but arose from differences in cooperativity within the corresponding ternary complexes comprising DP-005, 14-3-3, and the phosphorylated client, highlighting how a local PTM can reshape higher-order protein assemblies through reader-mediated interactions.
Given that dysregulation of the NF-κB pathway is related to cancer proliferation and inflammatory diseases,49 selective stabilization of the 14-3-3/p65 interaction can offer a therapeutic strategy, although its translational potential remains to be established.50 Subsequent studies on other 14-3-3 clients, including ERα, C-RAF, and CFTR, further support reader proteins as targets for manipulating PTM-dependent interaction networks.47
2.5. PTM-driven regulation of larger assemblies
A combination of these intramolecular structural changes or intermolecular interaction changes regulates larger assemblies. In this part, we introduce the representative ones, chromatin, amyloid, cytoskeleton, or biomolecular condensates.Electrostatic interactions also play a crucial role in the chromatin state. For example, acetylation of histones neutralizes the positive charge of Lys on histone tails, weakening electrostatic interactions with DNA and inter-nucleosome interactions (Fig. 8c); H4K16ac in particular disrupts higher-order chromatin fiber compaction and increases accessibility.58–60 Thus, PTMs tune chromatin both through molecular recognition by reader motifs and by physicochemical remodeling of nucleosomes.
In recent years, the histone code has expanded into a biophysical framework (Fig. 8d). HP1-rich heterochromatin domains undergo liquid–liquid phase separation (LLPS), with PTMs such as H3K9 methylation and H3S10 phosphorylation modulating material properties, dynamics, and compartmentalization.61,62 Chromatin itself also possesses an intrinsic propensity to phase separate; acetylation and other marks tune nucleosome–nucleosome cohesion and thereby shift condensate behavior.63 Together, these findings emphasize that PTMs couple molecular recognition to emergent physical properties to organize transcription in both space and time.
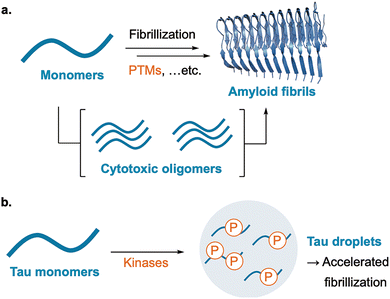
Phosphorylation of tau is among the best-studied PTMs in amyloid biology.67 Hyperphosphorylation is a defining pathological feature. Under near-physiological conditions, phosphorylated tau undergoes LLPS, and the resulting droplets can mature toward amyloid fibrils, positioning LLPS as an intermediate between soluble tau and insoluble aggregate (Fig. 9b).68
Similar PTM-driven regulation is reported for other amyloidogenic proteins. For amyloid-β (Aβ), phosphorylation at S8 promotes early oligomer formation and accelerates fibrillization, illustrating how a single site-specific modification can bias the aggregation pathway.69 For α-synuclein, aggregation is a pathological hallmark of Parkinson's disease, with α-synuclein deposited in Lewy bodies and Lewy neurites. In this context, phosphorylation at S129 is enriched in these lesions and widely used as a marker of pathological α-synuclein, whereas proteolytic cleavage and truncation have also been implicated in modulating aggregation and disease-associated species.70,71 For TDP-43, acetylation impairs RNA binding and increases aggregation propensity, yielding insoluble species reminiscent of ALS/FTLD pathology.72 More recent work showed that acetylation of TDP-43 at K136 promoted LLPS that evolves toward less fluid, pathological assemblies, an effect counteracted by SIRT1 deacetylation, while RNA binding antagonizes these neurotoxic phase transitions.
Together, these examples illustrate a shared principle: PTMs act as molecular switches that rebalance dynamic condensates and stable amyloid assemblies, linking chemical modification to disease-relevant structural outcomes.
2.6. Short summary
In this section, we grouped PTM effects on protein higher-order structures into two classes: intramolecular structural changes and intermolecular interaction changes. At the molecular scale, PTMs reweight local folding, dynamics, and binding networks to regulate activity, stability, and localization. These effects propagate mesoscale architectures, including chromatin, amyloid, and biomolecular condensates, through multivalency and cooperative assembly. This framework motivates the next sections on catalytic strategies to decipher and control PTM-driven structure and function in living systems.3. Biocatalysis to decipher effects of PTMs
3.1. Overview
As discussed above, several methodologies have been developed to explore the role of PTMs. In this section, we focus on enzyme-based tools for in situ PTM editing to enable cause-and-effect studies of PTMs on higher-order protein structures. Two complementary strategies are considered: (i) leveraging native enzymes as modulators of PTMs in cells and organisms, and (ii) deploying engineered enzymes that expand substrate/site selectivity and provide tighter spatial and temporal control. Together, these biocatalytic approaches connect residue-level editing with structural and systems-level understanding of how PTMs regulate higher-order protein structures.3.2. Native enzyme-based catalysis
Leveraging native writer or eraser enzymes without protein engineering is the most straightforward way to perturb PTMs. Although the substrate scope of a given enzyme can limit the range of proteins that can be targeted, native-enzyme perturbations can still yield mechanistic insights when coupled with quantitative proteomic or biophysical analyses that capture the global consequences of PTM introduction or removal. Such combined approaches enable causal links between PTMs and proteome-wide properties, including protein stability, interaction networks, and localization, depending on the analytical context.3.3. Engineered enzyme-based catalysis
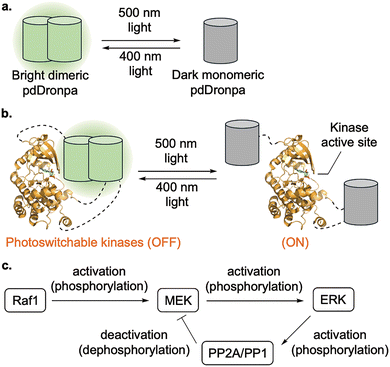
To overcome these limitations, the authors developed pdDronpa, a reversibly dimerizing fluorescent protein that associates under violet (∼400 nm) light and dissociates under cyan/green (∼500 nm) light irradiation (Fig. 11a). Fusing pdDronpa to kinase scaffolds yielded light-gated kinase variants such as psRaf-1, psMEK1/2, and psCDK5, whose kinase activities are caged or uncaged by light-linked pdDronpa association or dissociation (Fig. 11b). Light irradiation switched kinase activity ON/OFF within minutes, enabling precise programming of phosphorylation ability in cultured cells and in vivo. Using short pulses (<minutes) of psRaf-1 activation, the authors uncovered a fast negative feedback mechanism in the Raf-MEK signaling whereby ERK activity promotes PP2A/PP1-mediated dephosphorylation of MEK (Fig. 11c). Because this feedback operates on a minute timescale, its identification required rapid, reversible kinase switching that is inaccessible to conventional genetic perturbations. Collectively, these results have established an experimental blueprint for dynamics-resolved PTM causality using photoactivatable enzyme catalysis and motivate biological applications in which the timing of phosphorylation encodes function. For example, subsequent work leveraged psMEK-based control of ERK to demonstrate that precise phosphorylation/dephosphorylation timing determines long-term memory outcomes in a neural circuit model.80
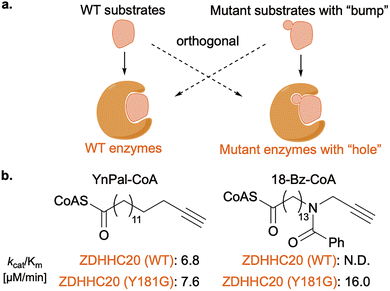
Applying this concept, Tate and co-workers generated engineered ZDHHC S-acyltransferases matched to uniquely modified fatty-acyl probes, and performed proteome-wide, isoform-resolved mapping of S-acylation in live cells.82 Long-chain S-acylation on Cys is a well-known PTM that regulates protein localization in cells. The human ZDHHC family comprises 23 enzymes and is estimated to modify >3000 cysteine residues across ∼12% of the proteome.83 The substrate networks of individual ZDHHCs have been difficult to define due to (i) the absence of inhibitors highly selective to each subtype and (ii) compensation following overexpression/knockdown/knockout. To address this, the authors created mutants of five diverse ZDHHCs (3, 7, 11, 15, and 20) with a “hole” that transfer “bumped” fatty-acyl probes selectively, thereby labeling substrates of the engineered isoform in cells (Fig. 12b). Proteomic analyses revealed that each engineered ZDHHC possessed a largely non-overlapping substrate repertoire, indicating that individual S-acyltransferases engage distinct proteome subsets. Notably, the study identified IFITM3 as a high-confidence substrate of ZDHHC20 and reconciled earlier knockout observations by demonstrating compensation among ZDHHCs. IFITM acylation has been implicated in antiviral defence and cancer biology,84–86 underscoring how subtype-level editing can interrogate how site/isoform selectivity and lipid chain chemistry encode changes in localization, complex assembly, and function.
3.4. Short summary
Together, these biocatalytic approaches, ranging from native enzyme utilization to precision-engineered variants, are transforming how PTMs are studied in their natural context. Native enzymes provide realistic perturbations that reveal how bona fide catalysis reshapes protein characteristics, while engineered catalysts extend these principles to living cells with unprecedented spatial, temporal, and molecular precision. By directly linking catalysis to structure and function, such tools convert PTM research from correlative observation into causal experimentation. Ultimately, the convergence of enzymology, chemical biology, and structural proteomics is establishing a quantitative framework for how catalytic modification events dynamically organize the proteome and protein higher-order architectures.4. Abiotic catalysis to decipher effects of PTMs
4.1. Spatially defined PTM-introduction by an abiotic/enzymatic hybrid catalyst system
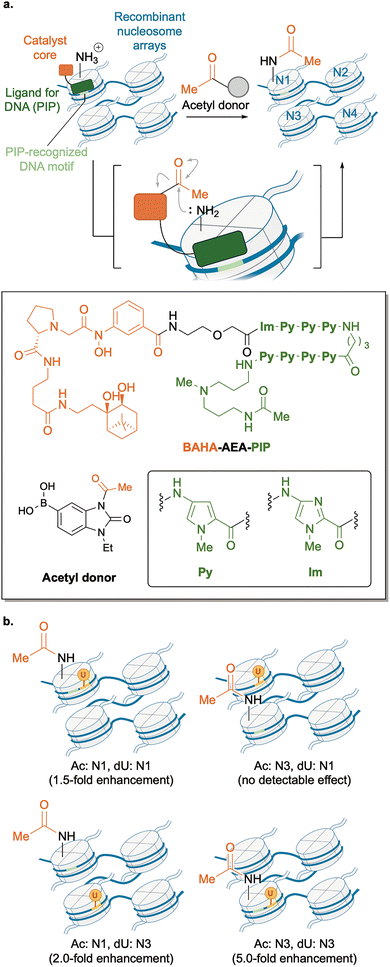
Beyond enzymes, synthetic molecules with enzyme-like activity represent a growing frontier for exploring the role of PTMs. Sczepanski and our group developed an abiotic/enzymatic hybrid catalyst system (ABEHCS) that enabled the construction of nucleosome arrays bearing spatially defined histone PTMs and DNA damage, thereby allowing systematic analysis of how the relative position of PTMs and DNA damages influences chromatin-associated processes.87 Using this approach, we uncovered a spatially dependent relationship between H3K56ac and base excision repair (BER) efficiency.In this system, we developed an abiotic nucleophilic catalyst, PIP–AEA–BAHA, which promoted lysine acetylation on histone H3 within recombinant nucleosome arrays using a stable acetyl donor (Fig. 13a). This reaction relies on ligand-directed chemistry88 combined with our original catalyst core (BAHA; boronate-assisted hydroxamic acid) for lysine acylation.89 Unlike canonical methods to generate homogeneously modified nucleosome arrays, this strategy enabled regioselective acetylation within a specific nucleosome embedded within the tetramer. Because the ligand motif, a pyrrole-imidazole polyamide (PIP), selectively binds the minor groove of DNA with a defined sequence preference,90 the catalyst selectively introduced acetylation into the nucleosome harboring the PIP-recognized DNA motif among four nucleosomes (N1–N4) in the array. Initial abiotic acetylation produced off-target modifications on histone tail lysines; however, subsequent enzymatic deacetylation using sirtuin histone deacetylases (Sirt3 and Sirt7) efficiently removed these unintended marks. This sequential abiotic/enzymatic editing yielded tetranucleosome arrays containing H3K56ac at a specific mononucleosome in high yield (∼61%), while minimizing acetylation on other lysine residues.
By combining ABEHCS with a plug-and-play strategy that allows site-specific insertion of isotope-labelled 2′-deoxyuridine (dU) into nucleosome arrays,91 we directly examined how the spatial arrangement of histone acetylation and DNA damage affects BER catalyzed by uracil DNA glycosylase (UDG) and apurinic/apyrimidinic endonuclease 1 (APE1) (Fig. 13b). These experiments revealed that H3K56ac enhanced BER within nucleosome arrays in a strongly position-dependent manner. Notably, the PTM-driven enhancement of the repair-promoting effect was most pronounced when DNA damage resided at internal nucleosomes (N3) within the array, where DNA accessibility is intrinsically limited.
Collectively, this study demonstrated that histone PTMs do not act merely as binary on/off signals but function as spatially encoded regulators whose biological consequences depend critically on their relative positioning within higher-order chromatin architectures. By enabling programmable, nucleosome-selective PTM installation without genetic manipulation, ABEHCS provides a chemical framework for deciphering how the spatial organization of PTMs governs chromatin-based phenomena.
5. Biocatalysis to control PTMs
5.1. Overview
Beyond elucidating PTM mechanisms, recent works seek to actively control PTMs as a therapeutic strategy through direct catalytic installation or removal of PTMs, rather than through inhibition of native PTM writers or erasers. Two complementary axes are emerging: (i) biological catalysis, including engineered enzymes (Section 5.2) and induced-proximity systems (Section 5.3), and (ii) abiotic catalysts based on synthetic small molecular catalysts (Section 6). Biological systems offer high selectivity based on the high molecular recognition ability of proteins, whereas abiotic systems provide broader chemical diversity in reaction space and potentially superior drug-like properties. In this section, we describe the design and utilities of biocatalysis for PTM manipulations.5.2. Engineered enzymes for epigenetic therapy
As outlined in Section 3, engineered enzymes can edit PTMs on specific proteins and residues within living cells. A major therapeutic concept is epigenome editing: installing or erasing histone/DNA marks at chosen genomic loci to reprogram transcription without introducing DNA breaks. This has been achieved by fusing writer/eraser domains to programmable DNA-binding proteins such as zinc finger proteins (ZFs), transcription activator-like effectors (TALEs), and more readily programmable nuclease-null deactivated Cas9 (dCas9).92Reddy, Gersbach, and colleagues pioneered a dCas9-based approach by fusing the catalytic core of histone acetyltransferase p300 to dCas9, creating a programmable acetyltransferase (Fig. 14a).93 Guide-RNA targeting deposited H3K27ac at promoters and enhancers of targeted genes, and activated transcription with high locus specificity, establishing that chromatin-localized catalytic editing can drive gene expression without DNA cleavage.
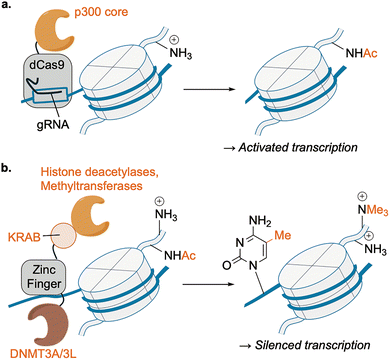 | ||
| Fig. 14 Engineered enzymes for epigenetic therapy. (a) dCas9–p300 approach for transcriptional activation of specific genes. (b) ZF-ETRs approach for transcriptional silencing of specific genes. | ||
Inspired by this dCas9–p300 system, many engineered enzyme systems were developed toward therapeutic purposes.94 Notably, Lombardo and co-workers demonstrated durable and efficient in vivo gene silencing through “hit-and-run” epigenome editing.95 The authors engineered zinc-finger-based epigenetic transcriptional repressors (ETRs) that combine KRAB (recruiter of histone deacetylases and methyltransferases), DNMT3A (DNA methyltransferase), and DNMT3L (co-factor of DNMT3A) domains within one scaffold (Fig. 14b). Delivered as mRNA in lipid nanoparticles to hepatocytes, ETRs induced targeted DNA methylation and repressive histone modifications at the Pcsk9 locus without DNA breaks, achieving robust silencing and a ∼50% reduction of circulating PCSK9 proteins that persisted for nearly one year in mice after a single administration. Importantly, this repression remained detectable even after surgically induced liver regeneration, supporting the high durability of this epigenome manipulation despite the absence of permanent genome modification and its potentially reversible nature.
Together, these studies established that biocatalytic editing by chromatin-localized, engineered enzymes can precisely manipulate transcriptional states. As the authors proposed, combining such systems with improved delivery technologies (DDS, viral or nanoparticle vectors) could yield gene-specific epigenetic therapies applicable far beyond oncology, including neurological and metabolic diseases.
5.3. PTM regulation with chemical inducers of dimerization
A complementary biological strategy is to chemically rewire PTM enzyme–substrate interactions using bifunctional small molecules. Chemical inducers of dimerization (CIDs) recruit a writer or eraser enzyme into proximity to a target non-native for the enzyme, enabling PTM installation or removal. The prototypical example is proteolysis-targeting chimeras (PROTACs), which recruit an E3 ubiquitin ligase to a protein of interest (POI) to induce ubiquitination and subsequent proteasomal degradation, a concept that revolutionized targeted protein degradation.96Building on this logic, chemical dimerizers have been generalized to other PTMs. In 2020, Choudhary and colleagues developed phosphorylation-inducing chimeric small molecules (PHICS) that recruit kinases, such as AMP-activated protein kinase (AMPK) or protein kinase C (PKC), to non-physiological substrates to promote their phosphorylation (Fig. 15a).97 PHICS1 and PHICS2 induced phosphorylation of BRD4 proteins in test tubes by dimerizing BRD4 with AMPK (β1 isoform) for PHICS1 and PKC for PHICS2, whereas negative-control analogues (iPHICS1/2) lacking BRD4-binding capability were inactive. In addition, PHICS3 targeting Bruton's tyrosine kinase (BTK) induced phosphorylation of overexpressed BTK-Flag via AMPK recruitment in living cells. These results demonstrated the feasibility of synthetic dimerizers for controlling phosphorylation.
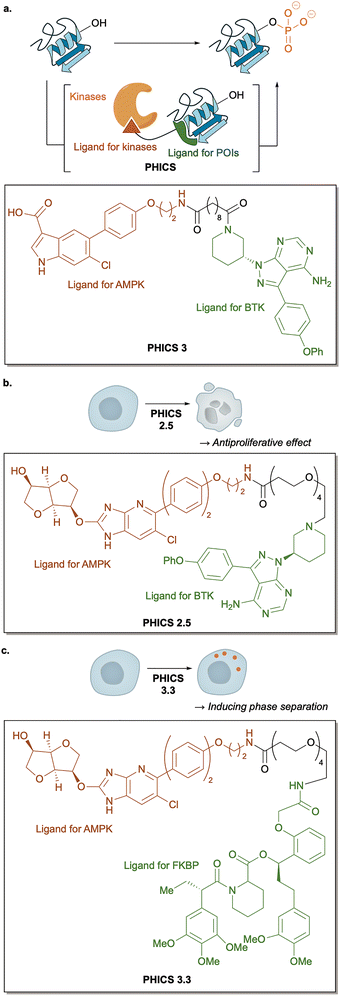 | ||
| Fig. 15 PTM regulation with chemical inducers of dimerization. (a) Schematics of PHICS. (b) and (c) Next-generation PHICS for regulating dynamic cellular processes. | ||
In 2025, the same group greatly advanced this platform by redesigning both the AMPK-recruiting moiety and the linker architecture to create next-generation PHICS with improved potency.98 First-generation PHICs recruited only a specific isoform of AMPK and were effective only in a subset of cell types. Additionally, they required target overexpression or nutrient stress for detectable phosphorylation. To overcome these limitations, the authors employed MK-8722, a pan-AMPK recruiter capable of engaging all major AMPK complexes.99 They also optimized linker length and composition to enhance ternary complex formation, resulting in efficient phosphorylation of endogenous BTK proteins under physiological culture conditions without the need for overexpression or nutrient stress. The most effective compound, PHICS 2.5, exhibited strong antiproliferative activity in BTK-dependent lymphoma cell lines (EC50 = 3.2 and 5.0 µM for Raji and Mino cells, respectively), outperforming ibrutinib (EC50 > 10 µM for Raji and Mino cells) and demonstrating the therapeutic potential of proximity-induced phosphorylation (Fig. 15b).
Notably, they also developed PHICS 3.3 for liprin-α3 fused an FKBP tag, a presynaptic scaffold protein whose phosphorylation drives LLPS. Proximity-induced phosphorylation of liprin-α3 using PHICS 3.3 triggered LLPS in living cells, showing that induced kinase recruitment can be used to control membraneless assemblies’ formation (Fig. 15c).
Together, the PHICS studies mark a progression from proof-of-principle to programmable control of dynamic cellular processes, underscoring that chemically induced-proximity biocatalysis extends beyond degradation to spatiotemporal rewiring of phosphorylation networks.
5.4. Short summary
In this section, we outlined biological strategies that achieve active control of PTMs by exploiting the high selectivity of enzymes. Biological systems, including engineered enzymes and chemical dimerizers, offer programmable and reversible modulation of PTMs through precise molecular recognition. With these platforms in hand, their therapeutic potential is increasingly being explored as novel drug modalities that access protein higher-order structures and functions via PTM control. Advancing translational aspects, including ADMET properties, together with further improvements in reactivity and selectivity, represents a critical next step toward practical applications.6. Abiotic catalysis to control PTMs
6.1. Overview
Abiotic catalysis for PTM control, as introduced in Section 4, can also be used for therapeutic purposes. In principle, this strategy is not constrained by naturally evolved enzymatic reaction and can therefore include virtually any chemical transformation, offering a broader reaction space than enzyme-dependent biocatalysis alone. In addition, abiotic approaches can employ synthetic small-molecule catalysts, which may exhibit superior drug-like properties.Our group is pursuing this direction by postulating the concept of “catalysis medicine,” in which abiotic catalysts manipulate post-translational modifications (PTMs) in living systems toward therapeutic applications.100,101 This approach enables a purely chemical way to probe and reprogram biological architectures without genetic manipulation.
6.2. Abiotic catalysis for epigenetic therapy
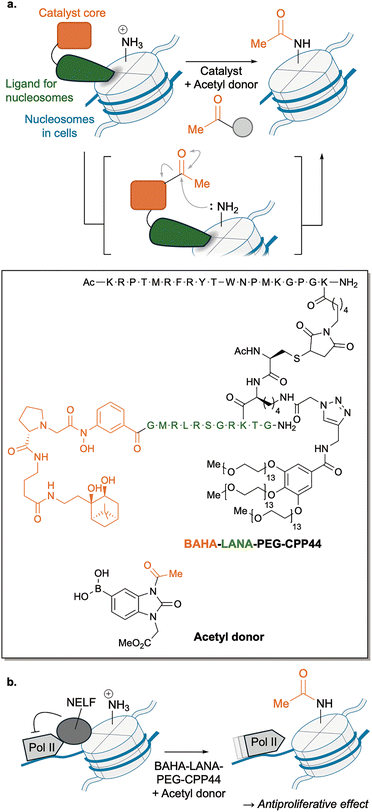
A recent advance demonstrated the potential of this strategy in epigenetic therapy. We developed an abiotic nucleophilic catalyst that promotes acetylation of histone proteins using a stable acetyl donor with high selectivity for H2BK120.102–104 This selective reaction is achieved through ligand-directed chemistry88 combined with our original catalyst core for lysine acylation,89,102,105 as discussed in Section 4. The LANA peptide serves as a nucleosome-binding ligand, positioning the catalytic core to acetylate the proximal lysine under physiological conditions (Fig. 16a). The optimized catalyst, BAHA–LANA–PEG–CPP44, which had selective cell-membrane permeability toward leukemia cells, induced acetylation of endogenous H2BK120 across multiple leukemia cell lines. Time-resolved analyses revealed that this site-specific acetylation displaced the negative transcription elongation factor (NELF) complex from chromatin, thereby activating some genes by releasing RNA polymerase II (Pol II) from NELF-mediated suppression (Fig. 16b). We also observed that this epigenome reprogramming suppressed leukemia cell proliferation and tumorigenicity in vivo, establishing a causal connection between a chemically installed PTM, local chromatin remodelling, and transcriptional control. The regioselectivity of the catalytic histone acylation was recently extended from H2BK120 to H2BK108 and H2BK43, promising further utility of this approach in furnishing other biological activities.1066.3. Abiotic catalysis for amyloid photooxygenation
Building on the concept of “catalysis medicine,” our group has also pursued the chemical regulation of protein higher-order assemblies, such as amyloids. Traditional therapeutic strategies of amyloidosis have focused on inhibiting amyloid formation or promoting clearance by biological means, such as anti-amyloid antibody drugs.107 In contrast, a purely chemical approach using small molecules to directly remodel or degrade amyloid assemblies remained largely unexplored, despite its advantages in availability and cost for therapeutic applications.To address this challenge, we developed a series of abiotic catalysts capable of selectively oxygenating His or Met on amyloid fibrils under physiological conditions and light irradiation.108 As a representative example, leuco ethyl violet (LEV) is a highly active, small-molecule, blood–brain barrier (BBB)-permeable catalyst enabling amyloid oxygenation in Alzheimer model mice (Fig. 17a).109 LEV is a prodrug form of the active catalyst, ethyl violet (EV), which is produced from LEV through an autocatalytic process. The catalytic photooxygenation of amyloids reduced their cytotoxicity and seeding ability, enhanced microglial uptake and degradation of modified species, thereby demonstrating that the chemical protein oxidation event can modulate complex proteostasis and biological clearance pathways.110,111 These catalysts operate through a photochemically driven mechanism in which the microenvironment of amyloid fibrils stabilizes the excited singlet state of the catalyst (1cat*), preventing rapid thermal relaxation (Fig. 17b). After intersystem crossing (ISC), the generated long-lived triplet state (3cat*) activates ground-state triplet oxygen (3O2) to singlet oxygen (1O2). The resulting locally generated singlet oxygen oxidizes the His112 and Met side chains of Aβ amyloid. The amyloid-selective photooxygenation has recently been extended to intracellular amyloid, α-synuclein, despite the reducing cellular environment.113 The controlled local generation of 1O2 in the vicinity of amyloid, driven by the catalyst's amyloid-sensing function, is the key to this phenomenon. Furthermore, we identified therapeutic effects of the catalytic photooxygenation within an animal model of transthyretin amyloidosis.114 Therefore, the small-molecule catalysis may be useful for therapeutics to treat various amyloidoses.
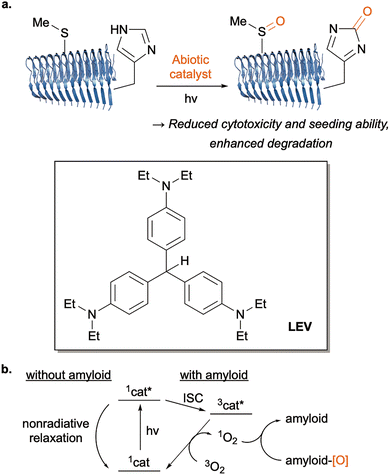 | ||
| Fig. 17 Abiotic catalysis for amyloid oxygenation. (a) Amyloid photooxygenation by abiotic catalyst. (b) Schematics of amyloid-selective photooxygenation via singlet oxygen generation. | ||
6.4. Short summary
In this section, we outlined abiotic strategies that manipulate histone epigenetics and induce amyloid degradation. Abiotic catalysts expand chemical possibilities by introducing reaction pathways beyond those available in nature, enabling direct manipulation of protein structures and assemblies. Furthermore, such catalysis studies will lead to a new modality for treating various diseases.7. Conclusion and perspectives
PTMs constitute a universal mechanism that couples the chemical state of proteins to their structural and functional organization. Through site-selective and reversible chemical reactions, PTMs dynamically reshape protein conformation, stability, intermolecular interactions, and ultimately the architecture of higher-order assemblies such as chromatin, amyloids, and phase-separated condensates. As accumulating evidence reveals, these structural transitions are not secondary byproducts but represent a physical basis through which biological systems process, transfer, and store information. Catalysis-based PTM control provides a powerful framework to decipher and control these processes with molecular precision. Enzyme-based biological catalysis and abiotic catalysis now enable direct interrogation of PTMs in living systems, bridging molecular chemistry, cellular biology, and medicine.Despite such progress, precise catalytic modulation of PTMs on dilute targets within crowded, heterogeneous cellular environments remains in its early stages. Although protein engineering has delivered tools with high spatial and temporal control, realizing analogous capabilities with small-molecule catalysts that operate in vivo remains a central challenge, particularly for therapeutic applications. Key hurdles include selective access to individual residues, resistance to redox and pH fluctuations, stability in cells and animals, and rigorous, structure-resolved evaluation of downstream biological consequences for distinguishing direct effects of catalytic PTM modulation from indirect cellular responses. In addition, because PTM-dependent structural transitions and their biological outcomes are often tissue- or cell-type specific, it is essential to study these processes in physiologically relevant systems, particularly in contexts such as neurodegenerative diseases where protein aggregation occurs in post-mitotic neurons. Advances in chemical synthesis, reaction design, and analytical platforms will be required to address these limitations. Mass-spectrometry-based methods and structural proteomics must evolve further to capture transient and substoichiometric PTM states, along with their conformational consequences. Equally important will be progress in molecular delivery technologies, including cell-permeable motifs and nanocarriers that can transport catalytic systems to defined intracellular or in vivo locations.
In summary, the convergence of catalysis, PTM chemistry, and structural biology is redefining how protein architectures can be understood and manipulated. By converting chemical reactivity into a mechanism capable of writing, erasing, and reorganizing protein structures, catalysis is poised to become a central principle for both fundamental biochemical analysis and therapeutic intervention. Continued development of catalytic strategies for PTM control will deepen our understanding of protein structural dynamics and may inaugurate a broader paradigm of catalysis-driven medicine.
Conflicts of interest
There are no conflicts to declare.Data availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
This work was supported by JPMJCR2572 (JST-CREST, MK), JP23H05466 (JSPS KAKENHI, MK), and JPMJAX242K (JST-ACT-X, YY).References
- S. Prabakaran, G. Lippens, H. Steen and J. Gunawardena, Wiley Interdiscip. Rev.: Syst. Biol. Med., 2012, 4, 565–583 CrossRef CAS PubMed.
- T. Klein, U. Eckhard, A. Dufour, N. Solis and C. M. Overall, Chem. Rev., 2018, 118, 1137–1168 CrossRef CAS PubMed.
- Y.-Y. Luo, J.-J. Wu and Y.-M. Li, Chem. Commun., 2021, 57, 13275–13287 RSC.
- F. Chu, S. Sharma, S. D. Ginsberg and G. Chiosis, Trends Biochem. Sci., 2025, 50(10), 892–905 CrossRef CAS PubMed.
- B. M. LeBlanc, R. Y. Moreno, E. E. Escobar, M. K. V. Ramani, J. S. Brodbelt and Y. Zhang, RSC Chem. Biol., 2021, 2, 1084–1095 RSC.
- S. A. G. Cuijpers and A. C. O. Vertegaal, Trends Biochem. Sci., 2018, 43, 251–268 CrossRef CAS PubMed.
- C. M. D. de Fenffe, J. Govers and F. Mattiroli, Biochemistry, 2025, 64, 2138–2153 CrossRef PubMed.
- B. MacTaggart and A. Kashina, Cytoskeleton, 2021, 78, 142–173 CrossRef PubMed.
- Q. Zhong, X. Xiao, Y. Qiu, Z. Xu, C. Chen, B. Chong, X. Zhao, S. Hai, S. Li, Z. An and L. Dai, MedComm, 2023, 4, e261 CrossRef CAS PubMed.
- M. den Ridder, P. Daran-Lapujade and M. Pabst, FEMS Yeast Res., 2019, 20, foz088 CrossRef PubMed.
- Y. Zhang, B. R. Fonslow, B. Shan, M.-C. Baek and J. R. Yates, Chem. Rev., 2013, 113, 2343–2394 CrossRef CAS PubMed.
- https://ftp.uniprot.org/pub/databases/uniprot/current_release/knowledgebase/complete/docs/ptmlist.txt .
- S. Doll and A. L. Burlingame, ACS Chem. Biol., 2015, 10, 63–71 CrossRef CAS PubMed.
- L. Zhai, K. Chen, B. Hao and M. Tan, Acta Pharmacol. Sin., 2022, 43, 3112–3129 CrossRef CAS PubMed.
- M. K. Tarrant and P. A. Cole, Annu. Rev. Biochem., 2009, 78, 797–825 CrossRef CAS PubMed.
- J. White, F. A. Derheimer, K. Jensen-Pergakes, S. O’Connell, S. Sharma, N. Spiegel and T. A. Paul, Trends Pharmacol. Sci., 2024, 45, 243–254 CrossRef CAS PubMed.
- K. W. Barber and J. Rinehart, Nat. Chem. Biol., 2018, 14, 188–192 CrossRef CAS PubMed.
- T. Peng, T. Das, K. Ding and H. C. Hang, Protein Sci., 2023, 32, e4618 CrossRef CAS PubMed.
- Y. David, M. Vila-Perelló, S. Verma and T. W. Muir, Nat. Chem., 2015, 7, 394–402 CrossRef CAS PubMed.
- A. J. Burton, M. Haugbro, E. Parisi and T. W. Muir, Proc. Natl. Acad. Sci. U. S. A., 2020, 117, 12041–12049 CrossRef CAS PubMed.
- J. Roca-Martinez, T. Lazar, J. Gavalda-Garcia, D. Bickel, R. Pancsa, B. Dixit, K. Tzavella, P. Ramasamy, M. Sanchez-Fornaris, I. Grau and W. F. Vranken, Front. Mol. Biosci., 2022, 9, 959956 CrossRef CAS PubMed.
- M. C. Marrero, V. H. Mello, P. Sartori and P. Beltrao, Nat. Commun., 2025, 16, 9407 CrossRef PubMed.
- M. J. Suskiewicz, BioEssays, 2024, 46, e2300178 CrossRef PubMed.
- M. D. Jackson and J. M. Denu, J. Biol. Chem., 2002, 277, 18535–18544 CrossRef CAS PubMed.
- Y. Shi, F. Lan, C. Matson, P. Mulligan, J. R. Whetstine, P. A. Cole, R. A. Casero and Y. Shi, Cell, 2004, 119, 941–953 CrossRef CAS PubMed.
- I. Maksimovic and Y. David, Trends Biochem. Sci., 2021, 46, 718–730 CrossRef CAS PubMed.
- T. Baldensperger, M. Preissler and C. F. W. Becker, RSC Chem. Biol., 2024, 6, 129–149 RSC.
- P. Beltrao, P. Bork, N. J. Krogan and V. van Noort, Mol. Syst. Biol., 2013, 9, MSB134521 CrossRef PubMed.
- X. Hong, N. Li, J. Lv, Y. Zhang, J. Li, J. Zhang and H.-F. Chen, Bioinformatics, 2022, 39, btac823 CrossRef PubMed.
- W. Li, J. Shen, A. Zhuang, R. Wang, Q. Li, A. Rabata, Y. Zhang and D. Cao, Cell. Mol. Biol. Lett., 2025, 30, 98 CrossRef PubMed.
- J. M. Lee, H. M. Hammarén, M. M. Savitski and S. H. Baek, Nat. Commun., 2023, 14, 201 CrossRef CAS PubMed.
- S. P. Moon, A. T. Balana and M. R. Pratt, Curr. Opin. Chem. Biol., 2021, 64, 76–89 CrossRef CAS PubMed.
- J. Li, M. Zhang, W. Ma, B. Yang, H. Lu, F. Zhou and L. Zhang, Mol. Biomed., 2022, 3, 13 CrossRef PubMed.
- P. Beltrao, V. Albanèse, L. R. Kenner, D. L. Swaney, A. Burlingame, J. Villén, W. A. Lim, J. S. Fraser, J. Frydman and N. J. Krogan, Cell, 2012, 150, 413–425 CrossRef CAS PubMed.
- M. Leutert, S. W. Entwisle and J. Villén, Mol. Cell. Proteomics, 2021, 20, 100129 CrossRef CAS PubMed.
- C. Janke and M. M. Magiera, Nat. Rev. Mol. Cell Biol., 2020, 21, 307–326 CrossRef CAS PubMed.
- P. Craveur, T. J. Narwani, J. Rebehmed and A. G. de Brevern, Amino Acids, 2019, 51, 1065–1079 CrossRef CAS PubMed.
- A. Bah, R. M. Vernon, Z. Siddiqui, M. Krzeminski, R. Muhandiram, C. Zhao, N. Sonenberg, L. E. Kay and J. D. Forman-Kay, Nature, 2015, 519, 106–109 CrossRef CAS PubMed.
- I. Chu, J. Sun, A. Arnaout, H. Kahn, W. Hanna, S. Narod, P. Sun, C.-K. Tan, L. Hengst and J. Slingerland, Cell, 2007, 128, 281–294 CrossRef CAS PubMed.
- S. L. Rath and S. Senapati, Sci. Rep., 2016, 6, 26450 CrossRef CAS PubMed.
- C. Johnson, S. Crowther, M. J. Stafford, D. G. Campbell, R. Toth and C. MacKintosh, Biochem. J., 2010, 427, 69–78 CrossRef CAS PubMed.
- M. Molzan and C. Ottmann, J. Mol. Biol., 2012, 423, 486–495 CrossRef CAS PubMed.
- G. Gogl, K. V. Tugaeva, P. Eberling, C. Kostmann, G. Trave and N. N. Sluchanko, Nat. Commun., 2021, 12, 1677 CrossRef CAS PubMed.
- K. Pennington, T. Chan, M. Torres and J. Andersen, Oncogene, 2018, 37, 5587–5604 CrossRef CAS PubMed.
- B. A. Somsen, P. J. Cossar, M. R. Arkin, L. Brunsveld and C. Ottmann, ChemBioChem, 2024, 25, e202400214 CrossRef CAS PubMed.
- M. Wolter, P. de Vink, J. F. Neves, S. Srdanović, Y. Higuchi, N. Kato, A. Wilson, I. Landrieu, L. Brunsveld and C. Ottmann, J. Am. Chem. Soc., 2020, 142, 11772–11783 CrossRef CAS PubMed.
- Y. Tian, L. Li, L. Wu, Q. Xu, Y. Li, H. Pan, T. Bing, X. Bai, A. V. Finko, Z. Li and J. Bian, J. Med. Chem., 2025, 68, 2124–2146 CrossRef CAS PubMed.
- Y. Liu and R. Hong, Cell Rep. Phys. Sci., 2024, 5, 102141 CrossRef CAS.
- Q. Guo, Y. Jin, X. Chen, X. Ye, X. Shen, M. Lin, C. Zeng, T. Zhou and J. Zhang, Signal Transduction Targeted Ther., 2024, 9, 53 CrossRef CAS PubMed.
- Y. Pan, S. Gangwar, M. Abbas and K. Raza, Sci. Rep., 2025, 15, 11708 CrossRef CAS PubMed.
- B. D. Strahl and C. D. Allis, Nature, 2000, 403, 41–45 CrossRef CAS PubMed.
- A. J. Bannister, P. Zegerman, J. F. Partridge, E. A. Miska, J. O. Thomas, R. C. Allshire and T. Kouzarides, Nature, 2001, 410, 120–124 CrossRef CAS PubMed.
- M. Lachner, D. O’Carroll, S. Rea, K. Mechtler and T. Jenuwein, Nature, 2001, 410, 116–120 CrossRef CAS PubMed.
- C. A. Musselman, M.-E. Lalonde, J. Côté and T. G. Kutateladze, Nat. Struct. Mol. Biol., 2012, 19, 1218–1227 CrossRef CAS PubMed.
- J. Jung and M. S. Werner, Trends Genet., 2026, 42(2), 126–136 CrossRef CAS PubMed.
- P. R. Nielsen, D. Nietlispach, H. R. Mott, J. Callaghan, A. Bannister, T. Kouzarides, A. G. Murzin, N. V. Murzina and E. D. Laue, Nature, 2002, 416, 103–107 CrossRef CAS PubMed.
- D. J. Owen, P. Ornaghi, J. Yang, N. Lowe, P. R. Evans, P. Ballario, D. Neuhaus, P. Filetici and A. A. Travers, EMBO J., 2000, 19, 6141–6149 CrossRef CAS PubMed.
- M. Shogren-Knaak, H. Ishii, J.-M. Sun, M. J. Pazin, J. R. Davie and C. L. Peterson, Science, 2006, 311, 844–847 CrossRef CAS PubMed.
- R. Collepardo-Guevara, G. Portella, M. Vendruscolo, D. Frenkel, T. Schlick and M. Orozco, J. Am. Chem. Soc., 2015, 137, 10205–10215 CrossRef CAS PubMed.
- M. Shvedunova and A. Akhtar, Nat. Rev. Mol. Cell Biol., 2022, 23, 329–349 CrossRef CAS PubMed.
- A. G. Larson, D. Elnatan, M. M. Keenen, M. J. Trnka, J. B. Johnston, A. L. Burlingame, D. A. Agard, S. Redding and G. J. Narlikar, Nature, 2017, 547, 236–240 CrossRef CAS PubMed.
- A. R. Strom, A. V. Emelyanov, M. Mir, D. V. Fyodorov, X. Darzacq and G. H. Karpen, Nature, 2017, 547, 241–245 CrossRef CAS PubMed.
- B. A. Gibson, L. K. Doolittle, M. W. G. Schneider, L. E. Jensen, N. Gamarra, L. Henry, D. W. Gerlich, S. Redding and M. K. Rosen, Cell, 2019, 179, 470–484.e21 CrossRef CAS PubMed.
- D. Willbold, B. Strodel, G. F. Schröder, W. Hoyer and H. Heise, Chem. Rev., 2021, 121, 8285–8307 CrossRef CAS PubMed.
- S. H. W. Scheres, B. Ryskeldi-Falcon and M. Goedert, Nature, 2023, 621, 701–710 CrossRef CAS PubMed.
- M. S. Hipp, P. Kasturi and F. U. Hartl, Nat. Rev. Mol. Cell Biol., 2019, 20, 421–435 CrossRef CAS PubMed.
- S. Wegmann, J. Biernat and E. Mandelkow, Curr. Opin. Neurobiol., 2021, 69, 131–138 CrossRef CAS PubMed.
- S. Wegmann, B. Eftekharzadeh, K. Tepper, K. M. Zoltowska, R. E. Bennett, S. Dujardin, P. R. Laskowski, D. MacKenzie, T. Kamath, C. Commins, C. Vanderburg, A. D. Roe, Z. Fan, A. M. Molliex, A. Hernandez-Vega, D. Muller, A. A. Hyman, E. Mandelkow, J. P. Taylor and B. T. Hyman, EMBO J., 2018, 37, EMBJ201798049 CrossRef PubMed.
- S. Kumar, N. Rezaei-Ghaleh, D. Terwel, D. R. Thal, M. Richard, M. Hoch, J. M. M. Donald, U. Wüllner, K. Glebov, M. T. Heneka, D. M. Walsh, M. Zweckstetter and J. Walter, EMBO J., 2011, 30, 2255–2265 CrossRef CAS PubMed.
- H. Park, T.-I. Kam, V. L. Dawson and T. M. Dawson, Nat. Rev. Neurol., 2025, 21, 32–47 CrossRef PubMed.
- A. Bluhm, S. Schrempel, S. von Hörsten, A. Schulze and S. Roßner, Int. J. Mol. Sci., 2021, 22, 5450 CrossRef CAS PubMed.
- T. J. Cohen, A. W. Hwang, C. R. Restrepo, C.-X. Yuan, J. Q. Trojanowski and V. M. Y. Lee, Nat. Commun., 2015, 6, 5845 CrossRef CAS PubMed.
- D. T. King, J. E. Serrano-Negrón, Y. Zhu, C. L. Moore, M. D. Shoulders, L. J. Foster and D. J. Vocadlo, J. Am. Chem. Soc., 2022, 144, 3833–3842 CrossRef CAS PubMed.
- M. M. Savitski, F. B. M. Reinhard, H. Franken, T. Werner, M. F. Savitski, D. Eberhard, D. M. Molina, R. Jafari, R. B. Dovega, S. Klaeger, B. Kuster, P. Nordlund, M. Bantscheff and G. Drewes, Science, 2014, 346, 1255784 CrossRef PubMed.
- I. Becher, A. Andrés-Pons, N. Romanov, F. Stein, M. Schramm, F. Baudin, D. Helm, N. Kurzawa, A. Mateus, M.-T. Mackmull, A. Typas, C. W. Müller, P. Bork, M. Beck and M. M. Savitski, Cell, 2018, 173, 1495–1507.e18 CrossRef CAS PubMed.
- A. Mateus, N. Kurzawa, I. Becher, S. Sridharan, D. Helm, F. Stein, A. Typas and M. M. Savitski, Mol. Syst. Biol., 2020, 16, MSB199232 CrossRef PubMed.
- X. X. Zhou, L. Z. Fan, P. Li, K. Shen and M. Z. Lin, Science, 2017, 355, 836–842 CrossRef PubMed.
- J. E. Toettcher, O. D. Weiner and W. A. Lim, Cell, 2013, 155, 1422–1434 CrossRef CAS PubMed.
- O. Dagliyan, M. Tarnawski, P.-H. Chu, D. Shirvanyants, I. Schlichting, N. V. Dokholyan and K. M. Hahn, Science, 2016, 354, 1441–1444 CrossRef CAS PubMed.
- N. V. Kukushkin, T. Tabassum and T. J. Carew, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2210478119 CrossRef CAS PubMed.
- K. Islam, Cell Chem. Biol., 2018, 25, 1171–1184 CrossRef CAS PubMed.
- C. A. Ocasio, M. P. Baggelaar, J. Sipthorp, A. L. de la Lastra, M. Tavares, J. Volarić, C. Soudy, E. M. Storck, J. W. Houghton, S. A. Palma-Duran, J. I. MacRae, G. Tomić, L. Carr, J. Downward, U. S. Eggert and E. W. Tate, Nat. Biotechnol., 2024, 42, 1548–1558 CrossRef CAS PubMed.
- M. S. Rana, P. Kumar, C.-J. Lee, R. Verardi, K. R. Rajashankar and A. Banerjee, Science, 2018, 359(6372), eaao6326 CrossRef PubMed.
- Y. Mo, Y. Han, Y. Chen, C. Fu, Q. Li, Z. Liu, M. Xiao and B. Xu, Mol. Cancer, 2024, 23, 274 CrossRef CAS PubMed.
- H. Zhang, Y. Sun, Z. Wang, X. Huang, L. Tang, K. Jiang and X. Jin, Nat. Commun., 2024, 15, 4642 CrossRef CAS PubMed.
- E. H. Garst, H. Lee, T. Das, S. Bhattacharya, A. Percher, R. Wiewiora, I. P. Witte, Y. Li, T. Peng, W. Im and H. C. Hang, ACS Chem. Biol., 2021, 16, 844–856 CrossRef CAS PubMed.
- Z. Liu, S. Xi, L. A. McGregor, K. Yamatsugu, S. A. Kawashima, J. T. Sczepanski and M. Kanai, Angew. Chem., Int. Ed., 2025, 64, e202500162 CrossRef CAS PubMed.
- S. Sakamoto and I. Hamachi, Isr. J. Chem., 2023, 63, e202200077 CrossRef CAS.
- C. Adamson, H. Kajino, S. A. Kawashima, K. Yamatsugu and M. Kanai, J. Am. Chem. Soc., 2021, 143, 14976–14980 CrossRef CAS PubMed.
- P. B. Dervan, Bioorg. Med. Chem., 2001, 9, 2215–2235 CrossRef CAS PubMed.
- D. R. Banerjee, C. E. Deckard, M. B. Elinski, M. L. Buzbee, W. W. Wang, J. D. Batteas and J. T. Sczepanski, J. Am. Chem. Soc., 2018, 140, 8260–8267 CrossRef CAS PubMed.
- M. Jinek, K. Chylinski, I. Fonfara, M. Hauer, J. A. Doudna and E. Charpentier, Science, 2012, 337, 816–821 CrossRef CAS PubMed.
- I. B. Hilton, A. M. D’Ippolito, C. M. Vockley, P. I. Thakore, G. E. Crawford, T. E. Reddy and C. A. Gersbach, Nat. Biotechnol., 2015, 33, 510–517 CrossRef CAS PubMed.
- L. Yuan, Y. Xiong, Y. Zhang, S. Gu and Y. Lei, Mol. Ther., 2026, 34, 46–67 CrossRef CAS PubMed.
- M. A. Cappelluti, V. M. Poeta, S. Valsoni, P. Quarato, S. Merlin, I. Merelli and A. Lombardo, Nature, 2024, 627, 416–423 CrossRef CAS PubMed.
- M. Békés, D. R. Langley and C. M. Crews, Nat. Rev. Drug Discovery, 2022, 21, 181–200 CrossRef PubMed.
- S. U. Siriwardena, D. N. P. M. Godage, V. M. Shoba, S. Lai, M. Shi, P. Wu, S. K. Chaudhary, S. L. Schreiber and A. Choudhary, J. Am. Chem. Soc., 2020, 142, 14052–14057 CrossRef CAS PubMed.
- R. Pergu, V. Sreekanth, P. Kokkonda, S. Lai, P. K. Tiwari, S. Singh, J. Kawai, S. Singha, W. Tian, V. Thimaradka, S. Siriwongsup, K. Tran and A. Choudhary, J. Am. Chem. Soc., 2025, 147, 25316–25324 CrossRef CAS PubMed.
- R. W. Myers, H.-P. Guan, J. Ehrhart, A. Petrov, S. Prahalada, E. Tozzo, X. Yang, M. M. Kurtz, M. Trujillo, D. G. Trotter, D. Feng, S. Xu, G. Eiermann, M. A. Holahan, D. Rubins, S. Conarello, X. Niu, S. C. Souza, C. Miller, J. Liu, K. Lu, W. Feng, Y. Li, R. E. Painter, J. A. Milligan, H. He, F. Liu, A. Ogawa, D. Wisniewski, R. J. Rohm, L. Wang, M. Bunzel, Y. Qian, W. Zhu, H. Wang, B. Bennet, L. L. Scheuch, G. E. Fernandez, C. Li, M. Klimas, G. Zhou, M. van Heek, T. Biftu, A. Weber, D. E. Kelley, N. Thornberry, M. D. Erion, D. M. Kemp and I. K. Sebhat, Science, 2017, 357, 507–511 CrossRef CAS PubMed.
- M. Kanai and Y. Takeuchi, Tetrahedron, 2023, 131, 133227 CrossRef CAS.
- M. Habazaki and M. Kanai, ACS Catal., 2025, 15, 21137–21151 CrossRef CAS.
- Y. Amamoto, Y. Aoi, N. Nagashima, H. Suto, D. Yoshidome, Y. Arimura, A. Osakabe, D. Kato, H. Kurumizaka, S. A. Kawashima, K. Yamatsugu and M. Kanai, J. Am. Chem. Soc., 2017, 139, 7568–7576 CrossRef CAS PubMed.
- Y. Fujiwara, Y. Yamanashi, A. Fujimura, Y. Sato, T. Kujirai, H. Kurumizaka, H. Kimura, K. Yamatsugu, S. A. Kawashima and M. Kanai, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2019554118 CrossRef CAS PubMed.
- Y. Yamanashi, S. Takamaru, A. Okabe, S. Kaito, Y. Azumaya, Y. R. Kamimura, K. Yamatsugu, T. Kujirai, H. Kurumizaka, A. Iwama, A. Kaneda, S. A. Kawashima and M. Kanai, Nat. Commun., 2025, 16, 887 CrossRef CAS PubMed.
- W. Hamajima, A. Fujimura, Y. Fujiwara, K. Yamatsugu, S. A. Kawashima and M. Kanai, ACS Chem. Biol., 2019, 14, 1102–1109 CrossRef CAS PubMed.
- T. Nozaki, M. Onoda, M. Habazaki, Y. Takeuchi, H. Ishida, Y. Sato, T. Kujirai, K. Hanada, K. Yamatsugu, H. Kurumizaka, H. Kimura, H. Kono, S. A. Kawashima and M. Kanai, J. Am. Chem. Soc., 2025, 147, 13732–13743 CrossRef CAS PubMed.
- E. Francis, S. Paylor, C. Van, B. Mathews, M. J. Sobhanian, D. Z. Mansour, G. Hennawi and N. J. Brandt, Expert Rev. Clin. Pharmacol., 2025, 1–16 Search PubMed , ahead-of-print.
- Y. Sohma, T. Sawazaki and M. Kanai, Org. Biomol. Chem., 2021, 19, 10017–10029 RSC.
- M. Furuta, S. Arii, H. Umeda, R. Matsukawa, K. Shizu, H. Kaji, S. A. Kawashima, Y. Hori, T. Tomita, Y. Sohma, H. Mitsunuma and M. Kanai, Adv. Sci., 2024, 11, 2401346 CrossRef CAS.
- N. Nagashima, S. Ozawa, M. Furuta, M. Oi, Y. Hori, T. Tomita, Y. Sohma and M. Kanai, Sci. Adv., 2021, 7, eabc9750 CrossRef CAS PubMed.
- S. Ozawa, Y. Hori, Y. Shimizu, A. Taniguchi, T. Suzuki, W. Wang, Y. W. Chiu, R. Koike, S. Yokoshima, T. Fukuyama, S. Takatori, Y. Sohma, M. Kanai and T. Tomita, Brain, 2021, 144, 1884–1897 CrossRef PubMed.
- R. Matsukawa, M. Yamane and M. Kanai, Chem. Rec., 2023, 23, e202300198 CrossRef CAS PubMed.
- A. Iwai, Y. Hwang, H. Umeda, T. Sawazaki, I. Tomizawa, M. Ono, M. Higuchi, Y. Sohma, Y. Hori, T. Tomita, H. Mitsunuma and M. Kanai, ChemistryEurope, 2025, 4(1), e202500192 CrossRef.
- M. Yamane, H. Umeda, M. Toyobe, A. Iwai, K. Ishihara, G. Kudo, H. Mitsunuma, Y. Hori, T. Tomita, M. Mizuguchi, M. Okada, M. Ueda, Y. Ando, S. A. Kawashima, Y. Sohma, H. Kaji, T. Hirokawa, K. Yamanaka and M. Kanai, J. Am. Chem. Soc., 2025, 147, 28860–28874 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |