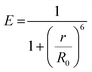Open Access Article
Open Access ArticleMechanistic insights into the design of fluorogenic molecules for wash-free biological applications
Rongrong
Huang
ab,
Syed Ali Abbas
Abedi
ab,
Guanyu
Jiang
ab,
Chao
Wang
c,
Lingya
Peng
d,
Tianruo
Shen
e,
Nabil
Mroweh
b,
Yu
Fang
 d and
Xiaogang
Liu
d and
Xiaogang
Liu
 *a
*a
aSchool of Chemistry, Chemical Engineering and Biotechnology, Nanyang Technological University, 637371, Singapore. E-mail: xiaogang.liu@ntu.edu.sg
bFluorescence Research Group, Singapore University of Technology and Design, 487372, Singapore
cState Key Laboratory of Fine Chemicals, College of Materials Science and Engineering, Shenzhen University, Shenzhen 518071, P. R. China
dKey Laboratory of Applied Surface and Colloid Chemistry, Ministry of Education; Shaanxi Provincial Key Laboratory of New Concept Sensors and Molecular Materials; School of Chemistry and Chemical Engineering, Shaanxi Normal University, Xi’an 710119, P. R. China
eNational Supercomputer Center in Tianjin, 10 West Xinhuan Road, Tianjin 300457, P. R. China
First published on 5th March 2026
Abstract
Conventional fluorescence microscopy is frequently constrained by wash-required labeling protocols. The mandatory removal of unbound probes complicates experimental workflows, perturbs fragile biological environments, and can eliminate weak or transient probe–target interactions. In addition, washing introduces time delays that obscure fast biological dynamics. Wash-free bioimaging has emerged as a powerful alternative, relying on fluorogenic probes that transition from a non-emissive to an emissive state upon target engagement. By eliminating washing steps, these strategies simplify operation, enhance contrast, preserve native biological environments, and enable sustained imaging through continuous exchange between bound and unbound fluorophores. This review establishes a mechanistic framework for the rational design of these wash-free imaging agents. We classify the dominant activation pathways as energy-transfer mechanisms, electron or charge-transfer processes, internal conversion to a dark state, structural isomerization (exemplified by spirocyclization in rhodamine scaffolds), and hydrogen-bond-induced quenching. Beyond these classical modes, we discuss phase-dependent effects such as aggregation-induced emission and disaggregation-induced emission, and highlight emerging paradigms, including in situ fluorophore formation, twisted intramolecular charge shuttle, and conical intersections. By linking photophysical mechanisms to molecular design principles and imaging performance, this review aims to guide the development of next-generation fluorogenic probes for high-contrast, real-time, and sustained imaging across molecular, cellular, and organismal scales.
1. Introduction
Fluorescence microscopy is a central tool in modern life sciences, enabling the visualization of biological structures and dynamic processes with high spatial and temporal resolution.1–11 Despite these strengths, its performance is often limited by conventional wash-required labeling protocols.12–14 These approaches rely on the physical removal of unbound probes, a step that not only increases experimental complexity but also perturbs biological systems. Washing introduces mechanical stress, osmotic changes, and time delays that hinder the observation of rapid or fragile physiological processes. Moreover, weakly bound or transient probe–target interactions are frequently lost during washing, leading to incomplete or biased readouts. In long-term experiments, signal dilution caused by cell division further compromises the ability of washed dyes to support sustained imaging.To address these limitations, bioimaging has increasingly shifted toward wash-free strategies, utilizing fluorogenic probes that remain optically silent (“OFF”, with negligible or weak emissions) until activated (“ON”, with enhanced emissions) by specific target binding.15–19 This OFF–ON behavior underlies wash-free imaging: unbound probes remain dark, minimizing background, while target engagement produces immediate, high-contrast signals without any washing step. Based on this principle, the benefits of wash-free imaging can be organized into four practical dimensions: simplified workflow, improved signal quality and biological integrity, enhanced temporal resolution for rapid dynamics, and support for sustained long-term imaging.
1.1. Simplified workflow
Wash-free protocols streamline experimental workflows by eliminating repeated medium exchanges, centrifugation, or rinsing. This reduction in manual handling minimizes sample loss, a benefit particularly critical when working with small cell populations. The simplified workflow also enhances compatibility with high-throughput, automated, and microfluidic platforms, making these probes ideal for large-scale screening and systems biology applications.20,21 Furthermore, wash-free strategies facilitate multiplexed imaging by enabling the simultaneous or sequential application of orthogonal chemistries without intermediate washing.22–24 Crucially, these protocols enable imaging in scenarios where washing is practically impossible, such as whole-animal or thick-tissue imaging, where unbound probes cannot be removed from systemic circulation.1.2. Improved signal quality and biological integrity
By remaining non-fluorescent until bound, fluorogenic probes yield exceptional signal-to-noise ratios (SNR), ensuring that fluorescence directly reflects the probe–target interaction rather than residual background.25 This mechanism is vital for detecting probes that bind weakly to their targets, as conventional washing steps often strip away these transient binders. Beyond optical fidelity, these protocols preserve the biological fidelity of the specimen. By avoiding the mechanical stress and osmotic shifts associated with washing, wash-free methods maintain cell viability and native morphology, which is essential for studying fragile systems like primary cells and organoids without introducing preparation artifacts.1.3. Enhanced temporal resolution
Wash-free methods offer superior temporal resolution by allowing imaging to commence immediately upon probe application.26–28 This capability eliminates the “blind spot” caused by washing delays, enabling the real-time visualization of rapid biological events such as receptor binding, enzymatic bursts, and ion fluxes. Because the protocol minimizes physical perturbation, these dynamic processes can be observed in their native state with high precision.1.4. Sustained long-term imaging
Finally, fluorogenic dyes are uniquely suited for extended imaging durations. Unlike washed samples, in which the dye pool is fixed and finite, wash-free systems use unbound probes in the medium as a “reservoir”.29,30 These non-emissive molecules can gradually replace photobleached bound fluorophores, thereby sustaining fluorescence over long periods. This reservoir effect is particularly advantageous for tracking proliferating cells. In washed samples, fluorescent dyes are progressively diluted with each cell division, whereas in wash-free systems, continuous uptake from the extracellular reservoir replenishes the intracellular pool, thereby enabling uninterrupted monitoring of cellular dynamics.The realization of these advantages critically depends on the fluorogenicity of the dye. This requirement has prompted a conceptual shift in how fluorescent labels and probes are defined. Traditionally, fluorescent labels are constitutive emitters used for structural visualization, and probes are responsive sensors. Fluorogenic dyes combine these attributes: they target specific structures like labels but remain silent until activated, like sensors. This built-in signal switching enhances contrast and allows for dynamic reporting, effectively merging labeling and sensing functions. As a result, the classical distinction between labels and probes is becoming increasingly fluid in wash-free bioimaging.
To ensure a high fluorogenicity, the molecular design must be grounded in precise photophysical control. Understanding the photophysical mechanisms that govern the fluorogenic “OFF–ON” behavior is essential for the rational design of effective fluorogenic imaging agents. These mechanisms determine how dyes remain non-emissive under basal conditions and activate upon specific molecular interactions.31 Several classical processes underpin this behavior. These include energy transfer mechanisms such as Förster resonance energy transfer (FRET), Dexter energy transfer (DET), and through-bond energy transfer (TBET); electron and charge transfer mechanisms, including photoinduced electron transfer (PET) and twisted intramolecular charge transfer (TICT); internal conversion to dark states (such as those induced by n–π* transitions); and reversible structural rearrangements, exemplified by spirocyclization reactions in rhodamine-based dyes. Hydrogen-bonding interactions between fluorophores and their solvent environment can also significantly influence fluorescence intensity in specific systems. In addition, phase-state transitions play a crucial role: aggregation-induced emission (AIE) activates fluorescence upon clustering, while disaggregation-induced emission (DIE) reverses the aggregation-caused quenching. More recently, emerging mechanisms such as twisted intramolecular charge shuttle (TICS), photoinduced charge centralization (PCC), and conical intersections (CI) have further expanded the molecular toolkit available for probe design. The in situ generation of fluorophores has garnered increasing attention in recent years. Mastering these principles enables chemists to tune the behavior of fluorophores for diverse applications in cellular, tissue, and organism-level imaging.
This review provides a systematic overview of the quenching and activation mechanisms underlying fluorogenic dyes and their applications in wash-free bioimaging (Scheme 1). We present representative molecular designs and highlight how specific photophysical mechanisms have been utilized to develop practical imaging agents. Where appropriate, these discussions are complemented by theoretical models and computational insights to illuminate the fundamental electronic processes involved. Furthermore, we offer a critical evaluation of current strategies, identifying both their advantages and remaining limitations. Our goal is to equip researchers with the conceptual and practical knowledge necessary to advance the design of fluorogenic dyes, concluding with a perspective on emerging opportunities in this rapidly evolving field.
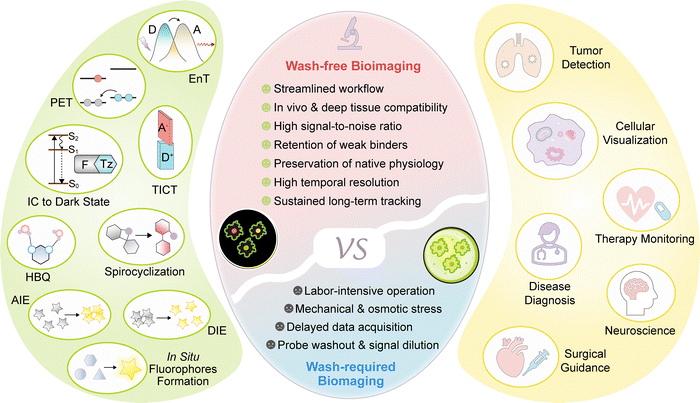 | ||
| Scheme 1 Schematic illustration of the key advantages of wash-free imaging over wash-required methods, the design principles of fluorogenic probes, and their diverse biological applications. | ||
2. Energy transfer
A powerful class of fluorogenic probes for wash-free bioimaging is based on energy transfer-mediated quenching and activation. In this approach, a fluorophore is covalently linked to a quencher, forming a fluorogenic probe. In the absence of the target analyte or triggering event, the excited-state energy of the fluorophore is efficiently transferred to the quencher, rendering the probe non-fluorescent (“dark”) and minimizing background signal. Upon encountering a specific biological target or undergoing a selective chemical reaction, the quencher is cleaved or destroyed. This disruption halts energy transfer, restoring the fluorescence of the probe and enabling sensitive detection or real-time monitoring of the target (Fig. 1a).2.1. Photophysical mechanisms of energy transfer
Three primary energy transfer mechanisms are relevant to the design of such probes: FRET, DET, and TBET.where r is the donor–acceptor distance and R0 is the Förster radius, defined as the distance at which energy transfer efficiency is 50%. The Förster radius depends on the extent of spectral overlap between the donor emission and acceptor absorption spectra, the quantum yield of the donor, and the relative orientation of the donor and acceptor transition dipoles. FRET is highly sensitive to changes in distance and orientation, making it ideal for designing probes where the quencher is removed or spatially separated upon target recognition.
Similar to FRET, DET requires spectral overlap between the donor's emission spectrum and the acceptor's absorption spectrum. This is the “resonance” condition necessary for energy transfer.
The key difference from FRET is the mechanism: DET relies on a quantum-mechanical exchange interaction (not on dipole–dipole coupling), which is highly dependent on spatial proximity and direct orbital overlap. This overlap governs the efficiency of DET and decays exponentially with distance, as described by:
| E ∝ e−2r/L |
DET is particularly relevant when the quencher is directly conjugated or fused to the fluorophore, as this configuration provides the necessary short-range overlap.
Unlike FRET and DET, TBET does not require substantial spectral overlap between the donor emission and acceptor absorption, as energy transfer proceeds through orbital coupling rather than resonance matching. As the donor–acceptor pair behaves as a single conjugated entity, the transfer process closely resembles intramolecular internal conversion.
TBET can be highly efficient and ultrafast when strong electronic coupling is established through a conjugated or fused π-system, enabling efficient quenching. Nevertheless, the intricate details of TBET are not yet fully established and frequently remain a topic of contention.
In practical donor–bridge–acceptor (D–B–A) systems, the distinction between through-bond and through-space energy transfer is often ambiguous. Through-bond (i.e., TBET) and through-space mechanisms (i.e., DET) frequently coexist and compete. The dominant pathway is simply the one with the fastest relative transfer rate, but experimentally disentangling these individual rate contributions is challenging. This ambiguity is more pronounced in flexible molecular systems. Conformational dynamics, such as intramolecular folding, can significantly alter the donor–acceptor distance and orientation, thereby modulating the rates of the distance-dependent FRET and DET pathways. This can accelerate the through-space mechanisms, making them competitive with the through-bond route.
Given the inherent ambiguity in distinguishing between FRET, DET, and TBET pathways, assigning a single dominant mechanism to a quenching phenomenon is often impractical or even misleading. To address this challenge, Liu and co-workers have proposed the term energy transfer to a dark state (ETDS). This term serves as a comprehensive, “mechanistically agnostic” descriptor that collectively accounts for the total energy transfer process without making an unsubstantiated claim about a specific pathway.34 However, when citing specific studies, we may use the nomenclature (e.g., DET, TBET) from the original publication for consistency, with the understanding that these labels represent a simplification of a more complex process.
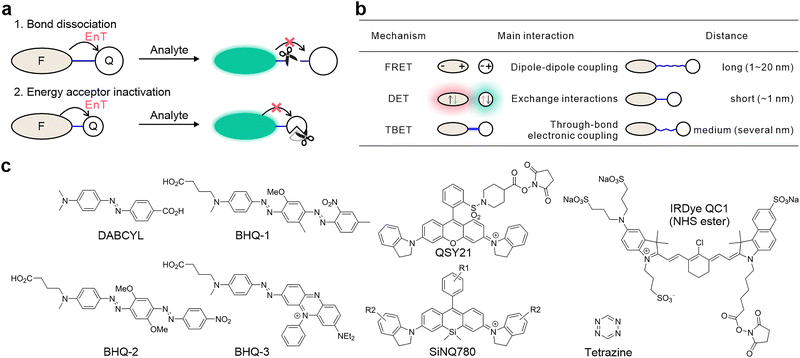
2.2. Commonly used quenchers
The design of fluorescence “OFF–ON” probes for wash-free bioimaging and biosensing relies fundamentally on the judicious selection of a quencher. Among the diverse classes of quenchers developed for this purpose, azo dyes, TICT-prone rhodamine and cyanine dyes, and tetrazine (n–π*) derivatives (Fig. 1c) have emerged as exceptionally versatile and practical.![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) N–), which imparts strong absorption in the visible region and a nonfluorescent nature. The low-lying n–π* states and the TICT rotation of the azo group in azo dyes enable them to efficiently dissipate the absorbed energy via nonradiative decay, making them ideal for FRET-based quenching. The broad absorption bands of azo dyes (typically spanning 400–700 nm) allow for effective quenching of many common fluorophores, especially those emitting in the visible range. Their chemical robustness and ease of conjugation further contribute to their popularity. Notable examples include DABCYL, BHQ-1, BHQ-2, and BHQ-3 (Fig. 1c), which are routinely employed in molecular beacon probes and enzyme substrates.
N–), which imparts strong absorption in the visible region and a nonfluorescent nature. The low-lying n–π* states and the TICT rotation of the azo group in azo dyes enable them to efficiently dissipate the absorbed energy via nonradiative decay, making them ideal for FRET-based quenching. The broad absorption bands of azo dyes (typically spanning 400–700 nm) allow for effective quenching of many common fluorophores, especially those emitting in the visible range. Their chemical robustness and ease of conjugation further contribute to their popularity. Notable examples include DABCYL, BHQ-1, BHQ-2, and BHQ-3 (Fig. 1c), which are routinely employed in molecular beacon probes and enzyme substrates.
A specific example is the system designed by Cornish and Jing, which utilizes a TMP–quencher–fluorophore triad (E1; Fig. 2a).36 In this probe, the binding of the TMP moiety to an engineered eDHFR (L28C) protein initiates a proximity-induced SN2 reaction. This reaction selectively displaces the quencher (tosylate-BHQ1), which achieves two goals simultaneously: it covalently labels the protein and activates the fluorophore (Fig. 2a). This “turn-on” mechanism is what enables the no-wash imaging of intracellular proteins.
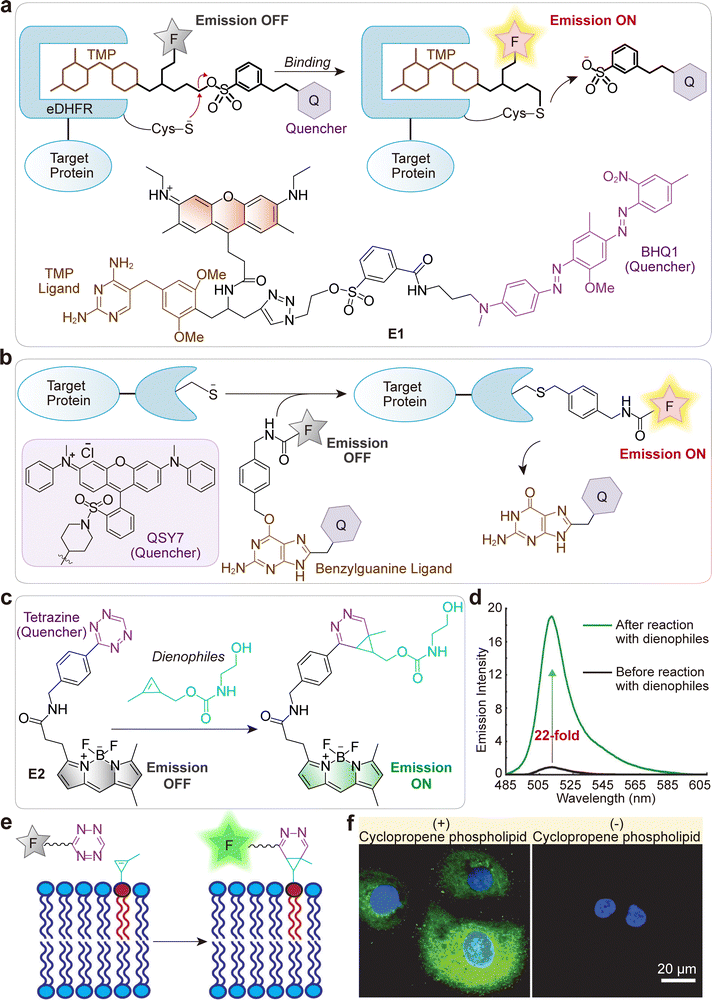 | ||
| Fig. 2 (a) Schematic illustration of the eDHFR protein labeling using fluorogenic TMP-tag E1, based on a trimeric TMP–quencher–fluorophore construct that undergoes a proximity-induced SN2 cleavage. The Cys residue acts as the nucleophile, displacing the tosylate linker upon TMP binding to the protein, thereby activating fluorescence. (b) Schematic representation of SNAP-tag fusion protein labeling using fluorogenic probes with dark quencher. (c) Reaction scheme for the fluorogenic tetrazine–BODIPY probe E2 with dienophiles. (d) Emission spectra of E2 before (black) and after (green) the reaction. (e) Reaction of E2 with membrane-bound cyclopropene phospholipid. The BODIPY chromophore is initially quenched by tetrazine, with fluorescence restored after cycloaddition reaction. (f) Live-cell confocal imaging of cyclopropene phospholipid distribution in SKBR3 cells using E2. Left: Cells incubated with cyclopropene phospholipid followed by probe E2 (green). Right: Control cells treated with E2 (green). Cells were treated with DAPI to visualize the nuclei (blue). Scale bars: 20 µm. (d)–(f) are reproduced from ref. 37 with permission from Wiley-VCH, copyright 2012. | ||
Notable examples include QSY21 and QC1, both of which act as effective quenchers in energy transfer systems (Fig. 1c). Urano and coworkers further developed SiNQ780 (λab ∼ 780 nm), an analog of QSY21 (λab ∼ 560 nm) with a red-shifted absorption,38 thereby expanding the applicability of TICT-based quenchers for near-infrared imaging.
For example, Correa and co-workers systematically developed benzylguanine fluorogenic probes using various dark quenchers (e.g., QSY7, Fig. 2b). This design strategy yielded probes with high quenching efficiencies, thereby permitting wash-free imaging.39
Because the absorbance of tetrazine is relatively weak,42 pronounced quenching effects often require the quencher to be positioned near the fluorophore, either through direct conjugation or via a short linker.
Notably, Devaraj and co-workers developed small, stable methylcyclopropene tags that function as minimal dienophiles for rapid, fluorogenic tetrazine cycloadditions (E2; Fig. 2c).37 These tags react rapidly with tetrazine dyes, a process that yields a strong fluorescence turn-on (Fig. 2d) suitable for live-cell imaging. The effectiveness of this approach was demonstrated by successfully imaging cyclopropene-tagged phospholipids in live human breast cancer cells (Fig. 2e and f).
2.3. Applications in bioimaging
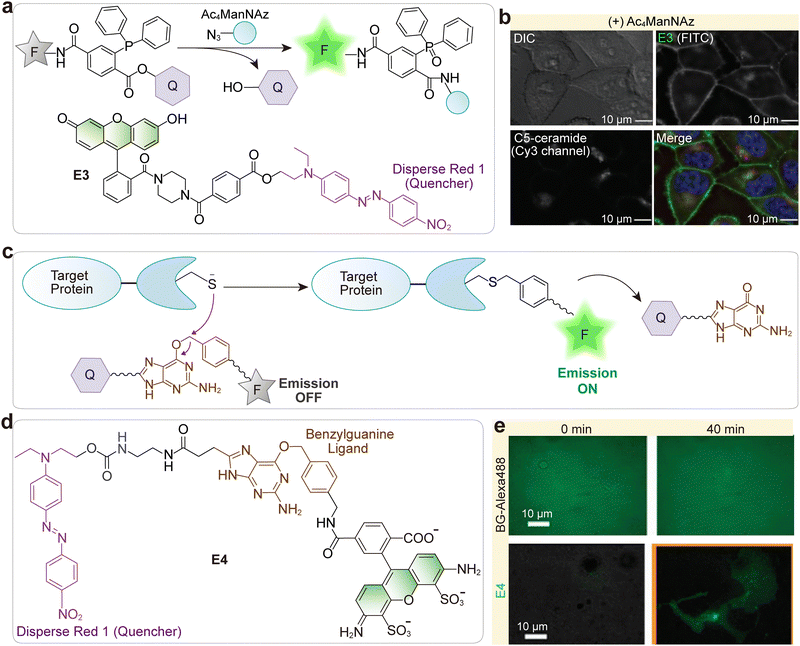 | ||
| Fig. 3 (a) Design of a quenched phosphine–fluorophore system that becomes fluorescent upon Staudinger ligation with azides, illustrated by E3 containing fluorescein and Disperse Red 1 quencher. (b) Fluorescence microscopy of Ac4ManNAz-treated HeLa cells labeled with E3, showing live-cell imaging in the FITC channel (E3) and Cyanine 3 channel (Golgi marker BODIPY TR C5-ceramide). (c) Design of a fluorescently activatable SNAP-tag labeling probe that becomes fluorescent upon binding to the SNAP-tag protein. (d) Chemical structure of the SNAP-tag labeling probe E4. (e) Comparison of the E4 (bottom) with a conventional protein-labeling probe (BG-Alexa488, top) in SNAP–EGFR-expressing COS7 cells. Cells were incubated with BG-Alexa488 or E4. For BG-Alexa488, cells were washed three times with PBS before imaging. (b) and (e) are adapted from ref. 43 and 44 with permission from Wiley-VCH and the American Chemical Society, copyright 2008 and 2011, respectively. | ||
Urano and co-workers designed a FRET-based fluorogenic probe (E4, Fig. 3c and d) for wash-free, real-time imaging of SNAP-tag proteins.44 The probe uses Disperse Red 1 to quench the fluorophore until covalent binding to the SNAP-tag occurs (Fig. 3c), achieving a high fluorescence activation ratio of over 300-fold. They confirmed that incubation with the cell-impermeable DRBG-488 enabled immediate visualization of SNAP-EGFR without the need for washing. In contrast, conventional protein labeling with a non-activatable dye, such as BG-Alexa488, exhibits a strong background signal and necessitates a prolonged washing step after incubation to obtain a clean fluorescence image (Fig. 3e). This FAPL (fluorescence activation-coupled protein labeling) method was successfully employed to continuously monitor the dynamics of protein trafficking, including EGFR endocytosis and exocytosis, in live cells.
Kikuchi and co-workers reported a no-wash, FRET-based labeling system using a mutant β-lactamase tag (BL-tag) and a hydrophilic azopyridinium quencher.45 This design achieved >98% quenching efficiency and the fastest bimolecular labeling rate (7.8 × 104 M−1 s−1) at the time, enabling labeling in under 15 minutes at 10 nM. This speed enabled real-time pulse-chase analysis of protein trafficking, including EGFR dynamics, in live cells.
Weissleder and co-workers reported bioorthogonal turn-on probes where a tetrazine moiety quenches various fluorophores (e.g., BODIPY, Oregon Green) via FRET (E5–E7; Fig. 4a).46 The probes react rapidly with strained dienophiles (i.e., trans-cyclooct-4-enol, TCO), resulting in a 15–21-fold turn-on of fluorescence (Fig. 4b–d). This strategy was demonstrated by the wash-free, high-contrast imaging of a trans-cyclooctene-modified taxol that can bind to the microtubular networks of PtK2 cells (Fig. 4e).
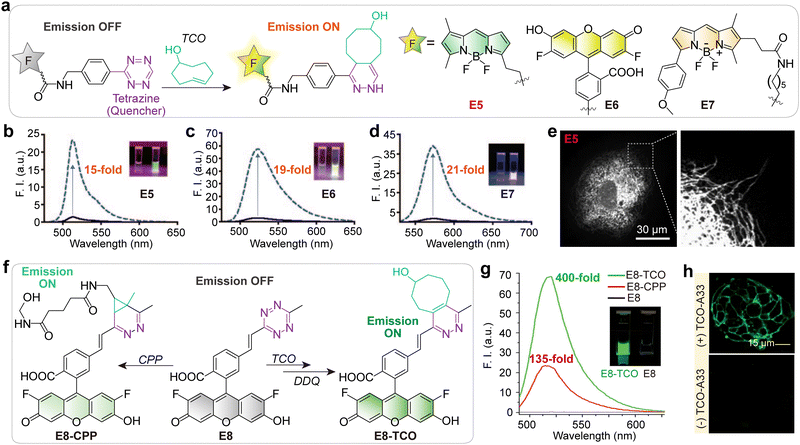 | ||
| Fig. 4 (a) Tetrazine–BODIPY dyes (E5–E7) that rapidly react with trans-cyclooct-4-enol (TCO) via an inverse-electron-demand Diels–Alder cycloaddition, yielding isomeric and emissive dihydropyrazine products. (b)–(d) Emission spectra of the tetrazine probes (black solid lines) and their corresponding dihydropyrazine products (blue dashed lines). Inset photographs compare the visible fluorescence of the tetrazine probes (left cuvettes) and the corresponding dihydropyrazine products (right cuvettes) under UV illumination. (e) Confocal microscopy of PtK2 cells after treatment with trans-cyclooctene taxol, followed by E5 (left), and the expansion of the section indicated by the dashed box reveals that tubular structures are clearly stained (right). Scale bars: 30 µm. (f) Fluorogenic reaction of E8 with cyclopropene (CPP) and trans-cyclooct-4-enol (TCO). Note that the reaction product of tetrazine with TCO can readily be aromatized by oxidation with 2,3-dichloro-5,6-dicyano-1,4-benzoquinone (DDQ). (g) Fluorescence emission spectra for E8 in PBS (gray line), E8-CPP (red line), and E8-TCO (green line) in PBS. Insets are fluorescent images of E8-TCO and E8 under UV light. (h) Live-cell imaging of LS174T cells. Top: Cells were labeled with TCO-conjugated A33 antibodies, washed, and imaged 30 min after addition of E8. Bottom: Cells not expressing TCO were treated with E8 for 30 min and imaged under identical conditions. Scale bars: 15 µm. (b)–(e), (g) and (h) are adapted from ref. 46 and 47 with permission from Wiley-VCH, copyright 2010 and 2014, respectively. | ||
To shorten the distance between the fluorophore and the tetrazine moiety, Wu and Devaraj reported an in situ cascade reaction for synthesizing highly π-conjugated tetrazine probes (E8; Fig. 4f), thereby maximizing TBET-based quenching.47 This method produced ultra-fluorogenic probes with turn-on ratios as high as 400-fold (for an Oregon Green tetrazine) upon reaction with dienophiles (Fig. 4g). The strategy was successfully demonstrated for the wash-free, live-cell imaging of the A33 antigen on the surface of human colon carcinoma cells (Fig. 4h).
Aktalay, Bossi, Hell, and co-workers developed compact, photoactivatable (PaX)–tetrazine dyads (E9–E14; Fig. 5a) for super-resolution nanoscopy.48 The tetrazine quenches both fluorescence (fluorogenicity) and photoactivation until bioorthogonal ligation (Fig. 5b), achieving a 3- to 9-fold fluorescence turn-on. This dual-quenching design enables minimal-linkage-error labeling via genetic code expansion, reaching a median localization precision of 2–3 nm in MINFLUX imaging of vimentin (Fig. 5c).
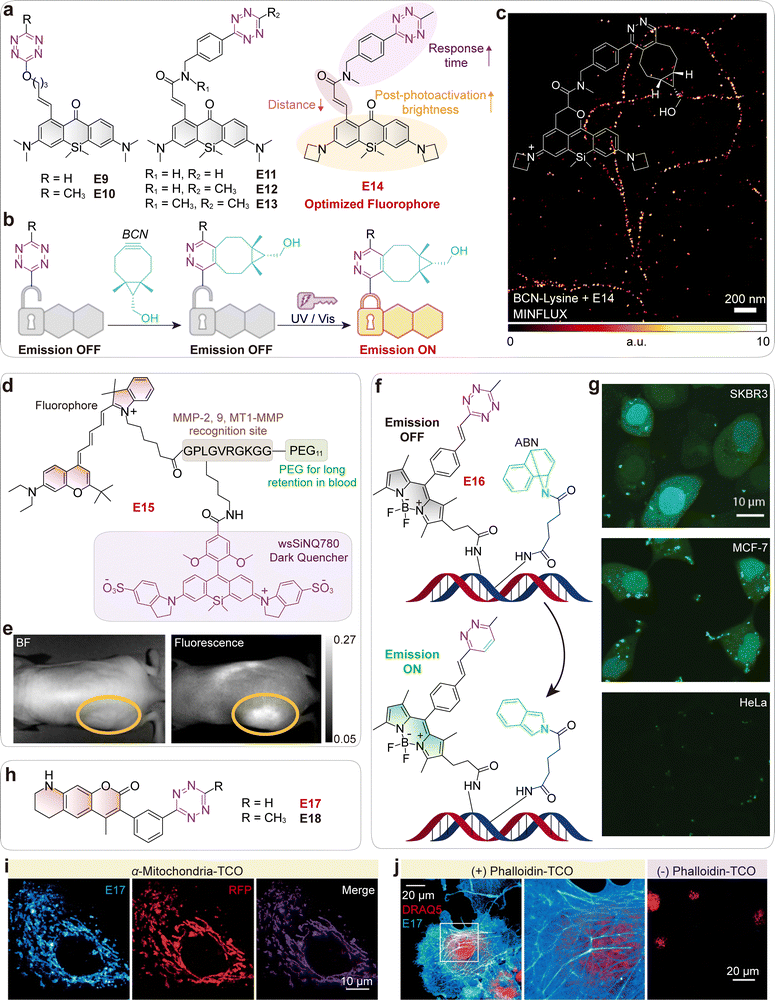 | ||
| Fig. 5 (a) Chemical structures and the structure optimization of photoactivatable (PaX)–tetrazine probes. (b) Schematic illustration of the turn-on of PaX–tetrazine probes via bioorthogonal reaction with bicyclo[6.1.0]nonyne (BCN), followed by photoactivation to the emissive closed-form pyronine fluorophore. (c) MINFLUX image of vimentin filaments in COS-7 cells incorporating endo BCN-L-lysine and labeled with compound E14. Inset shows the chemical structure of the emissive compound derived from E14 after a bioorthogonal reaction with BCN-lysine and subsequent photoactivation. (d) Chemical structure of MMP probe E15 with a SiNQ780 quencher. (e) Fluorescence images of an HT-1080 tumor-bearing nude mouse injected with MMP probe E15via the tail vein. (f) Schematic of a templated transfer reaction between ABN and Tz probe E16. (g) Live-cell detection of microRNA in human cancer cell lines using TMT chemistry. Scale bars: 10 µm. (h) Chemical structures of coumarin probes E17 and E18. (i) Fluorogenic imaging of RFP-tagged mitochondria in OVCA-429 cells. Cells were pretreated with anti-mitochondria–TCO antibody, rinsed, and imaged after addition of E17. (j) Fluorogenic imaging of actin in COS-1 cells. Cells were incubated with phalloidin–TCO and DRAQ5 (red), rinsed, and imaged after addition of E17 (blue). Control images were collected without phalloidin–TCO. (c), (e), (g), (i) and (j) are adapted from ref. 48, 38, 49 and 50 with permission from the American Chemical Society, American Chemical Society, American Chemical Society and Wiley-VCH, copyright 2023, 2015, 2014 and 2014. | ||
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000-fold.50 The design achieves this performance by optimizing TBET-based quenching through direct meta-substitution of the tetrazine on the coumarin (Fig. 5h), a mechanism confirmed to be dominant over FRET. These probes (E17, E18; Fig. 5h) enabled rapid, no-wash, real-time imaging of diverse biological targets, including mitochondria (Fig. 5i) and actin (Fig. 5j).
000-fold.50 The design achieves this performance by optimizing TBET-based quenching through direct meta-substitution of the tetrazine on the coumarin (Fig. 5h), a mechanism confirmed to be dominant over FRET. These probes (E17, E18; Fig. 5h) enabled rapid, no-wash, real-time imaging of diverse biological targets, including mitochondria (Fig. 5i) and actin (Fig. 5j).
Wieczorek and Wombacher synthesized a series of green-to-far-red-emitting fluorogenic tetrazine probes based on xanthene dyes (E19–E27; Fig. 6a and b).51 The design uses minimal interchromophore distance to maximize quenching (attributed to Dexter energy transfer), achieving turn-on ratios up to 109-fold (for E24). This work provided the first example of intracellular, no-wash live-cell protein imaging with tetrazine probes, targeting actin, mitochondria (Tom20-eDHFR, Fig. 6c), and nuclei (H2B-eDHFR, Fig. 6d).
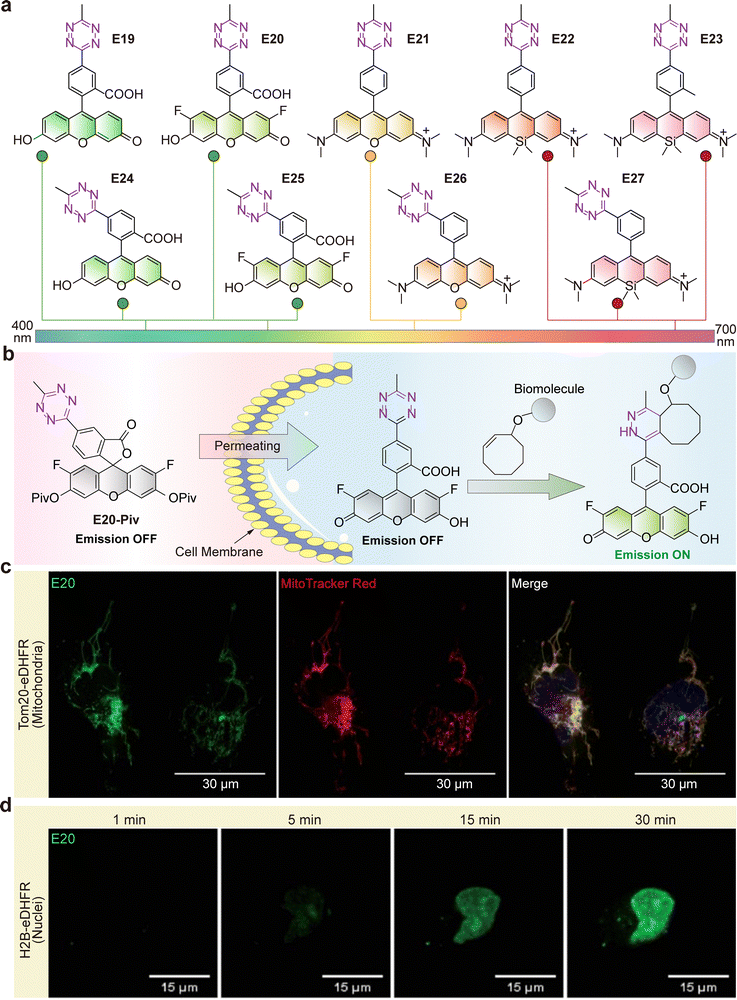 | ||
| Fig. 6 (a) Chemical structures of the green-to far-red-emitting fluorogenic xanthene–tetrazine probes (E19–E27). (b) Schematic representation of live cell labeling with E20-Piv. (c) and (d) No-wash confocal live cell imaging of (c) mitochondria and (d) nuclei in HeLa cells using E20-Piv. Live HeLa cells expressing Tom20-eDHFR or H2B-eDHFR were incubated with TMP-TCO and treated with E20-Piv. (c) and (d) are adapted from ref. 51 with permission from the Royal Society of Chemistry, copyright 2017. | ||
2.4. Advantages and design versatility
Energy transfer (EnT) based quenching mechanisms have emerged as a powerful solution for wash-free bioimaging, offering a class of fluorogenic probes that remain “dark” until they interact with their specific target.The foremost advantage of this strategy is the potential for exceptionally high signal-to-background ratios. In a well-designed probe, an efficient quencher held in proximity to the fluorophore can reduce background fluorescence to near-zero levels. Upon activation, the disruption of this quenching results in a massive increase in signal. While FRET-based systems offer substantial gains, mechanisms involving stronger electronic coupling, such as DET and TBET, afford substantially higher turn-on ratios, with reported signal enhancements in the hundreds to thousands.
EnT-based designs are remarkably modular. The selection of different donor–acceptor pairs provides fine-tuning of the spectral properties, enabling researchers to develop probes across various wavelengths and for multiplexed imaging.
This modularity has been ingeniously combined with bioorthogonal chemistry. Tetrazines, for example, function as highly efficient quenchers for many dyes. In this dual-purpose system, the tetrazine not only quenches the dye but also serves as a reactive handle. The bioorthogonal reaction with a dienophile, such as a cyclopropene, simultaneously removes the quenching effect and covalently links the dye to its target, providing an ideal mechanism for single-step, no-wash labeling.
Finally, EnT-based probes offer significant advantages in robustness. Unlike quenching mechanisms based on PET, which can be highly sensitive to local environmental changes (i.e., pH and solvent polarity), EnT is a through-space/bond mechanism that is comparatively stable. This makes EnT-based probes more reliable for quantitative measurements across the heterogeneous, diverse compartments of a living cell.
When paired with fast reaction kinetics, as seen in tetrazine cycloadditions, these probes can respond rapidly, enabling the real-time visualization of dynamic cellular processes.
2.5. Limitations and practical challenges
Despite these strengths, the practical implementation of energy-transfer-based quenching poses significant challenges, creating a persistent gap between theoretical design and robust biological performance.The primary bottleneck lies in the extreme spatial sensitivity of FRET-based systems. As FRET efficiency scales with the inverse sixth power of the distance (E ∝ 1/r6), even sub-nanometer fluctuations in the molecular scaffold can lead to dramatic signal instability. This sensitivity is compounded by the current lack of predictive models that can accurately account for the relative orientation of dyes within complex cellular environments, making the rational design of high-performance FRET probes a process of trial and error rather than precision engineering.
This structural sensitivity extends to the linker design, which presents a fundamental mechanical paradox: the linker must be rigid enough to maintain an efficient quenching geometry in the “OFF” state, yet sufficiently flexible to undergo the rapid conformational or chemical changes required for activation. For TBET or other covalently coupled systems, this often requires conjugated or rigid linkers that properly align the electronic orbitals, which can complicate synthesis and potentially reduce probe solubility or cell permeability.
Furthermore, achieving a near-zero background signal remains an elusive goal. Even with optimized designs, achieving a perfect “OFF” state is difficult. Incomplete quenching, particularly for fluorophores with high intrinsic quantum yields, can result in a measurable residual background that limits the probe's dynamic range, particularly for FRET systems.
Lastly, a critical but often overlooked bottleneck is the chemical and photochemical stability of the quencher. There is a persistent trade-off between reactivity and stability. For instance, the high electrophilicity of tetrazines, while beneficial for rapid bioorthogonal activation, renders them susceptible to nucleophilic attack by intracellular thiols. This chemical instability limits the longevity of these probes, preventing the continuous monitoring of biological processes over extended periods.
2.6. Outlook and future directions
Future research aims to overcome these limitations. A primary goal is to develop more effective quenchers with enhanced chemical and photochemical stability. Recent work on stabilized difluoroboronated tetrazines and other novel quencher motifs represents a promising step toward this goal.52To move toward in vivo applications, there is a strong emphasis on advancing EnT strategies into the near-infrared (NIR) I and II windows, which requires developing new, bright, stable long-wavelength dyes and long-wavelength quenchers to enable deep-tissue imaging.
The simultaneous imaging of multiple cellular processes is a key frontier that requires the development of orthogonal probe systems. This orthogonality must be both spectral, utilizing distinct colors such as green and red, and employing activation mechanisms that do not cross-react. Such a design enables the independent, wash-free visualization of multiple distinct molecular events within the same cell.
Finally, to move beyond trial and error, computational design is becoming increasingly essential. The routine calculation of Förster radii (R0), excited-state energies, and electronic coupling constants will expedite the rational design of new probes. Furthermore, molecular dynamics simulations could help predict the effects of linker flexibility and target-environment interactions on probe conformation and quenching efficiency.
It is also essential to recognize that tetrazines can induce quenching mechanisms beyond classical energy transfer, including electron transfer and internal conversion pathways.41 The diversity of quenching mechanisms, combined with the small size and chemical versatility of tetrazines, makes them uniquely suited for constructing compact, highly responsive fluorogenic probes. Further details on the photophysical behaviours of tetrazine quenchers can be found in recent comprehensive reviews.40,41
3. Photo-induced electron transfer
Whereas the energy transfer mechanisms discussed above rely on the redistribution of excitation energy between a fluorophore and a quencher, PET modulates fluorescence through transient electron transfer between the fluorophore and an electronically coupled donor or acceptor. This shift from energy-centric to electron-centric quenching introduces redox properties as a central design parameter, enabling more precise and predictable tuning of fluorescence responses.A typical PET probe consists of a fluorescent reporter (fluorophore) and a quencher unit that are electronically coupled. In its initial “OFF” state, excitation of the fluorophore leads to rapid electron transfer with the quencher, a non-emissive pathway that effectively darkens the probe. This quenching process can be disrupted by a specific trigger, such as target binding or enzymatic cleavage, which alters the electronic properties of the system.53,54 This disruption inhibits electron transfer, forcing the excited fluorophore to relax through its emissive pathway and restoring bright fluorescence.55,56
3.1. The mechanistic framework of PET
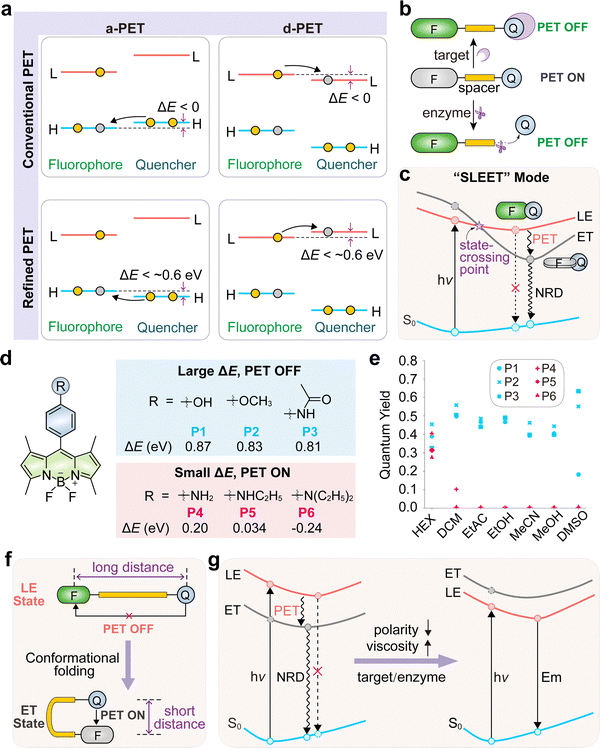
The operational principle of PET hinges on a finely balanced competition between two distinct excited states. The first is a radiative, locally excited (LE) state confined to the fluorophore, which gives rise to fluorescence. The second is a non-radiative electron-transfer (ET) state that forms between the fluorophore and the quencher, thereby silencing emission. The relative stability of these LE and ET states thus dictates the probe's fluorescence output. This property can be rationally engineered through molecular design and modulated by the local microenvironment.PET systems are generally classified based on two key features: the direction of electron flow and the structural linkage between the active components. In acceptor-PET (a-PET), the excited fluorophore accepts an electron from the quencher. Conversely, in donor-PET (d-PET), the excited fluorophore donates an electron to the quencher (Fig. 7a).57,58 This distinction is fundamental to probe design, as it dictates the required redox properties of the fluorophore and quencher components. Furthermore, the fluorophore and quencher can be joined by a flexible covalent spacer or linked directly.59 Spacer-linked systems offer modularity, allowing for tunable electronic coupling and the incorporation of cleavable moieties for reaction-based sensing (Fig. 7b).60–62 In directly linked systems, the components often adopt a pre-twisted or near-orthogonal geometry that facilitates electron transfer upon excitation (Fig. 7c). The short donor–acceptor distance further enhances transfer efficiency.63,64
The efficiency of PET quenching is not static but can be dynamically tuned by several physical and chemical factors, providing multiple avenues for designing environmentally responsive probes. The ET state is characterized by significant charge separation, making it highly stabilized by polar solvents. Consequently, PET quenching is most efficient in polar environments like water, rendering the probes dark. In contrast, non-polar microenvironments, such as the interior of lipid droplets or certain protein cavities, destabilize the ET state relative to the LE state, thereby suppressing PET and switching fluorescence on.65–67 For instance, Liu, Xu, and co-workers systematically demonstrated this effect with a series of PET probes whose fluorescence quantum yields were near zero in polar solvents but increased dramatically in nonpolar solvents (P1–P6; Fig. 7d and e), confirming the critical role of polarity in regulating emission.68
Conformational dynamics are also often coupled to the PET process. In directly linked systems, rotation of the receptor relative to the fluorophore enables near-orthogonal alignment, which maximizes charge separation (Fig. 7c).55,69 In spacer-linked systems, electron transfer generates oppositely charged fluorophore and quencher fragments, which can fold towards each other due to electrostatic attraction (Fig. 7f).70–72 High viscosity or steric confinement restricts the molecular motions required for optimal electron transfer, impeding the formation of the ET state and leading to fluorescence enhancement. Studies in solvent mixtures of constant polarity but varying viscosity have confirmed that restricting intramolecular motion is sufficient to suppress PET.63,68
In biological systems, polarity and viscosity are often interconnected. The hydrophobic pockets of proteins, for instance, are both less polar and more sterically confined than the aqueous cytoplasm. This combination works synergistically to suppress PET (Fig. 7g), making PET-based probes highly effective for reporting on protein binding.
Finally, the fundamental driving force for PET—the difference in redox potentials between the donor and acceptor—can be chemically modulated, such as cleavage of the quencher from the fluorophore (terminating PET) or chemical modification of the quencher to shift its redox potential.62,73,74 A prime example is pH sensing, where the protonation state of a quencher, such as an aniline group, modulates its ability to donate electrons, as demonstrated by Urano and co-workers in designing probes for acidic organelles.75
3.2. Computational modeling of PET processes
Theoretical models are indispensable for understanding PET and guiding the rational design of new probes. These models range from simple thermodynamic estimates to sophisticated quantum chemical calculations. The Rehm–Weller equation provides a foundational thermodynamic estimate of the free energy change for electron transfer (ΔGET):76| ΔGET = Eox – Ered – E00 – C |
Marcus’ theory offers a more complete kinetic description by incorporating the reorganization energy (λ) and the electronic coupling (V):78,79
kET ∝ V2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) exp[−(ΔGET + λ)2/4λkBT] exp[−(ΔGET + λ)2/4λkBT] |
A simplified yet popular approach is the frontier molecular orbital (FMO) model, which predicts PET feasibility based on the relative energies of the donor and acceptor orbitals.53 The frontier molecular orbital (FMO) approach represents the simplest and therefore the most widely used method for semi-quantitative prediction of PET tendencies. In this framework, donor-PET (d-PET) is expected when the LUMO of the quencher lies below the LUMO of the fluorophore. Conversely, acceptor-PET (a-PET) is predicted to occur when the HOMO of the quencher is positioned above the HOMO of the fluorophore. While this method offers an intuitive orbital-based criterion, it is accompanied by several limitations. Chief among them is the neglect of solvent effects: stabilization of the charge-separated ET state relative to the LE state can dramatically shift the balance between emissive and quenched states in polar versus nonpolar environments.68 In addition, the approach overlooks conformational changes that often occur upon excitation. These structural rearrangements, especially in flexible fluorophore–spacer–quencher systems, can play a decisive role in determining whether electron transfer is feasible.72
For reliable predictions, time-dependent density functional theory (TD-DFT) calculations are essential to accurately map the excited-state energy landscape. Specifically, long-range corrected functionals (e.g., CAM-B3LYP, ωB97X-D) or hybrid functionals with high Hartree–Fock exchange (e.g., M06-2X) must be employed to mitigate the self-interaction errors inherent in standard hybrid functionals, which notoriously underestimate the energy of charge-transfer states. Theoretical frameworks such as the SLEET model (state-crossing from a locally excited to an electron transfer state), proposed by Liu and co-workers, aim to provide a quantitative assessment of PET thermodynamics. However, the accuracy of such models relies heavily on careful benchmarking and the rigorous treatment of environmental effects.81
3.3. Applications in bioimaging
The versatility of the PET mechanism has enabled its successful application in imaging a wide range of biological targets and processes.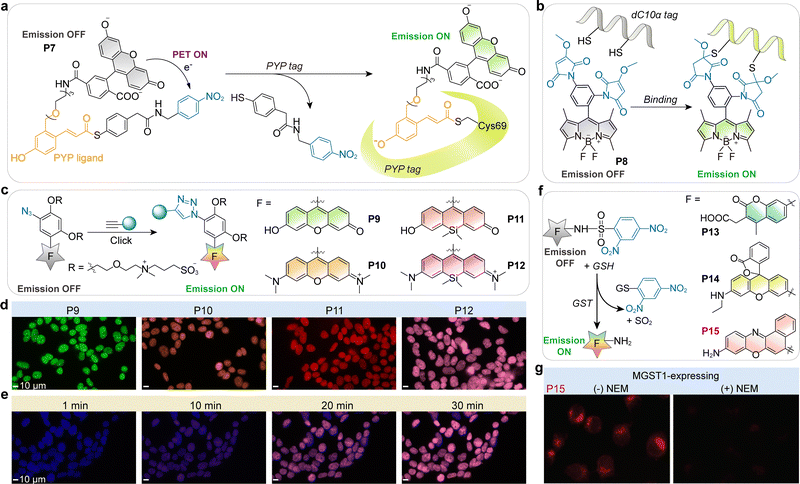 | ||
| Fig. 8 (a) Schematic illustration of the turn-on response of P7 upon binding to PYP-tag. (b) Protein labeling-triggered emission turn-on of a BODIPY dimaleimide fluorophore P8. (c) PET-based fluorogenic CalFluor probes P9–P12 activated by click chemistry. (d) No-wash labeling of EdU-labeled HEK 293T cells treated with CalFluor probes P9–P12. EdU is incorporated into DNA during replication, allowing selective detection of proliferating cells. Scale bars: 10 µm. (e) Time-dependent two-color imaging of HEK 293T cells using Hoechst 33342 (blue) for nuclear staining and P10 (magenta) for click-based fluorescent labeling. Scale bars: 10 µm. (f) Fluorogenic glutathione transferase (GST) substrates (P13–P15), and the switch-on enzymatic cleavage reaction triggered by glutathione (GSH). (g) Fluorescence microscopic imaging of MGST1-expressing live MCF7 cells incubated with P15. The left panel shows cells without N-ethylmaleimide (NEM) treatment, while the right panel shows cells pretreated with NEM to enhance MGST1 enzymatic activity. (d), (e) and (g) are adapted from ref. 83 and 84 with permission from the American Chemical Society, copyright 2015 and 2011, respectively. | ||
Building on this concept, Keillor and colleagues engineered thiol-reactive BODIPY dimaleimides (P8; Fig. 8b). In their unreacted state, the maleimide groups acted as strong electron acceptors, quenching BODIPY emission through PET.55 Covalent reaction with cysteine residues neutralized this quenching pathway, yielding bright intracellular signals. It is also worth noting that changes in the local environment induced by protein tag binding significantly suppress the PET mechanism. The fluorogenic nature of these probes significantly reduces background fluorescence, making them particularly effective for labeling cysteine-containing proteins, such as dC10α-tagged constructs, in live cells.
Vendrell and colleagues expanded the scope further, showing that direct conjugation of BODIPY to the tryptophan residue of an antifungal peptide produced probes (e.g., P16, Fig. 9a) that were non-emissive in aqueous solution but became strongly fluorescent upon insertion into fungal membranes.85 This spacer-free design ensured efficient PET quenching in polar media while switching on in the hydrophobic, viscous membrane environment. The dual influence of reduced polarity and increased viscosity suppressed PET and non-radiative decay, enabling high-contrast, wash-free imaging of fungal infections (Fig. 9b).
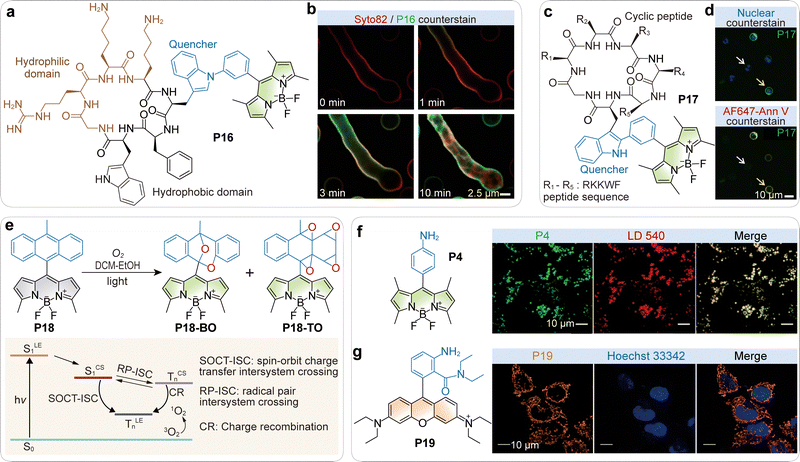 | ||
| Fig. 9 (a) Chemical structure of the cyclic BODIPY-labeled peptide P16. (b) Time-lapse high-resolution fluorescent imaging of A. fumigatus upon incubation with compound P16 (green) and a cell membrane counterstain (red). (c) Chemical structure of the cyclic BODIPY-labeled peptide P17. (d) Representative fluorescence confocal microscopy images of viable (white arrows) and apoptotic cells (yellow arrows) after incubation with P17 (green). Cells were co-stained with Hoechst 33342 (blue, top) and AF647-Annexin V (red, bottom) as nuclei and apoptosis markers, respectively. (e) Photoinduced transformation of P18, and the schematic illustration of the PET-initiated generation of singlet oxygen. (f) Chemical structures of BODIPY-based PET probe P4, and the co-staining images of HeLa cells using P4 (green) and LD 540 (red). (g) Chemical structures of rhodamine-based PET probe P19, and the co-staining of HeLa cells using P19 and Hoechst 33342. Orange channel, P19; blue channel, Hoechst 33342. (b), (d), (f) and (g) are adapted from ref. 85, 64 and 68 with permission from Springer Nature, Springer Nature and the American Chemical Society, copyright 2016, 2020 and 2020, respectively. | ||
In later work, the same group reported P17 (Fig. 9c), a BODIPY–cyclic peptide conjugate that selectively binds phosphatidylserine (PS) exposed on apoptotic cell membranes.64 In solution, the probe remained quenched, but binding to PS-rich domains triggered a strong turn-on response. This environment-driven switching provided a robust, wash-free alternative to annexin V staining for apoptosis imaging (Fig. 9d).
Other innovative PET-based mechanisms have also emerged. Senge and co-workers developed BODIPY–anthracene dyads (e.g., P18; Fig. 9e) in which PET initiated charge separation, followed by spin–orbit charge transfer intersystem crossing to generate triplet states (Fig. 9e).86 Subsequent singlet oxygen production led to oxidative reactions at the anthracene moiety, yielding strongly fluorescent products. Although not designed strictly as no-wash probes, these systems offered fluorogenic reporters for singlet oxygen generation in living cells, informing the development of heavy-atom-free triplet sensitizers.
Wu and co-workers extended this concept into the far-red and near-infrared (NIR) region by introducing an amino-tetrazine substituent at the meso-position of various BODIPY, rhodamine, and cyanine fluorophores (P20–P26; Fig. 10a).81 This created a PET pathway that efficiently quenched emission until a click reaction abolished PET, achieving unprecedented turn-on ratios of up to 1459-fold. These probes proved highly effective for wash-free mitochondrial imaging (Fig. 10b–f) and in vivo tumor imaging (Fig. 10g), showcasing the robustness of PET-based bioorthogonal chemistry.
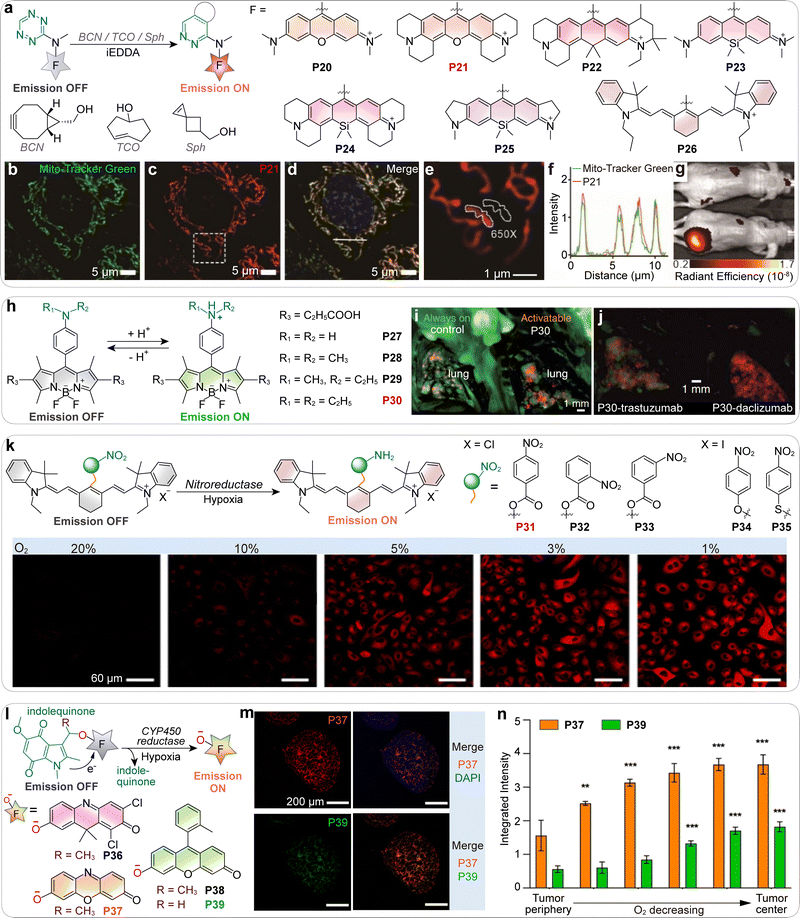 | ||
| Fig. 10 (a) Fluorogenic reactions between tetrazine probes P20–P26 and dienophiles. (b)–(d) Colocalization imaging of mitochondria using Mito-Tracker Green (green) and P21 (red). Scale bars: 5 µm. (e) Comparison of pixel intensity between regions of interest located within and outside mitochondria, in the boxed area shown in panel (c). Scale bar: 1 µm. (f) Plot of fluorescence intensity along the white line shown in panel (d). (g) In vivo imaging of mice using P22. Mice bearing tumor xenografts were intratumorally injected with saline (top) or TCO-TPP (bottom), followed by administration of P22 after 5 min. (h) The reversible and acidic pH-induced fluorescence activation of P27–P30. (i) Composite overlapped images of the mouse lung 1 day after injection of always-on probe (left) and pH-activatable probe P30 (right). (j) Spectrally unmixed and composite overlapped images of the lung 1 day after injection of pH-activatable antibody probes. Left: HER2-specific probe (P30-trastuzumab) selectively highlights HER2+ tumors (green) in the lung, clearly distinguishing them from RFP-labeled HER2− tumors (red). Right: Control probe (P30-daclizumab, targeting CD25) shows no green signal, indicating a lack of HER2+ tumor labeling; only RFP+HER2− tumors (red) are visible. This comparison demonstrates that fluorescence activation occurs only when the probe binds and is internalized by HER2+ cells, confirming the probe's target specificity and pH-dependent activation. (k) Oxygen-dependent nitroreductase-activated fluorescence turn-on of probes P31–P35, and the confocal fluorescence microscopy imaging of A549 cells incubated with P31 under different oxygen concentration conditions. Scale bars: 60 µm. (l) Oxygen-dependent cytochrome P450 (CYP450) reductase (PH51)-catalyzed reduction of P36–P39 to give the corresponding emissive fluorophores. (m) Representative confocal fluorescence images of cryo-sectioned multicellular spheroids incubated with hypoxia-sensitive probes. Top left: Fluorescence signal from compound P37, indicating regions of moderate hypoxia (∼4% O2). Top right: Merged image of compound P37 (green) and DAPI (blue), showing probe distribution relative to cell nuclei. Bottom left: Fluorescence signal from compound P39, indicating severe hypoxia (<0.5% O2). Bottom right: Merged image of compound P37 (green) and compound P39 (red), highlighting spatial differences in hypoxia levels within the spheroid. Scale bars: 200 µm. (n) Quantification of the fluorescence intensity of compounds P37 and P39 in spheroids treated as in (m). (b)–(g), (i), (j), (k), (m) and (n) are adapted from ref. 81, 75, 87 and 88 with permission from Wiley-VCH, Springer Nature, the American Chemical Society and the American Chemical Society, copyright 2022, 2009, 2015 and 2023, respectively. | ||
Li and co-workers later developed nitroaromatic heptamethine cyanines as hypoxia-sensitive probes (P31–P35; Fig. 10k).87 In these systems, PET quenching by the nitro group was relieved upon enzymatic reduction under hypoxia, restoring NIR fluorescence and providing strong tumor-to-background contrast. More recently, Wallabregue and colleagues introduced indolequinone-based probes (P36–P39; Fig. 10l) capable of distinguishing mild from severe hypoxia in tumors.88 Reduction of the electron-withdrawing indolequinone restored fluorescence (Fig. 10l), with different scaffolds tuned to specific oxygen thresholds. This enabled two-color wash-free hypoxia imaging (Fig. 10m), providing a powerful means of mapping tumor oxygen heterogeneity without additional staining (Fig. 10n).
3.4. Advantages and design versatility
PET stands as one of the most rationally designable mechanisms for wash-free bioimaging, primarily because well-defined thermodynamic principles govern its activation. The feasibility of electron transfer can typically be predicted using the Rehm–Weller equation or by analyzing frontier orbital energies. This predictive power enables a modular design strategy in which diverse recognition units—ranging from cleavable linkers to protonatable amines—can be “plugged into” fluorophores to generate reliable “OFF–ON” switches. In the “OFF” state, fluorescence is efficiently quenched by rapid electron transfer, ensuring low background signals and exceptional imaging contrast upon analyte binding.The defining strength of PET is its structural economy. Because modulation can be achieved with small substituents such as aliphatic amines or aniline derivatives, PET probes generally retain favorable physicochemical properties, including membrane permeability and solubility. This compact footprint contrasts sharply with energy-transfer-based approaches, which often require bulky quenchers, making PET particularly advantageous for intracellular sensing of pH and small ions, such as Ca2+, Zn2+, and ClO−.
Furthermore, PET is highly adaptable to diverse biological triggers. It can be activated by irreversible reactions, such as tetrazine click chemistry or enzymatic reduction of nitro groups, as well as by reversible interactions, such as pH-dependent protonation. This versatility supports the development of probes for a wide array of targets, including specific enzymes (e.g., GST, nitroreductase), physiological states (e.g., hypoxia, acidosis), and cell death pathways. Additionally, because PET is inherently sensitive to local polarity and viscosity, it can be exploited to image hydrophobic microenvironments—such as lipid droplets and protein pockets—without requiring chemical cleavage.
3.5. Limitations and practical challenges
Despite these strengths, implementing PET probes faces distinct challenges, particularly the inherent trade-off between sensitivity and environmental cross-reactivity. A major bottleneck is the “microenvironment-induced false positive” effect. The same solvent-polarity and viscosity sensitivity that enables membrane imaging often leads to non-specific activation in hydrophobic or highly viscous compartments, such as lipid bilayers or serum albumin aggregates. These environments can unintentionally destabilize the ET state, suppressing electron transfer regardless of the analyte's presence. This lack of selectivity creates a significant barrier to achieving high signal-to-noise ratios in heterogeneous biological systems, where non-specific accumulation is often indistinguishable from target binding.Protonation-driven suppression is a secondary yet equally critical bottleneck in rational design. Because amine-based quenchers are inherently basic, their lone pair availability—and thus their quenching efficiency—is tightly coupled to local pH. This dependency introduces a “location-dependent liability”: probes targeting non-pH analytes often undergo spontaneous activation when sequestered in acidic organelles (e.g., lysosomes). Decoupling the redox potential from the pKa of the receptor remains a fundamental challenge in expanding the utility of PET probes in complex intracellular environments.
Furthermore, the field suffers from a “kinetic-thermodynamic gap”. While the thermodynamic feasibility (determined by the Rehm–Weller equation) is easily calculated, the actual quenching efficiency is governed by electron transfer kinetics, which are sensitive to solvent reorganization and conformational dynamics. The current lack of computationally efficient models to predict these kinetic rates often relegates PET probe development to an empirical “trial-and-error” approach, hindering the precise tuning of “OFF” state leakiness and “ON” state recovery.
Finally, the photostability of PET probes is often compromised by the electron transfer mechanism itself. The radical ion pair intermediates generated during this process are chemically reactive and susceptible to irreversible degradation. Crucially, the inevitable population of long-lived triplet states via charge recombination (CR) acts as a “photochemical trap”, facilitating the generation of reactive oxygen species (ROS). This mechanism-induced photobleaching creates a fundamental paradox: the more efficient the quenching in the “OFF” state, the higher the risk of the probe degradation during continuous imaging.
3.6. Outlook and future directions
Future progress in PET probe design will rely on the rigorous integration of advanced computational methods with synthesis. Moving beyond simple thermodynamic estimates, the routine application of TD-DFT and molecular dynamics simulations will allow for more accurate predictions of excited-state energetics and conformational gating. This shift from empirical screening to predictive modeling is essential for minimizing trial and error in probe development.Extending PET mechanisms into the near-infrared (NIR) region remains a critical frontier. Long-wavelength dyes inherently possess small energy gaps, severely limiting the range of orbital energies available for effective donor–acceptor alignment. Overcoming this constraint will require the engineering of new electron-deficient NIR scaffolds and the identification of tailored quenchers through theory-driven screening. Additionally, exploring inter-fluorophore PET processes, where fluorophores act as mutual quenchers, offers a promising pathway to efficient long-wavelength switching.
Finally, the development of logic-gated PET systems presents a robust solution to the challenges of specificity. By coupling PET with complementary mechanisms such as FRET or caging groups that require coincident biological triggers, researchers can effectively filter out environmental noise. These next-generation designs will ensure that PET probes report strictly on the intended biological event, significantly improving reliability in complex in vivo settings.
4. Twisted intramolecular charge transfer
Whereas PET typically involves electron exchange between a fluorophore and an auxiliary quencher unit, electronic quenching can also arise from structural movements within the fluorophore itself. The most prominent example is TICT, where bond rotation creates a non-emissive state.Specifically, TICT describes the process in which an excited-state fluorophore undergoes intramolecular bond rotation to form a perpendicular, charge-separated state. This TICT state is typically non-emissive and serves as an intrinsic quenching pathway (Fig. 11a). In aqueous or polar solutions, fluorophores with strong TICT tendencies are therefore weakly fluorescent or completely dark.89,90
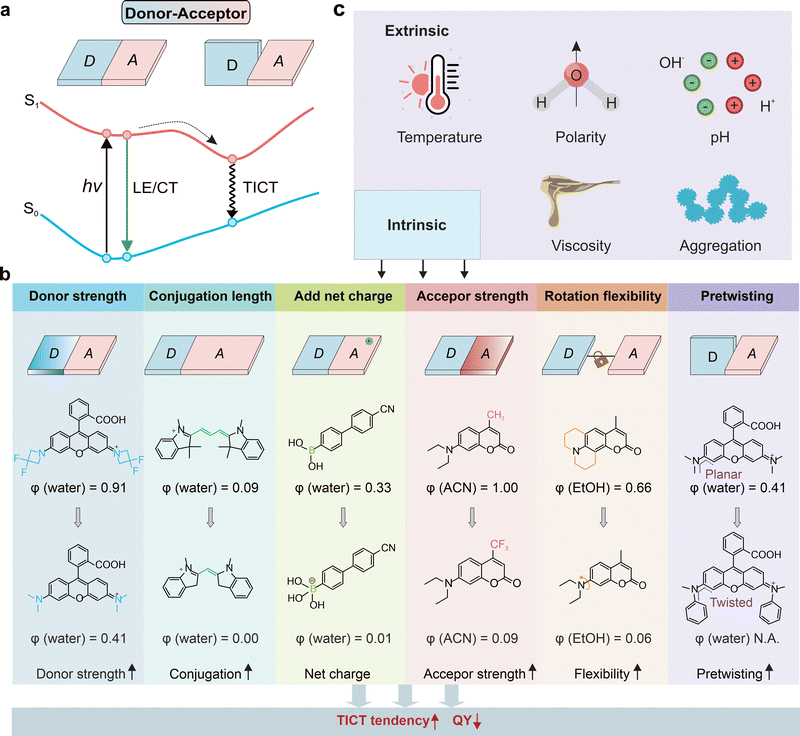 | ||
| Fig. 11 (a) Schematic illustration of the TICT mechanism. (b) Various molecular design strategies to modulate the TICT tendency. (c) Extrinsic factors that affect TICT tendency. | ||
By contrast, when conformational flexibility is restricted—for example, through biomolecular binding, partitioning into viscous compartments, or aggregation—the rotation into the TICT state is suppressed. This inhibition arises from two independent effects: reduced local polarity,6,91 which destabilizes the charge-separated state, and steric hindrance,92,93 which physically prevents bond twisting.
Additionally, chemical reactions between TICT fluorophores and biological molecules (e.g., enzymes, ROS, biothiols, metabolites, or amino acids) could also inhibit TICT. These reactions may either replace strong electron donors or acceptors with weaker ones or cleave the rotatable moieties, thereby preventing the formation of the non-emissive state.
Under these conditions, the fluorophore is locked in an emissive state, whether LE or intramolecular charge transfer (ICT). Since all excited states exhibit varying degrees of charge transfer, the LE and ICT labels are often used interchangeably in practice. This dark-to-bright switching mechanism directly underpins the power of TICT probes for wash-free bioimaging.
4.1. Design strategies for tuning TICT tendency
The ability to tune fluorogenicity depends on precise modulation of a fluorophore's TICT propensity. Liu, Xu, and co-workers established a structure–property framework, identifying several key parameters, such as donor strength, acceptor strength, pre-twisting, net charge, and π-conjugation length (Fig. 11b).89,94(1) Donor strength. Stronger electron donors promote charge separation, stabilizing the twisted charge-transfer state and thus increasing TICT tendency.
(2) π-Conjugation. Extending the conjugated backbone delocalizes orbitals and reduces charge localization, which suppresses TICT. Conversely, shorter conjugation enhances TICT.
(3) Net charge. Introducing cationic or anionic groups in the π-conjugation increases the effective acceptor or donor strength, amplifying the driving force for TICT.
(4) Acceptor strength. Stronger electron-withdrawing substituents likewise stabilize the charge-separated state, enhancing TICT.
(5) Structural flexibility. Restricting donor–acceptor rotations, by steric modification, rigidification, or rotor cleavage, blocks the LE/ICT-to-TICT transition and thereby suppresses TICT.
(6) Pre-twisting. Sterically enforced twisting destabilizes the emissive LE/ICT state, lowering the barrier for bond rotation and making TICT formation more favorable.
Together, these parameters provide a design map for systematically engineering fluorogenic probes. In practice, multiple strategies are often combined to achieve strong wash-free contrast.
Importantly, Liu et al. also highlighted the value of computational approaches for the rational design of TICT probes. By utilizing TD-DFT calculations, key parameters such as the TICT-driving energy (EDE) and rotational barriers (ERB) can be quantitatively estimated, providing predictive insight into TICT propensity. Accurate modeling of TICT, however, requires addressing the well-known limitations of standard hybrid density functionals, which tend to underestimate the energies of charge-transfer states. To overcome this issue, range-separated hybrid functionals (e.g., CAM-B3LYP and ωB97X-D) or hybrid functionals with a high Hartree–Fock exchange contribution (e.g., M06-2X), when combined with state-specific solvation treatments such as corrected linear response (cLR), are essential for reliably capturing TICT energetics and trends.100
Beyond intrinsic molecular design parameters, the external environment plays a critical role in modulating the propensity for TICT (Fig. 11c). Elevated temperature can help fluorophores overcome rotational energy barriers, facilitating access to the TICT state and concomitantly reducing fluorescence intensity, a principle widely exploited in fluorescence-based temperature sensing.101–103 Increased solvent polarity similarly stabilizes the highly polarized TICT state and promotes TICT formation. Changes in pH can induce protonation or deprotonation, introducing net charge redistribution within the molecule and thereby favoring TICT formation. In contrast, increased viscosity—such as in dense media or molecular aggregates—restricts intramolecular rotation, suppressing TICT and enhancing emission. Collectively, these environmental factors provide powerful means to tune TICT behavior and report on local physicochemical conditions. In the following sections, however, we focus primarily on intrinsic molecular parameters, which are more directly relevant to fluorophore design. At the same time, environmental effects are discussed mainly as guiding principles for deploying TICT probes in specific applications.
Representative studies illustrate these principles. Fang and co-workers replaced conventional dialkylamine donors with N-methylpyrrole groups across multiple scaffolds, including naphthalimide, phthalimide, coumarin, and rhodamine (T1 and T2; Fig. 12a).95 This strategy both enhanced donor strength and introduced steric hindrance, biasing molecules toward pre-twisted conformations. The resulting dyes were nearly non-emissive in polar (Fig. 12b) and low viscosity solutions (Fig. 12c), but showed strong turn-on signals in lipid droplets and protein environments. This study demonstrates that combining donor enhancement with pre-twisting can yield highly responsive, wash-free probes.
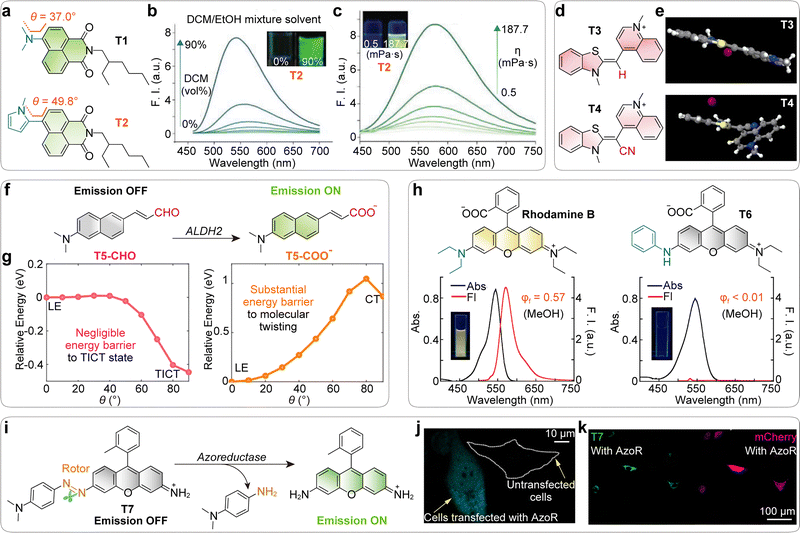 | ||
| Fig. 12 (a) Chemical structures of naphthalimide probes with different donors, and the corresponding pre-twisting angles between donors and fluorophore planes. (b) Emission spectra of T2 in solvent mixtures with varying DCM/MeOH volume ratios. Inset shows the corresponding fluorescence images. (c) Emission spectra of T2 in the mixture solvent of MeOH and glycerol with varying viscosities. Inset shows the corresponding fluorescence images. (d) Chemical structures of cyanine probes. (e) The crystal structures of T3 and T4, including an iodide counterion. (f) Schematic illustration of the turn-on fluorescence response of the ALDH2 probe triggered by enzymatic oxidation. (g) Excited-state (S1) potential energy surface (PES) scan of T5-CHO (left) and T5-COO− (right). (h) Chemical structures of rhodamine B and its amino-phenyl-substituted derivative (T6), together with their absorption spectra, emission spectra, quantum yield values in methanol, and fluorescence images under UV light. (i) Schematic representation of azo-rhodamine and its turn-on fluorescence response upon reduction catalyzed by azoreductase (AzoR). (j) Fluorescence confocal microscopy images of HeLa cells with T7. Cells were transfected with ECFP-fused AzoR. Scale bar: 10 µm. (k) Confocal fluorescence images showing the activation of T7 in HeLa cells expressing AzoR. HeLa cells were co-transfected with AzoR and mCherry plasmids. mCherry (red channel) serves as a cotransfection marker to identify cells that successfully express AzoR. Scale bar: 100 µm. (b), (c), (e), (g), (h), (j) and (k) are adapted from ref. 95–99 with permission from Wiley-VCH, the American Chemical Society, the American Chemical Society, the American Chemical Society and the American Chemical Society, copyright 2022, 2013, 2025, 2022 and 2017, respectively. | ||
Armitage and co-workers demonstrated that cyano (–CN) substitution on the polymethine bridge of cyanine dyes (T3 and T4; Fig. 12d) increased acceptor strength while enforcing non-planarity of the backbone (Fig. 12e).96 This dual action promoted TICT formation, effectively suppressing background fluorescence and improving photostability by shortening the excited-state lifetime of free dyes. Building on this strategy, cyano-bridged scaffolds have become widely adopted, even serving as a popular design for AIEgens that exploit their strong TICT sensitivity.
Li and collaborators presented a rational strategy for constructing an ALDH2 probe library, in which enzymatic oxidation of the aldehyde (–CHO) to a carboxylate (–COO−) suppresses TICT, via reducing the electron-withdrawing strength of the acceptor moiety (T5; Fig. 12f and g).97 This reaction-driven modulation transforms the probe from a non-emissive state into an emissive one, resulting in a pronounced fluorescence turn-on.
The Urano group developed rhodamine-based probes incorporating amino-phenyl substituents (T6; Fig. 12h).98,99 The phenyl group introduced steric repulsion, driving the dyes into pre-twisted conformations and lowering the barrier to TICT. This design highlights how pre-twisting can act as a powerful driver of fluorogenicity.
Liu, Xu, and co-workers explored the impact of introducing net charges into hemicyanine derivatives.94 Cationic substituents effectively enhanced the acceptor strength, thereby increasing the driving force for TICT and improving environmental sensitivity. However, introducing a net charge can also compromise membrane permeability. This trade-off can be mitigated by introducing additional substituents to neutralize or shield the charge.
Liu, Xu, and co-workers systematically investigated a family of hemicyanines with varying π-conjugation lengths.94 They demonstrated that extended conjugation delocalized the frontier orbitals and suppressed TICT formation, thereby reducing the sensitivity to viscosity. Similar observations have also been reported by several other groups, reinforcing π-conjugation as a critical design lever in TICT fluorophore engineering.104,105
Finally, Urano et al. introduced a fluorogenic probe design that leverages enzymatic removal of a rotary fragment to suppress TICT (T7; Fig. 12i). This reaction-driven modulation switches the probe from a non-emissive state to a highly fluorescent form, thereby enabling precise visualization of enzyme activity in living systems (Fig. 12j and k).99
4.2. Applications in bioimaging
The TICT mechanism has enabled a broad spectrum of wash-free bioimaging applications, spanning organelles,29,106 nucleic acids,107,108 proteins,94,109 and pathogens.110,111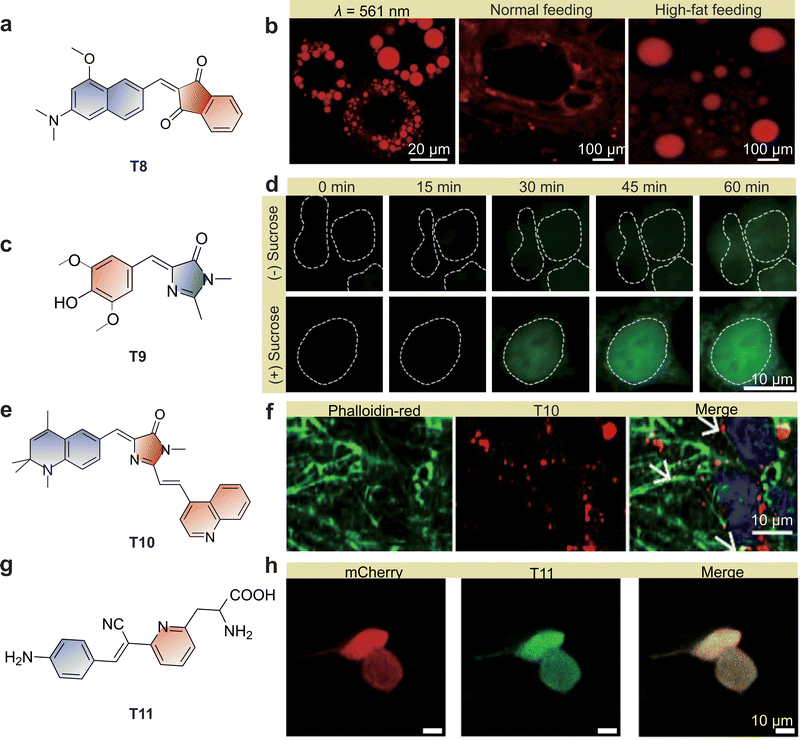 | ||
| Fig. 13 (a) Chemical structure of T8. (b) Confocal laser scanning microscopy images of differentiated 3T3-L1 preadipocyte cells under 561 nm excitation (left, scale bar: 20 µm), normal (middle, scale bar: 100 µm), and high-fat fed (right, scale bar: 100 µm) guinea pig liver tissues using T8. (c) Chemical structure of GFP-like T9. (d) Fluorogenic imaging of 5S-Spinach RNA induction under osmotic stress using T9. HEK293T cells expressing 5S-Spinach were pretreated with ML-60128 and subsequently exposed to vehicle or sucrose to induce osmotic stress. Sucrose treatment markedly increases 5S-Spinach RNA levels compared with control. (e) Chemical structure of T10. (f) Confocal imaging of GG2 cells stained with TRICT-Phalloidin in the presence of T10. Scale bars: 10 µm. (g) Chemical structure of T11. (h) Confocal imaging of HEK293T cells expressing FRB-87PAPd–mCherry fusion protein and wild-type FKBP using T11, with fluorescence activation upon rapamycin addition. Scale bars: 10 µm. (b), (d), (f) and (h) are adapted from ref. 112–115 with permission from the American Chemical Society, the American Association for the Advancement of Science, the American Chemical Society, and Springer Nature, copyright 2022, 2011, 2021 and 2024, respectively. | ||
Protein misfolding and aggregation represent another fertile ground for TICT-based design. Liu, Xu, and co-workers devised a sensor array of hemicyanines (e.g., T12; Fig. 14a) with systematically varied TICT propensities, enabling sequential detection of amyloid-β aggregation from monomers to fibrils (Fig. 14b).94 The graded turn-on responses provided a continuous, wash-free readout of aggregation states in living cells, surpassing conventional amyloid stains such as Thioflavin T. Complementarily, Venkatesh and co-workers developed bimane-based molecular rotors (e.g., T13, Fig. 14c) tailored to detect α-synuclein condensates and fibrils (Fig. 14d).104 These probes respond to changes in viscosity and polarity associated with Parkinson's disease pathology, offering wash-free monitoring of both early oligomers and mature aggregates. Together, these studies demonstrate how TICT fluorophores can illuminate diverse aggregation pathways, with direct relevance to neurodegenerative diseases.
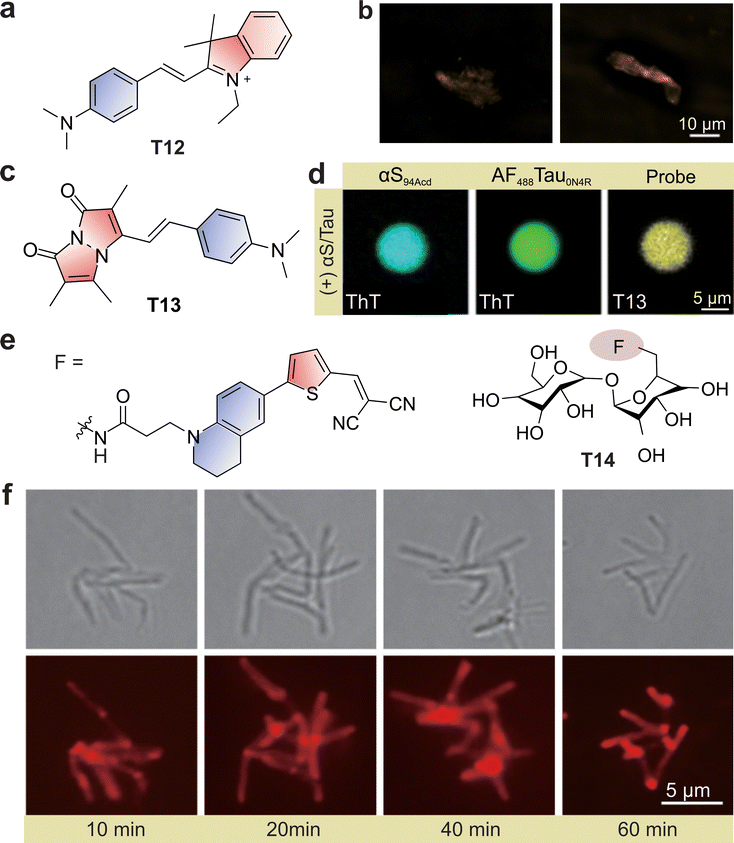 | ||
| Fig. 14 (a) Chemical structure of hemicyanine probe T12. (b) Structured illumination microscopy (SIM) fluorescence imaging of Aβ protein fibrils using T12. (c) Chemical structure of T13. (d) Confocal laser scanning microscopy (CLSM) images of αS/Tau0N4R condensates stained with T13 and ThT doped with 5% AF488-labeled Tau0N4R and 5% Acd-labeled αS. (e) Chemical structure of the bacterial probe T14. (f) Fluorogenic imaging of M. smegmatis incubated with T14 for the indicated durations. (b), (d) and (f) are adapted from ref. 94, 104 and 111 with permission from the Royal Society of Chemistry, the American Chemical Society and Wiley-VCH, copyright 2023, 2025 and 2023, respectively. | ||
4.3. Advantages and design versatility
The primary advantage of TICT-based probes lies in their intrinsic “dark-to-bright” switching mechanism, which simplifies probe architecture. Unlike FRET or PET systems, which often require the covalent linkage of a separate quencher or a specific receptor unit, TICT probes are frequently monochromophoric structures containing integrated molecular rotors. This structural simplicity minimizes perturbation to the biological target and facilitates cell permeability.A significant strength of TICT dyes is their exquisite sensitivity to local physicochemical environments. Because their emission increases when molecular rotation is restricted, they serve as powerful indicators of viscosity, rigidity, and polarity. This property has made TICT dyes gold-standard tools for visualizing membrane fluidity, macromolecular crowding, and the liquid–liquid phase separation (LLPS) of biomolecular condensates. When combined with targeting motifs, they can report both spatial location and microenvironment, adding functional depth to wash-free imaging.
TICT behavior is also highly tunable through the “structure–property” framework established by Liu, Xu, and others. Adjusting donor–acceptor strength, pre-twisting, net charge, and π-conjugation allows precise control over rotational barriers and driving energies. This tunability supports the development of probes with tailored sensitivity, brightness, and switching efficiency across diverse applications, from small-molecule sensors to engineered protein-based fluorogens.
Finally, the TICT principle is compatible with virtually all major fluorophore families, including coumarins, naphthalimides, rhodamines, and cyanines. Modifying donor–acceptor groups or bridge structures enables systematic tuning of emission wavelength, brightness, and fluorogenicity from the visible to the near-infrared range.
4.4. Limitations and practical challenges
Despite their versatility, TICT probes face several practical challenges. While their high environmental sensitivity enables powerful sensing, it creates a critical bottleneck in target specificity. A probe designed for a specific protein aggregate may also be activated in viscous lipid droplets or hydrophobic protein pockets, making it difficult to distinguish specific binding from nonspecific accumulation. As a result, each probe requires careful validation in the intended biological context.Achieving an optimal ON/OFF contrast also involves trade-offs. There is an inherent tension between quenching efficiency and brightness: a strong TICT tendency effectively suppresses background fluorescence in aqueous media but often results in a dim “ON” state if the biological target cannot provide sufficient steric hindrance to fully arrest molecular rotation. Conversely, reducing the rotational freedom to boost brightness inevitably raises the “OFF” state leakiness. This “dynamic range ceiling” remains a major hurdle for high-contrast, wash-free imaging in complex intracellular milieus.
From a photophysical standpoint, the susceptibility of TICT states to thermal and pH fluctuations complicates quantitative imaging. Because rotational kinetics are governed by the Arrhenius-like dependence on temperature and viscosity, and donor/acceptor strengths are sensitive to local pH/polarity, TICT-based signals are often “multi-parametric”. The current difficulty in isolating a single environmental variable from this convoluted signal prevents TICT probes from being used as truly quantitative “molecular rulers” in dynamic live-cell experiments.
In addition, the development of NIR-emissive TICT dyes faces a structural bottleneck. Extending conjugation to reach longer wavelengths often increases the rotational energy barrier, making it difficult to maintain efficient quenching in the unbound state.
Finally, photostability is fundamentally compromised by the electronic nature of the TICT state. Although some improved designs exhibit good stability, many TICT dyes are prone to bleaching or photooxidation. The charge-separated TICT state often facilitates efficient intersystem crossing (ISC) to long-lived triplet states.116 These triplets act as “reactive centers” that sensitize the formation of ROS, leading to irreversible self-destruction of the fluorophore. Although engineering non-radiative decay through accessible conical intersections can theoretically bypass these triplet traps, it introduces a “stability-brightness paradox”: an overly dominant CI pathway can lead to excessive internal conversion, potentially quenching the desired fluorescence even in the “ON” state. This remains one of the most formidable challenges in the field.
4.5. Outlook and future directions
A major future direction is the seamless integration of TICT rotors into biological scaffolds. Expanding the genetic code to incorporate distinct TICT-capable amino acids will enable the “wash-free” imaging of endogenous protein folding and conformational dynamics with minimal background, thereby moving beyond simple localization imaging.While most existing TICT modulations rely on physical properties such as viscosity and polarity, introducing chemical reactions, such as enzyme-cleavable rotors or reactive handles that lock dyes into emissive states only upon specific biochemical events, adds a powerful new dimension. This chemically gated approach offers an extra layer of specificity, delivering high fluorogenic contrast and enabling real-time imaging of enzymatic activity in live cells without the need for washing steps.
Developing sensitive TICT scaffolds in the NIR-I and, ultimately, NIR-II remains a significant goal. Achieving long-wavelength emission without sacrificing rotor function will require improved modeling of electronic states, torsional barriers, and solvent effects. Pre-twisting strategies and alternative donor–acceptor architectures may help overcome many of these design challenges.
5. Internal conversion to dark states
Suppression of bond rotation in TICT probes represents a powerful way to recover fluorescence. Beyond these large-scale motions, fluorogenicity can also be encoded through the relative ordering of electronic states, specifically by facilitating internal conversion to a dark state within the molecule's own energy landscape.Internal conversion to a dark state (ICDS) is a distinct fluorogenic strategy. Unlike FRET or PET, which rely on external quenchers, ICDS encodes quenching within the fluorophore's own electronic structure. After photoexcitation, the fluorophore first occupies a bright π–π* state, which rapidly relaxes (via internal conversion) into a lower-lying dark state, usually of n–π* character. In some systems, a charge-separated state such as TICT serves a similar role. The dark state dissipates energy non-radiatively, leaving unbound or unreacted probes essentially nonfluorescent. Because TICT is covered in Section 4, this section focuses mainly on n–π* dark states, with a brief mention of charge-transfer states accessed through IC.
Fluorescence arises only when the dark state is destabilized relative to the π–π* state. This can be achieved through covalent modifications, such as introducing donor groups or removing heteroatom fragments, or through environmental effects, such as polarity, proticity, or binding. In both cases, the outcome is a selective “OFF–ON” response, where emission is switched on only after reaction or binding.
5.1 Design strategies for IC-driven quenching
One approach to harnessing IC is to modulate donor strength within a conjugated fluorophore (Fig. 15). Weak donors destabilize the π–π* state, making the n–π* state energetically favored and quenching fluorescence. Strong donors stabilize the π–π* state as the lowest excited state, allowing radiative decay and bright emission (Fig. 15a, d and e). This simple electronic tuning offers a versatile approach for designing probes that are responsive to structural or environmental changes.118–120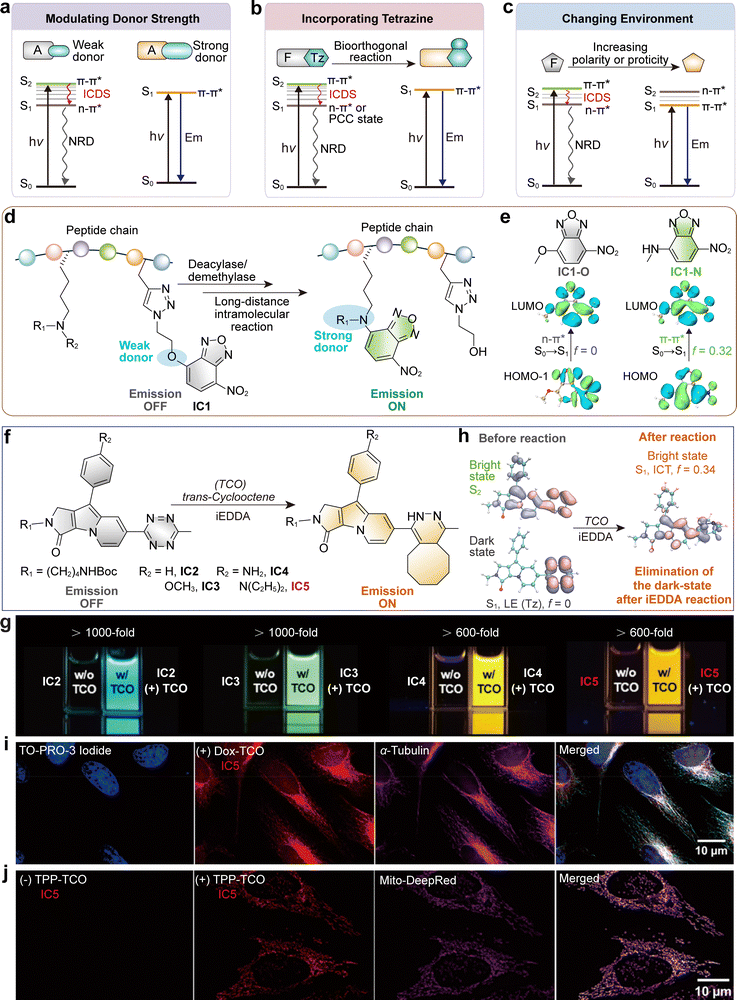 | ||
| Fig. 15 (a)–(c) Schematic representations of three strategic approaches to facilitate internal conversion to a dark state (ICDS) in the context of fluorogenic probe design. (d) Illustration of the fluorescence turn-on mechanism of IC1 in response to deacetylase, desuccinylase, and demethylase activities. (e) Molecular orbital distributions and oscillator strength (f) values of the model compounds IC1-O (left) and IC1-N (right). The S0 → S1 photoexcitation of IC1-O is dominated by a dark n–π* transition, with a zero-oscillator strength value, while that of IC1-N is dominated by a bright π–π* transition, with a considerable oscillator strength value of 0.32. All calculations employed the M06-2X/Def2-SVP level of theory in vacuum. (f) Schematic representation of the fluorescence turn-on mechanism of probes IC2–IC5, triggered by bioorthogonal cycloaddition with trans-cyclooct-4-enol (TCO). (g) Fluorescence images and the turn-on ratios of IC2–IC5 under UV light before and after the bioorthogonal reaction with TCO. (h) Hole (ice blue) and electron (pink) distributions of Seoul-Fluor–tetrazine conjugates before and after the reaction with TCO. Before reaction, tetrazine generates a low-lying, dark LE state, denoted LE(Tz), characterized by zero oscillator strength. After the bioorthogonal reaction with TCO, the low-lying state becomes a bright state characterized by a π–π* transition. Calculations were performed using IC2 as a representative example, omitting the methyl group on tetrazine. Calculations employed the M06-2X functional with the Def2-SVP basis set in water. (i) Fluorogenic bioorthogonal imaging of microtubules with IC5 in fixed cells pretreated with Dox–TCO (red). TO-PRO-3 iodide (blue) was used for nucleus staining. Selective microtubule staining was further confirmed with immunofluorescence using α-tubulin antibody (magenta). (j) Fluorogenic bioorthogonal imaging of mitochondria with IC5 (red) in live cell conditions without washing steps. Cells were treated with DMSO (no fluorescence) or TPP–TCO. MitoTracker Deep Red (magenta) was used as a reference control for cell staining. (g), (i) and (j) are adapted from ref. 117 with permission from the American Chemical Society, copyright 2018. | ||
A second strategy involves incorporating tetrazines (or other fragments) directly into the conjugated backbone of a fluorophore, rather than attaching them to fluorophores via linkers. These fragments introduce n–π* states that quench fluorescence efficiently. When the probe undergoes a bioorthogonal reaction—such as an inverse electron-demand Diels–Alder reaction with strained alkenes or alkynes—the tetrazine is consumed or destabilized, eliminating the dark pathway and restoring emission (Fig. 15b, f and h).121,122 This mechanism can produce dramatic turn-on ratios, sometimes several hundred-fold, and is highly effective in live-cell imaging. In the red and near-infrared (NIR) regions, due to the stabilization of the π–π* state, however, the n–π* state is often too high to serve as the primary quencher. Instead, the electron-withdrawing tetrazine stabilizes intramolecular charge-transfer states, producing a photoinduced charge centralized (PCC) state that dissipates energy non-radiatively (Fig. 16).41,123 Although reminiscent of PET, this quenching is fully intramolecular and best regarded as a special form of IC. PCC extends fluorogenicity into NIR, enabling probes with improved tissue penetration, reduced phototoxicity, and broader imaging scope.
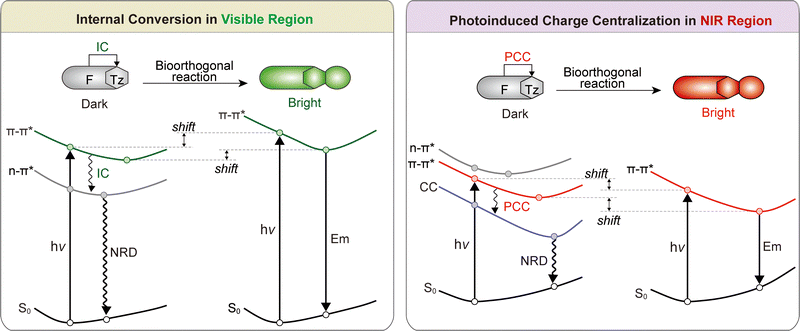 | ||
| Fig. 16 Schematic illustrations of the photophysical mechanisms involved in the Tz-based fluorogenic compounds working in the visible region (left) and the NIR region (right). | ||
A third, less bioimaging-relevant pathway is environment-sensitive state switching. Certain small fluorophores display closely spaced π–π* and n–π* states whose order changes with solvent polarity or proticity. In nonpolar solvents, the n–π* state is stabilized and quenches emission; in polar or protic solvents, the π–π* state dominates and emission returns (Fig. 15c). While valuable for understanding IC photophysics, these dyes usually emit in the blue region, where autofluorescence and phototoxicity limit biological use.124 For this reason, the discussion here emphasizes donor modulation and tetrazine incorporation, which provide more versatile strategies for bioimaging.
5.2. Applications in bioimaging
Embedding n–π* or PCC states into fluorophores has enabled the design of highly fluorogenic probes. Nitrobenzoxadiazole (NBD) derivatives are a well-studied example. In their intact form, the n–π* state quenches fluorescence (Fig. 15e).119 Enzymatic transformations such as lysine deacylation or demethylation convert O-NBD into N-NBD, stabilizing the π–π* state and switching on emission (Fig. 15d and e). These probes are valuable for studying lysine deacylase and demethylase activity, including distinctions among histone demethylation states. NBD derivatives also undergo hydrogen bond–induced quenching (HBQ), as discussed in Section 7, which adds another layer of sensitivity to biochemical environments.Tetrazine-based probes offer perhaps the most versatile applications. While tetrazines are covered extensively elsewhere,40,41 it is worth noting that these systems exemplify how IC-driven quenching integrates seamlessly with bioorthogonal chemistry to produce generalizable fluorogenic scaffolds. Park and colleagues developed Seoul-Fluor–tetrazine conjugates (Fig. 15f) with monochromophoric designs, producing color-tunable probes for multiplex microtubule and mitochondria labeling in live cells.117 Mechanistically, tetrazines introduce low-lying n–π* dark states that funnel excitation energy into nonradiative decay. After the iEDDA reaction, this pathway is eliminated, and emission is restored (Fig. 15g), yielding robust wash-free imaging of microtubules and mitochondria (Fig. 15i and j) across visible wavelengths. Kim and co-workers extended this strategy to BODIPY–tetrazine conjugates (Fig. 17a), which remained essentially dark until ligated to bicyclo[6.1.0]nonyne (BCN)-modified biomolecules.125 These enabled live-cell imaging of mitochondria and lysosomes with fluorescence enhancements exceeding 1000-fold (Fig. 17a, bottom panel). More recently, Shen and Liu showed that for such monochromophoric designs with an integrated π-conjugation, IC to the low-lying dark n–π* state is responsible for fluorescence quenching in the visible regime (Fig. 16, left panel).126 In the NIR region, tetrazine conjugates operate through PCC rather than n–π* quenching (Fig. 16, right panel).123 Excitation centralizes charge on the tetrazine moiety, creating a quasi-charge-separated, non-emissive state. After the bioorthogonal reaction, this pathway is abolished, restoring deep red or NIR emission. This extension into the NIR expands IC-based probes to deep-tissue and multiplex imaging.
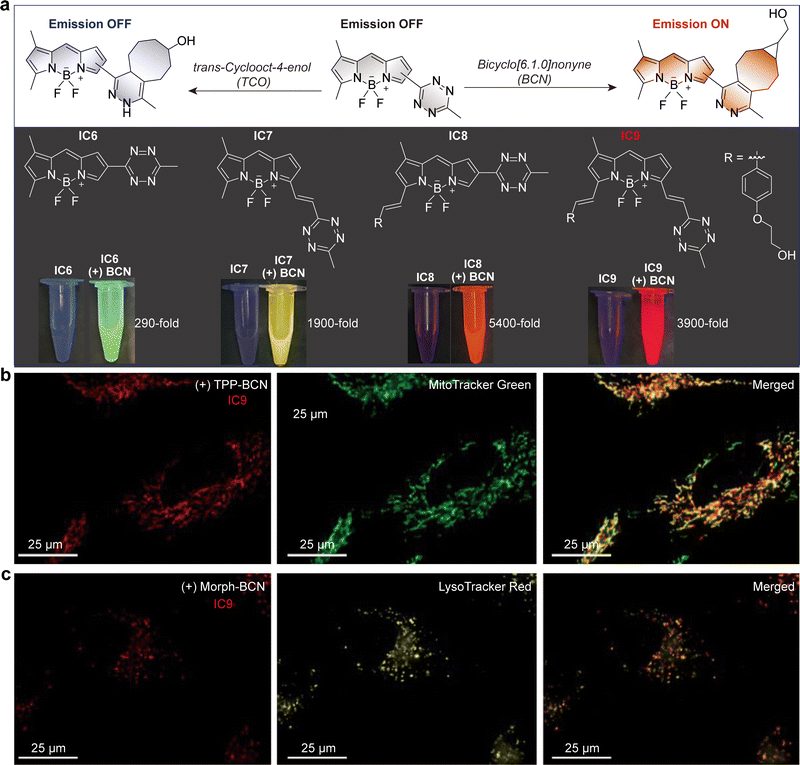 | ||
| Fig. 17 (a) Schematic representation of the bioorthogonal reactions of the probes (IC6–IC9) with trans-cyclooct-4-enol (TCO) and bicyclo[6.1.0]nonyne (BCN). The dihydropyridazine product formed by the reaction with TCO is non-fluorescent, while the reaction with BCN can trigger the emission of IC6–IC9. The bottom panel shows the corresponding fluorescence images, and the turn-on ratios of IC6–IC9 under UV light before and after the bioorthogonal reaction with BCN. (b) and (c) Fluorogenic bioorthogonal imaging of (b) mitochondria and (c) lysosomes with IC9 (red) without washing steps. Cells were treated with TPP-BCN or Morph-BCN. MitoTracker Green (green) or LysoTracker Red (yellow) was used to stain cells as the corresponding reference controls. (a)–(c) are adapted from ref. 125 with permission from Wiley-VCH, copyright 2023. | ||
5.3. Advantages and design versatility
Internal conversion (IC) strategies stand out for their ability to generate monochromophoric fluorogenic probes. Unlike FRET or PET systems, which typically require an appended quencher, IC encodes the quenching element directly within the fluorophore's electronic structure. This produces compact scaffolds with minimal disruption to biological interactions. When groups such as tetrazines or nitro moieties are incorporated directly into the π-conjugated backbone, their orbital overlap efficiently funnels excitation energy into a low-lying dark state. The resulting “OFF” state often exhibits near-zero quantum yield, enabling exceptionally high turn-on ratios that can exceed 1000-fold.IC-based quenching is also highly compatible with bioorthogonal chemistry. In many tetrazine systems, a single structural unit serves as both the reactive handle and the quencher. Because fluorescence recovery occurs only after a specific reaction, such as an inverse electron-demand Diels–Alder cycloaddition, the probability of false activation is essentially eliminated. This tight coupling of reactivity and quenching simplifies probe design and strengthens selectivity.
A further strength of IC is its applicability across the visible and near-infrared (NIR) spectral regions. In visible fluorophores, n–π* states provide efficient pathways to dark states. For red and NIR chromophores, the recently recognized photoinduced charge centralization (PCC) mechanism enables analogous quenching by stabilizing a charge-transfer state. These design principles are applicable across diverse fluorophore scaffolds, supporting the rational development of fluorogenic probes over a broad spectral range.
5.4. Limitations and practical challenges
Despite these advantages, the implementation of IC-based probes is governed by a dedicated energetic equilibrium, imposing several fundamental constraints. A primary bottleneck is the extreme sensitivity of state-ordering: for effective quenching, the dark state (e.g., n–π* or PCC) must be positioned with sub-electron Volt precision immediately below the bright π–π* state. This narrow energetic window renders the “OFF” state vulnerable to environmental factors, including minor perturbations in solvent polarity, proticity, or local hydrogen bonding. These fluctuations can reorder the relative ordering of bright π–π* and dark n–π* or PCC states, triggering premature fluorescence. The current lack of molecular scaffolds capable of “locking” this energy hierarchy against fluctuating biological environments remains a significant barrier to developing robust, non-fluorescent precursors.The development of IC-based probes also encounters a “red-shift deadbolt” within the NIR window. As π-conjugation is extended to achieve longer wavelengths, the delocalized π–π* state undergoes more rapid energetic stabilization compared to the localized n–π* or PCC states. This divergent energy scaling often leads to a “mechanism collapse”, where the bright state inevitably drops below the dark quenching channel, fundamentally precluding a dark “OFF” state in NIR dyes.
Moreover, incorporating heteroatoms or highly electron-poor fragments to generate n–π* or PCC states can compromise photostability.127 These states may enhance intersystem crossing or accelerate photooxidation, thereby accelerating bleaching during high-intensity imaging.
Reaction-based activation introduces additional limitations. Tetrazine systems, for example, can yield different photophysical outcomes depending on the reaction partner. This product-dependent photophysics means that a probe optimized for one bioorthogonal pair may perform poorly with another. Consequently, researchers may face limited flexibility in tuning reaction kinetics without risking loss of fluorescence contrast, thereby hindering the broader utility of IC-based “turn-on” platforms.
5.5. Outlook and future directions
Future progress in IC-based fluorogenic probe design will hinge on the rigour of computational modeling. High-level quantum chemical tools, such as TD-DFT with explicit solvation or mixed quantum–classical dynamics, are becoming essential for accurately predicting the energy gaps between π–π*, n–π*, and PCC states. These capabilities will support the de novo construction of fluorophores in which dark-state properties are engineered with higher precision.Furthermore, expanding the PCC mechanism with new electron-deficient, chemically robust quenching motifs may enable IC-based probes to operate in the NIR window. The systematic development of new quenching fragments, along with precise control over de-excitation pathways, is expected to underpin the advancement of IC-driven fluorogenic probes compatible with deep-tissue imaging.
Finally, another promising direction is the development of orthogonal, multiplexable IC scaffolds. By developing distinct quenching triggers beyond tetrazines that respond to different bioorthogonal pairs, it should be possible to create families of probes that enable simultaneous, wash-free imaging of multiple distinct targets. Such multiplexing would significantly expand the ability to map organelles, protein complexes, and metabolic pathways simultaneously.124
6. Structural isomerization
While internal conversion relies on the tuning of existing excited-state manifolds, another robust approach to fluorogenicity involves a reversible chemical transformation. This is exemplified by structural isomerization, specifically spirocyclization, which toggles between a non-emissive closed form and a highly fluorescent open state, such as in rhodamines, fluoresceins, and rhodols. In these scaffolds, the reversible formation of spirocyclic structures leads to dramatic changes in absorption and fluorescence properties.19,128 This process not only governs the fundamental photophysics of the fluorophores but also affects their cell–membrane permeability and interactions with biological targets.11,19,91,129,130 Understanding and controlling these structural transformations is therefore essential for optimizing fluorogenic probes and developing effective imaging tools.6.1. Design strategies for rhodamine-based fluorogenic probes
Rhodamines are among the most important classes of organic fluorophores, widely used in biological imaging due to their high molar extinction coefficients, excellent fluorescence quantum yields, and broad emission tunability. They also possess strong biological compatibility and membrane permeability. However, what sets rhodamines apart from many other fluorophores is their unique spirocyclization equilibrium.130,131 This reaction allows rhodamines to reversibly switch between two distinct forms: a colorless, non-fluorescent, membrane-permeable closed form, and a highly fluorescent, zwitterionic (or cationic) open form (Fig. 18a). This ring-opening transition enables the development of “turn-on” probes with significant fluorescence contrast in response to environmental stimuli or target binding.One key parameter used to quantify this equilibrium is KL–Z, or its logarithmic form log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z, which measures the ratio between the populations of the open and closed forms (Fig. 18b). A high KL–Z (or more positive log
KL–Z, which measures the ratio between the populations of the open and closed forms (Fig. 18b). A high KL–Z (or more positive log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z values) means that the open, fluorescent form is favored, whereas a low KL–Z (or more negative log
KL–Z values) means that the open, fluorescent form is favored, whereas a low KL–Z (or more negative log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z values) suggests dominance of the closed, non-emissive state. In a seminal study, Lavis and co-workers proposed that log
KL–Z values) suggests dominance of the closed, non-emissive state. In a seminal study, Lavis and co-workers proposed that log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z values above −2 indicate that many dyes are in the open form, leading to high background.132 On the other hand, values below −4 lock the dye in the closed form, rendering it too dim to be useful. They identified a functional “sweet spot” between −3 and −2, where dyes can exhibit significant ON/OFF contrast upon activation. While these thresholds may be specific to the rhodamine scaffolds used in their studies, the concept of tuning log
KL–Z values above −2 indicate that many dyes are in the open form, leading to high background.132 On the other hand, values below −4 lock the dye in the closed form, rendering it too dim to be useful. They identified a functional “sweet spot” between −3 and −2, where dyes can exhibit significant ON/OFF contrast upon activation. While these thresholds may be specific to the rhodamine scaffolds used in their studies, the concept of tuning log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z remains broadly relevant for designing responsive fluorogenic probes.
KL–Z remains broadly relevant for designing responsive fluorogenic probes.
Another important parameter used to describe rhodamine switching behavior is the pKcycl (Fig. 18b), which denotes the pH at which the concentrations of the closed spirolactone (or spirolactam) form and the open zwitterionic/cationic form are equal.9 Unlike the standard pKa that refers to dissociation of a specific functional group, pKcycl captures a global conformational switch between emissive and non-emissive states. Acidic environments often stabilize the open form by protonating the ring lock group, thereby enhancing fluorescence (Fig. 18b). This protonation-driven equilibrium explains the pH-sensitive switching and blinking behavior exploited in super-resolution microscopy.
Although pKa and pKcycl are sometimes reported interchangeably, it is more precise to use pKcycl when referring specifically to the ring-closing equilibrium. A high pKcycl means that the dye favors the open state even under near-neutral conditions, while a low pKcycl indicates closure unless triggered by acidification or specific binding events.9,11,133 In this review, we retain both pKa and pKcycl notations, as reported in the original literature, to ensure clarity and consistency with the cited sources.
Both intrinsic molecular design and external environmental conditions can modulate the ring-opening behavior of rhodamines by stabilizing or destabilizing the zwitterionic open form (Fig. 18b–f). Liu and co-workers proposed a unified push–pull model that explains how electron-donating and electron-withdrawing groups influence this equilibrium.134,135 Enhancing the electron-donating strength of the donor group on the xanthene core (e.g., R1) increases electron density toward the spiro ring, thereby favoring ring opening. At the same time, installing stronger electron-withdrawing groups at the meso-phenyl ring (R3 and R4) can further stabilize the zwitterionic state, shifting the equilibrium toward the open form. On the other hand, modifications at the bridging atom (R2) significantly affect the ring-opening tendency: replacing the oxygen atom with bulkier or more electropositive groups such as –C(CH3)2, –Si(CH3)2, or phosphine oxide tends to reduce the zwitterion stability and thereby suppress ring opening.
In parallel, external factors such as increased solvent polarity, hydrogen bonding, lowered pH, or the presence of metal ions (acting as Lewis acids) also stabilize the charge-separated open form and promote fluorescence (Fig. 18b).136,137 Acidic media, in particular, can protonate the spirolactam nitrogen or its surrounding groups, tilting the equilibrium toward the open state.
In summary, rational tuning of both molecular architecture and environmental conditions allows precise control over rhodamine ring-opening behavior. By optimizing parameters such as log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z and pKcycl, chemists can design rhodamine-based fluorophores with desired levels of responsiveness, contrast, and emission properties, enabling a wide array of applications in bioimaging and sensing.
KL–Z and pKcycl, chemists can design rhodamine-based fluorophores with desired levels of responsiveness, contrast, and emission properties, enabling a wide array of applications in bioimaging and sensing.
The design of fluorogenic rhodamine dyes for wash-free bioimaging has been systematically advanced by the work of Lavis, Johnsson, and others, who collectively demonstrated how electronic tuning and scaffold engineering can optimize the lactone–zwitterion equilibrium.
Lavis and co-workers introduced the Janelia Fluor (JF) series, establishing that reducing the electron-donating strength of the amino donor (i.e., using difluoroazetidines) stabilizes the closed lactone form and reduces background fluorescence in solution (Fig. 18g).138 Upon binding to biomolecular targets, the local environment stabilizes the zwitterionic state—likely through hydrogen bonding or electrostatic interactions with nearby cations—thereby switching on bright emission. JF635 (a Si-rhodamine derivative, C1) exemplifies this concept, with a log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z of ∼−3.1 that biases it toward the closed form and yields strong fluorogenicity.
KL–Z of ∼−3.1 that biases it toward the closed form and yields strong fluorogenicity.
To further lower the log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z value of O-rhodamines (typically ∼0.5), Lavis and co-workers demonstrated that, in addition to fluorinating the azetidine substituents, direct installation of fluorine atoms onto the xanthene scaffold decreases its electron-donating strength.4,132 This modification yielded JF526 (C2, Fig. 18g), a green O-rhodamine analogue with a log
KL–Z value of O-rhodamines (typically ∼0.5), Lavis and co-workers demonstrated that, in addition to fluorinating the azetidine substituents, direct installation of fluorine atoms onto the xanthene scaffold decreases its electron-donating strength.4,132 This modification yielded JF526 (C2, Fig. 18g), a green O-rhodamine analogue with a log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z ≈ −2.3. The resulting equilibrium achieves a desirable balance—retaining cell permeability in the closed form while enabling robust binding-induced fluorescence activation in the open form.
KL–Z ≈ −2.3. The resulting equilibrium achieves a desirable balance—retaining cell permeability in the closed form while enabling robust binding-induced fluorescence activation in the open form.
In parallel, Johnsson and co-workers developed a systematic framework linking rhodamine fluorogenicity to the electronic properties of both the bridging atom and the spiro-substituents.139 By substituting the bridging oxygen with carbon, silicon, phosphorus, or sulfur groups, they tuned the scaffold's electronic balance and quantified the effects of substituents using Hammett σ correlations. Significantly, reducing the electron-withdrawing strength of the spiro-substituents lowers the log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z, thereby shifting the equilibrium toward the closed form. Within this design strategy, dyes such as MaP618 (a C-rhodamine derivative C3; Fig. 18g; log
KL–Z, thereby shifting the equilibrium toward the closed form. Within this design strategy, dyes such as MaP618 (a C-rhodamine derivative C3; Fig. 18g; log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z ≈ −2.5) and MaP700 (a Si-rhodamine derivative C4; Fig. 18g; log
KL–Z ≈ −2.5) and MaP700 (a Si-rhodamine derivative C4; Fig. 18g; log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z ≈ −2.7) emerged as representative examples (Fig. 18g), optimized to combine intense protein-induced fluorescence turn-on with good cell permeability. In contrast to Lavis’ Janelia Fluor series,138,140 which primarily achieved tuning by introducing fluorinated substituents on the xanthene scaffold (or its amino donors), Johnsson's approach emphasizes systematic variation of the spiro-substituents, providing a complementary framework for modulating rhodamine fluorogenicity.
KL–Z ≈ −2.7) emerged as representative examples (Fig. 18g), optimized to combine intense protein-induced fluorescence turn-on with good cell permeability. In contrast to Lavis’ Janelia Fluor series,138,140 which primarily achieved tuning by introducing fluorinated substituents on the xanthene scaffold (or its amino donors), Johnsson's approach emphasizes systematic variation of the spiro-substituents, providing a complementary framework for modulating rhodamine fluorogenicity.
Beyond equilibrium tuning, Lavis and colleagues pioneered the first photoactivatable Si-rhodamine (C5, C6; Fig. 19a).141 By caging the rhodamine amines with nitroveratryl oxycarbonyl (NVOC) groups, they locked the dye in a non-fluorescent spirolactone state. Upon UV illumination at 405 nm, the NVOC groups were photolyzed, releasing the amino donors, restoring donor strength, and enabling fluorogenic activation (Fig. 19a). Notably, substitution of the xanthene oxygen with a dimethylsilicon bridge introduced a ∼100 nm red-shift (λabs = 637 nm, λem = 654 nm), allowing for far-red photoactivatable imaging.
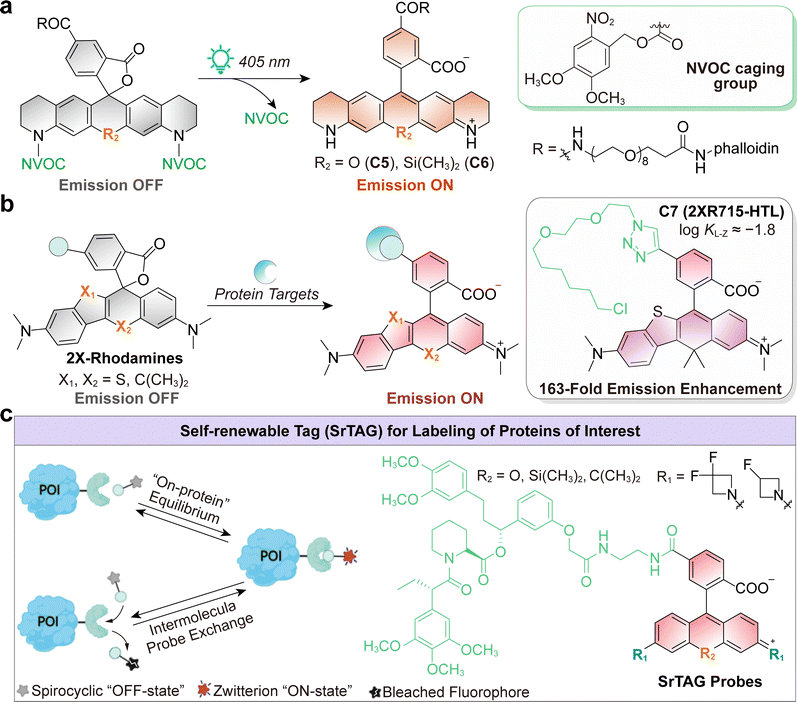 | ||
| Fig. 19 (a) Photo-activated cage release and subsequent ring opening of the rhodamine system. (b) The 2X rhodamine molecular system showing protein-binding-induced ring opening. (c) Schematic representation of the self-renewable tag (srTAG) probes for protein labeling. (c) is reproduced from ref. 129 with permission from the American Chemical Society, copyright 2023. | ||
Zhang and co-workers extended scaffold engineering further by designing 2X-rhodamine (2XR), which incorporates a dual atomic bridge (S + C(CH3)2) and a vinylene-extended π-system (Fig. 19b).142 This architecture rigidifies the chromophore, stabilizing the lactone–zwitterion equilibrium (log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z ≈ −1.8) and yielding bright NIR-I (centered at 765 nm, φf = 0.27 in EtOH) and measurable NIR-II emission above 1000 nm (φf ≈ 0.92%). Functionalization with a HaloTag ligand (2XR715-HTL, C7) produced an exceptional 163-fold fluorescence enhancement upon binding, demonstrating both long-wavelength coverage and strong fluorogenicity.
KL–Z ≈ −1.8) and yielding bright NIR-I (centered at 765 nm, φf = 0.27 in EtOH) and measurable NIR-II emission above 1000 nm (φf ≈ 0.92%). Functionalization with a HaloTag ligand (2XR715-HTL, C7) produced an exceptional 163-fold fluorescence enhancement upon binding, demonstrating both long-wavelength coverage and strong fluorogenicity.
Chu and colleagues introduced an orthogonal concept of “self-renewable” labeling by developing srTAG, a system that enables reversible, noncovalent protein labeling using FKBP-linked rhodamine dyes (Fig. 19c).129 Built on the JF635 scaffold, these dyes retain the ability to undergo spirocyclic ring-opening upon target binding, enabling direct fluorogenic activation on the protein. In parallel, unbound probes in solution can replace photobleached or damaged dyes through dynamic exchange, providing a continuous supply of functional fluorophores. Together, these two mechanisms—on-protein equilibrium and probe exchange—significantly extend imaging durations and enhance photostability, achieving two- to six-fold improvements over conventional HaloTag and SNAP-tag systems.
6.2. Design strategies for fluorescein-based fluorogenic probes
Fluorescein derivatives, long recognized for their high quantum yields and pH-dependent spirocyclization, have recently been re-engineered into fluorogenic probes with dual-gated activation and selective cellular retention. Two complementary strategies exemplify this trend.Kasteren and co-workers developed a bioorthogonal nutrient uptake probe (C8; Fig. 20a) based on carboxyfluorescein diacetate succinimidyl ester (CFSE) conjugated to a tetrazine quencher (Fig. 20a).143 In this design, fluorescence remains suppressed until two sequential events occur: (i) esterase-mediated hydrolysis in the cytosol restores the fluorescein scaffold, and (ii) a bioorthogonal iEDDA reaction between the tetrazine and strained-alkene–modified nutrients removes the quencher (Fig. 20a). This two-step gating strategy eliminates both extracellular background and intracellular leakage, producing a 20–27-fold fluorescence turn-on. By coupling fluorescein's classic spirocyclic OFF/ON switch with a tetrazine quencher, the probe enables real-time tracking of nutrie nt uptake at single-cell resolution.
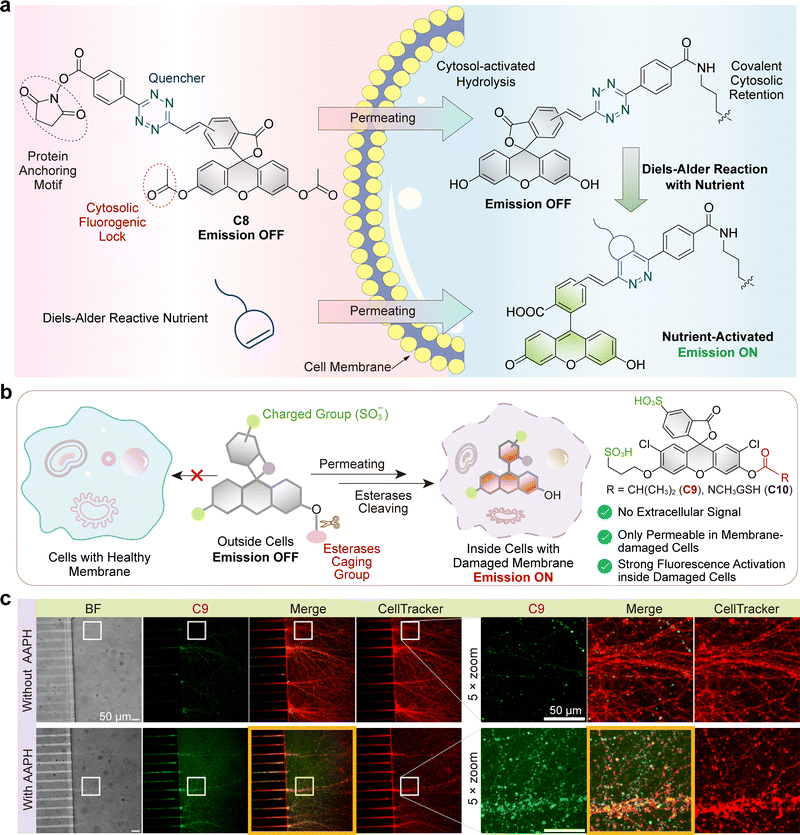 | ||
| Fig. 20 (a) Mechanism for the dual-quenched, live-cell-compatible nutrient probe C8. Carboxyfluorescein diacetate modified with a tetrazine quencher becomes fluorogenic upon cellular uptake and fluorescent after an iEDDA reaction with nutrient analogues bearing strained alkenes, enabling real-time visualization of nutrient uptake with the temporal resolution of the iEDDA reaction. (b) Schematic representation of the fluorogenic imaging of cell membrane damage using MDG probes C9 and C10. (c) Axonal staining with C9 (green) in the absence (top) and presence (bottom) of AAPH, a radical initiator compromising cell membrane integrity and increasing permeability. CellTracker (red) was used as a reference to stain live cells, providing a stable fluorescent signal to visualize cell morphology and confirm viability. (c) is adapted from ref. 19 with permission from the American Chemical Society, copyright 2024. | ||
Thorn-Seshold and colleagues introduced MDG (Membrane Damage Green) probes (C9, C10; Fig. 20b), which exploit fluorescein sulfonation to enforce cell impermeability.19 In healthy cells, the probes remain excluded. However, when membrane integrity is compromised during processes such as ferroptosis, axonal degeneration, or necrosis, intracellular esterases remove protective groups, thereby activating the dye (Fig. 20b). The spirocyclization equilibrium keeps the probes non-fluorescent until unmasking, at which point green emission is irreversibly retained within damaged cells (Fig. 20c). This ensures accurate wash-free imaging, since the unbound probe cannot penetrate intact cells and thus does not contribute to the background.
In both strategies, esterase cleavage plays a central role: by restoring the electron-donating group, it shifts the equilibrium from the spirocyclic (closed) form to the open, fluorescent form, thereby activating fluorescence.
It is worth mentioning that, in classical fluoresceins, basic conditions typically promote the ring-opening reaction. This occurs because deprotonation enhances the electron-donating strength of the phenolic hydroxyl group—shifting from –OH to –O−—thereby stabilizing the open, fluorescent form. This mechanism contrasts with the acid-activated ring-opening observed in rhodamines and related dye families.
6.3. Design strategies for rhodol-based fluorogenic probes
Recent advances in rhodol-based scaffolds have established a unifying strategy that combines spirocyclization-driven quenching with covalent trapping to achieve localized, high-contrast imaging. This approach enables the design of enzyme-activated fluorogenic probes that deliver both high signal contrast and robust cellular retention.Urano and co-workers first reported the SPiDER-bGal platform (C11, C12; Fig. 21a), in which β-galactosidase cleavage unmasks an intramolecular nucleophile that undergoes spirocyclization to restore xanthene conjugation.144 At the same time, a quinone methide intermediate is generated, reacting with nearby nucleophiles to immobilize the activated dye (Fig. 21a) within cells and tissues covalently. This dual mechanism produced a strong turn-on response (up to ∼990-fold) with minimal leakage, enabling functional imaging of LacZ reporters at single-cell resolution (Fig. 21b and c).
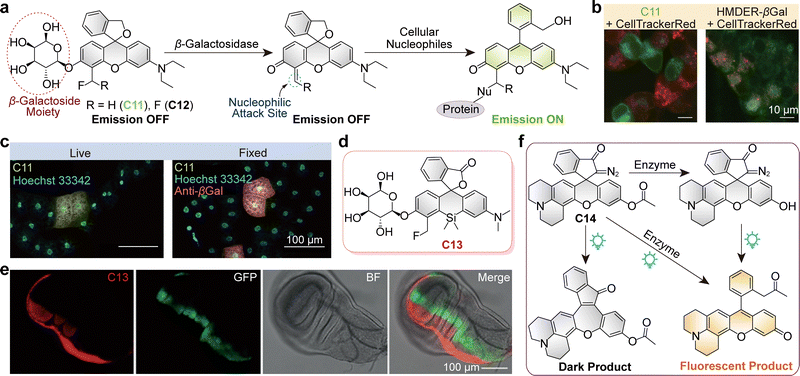 | ||
| Fig. 21 (a) Schematic illustration of enzyme-triggered β-galactoside cleavage and protein-binding-induced ring opening, leading to the fluorescence turn-on of SPiDER-bGal platform (probes C11, C12). Upon enzymatic cleavage of the β-galactoside moiety, fluorescence is activated, and intracellular protein binding occurs concurrently, immobilizing the fluorescent product within living cells. (b) Live imaging of HEK-lacZ(+) and HEK-lacZ(−) cells with C11 (green, left) and HMDER-βGal (green, right). The HEK-lacZ(−) cells were pretreated with CellTrackerRed (red) to distinguish lacZ-negative cells in co-culture imaging. Scale bars: 10 µm. (c) Fluorescent labeling of HEK-lacZ(+) flip-out clones in larval fat body under live and fixed conditions, with nuclear counterstaining using Hoechst 33342 (blue). Immunohistochemical staining of β-galactosidase was performed after fixation (pink, right). Scale bars: 100 µm. (d) Chemical structure of SPiDER-Red-bGal probe C13. (e) Fluorescence imaging of live en-lacZ/dpp-GFP Drosophila third larval wing discs after incubation with C13. Scale bars: 100 µm. β-Galactosidase expression was driven in the posterior compartment of wing discs by the engrailed promoter, whereas GFP expression was restricted to anterior border cells under the control of the decapentaplegic promoter. (f) Proposed mechanism of enzyme- and photoirradiation-co-triggered ring opening leading to fluorescence activation of C14. (b), (c) and (e) are adapted from ref. 144 and 145 with permission from Wiley-VCH, copyright 2016 and 2018, respectively. | ||
Building on this design, Urano and colleagues developed SPiDER-Red-bGal (C13; Fig. 21d), which replaced the rhodol core with a silicon rhodol scaffold to red-shift emission to ∼630 nm.145 The probe retains the same dual mechanism—spirocyclization for OFF/ON control and quinone methide-mediated covalent trapping—but now enables orthogonal multicolor imaging alongside GFP reporters in tissues and live animals (Fig. 21e).
In parallel, Rivera-Fuentes and co-workers expanded the concept of self-immobilization by combining enzymatic activation with photoactivation chemistry (Fig. 21f).146 Their probe (C14) employed a diazoindanone-protected rhodol, which remained non-fluorescent until hydrolyzed by carboxylesterases. Upon light irradiation, the deacetylated intermediate undergoes a Wolff rearrangement, yielding a highly fluorescent rhodol dye. The transient ketene intermediate reacts with nearby macromolecules, covalently trapping the fluorophore and thereby preventing diffusion. This dual-gated (enzyme + photo) system enabled single-molecule localization microscopy (SMLM) of esterase activity with ∼36 nm resolution in live cells.
Together, these examples demonstrate the underlying principle of wash-free bioimaging by coupling fluorogenicity with covalent immobilization. However, the original studies did not explicitly perform wash-free imaging experiments with these rhodol dyes.
6.4. Design strategies for cyanine-based fluorogenic probes
The concept of spirocyclization, originally central to rhodamine fluorogenicity, has been successfully extended to the cyanine dye family, particularly with the goal of improving fluorogenic contrast in the near-infrared (NIR) region. Sustained efforts have focused on modulating the ring-opening equilibrium to suppress background fluorescence while retaining high turn-on signals upon target engagement.Inspired by the rhodamine spirocyclization mechanism, Lin and co-workers first demonstrated that introducing nucleophilic substituents at the meso-position of heptamethine cyanines could trigger intramolecular spirocyclization (C15–C17; Fig. 22a, top panel). This reaction disrupts the polymethine π-conjugation, quenching fluorescence until ring-opening is triggered under appropriate conditions.147 This straightforward “capping” strategy established the feasibility of OFF/ON switching for NIR cyanines. However, early designs suffered from limitations, including residual baseline fluorescence and limited tunability of response thresholds.
Building on this foundation, later studies introduced more refined strategies to tune the cyclization equilibrium. Hiruta and colleagues varied the nucleophile moiety from a hydroxy group to an amide moiety, switching the pKcycl values from ∼10.0 and ∼7.0 (C19, C20; Fig. 22a, bottom panel).148 Furthermore, regulating the probe's hydrophilicity (C21–C24, Fig. 22b, left panel) or micelle encapsulation tuned the effective response window to pH 6.2–8.3, thereby enhancing compatibility with physiological conditions. While these modifications improve environmental sensitivity, the relatively high pKcycl values still limit their use in fluorogenic applications at neutral or mildly acidic pH.
To overcome these constraints, Schnermann and co-workers developed a modular PyBox (pyridinium–benzoxazole) strategy to functionalize heptamethine indocyanines (C25–C28; Fig. 22b, right panel).149 They introduced trifluoroethyl-substituted indolenine units to attenuate the electron-donating strength of the polymethine core (C26) and/or C4′-phenyl-2-acyl-sulfamide moieties to inhibit the electron-withdrawing ring-locking group (C27). This strategy produced an optimized dye (C28) that predominantly adopts the closed form in aqueous media. These modifications significantly enhanced fluorogenicity by promoting selective activation in polar environments.
In a parallel approach, Miki and co-workers developed indocyanine green (ICG) derivatives featuring nucleophilic groups such as amines, hydroxyls, thiols, or carboxylates (C29–C31; Fig. 22c).150 These substituents undergo base-induced intramolecular cyclization to form a non-emissive spirocyclic structure. The pKa of this transition can be tuned by the nature of the nucleophile. Among the tested series, C31 demonstrated reversible ON/OFF switching between pH 3 and 7, indicating its potential for dynamic pH-responsive imaging.
A notable conceptual leap came from Rivera-Fuentes and colleagues, who redesigned the cyclization geometry to favor a 5-exo-trig mechanism, instead of the less favorable 5-endo-trig or 6-endo-trig pathways (Fig. 22d–f).151 This geometric optimization can stabilize the closed form and effectively suppress background fluorescence in cyanine dyes spanning the Cy3, Cy5, and Cy7 scaffolds. By lowering the pKa values below physiological pH (e.g., 6.4 for Cy5 analogues), these dyes remain quenched under resting conditions but exhibit strong fluorescence activation upon binding or environmental shift.
Building on this principle, Wu and co-workers established a general synthetic platform for creating structurally diverse heptamethine cyanines with optimized OFF states (Fig. 22g).152 Their dyes exhibited more than a 100-fold lower baseline fluorescence than conventional Cy7 analogues. They exhibited tunable responsiveness to pH, polarity, and enzymatic activity, while retaining photostability and biocompatibility.
Although many of these spirocyclic cyanines remain mainly in the closed, non-fluorescent form under ambient conditions, binding to target proteins or biomolecular environments—via hydrogen bonding or cation interactions—can shift the equilibrium toward the open, emissive state. This environment-triggered transition underlies their utility as high-contrast, wash-free fluorogenic probes.
Collectively, these advances have transformed cyanine dyes from purely “always-on” NIR fluorophores into a versatile platform for stimulus-responsive imaging. Through innovations in spirocyclization chemistry, fluorogenic cyanines now offer high spatial and temporal precision in biological imaging across the visible and NIR windows.
Across rhodamines, fluoresceins, rhodols and cyanines, spirocyclization has emerged as a unifying mechanism for the design of fluorogenic probes. By tuning log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z, pKcycl, and substituent effects—or by coupling ring opening with enzymatic or covalent strategies—chemists can engineer dyes with tailored responsiveness, low background, and high contrast. These principles underpin the development of next-generation probes for wash-free imaging and super-resolution microscopy.
KL–Z, pKcycl, and substituent effects—or by coupling ring opening with enzymatic or covalent strategies—chemists can engineer dyes with tailored responsiveness, low background, and high contrast. These principles underpin the development of next-generation probes for wash-free imaging and super-resolution microscopy.
6.5. Applications in bioimaging
Recent advances across major scaffolds—fluoresceins, cyanines, and rhodamines—have yielded probes that enable imaging diverse biological targets with high contrast and mechanistic precision. Below, we summarize representative applications, organized by biological targets.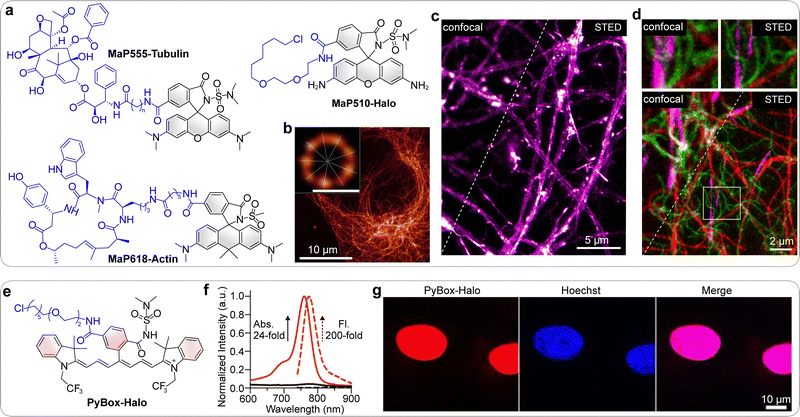 | ||
Fig. 23 (a) Chemical structures of various MaP labels. (b) Wash-free STED imaging of microtubules in live U2OS cells labelled with MaP555-tubulin. Scale bar: 10 µm. Inset shows the deconvoluted image of a selected centriole, with dotted lines indicating the nine intensity maxima corresponding to the nine microtubule triples. Scale bar: 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) nm. (c) Wash-free confocal and STED images revealing periodic actin structures in the axons of rat primary hippocampal neurons labeled with MaP618-actin. Scale bar: 5 nm. (c) Wash-free confocal and STED images revealing periodic actin structures in the axons of rat primary hippocampal neurons labeled with MaP618-actin. Scale bar: 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) µm. (d) No-wash three-color confocal and STED images of live U2OS Vimentin-Halo-expressing cells stained with MaP510-Halo (green), MaP555-tubulin (red), MaP618-actin (magenta), and verapamil for 2 h. The top panels show magnified confocal and STED images of the region indicated by the white box in the bottom panel. Image data were smoothed using a one-pixel Gaussian low-pass filter. Scale bar: 2 µm. (d) No-wash three-color confocal and STED images of live U2OS Vimentin-Halo-expressing cells stained with MaP510-Halo (green), MaP555-tubulin (red), MaP618-actin (magenta), and verapamil for 2 h. The top panels show magnified confocal and STED images of the region indicated by the white box in the bottom panel. Image data were smoothed using a one-pixel Gaussian low-pass filter. Scale bar: 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) µm. (e) Chemical structure of PyBox-Halo label. (f) Absorption (solid line) and fluorescence (dashed line) spectra of PyBox-Halo (black) and PyBox-Halo with HaloTag7 (red) in pH 7.3 HEPES buffer. (g) No-wash live-cell confocal images of U-2 OS cells expressing HaloTag-histone H2B labeled with PyBox-Halo. Scale bars: 10 µm. (e) Chemical structure of PyBox-Halo label. (f) Absorption (solid line) and fluorescence (dashed line) spectra of PyBox-Halo (black) and PyBox-Halo with HaloTag7 (red) in pH 7.3 HEPES buffer. (g) No-wash live-cell confocal images of U-2 OS cells expressing HaloTag-histone H2B labeled with PyBox-Halo. Scale bars: 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) µm. (b)–(d), (f) and (g) are adapted from ref. 139 and 149 with permission from Springer Nature and the American Chemical Society, copyright 2020 and 2023, respectively. µm. (b)–(d), (f) and (g) are adapted from ref. 139 and 149 with permission from Springer Nature and the American Chemical Society, copyright 2020 and 2023, respectively. | ||
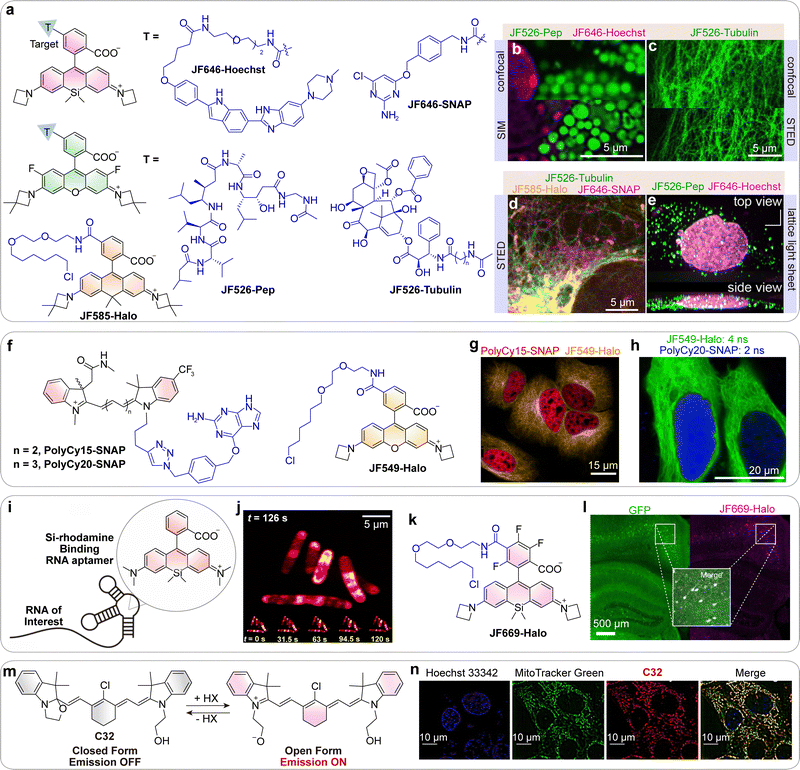 | ||
Fig. 24 (a) Chemical structures of various JF labels. (b) Confocal and SIM images of mouse primary hippocampal neurons labeled with JF526-Pep (green) and JF646-Hoechst (magenta). (c) Confocal and STED microscopy images of U2OS cells labeled with JF526-Tubulin (green). (d) Three-color live-cell STED imaging of U2OS cells expressing Sec61β–SNAP-tag labeled with JF646-SNAP-tag ligand (magenta), TOMM20-HaloTag stained with JF585-Halo-tag ligand (yellow), and microtubules stained with JF526-Tubulin (green). (e) Lattice light-sheet microscopy image of U2OS cells stained with JF526-Pep (green) and JF646-Hoechst (magenta). Scale bars: 5 µm. (f) Chemical structures of the polymethine cyanine label and the JF label used to co-stain cells. (g) Multicolour imaging of live HeLa cells co-transfected with H2B-SNAPf-mTurquoise2 and TUBB5-Halo, and labeled with PolyCy15-SNAP (red) and JF549-Halo (orange). Scale bar: 15![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) µm. (h) Fluorescence lifetime multiplexing of the cells co-transfected with H2B-SNAPf-mTurquoise2 and TUBB5-Halo, and incubated with PolyCy20-SNAP (blue) and JF549-Halo (green). Scale bar: 20 µm. (h) Fluorescence lifetime multiplexing of the cells co-transfected with H2B-SNAPf-mTurquoise2 and TUBB5-Halo, and incubated with PolyCy20-SNAP (blue) and JF549-Halo (green). Scale bar: 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) µm. (i) General RNA labeling strategy using a SiR-binding aptamer. (j) STED imaging of live E. coli expressing GFP-SiR mRNA, showing time-lapse images acquired at 31.5 s intervals over 126 s (bottom) and a magnified view at 126 s (top). (k) Chemical structure of JF669-Halo label. (l) Fluorescent image of a fixed coronal mouse brain slice from an animal expressing GFP-HaloTag fusion protein in neurons after intravenous administration of JF669-Halo. (m) The pH-modulated spirocyclization equilibrium of C32. (n) SIM images of HeLa cells co-stained by Hoechst 33342 (blue), MitoTracker Green (green), and C32 (red). Scale bars: 10 µm. (b)–(e), (g), (h), (j), (l) and (n) are adapted from ref. 132, 151, 153, 140 and 30 with permission from the American Chemical Society, Springer Nature, the American Chemical Society, Springer Nature and the Chinese Chemical Society, copyright 2019, 2023, 2019, 2020 and 2025, respectively. µm. (i) General RNA labeling strategy using a SiR-binding aptamer. (j) STED imaging of live E. coli expressing GFP-SiR mRNA, showing time-lapse images acquired at 31.5 s intervals over 126 s (bottom) and a magnified view at 126 s (top). (k) Chemical structure of JF669-Halo label. (l) Fluorescent image of a fixed coronal mouse brain slice from an animal expressing GFP-HaloTag fusion protein in neurons after intravenous administration of JF669-Halo. (m) The pH-modulated spirocyclization equilibrium of C32. (n) SIM images of HeLa cells co-stained by Hoechst 33342 (blue), MitoTracker Green (green), and C32 (red). Scale bars: 10 µm. (b)–(e), (g), (h), (j), (l) and (n) are adapted from ref. 132, 151, 153, 140 and 30 with permission from the American Chemical Society, Springer Nature, the American Chemical Society, Springer Nature and the Chinese Chemical Society, copyright 2019, 2023, 2019, 2020 and 2025, respectively. | ||
6.6. Advantages and design versatility
Cyclization has emerged as one of the most widely adopted strategies for engineering fluorogenic dyes, particularly in the rhodamine family. This mechanism has a proven track record: established scaffolds such as the JF series, and MaP dyes have already been optimized for super-resolution imaging, in vivo labeling, and biosensing, demonstrating both versatility and reliability.132,138–140,154Spirocyclization offers exceptionally high ON/OFF contrast. The closed spirolactone is nearly colorless and nonfluorescent, whereas the open zwitterion absorbs strongly (ε ∼ 105 M−1 cm−1) and can reach quantum yields as high as 0.8–0.9.132,155 This stark photophysical disparity underpins the excellent signal-to-background ratios that can be achieved in wash-free imaging experiments. Notably, this feature has been exploited in self-labeling tags, including SNAP-, CLIP-, and HaloTag systems.132,156
Spirocyclization also governs cell permeability. The neutral closed form efficiently crosses lipophilic membranes, but once inside the cell, binding-induced ring opening traps the dye in its charged zwitterionic state, enhancing intracellular retention. This mechanism solves a fundamental paradox in probe design: reconciling the need for membrane permeability with the requirement for high polarity and brightness in the bound state.
Another strength is the scaffold's design flexibility. The equilibrium can be shifted by systematic modifications of substituents that exploit electronic push–pull effects, or by altering the bridging atom from oxygen to carbon, silicon, or even phosphorus and sulfur.138,157–162 These strategies have enabled the development of probes that span the visible to near-infrared range. Beyond electronics, steric effects also provide a tunable design axis. Harbron and co-workers demonstrated that bulky substituents at the spirocyclic junction destabilize the closed form, increasing pKcycl and thereby tuning fluorogenicity,163 suggesting further opportunities for rational scaffold engineering.
Furthermore, the spirocyclization of xanthene dyes can also be modulated by light through direct photochemical mechanisms or indirect, protein-mediated strategies.146,164 In certain xanthenes, light irradiation can directly induce structural changes within the dye molecule.165,166 These changes often involve the cleavage or rearrangement of specific functional groups or substituents,146,164 which in turn alters the electronic distribution of the molecule, enabling light-triggered ring opening and fluorescence activation.
In contrast to these direct photochemical strategies, Walterspiel et al. developed an indirect light-controlled system in which fluorescence activation is mediated by protein conformational changes (Fig. 25).167 Their psHaloTag construct integrates a LOV2 domain that responds to 450 nm light, triggering a structural rearrangement that alters the dye-binding environment and promotes ring opening. This enables reversible, deep-red fluorescence turn-on with excellent spatiotemporal control in living cells, as well as compatibility with super-resolution imaging techniques such as SMLM.
6.7. Limitations and practical challenges
Despite these clear advantages, the implementation of these spirocyclization mechanism-based probes is constrained by a restrictive equilibrium “sweet spot”. A primary bottleneck is the thermodynamic fragility of the ring-opening/closing balance. Optimal performance typically necessitates a narrow log![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) KL–Z window of −3 to −2.132,154 Achieving this precise equilibrium requires iterative and exhaustive molecular tuning, as even minor structural modifications can shift the population entirely toward the non-fluorescent lactone or the chronically “ON” zwitterion. This narrow functional range renders the rational design of new fluorophore scaffolds inherently inefficient and labor-intensive.
KL–Z window of −3 to −2.132,154 Achieving this precise equilibrium requires iterative and exhaustive molecular tuning, as even minor structural modifications can shift the population entirely toward the non-fluorescent lactone or the chronically “ON” zwitterion. This narrow functional range renders the rational design of new fluorophore scaffolds inherently inefficient and labor-intensive.
Another critical limitation is environmental variability. Because the equilibrium is highly sensitive, factors such as pH, ionic strength, and macromolecular crowding can shift the balance unpredictably across different intracellular compartments.135 This lack of environmental robustness represents a significant barrier to quantitative imaging. Coupling these dyes to well-defined target groups (e.g., HaloTag or SNAP-tag) can minimize variability and enhance specificity.
Furthermore, current photoactivated spirocyclization systems face a “spectral bottleneck” regarding their activation wavelengths. Most existing platforms rely on high-energy, short-wavelength light (UV or blue) for ring-opening, which induces significant phototoxicity and limits tissue penetration. A key identification of the field's current stagnation is the lack of red- or NIR-responsive molecular switches that maintain efficient spirocyclization kinetics. Overcoming this limitation through the engineering of low-energy light-responsive motifs or dye–protein interfaces is essential for extending the utility of these probes to deep-tissue and long-term live-cell imaging.
Finally, the synthetic complexity of spirocyclization-tuned dyes should not be overlooked. Fine control over KL–Z often requires precise substituent modifications or bridging-atom replacements, which may demand multi-step synthesis and expertise in complex organic synthesis,138 restricting the rapid expansion of the spirocyclization toolbox.
6.8. Outlook and future directions
Although rhodamines, cyanines, fluoresceins, and rhodols have long dominated spirocyclic fluorogenic probe design, extending this concept to alternative dye families offers a compelling opportunity for innovation. Recent work by Hao et al. demonstrated that certain BODIPY derivatives can undergo spirocyclization to modulate fluorescence in response to environmental cues (C33–C38; Fig. 26), highlighting the potential of alternative dye families to diversify fluorogenic systems and enable new photophysical behaviors.168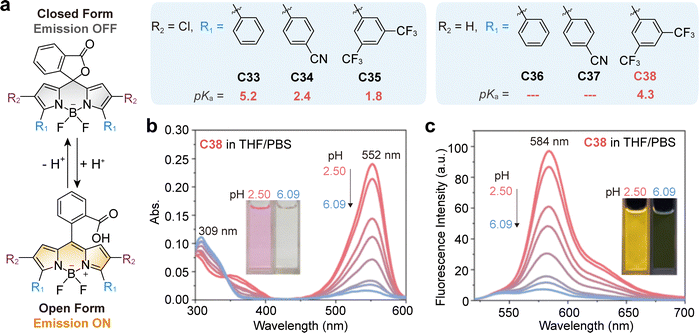 | ||
Fig. 26 (a) Chemical structures and the acid–base induced structure conversion of BODIPY derivatives between a non-emissive closed form and an emissive open form. (b) and (c) Absorption and fluorescence spectra of C38 at various pH values in THF-PBS mixtures (V![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) V = 1 V = 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) : :![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1). Insets show the corresponding images of C38 solutions with different pH under daylight or UV light. (b) and (c) are adapted from ref. 168 with permission from Wiley-VCH, copyright 2025. 1). Insets show the corresponding images of C38 solutions with different pH under daylight or UV light. (b) and (c) are adapted from ref. 168 with permission from Wiley-VCH, copyright 2025. | ||
A major challenge ahead is translating spirocyclization into the deep near-infrared (NIR-II) region (>1000 nm), which is critical for clinical imaging and long-path tissue penetration. While hybrid rhodamine-cyanine architectures (e.g., 2X-rhodamine) have achieved significant milestones, designing fully spirocyclic dyes that combine NIR-II emission with high brightness and chemical stability remains a synthetic frontier. Approaches such as enforcing geometric constraints (e.g., 5-exo-trig cyclization), rigidifying frameworks, and introducing steric or electronic shielding to manage hydration and nucleophilic attack will be essential for stabilizing these chromophores under physiological conditions.
Another priority is optimizing the kinetics of spirocyclization for super-resolution microscopy. Spontaneously blinking dyes, such as cyanines and rhodamine probes, illustrate how spirocyclization equilibria inherently generate the “ON–OFF” duty cycles required for single-molecule localization microscopy (SMLM).9,151,169,170 Future designs should move beyond thermodynamic tuning to systematically control switching rates, which govern localization precision and temporal resolution in live-cell nanoscopy without reliance on toxic redox buffers.
Finally, side-chain engineering provides a versatile approach for tailoring probe performance in complex biological environments.171 Strategic incorporation of small polar or charged substituents can enhance solubility, reduce nonspecific adhesion, and enable target-specific activation, thereby improving both permeability and functional precision.172,173
7. Hydrogen bond-induced quenching
Spirocyclization provides a powerful intramolecular switch triggered by target binding. In contrast, fluorogenicity can also be driven by intermolecular solvent effects, such as hydrogen bond-induced quenching (HBQ), where the presence of water (or other protic solvents) serves as the quenching stimulus. In aqueous or other protic environments, fluorophores bearing hydrogen-bond acceptor sites interact strongly with solvent molecules. These interactions open nonradiative decay pathways that suppress fluorescence, leaving the dyes weakly emissive or completely dark. In contrast, when the same molecules partition into aprotic or hydrophobic microenvironments, such quenching channels are minimized, and intrinsic emission is restored. This environmentally triggered “OFF–ON” response enables probes to remain silent in bulk cytoplasm. At the same time, they selectively light up in compartments where water is excluded, such as lipid droplets, lipid bilayer membranes, or hydrophobic protein pockets.7.1. Design principles and mechanistic insights
The defining feature of HBQ-active fluorophores is the presence of functional groups that engage strongly with protic solvents. Such interactions open nonradiative deactivation pathways that efficiently suppress emission. Many HBQ probes adopt a donor–acceptor (“push–pull”) architecture, where electron-donating substituents (e.g., dialkylamino groups) and electron-withdrawing groups (e.g., carbonyl or cyano moieties) are positioned on a conjugated scaffold. Upon photoexcitation, this arrangement produces a polar ICT state. Hydrogen bonding at electron-accepting sites stabilizes non-emissive states or promotes vibrational relaxation, leading to quenching (Fig. 27a).174–176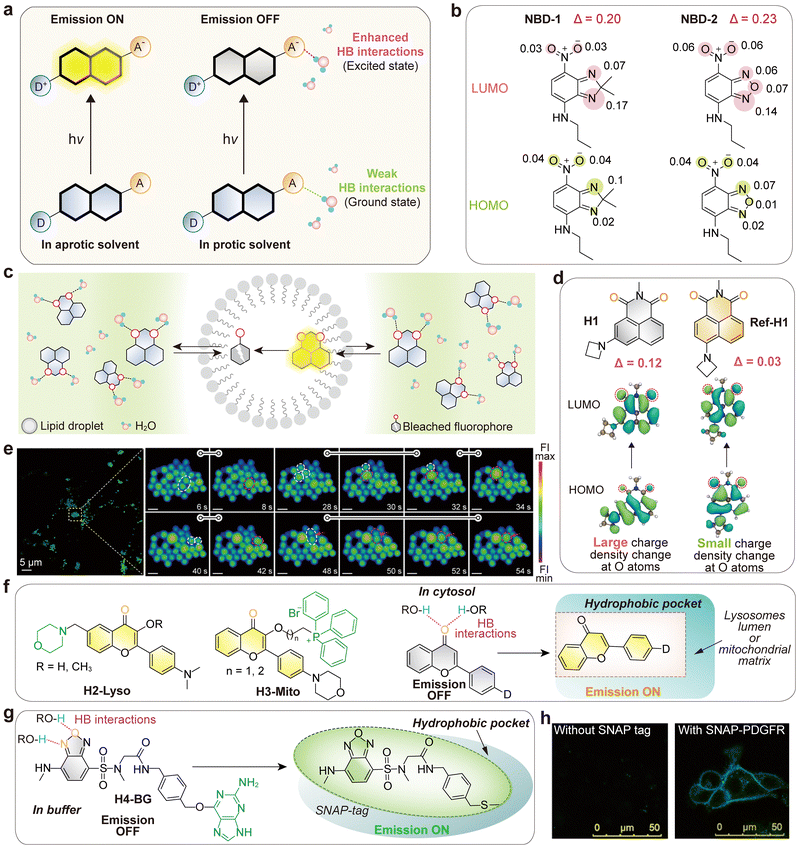 | ||
| Fig. 27 (a) Schematic illustrations of the hydrogen bond-induced quenching in protic solvents. (b) Atomic contributions to HOMO and LUMO of NBD-1 (left) and NBD-2 (right), calculated from the optimized S0 structure in ethanol. Green and pink regions indicate atomic contributions to the HOMO and LUMO, respectively. Δ denotes the sum of the absolute change of atomic contributions of HOMO and LUMO at each hydrogen bond accepting site (highlighted in green and pink). All calculations were performed at the M06-2X/Def2-SVP level in ethanol. (c) Schematic illustration of the cytosolic probe pool to act as a quenched reservoir that replenishes photobleached LD-localized probes. (d) Distributions of HOMO and LUMO of H1 and its reference during S1 de-excitation in water. The corresponding hydrogen-bond-accepting sites and Δ values were labeled. All calculations were performed at the CAM-B3LYP/Def2-SVP level. (e) SIM imaging of live HeLa cells stained with H1, together with locally enlarged views of the boxed region showing dynamic LD coalescence over 1 min. Five LD–LD coalescence events were analysed separately and are indicated by pin labels. Between 48 and 54 s, the intermediate state of LD coalescence is highlighted by a red arrow. (f) Chemical structures of flavone-based probes (H2-Lyso, H3-Mito) exhibiting hydrogen bond-induced quenching in the cytosol and their fluorescence turn-on upon localization in hydrophobic organelle pockets. (g) Chemical structure of H4-BG showing hydrogen bond-induced quenching in aqueous buffer, and its fluorescence turn-on upon covalent reaction with SNAP-tag protein. (h) Fluorescence imaging of living cells without exogenous SNAP-tag expression (left), and with SNAP-PDGFR on the cell surface (right). (e), (d) and (h) are adapted from ref. 29 and 177 with permission from Wiley-VCH and the American Chemical Society, copyright 2021 and 2014, respectively. | ||
In the ground state, these dyes establish a dynamic hydrogen-bonding equilibrium with surrounding protic solvents. Upon excitation, charge redistribution disrupts this equilibrium, and the extent of the disruption correlates with quenching efficiency. To quantify this effect, Liu and co-workers introduced the HBQ index (Δ).178 This index is derived from computational analyses of changes in orbital contributions at hydrogen-bond-accepting atoms between the HOMO (approximating the ground state) and the LUMO (approximating the excited state). A considerable Δ value reflects significant charge redistribution, which destabilizes hydrogen bond equilibria and enhances nonradiative decay. Using this descriptor, they rationalized the diverse emission behaviors of nitrobenzoxadiazole (NBD, Fig. 27b) derivatives, also known as SCOTfluors, showing that probes with higher Δ values are dark in water but strongly emissive in lipid membranes.178 Conversely, fluorophores with small Δ values typically resist HBQ. Consequently, these compounds generally lack this specific environmental sensitivity if no alternative relaxation pathways exist.
Despite these advances, the mechanistic understanding of hydrogen bond-induced quenching remains limited compared with more established processes such as PET and TICT. While descriptors such as the HBQ index provide a helpful first step toward quantification, the precise quenching channels remain unresolved. It is plausible that strong hydrogen-bonding interactions may distort the excited-state geometry of a fluorophore, facilitating access to nonradiative decay funnels, such as conical intersections. Similarly, hydrogen bonding could cooperate with other pathways, such as TICT formation or PET, suggesting that HBQ is better understood as a synergistic effect rather than a standalone mechanism. Moreover, the HBQ index focuses primarily on changes in charge density at hydrogen-bond-accepting sites, yet additional structural factors may also play decisive roles. Rotations or torsional motions induced by photoexcitation, for example, could further destabilize the delicate ground-state equilibrium and accelerate quenching. These open questions highlight that HBQ remains an emerging area, with considerable room for mechanistic refinement and probe innovation.
7.2. Applications in bioimaging
HBQ-based fluorophores have been applied to a broad range of biomolecular targets, leveraging their silent cytosolic background and sharp fluorogenic response in hydrophobic niches.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) P) allowed the cytosolic probe pool to act as a quenched reservoir that replenished photobleached LD-localized probes (Fig. 27c), sustaining long-term imaging under super-resolution conditions. More recently, Huang and colleagues extended HBQ into the NIR region by designing compact, rigidified dyes with strategically placed HB-acceptor sites.174 These probes were dark in water but emitted brightly in lipid droplets and bacterial membranes, offering deep red/NIR emission suitable for translational applications.
P) allowed the cytosolic probe pool to act as a quenched reservoir that replenished photobleached LD-localized probes (Fig. 27c), sustaining long-term imaging under super-resolution conditions. More recently, Huang and colleagues extended HBQ into the NIR region by designing compact, rigidified dyes with strategically placed HB-acceptor sites.174 These probes were dark in water but emitted brightly in lipid droplets and bacterial membranes, offering deep red/NIR emission suitable for translational applications.
Collectively, these examples illustrate the versatility of HBQ-based probes. From organelle-specific trackers to protein-labeling reagents, they provide wash-free imaging with high signal-to-background ratios and have been successfully extended into the NIR window for in vivo applications.
7.3. Advantages and design versatility
The defining advantage of HBQ lies in its direct utilization of water, the most abundant cellular component, as the switching element.181 Unlike PET, which requires precise redox matching, or EnT, which depends on a paired acceptor, HBQ provides a robust, intrinsic readout of local hydrophobicity. This mechanism is therefore highly effective for distinguishing the aqueous cytosol from lipid-rich organelles, membranes, or hydrophobic protein pockets.From a synthetic perspective, accessing HBQ probes is remarkably efficient. It typically requires only the strategic installation of hydrogen-bond acceptor groups (e.g., carbonyl, nitro, or cyano substituents) onto a donor–acceptor scaffold, obviating the need for bulky external quenchers or complex linkers. Crucially, not all hydrogen-bond acceptors are susceptible to HBQ; only those that undergo significant charge transfer form strong hydrogen-bond interactions and experience considerable quenching. The recent development of the HBQ index (Δ) has further refined this process, offering a computational descriptor that aids in rationalizing quenching efficiency and advancing the field toward more targeted molecular engineering.
Consequently, the design space for HBQ is broad. Rigidified HBQ dyes have recently extended this mechanism into the deep-red and near-infrared (NIR) regions. At the same time, applications have expanded to include lipid droplets, lysosomes, mitochondria, and SNAP-tag protein labelling—all achieved under wash-free conditions.
Perhaps the most distinct benefit of HBQ is its reversibility.174,180,182,183 Unlike enzyme-activated probes that rely on irreversible cleavage, HBQ probes maintain a dynamic equilibrium with their environment. This reversibility enables a “self-healing reservoir effect” during long-term, high-intensity imaging. Because the probe remains dark and stable in the aqueous cytosol, this cytosolic pool continuously partitions into the target compartment to replenish fluorophores photobleached under STED or SIM illumination. This dynamic replenishment enables extended super-resolution imaging, which is often unachievable with permanently tethered or irreversible reaction-based systems.
7.4. Limitations and practical challenges
Despite these strengths, the practical implementation of HBQ is hindered by an unresolved mechanistic foundation. While the HBQ index (Δ) correlates well with quenching efficiency, HBQ rarely operates as a standalone mechanism. A primary bottleneck is the unresolved cooperative interplay between HBQ and other competing pathways, such as conical intersections, TICT, and PET, but the relative contributions of these channels remain unclear. This mechanistic ambiguity complicates predictive design, often necessitating empirical screening to fine-tune performance.Specificity presents a second challenge. Because HBQ responds to a physical property (hydrophobicity) rather than a specific molecular structure, off-target activation is an inherent risk. Semi-hydrophobic environments, such as plasma membranes, serum albumin, or crowded cytosolic assemblies, can partially shield hydrogen-bonding sites, thereby restoring fluorescence. Achieving high-contrast imaging of a specific hydrophobic pocket, therefore, requires meticulous validation to distinguish signal from non-specific background.
Furthermore, the reliance on hydrogen-bonding networks introduces a “pH-convolution liability”. Protonation directly alters hydrogen-bond strengths and the probe's interaction with water. In acidic organelles like lysosomes, fluorescence intensity becomes a convolution of both water content and pH, potentially complicating quantitative interpretation.
Ultimately, the development of HBQ probes is hindered by an inherent trade-off associated with solubility. To function as a “dark” reservoir, the probe must be soluble in aqueous cytosol; however, it must also exhibit sufficient lipophilicity to partition into hydrophobic targets. Overly lipophilic structures may aggregate, excluding water and triggering non-specific fluorescence (false positives), while excessive hydrophilicity may prevent target integration. Precise tuning of lipophilicity (clogP) is therefore essential to balance solubility with quenching efficiency.
7.5. Outlook and future directions
A primary frontier in HBQ research is to resolve the mechanistic ambiguity surrounding it. The current HBQ index (Δ) provides a useful but static descriptor that fails to capture the dynamic interplay between excited-state reorganization and fluctuations in the water network. Integrating femtosecond transient absorption spectroscopy (fs-TAS) with TD-DFT and nonadiabatic dynamics in explicit-solvent environments would enable the field to map hydrogen-bond-induced funnels and quantify coupling to conical intersections. Such mechanistic clarity is a prerequisite for the de novo design of probes with activation thresholds precisely tailored to specific hydrophobic niches.Expanding HBQ into the NIR window remains a critical synthetic challenge. Long-wavelength fluorophores typically require extended conjugation, which increases lipophilicity and promotes aggregation—factors that erode the aqueous “dark state.” Future architectures will likely prioritize compact, rigid cores with electronically coupled H-bond acceptor sites that sustain NIR emission without compromising water solubility.
Another promising direction is to hybridize HBQ with reactive trapping strategies. By incorporating electrophilic warheads, HBQ probes could be designed to form covalent bonds upon entering a target site. This “lock-in” strategy would combine the low background of HBQ with the spatial fidelity of covalent labeling, preventing diffusion and reducing background from reversible partitioning.
Finally, optimizing the kinetics of the “reservoir effect” represents a significant opportunity for super-resolution microscopy. Future efforts should focus on tuning the ON/OFF exchange rates to maximize the replenishment of photobleached dyes, thereby establishing HBQ probes as the premier tool for long-duration, high-intensity live-cell imaging.
8. Aggregation-induced emission
HBQ probes excel at reporting on local hydrophobicity by interacting with water. A related but distinct strategy for wash-free imaging leverages phase-state transitions, such as AIE. AIE is a photophysical phenomenon observed in specific fluorophores, known as AIE luminogens (AIEgens). These molecules exhibit minimal fluorescence when freely dissolved but display intense emission upon aggregation. In bioimaging, aggregation can be induced when binding to biomacromolecules, localization within organelles, or partitioning into lipid-rich environments, thereby triggering fluorescence activation through motion restriction.8.1. Photophysical mechanisms of AIE
Significant efforts have been dedicated to elucidating AIE mechanisms, which are generally rationalized in terms of either geometrical or electronic structural changes responsible for fluorescence enhancement upon aggregation.184–187 From a geometrical structure perspective, the restriction of intramolecular motion (RIM) mechanism proposed by Tang and co-workers has been widely accepted as the dominant origin of AIE behavior (Fig. 28a and b).188,189 Such motions can be further classified into rotations (restriction of intramolecular rotations, RIR, Fig. 28a), vibrations (restriction of intramolecular vibrations, RIV, Fig. 28b), and even bending (restriction of intramolecular bending, RIB, Fig. 28c).190 These molecular motions dissipate the excited-state energy via non-radiative decays, leading to no (or weak) fluorescence in solution. However, upon aggregation of AIEgens, intramolecular rotations and vibrations are restricted, enabling bright fluorescence.191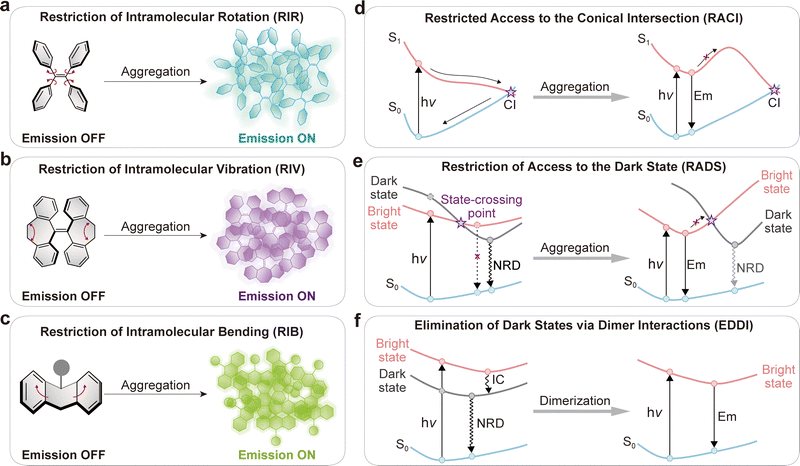 | ||
| Fig. 28 Schematic illustrations of various underlying photophysical insights involved in the AIE mechanism. | ||
From an electronic-structure standpoint, the restricted access to a conical intersection (RACI, Fig. 28d) mechanism has emerged as a widely invoked model to rationalize the AIE effect.192 The CI is the region of the potential energy surface (PES) where the ground and excited states become quasi-degenerate, resulting in a high probability of non-radiative internal conversion.193,194 Excessive access to CI in dilute solution is responsible for efficient fluorescence quenching. By contrast, in the aggregate or solid state, CI often becomes energetically unfavorable owing to restricted geometrical relaxation, thereby suppressing non-radiative decay pathways and leading to fluorescence recovery. Restriction of TICT,195,196 suppression of PET,68 restriction of access to the dark state (RADS, Fig. 28e),197 and elimination of dark states via dimer interactions (EDDI, Fig. 28f) mechanisms198 can also be employed to account for the AIE phenomenon of certain molecules. In these cases, the formation of a stable, complete charge-transfer (CT) state, or other low-lying dark states (e.g., n–π*), can efficiently quench the fluorescence of AIEgen monomers. Upon aggregation, restricted molecular motions and/or reduced local polarity hinder the population of these dark states, thereby restoring radiative decay and enabling intense fluorescence.
Readers interested in detailed theoretical discussions are encouraged to consult comprehensive reviews by Tang and colleagues.191,199,200
8.2. Applications in bioimaging
The discovery of fluorophores with AIE characteristics has opened new opportunities to develop fluorogenic probes for specific biological applications, such as organelle imaging (mitochondrial tracking,201–203 lysosome imaging,204 lipid droplet imaging205), cell membrane staining,205,206 protein dynamics tracking,207,208 enzymatic activity monitoring,209–211 bacterial imaging212,213 and in vivo tumor imaging.214Tetraphenylethene (TPE), a prototypical AIEgen, has been widely employed in organelle imaging. Tang and co-workers reported A1 (Fig. 29a), a TPE–triphenylphosphonium (TPP) conjugate that enables long-term visualization of mitochondrial morphology with high photostability (Fig. 29b).201 Unlike commercial dyes such as MitoTracker Red FM (MT), which lose specificity under carbonyl cyanide m-chlorophenylhydrazone (CCCP)-induced depolarization, A1 retains targeting fidelity due to its dual positive charges and lipophilic TPE core, which together stabilize mitochondrial localization and broaden responsiveness to membrane potential fluctuations (Fig. 29c). However, its short blue emission limits imaging contrast and resolution in deep tissue.
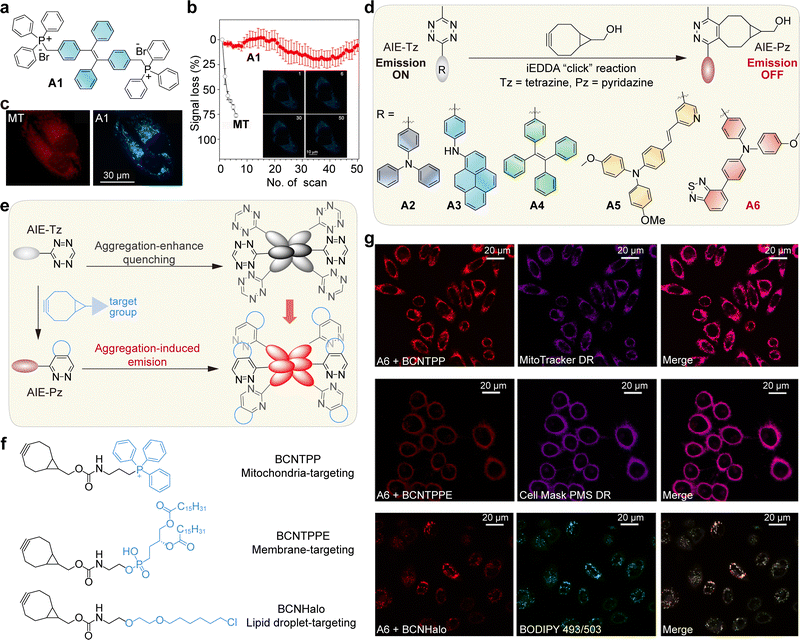 | ||
| Fig. 29 (a) Chemical structures of TPE-based mitochondria-targeted probe A1. (b) Signal loss (%) of the fluorescence intensity of A1 (red solid circle) and MitoTracker red FM (MT, black open circle) as a function of scan number. Inset: Fluorescent images of living HeLa cells labeled with A1 with an increasing number of scans (1–50 scans; the number of scans shown in the upper right corner). (c) Fluorescent images of carbonyl cyanide m-chlorophenylhydrazone (CCCP) treated HeLa cells stained with MT (red, left) and A1 (blue, right). CCCP is a protonophore uncoupler that can induce rapid mitochondrial acidification and impairs ATP synthase activity, leading to a reduction in the mitochondrial membrane potential (ΔΨm). (d) Schematic illustration of the fluorescence turn-on mechanism of the AIE-Tz probes (A2–A6), triggered by bioorthogonal cycloaddition with BCN-OH. (e) Proposed working mechanism for the “Matthew effect” in the aggregate emission of AIE-Tz probes. Aggregation quenches the emission of AIE-Tz but enhances the emission of AIE-Pz after tetrazine removal by click reactions, resulting in high fluorescence enhancement. (f) Chemical structures of bicyclononyne (BCN) adducts carrying various targeting units. (g) Imaging of the living cells incubated with A6. Fluorescence activation occurs via in situ reactions of A6 with BCN adducts equipped with various targeting units for mitochondria (red, top), cell membrane (red, middle), and lipid droplet (red, bottom). MitoTracker DR (deep red), Cell Mask PMS DR (deep red), or BODIPY 493/503 (green) were used to stain cells as the corresponding reference controls. (b), (c) and (g) are adapted from ref. 201 and 218 with permission from the American Chemical Society and Wiley-VCH, copyright 2013 and 2024, respectively. | ||
Tetrazine-based fluorogenic dyes have been widely explored, with significant efforts focused on improving the emission enhancement ratio (IAC/IBC) of the post-click product.218 To address this challenge, Tang and co-workers developed tetrazine-modified AIE-Tz probes (A2–A6; Fig. 29d) that exhibit weak emission across the blue-to-red region in both monomeric and aggregated states (Fig. 29e). Theoretical studies revealed a TICT–ICDS quenching pathway, intensified upon aggregation (due to intermolecular interactions). Notably, the iEDDA click reaction does not restore monomer fluorescence due to persistent TICT, but the resulting pyridazine products show strong AIE. This “Matthew effect” arises from synergistic tetrazine-induced quenching and intrinsic AIE (Fig. 29e). Building on this, a red-emissive AIEgen was conjugated with targeting ligands for selective imaging of mitochondria, membranes, and lipid droplets (Fig. 29f and g). This strategy offers a general approach to achieving ultrahigh fluorogenicity.
Guo and co-workers reported an AIE probe (A7; Fig. 30) that remains non-emissive in the Aβ-unbound state due to a free-rotation-facilitated TICT process.221 Compared to commercial probes (e.g., ThT), A7 avoids ACQ and offers NIR emission, a large Stokes shift, ultrahigh signal-to-noise ratio, strong Aβ binding, and blood–brain barrier permeability (Fig. 30), enabling high-fidelity in vivo imaging of Aβ plaques in Alzheimer's disease.
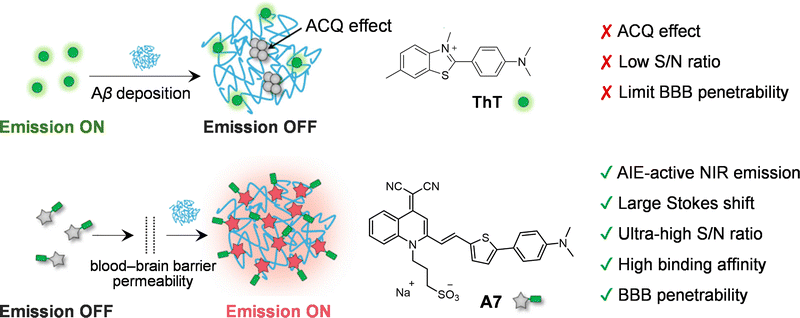 | ||
| Fig. 30 Comparison of A7 with widely used ThT for Aβ aggregates sensing. ThT, a commercial gold standard for the detection of Aβ protein aggregates, fails to detect Aβ plaques in living animals because of its ACQ effect, low S/N ratio, and limited blood–brain barrier (BBB) permeability. Probe A7 is a near-infrared AIE–active probe enabling in vivo and in situ ultrasensitive lighting up of Aβ plaques. Fig. 30 is reproduced from ref. 221 with permission from Springer Nature, copyright 2023. | ||
Yu, Kim, and co-workers reported a water-soluble AIE probe (A8; Fig. 31a) with a hydrophobic tail and cationic head, enabling rapid, selective plasma membrane staining (Fig. 31b–e) via dual hydrophobic and electrostatic interactions (Fig. 31a).206 This design allowed non-invasive imaging of neuronal and erythrocyte membranes in the brain (Fig. 31c and d). However, the fluidity of phospholipid bilayers often leads to probe displacement, limiting long-term visualization of dynamic structures such as migrasomes.
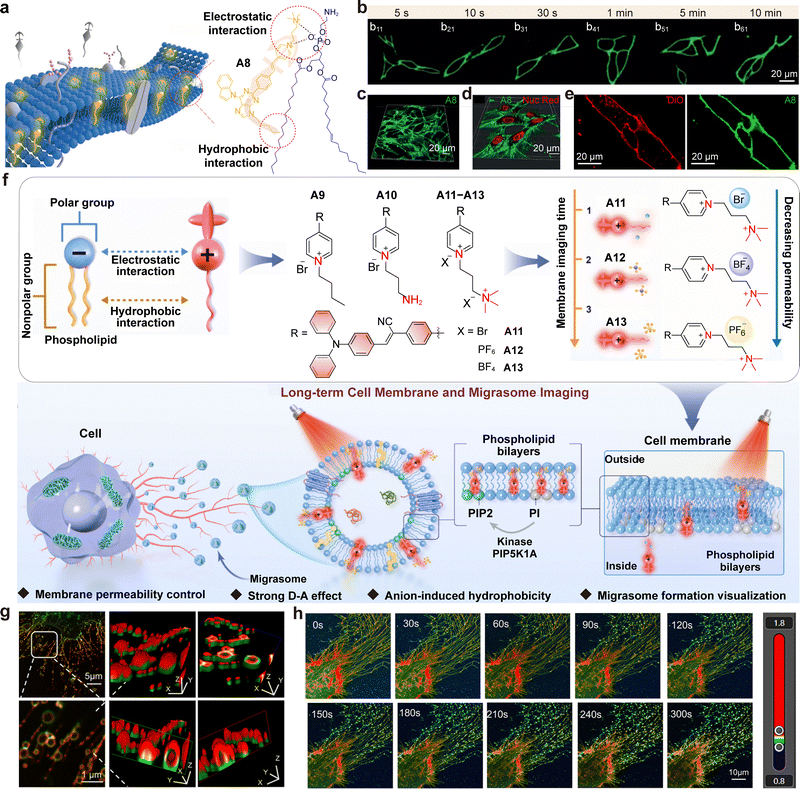 | ||
| Fig. 31 (a) Schematic representation of plasma membrane imaging using AIE-active probe A8, featuring a hydrophilic head group and a hydrophobic tail group. (b) Imaging of B16 cells stained for different times using A8. (c) and (d) The 3D reconstructed image of primary hippocampal neurons stained with (c) A8, and (d) A8 (green) and NucRed (red). NucRed was used to stain the cell nucleus. (e) Imaging of live B16 cells incubated with DiO (red, left) and A8 (green, right). DiO was used to stain the cell membrane as a reference control. (f) Schematic illustration of the molecular design strategy and membrane permeability modulation of probes A9–A13 for migrasome imaging. Top: The schematic shows stepwise modifications of the probe structures aimed at optimizing long-term imaging capability, including the addition of alkyl chains to the pyridinium moiety, terminal amine functionalization, and conversion to tertiary ammonium salts with varying counterions (Br−, BF4−, and PF6−). Membrane permeability is modulated by alkyl chain design and changes in counterion size. Bottom: The schematic highlights the probe's preferential binding to phosphoinositide-rich membrane regions, particularly phosphatidylinositol 4,5-bisphosphate (PIP2), within the plasma membrane and migrasomes. (g) Confocal 3D microscopy of H1299 cells stained with A12 (red) and co-labeled with WGA (green), a plasma membrane marker used to validate A12 localization. (h) Confocal microscopy lifetime imaging of H1299-TSPAN4 cells labeled with A12 during migrasome formation. (a)–(e) and (f)–(h) are adapted from ref. 206 and 222 with permission from Wiley-VCH and the American Chemical Society, copyright 2019 and 2025, respectively. | ||
To overcome this, Tang and colleagues developed a series of NIR-emissive AIE probes (A9–A13; Fig. 31f, top panel) incorporating dicationic donor–π–acceptor motifs for selective and long-term membrane and migrasome imaging (Fig. 31f, bottom panel).222A12 stood out due to its bulky PF6− counterion, which mitigated ACQ and enhanced lipophilicity, enabling dual anchoring via hydrophobic and electrostatic interactions. A12 rapidly accumulated in phosphatidylinositol 4,5-bisphosphate (PIP2)-rich migrasomes within 5 minutes, showing strong lipid affinity (Fig. 31g). Fluorescence lifetime imaging revealed a shift from 1.8 to 0.9 ns during migrasome formation, allowing real-time tracking of membrane remodeling (Fig. 31h). This sensitivity to lipid composition positions A12 as a robust tool for studying lipid dynamics, cellular signaling, and phospholipid-rich organelles under physiological and pathological conditions.
These membrane probes generally share a common structural motif comprising a positively charged headgroup and a hydrophobic alkyl tail, which together enable anchoring through electrostatic and hydrophobic interactions. For effective wash-free imaging, a PM probe should remain confined to the extracellular leaflet, either by being membrane-impermeable or by exhibiting selective affinity, thereby avoiding undesired intracellular entry. In this context, highly charged or polar structures typically display low membrane permeability, whereas small, lipophilic molecules are prone to diffuse across the bilayer, compromising membrane-specific labelling. Increasing molecular size, for example by incorporating bulky substituents, can effectively suppress passive permeation by reducing bilayer crossing efficiency, thereby improving surface specificity.
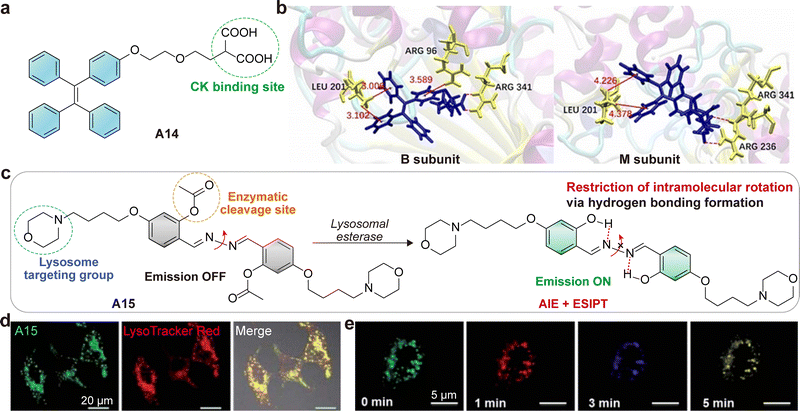 | ||
| Fig. 32 (a) Chemical structure of A14 enabling light-up and selective visualization of CK-B, with very low interference from CK-M. (b) Molecular dynamics simulations illustrate A14 binding within the cavities of CK-B (left) and CK-M (right) subunits. In both subunits, the ethyl malonic acid fragments of A14 are stabilized through hydrogen bonding. Notably, in CK-B, strong C–H⋯π interactions between side-chain methyl groups and the phenyl rings of A14 restrict molecular motion, resulting in markedly enhanced emission. In contrast, such interactions are virtually absent in CK-M due to the long interaction distance (>4.2 Å), which fails to constrain the probe's motion and leads to weak fluorescence. (c) Design of probe A15 for specific fluorogenic detection of lysosomal esterase. (d) Confocal images of MCF-7 cells stained with A15 (green) and LysoTracker Red (red), a lysosome marker used to validate A15 localization. Scale bars: 20 µm. (e) Confocal images of an MCF-7 cell stained with A15 and stimulated using chloroquine, a mild stimulant to induce lysosomal movement. Different pseudo-colors were used to illustrate the fluorescence images at different stimulation times. Scale bars: 5 µm. (b), (d) and (e) are adapted from ref. 223 and 210 with permission from Wiley-VCH and the Royal Society of Chemistry, copyright 2020 and 2014, respectively. | ||
Liu and co-workers developed a lysosome-targeted esterase probe (A15; Fig. 32c) by integrating AIE and excited-state intramolecular proton transfer (ESIPT) mechanisms.210 The salicyladazine fluorophore, modified with acetoxyl and morpholine groups, remained non-emissive due to disrupted hydrogen bonding and N–N bond rotation. Lysosomal esterase-mediated cleavage restored hydroxyl groups, triggering ESIPT, while concurrent aggregation in aqueous activated AIE (Fig. 32c). This dual-channel synergy stabilized proton transfer even in polar environments, yielding bright emission with a large Stokes shift and high signal-to-noise ratio.
The AIE–ESIPT strategy enables sensitive and selective fluorescence activation, exemplified by real-time imaging of lysosomal esterase activity in live cells (Fig. 32d and e). This dual-mechanism design offers a general framework for enhancing enzymatic probe performance beyond conventional AIE limitations.
A representative example is A16 (Fig. 33a), an amphiphilic AIEgen that is weakly emissive in water but exhibits a pronounced fluorescence turn-on upon binding bacteria, thereby eliminating washing steps.225 Its cationic and lipophilic groups anchor into bacterial membranes, inducing aggregation and selective envelope labeling (Fig. 33b). Notably, A16 also acts as a theranostic agent, generating ROS under light to kill bound bacteria (Fig. 33c). This “image-and-destroy” strategy showcases the potential of dual-functional AIEgens for microbial detection, infection monitoring, and light-triggered antimicrobial therapy.
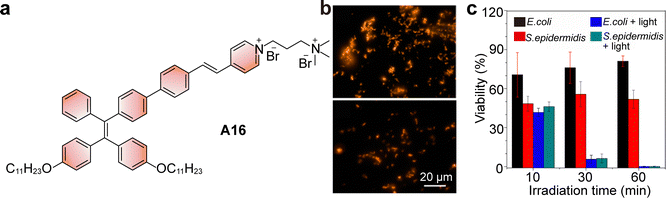 | ||
| Fig. 33 (a) Chemical structure of probe A16, which exhibits turn-on emission upon interaction with bacteria and also functions as a bactericide for bacterial elimination. (b) Fluorescence images of S. epidermidis (top) and E. coli (bottom) incubated with A16. Scale bar: 20 µm. (c) Killing efficiency of A16 against E. coli and S. epidermidis with and without room light irradiation at different time points. (b) and (c) are adapted from ref. 225 with permission from the American Chemical Society, copyright 2015. | ||
Tang et al. reported a water-soluble AIEgen (A17; Fig. 34a) featuring a donor–acceptor framework with piperazine as the donor and a pyridinium salt as the acceptor, resulting in red-shifted absorption and emission, along with improved hydrophilicity.227A17 showed weak red fluorescence (640 nm, φf = 0.9%) in water. For bioconjugation, A17 was functionalized with an NHS ester and linked to cetuximab, forming mAb-A17 (Fig. 34a), which retained bioactivity and selectively targeted EGFR-overexpressing cancer cells.
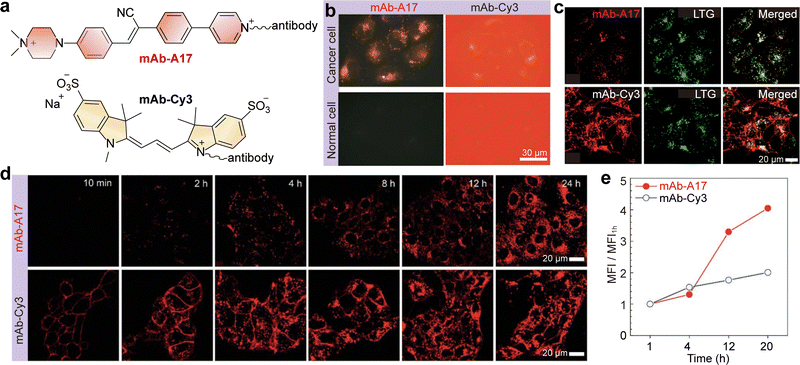 | ||
| Fig. 34 (a) Chemical structures of fluorogenic probe mAb-A17 and the “always-on” probe mAb-Cy3. (b) Fluorescence images of HCC827 cancer cells and normal cells incubated with mAb-A17 and mAb-Cy3. mAb-A17 enables wash-free and high-contrast cancer cell imaging, whereas mAb-Cy3, due to its always-on fluorescence, suffers from strong background interference, making it difficult to distinguish cancer cells from normal cells. Scale bars: 30 µm. (c) Confocal imaging of HCC827 cells stained with mAb-A17 and mAb-Cy3 conjugates for 4 h and then co-stained with LysoTracker Green (LTG) for 5 min. Scale bars: 20 µm. (d) Confocal imaging of HCC827 cells stained with mAb-A17 at different incubation times without PBS washing, and mAb-Cy3 at different incubation times after PBS washing. Scale bars: 20 µm. (e) Flow cytometric analysis of HCC827 cells after incubation with mAb-A17 and mAb-Cy3 at different time intervals. (b)–(e) are adapted from ref. 227 with permission from the Royal Society of Chemistry, copyright 2017. | ||
Compared to conventional “always-on” probes (e.g., mAb-Cy3), which show non-specific fluorescence across membranes and intracellular compartments, mAb-A17 offers minimal background and significantly improved tumor-to-background contrast (Fig. 34b and c). Notably, mAb-A17 fluorescence was absent on the cell surface but gradually increased within HCC827 cells (Fig. 34d and e), reflecting endocytosis-driven internalization.
By selectively targeting EGFR-overexpressing cells and responding to intracellular trafficking, mAb-A17 enables precise tumor delineation and holds strong potential for image-guided surgery.
Finally, it is worth mentioning that certain hydrophobic AIE probes, which can offer advantages such as higher photostability or longer emission wavelengths, exhibit poor solubility in aqueous media. To overcome this limitation, hydrophilic carriers, including polymeric micelles, liposomes, proteins, cyclodextrins, and PEGylated nanoparticles, are frequently employed to encapsulate or modify the probes. Such formulations improve dispersibility without sacrificing imaging performance and extend the utility of hydrophobic AIEgens for biological applications.228
8.3. Advantages and design versatility
The primary allure of AIE-based imaging techniques lies in their exceptional fluorescence enhancement ratios. These dramatic turn-on ratios, spanning tens to thousands, provide superior signal-to-noise ratios that are crucial for detecting low-abundance cellular targets and enabling sensitive single-cell-level imaging. Because the freely diffusing unbound probes remain non-emissive, AIE is intrinsically suited for high-contrast, wash-free imaging.A unique strength of AIE is its suitability for long-term and high-concentration imaging. Upon aggregation, the high molecular density within nano-aggregates ensures that the overall brightness remains robust even if individual fluorophores bleach. This “collective brightness” makes these probes especially suited for long-term time-lapse applications.
Furthermore, AIEgens offer broad structural tunability. By adjusting electron donors and acceptors, introducing heteroatoms, or extending π-conjugation, researchers can tailor emissions across a wide spectral range from the visible to the near-infrared (NIR-II). Beyond spectral tuning, the installation of site-specific anchoring groups enables precise subcellular targeting.
Additionally, many AIEgens possess efficient intersystem crossing pathways, enabling them to act as photosensitizers for image-guided antimicrobial or anticancer phototherapy.
Finally, the development of hydrophilic AIEgens and colloidal formulations (micelles, liposomes, proteins, PEGylated nanoparticles) has overcome early solubility issues. These strategies maintain dispersion in blood, minimize nonspecific aggregation, and retain wash-free contrast in complex animal imaging and image-guided surgery contexts.
8.4. Limitations and practical challenges
Despite their strengths, AIE probes are fundamentally constrained by a lack of specificity. The reliance on aggregation or physical restriction means that the “turn-on” event is inherently sensitive to the environment's physical state (viscosity, crowding) rather than solely to a specific chemical recognition event. This can lead to a non-specific background if the probe aggregates in unrelated hydrophobic compartments or binds non-specifically to sticky proteins. Without secondary recognition elements (ligand, enzyme, or bioorthogonal step), wash-free specificity can drift.The kinetic control of aggregate morphology within complex biological media presents a second challenge. Uncontrolled large aggregates can be taken up by cells via endocytosis, accumulate in lysosomes, and generate false-positive signals distinct from the intended target. Furthermore, these large aggregates may cause steric hindrance, interfering with biomolecule functionality or blocking interactions with other cellular components, thereby introducing artifacts into biological studies.
From a synthetic perspective, balancing solubility is a critical trade-off. To function as a wash-free probe, the AIEgen needs to be soluble enough to remain molecularly dissolved (and thus dark) in the culture medium, yet hydrophobic enough to aggregate avidly at the target site. Achieving this “metastable” solubility window often requires careful tuning of ionic groups and alkyl chains. Poor aqueous solubility can lead to the formation of colloidal or protein-corona aggregates in serum-containing media, thereby activating AIE before target engagement.
Finally, the unintentional generation of ROS represents a persistent photochemical bottleneck. Although AIEgens typically exhibit good photostability, some aggregates may generate ROS under prolonged illumination, causing inadvertent cellular damage. Thus, careful evaluation of phototoxic effects is necessary, particularly when distinguishing between diagnostic imaging and therapeutic applications.
8.5. Outlook and future directions
The future of AIE hinges on transitioning from water-soluble, target-specific AIEgens that avoid nonspecific aggregation to precise, binding-induced restriction of motion. The “artificial fluorescent protein” concept, where AIEgens light up only when bound to a specific protein pocket, represents the ideal paradigm for this approach.To further refine this specificity, a robust strategy is to combine the RIM mechanism with an additional chemical gating event, such as bioorthogonal ligation or enzyme-mediated cleavage. In this “AND-logic” dual-trigger design, fluorescence activation requires both aggregation (physical restriction) and a precise biochemical reaction (chemical unmasking). This multi-gated architecture ensures that the probe remains silent in non-target environments, maximizing contrast and minimizing off-target activation in complex biological systems.
Simultaneously, expanding AIE into the NIR-II window remains a significant priority for in vivo imaging. While classic TPE derivatives provide robust signals, they predominantly emit in the blue/green region. The synthetic frontier now lies in developing red-shifted AIE cores that maintain the necessary rotor mechanism without becoming overly lipophilic, a difficult balance given that extended conjugation often reduces solubility.
Finally, the field is increasingly capitalizing on the theranostic potential of AIEgens. As demonstrated by the bacterial “image-and-destroy” probes, AIEgens can effectively serve dual roles as imaging agents and photosensitizers (ROS generators). Future designs will likely focus on tuning the excited-state energy levels to optimize ROS generation for tumor therapy (photodynamic therapy) while maintaining high-contrast imaging performance.
For comprehensive coverage of AIE research, readers are encouraged to consult dedicated reviews.191,199,200
9. Disaggregation-induced emission
DIE is an increasingly crucial photophysical phenomenon in wash-free fluorescence imaging. Unlike AIE, where fluorophores become emissive upon clustering, DIE describes the reverse behavior: fluorophores that are non-emissive or weakly emissive in their aggregated state exhibit bright fluorescence upon disaggregation into monomeric units (Fig. 35a). This distinctive feature provides a powerful mechanism for designing fluorescence “turn-on” probes and fluorogenic dyes.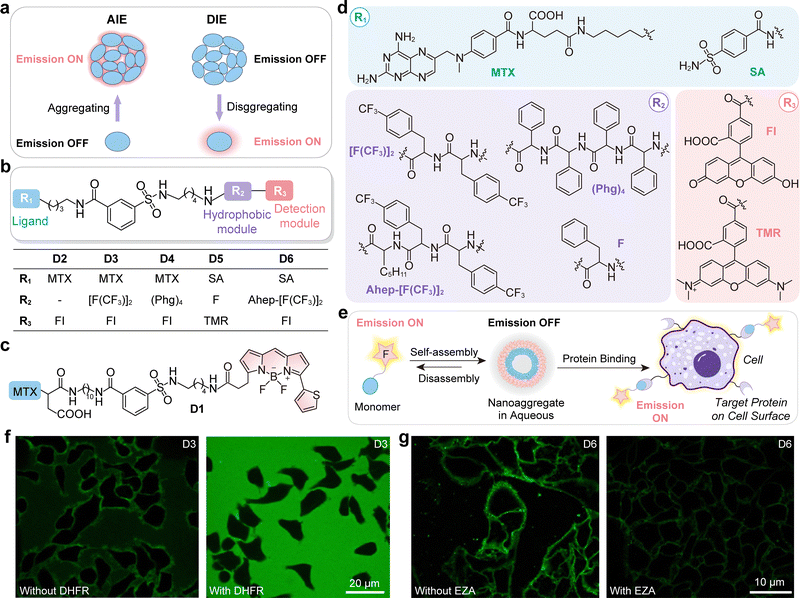 | ||
| Fig. 35 (a) Schematic illustration comparing the AIE and DIE phenomena. (b–d) Chemical structures of the self-assembling probes D1–D6. Each probe comprises three functional modules: a targeting ligand (R1) for protein recognition, a hydrophobic segment (R2) to modulate the hydrophilic/hydrophobic balance, and a reporting fluorophore (R3). (e) Schematic representation of self-assembling turn-on fluorogenic probes designed for cell-surface protein imaging. (f) Fluorescence images of KB cells treated with probe D3 in the absence (left) or presence (right) of dihydrofolate reductase (DHFR). The addition of DHFR could modulate competitive interactions between the probe and folate receptors. Scale bars: 20 µm. (g) Transmembrane-type hCA imaging of A549 cells using probe D6 without (left) or with (right) ethoxzolamide (EZA), cultured under hypoxic conditions. EZA is a carbonic anhydrase inhibitor. Scale bars: 10 µm. (f) and (g) are adapted from ref. 233 with permission from the American Chemical Society, copyright 2012. | ||
9.1. Photophysical mechanisms of DIE
The underlying mechanism of DIE is closely related to the structural and electronic properties of the fluorophore in aggregation states. In the aggregated state, many planar and π-conjugated fluorophores experience strong intermolecular interactions, such as π–π stacking, intermolecular PET, or CT processes.229,230 These interactions lead to excitonic coupling and enhanced nonradiative decay pathways, all of which contribute to fluorescence quenching.231,232 Furthermore, phenomena such as self-absorption and inner-filter effects in aggregates can lead to the reabsorption of emitted photons, thereby diminishing the overall fluorescence signal.Upon disaggregation, these quenching mechanisms are disrupted. Physical separation of the chromophores reduces π–π interactions, limits intermolecular PET and CT pathways, and eliminates the inner filter effect. As a result, the excited-state energy is more likely to decay radiatively, restoring the fluorescence signal. This switch-like behaviour forms the basis for using DIE as a sensitive reporter mechanism in wash-free imaging, particularly under conditions where target-specific interactions induce probe disassembly.233,234
Structurally, many DIE-active fluorophores are based on planar π-conjugated systems, such as BODIPY,235 fluorescein,15 rhodamines,236 cyanines,237 and squaraines.238 These dyes tend to form aggregates in aqueous or polar environments due to their hydrophobic cores. By careful molecular design—often involving hydrophobic–hydrophilic balance or responsive linkers—these dyes can be tuned to remain aggregated under normal conditions and disaggregate upon interacting with specific biological targets.196 For instance, binding to membrane surfaces,239 proteins,240,241 enzyme,235 or nucleic acids242,243 may induce disaggregation. Alternatively, enzymatic cleavage of a self-assembling moiety can convert non-emissive aggregates into highly fluorescent monomers.
9.2. Applications in bioimaging
DIE probes have been successfully applied in various wash-free bioimaging contexts, leveraging their “turn-on” property to achieve high signal-to-noise ratios.This modular architecture enables precise control over the probe's aggregation behavior and fluorescence output. For instance, probe D3, a fluorescein-labeled construct with a Val–Phe dipeptide tail, forms 10–50 nm aggregates that are non-fluorescent in solution but show a ten-fold fluorescence enhancement upon binding to folate receptors on KB cell membranes.233 Signal reduction upon folate competition confirms both specificity and reversibility (Fig. 35f). Similarly, probe D6 incorporates a Phe–Phe–Leu tripeptide to target transmembrane carbonic anhydrases (CAs) in A549 cells. Under hypoxic conditions, it undergoes disassembly and yields a nine-fold signal increase, which is diminished in the presence of CA inhibitors—demonstrating its utility for imaging-based drug screening (Fig. 35g).
Equally important are the lessons learned from less successful designs, which highlight the delicate balance required in tuning hydrophilicity and hydrophobicity. Probe D1, based on BODIPY with a long alkyl tail, forms large (100–200 nm) aggregates that exhibit excellent turn-on response in vitro but suffer from nonspecific binding and poor cell imaging performance. At the other extreme, probe D2 is overly hydrophilic and fails to aggregate, resulting in constant background fluorescence. Conversely, probe D4 is too hydrophobic to disassemble upon target binding, leading to a permanently “OFF” state. These examples underscore the critical importance of modular fine-tuning, as even minor design deviations can render the probe ineffective.
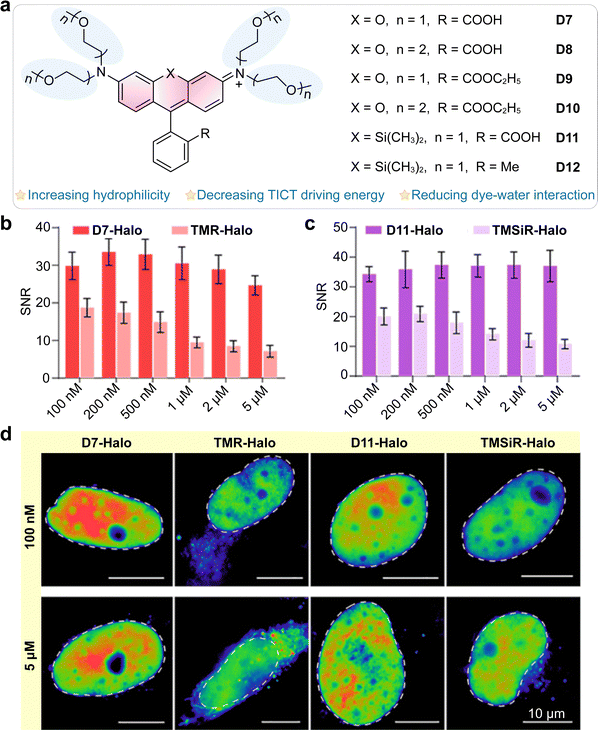 | ||
| Fig. 36 (a) Design and chemical structures of Si-rhodamine dyes D7–D12. (b) and (c) The comparison of SNR between (b) D7-Halo and TMR-Halo, and (c) D11-Halo and TMSiR-Halo, under different probe concentrations. (d) Structured illumination microscopy imaging of nuclei using D7-Halo, TMR-Halo, D11-Halo, and TMSiR-Halo under different probe concentrations. (e) Confocal imaging of nuclei using D7-Halo, TMR-Halo, D11-Halo, and TMSiR-Halo under different probe concentrations. (b)–(d) are adapted from ref. 171 with permission from the American Chemical Society, copyright 2025. | ||
To mitigate these issues, the researchers replaced conventional N-alkyl substituents with hydrophilic ether chains (Fig. 36a). This ether modification significantly improves water solubility and reduces aggregate size. The resulting smaller aggregates are more stable and remain non-fluorescent until they encounter their specific molecular targets.
In live-cell imaging, ether-modified probes such as D7-Halo display minimal intracellular background even at 5 µM. In contrast, traditional probes like TMR-Halo show pronounced off-target staining at concentrations as low as 100 nM (Fig. 36d, especially in organelles such as the nucleus, lysosomes, and mitochondria).
By lowering nonspecific activation and enabling the use of higher probe concentrations, ether modification enhances the SNR of rhodamines/Si-rhodmines by 1.7–3.4× (Fig. 36b and c). This improvement facilitates wash-free labeling and supports long-term super-resolution imaging techniques, such as SIM and SMLM. It enables the visualization of dynamic processes, including lysosomal oscillations and actin remodeling, with high fidelity and minimal background interference.
The BF2-bridged cyanine (BCy) platform (Fig. 37a) represents a significant advance in wash-free fluorescent probe design by leveraging DIE.245 These dyes form non-emissive aggregates in water but “turn on” upon disaggregation in hydrophobic environments, enabling high-contrast imaging without washing.
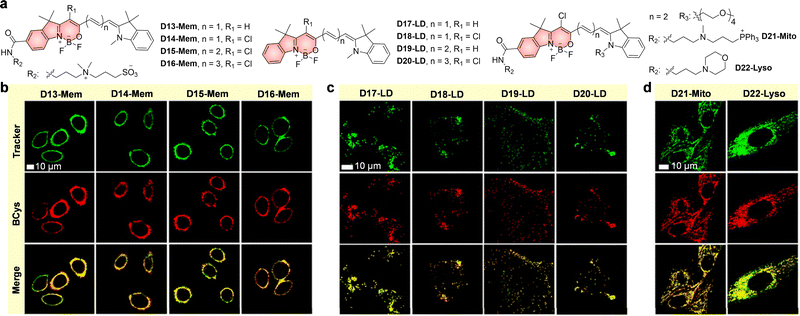 | ||
| Fig. 37 (a) Chemical structures of BF2-bridged cyanine probes. (b) Plasma membrane imaging of A549 cells treated with D13-Mem (red), D14-Mem (red), D15-Mem (red), D16-Mem (red), and CellMask Green (green). (c) Lipid droplet imaging of A549 cells stained with D17-LD (red), D18-LD (red), D19-LD (red), D20-LD (red) probes and BODIPY493/503 (green). (d) Colocalization of D21-Mito (red) with MitoTracker Green (green), and D22-Lyso (red) with LysoTracker Green (green) in live A549 cell imaging. Scale bars: 10 µm. (b)–(d) are adapted from ref. 245 with permission from the American Chemical Society, copyright 2025. | ||
This aggregation-dependent fluorogenic behavior is exemplified by MemBCy probes (D13-Mem–D16-Mem), which target the plasma membrane (Fig. 37b). Their zwitterionic sulfobetaine groups guide dyes to insert into the lipid bilayer, triggering disaggregation and bright fluorescence. MemBCy stains rapidly (1–30 minutes), remains membrane-bound for up to 6 hours, and works at nanomolar concentrations. Their low permeability also makes them useful for monitoring membrane integrity during cell death.
Similarly, targeting lipid droplets was achieved with LD probes (D17-LD–D20-LD) by exploiting the scaffold's hydrophobicity. These probes are non-fluorescent in water but become highly emissive in the lipid-rich droplet interior (Fig. 37c). This turn-on response yields excellent signal-to-background contrast, suitable for wash-free imaging. LDBCy dyes also show superior photostability and compatibility with STED microscopy, enabling sub-diffraction imaging and dual-color tracking of organelle interactions.
Using the same strategy, Mao and co-workers extended the BCy platform to develop wash-free fluorogenic dyes (D21-Mito, D22-Lyso) for labeling mitochondria and lysosomes (Fig. 37d). By attaching appropriate organelle-targeting moieties to the BCy scaffold, they created probes that retain environment-responsive fluorescence activation, enabling specific, high-contrast imaging without the need for washing.
Klymchenko et al. developed a fluorogenic probe (D23; Fig. 38b) composed of two cyanine units linked by flexible cadaverine or lysine spacers and equipped with low-affinity membrane anchors (Fig. 38a).237 In aqueous solution, the two cyanine units form intramolecular H-aggregates, inducing strong self-quenching, while membrane binding disrupts aggregation and restores fluorescence (Fig. 38a). This reversible binding enables spontaneous ON/OFF switching without washing steps. The dimeric design provides >2-fold higher single-molecule brightness and ∼10-fold slower diffusion compared to monomeric dyes. These properties minimize motion blur and enhance localization precision for PAINT-based super-resolution imaging of live-cell membranes, enabling nanoscale resolution and dynamic 3D visualization of membrane structures (Fig. 38c–g). Notably, this intramolecular “dimeric” arrangement forms the minimal structural motif required to achieve DIE.
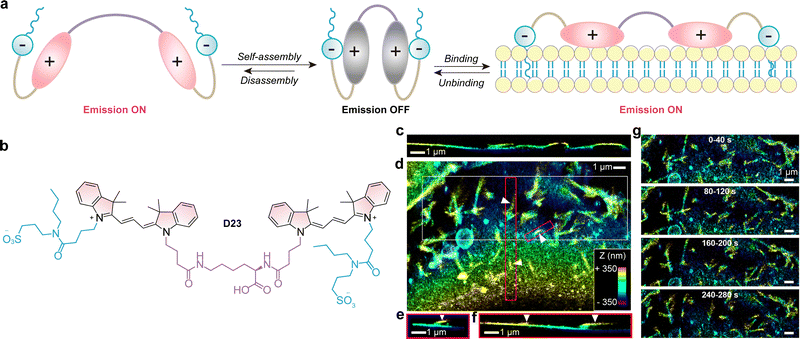 | ||
| Fig. 38 (a) Concept of switchable cyanine-based dimeric probe that lights up after reversible binding to biomembranes. (b) Chemical structure of D23. (c)–(g) 3D-PAINT imaging with D23 reveals rich 3D plasma membrane structures and dynamics in living cells. The sample was illuminated with 560 nm light (∼1 kW cm−2) in wide-field mode and recorded at 110 frames s−1 with a 9 ms exposure time. (c) Vertical cross-sectional view of 3D-PAINT-reconstructed plasma membrane in a COS-7 cell with undulating membrane heights. (d) In-plane (top-down) view of a different cell, showing numerous nanoscale tubules. (e) and (f) Vertical cross-sectional views taken along the (e) magenta and (f) red boxes in (d), revealing tubules extending away from the cell surface (arrowheads). For (c), (e) and (f), the vertical dimension denotes depth into the sample. (g) Time-resolved 3D-PAINT data acquired in 40 s intervals at four time points from the white box region in (d). For all images, color presents depth (Z) according to the color scale in (d). (c)–(g) are adapted from ref. 237 with permission from the American Chemical Society, copyright 2022. | ||
 | ||
| Fig. 39 Schematic illustration showing the fluorogenic turn-on response of D24 upon CuAAC reaction with azido DNA. | ||
Although this strategy is powerful, its reliance on a copper catalyst limits its use to fixed samples due to cytotoxicity. Future development of catalyst-free bioorthogonal triggers, such as strain-promoted azide–alkyne cycloaddition (SPAAC) or vinyl–tetrazine ligation, would extend DIE probes to live-cell and in vivo imaging, allowing real-time visualization of dynamic biological processes without washing steps.
9.3. Advantages and design versatility
A fundamental advantage of DIE is its ability to leverage the vast library of existing planar fluorophores. Unlike AIE, which often requires the synthesis of specific rotor-based scaffolds (like TPE), DIE allows researchers to repurpose classic, high-quantum-yield dyes (e.g., fluorescein, cyanines, BODIPY) into “turn-on” probes simply by engineering their aggregation states. This grants immediate access to diverse spectral properties without requiring de novo chromophore design.DIE systems are uniquely capable of achieving an extremely dark “OFF” state due to the high efficiency of intermolecular quenching (e.g., H-aggregation) within the supramolecular assembly. This property contributes a negligible background signal, enabling wash-free imaging with exceptional contrast.
Furthermore, unlike enzyme-activated probes that rely on irreversible covalent cleavage, DIE fluorophores activate through non-destructive supramolecular disassembly. This mechanism is often faster and can preserve the chemical integrity of the fluorophore. It offers a robust alternative for real-time sensing and, in some cases, enables the design of reversible probes capable of monitoring dynamic biological fluctuations.
Additionally, the modular nature of these assemblies facilitates integration with functional ligands. As demonstrated by Hamachi's amphiphilic strategy (target ligand + hydrophobic tail + reporter), the recognition moiety and the signaling unit can be independently optimized. This architecture enables tunable affinity and kinetics, allowing for no-wash cell-surface imaging and drug screening. Similarly, BF2-bridged cyanines (BCy) exploit specific hydrophobic interactions to label plasma membranes, lipid droplets, and mitochondria with nanomolar concentrations, demonstrating that DIE can achieve organelle specificity purely through physicochemical tuning.
9.4. Limitations and practical challenges
The biological deployment of DIE-based probes is fundamentally constrained by a precarious thermodynamic stability trade-off. A primary bottleneck is the restrictive “disaggregation window”: if the aggregate is too stable (too hydrophobic), it fails to disassemble upon target binding (permanently OFF). Conversely, if it is too unstable (too hydrophilic), it fails to aggregate initially (permanently ON/high background). Achieving this precise balance typically requires extensive empirical screening of alkyl tail lengths or peptide sequences, as predictive computational models for these large assemblies are still lacking.Furthermore, the disassembly kinetics of DIE probes represent a significant temporal bottleneck. Unlike small-molecule binding, which is typically diffusion-limited, the disassembly of a supramolecular aggregate is a complex, multi-step process that may be kinetically slow, potentially limiting the temporal resolution for tracking dynamic biological events.
Additionally, the reliance on physical disassembly also introduces the risk of “false positives”: non-specific disaggregation can occur if the probe encounters unrelated surfactants, lipid membranes, or hydrophobic pockets on serum albumin. Quantification presents another hurdle. Because brightness scales with the local concentration of monomers released after disassembly, differences in critical aggregation concentration, aggregate size, and dispersity can complicate the absolute quantification of analytes across different cell types or time points.
From a biological perspective, introducing aggregates into cells carries inherent risks. Large aggregates (>100 nm) are prone to endocytosis and lysosomal accumulation, which generates background noise distinct from the specific target. Moreover, the hydrophobic tails or cationic surfaces used to drive self-assembly can perturb membrane integrity (surfactant effects). Therefore, rigorous cytotoxicity and electrophysiology control remain critical challenges, especially for long-term imaging or when using high local probe concentrations.
9.5. Outlook and future directions
A critical frontier for DIE is the development of catalyst-free bioorthogonal probes. Transitioning to strain-promoted azide–alkyne cycloaddition (SPAAC) or tetrazine ligation to trigger disassembly would enable live-cell applications by eliminating the cytotoxic copper catalysts required by current generations.To enhance specificity, future designs should pursue “Dual-Gated” DIE systems. By combining disassembly with a specific chemical event (e.g., enzyme cleavage plus bioorthogonal ligation), fluorescence activation can be governed by the coincidence of multiple triggers, thereby minimizing off-target disaggregation. This logic parallels the “AND-gate” strategies successfully employed in AIE.218
Simultaneously, there is a need for rigorous “aggregate engineering” via smart polymers. A promising avenue is to encapsulate DIE probes within stimuli-responsive “smart” coatings. These shells sterically stabilize the quenched aggregates during circulation, preventing a non-specific background. The coating is engineered to degrade only in response to a specific microenvironmental trigger (e.g., tumor acidity or intracellular enzymes), thereby facilitating controlled dye release and disaggregation.247 Coupling this “responsive gating” with active targeting ligands creates a robust, high-fidelity platform. This platform integrates the selectivity of biological recognition with the high contrast offered by the DIE mechanism.
Finally, the integration of DIE with theranostics represents a high-value direction. Since many DIE-active dyes (like BODIPY and cyanines) are potent photosensitizers, the aggregated “OFF” state effectively suppresses phototoxicity, while the disaggregated “ON” state restores both fluorescence and ROS generation.248,249 This “activatable photodynamic therapy” (aPDT) paradigm offers a route to precise tumor treatment with minimal off-target side effects.
10. In situ formation of fluorophores
While the preceding mechanisms rely on modulating the emissive state of a pre-existing fluorophore, wash-free imaging can also be achieved by constructing the chromophore itself within the biological environment. This approach, known as in situ formation of fluorophores, relies on chemical or metabolic triggers to generate the reporter from silent precursors. This approach can also enable simultaneous therapeutic interventions, such as photodynamic therapy (PDT).In this section, we highlight several representative classes of in situ-generated fluorophores. Some examples naturally overlap with mechanisms discussed earlier, reflecting the structural diversity of their precursors. Nevertheless, we include them here for completeness. In many cases, the underlying photophysical processes remain insufficiently characterized. This limitation motivates classifying these systems under in situ formation rather than within sections focused on defined photophysical mechanisms.
10.1. Representative examples and applications
A well-established example is the metabolic conversion of 5-aminolevulinic acid (ALA) into protoporphyrin IX (PpIX, ISF-1; Fig. 40a).250 ALA is a natural intermediate in the heme biosynthetic pathway. When supplied exogenously, it bypasses the rate-limiting enzyme ALA synthase (ALAS), thereby accelerating the downstream formation of porphyrin intermediates (Fig. 40a). Crucially, this process exploits a metabolic imbalance often present in malignant cells. Many tumor types display elevated activity of enzymes that generate PpIX but reduced ferrochelatase activity, which normally inserts ferrous iron to produce non-fluorescent heme (Fig. 40a). As a result, tumor cells selectively accumulate photoactive PpIX, whereas healthy cells efficiently convert it to heme and avoid phototoxic buildup.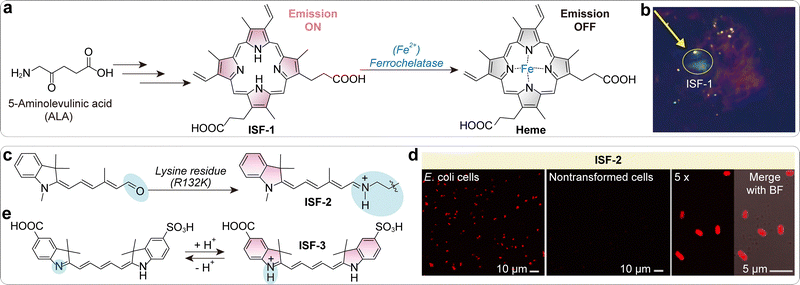 | ||
| Fig. 40 (a) Formation of emissive protoporphyrin IX (PpIX, ISF-1), and its turn-off response to ferrochelatase. (b) Quantitative fluorescence imaging reveals residual ISF-1 in the surgical field near the end of glioblastoma resection. The imaging map indicates significant ISF-1 concentration at the center (green area within the yellow circle) of the field of view, corroborated by histopathology, which classifies the tissue as a tumor. (c) Formation of emissive ISF-2 through lysine residue reaction upon protein binding. (d) Fluorescence imaging of E. coli cells following incubation with the precursor of ISF-2. Cells expressing KLE:R59W:L28W display pronounced fluorescence under 594 nm excitation, with emission collected using a 615 nm long-pass filter (scale bars: 10 µm), whereas non-transformed control cells exhibit no detectable signal under identical conditions. A magnified fluorescence image of the KLE:R59W:L28W/probe complex (5-fold enlargement; scale bar: 5 µm) is presented alongside a brightfield image overlaid with the corresponding fluorescence distribution. (e) Formation of the emissive ISF-3via protonation under the acidic environment. (b) and (d) are adapted from ref. 252 and 255 with permission from Springer Nature and the American Chemical Society, copyright 2012 and 2015, respectively. | ||
The resulting PpIX exhibits dual photophysical functions central to its biomedical applications. As a red-emissive fluorophore (λem ≈ 635 nm), it provides high-contrast visualization in fluorescence-guided surgery (FGS, Fig. 40b).251,252 Under blue-light excitation, PpIX-enriched tumor margins emit bright red fluorescence, allowing real-time delineation during resection. A landmark phase III clinical trial in malignant glioma established ALA-FGS as a standard of care by demonstrating significant improvement in both resection completeness and progression-free survival.253
Beyond imaging, PpIX also acts as a potent photosensitizer for PDT.254 Upon excitation, it produces reactive oxygen species, primarily singlet oxygen, inducing mitochondria-mediated apoptosis and localized necrosis. This metabolic “Trojan Horse” concept, where a benign biosynthetic precursor is converted into a tumor-specific fluorescent and therapeutic agent, illustrates the translational power of molecular design guided by endogenous metabolism.
Yapici and co-workers developed a hybrid “turn-on” imaging strategy that couples a small molecule precursor with a genetically encoded protein scaffold to generate a fluorescent cyanine dye (ISF-2; Fig. 40c) in situ.255 The design utilizes a re-engineered cellular retinoic acid binding protein II (CRABPII) to host a non-fluorescent merocyanine aldehyde. The activation mechanism is driven by the covalent formation of a protonated Schiff base (PSB) and subsequent structural rigidification. Upon binding, the precursor reacts with an engineered lysine residue (R132K) to form a Schiff base, which is protonated to yield a resonating iminium cation. This transformation results in a bathochromic shift and the generation of fluorescence. Furthermore, the protein cavity restricts the torsional motion of the dye, thereby suppressing non-radiative decay and enhancing the quantum yield to 39%. In terms of biological applications, this system facilitated the rapid, wash-free imaging of E. coli (Fig. 40d), allowing for the visualization of basal expression levels within one minute of probe addition with negligible background.
Similarly, many cyanine-based probes utilize the in situ formation of fluorescent species through pH-dependent protonation of a neutral precursor. In 2002, Cooper et al. introduced CypHer5, a functionalized pentamethine cyanine designed to monitor receptor internalization via pH-dependent activation.256 The probe is non-fluorescent at physiological pH (7.4) because the indolenine nitrogen is deprotonated; under these conditions, it absorbs near ∼500 nm. Upon internalization into acidic endosomes (pH < 6), protonation restores conjugation and switches the dye on (Fig. 40e, ISF-3, λex ∼ 645 nm, λem ∼ 665 nm). CypHer5 has a reported pKa of 6.1 and is effective for high-throughput screening of G-protein-coupled receptor internalization. Building on this concept, Grover and colleagues developed a genetically encoded pH sensor to track surface proteins during endocytosis.257
More recently, researchers have integrated pH sensing with specific chemical triggers to enhance selectivity. Schnermann and co-workers developed the CyBam (cyanine carbamate) platform, in which a caged heptamethine norcyanine is activated through a two-step process: cleavage of a carbamate linker by a trigger (e.g., GGT or ROS) followed by protonation of the released norcyanine (pKa ∼ 5.2) to yield a bright NIR fluorophore (ISF-4 and ISF-5; Fig. 41a).258,259 CyBam probes showed ∼170-fold fluorescence enhancement (Fig. 41b) and enabled imaging of GGT activity (Fig. 41c–e) in ovarian cancer and ROS in prostate cancer cells.
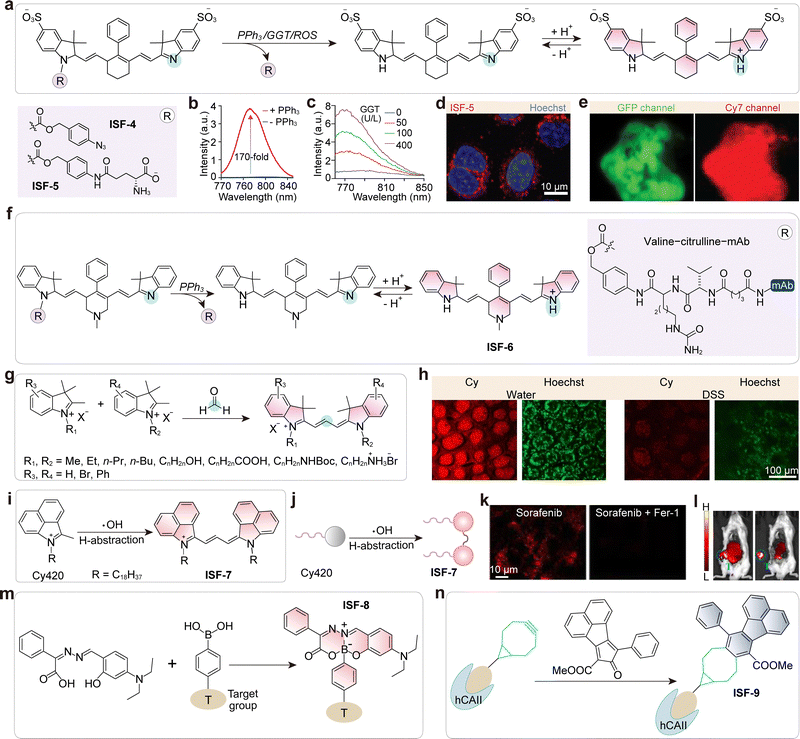 | ||
| Fig. 41 (a) Molecular design and turn-on mechanism of CyBam dyes (ISF-4, ISF-5). The activation process involves two steps: (i) cleavage of the carbamate linker and (ii) indolenine protonation. (b) Fluorescence changes of ISF-4 upon reaction with PPh3. (c) Increase in fluorescence intensity of ISF-5 after incubation with increasing concentrations of γ-glutamyl transpeptidase (GGT). (d) Confocal images showing activation of ISF-5 in SHIN-3 cells. Fluorescent signals from the probe (ISF-5, red) and nucleus (Hoechst, blue) are displayed. (e) In vivo imaging of SHIN-3-ZsGreen metastatic tumors using ISF-5. Green and red pseudocolors represent signals from GFP and Cy7 channels, respectively. (f) Schematic illustration for the generation of emissive ISF-6 from CyLBam-based antibody–drug conjugate. The inset shows the cathepsin-cleavable dipeptide (valine–citrulline) linker conjugated to a monoclonal antibody (mAb). (g) Synthetic scheme for Cy3 synthesis in the presence of formaldehyde. (h) Fluorescence imaging of small intestine tissues from healthy and IBD mice stained with Cy precursor (red) and Hoechst (green). Dextran sulfate sodium (DSS) induces colitis, a model of inflammatory bowel disease (IBD), characterized by intestinal inflammation and epithelial barrier damage. (i and j) Schematic illustration of hydroxyl radical-triggered NIR cyanine ISF-7 generation from monomer Cy420. (k) Confocal fluorescence imaging of Huh7 cells with Cy420 after ferroptosis-induced ˙OH generation, with and without the ferroptosis inhibitor Ferrostatin-1 (Fer-1). (l) Anatomical fluorescence imaging of tumor-bearing mice at 48 h post-sorafenib treatment and Cy420 injection. (m) Design strategy for constructing organelle-specific boronic acid-derived salicylidenehydrazone labeling platform ISF-8. (n) Schematic representation of selective labeling of hCAII using the bioorthogonally activated probe ISF-9. (b)–(e), (h), (k) and (l) are adapted from ref. 259–261 with permission from the American Chemical Society, Wiley-VCH and Wiley-VCH, copyright 2021, 2025 and 2026, respectively. | ||
Building on this concept, Usama et al. introduced CyLBams (cyanine lysosome-targeting carbamates) as fluorogenic mimics of antibody–drug conjugates.259 A cathepsin B–cleavable carbamate releases N-Me-NorCy7 (ISF-6, Fig. 41f), whose tertiary amine (pKa ≈ 4.6) promotes protonation and lysosomal retention. This design achieved ∼187-fold higher cellular uptake and tumor-to-background ratios of 4–5, allowing imaging of EGFR+ and CD276+ breast cancer models and highlighting the potential of cathepsin-cleavable linkers for targeted tumor imaging.
Chang and co-workers developed an innovative in situ fluorescence activation strategy for intracellular formaldehyde sensing.260 This approach leverages formaldehyde-mediated condensation to generate fluorescent cyanine dyes directly within living cells (Fig. 41g), eliminating the need for external probe delivery. This method offers a highly efficient and sustainable approach for real-time intracellular imaging under physiological conditions (Fig. 41h), minimizing background interference and expanding tools for dynamic cellular monitoring.
Qu et al. introduced a zero-background imaging strategy utilizing a non-emissive precursor, Cy420. Cy420 self-assembles into nanoparticles that selectively accumulate in tumors via the enhanced permeability and retention (EPR) effect.261 The imaging activation is triggered by sorafenib-induced ferroptosis, where generated hydroxyl radicals catalyze the in situ dimerization of Cy420 into ISF-7 (Fig. 41i and j), a potent NIR fluorophore. This tumor-specific activation effectively eliminates background noise (Fig. 41k), enabling high-contrast imaging for precise tumor localization and surgical guidance (Fig. 41l).
Zhang and co-workers developed intrinsically wash-free fluorescent probes based on boronic acid-derived salicylidenehydrazones (ISF-8, Fig. 41m).262 Their modular architecture comprised a salicylidenehydrazone ligand, a phenylboronic acid moiety, and an organelle-targeting group—morpholine for lysosomes or triphenylphosphonium for mitochondria.
These fluorophores exhibit pronounced environmental sensitivity: they are weakly emissive in polar and protic solvents, likely due to TICT and hydrogen-bonding interactions. However, upon partitioning into hydrophobic or aprotic microenvironments such as organelle membranes, they undergo a dramatic fluorescence “turn-on,” increasing from nearly 0% in aqueous buffer to as high as 68% in hydrophobic solvents (e.g., Lyso-BS in chloroform). The probes are excited in the blue-green region (λex ≈ 470 nm) and emit in the green channel (λem ≈ 520–550 nm).
Biological validation demonstrated that ISF-8-Lyso and ISF-8-Mito enable high-fidelity, wash-free staining of lysosomes (Pearson's coefficient = 0.93) and mitochondria (Pearson's coefficient = 0.91) in HeLa cells after only one minute of incubation.
Ji and co-workers introduced a “click and fluoresce” strategy that enables the bioorthogonal generation of a fluorophore in situ.263 This system pairs a biomolecule-tagged strained alkyne (endo-BCN) with a non-fluorescent precursor based on a cyclopentadienone scaffold fused to a naphthalene group (Fig. 41n). A cascade reaction drives the fluorescence turn-on: the non-emissive cyclopentadienone undergoes an iEDDA cycloaddition with the alkyne, followed by cheletropic elimination of carbon monoxide (CO).
This irreversible transformation converts the silent reactants into a stable, highly fluorescent naphthalene-fused benzene derivative (ISF-9, Fig. 41n). The product emits in the blue region (λem ≈ 461–465 nm) with quantum yields of 0.13–0.17. Because the starting material is optically silent, the system offers near-zero background fluorescence. This capability was demonstrated through wash-free imaging of membrane lipids (via DOPE–BCN conjugates) and human carbonic anhydrase II (hCAII) in live HeLa cells, underscoring its potential for high-throughput screening applications.
Wang and co-workers introduced a bioorthogonal synthesis strategy that constructs the fluorophore scaffold directly within the biological system (Fig. 42a).264 This design relies on the chemoselective condensation between electron-deficient β-chloroacroleins (β-CAs) and meta-aminothiophenol (m-AT). The “turn-on” mechanism is driven by the de novo formation of the chromophore: the precursors are optically silent, but their reaction generates a benzothiazole-fused polycyclic system. This structural transformation establishes an extended π-conjugation network, resulting in intense fluorescence (i.e., ISF-10 and ISF-11; Fig. 42a).
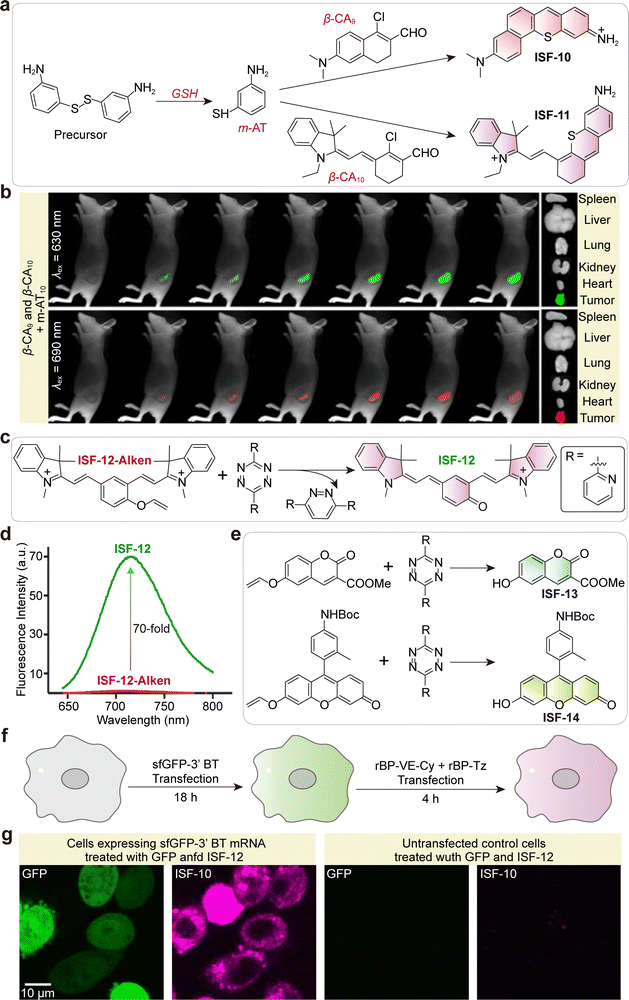 | ||
| Fig. 42 (a) Mechanisms of the GSH-activated bioorthogonal reactions for the imaging of cancer cells and tumors in vivo. (b) In vivo NIR fluorescence imaging of HepG2 tumor-bearing mice following GSH-activated bioorthogonal reactions. Mice were intratumorally injected with a mixed solution of β-CA9 and β-CA10, followed by the tail vein injection of the precursor and real-time imaging over a 6 h period. (c) Bioorthogonal reaction between vinyl ether-caged cyanine and tetrazine derivative, yielding the uncaged fluorophore ISF-12. (d) Fluorescence comparison and turn-on ratio between caged ISF-12-Alken and uncaged ISF-12. (e) Representative vinyl ether-caged molecules and their emissive products upon tetrazine-mediated uncaging. (f) Schematic representation of the transfection and imaging workflow. CHO cells were first transfected with the sfGFP-3′ BT plasmid encoding the target sequence, followed by treatment with rBP-VE-Cy and rBP-Tz. (g) Confocal fluorescence images showing cytoplasmic localization of the activated NIR signal in cells expressing the target mRNA (top) compared to controls lacking either the complementary sequence or tetrazine partner (bottom). Scale bars: 10 µm. (b), (d) and (g) are adapted from ref. 264 and 265 with permission from the American Chemical Society, copyright 2025 and 2016, respectively. | ||
A key advantage of this approach is the tunability of the emission wavelengths, which range from 627 nm to 778 nm depending on the conjugation length of the β-CA precursor. Because the starting materials exhibit negligible background fluorescence, the specific generation of the product yields exceptional signal-to-noise ratios—most notably, the β-CA9 variant achieved a massive 22![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 575-fold fluorescence enhancement. This high-contrast performance facilitated simultaneous, wash-free two-color imaging of lysosomes and mitochondria in HepG2 cells. Furthermore, the authors demonstrated the therapeutic relevance of the system by designing a glutathione (GSH)-responsive precursor. This precursor enabled logic-gated, tumor-specific imaging in vivo by exploiting the elevated GSH levels characteristic of the cancer microenvironment (Fig. 42b).
575-fold fluorescence enhancement. This high-contrast performance facilitated simultaneous, wash-free two-color imaging of lysosomes and mitochondria in HepG2 cells. Furthermore, the authors demonstrated the therapeutic relevance of the system by designing a glutathione (GSH)-responsive precursor. This precursor enabled logic-gated, tumor-specific imaging in vivo by exploiting the elevated GSH levels characteristic of the cancer microenvironment (Fig. 42b).
A more straightforward approach to in situ fluorophore generation involves photochemical or chemical “uncaging” reactions, in which a non-fluorescent precursor is converted into a fluorescent species upon activation. Wu and colleagues developed a bioorthogonal approach for generating cyanine dyes, enabling high-contrast, wash-free imaging of mRNA.265 Their design leverages a nucleic acid-templated reaction, in which two antisense probes hybridize to adjacent sequences on the target mRNA: one bearing a vinyl ether-caged cyanine fluorophore and the other a tetrazine moiety. In the latent “OFF” state, the vinyl ether masks the phenoxide electron donor of the cyanine scaffold, disrupting π-conjugation and quenching fluorescence. Upon hybridization, the proximity of the probes induces an iEDDA reaction, followed by elimination, which liberates the phenoxide and restores π-conjugation, switching the dye to its emissive state (Fig. 42c and e). The resulting near-infrared probe (ISF-12, λem ≈ 710 nm; Fig. 42c) exhibited over 70-fold fluorescence enhancement (Fig. 42d) in DNA-templated assays, underscoring the robustness of this iEDDA-mediated uncaging strategy for live-cell applications. Because fluorescence is activated only upon specific target binding and reaction (Fig. 42f), this method enables direct visualization of intracellular mRNA (Fig. 42g) without washing steps to remove unbound probes.
In 2025, Shen and co-workers addressed the challenge of visualizing cell viability and photodamage with “Rhodalive”, a zero-background rhodamine probe explicitly activated by the mitochondrial electron transport chain (ETC; Fig. 43a).266 Unlike probes that rely on hydrolytic enzymes or simple pH changes, Rhodalive operates through a unique reduction–oxidation sequence. In the latent “OFF” state, the fluorescence of the rhodamine scaffold is quenched by a 4-nitrobenzyl methanoate moiety attached at the C9 position, which disrupts the strictly conjugated system.
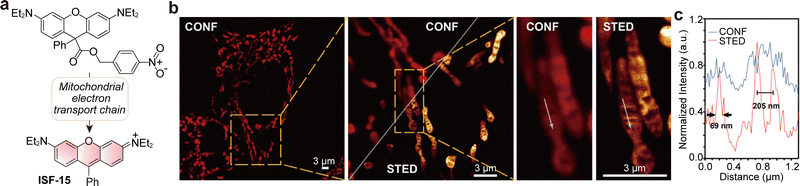 | ||
| Fig. 43 (a) Design rationale of a reduction-activated turn-on fluorescent probe (ISF-15) for monitoring mitochondrial ETC activity. (b) Confocal and STED imaging of live HeLa cells via the formation of ISF-15. (c) Intensity profile along the white arrow in panel (b). Scale bars: 3 µm. (b) and (c) are adapted from ref. 266 with permission from the American Chemical Society, copyright 2025. | ||
The “turn-on” mechanism is initiated by a single-electron reduction via ETC leakage, which converts the nitro group into a radical anion. This triggers a single-electron transfer (SET) cascade, resulting in the release of CO2 and the formation of a triphenylmethyl radical intermediate. Uniquely, this intermediate is subsequently oxidized by molecular oxygen to yield the highly fluorescent rhodamine species, ISF-15 (Fig. 43a, λem = 580 nm). This specific requirement for active electron transport allows the probe to distinguish live cells from fixed or dead cells with high fidelity. The probe's robust signal-to-noise ratio enabled wash-free stimulated emission depletion (STED) microscopy of mitochondrial cristae with ∼69 nm resolution (Fig. 43b and c), as well as the real-time visualization of single-cell photodamage induced by photodynamic therapy.
10.2. Advantages and design versatility
In situ fluorophore generation provides an elegant alternative to conventional quenching-based “turn-on” mechanisms, as it creates the emissive species directly within the biological milieu. This intrinsic activation process, which transforms optically silent precursors into fluorescent products, confers several key advantages for wash-free bioimaging.The primary benefit of this strategy is an exceptional signal-to-background ratio. Because the precursors are non-emissive, fluorescence only appears after a specific chemical or enzymatic reaction, virtually eliminating background signals from unreacted probes. This “zero-background” approach enables massive signal enhancements, with systems such as bioorthogonal “click-and-fluoresce” or β-chloroacrolein condensations achieving turn-on ratios of 100-fold to 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000-fold. This high contrast is ideal for the real-time visualization of dynamic processes, such as mRNA hybridization or receptor internalization, as it enables continuous monitoring without the need for washing steps that would otherwise interrupt the experiment.
000-fold. This high contrast is ideal for the real-time visualization of dynamic processes, such as mRNA hybridization or receptor internalization, as it enables continuous monitoring without the need for washing steps that would otherwise interrupt the experiment.
Furthermore, this method achieves high target specificity by leveraging endogenous pathways. The activation mechanism itself, whether driven by metabolism (e.g., ALA to PpIX), specific enzyme activity (e.g., cathepsins, GGT), or unique physiological states (e.g., acidic pH, ETC redox leakage), co-encodes biological function into the fluorescent readout.
This functional specificity can also be harnessed for theranostic applications. The ALA–PpIX system, for example, is combined with fluorescence-guided surgery and photodynamic therapy, enabling simultaneous diagnosis and treatment through the same endogenously generated fluorophore.
10.3. Limitations and practical challenges
Despite these benefits, several practical and conceptual challenges remain.A primary bottleneck is the dependency on endogenous metabolic machinery (e.g., in ALA–PpIX system), which introduces significant inter-patient and intra-tissue variability. Because enzyme expression levels vary across biological contexts, the resulting signal becomes a convoluted function of metabolic capacity rather than purely analyte abundance, limiting the diagnostic reliability.
The challenge of precursor design is significant, as achieving high reactivity while maintaining stability and biocompatibility is nontrivial: high-reactivity precursors, such as aldehydes or carbamates, are frequently susceptible to spontaneous hydrolysis or non-specific side reactions within the complex cellular milieu, leading to diminished biocompatibility and off-target background.
From a photophysical standpoint, in situ formation often limits the tunability of the resulting optical properties. Unlike modular quenching-based designs, chemical reactions typically fix the chromophore structure, limiting emission color control and multiplexing potential.
Additionally, delivering multiple components and ensuring they co-localize at the same subcellular site remains a significant challenge, as differential uptake and diffusion rates can lead to spatially decoupled reaction pairs. Finally, the irreversible nature of fluorophore generation complicates quantitative interpretation. Because fluorophore generation is often irreversible and depends on local reaction rates, the resulting fluorescence intensity reflects both probe distribution and chemical conversion efficiency, which complicates direct correlation with a biological state. This lack of “dynamic reversibility” represents a major barrier to high-fidelity quantification, as the signal often reflects the cumulative history of the reaction rather than the real-time concentration of the target analyte.
10.4. Outlook and future directions
Future development of in situ fluorophore generation should focus on several key areas to overcome current limitations.Priority should be placed on the continued optimization of reaction chemistry to design faster, more orthogonal, and highly selective reactions that operate efficiently at low physiological concentrations. This includes the development of modular reaction platforms that enable facile spectral tuning, particularly in the near infrared, without compromising reactivity.
The next level of precision will be achieved by integrating chemical activation with genetic or metabolic engineering. This will enable cell-type-specific fluorogenesis and the design of dual-trigger “AND-gate” probes (e.g., requiring both an enzyme and a specific pH) to mitigate off-target activation.
Finally, unlocking the quantitative potential of this strategy requires standardized kinetic models to separate signal evolution from reaction conversion, which enables more accurate correlation of the fluorescent signal with the underlying biological state.
In summary, in situ fluorophore generation transforms the microenvironment into a chemical reactor, providing unrivaled specificity and minimal background for wash-free imaging. The next frontier lies in expanding the reaction toolbox, improving control over spatial and spectral properties, and coupling fluorogenesis with biological function to create truly responsive, multi-functional imaging systems.
11. Other mechanisms
The de novo generation of fluorophores offers exceptional contrast by coupling imaging to metabolism. Beyond these frameworks, several emerging quenching mechanisms, such as twisted intramolecular charge shuttle (TICS) and access to conical intersections (CI), provide additional opportunities for precise fluorogenic control.11.1. Twisted intramolecular charge shuttle
Wash-free bioimaging can also be achieved by suppressing the TICS mechanism, a distinct quenching pathway from the canonical TICT mechanism (Fig. 44a).267 Unlike TICT, where charge transfer is unidirectional, TICS involves a dynamic, bidirectional charge exchange coupled with intramolecular rotation. In this model, a dialkylamine or aniline moiety acts initially as an electron-withdrawing group but switches to an electron donor upon a ∼90° twist, generating a non-emissive “dark” state with nearly zero oscillator strength. Fluorescence is restored when this rotational motion is physically restricted (e.g., in viscous media) or chemically blocked (e.g., by reacting with thiols such as glutathione).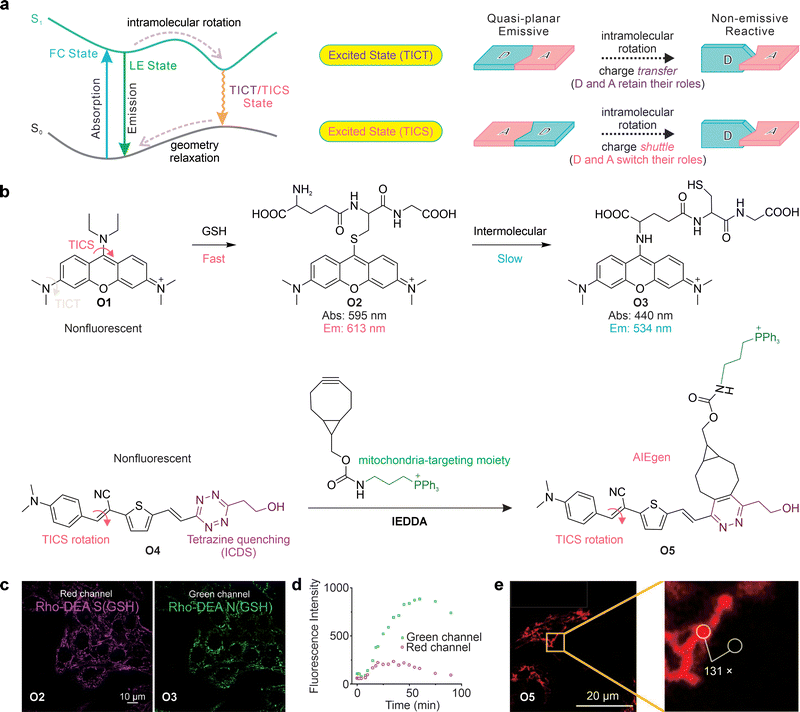 | ||
| Fig. 44 (a) Conceptual comparison of TICT and TICS. (b) Chemical structures and key reactions underlying TICS-based fluorescent probes. (c) Dual-color confocal images of HeLa cells incubated with 0.5 µM probe O1 for 4 min. (d) Corresponding fluorescence intensity changes over time in the red (O2) and green (O3) channels. (e) Wash-free mitochondrial imaging enabled by the in situ formation of AIEgen O5 from probe O4, with the right panel showing fluorescence intensity comparisons inside and outside the organelle regions of interest. (c), (d) and (e) are adapted from ref. 268 and 269 with permission from Elsevier and Wiley-VCH, copyright 2022 and 2025, respectively. | ||
Based on this concept, Xu and co-workers designed Rho-DEA, a pyronine-based TICS probe capable of selectively detecting glutathione (GSH) over similar biothiols (O1; Fig. 44b–d).268 In its native form, Rho-DEA is nearly non-fluorescent (φf <0.01) due to efficient TICS quenching. Reaction with GSH proceeds through sequential nucleophilic substitutions that disrupt the TICS pathway and generate two fluorescent products: a red-emitting intermediate (Rho-S or O2; λem ∼ 613 nm) and a green-emitting final product (Rho-N or O3; λem ∼ 534 nm). This red-to-green temporal shift yields a radiometric signature specific to GSH, enabling dynamic tracking of mitochondrial thiol homeostasis in live HeLa cells.
Wu and co-workers further advanced the concept by integrating TICS with a dual-lock bioorthogonal platform combining tetrazine chemistry and AIE (O4; Fig. 44b and e).269 The probe contains two sequential activation barriers: (Lock 1) a tetrazine quencher removed via reaction with strained alkynes, and (Lock 2) a TICS-active molecular rotor that remains dark until aggregation restricts its motion. Only upon target accumulation and aggregation is the TICS pathway suppressed, producing intense fluorescence (from O5). This design yielded sigmoidal response behavior, an exceptionally high turn-on ratio (up to 1033-fold), and a significant Stokes shift (∼200 nm). The ultra-low background enabled wash-free, multiplexed imaging of cellular organelles and precise detection of ischemic injury via αvβ3 integrin upregulation, illustrating the potential of dual-lock TICS systems for imaging low-abundance biomarkers.
TICS-based wash-free imaging introduces a distinctive mechanistic route to fluorescence control, one that exploits the interplay between charge shuttling and molecular dynamics. Its ability to produce ultra-low background and amplify local molecular events makes it a promising next-generation strategy. However, further structural generalization and quantitative understanding are needed for widespread bioimaging applications.
11.2. Conical intersections
Wash-free bioimaging can be achieved using molecular rotors or environment-sensitive dyes that undergo efficient non-radiative decay through intramolecular rotation and access to conical intersections (CI; Fig. 45a).270,271 CI is a special point where two electronic energy states (i.e., S0 and S1) of a molecule become exactly equal in energy and intersect. Access to a CI is the dominant nonradiative decay route in molecular rotors. The large-amplitude structural motions required to reach this funnel enhance vibronic coupling. This enhancement, in turn, opens additional vibrational relaxation pathways. In aqueous or low-viscosity environments, free rotation of groups such as meso-phenyl rings enables rapid internal conversion via CI, quenching fluorescence. When the probe binds to a biomolecule or enters a viscous or hydrophobic environment, this motion is restricted, forcing radiative relaxation and restoring strong emission.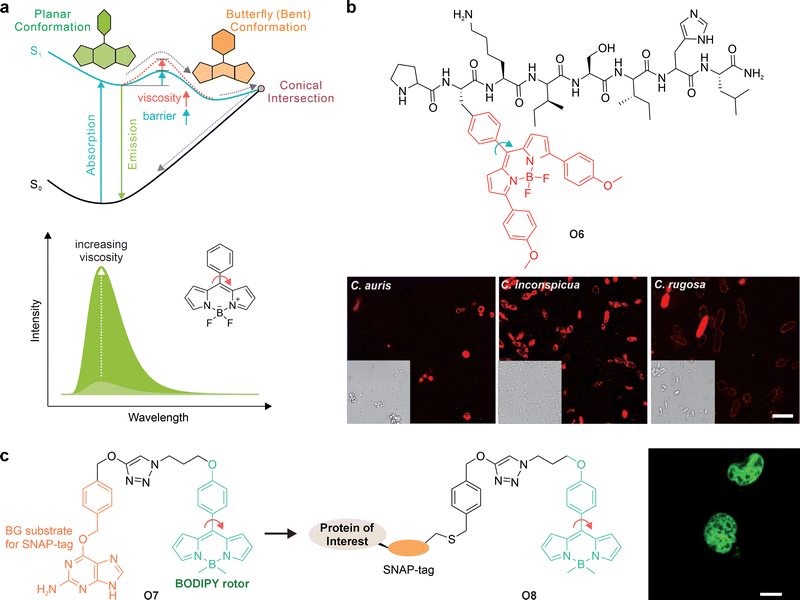 | ||
| Fig. 45 (a) Schematic representation of the potential energy surfaces of molecular rotors, illustrating the approach toward a CI upon photoexcitation and the progressive increase in the energy barrier for CI access as medium viscosity rises (top panel), along with the corresponding changes in fluorescence intensity (bottom panel). (b) Molecular structure of peptide O6 and wash-free live-cell confocal fluorescence imaging of various Candida strains after incubation with O6. (c) Molecular structures of O7 and O8 (left panel) and wash-free fluorescence imaging of O7 conjugated to SNAP-tagged nuclear histone H2B (O8) in live cells (right panel). Scale bars: 10 µm. (b) and (c) are adapted from ref. 271 and 272 with permission from Wiley-VCH and Elsevier, copyright 2022 and 2017, respectively. | ||
A representative example is the phenylalanine–BODIPY (Phe–BODIPY; O6) series developed by Vendrell and co-workers as fluorogenic amino acids for peptide synthesis (Fig. 45b, top panel).271 Removal of the 1,7-methyl groups on the BODIPY core lowered the rotational barrier of the meso-phenyl group, promoting non-emissive CI access in aqueous media (φf <0.1%). When incorporated into antimicrobial peptides such as Jelleine-I analogues, these probes remained dark in buffer or urine but became highly emissive (φf ≈ 0.15) upon interacting with fungal membranes. The optimized probe (Peptide 17) enabled wash-free detection of Candida in urine (Fig. 45b, bottom panel) with a detection limit of 1.8 × 105 CFU mL−1.
Xiao and co-workers designed BDP-V BG, a hybrid sensor coupling a viscosity-sensitive BODIPY rotor (BDP-V) with an O6-benzylguanine (BG) SNAP-tag substrate (O7; Fig. 45c).272 The free probe exhibited weak emission (φf ≈ 0.01) due to rotational quenching. However, SNAP-tag binding restricted rotation, resulting in a 53-fold fluorescence enhancement (in O8). This design directly converts protein labeling into a fluorescence “turn-on” event, eliminating the need for washing steps.
Molecular rotors feature high signal-to-noise ratios, straightforward activation, and real-time responsiveness, making them valuable tools for visualizing dynamic biological processes. However, current molecular rotor designs are mainly limited to the BODIPY family, which offers a relatively narrow emission range. Future research should focus on extending the emission into the near-infrared (NIR) region for deep-tissue imaging by modifying the rotor architecture (i.e., via managing steric hindrance) and integrating it into other established fluorophore scaffolds such as rhodamines and cyanines.
12. Conclusion
The development of fluorogenic molecules has fundamentally transformed modern bioimaging. As detailed in this review, the transition from “always-on” dyes to intelligent, responsive fluorogenic dyes enables interrogation of complex biological systems with unprecedented signal-to-noise ratios and minimal physical perturbation. The field has matured significantly, evolving from serendipitous discoveries to rational design guided by deep photophysical insights.A broad spectrum of mechanisms now underpins this progress. From energy transfer to dark states, PET-driven quenching, IC to dark states, to canonical TICT and TICS, each offers unique advantages for governing emissive dynamics in complex biological environments. Additional strategies such as spirocyclization, HBQ, AIE, DIE, and conical intersection further expand the design space, enabling probes that respond to subtle changes in biomarker concentrations, specific protein/enzymatic activities, cellular microenvironment parameters (e.g., pH, polarity, and viscosity). Recent innovations, including in situ fluorophore formation, underscore the trend toward intelligent, context-specific imaging systems.
Despite these advances, several opportunities remain to propel the field forward:
1. Integration of multiple quenching mechanisms for enhanced specificity
Current designs typically rely on a single fluorogenic mechanism, which limits both dynamic range and selectivity. Incorporating multiple quenching pathways, each responsive to distinct chemical or biological triggers, could enable orthogonal activation modes, dramatically improving sensitivity and specificity in complex environments.
2. Expanding beyond intensity-based imaging to lifetime modulation
Most wash-free strategies exploit changes in fluorescence intensity, yet many mechanisms also modulate excited-state lifetimes. Leveraging fluorescence lifetime imaging microscopy (FLIM) in combination with intensity-based readouts provides a powerful route to multiplexed imaging and deeper insights into dynamic cellular processes. Notably, the lifetime is independent of probe concentration, making it an intrinsic and reliable calibration metric.
3. Adding multifunctionality: from imaging to theranostics
Future probes should extend beyond imaging roles by integrating diagnostic and therapeutic capabilities, such as PDT or controlled drug release. This convergence of imaging and treatment could enable real-time monitoring of therapeutic efficacy and the delivery of “customized” interventions.
4. Overcoming optical penetration limits via hybrid modalities
Even with NIR-I and NIR-II emission, light penetration remains a bottleneck for in vivo imaging. Combining fluorogenic probes with complementary modalities, such as X-ray-activated luminescence, and other imaging modalities (such as ultrasound imaging, magnetic resonance imaging, computed tomography, positron emission tomography, and single photon emission computed tomography) could overcome depth limitations and provide multimodal datasets for comprehensive biological analysis.
5. Computationally guided design and predictive modeling
The complexity of multi-mechanism systems demands rational design strategies. Integrating quantum chemical calculations, molecular dynamics simulations, and machine learning models will accelerate the discovery of probes with optimized photophysical properties and activation kinetics.
6. Addressing practical constraints beyond photophysics
This review focused on photophysical mechanisms for wash-free imaging, but translating these concepts into real-world tools requires tackling additional challenges. Chemical selectivity, cytotoxicity, solubility, photostability, and synthetic feasibility are critical for in vivo applications and must be carefully optimized to ensure reliable performance.
Finally, although our emphasis has been on wash-free imaging, the mechanisms discussed here have broader relevance. They can inform the design of fluorescent sensors and innovative luminescent materials for a wide range of applications. We hope this review serves as a conceptual foundation and a catalyst for innovation across disciplines.
Conflicts of interest
There are no conflicts to declare.Abbreviations
| a-PET | Acceptor-photoinduced electron transfer |
| AIE | Aggregation-induced emission |
| ALA | 5-Aminolevulinic acid |
| ALAS | ALA synthase |
| ALDH2 | Aldehyde dehydrogenase 2 |
| aPDT | Activatable photodynamic therapy |
| BCN | Bicyclo[6.1.0]nonyne |
| BG | O 6-Benzylguanine |
| BHQ | Black hole quencher |
| BL | β-Lactamase |
| BODIPY | Boron-dipyrromethene |
| CA | Carbonic anhydrase |
| CCCP | Carbonyl cyanide m-chlorophenylhydrazone |
| CFSE | Carboxyfluorescein diacetate succinimidyl ester |
| CI | Conical intersection |
| CK-B | B subunit of cytosolic creatine kinase isoenzyme |
| CO | Carbon monoxide |
| CR | Charge recombination |
| CRABPII | Cellular retinoic acid binding protein II |
| CT | Charge transfer |
| CuAAC | Copper-catalyzed azide–alkyne cycloaddition |
| DABCYL | 4-(4-Dimethylaminophenylazo)benzoic acid |
| DET | Dexter energy transfer |
| DIE | Disaggregation-induced emission |
| d-PET | Donor-photoinduced electron transfer |
| EDDI | Elimination of dark states through dimer interactions |
| eDHFR | Engineered dihydrofolate reductase |
| EGFR | Epidermal growth factor receptor |
| EnT | Energy transfer |
| ESIPT | Excited-state intramolecular proton transfer |
| ET | Electron transfer |
| ETC | Electron transport chain |
| ETDS | Energy transfer to a dark state |
| FAPL | Fluorescence activation-coupled protein labeling |
| FGS | Fluorescence-guided surgery |
| FLIM | Fluorescence lifetime imaging microscopy |
| FMO | Frontier molecular orbital |
| FRET | Förster resonance energy transfer |
| fs-TAS | Femtosecond transient absorption spectroscopy |
| GGT | Gamma-glutamyl transferase |
| GSH | Glutathione |
| GST | Glutathione transferase |
| HBQ | Hydrogen-bond-induced quenching |
| hCAII | Human carbonic anhydrase II |
| IC | Internal conversion |
| ICG | Indocyanine green |
| ICT | Intramolecular charge transfer |
| iEDDA | Inverse electron-demand Diels–Alder |
| igMFPs | Mimics of fluorescent proteins |
| ISC | Intersystem crossing |
| LD | Lipid droplet |
| LE | Locally excited state |
| m-AT | meta-Aminothiophenol |
| MDG | Membrane damage green |
| MINFLUX | Minimal photon fluxes nanoscopy |
| miR | microRNA |
| MMP | Matrix metalloproteinase |
| NBD | Nitrobenzoxadiazole |
| NIR | Near-infrared |
| NIR-I | Near-infrared I window |
| NIR-II | Near-infrared II window |
| NRD | Non-radiative decay |
| NVOC | Nitroveratryl oxycarbonyl |
| PCC | Photoinduced charge centralization |
| PDT | Photodynamic therapy |
| PES | Potential energy surface |
| PET | Photoinduced electron transfer |
| PIP2 | Phosphatidylinositol 4,5-bisphosphate |
| PpIX | Protoporphyrin IX |
| PS | Phosphatidylserine |
| PSB | Protonated Schiff base |
| RACI | Restricted access to a conical intersection |
| RADS | Restriction of access to the dark state |
| RIB | Restriction of intramolecular bending |
| RIM | Restriction of intramolecular motion |
| RIV | Restriction of intramolecular vibrations |
| ROS | Reactive oxygen species |
| SET | Single-electron transfer |
| SIM | Structured illumination microscopy |
| SLEET | State-crossing from a locally excited to an electron transfer state |
| SNR | Signal-to-noise ratio |
| SPAAC | Strain-promoted azide–alkyne cycloaddition |
| STED | Stimulated emission depletion |
| TBET | Through-bond energy transfer |
| TCO | trans-Cyclooctene |
| TD-DFT | Time-dependent density functional theory |
| TICT | Twisted intramolecular charge transfer |
| TICS | Twisted intramolecular charge shuttle |
| TMP | Trimethoprim |
| TMT | Tetrazine-mediated transfer |
| TPE | Tetraphenylethene |
| TPP | Triphenylphosphonium |
| β-CAs | β-Chloroacroleins |
Data availability
Additional computational data for this article are available at DR-NTU (Data) at https://doi.org/10.21979/N9/UNEWBM.Acknowledgements
The authors are thankful for the financial support from the Ministry of Education, Singapore (MOE-T2EP10222-0001), the Natural Science Foundation of China (22427802 and 22132002), and the Ministry of Science and Technology of China (2022YFA1205502).References
- L. D. Lavis and R. T. Raines, ACS Chem. Biol., 2008, 3, 142–155 CrossRef CAS PubMed.
- M. Y. Berezin and S. Achilefu, Chem. Rev., 2010, 110, 2641–2684 CrossRef CAS PubMed.
- X. Chen, T. Pradhan, F. Wang, J. S. Kim and J. Yoon, Chem. Rev., 2012, 112, 1910–1956 CrossRef CAS PubMed.
- L. D. Lavis and R. T. Raines, ACS Chem. Biol., 2014, 9, 855–866 CrossRef CAS PubMed.
- X. Li, X. Gao, W. Shi and H. Ma, Chem. Rev., 2014, 114, 590–659 CrossRef CAS PubMed.
- A. S. Klymchenko, Acc. Chem. Res., 2017, 50, 366–375 CrossRef CAS PubMed.
- Y. M. Poronik, K. V. Vygranenko, D. Gryko and D. T. Gryko, Chem. Soc. Rev., 2019, 48, 5242–5265 RSC.
- S. Samanta, K. Lai, F. Wu, Y. Liu, S. Cai, X. Yang, J. Qu and Z. Yang, Chem. Soc. Rev., 2023, 52, 7197–7261 RSC.
- S.-n Uno, M. Kamiya, T. Yoshihara, K. Sugawara, K. Okabe, M. C. Tarhan, H. Fujita, T. Funatsu, Y. Okada, S. Tobita and Y. Urano, Nat. Chem., 2014, 6, 681–689 CrossRef CAS PubMed.
- H. Takakura, Y. Zhang, R. S. Erdmann, A. D. Thompson, Y. Lin, B. McNellis, F. Rivera-Molina, S.-N. Uno, M. Kamiya, Y. Urano, J. E. Rothman, J. Bewersdorf, A. Schepartz and D. Toomre, Nat. Biotechnol., 2017, 35, 773–780 CrossRef CAS PubMed.
- W. Chi, D. Tan, Q. Qiao, Z. Xu and X. Liu, Angew. Chem., Int. Ed., 2023, 62, e202306061 CrossRef CAS PubMed.
- K. Torii, S. Benson, Y. Hori, M. Vendrell and K. Kikuchi, Chem. Sci., 2024, 15, 1393–1401 RSC.
- A. K. Rudd, N. Mittal, E. W. Lim, C. M. Metallo and N. K. Devaraj, J. Am. Chem. Soc., 2020, 142, 17887–17891 CrossRef CAS PubMed.
- L. Li and H. Sun, Biochemistry, 2020, 59, 216–217 CrossRef CAS PubMed.
- P. Shieh, M. J. Hangauer and C. R. Bertozzi, J. Am. Chem. Soc., 2012, 134, 17428–17431 CrossRef CAS PubMed.
- Y. Tsubono, Y. Kawamoto, T. Hidaka, G. N. Pandian, K. Hashiya, T. Bando and H. Sugiyama, J. Am. Chem. Soc., 2020, 142, 17356–17363 CrossRef CAS PubMed.
- M. Minoshima, S. I. Reja, R. Hashimoto, K. Iijima and K. Kikuchi, Chem. Rev., 2024, 124, 6198–6270 CrossRef CAS PubMed.
- X. Wu, R. Wang, N. Kwon, H. Ma and J. Yoon, Chem. Soc. Rev., 2022, 51, 450–463 RSC.
- P. Mauker, D. Beckmann, A. Kitowski, C. Heise, C. Wientjens, A. J. Davidson, S. Wanderoy, G. Fabre, A. B. Harbauer, W. Wood, C. Wilhelm, J. Thorn-Seshold, T. Misgeld, M. Kerschensteiner and O. Thorn-Seshold, J. Am. Chem. Soc., 2024, 146, 11072–11082 CAS.
- J. Zhang, X. Chai, X. P. He, H. J. Kim, J. Yoon and H. Tian, Chem. Soc. Rev., 2019, 48, 683–722 RSC.
- M. Bojtár, K. Németh, F. Domahidy, G. Knorr, A. Verkman, M. Kállay and P. Kele, J. Am. Chem. Soc., 2020, 142, 15164–15171 CrossRef PubMed.
- D. Kim, J. H. Lee, J. Y. Koo, H. M. Kim and S. B. Park, Bioconjugate Chem., 2020, 31, 1545–1550 CrossRef CAS PubMed.
- B. L. Oliveira, Z. Guo, O. Boutureira, A. Guerreiro, G. Jimenez-Oses and G. J. L. Bernardes, Angew. Chem., Int. Ed., 2016, 55, 14683–14687 CrossRef CAS PubMed.
- K. Fujita and Y. Urano, Chem. Rev., 2024, 124, 4021–4078 Search PubMed.
- E. Kozma and P. Kele, Org. Biomol. Chem., 2019, 17, 215–233 RSC.
- P. Werther, K. Yserentant, F. Braun, K. Grußmayer, V. Navikas, M. Yu, Z. Zhang, M. J. Ziegler, C. Mayer, A. J. Gralak, M. Busch, W. Chi, F. Rominger, A. Radenovic, X. Liu, E. A. Lemke, T. Buckup, D.-P. Herten and R. Wombacher, ACS Cent. Sci., 2021, 7, 1561–1571 CrossRef CAS PubMed.
- F. de Moliner, Z. Konieczna, L. Mendive-Tapia, R. S. Saleeb, K. Morris, J. A. Gonzalez-Vera, T. Kaizuka, S. G. N. Grant, M. H. Horrocks and M. Vendrell, Angew. Chem., Int. Ed., 2023, 62, e202216231 CrossRef CAS PubMed.
- F. Nadal-Bufi, R. V. Nithun, F. de Moliner, X. Lin, S. Habiballah, M. Jbara and M. Vendrell, ACS Cent. Sci., 2025, 11, 66–75 CrossRef CAS PubMed.
- J. Chen, C. Wang, W. Liu, Q. Qiao, H. Qi, W. Zhou, N. Xu, J. Li, H. Piao, D. Tan, X. Liu and Z. Xu, Angew. Chem., Int. Ed., 2021, 60, 25104–25113 CrossRef CAS PubMed.
- Q. Qi, J. Li, Q. Qiao, C. Yan, M. Izadyar, C. Wang, S. A. A. Abedi, X. Liu, Z. Guo and Z. Xu, CCS Chem., 2025, 7, 3409–3420 CrossRef CAS.
- J. Tan, C. Wang, Z. Hu and X. Zhang, Exploration, 2025, 5, 20230094 CrossRef CAS PubMed.
- P. Klán and J. Wirz, Photochemistry of organic compounds: from concepts to practice, John Wiley & Sons, Hoboken, 2009 Search PubMed.
- V. Balzani, P. Ceroni and A. Juris, Photochemistry and photophysics: concepts, research, applications, Wiley-VCH, Weinheim, 2014 Search PubMed.
- W. Chi, L. Huang, C. Wang, D. Tan, Z. Xu and X. Liu, Mater. Chem. Front., 2021, 5, 7012–7021 RSC.
- B. Fang, Y. Shen, B. Peng, H. Bai, L. Wang, J. Zhang, W. Hu, L. Fu, W. Zhang, L. Li and W. Huang, Angew. Chem., Int. Ed., 2022, 61, e202207188 CrossRef CAS PubMed.
- C. Jing and V. W. Cornish, ACS Chem. Biol., 2013, 8, 1704–1712 CrossRef CAS PubMed.
- J. Yang, J. Šečkutė, C. M. Cole and N. K. Devaraj, Angew. Chem., Int. Ed., 2012, 51, 7476–7479 CrossRef CAS PubMed.
- T. Myochin, K. Hanaoka, S. Iwaki, T. Ueno, T. Komatsu, T. Terai, T. Nagano and Y. Urano, J. Am. Chem. Soc., 2015, 137, 4759–4765 CrossRef CAS PubMed.
- X. Sun, A. Zhang, B. Baker, L. Sun, A. Howard, J. Buswell, D. Maurel, A. Masharina, K. Johnsson, C. J. Noren, M. Q. Xu and I. R. Corrêa Jr, ChemBioChem, 2011, 12, 2217–2226 CrossRef CAS PubMed.
- A. Yu, X. He, T. Shen, X. Yu, W. Mao, W. Chi, X. Liu and H. Wu, Chem. Soc. Rev., 2025, 54, 2984–3016 RSC.
- T. Shen and X. Liu, Chem. Sci., 2025, 16, 4595–4613 RSC.
- A.-C. Knall, M. Hollauf and C. Slugovc, Tetrahedron Lett., 2014, 55, 4763–4766 Search PubMed.
- M. J. Hangauer and C. R. Bertozzi, Angew. Chem., Int. Ed., 2008, 47, 2394–2397 CrossRef CAS PubMed.
- T. Komatsu, K. Johnsson, H. Okuno, H. Bito, T. Inoue, T. Nagano and Y. Urano, J. Am. Chem. Soc., 2011, 133, 6745–6751 CrossRef CAS PubMed.
- S. Mizukami, S. Watanabe, Y. Akimoto and K. Kikuchi, J. Am. Chem. Soc., 2012, 134, 1623–1629 CrossRef CAS PubMed.
- N. K. Devaraj, S. Hilderbrand, R. Upadhyay, R. Mazitschek and R. Weissleder, Angew. Chem., Int. Ed., 2010, 49, 2869–2872 CrossRef CAS PubMed.
- H. Wu, J. Yang, J. Šečkutė and N. K. Devaraj, Angew. Chem., Int. Ed., 2014, 53, 5805–5809 CrossRef CAS PubMed.
- A. Aktalay, R. Lincoln, L. Heynck, M. A. D. R. Lima, A. N. Butkevich, M. L. Bossi and S. W. Hell, ACS Cent. Sci., 2023, 9, 1581–1590 Search PubMed.
- H. Wu, B. T. Cisneros, C. M. Cole and N. K. Devaraj, J. Am. Chem. Soc., 2014, 136, 17942–17945 CrossRef CAS PubMed.
- L. G. Meimetis, J. C. Carlson, R. J. Giedt, R. H. Kohler and R. Weissleder, Angew. Chem., Int. Ed., 2014, 53, 7531–7534 CrossRef CAS PubMed.
- A. Wieczorek, P. Werther, J. Euchner and R. Wombacher, Chem. Sci., 2017, 8, 1506–1510 RSC.
- X. Zhu, J. Zhang, H. Geng, K. Wang, Y. Li, X. Yang, H. Shi, J. Tai, X. Yan, Y. Yang, H. Sun and Q. Zhu, Sci. Adv., 2025, 11, eadw1098 CrossRef CAS PubMed.
- B. Daly, J. Ling and A. P. de Silva, Chem. Soc. Rev., 2015, 44, 4203–4211 RSC.
- H. Niu, J. Liu, H. M. O'Connor, T. Gunnlaugsson, T. D. James and H. Zhang, Chem. Soc. Rev., 2023, 52, 2322–2357 RSC.
- Y. Chen, K. Tsao, S. L. Acton and J. W. Keillor, Angew. Chem., Int. Ed., 2018, 57, 12390–12394 CrossRef CAS.
- Y. Wu, G. Guo, J. Zheng, D. Xing and T. Zhang, ACS Sens., 2019, 4, 44–51 CrossRef CAS PubMed.
- R. Huang, T. Liu, H. Peng, J. Liu, X. Liu, L. Ding and Y. Fang, Chem. Soc. Rev., 2024, 53, 6960–6991 RSC.
- M. Xiao, W. Sun, J. Fan, J. Cao, Y. Li, K. Shao, M. Li, X. Li, Y. Kang, W. Zhang, S. Long, J. Du and X. Peng, Adv. Funct. Mater., 2018, 28, 1805128 CrossRef.
- X. Zhang, L. Chi, S. Ji, Y. Wu, P. Song, K. Han, H. Guo, T. D. James and J. Zhao, J. Am. Chem. Soc., 2009, 131, 17452–17463 CrossRef CAS PubMed.
- L. E. Greene, R. Godin and G. Cosa, J. Am. Chem. Soc., 2016, 138, 11327–11334 CrossRef CAS PubMed.
- L. E. Greene, R. Lincoln and G. Cosa, J. Am. Chem. Soc., 2017, 139, 15801–15811 CrossRef CAS PubMed.
- S. U. Hettiarachchi, B. Prasai and R. L. McCarley, J. Am. Chem. Soc., 2014, 136, 7575–7578 CrossRef CAS PubMed.
- A. S. Walker, P. R. Rablen and A. Schepartz, J. Am. Chem. Soc., 2016, 138, 7143–7150 CrossRef CAS PubMed.
- N. D. Barth, R. Subiros-Funosas, L. Mendive-Tapia, R. Duffin, M. A. Shields, J. A. Cartwright, S. T. Henriques, J. Sot, F. M. Goni, R. Lavilla, J. A. Marwick, S. Vermeren, A. G. Rossi, M. Egeblad, I. Dransfield and M. Vendrell, Nat. Commun., 2020, 11, 4027 CrossRef CAS PubMed.
- T. C. Owyong, P. Subedi, J. Deng, E. Hinde, J. J. Paxman, J. M. White, W. Chen, B. Heras, W. W. H. Wong and Y. Hong, Angew. Chem., Int. Ed., 2020, 59, 10129–10135 CrossRef CAS PubMed.
- S. Jeon, T. I. Kim, H. Jin, U. Lee, J. Bae, J. Bouffard and Y. Kim, J. Am. Chem. Soc., 2020, 142, 9231–9239 CrossRef CAS PubMed.
- H. Li, L. Guo, W. Miao, Z. Wang, Z. Kang, S. Wang, X. Guo, C. Yu, L. Jiao and E. Hao, Angew. Chem., Int. Ed., 2025, 64, e202509567 CrossRef CAS PubMed.
- W. Chi, J. Chen, W. Liu, C. Wang, Q. Qi, Q. Qiao, T. M. Tan, K. Xiong, X. Liu, K. Kang, Y. T. Chang, Z. Xu and X. Liu, J. Am. Chem. Soc., 2020, 142, 6777–6785 CrossRef CAS PubMed.
- B. Shen, K. H. Jung, S. Ye, C. A. Hoelzel, C. H. Wolstenholme, H. Huang, Y. Liu and X. Zhang, Aggregate, 2023, 4, e301 CrossRef CAS.
- H. Zhang, J. Fan, J. Wang, B. Dou, F. Zhou, J. Cao, J. Qu, Z. Cao, W. Zhao and X. Peng, J. Am. Chem. Soc., 2013, 135, 17469–17475 CrossRef CAS PubMed.
- X. Y. Lauteslager, B. Wegewijs, J. W. Verhoeven and A. M. Brouwer, J. Photochem. Photobiol., A, 1996, 98, 121–126 CrossRef CAS.
- S. Huang, S. A. A. Abedi, Z. Li, R. Huang, X. Yan, M. Izadyar, Q. Qiao, Y. Fang, Z. Xu and X. Liu, CCS Chem., 2024, 6, 2804–2813 CrossRef CAS.
- S. Leng, Q. Qiao, L. Miao, W. Deng, J. Cui and Z. Xu, Chem. Commun., 2017, 53, 6448–6451 RSC.
- Z. Ma, Z. Fu, B. Hou, X. Zeng, J. Wang, Y. Tan, Y. Jiang, N. Xu and C. Tan, J. Am. Chem. Soc., 2025, 147, 37005–37011 CrossRef CAS PubMed.
- Y. Urano, D. Asanuma, Y. Hama, Y. Koyama, T. Barrett, M. Kamiya, T. Nagano, T. Watanabe, A. Hasegawa, P. L. Choyke and H. Kobayashi, Nat. Med., 2009, 15, 104–109 CrossRef CAS PubMed.
- A. Weller, Pure Appl. Chem., 1968, 16, 115–124 CrossRef CAS.
- G. A. Parada, Z. K. Goldsmith, S. Kolmar, B. Pettersson Rimgard, B. Q. Mercado, L. Hammarström, S. Hammes-Schiffer and J. M. Mayer, Science, 2019, 364, 471–475 CrossRef CAS PubMed.
- R. A. Marcus, J. Chem. Phys., 1956, 24, 966–978 CrossRef CAS.
- R. A. Marcus, Angew. Chem., Int. Ed. Engl., 1993, 32, 1111–1121 CrossRef.
- C.-P. Hsu, Acc. Chem. Res., 2009, 42, 509–518 CrossRef CAS PubMed.
- W. Mao, W. Chi, X. He, C. Wang, X. Wang, H. Yang, X. Liu and H. Wu, Angew. Chem., Int. Ed., 2022, 61, e202117386 CrossRef CAS PubMed.
- Y. Hori, K. Nakaki, M. Sato, S. Mizukami and K. Kikuchi, Angew. Chem., Int. Ed., 2012, 51, 5611–5614 CrossRef CAS PubMed.
- P. Shieh, V. T. Dien, B. J. Beahm, J. M. Castellano, T. Wyss-Coray and C. R. Bertozzi, J. Am. Chem. Soc., 2015, 137, 7145–7151 CrossRef CAS PubMed.
- J. Zhang, A. Shibata, M. Ito, S. Shuto, Y. Ito, B. Mannervik, H. Abe and R. Morgenstern, J. Am. Chem. Soc., 2011, 133, 14109–14119 CrossRef CAS PubMed.
- L. Mendive-Tapia, C. Zhao, A. R. Akram, S. Preciado, F. Albericio, M. Lee, A. Serrels, N. Kielland, N. D. Read, R. Lavilla and M. Vendrell, Nat. Commun., 2016, 7, 10940 CrossRef CAS PubMed.
- M. A. Filatov, S. Karuthedath, P. M. Polestshuk, H. Savoie, K. J. Flanagan, C. Sy, E. Sitte, M. Telitchko, F. Laquai, R. W. Boyle and M. O. Senge, J. Am. Chem. Soc., 2017, 139, 6282–6285 CrossRef CAS PubMed.
- Y. Li, Y. Sun, J. Li, Q. Su, W. Yuan, Y. Dai, C. Han, Q. Wang, W. Feng and F. Li, J. Am. Chem. Soc., 2015, 137, 6407–6416 CrossRef CAS PubMed.
- A. L. D. Wallabregue, H. Bolland, S. Faulkner, E. M. Hammond and S. J. Conway, J. Am. Chem. Soc., 2023, 145, 2572–2583 CrossRef CAS PubMed.
- C. Wang, W. Chi, Q. Qiao, D. Tan, Z. Xu and X. Liu, Chem. Soc. Rev., 2021, 50, 12656–12678 Search PubMed.
- R. Huang, M. Li, D. Lin, Y. Shao, C. Shang, Q. Liu, G. Liu, N. Li, R. Miao, H. Peng, Y. Tang and Y. Fang, Aggregate, 2023, 4, e203 Search PubMed.
- Q. Qiao, A. Song, G. Jiang, Y. Zhou, Y. Ruan, W. Jia, X. Liu and Z. Xu, Angew. Chem., Int. Ed., 2025, 64, e202503177 CrossRef CAS PubMed.
- R. Huang, H. Liu, K. Liu, G. Wang, Q. Liu, Z. Wang, T. Liu, R. Miao, H. Peng and Y. Fang, Anal. Chem., 2019, 91, 14451–14457 CrossRef CAS PubMed.
- J. V. Jun, E. J. Petersson and D. M. Chenoweth, J. Am. Chem. Soc., 2018, 140, 9486–9493 CrossRef CAS PubMed.
- C. Wang, W. Jiang, D. Tan, L. Huang, J. Li, Q. Qiao, P. Yadav, X. Liu and Z. Xu, Chem. Sci., 2023, 14, 4786–4795 RSC.
- R. Miao, J. Li, C. Wang, X. Jiang, Y. Gao, X. Liu, D. Wang, X. Li, X. Liu and Y. Fang, Adv. Sci., 2022, 9, e2104609 CrossRef PubMed.
- N. I. Shank, H. H. Pham, A. S. Waggoner and B. A. Armitage, J. Am. Chem. Soc., 2013, 135, 242–251 Search PubMed.
- R. Tao, Y. Chen, T. Yang, S. Hu, W. Lv, X. Li, Z. Wang, R. Zhang, Z. Wu, T. Hou, Z. Xu, Y. Wang, X. Liu and X. Li, J. Am. Chem. Soc., 2025, 147, 41768–41778 CrossRef CAS PubMed.
- K. Hanaoka, S. Iwaki, K. Yagi, T. Myochin, T. Ikeno, H. Ohno, E. Sasaki, T. Komatsu, T. Ueno, M. Uchigashima, T. Mikuni, K. Tainaka, S. Tahara, S. Takeuchi, T. Tahara, M. Uchiyama, T. Nagano and Y. Urano, J. Am. Chem. Soc., 2022, 144, 19778–19790 CrossRef CAS PubMed.
- N. Shin, K. Hanaoka, W. Piao, T. Miyakawa, T. Fujisawa, S. Takeuchi, S. Takahashi, T. Komatsu, T. Ueno, T. Terai, T. Tahara, M. Tanokura, T. Nagano and Y. Urano, ACS Chem. Biol., 2017, 12, 558–563 CrossRef CAS PubMed.
- C. Wang, Q. Qiao, W. Chi, J. Chen, W. Liu, D. Tan, S. McKechnie, D. Lyu, X.-F. Jiang, W. Zhou, N. Xu, Q. Zhang, Z. Xu and X. Liu, Angew. Chem., Int. Ed., 2020, 59, 10160–10172 CrossRef CAS PubMed.
- J. J. Shah, M. Gaitan and J. Geist, Anal. Chem., 2009, 81, 8260–8263 CrossRef CAS PubMed.
- J. Sakakibara and R. J. Adrian, Exp. Fluids, 1999, 26, 7–15 CrossRef CAS.
- X.-D. Wang, O. S. Wolfbeis and R. J. Meier, Chem. Soc. Rev., 2013, 42, 7834–7869 RSC.
- Y. Venkatesh, K. B. Narayan, T. Baumgart and E. J. Petersson, J. Am. Chem. Soc., 2025, 147, 15115–15125 CrossRef CAS PubMed.
- Y. Liu, C. H. Wolstenholme, G. C. Carter, H. Liu, H. Hu, L. S. Grainger, K. Miao, M. Fares, C. A. Hoelzel, H. P. Yennawar, G. Ning, M. Du, L. Bai, X. Li and X. Zhang, J. Am. Chem. Soc., 2018, 140, 7381–7384 CrossRef CAS PubMed.
- X. Zhang, L. Yuan, J. Jiang, J. Hu, A. du Rietz, H. Cao, R. Zhang, X. Tian, F. Zhang, Y. Ma, Z. Zhang, K. Uvdal and Z. Hu, Anal. Chem., 2020, 92, 3613–3619 CrossRef CAS PubMed.
- F. Hovelmann, I. Gaspar, J. Chamiolo, M. Kasper, J. Steffen, A. Ephrussi and O. Seitz, Chem. Sci., 2016, 7, 128–135 RSC.
- X. Luo, B. Xue, G. Feng, J. Zhang, B. Lin, P. Zeng, H. Li, H. Yi, X. L. Zhang, H. Zhu and Z. Nie, J. Am. Chem. Soc., 2019, 141, 5182–5191 CrossRef CAS PubMed.
- Y. Hori, T. Norinobu, M. Sato, K. Arita, M. Shirakawa and K. Kikuchi, J. Am. Chem. Soc., 2013, 135, 12360–12365 CrossRef CAS PubMed.
- T. Gao, X. Cao, J. Dong, Y. Liu, W. Lv, C. Li, X. Feng and W. Zeng, Dyes Pigm., 2017, 143, 436–443 CrossRef CAS.
- N. Banahene, D. M. Gepford, K. J. Biegas, D. H. Swanson, Y. P. Hsu, B. A. Murphy, Z. E. Taylor, I. Lepori, M. S. Siegrist, A. Obregon-Henao, M. S. Van Nieuwenhze and B. M. Swarts, Angew. Chem., Int. Ed., 2023, 62, e202213563 CrossRef CAS PubMed.
- C. J. Wu, X. Y. Li, T. Zhu, M. Zhao, Z. Song, S. Li, G. G. Shan and G. Niu, Anal. Chem., 2022, 94, 3881–3887 CrossRef CAS PubMed.
- J. S. Paige, K. Y. Wu and S. R. Jaffrey, Science, 2011, 333, 642–646 CrossRef CAS PubMed.
- J. Zhang, H. Li, B. Lin, X. Luo, P. Yin, T. Yi, B. Xue, X. L. Zhang, H. Zhu and Z. Nie, J. Am. Chem. Soc., 2021, 143, 19317–19329 CrossRef CAS PubMed.
- L. Hu, W. Cao, Y. Jiang, W. Cai, X. Lou and T. Liu, Nat. Chem., 2024, 16, 1960–1971 CrossRef CAS PubMed.
- S. Chanmungkalakul, S. A. A. Abedi, F. J. Hernández, J. Xu and X. Liu, Chin. Chem. Lett., 2024, 35, 109227 CrossRef CAS.
- Y. Lee, W. Cho, J. Sung, E. Kim and S. B. Park, J. Am. Chem. Soc., 2018, 140, 974–983 CrossRef CAS PubMed.
- W. Mao, J. Tang, L. Dai, X. He, J. Li, L. Cai, P. Liao, R. Jiang, J. Zhou and H. Wu, Angew. Chem., Int. Ed., 2021, 60, 2393–2397 CrossRef CAS PubMed.
- W. Wei, J. Zhang, Z. Xu, Z. Liu, C. Huang, K. Cheng, L. Meng, Y. Matsuda, Q. Hao, H. Zhang and H. Sun, ACS Sens., 2023, 8, 28–39 CrossRef CAS PubMed.
- H. Wang, H. Wang, T. Xiu, X. Zhang, Y. Tang, W. Zhang, W. Zhang, P. Li and B. Tang, Angew. Chem., Int. Ed., 2025, 64, e202506728 CrossRef CAS PubMed.
- H. Son, D. Kim, S. Kim, W. G. Byun and S. B. Park, Angew. Chem., Int. Ed., 2025, 64, e202421982 CrossRef CAS PubMed.
- Y. Deng, T. Shen, X. Yu, J. Li, P. Zou, Q. Gong, Y. Zheng, H. Sun, X. Liu and H. Wu, Angew. Chem., Int. Ed., 2024, 63, e202319853 CrossRef CAS PubMed.
- T. Shen and X. Liu, Bioconjug. Chem., 2024, 35, 1024–1032 CrossRef CAS PubMed.
- T. Shen, D. Tan, M. Shanmugham and X. Liu, Sens. Diagn., 2022, 1, 714–718 RSC.
- D. Kim, H. Son and S. B. Park, Angew. Chem., Int. Ed., 2023, 62, e202310665 CrossRef CAS PubMed.
- T. Shen, W. Zhang, P. Yadav, X. W. Sun and X. Liu, Mater. Chem. Front., 2023, 7, 1082–1092 RSC.
- S. Uchiyama, K. Takehira, S. Kohtani, K. Imai, R. Nakagaki, S. Tobita and T. Santa, Org. Biomol. Chem., 2003, 1, 1067–1072 RSC.
- A. N. Butkevich, G. Y. Mitronova, S. C. Sidenstein, J. L. Klocke, D. Kamin, D. N. H. Meineke, E. D'Este, P. T. Kraemer, J. G. Danzl, V. N. Belov and S. W. Hell, Angew. Chem., Int. Ed., 2016, 55, 3290–3294 CrossRef CAS PubMed.
- Z. Du, W. Wang, S. Luo, L. Zhang, S. Yuan, Y. Hei, Z. Bao, C. Chen, Y. Lin and L. Chu, J. Am. Chem. Soc., 2023, 145, 18968–18976 CrossRef CAS PubMed.
- D. Si, Q. Li, Y. Bao, J. Zhang and L. Wang, Angew. Chem., Int. Ed., 2023, 62, e202307641 CrossRef CAS PubMed.
- Q. Wang, Y. Ma, Y. Shang, T. Liu, X. Liu, J. Liu, L. Ding, R. Miao and Y. Fang, Angew. Chem., Int. Ed., 2025, 64, e202514295 CrossRef CAS PubMed.
- Q. Zheng, A. X. Ayala, I. Chung, A. V. Weigel, A. Ranjan, N. Falco, J. B. Grimm, A. N. Tkachuk, C. Wu, J. Lippincott-Schwartz, R. H. Singer and L. D. Lavis, ACS Cent. Sci., 2019, 5, 1602–1613 CrossRef CAS PubMed.
- W. Chi, Q. Qiao, C. Wang, J. Zheng, W. Zhou, N. Xu, X. Wu, X. Jiang, D. Tan, Z. Xu and X. Liu, Angew. Chem., Int. Ed., 2020, 59, 20215–20223 CrossRef CAS PubMed.
- X. Wu, Y. Gao, W. Chi, C. Wang, Z. Xu and X. Liu, Mater. Chem. Front., 2023, 7, 1137–1145 RSC.
- W. Chi, Q. Qi, R. Lee, Z. Xu and X. Liu, J. Phys. Chem. C, 2020, 124, 3793–3801 CrossRef CAS.
- F. Deng, Q. Qiao, J. Li, W. Yin, L. Miao, X. Liu and Z. Xu, J. Phys. Chem. B, 2020, 124, 7467–7474 CrossRef CAS PubMed.
- M. Qiao, Y. Cao, T. Liu, L. Ding and Y. Fang, Sens. Actuators, B, 2021, 331, 129408 CrossRef CAS.
- J. B. Grimm, A. K. Muthusamy, Y. Liang, T. A. Brown, W. C. Lemon, R. Patel, R. Lu, J. J. Macklin, P. J. Keller, N. Ji and L. D. Lavis, Nat. Methods, 2017, 14, 987–994 CrossRef CAS PubMed.
- L. Wang, M. Tran, E. D’Este, J. Roberti, B. Koch, L. Xue and K. Johnsson, Nat. Chem., 2020, 12, 165–172 CrossRef CAS PubMed.
- J. B. Grimm, A. N. Tkachuk, L. Xie, H. Choi, B. Mohar, N. Falco, K. Schaefer, R. Patel, Q. Zheng, Z. Liu, J. Lippincott-Schwartz, T. A. Brown and L. D. Lavis, Nat. Methods, 2020, 17, 815–821 CrossRef CAS PubMed.
- J. B. Grimm, T. Klein, B. G. Kopek, G. Shtengel, H. F. Hess, M. Sauer and L. D. Lavis, Angew. Chem., Int. Ed., 2016, 55, 1723–1727 CrossRef CAS PubMed.
- W. Wu, K. Yan, Z. He, L. Zhang, Y. Dong, B. Wu, H. Liu, S. Wang and F. Zhang, J. Am. Chem. Soc., 2024, 146, 11570–11576 CAS.
- Y. Wang, D. Torres-Garcia, T. P. Mostert, L. Reinalda and S. I. Van Kasteren, Angew. Chem., Int. Ed., 2024, 63, e202401733 CrossRef CAS PubMed.
- T. Doura, M. Kamiya, F. Obata, Y. Yamaguchi, T. Y. Hiyama, T. Matsuda, A. Fukamizu, M. Noda, M. Miura and Y. Urano, Angew. Chem., Int. Ed., 2016, 55, 9620–9624 CrossRef CAS PubMed.
- H. Ito, Y. Kawamata, M. Kamiya, K. Tsuda-Sakurai, S. Tanaka, T. Ueno, T. Komatsu, K. Hanaoka, S. Okabe, M. Miura and Y. Urano, Angew. Chem., Int. Ed., 2018, 57, 15702–15706 CrossRef CAS PubMed.
- E. A. Halabi, Z. Thiel, N. Trapp, D. Pinotsi and P. Rivera-Fuentes, J. Am. Chem. Soc., 2017, 139, 13200–13207 CrossRef CAS PubMed.
- L. He, W. Lin, Q. Xu, M. Ren, H. Wei and J. Y. Wang, Chem. Sci., 2015, 6, 4530–4536 RSC.
- A. Sakama, H. Seo, J. Hara, Y. Shindo, Y. Ikeda, K. Oka, D. Citterio and Y. Hiruta, Chem. Commun., 2024, 60, 5984–5987 RSC.
- S. M. Usama, S. C. Marker, D.-H. Li, D. R. Caldwell, M. Stroet, N. L. Patel, A. G. Tebo, S. Hernot, J. D. Kalen and M. Schnermann, J. Am. Chem. Soc., 2023, 145, 14647–14659 CrossRef CAS PubMed.
- K. Miki, K. Kojima, K. Oride, H. Harada, A. Morinibu and K. Ohe, Chem. Commun., 2017, 53, 7792–7795 RSC.
- A. Martin and P. Rivera-Fuentes, Nat. Chem., 2024, 16, 28–35 CrossRef CAS PubMed.
- F. Wu, B. Zheng, X. Qiu, T. Hu, Y. Yang, K. Yang, B. Zhou, Q. Ma, D. W. Pang and H. Xiong, Angew. Chem., Int. Ed., 2025, 64, e202423059 CrossRef CAS PubMed.
- R. Wirth, P. Gao, G. U. Nienhaus, M. Sunbul and A. Jaschke, J. Am. Chem. Soc., 2019, 141, 7562–7571 CrossRef CAS PubMed.
- N. Lardon, L. Wang, A. Tschanz, P. Hoess, M. Tran, E. D’Este, J. Ries and K. Johnsson, J. Am. Chem. Soc., 2021, 143, 14592–14600 CrossRef CAS PubMed.
- J. B. Grimm, B. P. English, J. Chen, J. P. Slaughter, Z. Zhang, A. Revyakin, R. Patel, J. J. Macklin, D. Normanno, R. H. Singer, T. Lionnet and L. D. Lavis, Nat. Methods, 2015, 12, 244–250 CrossRef CAS PubMed.
- G. Lukinavicius, L. Reymond, K. Umezawa, O. Sallin, E. D'Este, F. Gottfert, H. Ta, S. W. Hell, Y. Urano and K. Johnsson, J. Am. Chem. Soc., 2016, 138, 9365–9368 CrossRef CAS PubMed.
- G. Lukinavičius, K. Umezawa, N. Olivier, A. Honigmann, G. Yang, T. Plass, V. Mueller, L. Reymond, I. R. Corrêa Jr, Z.-G. Luo, C. Schultz, E. A. Lemke, P. Heppenstall, C. Eggeling, S. Manley and K. Johnsson, Nat. Chem., 2013, 5, 132–139 CrossRef PubMed.
- F. Deng and Z. Xu, Chin. Chem. Lett., 2019, 30, 1667–1681 CrossRef CAS.
- J. Liu, Y.-Q. Sun, H. Zhang, H. Shi, Y. Shi and W. Guo, ACS Appl. Mater. Interfaces, 2016, 8, 22953–22962 CrossRef CAS PubMed.
- X. Chai, X. Cui, B. Wang, F. Yang, Y. Cai, Q. Wu and T. Wang, Chem. – Eur. J., 2015, 21, 16754–16758 CrossRef CAS PubMed.
- Y. Koide, Y. Urano, K. Hanaoka, W. Piao, M. Kusakabe, N. Saito, T. Terai, T. Okabe and T. Nagano, J. Am. Chem. Soc., 2012, 134, 5029–5031 CrossRef CAS PubMed.
- Y. Koide, Y. Urano, K. Hanaoka, T. Terai and T. Nagano, J. Am. Chem. Soc., 2011, 133, 5680–5682 CrossRef CAS PubMed.
- S. G. Stratton, G. H. Taumoefolau, G. E. Purnell, M. Rasooly, W. L. Czaplyski and E. J. Harbron, Chem. – Eur. J., 2017, 23, 14064–14072 CrossRef CAS PubMed.
- X. Chen, J. Liu, M. Chen, J. Zhou, Y. Zhang, X. Hu, W. Geng, Q. Mao, H. Kitagishi, J. Chen, X. Qian, Y. Yang, Y. Lei and X. Luo, J. Am. Chem. Soc., 2024, 146, 30361–30371 CrossRef CAS PubMed.
- M. K. Lee, P. Rai, J. Williams, R. J. Twieg and W. E. Moerner, J. Am. Chem. Soc., 2014, 136, 14003–14006 CrossRef CAS PubMed.
- Q. Qi, W. Chi, Y. Li, Q. Qiao, J. Chen, L. Miao, Y. Zhang, J. Li, W. Ji, T. Xu, X. Liu, J. Yoon and Z. Xu, Chem. Sci., 2019, 10, 4914–4922 RSC.
- F. Walterspiel, B. Ugarte-Uribe, J. Weidenhausen, M. Vincent, K. K. Narayanasamy, A. Dimitriadi, A. U. M. Khan, M. Fritsch, C. W. Muller, T. Zimmermann and C. Deo, Angew. Chem., Int. Ed., 2025, 64, e202424955 CrossRef CAS PubMed.
- H. Zuo, Y. Liu, L. Wang, R. Li, X. Guo, L. Guo, H. Li, M. Li, X. Zhang, L. Jiao, Y. Xiao and E. Hao, Angew. Chem., Int. Ed., 2025, 64, e202518973 Search PubMed.
- Y. Zheng, Z. Ye, X. Zhang and Y. Xiao, J. Am. Chem. Soc., 2023, 145, 5125–5133 CrossRef CAS PubMed.
- Z. Ye, H. Yu, W. Yang, Y. Zheng, N. Li, H. Bian, Z. Wang, Q. Liu, Y. Song, M. Zhang and Y. Xiao, J. Am. Chem. Soc., 2019, 141, 6527–6536 CrossRef CAS PubMed.
- X. Fang, Q. Qiao, Z. Li, H.-K. Li, Y. Huang, D. Hou, J. Chen, N. Xu, K. An, W. Jiang, Y. Tao, P. Bao, Y. Zhang, Z. Wu, X. Liu and Z. Xu, J. Am. Chem. Soc., 2025, 147, 22253–22267 CrossRef CAS PubMed.
- L. G. Wang, A. R. Montaño, J. R. Combs, N. P. McMahon, A. Solanki, M. M. Gomes, K. Tao, W. H. Bisson, D. A. Szafran, K. S. Samkoe, K. M. Tichauer and S. L. Gibbs, Nat. Chem., 2023, 15, 729–739 CrossRef CAS PubMed.
- T. Kamikawa, A. Hashimoto, N. Yamazaki, J. Adachi, A. Matsushima, K. Kikuchi and Y. Hori, Chem. Sci., 2024, 15, 8097–8105 RSC.
- R. Huang, Q. Qiao, D. Seah, T. Shen, X. Wu, F. de Moliner, C. Wang, N. Ding, W. Chi, H. Sun, M. Vendrell, Z. Xu, Y. Fang and X. Liu, J. Am. Chem. Soc., 2025, 147, 5258–5268 CrossRef CAS PubMed.
- N. Barman, D. Singha and K. Sahu, J. Phys. Chem. A, 2013, 117, 3945–3953 CrossRef CAS PubMed.
- G.-J. Zhao and K.-L. Han, J. Phys. Chem. A, 2007, 111, 9218–9223 CrossRef CAS PubMed.
- T.-K. Liu, P.-Y. Hsieh, Y.-D. Zhuang, C.-Y. Hsia, C.-L. Huang, H.-P. Lai, H.-S. Lin, I. C. Chen, H.-Y. Hsu and K.-T. Tan, ACS Chem. Biol., 2014, 9, 2359–2365 CrossRef CAS PubMed.
- Y. Gao, C. Wang, W. Chi and X. Liu, Chem. – Asian J., 2020, 15, 4082–4086 CrossRef CAS PubMed.
- K. A. Bertman, C. S. Abeywickrama, H. J. Baumann, N. Alexander, L. McDonald, L. P. Shriver, M. Konopka and Y. Pang, J. Mater. Chem. B, 2018, 6, 5050–5058 RSC.
- B. Liu, M. Shah, G. Zhang, Q. Liu and Y. Pang, ACS Appl. Mater. Interfaces, 2014, 6, 21638–21644 CrossRef CAS PubMed.
- A. S. Klymchenko, Acc. Chem. Res., 2023, 56, 1–12 CrossRef CAS PubMed.
- M. Collot, T. K. Fam, P. Ashokkumar, O. Faklaris, T. Galli, L. Danglot and A. S. Klymchenko, J. Am. Chem. Soc., 2018, 140, 5401–5411 Search PubMed.
- M. Collot, S. Bou, T. K. Fam, L. Richert, Y. Mely, L. Danglot and A. S. Klymchenko, Anal. Chem., 2019, 91, 1928–1935 CrossRef CAS PubMed.
- P. Zhou, P. Li, Y. Zhao and K. Han, J. Phys. Chem. Lett., 2019, 10, 6929–6935 CrossRef CAS PubMed.
- J. Guo, J. Fan, X. Liu, Z. Zhao and B. Z. Tang, Angew. Chem., Int. Ed., 2020, 59, 8828–8832 CrossRef CAS PubMed.
- L. Viglianti, N. Xie, H. H. Y. Sung, A. A. Voityuk, N. L. C. Leung, Y. Tu, C. Baldoli, I. D. Williams, R. T. K. Kwok, J. W. Y. Lam, E. Licandro, L. Blancafort and B. Z. Tang, Angew. Chem., Int. Ed., 2020, 59, 8552–8559 CrossRef CAS PubMed.
- H. Wang, Q. Gong, G. Wang, J. Dang and F. Liu, J. Chem. Theory Comput., 2019, 15, 5440–5447 CrossRef CAS PubMed.
- Z. Zhao, P. Lu, J. W. Y. Lam, Z. Wang, C. Y. K. Chan, H. H. Y. Sung, I. D. Williams, Y. Ma and B. Z. Tang, Chem. Sci., 2011, 2, 672–675 RSC.
- J. Chen, B. Xu, X. Ouyang, B. Z. Tang and Y. Cao, J. Phys. Chem. A, 2004, 108, 7522–7526 CrossRef CAS.
- W. Chi, J. Dai, C. Yan, D. Tan, Z. Guo and X. Liu, J. Mater. Chem. C, 2023, 11, 10205–10214 RSC.
- Y. Chen, J. W. Y. Lam, R. T. K. Kwok, B. Liu and B. Z. Tang, Mater. Horiz., 2019, 6, 428–433 RSC.
- Q. Li and L. Blancafort, Chem. Commun., 2013, 49, 5966–5968 RSC.
- R. Crespo-Otero, Q. Li and L. Blancafort, Chem. – Asian J., 2019, 14, 700–714 CrossRef CAS PubMed.
- W. L. Ding, X. L. Peng, G. L. Cui, Z. S. Li, L. Blancafort and Q. S. Li, ChemPhotoChem, 2019, 3, 814–824 CrossRef CAS.
- J. Li, Y. Qian, L. Xie, Y. Yi, W. Li and W. Huang, J. Phys. Chem. C, 2015, 119, 2133–2141 CrossRef CAS.
- S. A. A. Abedi, W. Chi, D. Tan, T. Shen, C. Wang, E. C. X. Ang, C.-H. Tan, F. Anariba and X. Liu, J. Phys. Chem. A, 2021, 125, 8397–8403 CrossRef CAS PubMed.
- Y. Tu, J. Liu, H. Zhang, Q. Peng, J. W. Y. Lam and B. Z. Tang, Angew. Chem., Int. Ed., 2019, 58, 14911–14914 CrossRef CAS PubMed.
- Q. Qi, L. Huang, R. Yang, J. Li, Q. Qiao, B. Xu, W. Tian, X. Liu and Z. Xu, Chem. Commun., 2019, 55, 1446–1449 RSC.
- J. Mei, N. L. C. Leung, R. T. K. Kwok, J. W. Y. Lam and B. Z. Tang, Chem. Rev., 2015, 115, 11718–11940 CrossRef CAS PubMed.
- R. T. K. Kwok, C. W. T. Leung, J. W. Y. Lam and B. Z. Tang, Chem. Soc. Rev., 2015, 44, 4228–4238 RSC.
- C. W. T. Leung, Y. Hong, S. Chen, E. Zhao, J. W. Y. Lam and B. Z. Tang, J. Am. Chem. Soc., 2013, 135, 62–65 CrossRef CAS PubMed.
- F. Ren, P. Liu, Y. Gao, J. Shi, B. Tong, Z. Cai and Y. Dong, Mater. Chem. Front., 2019, 3, 57–63 RSC.
- H. Wu, X. Guo, C. Yu, W.-Y. Wong, E. Hao and L. Jiao, Dyes Pigm., 2020, 176, 108209 CrossRef CAS.
- J. Wang, X. Gu, P. Zhang, X. Huang, X. Zheng, M. Chen, H. Feng, R. T. K. Kwok, J. W. Y. Lam and B. Z. Tang, J. Am. Chem. Soc., 2017, 139, 16974–16979 CrossRef CAS PubMed.
- S. M. Sayed, H. R. Jia, Y. W. Jiang, Y. X. Zhu, L. Ma, F. Yin, I. Hussain, A. Khan, Q. Ma, F. G. Wu and X. Lu, J. Mater. Chem. B, 2021, 9, 4303–4308 RSC.
- L. Shi, Y. H. Liu, K. Li, A. Sharma, K. K. Yu, M. S. Ji, L. L. Li, Q. Zhou, H. Zhang, J. S. Kim and X. Q. Yu, Angew. Chem., Int. Ed., 2020, 59, 9962–9966 CrossRef CAS PubMed.
- S. Xie, A. Y. H. Wong, S. Chen and B. Z. Tang, Chem. – Eur. J., 2019, 25, 5824–5847 Search PubMed.
- H. Shi, J. Liu, J. Geng, B. Z. Tang and B. Liu, J. Am. Chem. Soc., 2012, 134, 9569–9572 Search PubMed.
- L. Peng, M. Gao, X. Cai, R. Zhang, K. Li, G. Feng, A. Tong and B. Liu, J. Mater. Chem. B, 2015, 3, 9168–9172 Search PubMed.
- M. Gao, Q. Hu, G. Feng, B. Z. Tang and B. Liu, J. Mater. Chem. B, 2014, 2, 3438–3442 RSC.
- H. W. Liu, K. Li, X. X. Hu, L. Zhu, Q. Rong, Y. Liu, X. B. Zhang, J. Hasserodt, F. L. Qu and W. Tan, Angew. Chem., Int. Ed., 2017, 56, 11788–11792 Search PubMed.
- H. Bai, W. He, J. H. C. Chau, Z. Zheng, R. T. K. Kwok, J. W. Y. Lam and B. Z. Tang, Biomaterials, 2021, 268, 120598 Search PubMed.
- G. Feng, Y. Yuan, H. Fang, R. Zhang, B. Xing, G. Zhang, D. Zhang and B. Liu, Chem. Commun., 2015, 51, 12490–12493 RSC.
- D. Wang, H. Su, R. T. K. Kwok, X. Hu, H. Zou, Q. Luo, M. M. S. Lee, W. Xu, J. W. Y. Lam and B. Z. Tang, Chem. Sci., 2018, 9, 3685–3693 Search PubMed.
- F. Y. Zhu, L. J. Mei, R. Tian, C. Li, Y. L. Wang, S. L. Xiang, M. Q. Zhu and B. Z. Tang, Chem. Soc. Rev., 2024, 53, 3350–3383 RSC.
- S. Bachollet, Y. Shpinov, F. Broch, H. Benaissa, A. Gautier, N. Pietrancosta, J. M. Mallet and B. Dumat, Org. Biomol. Chem., 2022, 20, 3619–3628 Search PubMed.
- Z. Chen, C. Jing, S. S. Gallagher, M. P. Sheetz and V. W. Cornish, J. Am. Chem. Soc., 2012, 134, 13692–13699 Search PubMed.
- S. Segawa, X. Ou, T. Shen, T. Ryu, Y. Ishii, H. H. Y. Sung, I. D. Williams, R. T. K. Kwok, K. Onda, K. Miyata, X. He, X. Liu and B. Z. Tang, Aggregate, 2024, 5, e499 Search PubMed.
- T. Ku, J. Swaney, J.-Y. Park, A. Albanese, E. Murray, J. H. Cho, Y.-G. Park, V. Mangena, J. Chen and K. Chung, Nat. Biotechnol., 2016, 34, 973–981 CrossRef CAS PubMed.
- T. Zhang, X. Chen, C. Yuan, X. Pang, P. Shangguan, Y. Liu, L. Han, J. Sun, J. W. Y. Lam, Y. Liu, J. Wang, B. Shi and B. Z. Tang, Angew. Chem., Int. Ed., 2023, 62, e202211550 CrossRef CAS PubMed.
- C. Yan, J. Dai, Y. Yao, W. Fu, H. Tian, W. H. Zhu and Z. Guo, Nat. Protoc., 2023, 18, 1316–1336 CrossRef CAS PubMed.
- Z. Ye, C. Qiu, Z. Liu, H. Wen, F. Gao, Y. Zhang, W. J. Wang, J. Zhang, Z. Qiu, W. Zhang, P. Alam, Z. Zhao and B. Z. Tang, J. Am. Chem. Soc., 2025, 147, 28442–28453 CrossRef CAS PubMed.
- T. Zang, Y. Xie, S. Su, F. Liu, Q. Chen, J. Jing, R. Zhang, G. Niu and X. Zhang, Angew. Chem., Int. Ed., 2020, 59, 10003–10007 CrossRef CAS PubMed.
- W. Liu, R. Li, F. Deng, C. Yan, X. Zhou, L. Miao, X. Li and Z. Xu, ACS Appl. Bio. Mater., 2021, 4, 2104–2112 CrossRef CAS PubMed.
- E. Zhao, Y. Chen, H. Wang, S. Chen, J. W. Y. Lam, C. W. T. Leung, Y. Hong and B. Z. Tang, ACS Appl. Mater. Interfaces, 2015, 7, 7180–7188 CrossRef CAS PubMed.
- Y. Wang, X. Pan, T. Dai, L. Wang, H. Shi, H. Wang and Z. Chen, Biomater. Sci., 2022, 10, 4866–4875 RSC.
- X. Shi, C. Y. Y. Yu, H. Su, R. T. K. Kwok, M. Jiang, Z. He, J. W. Y. Lam and B. Z. Tang, Chem. Sci., 2017, 8, 7014–7024 RSC.
- R. Jia, H. Xu, C. Wang, L. Su, J. Jing, S. Xu, Y. Zhou, W. Sun, J. Song, X. Chen and H. Chen, J. Nanobiotechnol., 2021, 19, 419 Search PubMed.
- X. Liu, J. M. Cole and K. S. Low, J. Phys. Chem. C, 2013, 117, 14723–14730 CrossRef CAS.
- Y. Shao, R. Huang, G. Jiang, Q. Shi, H. Wei, G. Wang, W. Chi, H. Peng, X. Liu and Y. Fang, Chem. Mater., 2024, 36, 3223–3232 CrossRef CAS.
- X. Liu, J. M. Cole, P. C. Y. Chow, L. Zhang, Y. Tan and T. Zhao, J. Phys. Chem. C, 2014, 118, 13042–13051 CrossRef CAS.
- X. Xu, H. Liu, Q. Sun, X. Fu, R. Huang and Y. Fang, J. Photochem. Photobiol., A, 2018, 355, 425–430 CrossRef CAS.
- K. Mizusawa, Y. Takaoka and I. Hamachi, J. Am. Chem. Soc., 2012, 134, 13386–13395 Search PubMed.
- D. Zhai, W. Xu, L. Zhang and Y.-T. Chang, Chem. Soc. Rev., 2014, 43, 2402–2411 RSC.
- K. T. Hong, S. B. Park, D. P. Murale, J. H. Lee, J. Hwang, W. Y. Jang and J. S. Lee, Angew. Chem., Int. Ed., 2024, 63, e202405525 Search PubMed.
- F. Bouhedda, K. T. Fam, M. Collot, A. Autour, S. Marzi, A. Klymchenko and M. Ryckelynck, Nat. Chem. Biol., 2020, 16, 69–76 CrossRef CAS PubMed.
- I. O. Aparin, R. Yan, R. Pelletier, A. A. Choi, D. I. Danylchuk, K. Xu and A. S. Klymchenko, J. Am. Chem. Soc., 2022, 144, 18043–18053 CrossRef CAS PubMed.
- I. A. Karpenko, M. Collot, L. Richert, C. Valencia, P. Villa, Y. Mely, M. Hibert, D. Bonnet and A. S. Klymchenko, J. Am. Chem. Soc., 2015, 137, 405–412 Search PubMed.
- K. Ohira, Y. Sato and S. Nishizawa, ACS Sens., 2023, 8, 522–526 Search PubMed.
- Q. Qiao, W. Liu, W. Chi, J. Chen, W. Zhou, N. Xu, J. Li, X. Fang, Y. Tao, Y. Zhang, Y. Chen, L. Miao, X. Liu and Z. Xu, Aggregate, 2022, 4, e258 Search PubMed.
- Y. Zhuang, T. Yin, J. Li, Y. Zang and X. Li, Anal. Chem., 2024, 96, 9034–9042 Search PubMed.
- A. Pandith, Y. Luo, Y. Jang, J. Bae and Y. Kim, Angew. Chem., Int. Ed., 2023, 62, e202215049 CrossRef CAS PubMed.
- C. Yan, L. Miao, Y. Zhang, X. Zhou, G. Wang, Y. Li, Q. Qiao and Z. Xu, Sens. Actuators, B, 2023, 386, 133731 Search PubMed.
- K. Mizusawa, Y. Ishida, Y. Takaoka, M. Miyagawa, S. Tsukiji and I. Hamachi, J. Am. Chem. Soc., 2010, 132, 7291–7293 CrossRef CAS PubMed.
- Z. Zhan, J. Zhang, T. Shen, J. Li, L. Chai, L. Pan, H. Yang, T. Liu, X. Liu and W. Mao, J. Am. Chem. Soc., 2025, 147, 22932–22943 CrossRef CAS PubMed.
- K. Basran and N. W. Luedtke, Chem. Commun., 2025, 61, 4172–4175 RSC.
- C. Yan, Z. Guo, Y. Liu, P. Shi, H. Tian and W. H. Zhu, Chem. Sci., 2018, 9, 6176–6182 RSC.
- J. Li, X. Du, X. Zhou and J. Yoon, Adv. Healthcare Mater., 2023, 12, e2301022 CrossRef PubMed.
- H. Huang, Q. Liu, J. H. Zhu, Y. Tong, D. Huang, D. Zhou, S. Long, L. Wang, M. Li, X. Chen and X. Peng, Aggregate, 2025, 6, e706 CrossRef CAS.
- K. M. Tewari and I. M. Eggleston, Photochem. Photobiol. Sci., 2018, 17, 1553–1572 Search PubMed.
- A. L. Vahrmeijer, M. Hutteman, J. R. Van Der Vorst, C. J. Van De Velde and J. V. Frangioni, Nat. Rev. Clin. Oncol., 2013, 10, 507–518 CrossRef CAS PubMed.
- P. A. Valdés, F. Leblond, V. L. Jacobs, B. C. Wilson, K. D. Paulsen and D. W. Roberts, Sci. Rep., 2012, 2, 798 Search PubMed.
- W. Stummer, U. Pichlmeier, T. Meinel, O. D. Wiestler, F. Zanella and H.-J. Reulen, Lancet Oncol., 2006, 7, 392–401 CrossRef CAS PubMed.
- J. C. Kennedy, R. H. Pottier and D. C. Pross, J. Photochem. Photobiol., B, 1990, 6, 143–148 CrossRef CAS PubMed.
- I. Yapici, K. S. S. Lee, T. Berbasova, M. Nosrati, X. Jia, C. Vasileiou, W. Wang, E. M. Santos, J. H. Geiger and B. Borhan, J. Am. Chem. Soc., 2015, 137, 1073–1080 CrossRef CAS PubMed.
- M. E. Cooper, S. Gregory, E. Adie and S. Kalinka, J. Fluoresc., 2002, 12, 425–429 CrossRef CAS.
- A. Grover, B. F. Schmidt, R. D. Salter, S. C. Watkins, A. S. Waggoner and M. P. Bruchez, Angew. Chem., Int. Ed., 2012, 51, 4838–4842 CrossRef CAS PubMed.
- S. M. Usama, F. Inagaki, H. Kobayashi and M. J. Schnermann, J. Am. Chem. Soc., 2021, 143, 5674–5679 CrossRef CAS PubMed.
- S. M. Usama, S. C. Marker, D. R. Caldwell, N. L. Patel, Y. Feng, J. D. Kalen, B. Croix and M. J. Schnermann, J. Am. Chem. Soc., 2021, 143, 21667–21675 CrossRef CAS PubMed.
- S. H. Lee, T.-J. Lee, S. Sarkar, H. Cho, Q. P. N. Nhu and Y.-T. Chang, Angew. Chem., Int. Ed., 2025, 64, e202413121 CrossRef CAS PubMed.
- X. Xie, Z. Lu, Y. Kong, C. Jin, Z. Sun, X. Yang, Z. Li, F. Qu and W. Tan, Angew. Chem., Int. Ed., 2026, 65, e20221 CrossRef CAS PubMed.
- B. Zhang, G. Feng, S. Wang and X. Zhang, Dyes Pigm., 2018, 149, 356–362 CrossRef CAS.
- X. Ji, K. Ji, V. Chittavong, R. E. Aghoghovbia, M. Zhu and B. Wang, J. Org. Chem., 2017, 82, 1471–1476 CrossRef CAS PubMed.
- L. Wang, Y. Huang, J. Wang, Y. Jiang, B.-P. Jiang, H. Chen, H. Liang and X.-C. Shen, J. Am. Chem. Soc., 2025, 147, 6707–6716 CrossRef CAS PubMed.
- H. Wu, S. C. Alexander, S. Jin and N. K. Devaraj, J. Am. Chem. Soc., 2016, 138, 11429–11432 CrossRef CAS PubMed.
- S. Shen, D. Hong, X. Qian and X. Luo, Anal. Chem., 2025, 97, 3696–3703 CrossRef CAS PubMed.
- W. Chi, Q. Qiao, R. Lee, W. Liu, Y. S. Teo, D. Gu, M. J. Lang, Y.-T. Chang, Z. Xu and X. Liu, Angew. Chem., Int. Ed., 2019, 58, 7073–7077 CrossRef CAS PubMed.
- W. Liu, J. Chen, Q. Qiao, X. Liu and Z. Xu, Chin. Chem. Lett., 2022, 33, 4943–4947 CrossRef CAS.
- X. Yu, X. Liu, H. Sun, T. Shen, Y. Deng, H. Ren, P. Zou, Y. Zheng, P. Xiao, Q. Gong, M. Qin, X. Liu and H. Wu, Angew. Chem., Int. Ed., 2025, 64, e202511705 CrossRef CAS PubMed.
- X. Liu, W. Chi, Q. Qiao, S. V. Kokate, E. P. Cabrera, Z. Xu, X. Liu and Y.-T. Chang, ACS Sens., 2020, 5, 731–739 CrossRef CAS PubMed.
- L. Mendive-Tapia, D. Mendive-Tapia, C. Zhao, D. Gordon, S. Benson, M. J. Bromley, W. Wang, J. Wu, A. Kopp, L. Ackermann and M. Vendrell, Angew. Chem., Int. Ed., 2022, 61, e202117218 CrossRef CAS PubMed.
- C. Wang, X. Song, L. Chen and Y. Xiao, Biosens. Bioelectron., 2017, 91, 313–320 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |