Investigating PVC polymer–plasticizer interactions with atomistic MD simulations and potential of mean force calculations
Sai Athmeeya G.
Shet
 a,
Jan-Michael Y.
Carrillo
a,
Jan-Michael Y.
Carrillo
 *b,
Jacek
Jakowski
*b,
Jacek
Jakowski
 c,
S. S. R. Kumar
Challa
d and
V. N. Ravi Kishore
Vutukuri
c,
S. S. R. Kumar
Challa
d and
V. N. Ravi Kishore
Vutukuri
 *a
*a
aDepartment of Chemistry, Sri Sathya Sai Institute of Higher Learning, Puttaparthi, Andhra Pradesh-515134, India. E-mail: vnrkishorevutukuri@sssihl.edu.in
bCenter for Nanophase Materials Sciences, Oak Ridge National Laboratory, Oak Ridge, Tennessee 37831, USA. E-mail: carrillojy@ornl.gov
cComputational Sciences and Engineering Division, Oak Ridge National Laboratory, Oak Ridge, Tennessee 37831, USA
dXheme Inc., 100 Carlson Ave, Newton, Massachusetts 02459, USA
First published on 2nd March 2026
Abstract
Atomistic molecular dynamics (MD) simulations coupled with potential of mean force (PMF) calculations were employed to investigate the interactions between PVC polymer chains containing 6–20 repeating units and various plasticizers, with the goal of identifying potential replacements for the toxic plasticizer di(2-ethylhexyl)phthalate (DEHP) used in blood bags. The selected plasticizers belong to various chemical families, such as orthophthalates, citrates, adipates, and the terephthalate DEHT. Both the polymer and the plasticizers lack ionizable groups, and their interactions are primarily governed by van der Waals and electrostatic forces. A model correlating PMF profiles with interaction forces was developed and validated across polyvinyl chloride (PVC) polymers of different lengths and all investigated plasticizers. This model provides insight into how structural variations in plasticizers influence their respective PMF values. The study ranks the investigated plasticizers based on binding affinity, identifying TOTM (tris(2-ethylhexyl) trimellitate, TEHTM), BTHC (butyryl trihexyl citrate, Citroflex B-6), ATHC (acetyl trihexyl citrate, Citroflex A-6, CA-6), and DEHT (bis(2-ethylhexyl)terephthalate, DOTP/DEHTP) as promising alternatives for further investigation. This comprehensive study encompasses PVC polymers of four different lengths and 14 plasticizers, with all data averaged over 30 independent simulations. The approach provides a deeper understanding of molecular interactions, enabling the tailoring of polymer–plasticizer systems for diverse applications, including extractables and leachables, with relevance spanning materials science to biomedical engineering, and also serves as a basis for developing coarse-grained simulation protocols. Overall, this study provides valuable insights for designing safer and more efficient plasticizer substitutes and is well supported by other studies.
1 Introduction
Plasticizers play a pivotal role in modifying the mechanical and thermal properties of polymers, particularly in PVC based blood bag formulations, where they constitute one-third of the additive market.1 Orthophthalates, notably DEHP, dominate this landscape, comprising up to 85% of the total plasticizer market.2 However, escalating environmental and health concerns—especially the endocrine-disrupting effects associated with orthophthalates—have led to regulatory restrictions and a pressing need for safer alternatives.3–5 Regulatory authorities have responded with tighter restrictions, especially in sensitive applications like medical devices (particularly blood bags) and children's toys.4,6,7 The European Union and the state of California have banned the use of blood bags that contain DEHP plasticizer.8–12 International regulations have outlined stringent limits and bans on certain derivatives, making the urgent exploration of safer alternatives imperative.13–15 Various studies have delved into PVC–plasticizer interactions, employing experimental techniques such as FTIR spectroscopy and differential thermal analysis.16,17The PVC polymer and plasticizer systems have been computationally investigated using full atomistic MD simulations by various research groups, with the aim of estimating thermodynamic, mechanical, and structure–property relationships. Zhou et al. investigated the density of the PVC–DEHP system with different force fields, such as COMPASS, OPLS, and OPLS-AA, and demonstrated that the OPLS-AA force field can accurately reproduce experimental and simulated properties by calculating both average and local shifts in glass transition temperatures of PVC and DEHP.18 Li et al. studied isoorthophthalates14 and various other plasticizers19 to gain molecular insights for designing plasticizers, and also developed a simulation protocol whose thermodynamic compatibility and plasticization efficacy were consistent with experimental results. For isoorthophthalates with the same side chain configuration, an increase in the length of the side chains leads to reduced compatibility and plasticization. Sahnoune et al. investigated the sorption of drug molecules on plasticized PVC, finding that the adsorption of diazepam and paracetamol was thermodynamically favorable and driven by van der Waals interactions. They used potential of mean force calculations to analyze the energy contributions of water desorption and drug dehydration.20 In another atomistic MD study by the same group, supported by experimental data, they studied leaching and migration rates of different plasticizers in different solvents and found that DINCH exhibited the highest leaching ratio, while TOTM showed the lowest in ethanol.21 Jagarlapudi et al. focused on bio-based plasticizers to replace DEHP, calculating properties such as tensile strength, mechanical performance, and diffusion in PVC–plasticizer systems, while assessing glass transition temperature, Young's modulus, shear modulus, fractional free volume, and diffusion. Their in silico study concluded that the performance of DBS (dibutyl sebacate) and DHS (diheptyl succinate) in PVC (20 monomers) was comparable to or better than that of DEHP.22 Computational methods like PMF calculations,23,24 coupled with tools like the weighted histogram analysis method (WHAM),25,26 aid in estimating the free energy of binding and, consequently, mutual solubility and compatibility.14
Molecular structures and interactions significantly influence plasticization efficiency, with parameters such as polarity, chemical groups, chain length, molecular weight, and dielectric constant identified as crucial factors affecting plasticizer–polymer compatibility.27 Empirical studies of diverse plasticizers aim to identify alternatives to DEHP, contributing to sustainable PVC–plasticizer formulations. For example, the presence of aromatic rings, ester groups and building blocks such as aliphatic chains play a pivotal role in optimizing plasticizer stability and compatibility.22,27 The intricate balance between aliphatic chain length, branching, and molecular weight emerges as a key consideration in designing an ideal plasticizer.
The comprehensive analysis presented in this article focuses on PVC polymers and aims to enhance the understanding of PVC–polymer and plasticizer interactions, as well as to provide insights for the development of improved plasticizers. Ways to design more sustainable alternative plasticizers are explored, addressing regulatory imperatives and aligning with the need to protect human health and the environment.2,28–35 In this article, the interaction strength between a single molecule of PVC polymer and a single molecule of plasticizer is studied through fully atomistic MD simulations coupled with PMF calculations for PVC polymers with lengths ranging from 6 to 20 repeating units and 14 different plasticizers that have the potential to replace DEHP are studied. Some of these plasticizers, such as BTHC (Citroflex B-6), TOTM (TEHTM), and DEHT (DOTP/DEHTP), which are already used commercially in non-blood bag applications, are considered promising and have the potential to replace DEHP.6,19,36–39
This study explores the structural features and functional groups of various plasticizers, including phthalates, which are characterized by a benzene ring with two ester groups; adipates, which contain a linear six-carbon aliphatic chain with two ester groups; citrates, which feature a central citric acid core with three esterified hydroxyl groups; and trimellitates, which consist of a benzene ring with three ester functional groups. See Table S1 of the SI for the chemical structures of the plasticizers and their categories. In classical MD simulations, different approaches can be used to computationally determine the binding energies between two molecules, such as steered MD simulations40–44 and umbrella sampling.45 This work focuses on calculating the binding energies by varying the distance between the centers of mass of the polymer and plasticizer, performing MD simulations, and determining PMF values through umbrella sampling. By analyzing normalized PMF (N-PMF) plots, this study aims to propose a model for interpreting simulation data to better understand the interactions between PVC polymers and plasticizers. The model will be validated to contribute to the design of environmentally sustainable plasticizer formulations.
The analysis was conducted on 14 different plasticizers, carefully selected from the literature as promising candidates to replace DEHP (see Table S1 of the SI). The goal is to examine the interactions between PVC and plasticizers and identify the plasticizer with the highest binding affinity. A stronger affinity between PVC and the plasticizer reduces leaching, which is advantageous for various applications. The length of the polymer chain was restricted to 20 monomer units because beyond this length, our studies showed increased chain folding and intrachain interactions that complicated the analysis. The current study is intended to be extended to longer polymer chain lengths in future work. The primary objective here is to understand the intermolecular interactions between a short polymer chain and a plasticizer, which could provide useful information for the development of coarse-grained simulations involving the polymer–plasticizer system. Furthermore, the Kuhn length of the PVC polymer is approximately 10 monomer units.46 For these reasons, the analysis presented in this manuscript has been restricted to PVC polymers containing 20 monomer units.
2 Simulation details
Molecular structures of PVC polymers of various lengths and plasticizer molecules were built using Avogadro software.47 AmberTools2148 was used to assign all-atom parameters to the molecules based on the general Amber force field (GAFF2).49 In the absence of ionizable groups and strong chemical bonding forces, the separation distances tend to be large, where both van der Waals and electrostatic interactions dominate. Due to the long-range nature of electrostatics, the use of classical force fields with their static point charge approximation is particularly well-suited to model these regions.50 This assumption was verified by performing DFT simulations, as discussed later in the manuscript. The functional form for the bond angles, dihedral angles, and non-bonded pairwise interactions of GAFF49,51 is given by: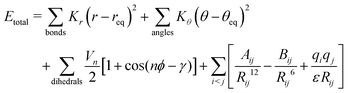 | (1) |
The initial molecular packing of the composite system (PVC and plasticizers) was generated using Packmol,52,53 and molecular dynamics simulations were performed using LAMMPS54–56 in a cubic simulation box with dimensions 128 × 128 × 128 Å3, under periodic boundary conditions. The Störmer-Verlet (Velocity-Verlet) time-stepping algorithm57 was used with a timestep of 1 fs for all calculations. A 12-6 Lennard-Jones potential with a cut-off radius of 10 Å was employed for short-range interactions. The typical cutoff for short-range Lennard-Jones nonbonded interactions in the AMBER force field is 10 Å, which is consistent with the GAFF2 parametrization for this force field.49 The Lorentz–Berthelot mixing rules58,59 were used to estimate intermolecular pair potentials. Long-range electrostatic interactions were handled using the particle–particle–particle–mesh (PPPM) method60,61 with a near-field cut-off radius of 10 Å. Initial equilibration was performed using the LAMMPS nve/limit command to relax overlapping atoms, followed by a Langevin thermostat that ran for 0.5 ns. For the subsequent potential of mean force (PMF) calculations investigating the interaction between a single polymer chain and a single plasticizer molecule, the polymer was restrained at its center of mass. Specifically, its translational motion was constrained while its rotational degrees of freedom were preserved. This restraint was implemented using a harmonic potential with a force constant of 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 kcal mol−1 Å−2. Additionally, a harmonic biasing potential with a force constant of 20 kcal mol−1 Å−2 was applied between the centers of mass of the polymer and the plasticizer. The 20 kcal mol−1 Å−2 was selected after systematically evaluating several force constants (1, 5, 20, and 100 kcal mol−1 Å−2) and choosing the one that provided optimal sampling efficiency, balancing force constant magnitude, simulation time, and integration time step. The corresponding histogram plot is shown in Fig. S3 in the SI, where the overlapping histograms indicate adequate sampling, enabling construction of the PMF using WHAM, as discussed below. Further equilibration was conducted using the Canonical (NVT) ensemble, running for 50 ps with a Nosé–Hoover thermostat and a 100 fs damping parameter.
000 kcal mol−1 Å−2. Additionally, a harmonic biasing potential with a force constant of 20 kcal mol−1 Å−2 was applied between the centers of mass of the polymer and the plasticizer. The 20 kcal mol−1 Å−2 was selected after systematically evaluating several force constants (1, 5, 20, and 100 kcal mol−1 Å−2) and choosing the one that provided optimal sampling efficiency, balancing force constant magnitude, simulation time, and integration time step. The corresponding histogram plot is shown in Fig. S3 in the SI, where the overlapping histograms indicate adequate sampling, enabling construction of the PMF using WHAM, as discussed below. Further equilibration was conducted using the Canonical (NVT) ensemble, running for 50 ps with a Nosé–Hoover thermostat and a 100 fs damping parameter.
The potential of mean force W(ξ) is given by,
 | (2) |
 as a function that depends on several degrees of freedom in the system, is expressed as:
as a function that depends on several degrees of freedom in the system, is expressed as: | (3) |
The PMF energy profiles were generated using the harmonic biasing umbrella sampling method, implemented through the weighted histogram analysis method (WHAM) algorithm.25,26 This method enables systematic sampling of configurations along a predefined coordinate, providing detailed insights into the interactions between the polymer and the plasticizer. The selected reaction coordinate for this study was the distance between the centers of mass of the polymer and the plasticizer. This choice facilitated a focused examination of the intermolecular interactions between these components and will also inform future coarse-grained modelling efforts, in which the plasticizer will be represented by a few connected Lennard-Jones (LJ) particles and the PVC polymer by connected LJ beads mapped at the Kuhn segment level. All the simulation details mentioned are summarized in a flowchart given in Fig. S1 in the SI.
To obtain an unbiased free energy profile along the chosen reaction coordinate of the PVC–plasticizer pair, WHAM was used to construct the PMF from 300 reaction coordinate steps, covering a range of 0 to 30 Å with an increment of 0.1 Å in center-to-center molecular separation and a runtime of 0.1 ns per step. To ensure the robustness and reliability of the results, each system was independently simulated thirty times. This ensured adequate sampling of all possible orientations between the polymer and the plasticizer, with each simulation initialized using random velocities assigned by LAMMPS. In the planned coarse-grained representation, the interactions will correspond to an ensemble average that is radially averaged.
The intermolecular free energy dissociation values from each PMF profile were extracted and subsequently averaged. The resulting PMF profiles exhibited slightly higher standard deviations due to the inherent stochastic nature of the systems studied (i.e., the molecules are free to rotate and separate randomly from any angle). Therefore, the standard deviations for all PMF values reported in this work were calculated using the bootstrapping method.62,63 The spring constant of 20 kcal mol−1 Å−2 used for the PMF calculations was optimized to ensure that fluctuations in the distance between the centers of mass of the two molecules were centered around the target value of the reaction coordinate and that sufficient overlap occurred between simulations (see Fig. S3 in the SI). All simulations were performed in vacuum. Although polymer chain collapse is a known possibility under such conditions, no evidence of collapse was observed in this study. The PVC chains considered here consisted of 6–20 monomer units; given that the Kuhn length of PVC is approximately 10 monomers, the polymer samples had both stretched and partially folded conformations.
To benchmark the relative contributions of van der Waals and electrostatic interactions and to validate the proposed model for interpreting the simulation data, potential of mean force (PMF) calculations were performed between the PVC polymer and the plasticizer with long-range electrostatic interactions turned off. This approach isolates the contribution of van der Waals interactions to the system. In general, the total interaction energy is expected to arise primarily from van der Waals and electrostatic interactions, with additional contributions from entropy. The conformational entropy contribution to the free energy of the PVC polymer–plasticizer system was further assessed through analysis of the dihedral angle distributions and the radii of gyration (Rg) for both the benchmark system and the system excluding electrostatic interactions. The entropic contribution to the free energy was quantified by averaging five independent PMF calculations performed at different temperatures (280 K to 360 K) and fitting the resulting free-energy data to a linear dependence on temperature. The entropy (ΔS) and enthalpy (ΔH) were estimated from the slope and y-intercept of the fitted line, respectively.
3 Results and discussion
PMF values were computed for PVC–plasticizer systems, with plasticizers considered as potential replacements for DEHP. The free energy barrier, defined as the difference between the lowest point and the saturation region of the PMF curve, was obtained from thirty independent simulations and averaged. A representative PMF curve is shown in Fig. S4, where the reaction coordinate corresponds to the separation distance between the plasticizer and polymer centers of mass. The lowest PMF value (set to zero) marks the starting configuration, and the curve rises incrementally as the molecules move apart, reaching a plateau at the dissociation free energy.The study was limited to PVC polymers of 6–20 monomer units, focusing on PVC segment–plasticizer interactions. This range reflects commercial relevance, as plasticizers (typically ∼30% loading7) are surrounded by Kuhn segments of PVC, ∼2.6 nm in length (≈10 monomer units).46 The simplified 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 interaction model effectively isolates PVC–plasticizer interactions while minimizing intramolecular PVC–PVC and plasticizer–plasticizer contributions. This framework also provides a basis for determining interaction energetics in the planned coarse-grained molecular dynamics (MD) simulations.
1 interaction model effectively isolates PVC–plasticizer interactions while minimizing intramolecular PVC–PVC and plasticizer–plasticizer contributions. This framework also provides a basis for determining interaction energetics in the planned coarse-grained molecular dynamics (MD) simulations.
To assess robustness, thirty independent simulations were carried out for each system. Free energy of dissociation values extracted from the PMF plots showed standard deviations ranging from 0.2 to 3.1 kcal mol−1. While variability arises from random initial conformations, velocities, and separation angles, results remained consistent across simulations. The larger deviations reflect stochasticity from random sampling, highlighting the importance of averaging over an ensemble of simulations to capture orientational variability in PVC–plasticizer separation.
Various types of plasticizers, including orthophthalates, terephthalates, adipates, citrates, and others, have been investigated as potential substitutes for DEHP.6,19,36–39 These plasticizers exhibit distinctive structural and chemical characteristics. Analyzing them from a structural standpoint involves considering factors such as alkyl chain length, functional groups, and the degree of branching, among others. However, comparing PMF values across plasticizers with differing structural properties poses a challenge. To account for this chemical diversity, normalized PMF values (N-PMF) were calculated by dividing the raw PMF values by the total molar mass of the system, including both the plasticizer and the PVC polymer. This approach allows for a more meaningful comparison among plasticizers with varying molecular weights and architectures. The normalization could, in principle, be performed in several ways—by dividing by the molar mass of the polymer alone, the molar mass of the plasticizer alone, or by the sum of both. Regardless of the normalization scheme chosen, the overall conclusions of this study remain unaffected.
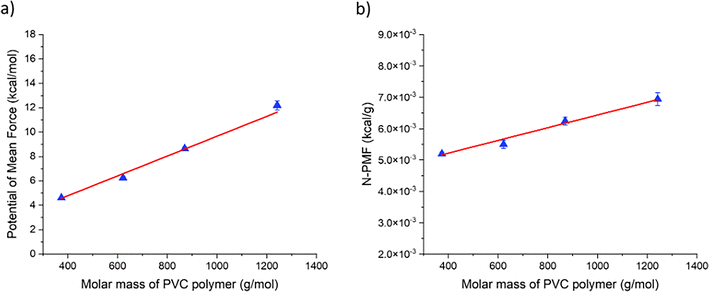
Fig. 1(a) illustrates the variation of PMF values for the interaction between the BTHC plasticizer and PVC polymer with molar masses ranging from 6 to 20 repeating units. The variation of PMF values was found to be linear, fitting well to a linear equation with a regression coefficient of 0.99. In Fig. 1(b), the N-PMF plot for the same system is shown, which also exhibited a linear relationship and could be fitted to a linear equation with a regression coefficient of 0.98. The interaction between the polymer and the plasticizer can be studied in two different ways: one involves examining the interaction of different plasticizers with a polymer of fixed length (Case I), and the other involves selecting one plasticizer and studying its interaction with polymers of different chain lengths (Case II). The plots obtained from both cases offer valuable insights into polymer–plasticizer interactions, which will be elaborated upon later. Additionally, N-PMF plots were generated, fitted with straight lines, and their slopes and y-intercepts interpreted. This analysis helps to shed light on the behavior of the system and offers insights into the underlying interaction mechanisms.
3.1 Case I: Fixed-length PVC polymers and plasticizer interaction
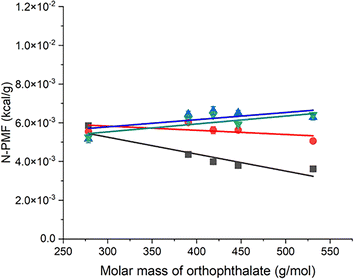
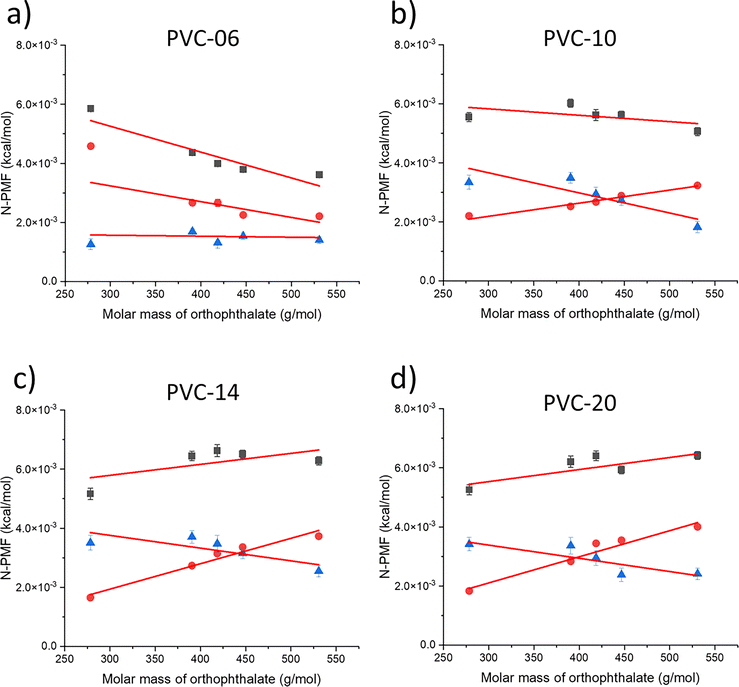
Fig. 2 depicts a plot of N-PMF versus the molar mass of orthophthalate plasticizers for PVC polymers of different lengths (6–20 repeat units). An increasing N-PMF value with increasing plasticizer molar mass was observed for PVC-20 (green inverted triangles). Since the only varying factor among the plasticizers is the length of the alkyl chain, this trend can be attributed to a larger increase in the ratio of van der Waals to electrostatic interactions. That is, the greater number of atoms comprising the plasticizer leads to a higher energy barrier. In contrast, black squares in Fig. 2 presents a plot of N-PMF versus the molar mass of orthophthalate plasticizers for PVC polymer with six repeat units (PVC-06), showing that the PMF value decreases with increasing plasticizer molar mass. PVC-06 is characterized by a significantly shorter chain length, and since only the alkyl chain length is increased in the plasticizer, the scenario shifts in favor of electrostatic interactions i.e., the intermolecular interactions become dominated by the electrostatic interactions. This shift is particularly pronounced due to the increased magnitude of polar interactions between PVC-06 and the orthophthalate plasticizers. In such instances, the variation of electrostatic interactions becomes more apparent, outweighing the variation of van der Waals forces. The interplay between alkyl chain length and the nature of functional groups in the plasticizer plays a pivotal role in dictating the prevalent interaction forces within the PVC-06 and orthophthalate plasticizer systems.
The contrasting characteristics observed in the N-PMF plots of PVC-20 and PVC-06, featuring positive and negative slopes, respectively, correspond to van der Waals and electrostatic interactions, revealing a distinct trend. Similarly, the slope of PVC-14 (blue triangles) is positive, and for PVC-10 (red circles) it is negative, and these lines lie between PVC-20 and PVC-06. These observations led to the development of a model wherein the N-PMF plot serves as a repository of vital information about the interactions within the system. According to this model, a positive slope in the plot indicates an increase in the ratio of van der Waals to electrostatic interactions between the polymer and plasticizer, while a negative slope signals a decrease in this ratio. However, an intriguing observation arises when the plot is extrapolated to zero mass. Typically, one would expect the y-intercept to approach zero, but this is not the case. This discrepancy suggests that factors beyond molar mass, such as electrostatics, come into play, indicating its nuanced role in understanding of the interactions in these systems. Thus, the y-intercept value in the N-PMF plot provides insights into the magnitude of the electrostatic interactions, as van der Waals interactions are directly related to the molar mass of the molecule.
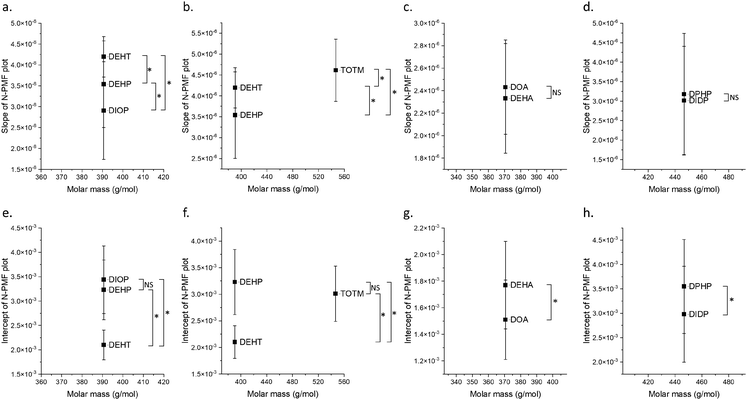
Fig. 2 (black squares) shows that for smaller PVC chain lengths, the slope is negative, and as the PVC chain length increases, the slope changes sign and becomes positive. The change in slope from negative to positive as the chain length increases establishes a direct connection between stronger van der Waals interactions and an increased slope, thereby reinforcing the proposed model. Subsequently, the slope and y-intercept values from the N-PMF plot of each polymer were plotted against the polymer molar mass, as seen in Fig. 3(a). This slope plot serves as a reliable indicator of the interactions prevailing between the PVC polymer and the plasticizer, which, in this case, is a combination of electrostatic and van der Waals forces. The difference in slope values between the points indicates the types of interactions. A higher slope value corresponds to an increased ratio of van der Waals to electrostatic interactions.
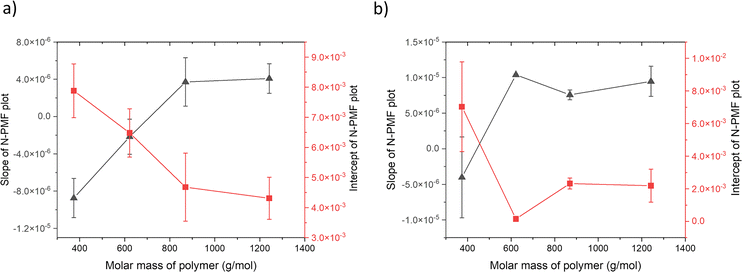 | ||
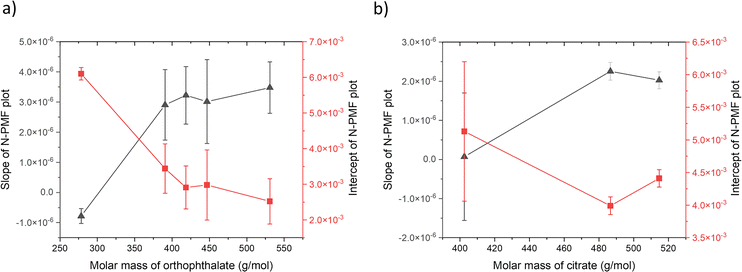
| Fig. 3 Plot of slope (primary y-axis) and intercept (secondary y-axis) values (a) obtained from linear fits of the N-PMF plots of orthophthalates shown in Fig. 2, against the molar mass of PVC polymer with various chain lengths (6–20 monomer units); (b) derived from a plot of N-PMF vs. molar mass of citrate plasticizers for PVC polymers of different chain lengths shown in Fig. S7 of the SI, plotted against the molar mass of PVC polymer. The solid lines serve as visual guides for the data points. | ||
Fig. 3(a) shows the plot of the slope and intercept of N-PMF values vs. the molar mass of PVC polymer. It reveals that the slopes and intercepts increase and decrease respectively with molar mass of the polymer. The increase in slope could be attributed to van der Waals interactions, as they increase with molar mass. Conversely, for smaller chain lengths, the slopes are negative, indicating the dominance of electrostatic interactions, as seen in the same figure. It is also observed that the intercept values decrease with increasing polymer molar mass. This trend can be explained as follows: as the molar mass of the PVC polymer increases, the magnitude of electrostatic interactions decreases due to the increase in the alkyl chain length of the orthophthalate plasticizer, leading to a reduction in the y-intercept. The plotted intercept values in Fig. 3(a) represent the relative extent of electrostatic interactions. The validity of the model is further reinforced by the correlation between the y-intercepts and 1/r scaling, as these intercepts are related to electrostatic interactions. Fig. S5 of SI shows the slope along with a guideline and Fig. S6 of SI shows the intercept along with 1/r fit to the plot using the equation y = c/x, where c = 3.51 ± 0.33, with a regression coefficient of 0.85.
In the combined plots shown in SI Fig. S8 and S9, which display the intercept and slope values obtained from linear fits of N-PMF vs. molar mass for both citrate and orthophthalate plasticizers, a distinct pattern emerges. From these figures, it is evident that the slopes for citrates are marginally higher than those for orthophthalates, suggesting a greater contribution of van der Waals interactions in citrates than in orthophthalates. This discrepancy could be attributed to differences in chemical structure, molecular geometry, multiple alkyl group substitutions on the central carbon of citrates, and higher molar mass, which lead to increased van der Waals contacts. However, the overall pattern of the plots remains consistent. This consistent trend across different plasticizers serves to validate the proposed model.
3.2 Case II: Plasticizer interaction with PVC polymers of varying chain lengths
The second set of plots (Case II), which involve N-PMF vs. molar mass of PVC polymer specific to each plasticizer, was plotted separately (see SI Fig. S10–S23) and analyzed in the same way as in the previous section. This additional analysis serves as further validation of the proposed model, interpreting the role of short-range van der Waals interactions and long-range electrostatic Coulomb interactions with the binding energy obtained from PMF calculations.3.3 PMF calculations without electrostatic interactions for orthophthalates
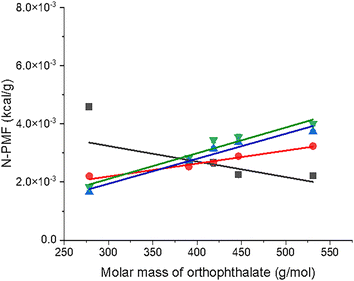
The model was further scrutinized by calculating the PMF values considering only van der Waals interactions during the MD simulations, with electrostatic interactions turned off. MD simulations were performed on the orthophthalate plasticizer and PVC polymers of different lengths, with long-range electrostatics turned off to better understand the origin of the linear behavior observed in the plots and the significance of the slopes and y-intercepts. This involved repeating the previously described simulations while following the same protocol, but with electrostatic interactions excluded. As in the earlier simulations, PMF values were averaged over 30 independent runs for each PVC and plasticizer system and plotted as a function of either the PVC molar mass or the orthophthalate plasticizer molar mass. Fig. 5 illustrates the N-PMF plot for orthophthalates interacting with PVC polymers of varying chain lengths, with electrostatics turned off. Remarkably, the slopes and y-intercepts extracted from these plots closely match those obtained when considering both van der Waals and electrostatic interactions, i.e., the total interaction energy.Fig. 6 illustrates the contributions of van der Waals, electrostatic, and total interaction energies to PVC–plasticizer interactions as a function of the molar mass of orthophthalates for PVC polymers with varying chain lengths. The electrostatic contribution was determined indirectly by subtracting the van der Waals interaction energy (obtained from simulations with electrostatic interactions disabled) from the total interaction energy (calculated from simulations including both electrostatic and van der Waals interactions). As the PVC chain length increases, the slope of the van der Waals curve becomes steeper, indicating a progressively greater contribution of van der Waals interactions to the overall interaction energy. Concurrently, a pronounced decrease in the electrostatic contribution is observed, further supporting the proposed model. Collectively, these findings highlight the increasing dominance of van der Waals interactions with increasing PVC chain length.
The subtraction of the van der Waals contribution from the total interaction energy assumes additivity of the interaction terms, as expressed by the fourth term on the right-hand side of eqn (1). Under this assumption, the procedure used to estimate the electrostatic contribution effectively cancels the entropic contribution, as demonstrated below. The conformational entropy of two systems—one including both Lennard-Jones and electrostatic interactions and the other including only Lennard-Jones interactions—was first evaluated by analyzing the distributions of backbone dihedral angles and the radii of gyration (Rg) of the polymer chain obtained during the PMF calculations. The backbone dihedral angle distributions (SI Fig. S24) and Rg values (SI Fig. S25 and S26) were found to be very similar, with only minor differences for the PVC polymer between PMF calculations performed with total interactions and those performed with van der Waals (Lennard-Jones) interactions alone.
When performing PMF calculations that include total interactions between the PVC polymer and plasticizer versus calculations that consider only van der Waals interactions, the results could be influenced by differences in the conformation of the PVC polymer chain. To assess whether the polymer conformation is consistent across calculations, additional MD simulations were performed, and the radius of gyration (Rg) was calculated for two PVC polymer and plasticizer combinations. The Rg values, an indicator of polymer conformation, were computed for the PVC-20 polymer and DIBP plasticizer, as shown in SI Fig. S25. The results revealed only minor differences, with no significant distinctions between the two sets of calculations. This suggests that the molecular configurations sampled in each simulation set are representative and consistent. A similar analysis was conducted for the DITP and PVC-20 pair, yielding a similar trend (see SI Fig. S26). DITP was selected due to its longer alkyl chain compared to other orthophthalates. The consistency in Rg values across simulations further validates our approach and confirms the reliability of our findings.
To further assess these differences, the backbone dihedral angle distributions for the two cases were analyzed using harmonic (Fourier) fitting, the Watson–Williams circular mean test,64 and the Watson–Wheeler circular distribution test.65 Harmonic (Fourier) fitting of the two datasets was carried out using an equation of the form
 | (4) |
This conclusion was further supported by a quantitative analysis in which the entropy of the PVC–plasticizer system was calculated for both interaction schemes using PMF calculations performed at multiple temperatures (280 K to 360 K). The resulting free-energy data were globally fit to a linear relationship to extract the enthalpy and entropy, as shown in SI Fig. S27. The entropy values, obtained from the slopes of the fitted lines, were identical within uncertainty for both interaction schemes (−0.030 ± 0.003 kcal mol−1 K−1), with an R2 value of 0.95.
3.4 Effect of structural variations on interaction strength across different plasticizers
The structure–property relationships of plasticizers interacting with PVC polymers of varying chain lengths were further investigated. Using the developed model, various classes of plasticizers were analyzed by plotting the slopes and y-intercept values of the plasticizers’ normalized potential of mean force (N-PMF) curves against PVC polymer chains ranging from 6 to 20 monomer units. The analysis involved comparing the slopes and y-intercepts to assess the impact of structural features, such as branching and substituent position on aromatic rings, across different families of plasticizers.3.5 Density functional theory calculations
The MD simulations discussed in this work are based on the assumption that the charge polarization effect between PVC and the plasticizers is minimal, and that the electrostatic interactions in eqn (1) can be approximated using static point charges. To verify this static point charge approximation used in classical MD simulations, density functional theory (DFT) calculations were performed, and partial atomic charges were evaluated from the corresponding electronic structure. Specifically, DFT calculations were conducted to investigate the interaction between polyvinyl chloride (PVC, 20 monomer units) and three plasticizers: butyryl trihexyl citrate (BTHC), di(2-ethylhexyl) phthalate (DEHP), and tris(2-ethylhexyl) trimellitate (TOTM). Additionally, the changes in the charge distributions of these systems as a function of the reaction coordinate were explored. Significant changes in atomic charge distribution as a function of the reaction coordinate may indicate non-negligible polarization, necessitating the inclusion of polarization effects in molecular dynamics force fields.The DFT calculations were performed using two range-separated hybrid functionals, CAM-B3LYP68 and ωB97X-D,69 along with the 6-31+G* basis set. Both functionals included Grimme's D3 dispersion correction to account for weak interactions, ensuring accurate treatment of weak interactions and long-range electrostatic effects.70–72 The input geometries for PVC–plasticizer complexes were obtained from molecular dynamics simulations at selected reaction coordinates. Single-point DFT energy calculations were performed without geometry optimization to retain the relative orientations and distances between PVC and the plasticizers as obtained from the molecular dynamics snapshots. Following the single-point DFT calculations, charge distributions were analyzed using Mulliken population analysis and the CHelpG (Charges from Electrostatic Potentials using a Grid) method with the Lebedev grid, using 50 angular points per heavy atom as well as hydrogen, and with radial spacing set to 0.5 Å. This approach parallels the Merz–Kollman methodology, which fits the ESP charges to concentric atomic spheres scaled by van der Waals radii to derive atom-centered charges.73 The results provide insights into the polarization and electrostatic interactions governing PVC–plasticizer binding.
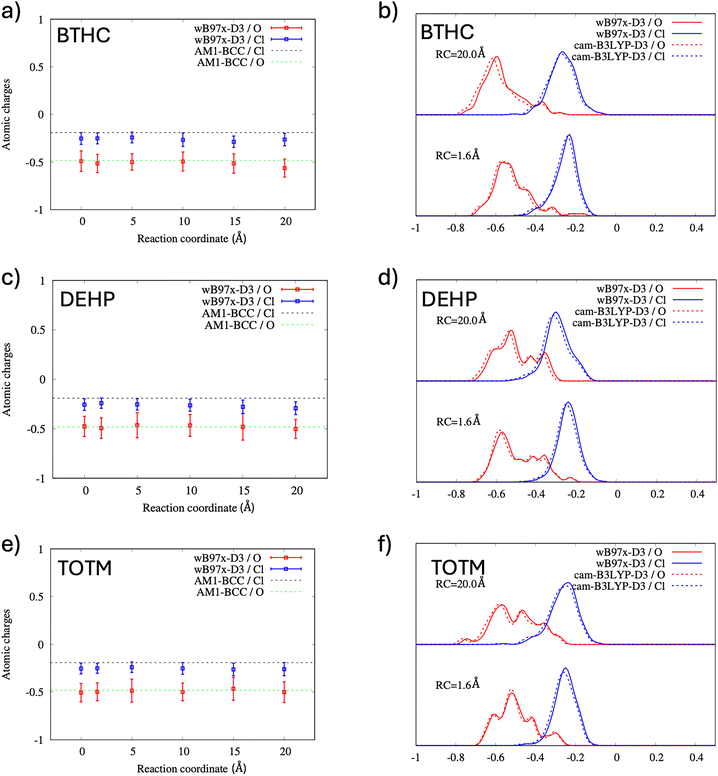
Charge distributions were evaluated for six reaction coordinate values: RC = 0, 1.6, 5.0, 10.0, 15.0, and 20.0 Å. Here, RC = 1.6 Å corresponds to the bound PVC–plasticizer system (approximately the minimum of the free energy from the PMF calculation), while RC = 20.0 Å represents the fully dissociated system. To account for statistical uncertainty, 11 randomly selected MD snapshots were considered for each RC value. The molecular structure of PVC interacting with plasticizers consisted of 208, 188, and 215 atoms for the BTHC, DEHP, and TOTM systems, respectively. In the charge analysis, only chloride atoms from PVC (20 Cl atoms) and oxygen atoms from the plasticizers (8, 4, and 6 O atoms for BTHC, DEHP, and TOTM, respectively) were analyzed. This choice reflects that Cl and ester O atoms carry the largest partial charges dominating any RC-dependent electrostatic changes relevant to the fixed-charge force field used in the MD simulations. Additional ωB97X-D3/CHELPG results including the PVC carbon adjacent to Cl and the plasticizer ester carbon, together with group-summed -CCl and ester charges, are reported in the SI Fig. S28, confirming that the changes in the mean charges with RC are small and show no systematic trend.
Fig. 8(a), (c) and (e) presents the average DFT charges on Cl and O atoms, along with the corresponding standard deviations (shown as error bars), calculated across all structures as a function of RC. The results indicate that the average DFT charges on Cl and O remain constant and have comparable values to those used in the MD simulations (see dashed lines in Fig. 8(a), (c) and (e)), which were derived from AM1-BCC calculations. This finding strongly suggests that polarization effects are minimal, supporting the use of non-polarizable force fields in molecular dynamics simulations. To further investigate polarization effects, charge distributions were calculated by applying a Gaussian convolution to the atomic charges of all Cl and O atoms from the 11 snapshots for RC = 1.6 Å and RC = 20 Å. Fig. 8(b), (d) and (f) shows that the mean charge on Cl (calculated over all Cl atoms from PVC-20 and across 11 random MD conformations) is independent of the plasticizer, with the charge distribution resembling a Gaussian function that changes minimally with the plasticizer or RC value. This observation supports the conclusion that Cl charges are minimally affected by polarization. For O atoms, the charge distribution varies significantly between BTHC, DEHP, and TOTM, consistent with their differing chemical structures. However, the changes in O charge distributions between RC = 1.6 Å and RC = 20 Å are relatively minor, further suggesting weak polarization effects. Overall, O atom charges exhibit slightly greater sensitivity to environmental changes than Cl charges. Interestingly, the charge distributions derived from CAM-B3LYP and ωB97X-D are very similar, further supporting the use of fixed partial charges, qi and qj, in the nonbonded pairwise interaction model (see eqn (1)), used in the current studies.
3.6 Discussion and ranking of the studied plasticizers
The systematic analysis presented in this paper focuses on the impact of structural variations on interaction strength across different plasticizers. The study examines how features such as ortho vs. para substitution in phthalate plasticizers, branching in adipates and orthophthalates influence the ratio of van der Waals to electrostatic interactions. These findings provide predictive insights that are crucial for optimizing polymer–plasticizer systems and facilitating informed material design strategies. A summary of the findings is presented as ranking of the plasticizers.Fig. 9 ranks the fourteen plasticizers studied according to their binding affinity with a PVC polymer consisting of 20 repeating units. Among the five iso-orthophthalates examined—DITP, DIDP, DINP, DIOP, and DIBP—across PVC polymers of varying lengths (6 to 20 repeating units), the PMF calculations indicate that DITP exhibits the strongest interaction with PVC, displaying the highest N-PMF value of 6.42 × 10−3 kcal g−1. Among the three citrates studied—ATBC, ATHC, and BTHC—the PMF calculations indicated that BTHC had the highest N-PMF value of 6.94 × 10−3 kcal g−1, making it the best among the citrates. Further analysis of all fourteen plasticizers identified TOTM as the most promising candidate, with the highest N-PMF value of 8.33 × 10−3 kcal g−1, suggesting it has the potential to replace DEHP. TOTM also showed the strongest interaction with PVC, implying its leachability is expected to be the lowest. However, it is important to note that factors such as mechanical properties, optical properties, biocompatibility, cytotoxicity, leachability, hemolysis, and other performance characteristics must also be considered when selecting an ideal replacement for DEHP. These factors will be investigated in a separate future study. The figures showing the ranking of plasticizers arising out of interaction of different plasticizers with PVC polymers of 14, 10, 6 repeating units is provided in SI Fig. S29 for reference. Small differences in the ranking of plasticizers were seen with polymer chains of different lengths. The figure with longer PVC polymer chain (PVC-20) is shown here as we expect the ranking of plasticizers to remain unchanged for longer chain length.
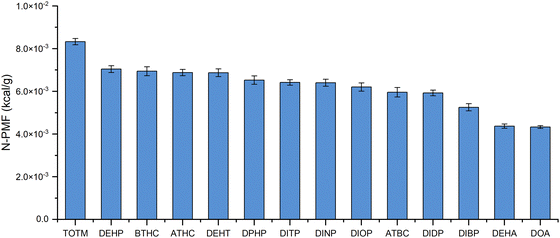 | ||
| Fig. 9 N-PMF values obtained from interaction studies between a PVC polymer with 20 repeating units and various plasticizers. | ||
3.7 Comparison with literature
The results presented in Fig. 9 were compared with similar simulations and experiments conducted by other groups on the systems studied here. A positive correlation was observed between the N-PMF values (plotted on the x-axis) and the heat of mixing (plotted on the y-axis), the latter of which was calculated by Li et al.14 using atomistic MD simulations, a method distinct from PMF. This correlation is shown in SI Fig. S30. The findings presented here align with those published by other groups, lending support to the methodology used and validating the results and model proposed. While the methodology presented here is not typically employed for studying polymer–plasticizer interactions, it complements such studies and offers the advantage of being computationally less expensive and faster compared to other methods. This could improve the precision of screening results by yielding better statistics from more simulation runs. Notably, the approach presented here requires only 25 node hours for one simulation on a commodity-type Linux cluster with 16 cores, in contrast to other approaches—such as full MD simulations of plasticizers in a PVC matrix—which take significantly longer to equilibrate and run. This method can be effectively used to screen candidates for next-generation plasticizers.Other research groups have also measured the migration rates of plasticizers in PVC-based tubing or plastic medical devices, and the results presented here are consistent with the trends observed in the current study and data from a few references is given below.74–77 Kambia et al. studied the release of TOTM and DEHP from PVC-based hemodialysis tubing plasticized with a TOTM-DEHP blend, and it was observed that TOTM leached less compared to DEHP.78 A similar trend is suggested in the PMF calculations presented here. TOTM had a higher N-PMF value of 8.33 × 10−3 kcal g−1 compared to DEHP (7.04 × 10−3 kcal g−1), implying a stronger interaction between TOTM and PVC. Furthermore, Millot et al.21 performed microsecond timescale simulations between PVC and TOTM/DEHT in ethanol to measure the leaching rates. The migration/leaching rates of the studied plasticizers were found to follow the order TOTM < DEHT, implying that TOTM interacts more strongly with PVC and hence has a lower migration rate, which is consistent with the results presented here. Consistently, migration assays under clinically relevant flow conditions showed DEHP to have the highest migration (up to 600 g dm−2 mL−1), while TOTM exhibited the lowest release (25 g dm−2 mL−1), corresponding to an approximately 350-fold reduction relative to DEHP.79 Therefore, the model presented, along with these studies, finds application in the field of extractables and leachables and could be used in designing and selecting the best extractable and leachable molecule.
Conclusions
The study utilizes atomistic MD simulations and PMF calculations to efficiently investigate interactions between PVC polymers of varying lengths (6–20 repeating units) and 14 different plasticizers, including orthophthalates, citrates, adipates, and the terephthalate DEHT. Since the molecules lack ionizable groups, interactions primarily occur through electrostatic and van der Waals forces. DFT calculations suggest that polarization effects are negligible, supporting the use of non-polarizable force fields in molecular dynamics simulations.A model based on N-PMF (dissociation energy of two molecules normalized by the sum of their molar masses) plots is proposed to analyze the slopes and y-intercepts for interpreting these interactions. Positive slopes with increasing molar mass indicate an increase in the ratio of van der Waals to electrostatic interactions, while negative slopes suggest a decrease in this ratio. Variations in y-intercepts reflect differences in the magnitude of electrostatic interactions. The model's validity is further confirmed through the examination of PVC–citrate interactions and individual plasticizer interactions with PVC polymers of different chain lengths.
This systematic analysis also investigates the impact of structural variations on interaction strength across different plasticizers. The study examines how features such as ortho vs. para substitution in phthalate plasticizers, branching in adipates and orthophthalates influence the ratio of van der Waals to electrostatic interactions. These findings offer predictive insights that are crucial for optimizing polymer–plasticizer systems and guiding informed material design strategies.
Additionally, the model can elucidate structural influences on PVC–plasticizer interactions, aiding in the design of novel plasticizers potentially replacing DEHP. Furthermore, interaction studies between the plasticizers and PVC chains revealed TOTM to be the most promising plasticizer based on binding affinity (N-PMF), as it interacts strongly with PVC and consequently leaches less, making it a potential replacement for DEHP. The top four plasticizers from our studies, based on their interaction with PVC-20 (excluding DEHP), are TOTM, BTHC, ATHC, and DEHT. TOTM was found to be better than DEHP, while BTHC, ATHC, and DEHT were found to be comparable to DEHP based on their binding affinities. These four plasticizers could be considered for further investigation as potential DEHP replacements. This work will also provide the necessary parameters for the development of coarse-grained molecular dynamics (CGMD) simulations.
Future work will involve exploring longer polymer chains, utilizing newer molecular dynamics force fields, and studying other polymers to enhance understanding and develop novel plasticizers. Overall, the model provides insights into PVC–plasticizer interactions, facilitates improved plasticizer design, and could be used in the development of new plasticizers, as well as in the study of extractables and leachables.
Author contributions
V. N. R. K. V. and S. S. R. K. C. conceived the idea; methodology designed by V. N. R. K. V. and J. M. Y. C.; software written by J. M. Y. C., S. A. G. S., and V. N. R. K. V.; data collection by S. A. G. S. and V. N. R. K. V.; data validation and formal analysis by V. N. R. K. V., J. M. Y. C and S. A. G. S.; resources provided by V. N. R. K. V. and J. M. Y. C.; data curation by S. A. G. S. and V. N. R. K. V.; J. J. performed DFT calculations; the first draft of the manuscript was prepared by S. A. G. S. and V. N. R. K. V. All authors have reviewed and approved the final version of the manuscript. V. N. R. K. V. supervised the project.Conflicts of interest
All the authors declare no other competing interests. One of the authors, SSSRKC, is the founder and Chief Executive Officer of Xheme Inc. The company's financial interests had no role in the study design; data collection, analysis, or interpretation; the decision to publish; or the preparation of the manuscript.Data availability
Data will be provided upon request. Please contact V N Ravi Kishore Vutukuri.The supporting information (SI) includes the chemical structures of all studied plasticizers; a simulation workflow schematic; PMF plot; N-PMF profiles for plasticizer–polymer systems; radius of gyration analyses under van der Waals-only and total interaction conditions; dihedral angle distributions; entropy estimation plot; DFT-derived atomic charges; and the ranking of plasticizers. See DOI: https://doi.org/10.1039/d5cp04798f.
Acknowledgements
Computational work was conducted under user projects CNMS2021-B-00956, CNMS2024-A-02400, and CNMS2026-R-03746 during the period August 2021 to May 2026 at the Center for Nanophase Materials Sciences (CNMS), a U.S. Department of Energy Office of Science User Facility at Oak Ridge National Laboratory. This research used resources of the Oak Ridge Leadership Computing Facility at Oak Ridge National Laboratory, which is supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC05-00OR22725. VNRKV thanks Jyotishman Dasgupta and Ravi Venkatramani from TIFR, Mumbai, for valuable discussions. JJ acknowledges computational resources provided by the ACCESS (Advanced Cyberinfrastructure Coordination Ecosystem: Services & Support) program through allocation TG-DMR110037.References
- T. Mekonnen, P. Mussone, H. Khalil and D. Bressler, J. Mater. Chem. A, 2013, 1, 13379–13398 RSC.
- A. Qadeer, K. L. Kirsten, Z. Ajmal, X. Jiang and X. Zhao, Environ. Sci. Technol., 2022, 56, 1482–1488 CrossRef CAS PubMed.
- J. A. Tickner, T. Schettler, T. Guidotti, M. McCally and M. Rossi, Am. J. Ind. Med., 2001, 39, 100–111 CrossRef CAS PubMed.
- F. Chiellini, M. Ferri, A. Morelli, L. Dipaola and G. Latini, Prog. Polym. Sci., 2013, 38, 1067–1088 CrossRef CAS.
- M. Hakkarainen, Chromatogr. Sustainable Polym. Mater.: Renewable, Degradable Recyclable, 2008, 159–185 CAS.
- L. Bernard, B. Décaudin, M. Lecoeur, D. Richard, D. Bourdeaux, R. Cueff, V. Sautou and A. S. Group, et al. , Talanta, 2014, 129, 39–54 CrossRef CAS PubMed.
- M. Rahman and C. S. Brazel, Prog. Polym. Sci., 2004, 29, 1223–1248 CrossRef CAS.
- The European Union, EU Phthalates Directive 2005/84/EC. 14 December 2005, https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=celex.
- Council of the European Union, Annex to the Commission Regulation (EU) Amending Annex XIV to Regulation (EC) No 1907/2006 of the European Parliament and of the Council Concerning the Registration, Evaluation, Authorisation and Restriction of Chemicals (REACH), 2021, https://Data.Consilium.Europa.Eu/Doc/Document/ST-6893-2021-ADD-1/En/Pdf (https://Data.Consilium.Europa.Eu/Doc/Document/ST-6893-2021-ADD-1/En/Pdf).
- The European Union, Regulation (EU) 2017/745 of the European Parliament and of the Council of 5 April 2017 on medical devices, amending Directive 2001/83/EC, Regulation (EC) No 178/2002 and Regulation (EC) No 1223/2009 and repealing Council Directives 90/385/EEC and 93/42/EEC, 2017, https://eur-lex.europa.eu/legal-content/EN/TXT/?uri=celex.
- The European Union, Commission Regulation (EU) 2023/2482 of 13 November 2023 amending Regulation (EC) No 1907/2006 of the European Parliament and of the Council as regards the substance bis(2-ethylhexyl) phthalate (DEHP) in medical devices, 2017, https://eur-lex.europa.eu/eli/reg/2023/2482.
- California Office of Environmental Health Hazard Assesment, Chemicals Considered or Listed Under Proposition 65, 2017, https://oehha.ca.gov/Proposition-65/Chemicals/Di2-Ethylhexylphthalate-Dehp.
- R. Jamarani, H. C. Erythropel, J. A. Nicell, R. L. Leask and M. Marić, Polymers, 2018, 10, 834 CrossRef PubMed.
- D. Li, K. Panchal, R. Mafi and L. Xi, Macromolecules, 2018, 51, 6997–7012 CrossRef CAS.
- R. Stringer, I. Labunska, D. Santillo, P. Johnston, J. Siddorn and A. Stephenson, Environ. Sci. Pollut. Res., 2000, 7, 27–36 CrossRef CAS PubMed.
- J. Drexler and J. Šimonk, Angew. Makromol. Chem., 1982, 102, 71–79 CrossRef CAS.
- N. González and M. Fernández-Berridi, J. Appl. Polym. Sci., 2006, 101, 1731–1737 CrossRef.
- Y. Zhou and S. T. Milner, Macromolecules, 2018, 51, 3865–3873 CrossRef CAS.
- D. Li, K. Panchal, N. K. Vasudevan, R. Mafi and L. Xi, Chem. Eng. Sci., 2022, 249, 117334 CrossRef CAS.
- M. Sahnoune, N. Tokhadzé, S. E. C. El Kettani, J. Devémy, F. Goujon, P. Chennell, A. Dequidt, C. Goutaudier, V. Sautou and P. Malfreyt, ACS Appl. Polym. Mater., 2022, 4, 4538–4550 CrossRef CAS.
- M. S. Millot, J. Devémy, P. Chennell, J. Pinguet, A. Dequidt, V. Sautou and P. Malfreyt, J. Mol. Liq., 2024, 396, 123965 CrossRef.
- S. S. Jagarlapudi, H. S. Cross, T. Das and W. A. Goddard III, ACS Appl. Mater. Interfaces, 2023, 15, 24858–24867 CrossRef CAS.
- J. G. Kirkwood, J. Chem. Phys., 1935, 3, 300–313 CrossRef CAS.
- B. Roux, Comput. Phys. Commun., 1995, 91, 275–282 CrossRef CAS.
- A. Grossfield, WHAM: the weighted histogram analysis method, 2012 Search PubMed.
- S. Kumar, J. M. Rosenberg, D. Bouzida, R. H. Swendsen and P. A. Kollman, J. Comput. Chem., 1992, 13, 1011–1021 CrossRef CAS.
- M. Bocqué, C. Voirin, V. Lapinte, S. Caillol and J.-J. Robin, J. Polym. Sci., Part A: Polym. Chem., 2016, 54, 11–33 CrossRef.
- Z.-M. Zhang, H.-H. Zhang, Y.-W. Zou and G.-P. Yang, Environ. Pollut., 2018, 240, 235–247 CrossRef CAS PubMed.
- Q. Zhang, J. Song, X. Li, Q. Peng, H. Yuan, N. Li, L. Duan and J. Ma, Mar. Pollut. Bull., 2019, 140, 107–115 CrossRef CAS.
- K. Khosravi and G. W. Price, Microchem. J., 2015, 121, 205–212 CrossRef CAS.
- K. M. Main, G. K. Mortensen, M. M. Kaleva, K. A. Boisen, I. N. Damgaard, M. Chellakooty, I. M. Schmidt, A.-M. Suomi, H. E. Virtanen and J. H. Petersen, et al. , Environ. Health Perspect., 2006, 114, 270–276 CrossRef CAS PubMed.
- Y.-M. Lee, J.-E. Lee, W. Choe, T. Kim, J.-Y. Lee, Y. Kho, K. Choi and K.-D. Zoh, Environ. Int., 2019, 126, 635–643 CrossRef CAS PubMed.
- R. Mankidy, S. Wiseman, H. Ma and J. P. Giesy, Toxicol. Lett., 2013, 217, 50–58 CrossRef CAS PubMed.
- J. Jurewicz and W. Hanke, Int. J. Occ. Med. Environ. Health, 2011, 24, 115–141 Search PubMed.
- E. G. Radke, A. Galizia, K. A. Thayer and G. S. Cooper, Environ. Int., 2019, 132, 104768 CrossRef CAS PubMed.
- H. D. Ozeren, M. Balçk, M. G. Ahunbay and J. R. Elliott, Macromolecules, 2019, 52, 2421–2430 CrossRef CAS.
- H. C. Erythropel, T. Brown, M. Maric, J. A. Nicell, D. G. Cooper and R. L. Leask, Chemosphere, 2015, 134, 106–112 CrossRef CAS PubMed.
- N. Kambia, A. Farce, K. Belarbi, B. Gressier, M. Luyckx, P. Chavatte and T. Dine, J. Enzyme Inhib. Med. Chem., 2016, 31, 448–455 CAS.
- H. C. Erythropel, A. Börmann, J. A. Nicell, R. L. Leask and M. Maric, Polymers, 2018, 10, 646 CrossRef PubMed.
- S. Izrailev, S. Stepaniants, B. Isralewitz, D. Kosztin, H. Lu, F. Molnar, W. Wriggers and K. Schulten, Computational Molecular Dynamics: Challenges, Methods, Ideas: Proceedings of the 2nd International Symposium on Algorithms for Macromolecular Modelling, Berlin, May 21-24, 1997, 1999, pp. 39–65.
- B. Isralewitz, M. Gao and K. Schulten, Curr. Opin. Struct. Biol., 2001, 11, 224–230 CrossRef CAS PubMed.
- S. Park and K. Schulten, J. Chem. Phys., 2004, 120, 5946–5961 CrossRef CAS PubMed.
- W. Zhang, M. L. Wang and S. W. Cranford, Sci. Rep., 2016, 6, 18659 CrossRef CAS PubMed.
- A. Yadav, S. Paul, R. Venkatramani and S. R. K. Ainavarapu, Sci. Rep., 2018, 8, 1989 CrossRef PubMed.
- W. You, Z. Tang and C.-E. A. Chang, J. Chem. Theory Comput., 2019, 15, 2433–2443 CrossRef CAS.
- J. H. Choe, J. Jeon, M. E. Lee, J. J. Wie, H.-J. Jin and Y. Yun, Nanoscale, 2018, 10, 2025–2033 RSC.
- M. D. Hanwell, D. E. Curtis, D. C. Lonie, T. Vandermeersch, E. Zurek and G. R. Hutchison, J. Cheminf., 2012, 4, 17 CAS.
- D. A. Case, H. M. Aktulga, K. Belfon, I. Ben-Shalom, S. R. Brozell, D. S. Cerutti, T. E. Cheatham III, V. W. D. Cruzeiro, T. A. Darden and R. E. Duke, et al., Amber 2021, University of California, San Francisco, 2021 Search PubMed.
- X. He, V. H. Man, W. Yang, T.-S. Lee and J. Wang, J. Chem. Phys., 2020, 153, 141502 Search PubMed.
- C. E. Dykstra, Chem. Rev., 1993, 93, 2339–2353 CrossRef CAS.
- J. Wang, R. M. Wolf, J. W. Caldwell, P. A. Kollman and D. A. Case, J. Comput. Chem., 2004, 25, 1157–1174 CrossRef CAS PubMed.
- L. Martnez, R. Andrade, E. G. Birgin and J. M. Martnez, J. Comput. Chem., 2009, 30, 2157–2164 CrossRef.
- J. M. Martnez and L. Martnez, J. Comput. Chem., 2003, 24, 819–825 CrossRef PubMed.
- S. Plimpton, J. Comput. Phys., 1995, 117, 1–19 CrossRef CAS.
- A. P. Thompson, H. M. Aktulga, R. Berger, D. S. Bolintineanu, W. M. Brown, P. S. Crozier, P. J. In't Veld, A. Kohlmeyer, S. G. Moore and T. D. Nguyen, et al. , Comput. Phys. Commun., 2022, 271, 108171 CrossRef CAS.
- W. M. Brown, P. Wang, S. J. Plimpton and A. N. Tharrington, Comput. Phys. Commun., 2011, 182, 898–911 CrossRef CAS.
- L. Verlet, Phys. Rev., 1967, 159, 98 CrossRef CAS.
- H. A. Lorentz, Ann. Phys., 1881, 248, 127–136 CrossRef.
- D. Berthelot, Compt. Rendus, 1898, 126, 15 Search PubMed.
- E. Pollock and J. Glosli, Comput. Phys. Commun., 1996, 95, 93–110 CrossRef CAS.
- P. P. Ewald, Ann. Phys., 1921, 369, 253–287 CrossRef.
- B. Efron, J. Am. Stat. Assoc., 1987, 82, 171–185 CrossRef.
- T. J. DiCiccio and B. Efron, Statist. Sci., 1996, 11, 189–228 Search PubMed.
- G. S. Watson and E. J. Williams, Biometrika, 1956, 43, 344–352 CrossRef.
- S. Wheeler and G. S. Watson, Biometrika, 1964, 51, 256–257 CrossRef.
- O. Guvench and A. D. J. MacKerell, J. Mol. Model., 2008, 14, 667–679 CrossRef CAS PubMed.
- K. V. Mardia, Sankhya, Ser. A, 1969, 31, 177–190 Search PubMed.
- T. Yanai, D. P. Tew and N. C. Handy, Chem. Phys. Lett., 2004, 393, 51–57 CrossRef CAS.
- Y.-S. Lin, G.-D. Li, S.-P. Mao and J.-D. Chai, J. Chem. Theory Comput., 2013, 9, 263–272 Search PubMed.
- S. Grimme, J. Comput. Chem., 2006, 27, 1787–1799 CrossRef CAS PubMed.
- S. Grimme, J. Antony, S. Ehrlich and H. Krieg, J. Chem. Phys., 2010, 132, 154104 CrossRef PubMed.
- S. Grimme, S. Ehrlich and L. Goerigk, J. Comput. Chem., 2011, 32, 1456–1465 CrossRef CAS PubMed.
- B. H. Besler, K. M. Merz Jr and P. A. Kollman, J. Comput. Chem., 1990, 11, 431–439 CrossRef CAS.
- L. Panneel, P. Cleys, C. Breugelmans, C. Christia, G. Malarvannan, G. Poma, P. G. Jorens, A. Mulder and A. Covaci, Int. J. Pharm., 2023, 631, 122472 CrossRef CAS PubMed.
- F. Münch, C. Höllerer, A. Klapproth, E. Eckert, A. Rüffer, R. Cesnjevar and T. Göen, Chemosphere, 2018, 202, 742–749 CrossRef.
- E. Eckert, F. Münch, T. Göen, A. Purbojo, J. Müller and R. Cesnjevar, Chemosphere, 2016, 145, 10–16 CrossRef CAS PubMed.
- D. Bourdeaux, M. Yessaad, P. Chennell, V. Larbre, T. Eljezi, L. Bernard, V. Sautou and A. S. Group, et al. , J. Pharm. Biomed. Anal., 2016, 118, 206–213 Search PubMed.
- K. Kambia, T. Dine, R. Azar, B. Gressier, M. Luyckx and C. Brunet, Int. J. Pharm., 2001, 229, 139–146 Search PubMed.
- L. Bernard, T. Eljezi, H. Clauson, C. Lambert, Y. Bouattour, P. Chennell, B. Pereira, V. Sautou and on behalf of the ARMED Study Group, PLoS One, 2018, 13, 1–15 Search PubMed.
| This journal is © the Owner Societies 2026 |