Open Access Article
Open Access ArticleShockwave-induced structural changes of lipid flat disk and transition to vesicle
Kenichiro Koshiyama
Graduate School of Technology, Industrial and Social Sciences Tokushima University, 2-1 Minamijosanjima-cho, Tokushima City, Tokushima, 770-8506, Japan. E-mail: koshiyama@tokushima-u.ac.jp
First published on 27th April 2026
Abstract
The transition from lipid flat disks to vesicles under shock waves is essential for producing nanosized vesicles during sonication. We perform non-equilibrium molecular dynamics simulations to examine how shock waves interact with a lipid flat disk. The lipid disk consists of coarse-grained saturated phospholipid models and is approximately 30 nm in diameter in the gel phase. Shock waves are simulated using a piston-driven method, with piston speeds limited to 1.0 km s−1 or less. When a planar shock wave strikes, the disk's structural changes depend on the impact angle and the shock intensity. A disk with its rotation axis parallel to the shock direction decreases in thickness while maintaining its circular shape. In contrast, a disk with its rotation axis perpendicular to the shock wave direction undergoes radial compression in the shock propagation direction, causing a temporary increase in ellipticity. Behind the shock front, lipid molecules become disordered, as indicated by a reduction in the average P2 order parameter of lipid chains and the gel fraction in the disk. This suggests that shock waves can trigger the phase transition of lipid disks from the gel to the liquid phase. The shock's intensity and the resulting structural changes influence subsequent vesicle formation. During recovery, vesicles often form from the disk after exposure to higher-intensity shock waves or after a temporary anisotropic disk induced by a side impact. This highlights the importance of impact angle. These structural changes in lipid flat disks caused by shock waves may help in understanding and controlling vesicle sizes through sonication.
1. Introduction
Small Unilamellar Vesicles (SUVs), also known as nanoliposomes, are spherical vesicles formed from a single lipid bilayer and range in size from several tens to hundreds of nanometers. Because they can encapsulate both hydrophobic and hydrophilic molecules, they have been widely studied as delivery vehicles for active substances in agriculture, food, cosmetics, and medicine.1–3 Additionally, SUVs are biologically produced as part of the extracellular vesicle family4 and play vital roles in intercellular communication.5 Therefore, producing engineered SUVs is crucial for designing molecular cargos that mimic natural biological processes and serve as models in fundamental biological research.A standard method for creating engineered SUVs involves sonication, which applies high ultrasound energy based on cavitation.3,6–10 During sonication, multilamellar or large unilamellar vesicles break into small bilayer fragments (flat disks), which then close into vesicles. Assuming spontaneous vesiculation, this process is explained by the balance between edge energy, which pulls the disk's edges, and elastic energy, related to the disk's bending.6,11 Additionally, the non-linear elastic properties of lipid disks have been implemented to understand the formation of nano-sized vesicles.12 Although the theoretical aspect of spontaneous vesiculation has been emphasized, the basic understanding of how sonication influences vesicle formation remains limited.7–9 In the sonication method, vesicles form from flat disks under the influence of cavitation. In contrast, conventional theory mainly focuses on spontaneous vesicle formation, assuming symmetrical shape changes and steady mechanical properties. Consequently, the high polydispersity of sonicated SUV size distributions remains unresolved, making it challenging to produce sonicated vesicles with optimal properties for many applications.
The sonication method relies on ultrasound cavitation, which involves various physical phenomena, including shear stress from microstreaming, microjetting, bubble oscillations, and shock waves.13,14 The shock wave is the initial and most intense phenomenon associated with cavitation and is known to cause permeabilization of biological cell membranes.15–17 Because the shockwave generated by cavitation in water occurs on a picosecond timescale,18,19 observing the dynamics of lipid nanoparticles during shockwave propagation experimentally is challenging. Non-equilibrium molecular dynamics (NEMD) simulations provide a valuable tool for numerically investigating picosecond lipid dynamics at nanometer spatial scales under the influence of shock waves. For example, NEMD studies have examined the interactions between shock waves and lipid bilayers20–23 and proteins24,25 Additionally, the effects of bubble jets induced by shock waves on cell membranes26–31 and the high-speed stretching of lipid bilayers associated with shock wave propagation32–35 have been explored. However, it remains unclear how shock waves influence lipid flat disks at the molecular scale, which are indispensable for understanding the vesiculation mechanism during sonication.
In this study, we investigate the behavior of lipid flat disks in response to plane shock waves and examine how impact angles and shock intensities influence the structural changes and the transition to vesicles using NEMD simulations. In the Methods section, we describe the preparation of the basic system using coarse-grained force fields for lipid systems and detail the implementation of the piston-driven shock wave technique. In the Results section, we verify the properties of plane shock waves and stable lipid flat disks, analyze the structural changes of the disks and lipid components under shock conditions, and the recovery processes after shock impact. We also examine the effects of impact angle and shock intensity. In the Discussion section, we present a possible scenario of vesiculation caused by shock waves during sonication and discuss methods to control the size of sonicated vesicles, along with limitations and future directions of this study.
2. Methods
The MARTINI force field version 3,36,37 a recently reparameterized coarse-grained (CG) molecular model of lipid systems, is used as the force field in CGMD calculations (Fig. 1(A)). Dipalmitoylphosphatidylcholine (DPPC) molecule is used as the lipid component of lipid flat disks and is represented in the CG model,36,37 where the choline, phosphate, and glycerol groups are mapped as NC3, PO4, and two GL beads, respectively. The two 16-carbon acyl chains are each modeled by four beads (C1–C4), with each bead representing four methylene units following the standard 4-to-1 mapping ratio.37 Additionally, four water molecules are represented as a single W bead. A rectangular computational system filled with CG water beads, subject to periodic boundary conditions in all directions, is prepared. A lipid flat disk with a diameter of approximately 30 nm, composed of 2400 DPPC molecules,38,39 is placed in the system (Fig. 1(B) and (C)). To prepare a stable DPPC disk, equilibration MD simulations are performed for 3 µs at 310 K and 1 bar, which is below the gel-liquid phase transition temperature of the MARTINI v3 DPPC lipid bilayer. The general setup36 is used for calculating CG interactions, except that the Verlet buffer tolerance is set to 0.0002 kJ mol−1 ps−1.40 The v-rescale algorithm41 with a time constant of 1.0 ps is used for the thermostat, and the Parrinello–Rahman algorithm42 with a time constant of 10 ps is used for the barostat. The time step for integrating the equations of motion using the leap-frog algorithm is 20 fs for the equilibrium MD simulations.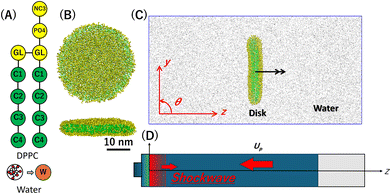 | ||
| Fig. 1 MARTINI v3 CG models of saturated lipid (DPPC) and water (A), top and side views of a stable lipid flat disk (B), the system configuration (C), and the schematic of a piston-driven plane shock wave simulation (D). The hydrophilic headgroup beads and hydrophobic tail beads are shown in yellow and green, respectively, with their names36 in Fig. 1(A) (see the text for details). The rotation axis of a lipid disk is indicated by the black arrow in Fig. 1(C). | ||
To apply a plane shock wave to the lipid systems, the periodic boundary condition is removed along the long axis of the system, and a Lennard-Jones potential wall, composed of coarse-grained water beads, is placed at one end. The generation and propagation of piston-driven plane shock waves are simulated by applying a piston velocity, Up, towarpd the potential wall to all beads, which are then reflected by the wall (Fig. 1(D)).43,44 During the shock wave MD simulations, temperature, pressure, and center of mass motion controls are not implemented, while the neighbor list is updated every step. The time step for integrating the equations of motion is set to 1 fs in shock wave MD simulations.
To investigate the effects of the shock impact angle on lipid flat disks, we prepare two systems in which the disk's rotation axis is set to be parallel (θ = 0°) or perpendicular (θ = 90°) to the direction of shock wave propagation (Fig. 1(C)). The initial distance between the potential wall and the center of a lipid disk is 36 nm. Shock wave MD simulations are conducted for three different initial configurations under each angle condition. The analysis of structural changes is limited to the period when the shock wave reflects at the vapor-liquid water interface (Fig. 1(D)). We perform MD simulations with GROMACS 2025 series in double precision45,46 and modify them to implement the piston-driven shock wave algorithm. The configuration pressure tensor is calculated using GROMACS-LS,47 and other analyses are carried out with GROMACS tools and in-house C and Python codes, partly supported by MDAnalysis.48
To verify how a lipid flat disk recovers after being impacted by a shock wave, we perform additional recovery simulations under controlled pressure and temperature conditions. We extract a shock-compressed region containing the lipid disk and run a short MD simulation to relax the system with pressure in the compressed direction, Pz = 1.0 bar, constant area A, and temperature T = 310 K, i.e., constant nPzAT MD simulation. Afterward, the pressure in all directions and temperature are relaxed for up to 1 µs. Note that the method for conducting recovery MD simulations within current computational limits remains controversial; a similar approach has been used to verify membrane recovery after shock wave impact.27
3. Results
3.1. Shock wave characteristics
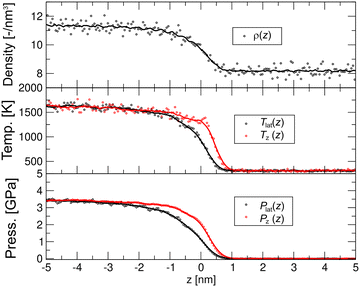
For shock wave CGMD simulations, we first verify the modeled shock waves by examining the relationship between particle velocity, Up, and shock wave propagation speed, Us, i.e., the Us−Up Hugoniot curve for water.49 Fig. 2 shows the Us−Up Hugoniot curve obtained with MARTIN v3 water beads. Here, Up represents piston velocity, and Us is measured directly from density-wave propagation. The CG water curve matches well with experimental data for Up ≤ 1.0 km s−1. Previous studies have shown that curves generated using the CG water model with MARTINI v227 or the all-atom water model26 also fit experimental data well. Although the water model in MARTINI v3 is slightly modified from MARTINI v2,36 it remains valid at least for Up values below 1.0 km s−1. Since the bubble wall speed of a cavitation bubble can reach about 1.0 km s−1 within micrometers from its center,18,19 we focus on lipid dynamics caused by shock waves with Up limited to 1.0 km s−1 or less. Additionally, we verify the density, temperature, and pressure profiles across the shock front for Up = 1.0 km s−1 (Fig. 3). In this range, pressure and temperature anisotropies become apparent, as reported in previous studies of strong shockwaves in simple Lennard-Jones liquids.43,50 Note that for NEMDs, the effects of computational parameters should be verified.20,21,32 We tested following parameters and algorithms of shock wave MD simulations: the time set of 0.5 fs, the wall model of hydrophobic beads, and the LAMMPS-like piston-driven method with momentum mirror.51 The obtained density profiles were unchanged at least with the above tested parameters and algorithms (see Fig. S1).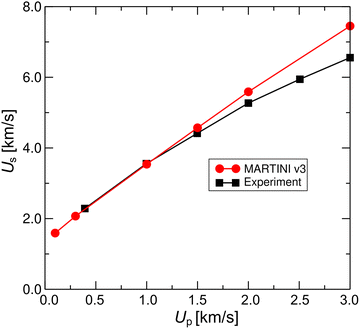 | ||
| Fig. 2 The relationship between the shock wave propagation speed, Us, and the particle velocity, Up, i.e., the Us–Up Hugoniot curve for MARTINI V3 water beads and water (experiment49). | ||
3.2. Stable lipid disk structure
The stable lipid disk structure forms the basis for controlling the transition to a vesicle. The radius of the 2400-lipid disk was 14.76 nm, based on the convex hull of lipid beads projected onto the disk plane. The thickness was determined to be 4.86 nm, measured as the distance between the averaged positions of PO4 beads in the upper and lower monolayers. The relative shape anisotropy estimated from the gyration tensor of C4 beads52,53 was approximately 0.25, indicating a well-flat and symmetric disk shape. The P2 order parameter,54 averaged over the lipid chains (C1–C2, C2–C3, and C3–C4; Fig. 1(A)) in the disk core region, was 0.93, indicating a highly ordered lipid chain structure. To quantify lipids in the gel phase of the disk, we analyzed the lipid arrangement, which is regularly packed in a honeycomb lattice, based on previous studies of the gel-liquid phase transition in MARTINI CG lipid systems.55 The lipids in the disk core were frequently identified as gel lipids (e.g., Fig. 4(C)), and the gel phase lipid fraction exceeded 68% of the total lipids. In this context, without the influence of shock waves, we consider the 2400 DPPC disk prepared at 310 K and 1 bar to be a stable gel-phase lipid disk when using the MARTINI v3 force field. | ||
| Fig. 4 Representative snapshots of lipid flat disks under the impact of shockwaves. The hydrophilic and hydrophobic beads are colored in yellow and green, respectively. The only C2 beads are shown in Fig. 4(c). | ||
3.3. Structural changes of lipid disk
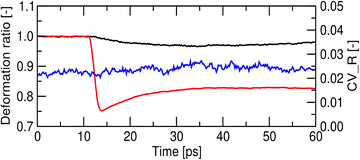
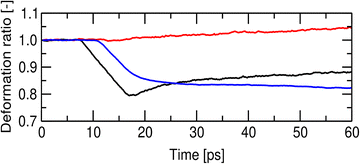
The shockwave-induced structural changes of a disk depend on its orientation. Fig. 4 shows typical structures of a disk aligned parallel (θ = 0°) and perpendicular (θ = 90°) to the shockwave's propagation direction. Before the shock wave impacts (left panels of Fig. 4(A)–(C)), the disk has an isotropic circular shape, with ordered lipid tails in the core region. After the impact, the shock wave compresses the disks along the propagation direction (right panels of Fig. 4(A) and (B)). For θ = 0°, the disk thickness quickly decreases and then slightly rebounds (red line in Fig. 5), which may be because of the anisotropy of the pressure tensor at the shock front (see Fig. 3). At the same time, its radius and shape remain mostly unchanged (black and blue lines in Fig. 5), indicating the compression maintaining its circular shape (Fig. 4(A) and (C)). Conversely, for θ = 90°, both the diameter along the propagation direction and the thickness decrease significantly (Fig. 6), leading to a temporary transition toward an anisotropic, ellipsoidal shape disk (Fig. 4(A) and (C)).Before the shock wave impact, the lipids in the core region of the disk are in the gel phase (Fig. 4(C) left). The shock wave distorts the lipid arrangement through disk compression and slight rebound. The distortion is most noticeable near the edge of the disk but develops into complex patterns (Fig. S2 and S3). For θ = 0°, the averaged P2 order parameter of lipid molecules drops suddenly and then rebounds, resembling the change in thickness (Fig. 4 and 7), while the gel fraction decreases monotonically. For θ = 90°, the P2 order parameter decreases monotonically. The gel fraction temporarily increases with the first compression but then decreases continuously (Fig. 7). For both θ = 0° and 90°, the decrease in gel fraction persists for at least 60 ps in the high-pressure and high-temperature region behind the shock front (Fig. 3).
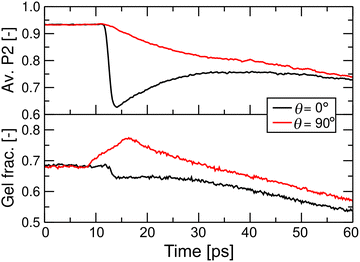 | ||
| Fig. 7 Representative changes of the average P2 order parameters of lipid chains and the gel fraction for Up = 0.8 km s−1. | ||
3.4. Effects of shock wave intensity
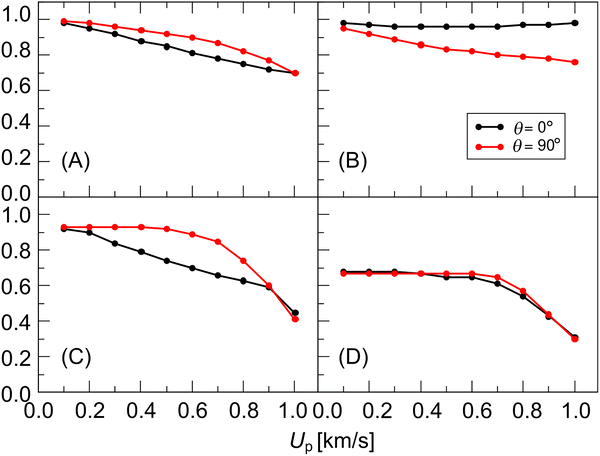
The structural changes of a lipid disk depend on the impact angle and shock wave intensity. Since shock wave intensity is proportional to Up, the changes are evaluated relative to Up. The normalized minimum thickness (Fig. 8(A)) decreases to about 0.7 as Up increases, and the values for θ = 0° are smaller than those for θ = 90°. The normalized minimum radius (Fig. 8(B)) appears to decrease with Up for θ = 90° and remains nearly unchanged for θ = 0°. This indicates volumetric compression of the disk occurs for both angle conditions. The minimum average P2 order parameter (Fig. 8(C)) decreases with Up, but the reduction becomes significant after Up = 0.7 km s−1, especially for θ = 90°. For both angle conditions, the minimum gel fraction (Fig. 8(D)) remains unaffected until Up reaches about 0.7 km s−1, after which it decreases markedly.3.5. Recovery from shock-compressed lipid disk
After experiencing a shock wave, the flat lipid disk either returns to its original shape or transforms into a vesicle during relaxation to 1 bar and 310 K (Fig. 9(A) and (B)). When a transition to a vesicle occurs, the lipid-order distortion caused by the shock wave (Fig. 4(C) and Fig. 8(C) and (D)) persists during relaxation, leading to vesicle formation (Fig. 9(B)). Fig. 9(C) displays the vesiculation rate for different Up values across three samples. Vesiculation takes place after a strong shock wave impact with Up ≥ 0.8 km s−1, roughly corresponding to the apparent decrease in the gel fraction (Fig. 8(D)). Furthermore, the threshold Up required to trigger vesiculation for θ = 90° (0.8 km s−1) is lower than for θ = 0° (0.9 km s−1). This suggests that shock impact from the disk side may be more effective in inducing vesiculation during relaxation. Note that we tested the effects of extracted regions on the vesiculation rate for recovery simulations (see the Method section). Although the threshold Up changes slightly, the qualitative result of the favor vesiculation for the disks when impacted from the side remains unchanged (see Fig. S4).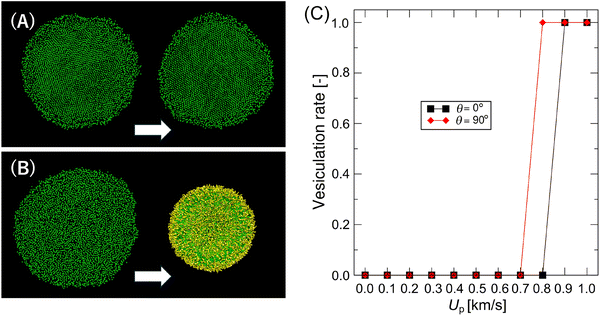 | ||
| Fig. 9 Representative snapshots of the recovery to disk (A) and the transition to vesicle (B) during relaxation MD simulations from SW-impacted lipid disk, and the vesiculation rate vs. Up (C). In Fig. 9(A) and (B), the results after short (left) and long (right) relaxation MD simulations are shown. All lipid beads are shown to visualize the vesicle clearly in Fig. 9(B). The vesiculation rate is calculated as the number of vesiculated samples in total (n = 3). | ||
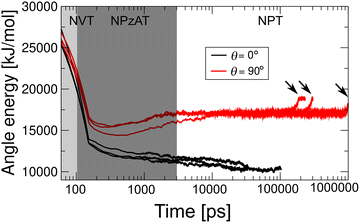
To confirm the recovery of the lipid disk from distortion, we further analyzed the temporal change in the angle potential energy, a sensitive measure of lipid phase changes, using MARTINI CG models. Fig. 10 shows representative results for Up = 0.8 km s−1. For θ = 0°, all samples revert to their original disk shape (Fig. 8(b)), and the energy relaxes back to its original value, where the disk is in the gel phase. In contrast, for = 90°, the energy relaxes to a higher level and then suddenly increases with vesiculation. Before vesiculation, the angle energy remains higher, suggesting that the lipid phase of the disk may be semi-stable in the liquid phase. Interestingly, when the gel-to-liquid phase transition is induced by shock waves, the semi-stable angle potential energy during recovery simulations is independent of the experienced impact angle and Up (Fig. S5). Once the liquid phase becomes semi-stable, the disk transforms into a vesicle within 1 us.
4. Discussion
4.1. Sonication method and vesiculation theory
We have demonstrated that shock waves temporarily alter the shape of lipid disks and the lipid arrangement. According to the classical vesiculation theory,6,11 the balance between the edge energy and the bending energy, thereby, the mechanical properties of the disk, is essential to the fate of disk shape. For example, lower bending modulus facilitates vesiculation, whereas higher bending modulus stabilizes the disk shape. Before shock wave impact, the lipid disk is in the gel phase (Fig. 4) and has a higher bending modulus.56 This explains the stable disk shape before the shockwaves impact. However, after high-intensity shock wave impact, the lipid arrangement becomes disordered, leading to a decrease in the gel fraction (Fig. 6). This suggests that shock waves can temporarily shift the lipid disk phase from the gel to the liquid phase. Because the bending modulus of the liquid phase is much lower than that of the gel phase,56 shock waves may set a favorable condition for vesiculation (Fig. S5). Also, the vesiculation theory states that the energy barrier for transitioning from a disk to a vesicle decreases as the bending energy decreases.11 This suggests that during sonication, shock waves from cavitation can trigger vesiculation by altering the phase of lipid disks and controlling their mechanical properties. Interestingly, shock-induced acceleration in molecular dynamics has been reported in protein binding.24,25 Furthermore, the shockwave-induced phase change in the lipid disk depends on impact angle and shock intensity (Fig. 8). Since the positions of cavitation bubbles and disks in their aqueous solutions vary significantly during the ultrasound exposure, the different sensitivities of lipid disks to shockwave impacts may explain the significant variation in the size of sonicated SUVs.7–94.2. Toward controlled vesiculation from lipid disk
Recently, a stable lipid flat disk, called a bicelle, has been extensively studied in biophysical chemistry.57 One of its unique features is its tendency to align either along or perpendicular to the magnetic field. In this paper, we revealed that the transition from lipid flat disks to vesicles can be favored in disks after shock-wave impact from the side (Fig. 8 and 9). This suggests that performing the sonication method under a magnetic field could control vesiculation. For instance, when aligned bicelles under a magnetic field are exposed to shock waves from their side generated by cavitation, the bicelles may be expected to transit to vesicles favorably. However, controlling cavitation under a magnetic field remains a challenging area of research.584.3. Limitations and future directions
Although we demonstrated structural changes in lipid flat disks induced by plane shock waves using NEMD, some limitations remain. The analysis was limited to several tens of picoseconds due to computational limits. However, some lipid dynamics (e.g., P2 order parameters or the gel fraction (Fig. 7)) would not reach convergence within this period. Additionally, we did not observe any obvious lipid flip-flop events during shock compression, which one might expect to occur because they could be more favorable at the edge in lipid bilayers.59 Likely, under extremely high-pressure and high-temperature conditions behind the shock front, lipid relaxation could take longer. Furthermore, analyzing detailed pressure distributions in the disk under shock waves can be valuable, as they may relate to vesiculation by altering the mechanical properties of lipid systems.60 Current nonequilibrium shockwave simulations restrict the sampling needed for detailed local pressure analysis. Such slow lipid dynamics and local pressure distributions can be studied using equilibrium MD simulations of shock waves,44 although applying this to biological systems is still challenging.The shock-induced distortion causing the phase change in the lipid disk develops into complex patterns (Fig. 4 and Fig. S2, and S3). We hypothesize that the phase transition triggered by shockwaves may have an intricate mechanism, since shockwaves significantly increase both temperature and pressure on picosecond time scales (Fig. 3). A detailed analysis of how the distortion occurs under shock waves requires further research, such as comparing it with results from heat-55,61 or pressure-induced62,63 phase transitions.
Bubble jets26–31 or negative pressure from pulsed shockwaves24,64 are known to influence lipid or protein dynamics, but we only considered plane shockwaves in this study. Additionally, cavitation bubbles emit spherical shockwaves whose intensities decay with distance.18,19 The interaction between shockwaves and bubbles, and the effect of shockwave pulse width on lipid disk dynamics, could be an interesting area of shockwave MD research and can be investigated using the piston-driven shockwave algorithm.44
Because we performed coarse-grained MD simulations, we did not examine more detailed atomistic features like water dynamics.20,21 All-atom MD simulations of lipid systems under shockwaves could provide valuable insights into shockwave phenomena in sonication, as water behavior during shock impacts is crucial to lipid membrane dynamics.65,66 Furthermore, although the MARTINI CG models effectively reproduce various lipid36,37 and shockwave characteristics (e.g., Fig. 2), they still need improvements in some lipid dynamics67 and phase behavior.61 All-atom MD simulations will support and improve the current understanding of lipid dynamics under shock waves achieved with the current CG models.
5. Conclusion
In this study, we performed non-equilibrium molecular dynamics simulations of lipid disks exposed to planar shock waves to understand how shock waves interact with a lipid flat disk at the molecular level. We found that shock waves compress the lipid disk in the direction of propagation and distort its lipid organization. A disk with its rotation axis parallel to the shock direction decreases in thickness while maintaining its circular shape. Conversely, a disk with its rotation axis perpendicular to the shock wave direction experiences radial compression along the shock propagation, causing a temporary increase in ellipticity. The shock wave alters the lipid arrangement through disk compression and a slight rebound, triggering a phase transition from the gel to the liquid state. During recovery, vesicles often form from the disk after exposure to higher-intensity shock waves or side impacts, which induce a temporarily anisotropic disk. Because the mechanical properties of lipid systems in the liquid phase are lower than in the gel phase, shock waves from cavitation during sonication can cause vesiculation by changing the phase of lipid disks through modulation of their mechanical properties. These molecular insights into the structural changes in lipid flat disks caused by shock waves could help in understanding and controlling vesicle size through sonication.Conflicts of interest
There are no conflicts to declare.Data availability
The modified GROMACS 2025 code, topology, computational parameters, and initial lipid disk configurations used in this paper are available in the institutional repository at [URL: https://doi.org/10.15000/0002013975]. The supporting data has been provided as part of the supplementary information (SI). Supplementary information: Fig. S1, S2, S3, S4, and S5. See DOI: https://doi.org/10.1039/d5cp04439a.Acknowledgements
This work was supported by JSPS KAKENHI Grant Number 24K22408. We thank Yugo Kakuno, Naoto Okuyama, Shuto Doi, and Taiki Shigematsu for fruitful discussions.References
- J. O. Eloy, M. Claro de Souza, R. Petrilli, J. P. A. Barcellos, R. J. Lee and J. M. Marchetti, Liposomes as carriers of hydrophilic small molecule drugs: Strategies to enhance encapsulation and delivery, Colloids Surf., B, 2014, 123, 345–363 CrossRef CAS PubMed.
- T. M. Taylor, P. M. Davidson, B. D. Bruce and J. Weiss, Liposomal nanocapsules in food science and agriculture, Crit. Rev. Food Sci. Nutr., 2005, 45, 587–605 CrossRef CAS PubMed.
- V. Weissig, Liposomes, Springer New York, New York, NY, 2nd edn, 2017, vol. 1522 Search PubMed.
- R. Kalluri and V. S. LeBleu, The biology, function, and biomedical applications of exosomes, Science, 2020, 367, eaau6977 CrossRef CAS PubMed.
- P. D. Robbins and A. E. Morelli, Regulation of immune responses by extracellular vesicles, Nat. Rev. Immunol., 2014, 14, 195–208 CrossRef CAS PubMed.
- W. Helfrich, The size of bilayer vesicles generated by sonication, Phys. Lett. A, 1974, 50, 115–116 CrossRef.
- G. Maulucci, M. De Spirito, G. Arcovito, F. Boffi, A. C. Castellano and G. Briganti, Particle size distribution in DMPC vesicles solutions undergoing different sonication times, Biophys. J., 2005, 88, 3545–3550 CrossRef CAS PubMed.
- D. J. Woodbury, E. S. Richardson, A. W. Grigg, R. D. Welling and B. H. Knudson, Reducing liposome size with ultrasound: Bimodal size distributions, J. Liposome Res., 2006, 16, 57–80 CrossRef CAS PubMed.
- A. A. Barba, S. Bochicchio, G. Lamberti and A. Dalmoro, Ultrasonic energy in liposome production: Process modelling and size calculation, Soft Matter, 2014, 10, 2574–2581 RSC.
- B. G. Tenchov, T. K. Yanev, M. G. Tihova and R. D. Koynova, A probability concept about size distributions of sonicated lipid vesicles, Biochim. Biophys. Acta, 1985, 816, 122–130 CrossRef CAS PubMed.
- P. Fromherz, Lipid-vesicle structure: Size control by edge-active agents, Chem. Phys. Lett., 1983, 94, 259–266 CrossRef CAS.
- C. Huang, D. Quinn, Y. Sadovsky, S. Suresh and K. J. Hsia, Formation and size distribution of self-Assembled vesicles, Proc. Natl. Acad. Sci. U. S. A., 2017, 114, 2910–2915 CrossRef CAS PubMed.
- F. Caupin and E. Herbert, Cavitation in water: a review, 2006, 7(9–10), 1000–1017 CAS.
- E. A. Neppiras, Acoustic Cavitation, Phys. Rep., 1980, 61, 159–251 CrossRef.
- L. M. López-Marín, A. L. Rivera, F. Fernández and A. M. Loske, Shock wave-induced permeabilization of mammalian cells, Phys. Life Rev., 2018, 26–27, 1–38 CrossRef PubMed.
- T. Kodama, Y. Tomita, K.-I. Koshiyama and M. J. K. Blomley, Transfection effect of microbubbles on cells in superposed ultrasound waves and behavior of cavitation bubble, Ultrasound Med. Biol., 2006, 32(6), 905–914 CrossRef PubMed.
- T. Kodama, Y. Tomita, Y. Watanabe, K. Koshiyama, T. Yano and S. Fujikawa, Cavitation bubbles mediated molecular delivery during sonoporation, J. Biomech. Sci. Eng., 2009, 4, 124–130 CrossRef.
- W. Lauterborn and T. Kurz, Physics of bubble oscillations, Rep. Prog. Phys., 2010, 73, 106501 CrossRef.
- W. Lauterborn and A. Vogel, Shock Wave Emission by Laser Generated Bubbles, 2013 Search PubMed.
- K. Koshiyama, T. Kodama, T. Yano and S. Fujikawa, Structural change in lipid bilayers and water penetration induced by shock waves: molecular dynamics simulations, Biophys. J., 2006, 91, 2198–2205 CrossRef CAS PubMed.
- K. Koshiyama, T. Kodama, T. Yano and S. Fujikawa, Molecular dynamics simulation of structural changes of lipid bilayers induced by shock waves: Effects of incident angles, Biochim. Biophys. Acta, 2008, 1778, 1423–1428 CrossRef CAS PubMed.
- S. Espinosa, N. Asproulis and D. Drikakis, Chemotherapy efficiency increase via shock wave interaction with biological membranes: A molecular dynamics study, Microfluid. Nanofluid., 2014, 16, 613–622 CrossRef CAS.
- R. Kfoury, B. Marzban, E. Makki, M. L. Greenfield and H. Yuan, Effect of pressure profile of shock waves on lipid membrane deformation, PLoS One, 2019, 14, 1–13 CrossRef PubMed.
- G. A. Kaminski, Computational studies of the effect of shock waves on the binding of model complexes, J. Chem. Theory Comput., 2014, 10, 4972–4981 CrossRef CAS PubMed.
- M. Araki, S. Matsumoto, G. J. Bekker, Y. Isaka, Y. Sagae, N. Kamiya and Y. Okuno, Exploring ligand binding pathways on proteins using hypersound-accelerated molecular dynamics, Nat. Commun., 2021, 12, 1–10 Search PubMed.
- A. Choubey, M. Vedadi, K. I. Nomura, R. K. Kalia, A. Nakano and P. Vashishta, Poration of lipid bilayers by shock-induced nanobubble collapse, Appl. Phys. Lett., 2011, 98, 127–129 CrossRef.
- K. P. Santo and M. L. Berkowitz, Shock wave interaction with a phospholipid membrane: Coarse-grained computer simulations, J. Chem. Phys., 2014, 140, 1–10 CrossRef PubMed.
- U. Adhikari, A. Goliaei and M. L. Berkowitz, Mechanism of membrane poration by shock wave induced nanobubble collapse: a molecular dynamics study, J. Phys. Chem. B, 2015, 119, 6225–6234 CrossRef CAS PubMed.
- N. Nan, D. Si and G. Hu, Nanoscale cavitation in perforation of cellular membrane by shock-wave induced nanobubble collapse, J. Chem. Phys., 2018, 149(7), 074902 CrossRef PubMed.
- T. Wei, M. Zhou, L. Gu, Y. Zhou and M. Li, How Shockwaves Open Tight Junctions of Blood–Brain Barrier: Comparison of Three Biomechanical Effects, J. Phys. Chem. B, 2022, 126, 5094–5102 CrossRef CAS PubMed.
- N. H. Linh, V. H. Man, M. S. Li, J. Wang, P. Derreumaux, T. L. Mai and P. H. Nguyen, Molecular dynamics simulation of cancer cell membrane perforated by shockwave induced bubble collapse, J. Chem. Phys., 2022, 157, 225102 CrossRef CAS PubMed.
- K. Koshiyama and S. Wada, Molecular dynamics simulations of pore formation dynamics during the rupture process of a phospholipid bilayer caused by high-speed equibiaxial stretching, J. Biomech., 2011, 44, 2053–2058 CrossRef PubMed.
- T. Shigematsu, K. Koshiyama and S. Wada, Effects of Stretching Speed on Mechanical Rupture of Phospholipid/Cholesterol Bilayers: Molecular Dynamics Simulation, Nat. Publ. Gr., 2015, 1–10 Search PubMed.
- L. Zhang, Z. Zhang, J. Jasa, D. Li, R. O. Cleveland, M. Negahban and A. Jérusalem, Molecular dynamics simulations of heterogeneous cell membranes in response to uniaxial membrane stretches at high loading rates, Sci. Rep., 2017, 7(1), 8316 CrossRef PubMed.
- M. A. Murphy, M. F. Horstemeyer, S. R. Gwaltney, T. Stone, M. Laplaca, J. Liao, L. Williams and R. Prabhu, Nanomechanics of phospholipid bilayer failure under strip biaxial stretching using molecular dynamics, Modell. Simul. Mater. Sci. Eng., 2016, 24(5), 055008 CrossRef.
- P. C. T. Souza, R. Alessandri, J. Barnoud, S. Thallmair, I. Faustino, F. Grünewald, I. Patmanidis, H. Abdizadeh, B. M. H. Bruininks, T. A. Wassenaar, P. C. Kroon, J. Melcr, V. Nieto, V. Corradi, H. M. Khan, J. Domański, M. Javanainen, H. Martinez-Seara, N. Reuter, R. B. Best, I. Vattulainen, L. Monticelli, X. Periole, D. P. Tieleman, A. H. de Vries and S. J. Marrink, Martini 3: a general purpose force field for coarse-grained molecular dynamics, Nat. Methods, 2021, 18, 382–388 CrossRef CAS PubMed.
- S. J. Marrink, L. Monticelli, M. N. Melo, R. Alessandri, D. P. Tieleman and P. C. T. Souza, Two decades of Martini: Better beads, broader scope, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2023, 13, 1–42 Search PubMed.
- K. Koshiyama, M. Taneo, T. Shigematsu and S. Wada, Bicelle-to-Vesicle Transition of a Binary Phospholipid Mixture Guided by Controlled Local Lipid Compositions: A Molecular Dynamics Simulation Study, J. Phys. Chem. B, 2019, 123(14), 3118–3123 CrossRef CAS PubMed.
- K. Koshiyama and K. Nakata, Effects of lipid saturation on bicelle to vesicle transition of a binary phospholipid mixture: a molecular dynamics simulation study, Soft Matter, 2023, 19, 7655–7662 RSC.
- H. Kim, B. Fábián and G. Hummer, Neighbor List Artifacts in Molecular Dynamics Simulations, J. Chem. Theory Comput., 2023, 19, 8919–8929 CrossRef CAS PubMed.
- G. Bussi, D. Donadio and M. Parrinello, Canonical sampling through velocity rescaling, J. Chem. Phys., 2007, 126, 014101 CrossRef PubMed.
- S. Nosé and M. L. Klein, Constant pressure molecular dynamics for molecular systems, Mol. Phys., 1983, 50, 1055–1076 CrossRef.
- B. L. Holian, W. G. Hoover, B. Moran and G. K. Straub, Shock-wave structure via nonequilibrium molecular dynamics and Navier-Stokes continuum mechanics, Phys. Rev. A, 1980, 22, 2798–2808 CrossRef CAS.
- P. Wen, G. Tao, D. E. Spearot and S. R. Phillpot, Molecular dynamics simulation of the shock response of materials: A tutorial, J. Appl. Phys., 2022, 131, 051101 CrossRef CAS.
- H. J. C. Berendsen, D. Vanderspoel and R. Vandrunen, Gromacs - a Message-Passing Parallel Molecular-Dynamics Implementation, Comput. Phys. Commun., 1995, 91, 43–56 CrossRef CAS.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindah, Gromacs: High performance molecular simulations through multi-level parallelism from laptops to supercomputers, SoftwareX, 2015, 1–2, 19–25 CrossRef.
- J. M. Vanegas and A. Torres-Sanchez, Examining the Mechanical Equilibrium of Microscopic Stresses in Molecular Simulations, Phys. Rev. Lett., 2015, 114, 258102 CrossRef PubMed.
- R. Gowers, M. Linke, J. Barnoud, T. Reddy, M. Melo, S. Seyler, J. Domański, D. Dotson, S. Buchoux, I. Kenney and O. Beckstein, MDAnalysis: A Python Package for the Rapid Analysis of Molecular Dynamics Simulations, Proc. 15th Python Sci. Conf., 2016, 98–105.
- A. P. Rybakov and I. A. Rybakov, Polymorphism of shocked water, Eur. J. Mech. B Fluids, 1995, 14, 323–332 Search PubMed.
- S. I. Anisimov, V. V. Zhakhovskii and V. E. Fortov, Shock wave structure in simple liquids, J. Exp. Theor. Phys. Lett., 1997, 65, 755–761 CrossRef.
- A. P. Thompson, H. M. Aktulga, R. Berger, D. S. Bolintineanu, W. M. Brown, P. S. Crozier, P. J. in't Veld, A. Kohlmeyer, S. G. Moore, T. D. Nguyen, R. Shan, M. J. Stevens, J. Tranchida, C. Trott and S. J. Plimpton, LAMMPS - a flexible simulation tool for particle-based materials modeling at the atomic, meso, and continuum scales, Comput. Phys. Commun, 2022, 271, 108171 CrossRef CAS.
- R. I. Dima and D. Thirumalai, Asymmetry in the shapes of folded and denatured states of proteins, J. Phys. Chem. B, 2004, 108, 6564–6570 CrossRef CAS.
- K. Koshiyama and S. Wada, Collapse of a lipid-coated nanobubble and subsequent liposome formation, Sci. Rep., 2016, 6, 28164 CrossRef CAS PubMed.
- T. J. Piggot, J. R. Allison, R. B. Sessions and J. W. Essex, On the Calculation of Acyl Chain Order Parameters from Lipid Simulations, J. Chem. Theory Comput., 2017, 13, 5683–5696 CrossRef CAS PubMed.
- D. Stelter and T. Keyes, Simulation of fluid/gel phase equilibrium in lipid vesicles, Soft Matter, 2019, 15, 8102–8112 RSC.
- D. Drabik, G. Chodaczek, S. Kraszewski and M. Langner, Mechanical Properties Determination of DMPC, DPPC, DSPC, and HSPC Solid-Ordered Bilayers, Langmuir, 2020, 36, 3826–3835 CrossRef CAS PubMed.
- E. J. Dufourc, Bicelles and nanodiscs for biophysical chemistry1, Biochim. Biophys. Acta, Biomembr., 2021, 1863, 183478 CrossRef CAS PubMed.
- T. Yoshimura, S. Watanabe, M. Ijiri and S. Ota, Development of processing technology using extremely high concentration cavitation energy by strong magnetic field, Results Mater., 2022, 14, 100289 CrossRef CAS.
- D. P. Tieleman and S. Marrink, Lipids Out of Equilibrium: Energetics of Desorption and Pore Mediated Flip-Flop, J. Am. Chem. Soc., 2006, 12462–12467 CrossRef CAS PubMed.
- O. H. S. Ollila, H. J. Risselada, M. Louhivuori, E. Lindahl, I. Vattulainen and S. J. Marrink, 3D Pressure field in lipid membranes and membrane-protein complexes, Phys. Rev. Lett., 2009, 102, 1–4 CrossRef PubMed.
- K. B. Pedersen, H. I. Ingólfsson, D. P. Ramirez-Echemendia, L. Borges-Araújo, M. D. Andreasen, C. Empereur-mot, J. Melcr, T. N. Ozturk, W. F. D. Bennett, L. R. Kjølbye, C. Brasnett, V. Corradi, H. M. Khan, E. A. Cino, J. Crowley, H. Kim, B. Fábián, A. C. Borges-Araújo, G. M. Pavan, G. Launay, F. Lolicato, T. A. Wassenaar, M. N. Melo, S. Thallmair, T. S. Carpenter, L. Monticelli, D. P. Tieleman, B. Schiøtt, P. C. T. Souza and S. J. Marrink, The Martini 3 Lipidome: Expanded and Refined Parameters Improve Lipid Phase Behavior, ACS Cent. Sci., 2025, 11, 1598–1610 CrossRef CAS PubMed.
- W. Ding, M. Palaiokostas, G. Shahane, W. Wang and M. Orsi, Effects of High Pressure on Phospholipid Bilayers, J. Phys. Chem. B, 2017, 121, 9597–9606 CrossRef CAS PubMed.
- M. Kato and R. Hayashi, Effects of high pressure on lipids and biomembranes for understanding high-pressure-induced biological phenomena, Biosci., Biotechnol., Biochem., 1999, 63, 1321–1328 CrossRef CAS PubMed.
- Y. Sliozberg and T. Chantawansri, Damage in spherical cellular membrane generated by the shock waves: Coarse-grained molecular dynamics simulation of lipid vesicle, J. Chem. Phys., 2014, 141, 184904 CrossRef PubMed.
- W. B. Han, S. J. Kim, H. H. An, H. S. Kim, Y. Kim and C. S. Yoon, Molecular dynamics simulation of interlayer water embedded in phospholipid bilayer, Mater. Sci. Eng., C, 2014, 36, 49–56 CrossRef CAS PubMed.
- K. Koshiyama, T. Yano and T. Kodama, Self-Organization of a Stable Pore Structure in a Phospholipid Bilayer, Phys. Rev. Lett., 2010, 105, 018105 CrossRef PubMed.
- O. Kroutil, L. Bartoš, I. Kabelka and R. Vácha, Martini 3 Limitations in Phospholipid Flip–Flop, J. Chem. Theory Comput., 2025, 21, 9227–9233 CrossRef CAS PubMed.
| This journal is © the Owner Societies 2026 |