Structural and catalytic consequences of active-site vs. distal mutations in human dehalogenase: insights from molecular dynamics simulations
Soumyajit
Karmakar
,
Biman
Giri
and
Sabyashachi
Mishra
 *
*
Department of Chemistry, Indian Institute of Technology Kharagpur, Kharagpur, West Bengal, India. E-mail: mishra@chem.iitkgp.ac.in; Tel: +91 3222 282328
First published on 24th February 2026
Abstract
Congenital hypothyroidism can result from mutations in human iodotyrosine deiodinase (hIYD), which catalyzes the deiodination of iodotyrosines (I-Tyr), a key step in thyroid hormone synthesis. Three homozygous mutations (R101W, F105–I106L, and I116T) are known causes of hypothyroidism. This computational study reveals that of the two loop I mutations in the flavin-binding domain (R101W and F105–I106L), F105–I106L has a stronger effect, causing greater structural distortion and weaker packing at the dimerization interface. These mutations reduce the binding energy of flavin and I-Tyr, compared to the wild type, due to a complete loss of R101 crown-like phosphate hydrogen bond in R101W and a partial loss of R101 and R279 hydrogen bonds in F105–I106L. In contrast, the distal I116T mutation has a marginal structural effect, but it alters the solvent-accessible surface area, van der Waals packing, and side-chain flexibility, which may explain its delayed clinical onset. Although the I116T mutation is far from the active site, it strengthens flavin and substrate binding via long-range effects. Protein-folding analysis via the Wako–Saitô–Muñoz–Eaton model shows that the wt-hIYD and R101W fold through the C-terminal region, while F105–I106L and I116T alter the folding pathway. Mutation-specific disruptions can impair electron transfer by altering I-Tyr alignment and flavin ring planarity. These findings reveal how hIYD mutations cause structural, energetic, and catalytic defects linked to hypothyroidism.
Introduction
Hypothyroidism is one of the most common congenital endocrine disorders, appearing in 1 out of 3000 newborns. It is associated with a deficiency of thyroxine (T4) and triiodothyronine (T3) hormones in the human thyroid gland.1,2 Iodine, a naturally scarce trace element, is essential for the biosynthesis of thyroid hormones (TH) and the prevention of hypothyroidism.3 The human iodotyrosine deiodinase (hIYD), a dehalogenase encoded by the DEHAL1 gene in the thyroid gland, recovers iodine from iodotyrosines formed during TH synthesis,4–13 thereby maintaining the iodine supply for continued hormone production.3,14,15 The hIYD enzyme belongs to the NADH oxidase/flavin reductase superfamily16 and catalyses the flavin assisted reductive deiodination of mono- and di-iodotyrosines.4,6,12,16,17 Malfunction of the hIYD enzyme causes severe hypothyroidism accompanied by goiter, accumulation of iodotyrosines in blood and urine, and variable degrees of cognitive impairment due to untreated hypothyroidism.5,7,8,18Various genetic defects are linked with both permanent and transient congenital hypothyroidism.1,2,5,19 In 2008, Moreno et al. reported three homozygous mutations, two missense (R101W and I116T) and one in-frame deletion of three base pairs (F105–I106L), as the molecular basis behind hypothyroidism from clinical studies on patients from different countries.4 Patients carrying these mutations developed severe goitrous hypothyroidism in infancy, with some experiencing cognitive delay due to untreated or late-treated thyroid hormone deficiency. Although parents were healthy, the disease inheritance was recessive, requiring two faulty copies of DEHAL1 to develop symptoms.4,5,8 Some mutations are completely inactivating and cause severe symptoms early in life, such as the F105–I106L deletion, while others, like the I116T variant, retain partial catalytic function and may only manifest as hypothyroidism several years later.4,5 The R101W mutation, although milder than the F105–I106L deletion, has been reported to appear at earlier stages of life.4 This variability also explains why the modern newborn-screening programs sometimes fail to detect the condition at birth, leading to symptom onset later in childhood.5 Notably, another mutation, A220T, identified in 2008, has been shown to cause hypothyroidism in humans and can remain asymptomatic until at least 14 years of age.5,7
In the functionally active form of hIYD, the active site region contains a flavin mononucleotide in its reduced hydroquinone state (FMNhq). Flavin interacts with the enzyme environment at three distinct regions:17,20 its phosphate group forms a crown-like structure by creating salt bridges with residues R100 and R101, its ribose moiety stabilizes itself through hydrogen bonds with S102 and S128* (* denotes residues coming from the alternate monomer), and its isoalloxazine ring is firmly held in place through a network of interactions with the zwitterionic portion of susbtrate I-Tyr, as well as the residues T239 and R104.20,21 The zwitterionic group of substrate I-Tyr anchors into the active site by forming crucial hydrogen bonds with K182, E157, and Y161.17 These key interactions in the hIYD active site enable its unusual catalytic reductive dehalogenation.
The R101W and F105–I106L mutations occur within the FMN-binding region of the hIYD enzyme.5 It is expected that these mutations would strongly affect how the enzyme binds to flavin. In contrast, the I116T mutation is located far from the active site. Traditionally, it has been assumed that mutations – particularly point mutations – exert their influence mainly in the immediate neighborhood of the altered residue,22–25 by changing local packing,26,27 surface charge, or conformational preferences.26 This localized view has guided decades of research in protein engineering,28 disease mechanism studies,29 and drug design.30 However, extensive experimental and theoretical work has shown that this view can be overly simplistic.31–33 Proteins are not rigid assemblies but densely packed, yet fluid-like networks of interactions where residues communicate through both direct contacts and indirect coupling mediated by the backbone and solvent.26,34,35 As a result, a perturbation at one site can be transmitted through this network, subtly or significantly influencing residues far away from the initial mutation site – a phenomenon known as long-range allosteric coupling.23,36–38 The behavior of proteins is thus an emergent property of a complex, correlated network of interactions.
While much of the previous studies on various proteins have focused on how mutations impact the active site or catalytic efficiency of a properly folded protein, mutations can also disrupt the delicate process of protein folding by altering the amino acid sequence, which governs intramolecular interactions.39,40 Even a single amino acid substitution can introduce steric clashes, disrupt hydrogen bonding networks, or change local hydrophobicity, leading to misfolding or aggregation. Misfolded proteins may fail to reach their functional conformation, become prone to degradation, or form toxic aggregates, as observed in diseases like cystic fibrosis or amyloidosis.41,42 Thus, understanding mutations requires not only studying functional deficits of the folded protein but also investigating how they may impair the folding pathway and structural stability from the start. In the absence of experimental unfolding curves or exhaustive atomistic simulations of hIYD, predicting mutation-induced stability changes remains a major challenge.
In this work, we aim to provide a comprehensive, multidimensional analysis of how mutations in the hIYD enzyme affect its structural and functional properties, potentially contributing to hypothyroidism by impairing thyroid hormone synthesis. Specifically, we investigate the long-range dynamic effects of mutational perturbations and how these propagate through the protein's interaction network to influence overall enzyme behavior. Using long-timescale all-atom explicit-solvent classical molecular dynamics (MD) simulations and binding free energy calculations, we characterize the wild-type (wt) enzyme and key mutants (R101W, F105–I106L, and I116T). Our approach integrates residue interaction network analysis to forecast how mutations impact global structural connectivity and long-range communication pathways within the protein. We further analyze the effects of mutations on protein stability, compactness, residue flexibility, active site hydrogen-bonding patterns, solvent accessible surface area (SASA), and substrate binding affinity. Together, these complementary analyses aim to elucidate the complex molecular mechanisms by which specific mutations perturb enzyme function.
Methods
Atomistic MD simulations with explicit solvation
The wild-type enzyme (wt-hIYD) was modeled from the crystal structure of hIYD (PDB ID: 4TTC),20 which features FMNox (in planar, oxidised state) and substrate I-Tyr. The flavin is modelled in its reduced, hydroquinone (FMNhq) state and I-Tyr in its phenolate form.20 The protonation states of all amino acids were determined at pH = 7.4 using the H++ server,43 using an external and internal dielectric constants of 80 and 4, respectively. All Glu, Asp, Lys, and Arg were treated as charged, and all His as neutral, with H147, H210, and H290 in δ-protonated form, whereas H73, H78, H80, H131, H167, H199, and H256 in the ε-protonated state. These assignments were cross-validated using PROPKA344,45 at pH 7.4 (see Table S2 in SI). The protein was solvated in a cubic periodic box (88.2 Å length) of TIP3P water molecules,46 and an ionic concentration of 150 mM was maintained with NaCl. The protein was treated with the AMBER ff14SB force field,47 while ligands are described using the general amber force field (GAFF).48 After performing a 1 µs MD simulation of the wt-hIYD enzyme, a representative snapshot closest to the centroid of the most populated, minimum-free-energy basin on the PCA-based free energy landscape (PC1–PC2) was selected and used as the initial model for generating the mutant systems.We employed the in silico mutagenesis tool of PyMOL49 to model the single-point mutants (R101W and I116T) and used ColabFold50 to model the deletion mutant (F105–I106L). The resulting model exhibited a high per-residue confidence score (pLDDT) of 92.5. The deep-learning-derived structure (ColabFold2) was validated, by comparing it with the template-based prediction, using Modeller51 with a backbone RMSD of 0.6 Å (for residues 82 to 275), which confirms the reliability of the ColabFold2-generated deletion mutant structure (Fig. S11 in SI). The starting structures of the mutants (R101W-hIYD, I116T-hIYD, and F105–I106L-hIYD) were protonated at a physiological pH of 7.4 and prepared for simulation following the same procedure described above.
To capture the anisotropic charge distribution of iodine in I-Tyr, we modeled the σ-hole of I52 with an explicit extra-point (EP) positioned 2.15 Å from the iodine atom along the C–I bond axis with a C–I–EP angle of 180°. The complete EP parameters53 are summarized in Table S1 in SI. The partial atomic charges for the FMN cofactor (FMNhq) and I-Tyr incorporating the extra-point (EP) charges were computed from the restrained electrostatic potential54 using B3LYP-D3/def2-TZVP level of theory.55 The simulation parameters have been benchmarked for different redox states of flavins in our previous work.17 Initial system setup, including topology and coordinate file generation, was accomplished with AmberTools18,56 while all subsequent molecular dynamics (MD) simulations were executed using the AMBER18 suite.56
The modelled structures were first minimized, followed by a gradual heating to 300 K over 1 ns within the NVT ensemble, where tmperature was maintained by a Langevin thermostat57 with a collision frequency of 5 ps−1. For mutants larger number of minimization steps are performed compared to wt-hIYD to ensure the removal of all bad contacts during the mutation of residues. Short-range non-bonded interactions were computed with a 10 Å cutoff, while long-range electrostatics were treated using the Particle Mesh Ewald (PME) method.58 The PME reciprocal-space calculations were performed on a 3D grid with default AMBER settings, which adapt grid points along each dimension according to the box size to achieve standard PME accuracy. After heating, a three-stage equilibration was performed: (1) 1 ns of NVT equilibration with 5 kcal mol−1 Å−2 restraints on the enzyme backbone, (2) 1 ns of NVT equilibration with the same restraint applied to the ligands, and (3) 10 ns of unrestrained equilibration under constant volume. From the equilibrated structures, production MD simulations of 1 µs were carried out for each system in the NPT ensemble at 300 K and 1 bar. A Berendsen barostat59 with isotropic scaling and a pressure relaxation time of 1 ps was used to maintain constant pressure. Despite its known limitations in NPT sampling,60 the Berendsen barostat yields equilibrium density sampling consistent with finite-size expectations, as supported by stable joint enthalpy–volume (H–V) distributions (see Fig. S13 in SI). The integration time step for production runs was set to 2 fs, with non-bonded interactions evaluated within a 10 Å cutoff. All trajectory analyses were conducted using cpptraj,61 gromacs tools, and in-house python scripts. The convergence of the MD simulations was validated using block-averaged backbone RMSD, energy drift analysis, and PCA cosine content, collectively demonstrating stable, well-converged trajectories for all systems (see Fig. S12 in SI).
Analysis protocols
Parameter sensitivity of MM–PBSA calculations were performed on six different models by varying the internal dielectric constant, atomic radii, and ionic strength (Fig. S14 in SI), which reveals sensitivity of ΔGbind to dielectric constant and radii but not ionic strength, consistent with the previous benchmarking.67 Comparative calculations for wt-hIYD and mutant proteins with the same ligand showed no significant change in ΔΔGbind, ensuring reliability of the results in terms of chosen MM–PBSA parameters (Fig. S14 in SI and Table S5). In its standard implementation, MM–PBSA relies on fixed-charge force fields and continuum solvation models, which restrict its ability to explicitly account for electronic polarization and can reduce accuracy, particularly for charged ligands, where strong polarization effects are present.67,68 The MM–PBSA energy across replicates are compared in Fig. S10 in SI, which demonstrates the accuracy and reproducibility of the calculations.
From the contact matrix ![[C with combining macron]](https://www.rsc.org/images/entities/i_char_0043_0304.gif) ij between residue pair i and j an undirected, weighted residue interaction graph G = (V,E) was constructed for each snapshot from the MD simulations, where each vertex (vi ∈ V) corresponds to a residue (i) and an edge (vi,vj) ∈ E with specific edge weight, reflecting the contact weight between residues. The betweenness centrality
ij between residue pair i and j an undirected, weighted residue interaction graph G = (V,E) was constructed for each snapshot from the MD simulations, where each vertex (vi ∈ V) corresponds to a residue (i) and an edge (vi,vj) ∈ E with specific edge weight, reflecting the contact weight between residues. The betweenness centrality
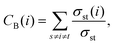 | (1) |
The WSME analyses were carried out using the MATLAB implementation of the WSME model75 (details provided in the SI). Here, we analyzed the folding landscape of a single monomeric domain spanning residues 71–290 using the SSA, DSA, and DSAw/L approximations.75 These approximations encompass a total of 19,34,58,980 microstates each, for the wt-hIYD, R101W, and I116T mutant systems, and 18,99,57,680 microstates for the F105–I106L deletion mutant. Notably, the block-level approximation implemented in the bWSME75 model was not included in the present analysis. The folding free-energy curves of the proteins are constructed at 300 K temperature. For an N-atomic system, the full free energy landscape has a dimensionality of 3N − 6. However, for protein folding problems, the backbone and the sidechain torsional degrees of freedom are mainly effective. This reduces the folding problem to 3–4 conformational degrees of freedom per residue.76 For the 220 residue hIYD protein monomer, this corresponds to approximately 660–880 conformational degrees of freedom. By splitting the protein into N-terminal and C-terminal halves, the folding process can be represented in terms of two reaction coordinates – the number of folded residues in each half – greatly reducing the complexity from a high-dimensional space to a 2D landscape. This projection, however, is only an approximation and cannot fully capture the richness of the underlying high-dimensional space. Such a coarse-graining makes it easier to identify intermediates where only one half is folded, detect sequential folding pathways, and assess coupling between the two halves.
Results and discussion
The flavoenzyme hIYD consists of three domains: an N-terminal trans-membrane domain, an intermediate domain, and a catalytic domain.20 The native enzyme belongs to the NADH oxidase/flavin reductase superfamily and is a homo-dimer with 290 amino acids per monomer. Each monomer comprises eight α-helices, eight β-strands, and six characteristic loops (see Table S4 in SI). The enzyme forms a domain-swapped dimer where α7 helices from each monomer form the dimer interface, which accommodates non-covalent binding of FMNhq. This architecture is essential for catalytic reductive dehalogenation. The genetic mutations occur in the I116T located in the α2-helix, and R101W and F105–I106L located in loop I (flavin-binding region), which plays a key role in flavin's binding and enzyme's activity (Fig. 1).Impact of mutations on conformational dynamics
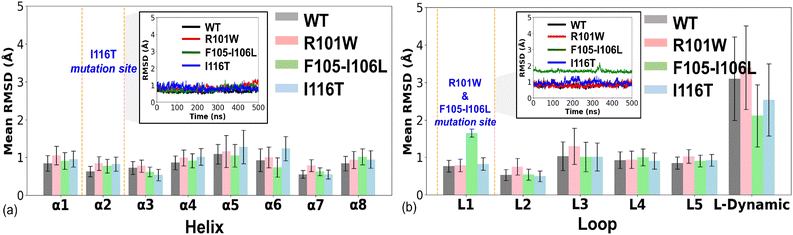
The Cα RMSD of the wt-hIYD and its three mutants is found to be generally low across the protein, indicating overall plasticity of the enzyme, even after mutation. Similarly, mutations lead to minor variation of the radius of gyration, suggesting no influence on the overall folded state of the enzyme (Fig. S2 in SI). In the I116T mutant, the site of mutation (α2-helix) does not show much change in the dynamics, whereas a greater RMSD change occurs in the active site lid helix α5 and in α6, as compared to the wild-type. The R101W and F105–I106L mutations, particularly the deletion mutation, cause a large dynamic effect in the loop I region, see Fig. 2(b). In a previous study, the deletion mutation has been termed as one of the most fatal mutations.4The wt-hIYD exhibits a single dominant equilibrium conformation in the free-energy-landscape spanned by the first two principal components (Fig. S3 in SI). In contrast, the R101W variant shows a more extended and asymmetric distribution (Fig. S3(f) in SI), suggesting increased flexibility or access to alternative conformational states along PC1. The narrow and highly directional distribution in the F105–I106L mutant indicates a constrained motion due to an increased rigidity. Meanwhile, the I116T variant resembles the wt in overall shape and range, with minor alterations (Fig. S3(h) in SI).
Helical composition analysis of hIYD shows that α1–α8 retain 98–100% helical content in wild-type and mutants, with the notable exception of the active site lid helix, α5. MD simulations shows that α5 consists of 61.6% α-helix, 28.6% turn, and 8.9% π-helix in wt-hIYD, indicating its structural flexibility. Previous studies suggest that this conformational flexibility may facilitate the product-release pathway.17 In the apo crystal structure (PDB ID: 4TTB), the electron density is missing for the α5 region and substrate analogs, such as halophenols, can induce conformational changes in this lid helix despite lacking zwitterionic groups.20,77 The R101W and F105–I106L mutations are found to increase α-helical content of α5, whereas I116T shows helical profile similar to wt (Fig. S4 in SI). These observations suggest that while the global structure of the enzyme remains largely intact, the mutations induce distinct local and allosteric changes in flexibility and collective motions, particularly in regions important for function, such as the active site lid and flavin-binding region.
The change in the betweenness centrality (|ΔCB|) in the protein interaction network after mutation is found to be small, indicating a very localized effect of the mutations. The small changes in |ΔCB| induced by R101W mutation decay beyond 40 Å distance from the site of mutation (R101), whereas continue over a larger distance for the deletion mutant (F105–I106L) from the site of mutation (F105) (see Fig. 3(b) and (c)). On the other hand, the I116T mutant does not show any pronounced effect in perturbing the residue interaction network (see Fig. 3(d)). This reinforces the idea that this mutation primarily affects local interactions without altering global network communication, except for the deletion mutation.
Mutations alter residue solvent exposure and local packing, as reflected by changes in solvent accessible surface area (SASA) and van der Waals (vdW) interaction energies. It is noteworthy that the residue burial inferred from SASA reflects the extent of solvent exposure, whereas vdW packing energy reports on the efficiency and favorability of short-range side-chain contacts within the protein interior, and the two are therefore related but not necessarily correlated. In the crystal structure (PDB ID: 4TTC), residues R101, F105, I106, and I116 show SASA values of 31.6, 41.7, 1.8, and 29.6%, respectively, consistent with the wt-hIYD MD simulations, where the mean relative SASA of R101 and I116 are 26.9 ± 3.8% and 20.6 ± 4.6%, indicating partial exposure of R101 while a more buried nature of I116. The F105 remains highly exposed (42.7 ± 8.5%) and I106 is essentially buried. In the R101W mutant, the mean SASA of residue 101 decreases to 17.3 ± 5.0%, indicating increased burial of the bulky tryptophan side chain relative to arginine; this enhanced burial is accompanied by a more favorable vdW interaction energy for W101 compared to R101 (Fig. 4a), reflecting improved local packing due to increased side-chain volume, although the modest change in mean energy suggests a local geometric adjustment rather than substantial energetic stabilization. In contrast, the F105–I106L mutant replaces a partially exposed (F105) and buried (I106) pair with a single leucine residue that becomes fully buried, leading to a net reduction in local SASA but a less favorable vdW interaction energy (Fig. 4b), consistent with reduced side-chain volume and poorer packing efficiency relative to the original aromatic–aliphatic pair. Thus, while mutations modulate solvent exposure, pronounced energetic destabilization arises primarily from volume loss and inefficient packing, particularly in the deletion-type mutant, in agreement with previous observations linking local hydrophobicity changes to mutant destabilization.5
Previous studies have not reported significant differences between the wt-hIYD and the I116T mutant, contributing to the delayed diagnosis of congenital hypothyroidism in affected patients.1,4 In the I116T mutant, residue T116 exhibits a modest increase in solvent exposure, with a mean SASA of 28.3 ± 8.7%. Consistent with this change, radial distribution analysis shows that the hydrophobic CD1 and CG1 atoms of I116 in the wt-hIYD lack structured hydration within ∼5 Å, whereas T116 displays well-defined water density peaks at 2–3 Å and sustained hydration beyond 4 Å, reflecting the introduction of polar interactions by the threonine hydroxyl group (see Fig. S1(a) in SI). This altered local environment is further accompanied by broader side-chain dihedral sampling in T116 relative to the rigid rotameric state of I116 in the wt-hIYD, indicating increased conformational flexibility (Fig. S1b, SI). In the wt-hIYD, I116 forms stabilizing hydrophobic contacts with L233 and V265 from the FMN-binding α7–α8 loop and the β3 strand, respectively, which are weakened upon mutation, leading to reduced vdW interaction energy for V265 (0.12 kcal mol−1) and a minor change in L233 (−0.03 kcal mol−1) interaction. Together, these effects indicate that the I116T mutation subtly perturbs the local hydrophobic core by increasing hydration and flexibility while losing hydrophobic interactions to the FMN-binding domain (Fig. 4c). The observed increase in local hydration and side-chain flexibility suggests a gradual weakening of the otherwise rigid hydrophobic environment. Polar substitutions at distal hydrophobic regions have been shown to alter protein stability and solvent coupling without measurably affecting catalytic function, effectively uncoupling enzyme activity from resilience to thermal or oxidative stress.78–84 We therefore hypothesize that the hydration and packing changes induced by I116T may reduce stress resilience under non-basal conditions which require further experimental validation.
To capture the spatial propagation of these effects, we computed per-residue changes in interaction energy (ΔEvdW = EmutantvdW − EwtvdW) and its fluctuations, see Fig. 4(d)–(f), where the residues are arranged on the basis of their Cα–Cα distance from the site of mutation. For the R101W mutation, vdW energy changes within 6 Å of the mutation site are minor, while fluctuations observed beyond 12 Å arise mainly from the flexible N- and C-termini rather than mutation-induced effects. Locally, R100 exhibits a small destabilization (0.08 kcal mol−1), whereas S102 shows a slight stabilization (−0.04 kcal mol−1), consistent with the disruption of the crown-like hydrogen-bonding network between R101 and the phosphate group of FMNhq, which weakens R100 packing while leaving the ribose-interacting (of flavin) S102 largely unaffected. The mutation also alters interactions within loop I by promoting a hydrogen bond between R104 and flavin in the FMNhq state, resembling the interaction previously observed after dehalogenation in the absence of I-Tyr (PDB ID: 4TTB), albeit at the cost of reduced local packing stability.
The F105–I106L deletion mutation shows a wider impact, affecting up to 20 Å from the site of mutation (Fig. 4(e)). Closer to the mutation site, V103 (−0.20 kcal mol−1) and R104 (−0.16 kcal mol−1) are stabilized, although R104 does not interact directly with flavin in this case. The K182, critical for I-Tyr binding, shows a strong packing stabilization (−0.42 kcal mol−1). The deletion mutation induces destabilization of the dimerization-interface (e.g., Q229 (0.14 kcal mol−1)), resulting in an overall ∼20 kcal mol−1 weakening of dimer packing (Fig. 4h). This mutation enhances local hydrophobic packing but introduces significant strain at the dimerization interface, which may impact hIYD's thermostability, solubility, and cooperative functioning.
The I116T mutation introduces localized effects (Fig. 4(f)). Unlike the F105–I106L deletion mutation, here, no long-range effects are seen. At the mutation site, the packing stabilization is the maximum (−0.67 kcal mol−1). Similar to R101W mutation, I116T mutation does not affect the packing of the dimerization interface (Fig. 4(g) and (i)). Finally, we compared wt and mutant EvdW distributions using a statistical test for all the mutants and corrected for multiple testing with Benjamini–Hochberg false discovery rate (FDR) (α = 0.05). The KS test revealed that >96% of residues exhibited significant differences (FDR < 0.05) in vdW energy distributions between wt and mutants. These results confirm that the observed differences are statistically significant.
The one-dimensional free-energy profile for the folding of wt-hIYD and its mutants is characterized by multiple partially structured states, with the fully folded native state never populated. This can be seen from the fact that the minimum of the free energy profiles occurs at a reaction coordinate value of 0.9, which corresponds to just 198 folded residues (out of 220), see Fig. S5 in SI. The folding pattern remains similar for different mutants to the wild-type protein, except for the R101W mutation, where the unfolded and the partially folded states have higher free energy than the rest. An intermediate state (denoted as I in Fig. S5 in SI) with a partially folded structure appears at 0.48 fraction of the structured residues.
For the wt-hIYD system, residue folding probability analysis indicates downhill folding pathway. Initially, residues 112–197 may fold to form a stable intermediate, comprising helices α2–α6 with their connecting loops, along with the β1 and β2 strands. Later, folding extend to the other active site regions like dynamic loop, helix α7, the α7–α8 loop, helix α8, and the β3 strand, accompanied by partial destabilization of some earlier-folded elements, particularly parts of helices α2 and α4 (see Fig. S6(a) in SI). Throughout this process, the α1 helix remains only partially folded, while the α1–α2 loop stays unfolded.
In the R101W mutant, the initial folded α2–α3 region gets destabilized and remains 70–80% folded, while the dynamic loop (residues 198–212) and β3 strand regions begin to fold earlier than in the wt. Because R101 is located in the α1–α2 loop (loop I) adjacent to helix α2, we hypothesize that substitution with bulky tryptophan perturbs local packing near the α2–α3 region, reducing its stability and thereby promoting earlier engagement of other residues during folding, at an increased energetic cost. The F105–I106L mutant and I116T exhibits distinct folding behaviors at the initial stage with the α2–α3 region remains only 30–40% folded and completely unfolded, respectively, altering the initial folding core of helices α2–α6 and their connecting loops (see Fig. S6(c) and (d) in SI).
Consistent with these trends, the WSME model predicts that folding in the R101W systems proceeds through slightly higher-energy intermediates dominated by C-terminal folding, whereas the F105–I106L and I116T mutants follow a distinct, lower-energy folding pathway initiated from the N-terminal region (Fig. 5 and Fig. S5, SI), comared to wt. Together, these results suggest that while the active-site region (α4–α6 and associated loops) acts as a conserved folding hotspot in hIYD, mutations that disrupt the early α2–α3 folding core can redirect the folding pathway and alter its energetic landscape, a hypothesis that can be tested experimentally.
Although the employed WSME model provides a topology-driven view of folding in hIYD monomer, it simplifies the system by neglecting inter-domain and membrane interactions, that may alter the folding landscape. Extending the model to multi-domain85,86 through extended-WSME86 models with virtual linkers is challenging, as these methods require a priori knowledge of linker placement and therefore necessitate further validation using experimental folding data, which, to the best of our knowledge, are not yet available. However, our predicted folding model including hotspots for wt-hIYD as well as mutants can be verified with future NMR relaxation measurements.
In the R101W mutant, electrostatic stabilization from R101 is noticeably reduced due to disruption of the crown-like salt-bridge network, leading to less favorable FMNhq binding, although partial compensation occurs through residues such as R279 and R104 (Fig. S7, SI). Substrate binding remains largely intact, but I-Tyr shows a modest change, consistent with altered FMNhq positioning. In the F105–I106L mutant, flavin binding is still primarily supported by R100 and R101; however, the strong electrostatic contribution from R104 seen in the wt-hIYD is diminished. These changes indirectly impact I-Tyr binding through rearrangement of electrostatic contributions from surrounding residues, including K207*, R104, and R100. In contrast, the I116T mutant shows flavin interactions largely unchanged relative to the wt-hIYD, while I-Tyr experiences slightly more favorable contributions from K182 and R104. Overall, these results indicate that mutations within the FMN-binding domain (R101W and F105–I106L) alter the phosphate crown and ribose stabilization network, affecting FMN binding and indirectly modulating substrate interactions, whereas the distal mutation (I116T) preserves cofactor binding while subtly enhancing substrate interactions (Fig. 6).
 | ||
| Fig. 6 Binding free energy of FMNhq (blue) and I-Tyr (pink) in wild-type and three mutants (R101W, F105–I106L, and I116T) of hIYD enzyme. | ||
Hydrogen-bond analysis (Table 1) provides complementary support to the binding free energy results. In R101W, the loss of R101 disrupts the phosphate crown, with S102 and T239 partially compensating for FMN stabilization, while R279 also contributes. F105–I106L shows a redistribution of flavin H-bonds: R100 and S102 remain dominant, R101 participation decreases, and T238 and R279 emerge as significant stabilizers. In I116T, FMN interactions are largely preserved, with R100, R101, and S102 dominating, and minor contributions from R104 and S128* supporting ribose and isoalloxazine contacts. Across all systems, I-Tyr H-bonds remain highly occupied with FMNhq, and key residues such as Y161 and A130* continue to anchor the substrate, indicating a robust cofactor-substrate network.
| Residue | % Occupancy | |||
|---|---|---|---|---|
| wt | R101W | F105–I106L | I116T | |
| FMNhq interactions | ||||
| R100 | 96.5 | 98.7 | 97.2 | 97.1 |
| R101 | 73.8 | 0 | 66.5 | 84.2 |
| S102 | 97.3 | 95.6 | 94.8 | 98.6 |
| R104 | 28.2 | 41.2 | 0 | 29.4 |
| T238 | 7.4 | 0 | 70.8 | 4.5 |
| T239 | 56.6 | 74.6 | 70.8 | 69.2 |
| R279 | 17.7 | 48.7 | 66.9 | 71.6 |
| S128* | 5.4 | 3.4 | 19.8 | 23.0 |
| H131* | 5.6 | 3.1 | 0.5 | 0 |
| I-Tyr | 100 | 100 | 100 | 100 |
| I-Tyr interactions | ||||
| E157 | 20.1 | 35.6 | 18.6 | 22.1 |
| Y161 | 95.4 | 97.0 | 95.9 | 96.6 |
| K182 | 16.4 | 17.4 | 19.8 | 18.6 |
| A130* | 77.8 | 78.9 | 74.9 | 80.4 |
| FMN | 100 | 100 | 100 | 100 |
Although the mutations do not drastically alter the active-site architecture and overall enzyme dynamics, they may subtly fine-tune the relative orientation of FMNhq and I-Tyr, thereby modulating catalytic turnover. In the wild-type (wt) enzyme, T239 maintains a stable interaction with the N5 atom of flavin (Fig. S8 in SI), which is essential for stabilizing the semiquinone state during single-electron transfer.87 The side-chain oxygen forms a dominant hydrogen bond at 3 Å, with a secondary shoulder peak around 3.7 Å. In the R101W mutant, this distribution shifts, showing a dominant peak near 4 Å, indicating a weakened interaction. The F105–I106L mutant, by contrast, preserves a similar interaction between the T239 side chain and flavin as observed in the wt system. For the I116T mutation, the distribution largely resembles that of the wt, but an alternate conformation appears at 6.2 Å. This alternative state may become more pronounced after multiple catalytic turnovers, potentially destabilizing the semiquinone form during catalysis. Notably, interactions with the backbone nitrogen are stronger in the mutants compared to the wt enzyme, whereas the recognition interaction between FMNhq and I-Tyr remains unaffected by mutation (Fig. S8 in SI). The position of the I-Tyr relative to flavin deviates modestly from the wt-hIYD conformation, with RMSD values of 0.64 Å, 0.83 Å, and 0.65 Å for the R101W, F105–I106L, and I116T mutants, respectively (Fig. S16 in SI).
For an effective reductive dehalogenation, the C–I bond of I-Tyr needs to be parallel to the C4a–N5 bond of flavin. In the crystal structure (PDB ID: 4TTC), the I–C–C4a–N5 dihedral is ∼22° for catalytically inactive FMNox state.17,20 It reduces to ∼ 8° for the catalytically active form of flavin (FMNhq), assisted by the puckering of the isoalloxazine ring. In all three mutants, this value remains close to 12–15°, although flavin remains in its hydroquinone state (Fig. 7 and Fig. S9 in SI). R101W shows the highest misalignment value of ∼14°. The puckering of flavin's isoalloxazine ring is also largely affected in the mutants. The butterfly angle17,88,89 (N1–N10–N5–C6 dihedral) remains ∼160° in the wt-hIYD enzyme. In all three mutants (R101W, F105–I106L, and I116T), this dihedral is near 174°, indicating planarity of the isoalloxazine ring (see Fig. 7(b), (d) and Fig. S9(b) and (d) in SI). A planar isoalloxazine ring reduces the aromatic stacking interaction between I-Tyr and FMNhq state, which may subsequently affect the through-space electron transfer rate in hIYD.
 | ||
| Fig. 7 The free energy profile projected along C4-N5-C6 angle and I-C-C4a-N5 dihedral angle of FMNhq and I-Tyr in the (a) wild-type, (c) R101W systems. Free energy profile along C4-N5-C6 angle and N1-N10-N5-C6 dihedral angle in the (b) wild-type and (d) R101W systems. The crystal structure values of these dihedrals are 22° and 177.2°, respectively, as shown by a black dot in the figure. The isoalloxazine planar conformation as present in the crystal structure (PDB ID: 4TTC),20 is also shown. The crossing point of the magenta lines shows the minimum energy basin in the free energy landscape. The interactions between the isoalloxazine ring of FMNhq and T239 and I-Tyr are shown in the side panel. | ||
QM-cluster calculations on the minimum free-energy structures (extracted from the most populated basin identified by PCA analysis) support these structural features. All catalytically relevant residues17,20 (K182, E157, Y161, R104, S102, S128, G129, A130*, and T239) were included in the QM region (209 atoms), treated at the DLPNO–CCSD(T)/def2-SVP level of theory and implicit solvation (dielectric constant 4.0) to approximate the protein environment. Topological analysis of the electron density within the quantum theory of atom-in-molecules (QTAIM) framework identifies distinct critical points at the flavin–I-Tyr interface in the wt-hIYD system. A bond critical point (BCP) is located along the interaction line connecting the iodine atom of I-Tyr and the N5 atom of the flavin isoalloxazine ring, and a ring critical point (RCP) between the pyrimidine portion of the isoalloxazine ring and the aromatic ring of I-Tyr, consistent with a stacked aromatic arrangement at the interface (Fig. 8). Although BCP and RCP are both present in I116T, the BCP reflects a weak interaction between iodine and flavin. The BCP was found missing in the R101W, while both BCP and RCP were missing in F105–I106L mutant. Hence, there is absence of a shared interaction topology and a weakened interaction between flavin and I-Tyr in the mutants.
Conclusions
In humans, congenital hypothyroidism can arise from three mutations (R101W, F105–I106L, and I116T) in the DEHAL1 gene, which encodes iodotyrosine deiodinase (hIYD), an enzyme required for thyroid functioning. While clinical studies have described the effects of these variants, their molecular consequences remain unclear. Using atomistic simulation, energetic decomposition, network analysis, and coarse-grained folding models, we have examined how these mutations perturb hIYD structure, dynamics, folding, and catalysis.The deletion mutation F105–I106L in loop I perturbs the residue interaction network to a longer distance from the site of mutation and decreases the vdW interaction energy (around 20 kcal mol−1) in the dimeric interface of the hIYD enzyme. On the other hand, I116T mutation, located in the distal α2 helix, leaves overall plasticity intact, while subtly remodelling the local hydrophobic core by increasing hydration, sidechain flexibility, and weakening stabilizing hydrophobic contacts with the FMN-binding domain. These modest structural and solvent-coupling changes are likely to compromise activity under stress conditions. The folding analysis by Wako–Saitô–Muñoz–Eaton (WSME) modeling shows that R101W follows a more energetically demanding pathway than wild type, whereas F105–I106L and I116T fold more readily. All mutants preserve the α2–α6 folding core, but destabilize the α2–α3 region to different extents: partially folded in R101W (70–80%), mostly unfolded in F105–I106L (30–40%), and completely unfolded in I116T. The flavin binding is impaired in R101W due to loss of R101 hydrogen bond and in F105–I106L due to disrupted hydrophobic contacts, but it is preserved in I116T.
Subtle geometric and electronic perturbations of mutation are seen from structural and QM analyses. The mutants show an increased misalignment of the I-Tyr C–I bond and the flavin C4a–N5 axis and a near-planar isoalloxazine ring, reducing aromatic stacking and through-space electron transfer. These structural effects are accompanied by mutation-dependent weakening of key stabilizing interactions, particularly involving T239 and flavin N5. The QTAIM analysis in in wt-hIYD reveals a well-defined interaction topology at the flavin–I-Tyr, including an iodine–N5 bond critical point and a ring critical point, consistent with efficient electronic coupling. The mutants lack these topological features to different extent, indicating a progressive loss of electronic communication despite minimal global structural disruption. We hope that these simulation-based molecular insights will motivate targeted biochemical and biophysical experiments, thereby advancing the medicinal community's understanding of DEHAL1-linked hypothyroidism and thyroid hormone metabolism.
Author contributions
S. K. conceived the original idea, curated the data, performed the formal analysis and investigation, developed the methodology and software, contributed resources, validated the results, prepared the visualizations, and wrote the original draft. B. G. contributed to data curation and formal analysis. S. M. acquired funding, administered the project, supervised the work, contributed to validation, and revised the manuscript.Conflicts of interest
There are no conflicts to declare.Data availability
Data for this article, including scripts and simulation files are available at github with DOI: https://doi.org/10.5281/zenodo.17615229.Supplementary information is available. See DOI: https://doi.org/10.1039/d5cp04422g.
Acknowledgements
S. K. acknowledges the Prime Minister's Research Fellowship, the Ministry of Education, Government of India, and Associate Membership of Royal Society of Chemistry for support. B. G. acknowledges IIT Kharagpur for the support. S. M. acknowledges the support from SERB, DST, Govt. of India (CRG/2022/004088 and SR/FST/CSII-026/2013). The authors thank Prof. Chitrangada Das Mukhopadhyay for useful discussion. This work used the resources of the Paramshakti supercomputing facility of IIT Kharagpur, established under the National Supercomputing Mission of the Government of India and supported by CDAC, Pune.References
- J. C. Moreno, Horm. Res., 2003, 60, 96–102 Search PubMed.
- X. Fan, C. Fu, Y. Shen, C. Li, S. Luo, Q. Li, J. Luo, J. Su, S. Zhang and X. Hu, et al. , Clin. Chim. Acta, 2017, 468, 76–80 CrossRef PubMed.
- C. González-Guerrero, M. Borsò, P. Alikhani, Y. Alcaina, F. Salas-Lucia, X.-H. Liao, J. Garca-Giménez, A. Bertolini, D. Martin and A. Moratilla, et al. , Thyroid, 2023, 33, 752–761 CrossRef PubMed.
- J. C. Moreno, W. Klootwijk, H. van Toor, G. Pinto, M. D'Alessandro, A. Lèger, D. Goudie, M. Polak, A. Grüters and T. J. Visser, N. Engl. J. Med., 2008, 358, 1811–1818 CrossRef PubMed.
- J. C. Moreno and T. J. Visser, Mol. Cell. Endocrinol., 2010, 322, 91–98 CrossRef PubMed.
- Z. Sun, Q. Su and S. E. Rokita, Arch. Biochem. Biophys., 2017, 632, 77–87 CrossRef PubMed.
- G. Afink, W. Kulik, H. Overmars, J. de Randamie, T. Veenboer, A. van Cruchten, M. Craen and C. Ris-Stalpers, J. Clin. Endocrinol. Metab., 2008, 93, 4894–4901 CrossRef PubMed.
- A. Burniat, I. Pirson, C. Vilain, W. Kulik, G. Afink, R. Moreno-Reyes, B. Corvilain and M. Abramowicz, J. Clin. Endocrinol. Metab., 2012, 97, E1276–E1283 CrossRef PubMed.
- A. Iglesias, L. Garca-Nimo, J. A. C. de Juan and J. C. Moreno, Best Pract. Res., Clin. Endocrinol. Metab., 2014, 28, 151–159 CrossRef PubMed.
- A. Phatarphekar and S. E. Rokita, Prot. Sci., 2016, 25, 2187–2195 CrossRef PubMed.
- S. R. Thomas, P. M. McTamney, J. M. Adler, N. LaRonde-LeBlanc and S. E. Rokita, J. Biol. Chem., 2009, 284, 19659–19667 CrossRef PubMed.
- J. Hu, Q. Su, J. L. Schlessman and S. E. Rokita, Prot. Sci., 2019, 28, 68–78 CrossRef PubMed.
- F. Ismail-Beigiy and M. Rahimifar, J. Clin. Endocrinol. Metab., 1977, 44, 499–506 CrossRef PubMed.
- K. Krause, S. Karger, O. Gimm, S.-Y. Sheu, H. Dralle, A. Tannapfel, K. W. Schmid, C. Dupuy and D. Fuhrer, Eur. J. Endocrinol., 2007, 156, 295–301 Search PubMed.
- A. Yoshihara, Y. Luo, Y. Ishido, K. Usukura, K. Oda, M. Sue, A. Kawashima, N. Hiroi and K. Suzuki, Endocr. J., 2019, 66, 349–357 CrossRef PubMed.
- J. E. Friedman, J. A. Watson, D. W.-H. Lam and S. E. Rokita, J. Biol. Chem., 2006, 281, 2812–2819 CrossRef PubMed.
- S. Karmakar and S. Mishra, Biochemistry, 2024, 63, 3310–3323 CrossRef PubMed.
- L. Chiovato, F. Magri and A. Carlé, Adv. Ther., 2019, 36, 47–58 CrossRef PubMed.
- L. Persani and M. Bonomi, Best Pract. Res., Clin. Endocrinol. Metab., 2017, 31, 255–263 CrossRef PubMed.
- J. Hu, W. Chuenchor and S. E. Rokita, J. Biol. Chem., 2015, 290, 590–600 CrossRef PubMed.
- N. Ingavat, J. M. Kavran, Z. Sun and S. E. Rokita, Biochemistry, 2017, 56, 1130–1139 CrossRef PubMed.
- R. Blake, S. T. Hess and J. Nicholson-Tuell, J. Mol. Evol., 1992, 34, 189–200 CrossRef PubMed.
- N. Rajasekaran, S. Suresh, S. Gopi, K. Raman and A. N. Naganathan, Biochemistry, 2017, 56, 294–305 CrossRef PubMed.
- A. N. Naganathan, J. Phys. Chem. B, 2013, 117, 4956–4964 CrossRef PubMed.
- A. N. Naganathan, Curr. Opin. Struct. Biol., 2019, 54, 1–9 CrossRef PubMed.
- G. R. Bowman and P. L. Geissler, J. Phys. Chem. B, 2014, 118, 6417–6423 CrossRef PubMed.
- J. C. Gaines, W. W. Smith, L. Regan and C. S. O'Hern, Phys. Rev. E, 2016, 93, 032415 CrossRef PubMed.
- F. H. Arnold, Curr. Opin. Biotechnol, 1993, 4, 450–455 CrossRef PubMed.
- S. Stefl, H. Nishi, M. Petukh, A. R. Panchenko and E. Alexov, J. Mol. Biol., 2013, 425, 3919–3936 CrossRef PubMed.
- J. A. Bikker, N. Brooijmans, A. Wissner and T. S. Mansour, J. Med. Chem., 2009, 52, 1493–1509 CrossRef PubMed.
- L. S. Bigman and Y. Levy, J. Phys. Chem. B, 2018, 122, 11450–11459 Search PubMed.
- M. W. Clarkson and A. L. Lee, Biochemistry, 2004, 43, 12448–12458 CrossRef PubMed.
- F. Xu, A. V. Colasanti, Y. Li and W. K. Olson, Nucl. Acids Res., 2010, 38, 6872–6882 Search PubMed.
- G. R. Bowman and P. L. Geissler, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 11681–11686 CrossRef PubMed.
- K. H. DuBay, G. R. Bowman and P. L. Geissler, Acc. Chem. Res., 2015, 48, 1098–1105 CrossRef PubMed.
- E. Guarnera and I. N. Berezovsky, Curr. Opin. Struc. Biol., 2019, 56, 18–27 CrossRef PubMed.
- S.-R. Tzeng and C. G. Kalodimos, Nature, 2009, 462, 368–372 CrossRef PubMed.
- R. B. Fenwick, S. Esteban-Martn and X. Salvatella, Eur. Biophys. J., 2011, 40, 1339–1355 Search PubMed.
- F. Henot, E. Rioual, A. Favier, P. Macek, E. Crublet, P. Josso, B. Brutscher, M. Frech, P. Gans and C. Loison, et al. , Nat. Commun., 2022, 13, 7601 CrossRef PubMed.
- M. D. Geschwind, Continuum, 2015, 21, 1612–1638 CrossRef PubMed.
- D. Fraser-Pitt and D. O’Neil, Future Sci. OA, 2015, 1, FSO57 CrossRef PubMed.
- S. Ventura, Future Sci. OA, 2015, 1, FSO38 CrossRef PubMed.
- J. C. Gordon, J. B. Myers, T. Folta, V. Shoja, L. S. Heath and A. Onufriev, Nucleic Acids Res., 2005, 33, W368–W371 CrossRef PubMed.
- C. R. Søndergaard, M. H. Olsson, M. Rostkowski and J. H. Jensen, J. Chem. Theo. Comput., 2011, 7, 2284–2295 Search PubMed.
- M. H. Olsson, C. R. Søndergaard, M. Rostkowski and J. H. Jensen, J. Chem. Theo. Comput., 2011, 7, 525–537 Search PubMed.
- P. Mark and L. Nilsson, J. Phys. Chem. A, 2001, 105, 9954–9960 Search PubMed.
- J. A. Maier, C. Martinez, K. Kasavajhala, L. Wickstrom, K. E. Hauser and C. Simmerling, J. Chem. Theory Comput., 2015, 11, 3696–3713 Search PubMed.
- J. Wang, R. M. Wolf, J. W. Caldwell, P. A. Kollman and D. A. Case, J. Comput. Chem., 2004, 25, 1157–1174 CrossRef PubMed.
- S. Yuan, H. S. Chan and Z. Hu, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2017, 7, e1298 Search PubMed.
- M. Mirdita, K. Schütze, Y. Moriwaki, L. Heo, S. Ovchinnikov and M. Steinegger, Nat. Methods, 2022, 19, 679–682 CrossRef PubMed.
- A. Šali and T. L. Blundell, J. Mol. Biol., 1993, 234, 779–815 CrossRef PubMed.
- R. S. Nunes, D. Vila-Viçosa and P. J. Costa, J. Am. Chem. Soc., 2021, 143, 4253–4267 CrossRef PubMed.
- M. A. Ibrahim, J. Phys. Chem. B, 2012, 116, 3659–3669 CrossRef PubMed.
- C. I. Bayly, P. Cieplak, W. Cornell and P. A. Kollman, J. Phys. Chem., 1993, 97, 10269–10280 CrossRef.
- S. Grimme, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2011, 1, 211–228 Search PubMed.
- D. Case, R. Walker, T. Cheatham, C. Simmerling, A. Roitberg, K. Merz, R. Luo and T. Darden, Amber 18, Univ, 2018 Search PubMed.
- R. L. Davidchack, R. Handel and M. Tretyakov, J. Chem. Phys., 2009, 130, 234101 CrossRef PubMed.
- R. E. Isele-Holder, W. Mitchell and A. E. Ismail, J. Chem. Phys., 2012, 137, 174107 Search PubMed.
- H. J. Berendsen, J. V. Postma, W. F. Van Gunsteren, A. DiNola and J. R. Haak, J. Chem. Phys., 1984, 81, 3684–3690 Search PubMed.
- Q. Ke, X. Gong, S. Liao, C. Duan and L. Li, J. Mol. Liq., 2022, 365, 120116 Search PubMed.
- D. R. Roe and T. E. Cheatham III, J. Chem. Theory Comput., 2013, 9, 3084–3095 Search PubMed.
- S. Genheden and U. Ryde, J. Comp. Chem., 2010, 31, 837–846 Search PubMed.
- S. Genheden and U. Ryde, Expert Opin. Drug Discovery, 2015, 10, 449–461 Search PubMed.
- E. Wang, H. Sun, J. Wang, Z. Wang, H. Liu, J. Z. Zhang and T. Hou, Chem. Rev., 2019, 119, 9478–9508 CrossRef PubMed.
- I. Massova and P. A. Kollman, Perspect. Drug Discovery Des., 2000, 18, 113–135 Search PubMed.
- B. R. Miller III, T. D. McGee Jr, J. M. Swails, N. Homeyer, H. Gohlke and A. E. Roitberg, J. Chem. Theory Comput., 2012, 8, 3314–3321 CrossRef PubMed.
- S. Genheden and U. Ryde, Expert Opin. Drug Discovery, 2015, 10, 449–461 Search PubMed.
- S. Wang, X. Sun, W. Cui and S. Yuan, Front. Pharmacol., 2022, 13, 1018351 Search PubMed.
- A. Hagberg, P. J. Swart and D. A. Schult, Exploring network structure, dynamics, and function using NetworkX, Los alamos national laboratory (lanl) technical report, 2007 Search PubMed.
- A. Hagberg and D. Conway, https://networkx.github.io, 2020, 1–48.
- R. T. McGibbon, K. A. Beauchamp, M. P. Harrigan, C. Klein, J. M. Swails, C. X. Hernández, C. R. Schwantes, L.-P. Wang, T. J. Lane and V. S. Pande, Biophys. J., 2015, 109, 1528–1532 CrossRef PubMed.
- H. Wako and N. Saitô, J. Phys. Soc. Jpn., 1978, 44, 1939–1945 Search PubMed.
- V. Muñoz and W. A. Eaton, Proc. Natl. Acad. Sci. U. S. A., 1999, 96, 11311–11316 CrossRef PubMed.
- A. N. Naganathan, J. Chem. Theo. Comput., 2012, 8, 4646–4656 Search PubMed.
- S. Gopi, A. Aranganathan and A. N. Naganathan, Curr. Res. Struct. Biol., 2019, 1, 6–12 Search PubMed.
- P. Minary and M. Levitt, J. Mol. Biol., 2008, 375, 920–933 Search PubMed.
- H. C. Greenberg, A. Majumdar, E. K. Cheema, A. Kozyryev and S. E. Rokita, Biochemistry, 2024, 63, 2225–2232 CrossRef PubMed.
- S. Akanuma, C. Qu, A. Yamagishi, N. Tanaka and T. Oshima, FEBS Lett., 1997, 410, 141–144 Search PubMed.
- S. Bershtein, W. Mu and E. I. Shakhnovich, Proc. Natl. Acad. Sci. U. S. A., 2012, 109, 4857–4862 Search PubMed.
- E. M. Adams, S. Pezzotti, J. Ahlers, M. Ruttermann, M. Levin, A. Goldenzweig, Y. Peleg, S. J. Fleishman, I. Sagi and M. Havenith, JACS Au, 2021, 1, 1076–1085 Search PubMed.
- K. Ghosh, A. M. de Graff, L. Sawle and K. A. Dill, J. Phys. Chem. B, 2016, 120, 9549–9563 CrossRef PubMed.
- I. Liguori, G. Russo, F. Curcio, G. Bulli, L. Aran, D. Della-Morte, G. Gargiulo, G. Testa, F. Cacciatore and D. Bonaduce, et al. , Clin. Interventions Aging, 2018, 757–772 Search PubMed.
- F. H. Niesen, H. Berglund and M. Vedadi, Nat. Prot., 2007, 2, 2212–2221 Search PubMed.
- G. Morreale de Escobar, M. J. Obregón and F. Escobar del Rey, Eur. J. Endocrinol., 2004, 151, U25–U37 Search PubMed.
- K. Ooka, R. Liu and M. Arai, Molecules, 2022, 27, 4460 CrossRef PubMed.
- T. Inanami, T. P. Terada and M. Sasai, Proc. Natl. Acad. Sci. U. S. A., 2014, 111, 15969–15974 CrossRef PubMed.
- A. Mukherjee and S. E. Rokita, J. Am. Chem. Soc., 2015, 137, 15342–15345 CrossRef PubMed.
- S. Nakai, F. Yoneda and T. Yamabe, Theor. Chem. Acc., 1999, 103, 109–116 Search PubMed.
- R. K. Kar, A. Miller and M. Mroginski, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2022, 12, e1541 Search PubMed.
| This journal is © the Owner Societies 2026 |