 Open Access Article
Open Access ArticleEvolutionary aspect of spike glycoprotein's conformational dynamics
Wentao
Xu
 ,
Tianyu
Guo
and
Haibin
Su
,
Tianyu
Guo
and
Haibin
Su
 *
*
Department of Chemistry, Laboratory of Theoretical and Computational Chemistry, The Hong Kong University of Science and Technology, Hong Kong, China. E-mail: haibinsu@ust.hkhaibinsu@ust.hk; Tel: +86 852 2358 7388
First published on 21st January 2026
Abstract
The conformational transition of the receptor binding domain (RBD) of the SARS-CoV-2 spike (S) protein from the “closed” to the “open” state is an important dynamic event influencing the infectivity of the virus. However, how accumulating mutations reshape these conformational dynamics during viral evolution remains to be clarified. In this study, we observed the motion of the RBD of fully glycosylated S protein trimers from wild type to Omicron (BA.2, BA.4&5) variants by using all-atomic molecular dynamics simulations. Our study indicates that, although a fully converged free energy landscape in all motion directions was not obtained, analysis of the molecular dynamics trajectories reveals significant differences in the RBD motional patterns among variants due to mutations. Studies have shown that specific surface mutations of the Omicron variant exhibit a strong coupling effect with glycan dynamics. This effect remodels the “glycan gate” conformation mediated by N165 and N343 glycans and subsequently regulates the motion of the RBD. These findings elucidate the molecular mechanism between viral surface mutations and glycan dynamic regulation, providing a new theoretical basis for understanding the evolutionary adaptability and biological function of the virus.
1. Introduction
Since the outbreak of the COVID-19 pandemic, significant progress has been made in understanding the evolution of SARS-CoV-2, with the spike protein remaining the focus of investigation.1,2 The dynamic conformational transition of the spike (S) protein—wherein its receptor binding domain (RBD) shifts from a protected “down” or “closed” state to an accessible “up” or “open” state—is a pivotal event governing viral infectivity.3 This is because RBD is directly responsible for binding to the host angiotensin-converting enzyme 2 (hACE2) receptor, the first step for viral entry.4,5 A vast amount of research tries to understand virus evolution impact, such as how mutations on the S protein surface alter its electrostatics and hydrophobicity to modulate receptor affinity and antibody evasion.6–9 However, how the conformational dynamics of the spike protein—as an independent, optimizable trait—is reshaped throughout viral evolution remains a critical unresolved question.Various experimental and computational methods have been employed to characterize the dynamic behavior of the S protein. Among them, the single-molecule fluorescence resonance energy transfer experiment observed and quantified the dynamic equilibrium of the RBD, revealing that viral variants (such as D614G) significantly shifted the conformational equilibrium towards an open state that is more conducive to binding than the wild-type (WT) strain.10,11 Meanwhile, cryo-electron microscopy studies also provided structural evidence that the D614G variants favor a more open conformation by disrupting an inter-protomer salt-bridge contact present in the WT strain.12–15 Other key single mutations, like K417N, were also shown to impact the stability of the S protein's open state.16 Furthermore, the hydrogen deuterium exchange mass spectrometry technique was used to analyse the flexible region of the S protein and further indicated that the Delta variant's spike tended to have an open conformation, but it was also found that the RBD of the early Omicron variant unexpectedly leaned more towards the closed conformation.17
Molecular dynamics (MD) simulation combined with enhanced sampling methods and free energy calculation is a powerful tool for studying the conformational dynamics of S proteins at the atomic level.18–24 For example, free energy calculations via 2D umbrella sampling have been used to explore the impact of a single mutation on this transition process, providing insights into the dynamic evolution of spike proteins, even when the specific mutations studied did not become prevalent variants.19 Other work focused on the influence of distant residues, demonstrating that the analysis of residue motion synergy from MD trajectories can provide profound insights for predicting the outcomes of distant mutations.18 Furthermore, research on the surface glycans of the spike protein has innovatively revealed their key regulatory role in the conformational dynamics of the RBD.25,26 Studies combining conventional molecular dynamics simulations with experimental validation first demonstrated that N-glycans at sites N165 and N234 play an essential structural role by stabilizing the RBD in the open conformation.26 Subsequently, a distinct mechanistic contribution was uncovered through weighted ensemble simulations, which showed that the N343 glycan acts as a “glycan gate”, actively facilitating the closed-to-open transition by interacting with the RBD surface.25 These findings highlight two different aspects of glycans' influence: stabilization of the open state and facilitation of the opening transition. This understanding has been quantitatively refined by a 2D free energy surface constructed using replica exchange umbrella sampling, which showed that glycans increase the energy barrier to opening and highlighted the roles of N165 and N343 in stabilizing both the up and down states.21 Further investigations have explored how the specific type and size of glycans at these key sites (N165, N234, N343) fine-tuned the stability of the open RBD, and examined the evolutionary implications of changes in the glycan shield topology, such as the loss of the N370 glycosylation site in SARS-CoV-2.22 These important studies have established that factors, from key mutations such as D614G to specific glycans such as N165, can play crucial regulatory roles in the S protein conformational dynamics.
However, virus evolution is not a simple accumulation of isolated factors.27,28 The challenge of current research lies in how these mutations can work in synergy with the dynamic regulation of glycans to jointly reshape the conformational landscape of the S protein of the variants of concern. Solving this challenge is precisely the key to answering the fundamental question mentioned in the beginning. In this study, we used all-atom metadynamic simulations of the fully glycosylated SARS-CoV-2 spike protein trimer to investigate how the RBD opening mechanism has evolved across major variants (from wild type to Omicron BA.2 and BA.4/5), and to analyze how surface mutations and glycan dynamics collectively shape this process, particularly at N165 and N343, reshapes the conformational landscape of the spike protein. Through this analysis, we aim to elucidate the connection between surface mutations and glycan dynamics, thereby establishing a mechanistic basis for understanding the structural and functional evolution of the virus.
2. Methods
2.1. Construction of fully glycosylated spike protein structures for VOCs
Following the framework developed by Woo et al.29 for fully glycosylated SARS-CoV-2 spike protein models, we constructed complete spike protein structures for the variants of concern (VOCs) studied herein. The precise set of amino acid mutations defining each VOC (Delta, BA.2, and BA.4/5) was determined based on the consensus data provided by authoritative genomic surveillance resources, specifically the covariants.org project, which tracks shared mutations across major viral lineages.With these defined sequences, we then generated the structures through homology modeling. Initially, utilizing wild-type models from the CHARMM-GUI COVID-19 Archive29,30 as templates, we incorporated the VOC-specific amino acid mutations using the MODELLER software package.31 For glycosylation patterns, we integrated recent comprehensive analyses by Shajahan et al.,32 which comparatively examined N- and O-glycosylation sites across wild-type, Delta, and Omicron spike proteins. In contrast to the wild-type strain, the N17 site in Delta and the N17, N74, and N149 sites in Omicron BA.2 were maintained in non-glycosylated states. Detailed information of mutations and glycan sequences are stored in 3_Additional_Information.xlsx. These structural modifications were implemented through the glycan reader & modeler module of the CHARMM-GUI online platform. The initial coordinates for the fully-glycosylated Spike protein system were prepared using the CHARMM-GUI. The protonation states of all titratable residues were assigned assuming a physiological pH of 7.0. The system was then solvated in a periodic boundary box with TIP3P water molecules, ensuring a minimum distance (20 Å) between the protein and the box edges. Potassium and chloride ions were added to neutralize the total system charge and to achieve a physiological salt concentration of 0.15 M. Both the protein and the attached N-glycans were described using the all-atom CHARMM36 force field.
2.2. System pre-equilibration and glycan relaxation
All subsequent MD simulations were performed using GROMACS 2023.2-plumed-2.10.0-dev version.33,34 To pre-equilibrate the glycosylated protein system and facilitate glycan relaxation, we initially performed energy minimization using the steepest descent algorithm with a convergence criterion of maximum force less than 1000 kJ mol−1 nm−1. Subsequently, the system entered a preliminary equilibration phase using the NVT ensemble, during which the Nose–Hoover thermostat35 maintained the system temperature at 300 K while position restraints were applied to both the protein backbone and side chains. Following this, a temperature equilibration phase was conducted, maintaining the NVT ensemble with position restraints applied exclusively to α-carbon, with temperature regulated at 300 K via the Nose–Hoover thermostat. To further enhance glycan relaxation, simulated annealing was implemented under the NVT ensemble with position restraints on protein heavy atoms, comprising four annealing cycles over 10 ns, each cycle linearly increasing temperature from 300 K to 600 K before cooling back to 300 K. Finally, the system underwent a terminal equilibration phase transitioning to the NPT ensemble, utilizing the Nose–Hoover thermostat and Parrinello–Rahman barostat36 to maintain temperature and pressure at 300 K and 1 bar, respectively. Throughout this protocol, van der Waals interactions were treated using force-switch correction with a cutoff radius of 1.2 nm, electrostatic interactions were handled via the Particle Mesh Ewald (PME) methodology,37–39 and hydrogen bond lengths were constrained using the LINCS algorithm40 across all simulation stages.2.3. Well-tempered metadynamics simulation
To enhance the sampling of open-to-closed conformational transitions, four collective variables (CVs) were selected and grouped into two categories. The first group comprises geometric descriptors modified based on Fallon's scheme: (i) CV1, RBD opening angle (see light blue dashes in Fig. 1A and B), and (ii) CV2, RBD torsional angle (see pink dashes in Fig. 1A and B). The second group includes root mean square deviation (RMSD)-based descriptors: (iii) RMSD-XA, the RMSD to a reference open structure, and (iv) RMSD-XB, the RMSD to a reference closed structure. In the metadynamics setup, CV1 and CV2 (the geometry CVs mentioned in the main text) were used for adding Gaussian hills. The hill widths were set as 0.08 rad, 0.05 rad, 1 Å and 1 Å. The hill height was set as 0.25 kcal mol−1 with a bias factor of 10. In the simulations, upper and lower wall restraints were applied to the collective variables RMSD-XA, RMSD-XB, CV1, and CV2 to confine their values within specified ranges (−1 to 30 for RMSD-XA and RMSD-XB, 0.8 to 2 for CV1, and −0.7 to 1 for CV2, in the unit of Å or rad) using harmonic potentials with a force constant of 500 kcal mol−1 Å2. Additionally, an upper wall restraint with a force constant of 1000 kcal mol−1 Å2 was applied to the RMSD of the RBD to limit its value below 2.5 Å to maintain RBD's structural stability.We conducted three independent repeated simulations for each variant, with each simulation lasting for 400 nanoseconds (400 ns * 3 replicas). Particularly for the BA.2 variant that exhibits a unique open mechanism, we performed an additional five repeated simulations to further verify its behavior. A detailed PLUMED file has been uploaded to 3-METAD.dat in the github repo.
2.4. Free energy surface reconstruction and reweighting
Herein, we employed the well-tempered metadynamics simulations considering the extensive and all-atom on the fully glycosylated S protein trimer to map the free energy landscape (FEL) governing RBD motion.41 The time-dependent bias potential accumulated during the metadynamics simulation is presented: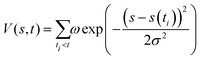 | (1) |
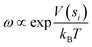 | (2) |
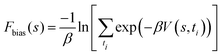 | (3) |
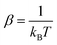 is the inverse thermal energy and V(s,ti) is the bias potential deposited at time.
is the inverse thermal energy and V(s,ti) is the bias potential deposited at time.
2.5. Low-energy state occupancy ratio
To quantitatively compare the thermodynamic stability and conformational diversity across variants, we calculated the occupancy from the RMSD-based FELs. The methodology first identifies a “low-energy conformational ensemble” by selecting all points on the FEL with a relative free energy below a set threshold (e.g., <8.0 kT). Within this ensemble of stable states, each conformation is then classified: it is defined as “open” if its RMSD to the closed reference is greater than its RMSD to the open reference, and “closed” otherwise. The occupancy of the open state is the percentage of conformations within this low-energy ensemble that are classified as “open”, with the closed state occupancy calculated analogously. These values, presented in Fig. 2 of the main text, serve as a direct measure of the system's thermodynamic preference for the open versus closed states, effectively integrating information about both the depth and breadth of the corresponding energy basins.2.6. Hydrogen bond contact analysis
Hydrogen bonds were identified using the mdtraj.baker_hubbard function, which implements the Baker-Hubbard geometric criteria.43 A hydrogen bond was defined to exist if the distance between the donor and acceptor heavy atoms was lower than 2.5 Å and the angle formed by donor-hydrogen-acceptor was greater than 120 degrees. Detailed information can be found in supporting script: 3_find_hbond_between.py.2.7. Calculations on glycan surface charge distribution
To investigate the electrostatic properties of the N165-glycan and validate our interpretation of its interaction with the RBD surface, we performed density functional theory (DFT) calculations. A representative structure of the sialylated N165-glycan was extracted from our MD trajectories. Single-point energy calculations were then carried out on this structure using the Gaussian 16 software package. The calculations employed the B3LYP hybrid functional44 with the 6-31G(d,p) basis set.45,46 The resulting molecular electrostatic potential (MEP)47 was subsequently mapped onto the electron density surface to visualize the charge distribution across the glycan, as presented in the SI (Fig. S5).3. Result and discussion
3.1. Comparative analysis strategy and semi-quantitative interpretation for free energy landscapes
To describe the conformational transition process of RBD (the red part in Fig. 1A) from the closed to the open state, we defined a set of geometric collective variables to define its opening angle and torsion dihedral (Fig. 1A). For metadynamics simulation, a strict convergence criterion of free energy calculation is that the CVs exhibit diffusion behavior along the sampled space over the simulation time. However, for a complex system such as the spike protein trimer, which is large and highly glycosylated, achieving ideal complete convergence within limited computing resources is extremely challenging. Our trajectory analysis indicates that some CVs have not yet achieved complete free diffusion throughout the conformational space. Therefore, interpreting the energy barrier height (e.g., the apparent energy barrier of![[1 with combining tilde]](https://www.rsc.org/images/entities/char_0031_0303.gif) 0.9 kcal mol−1) on the reconstructed free energy surface (such as the WT protein as shown in Fig. 1B) as accurate kinetic parameters is inappropriate.
0.9 kcal mol−1) on the reconstructed free energy surface (such as the WT protein as shown in Fig. 1B) as accurate kinetic parameters is inappropriate.
To this end, we adopted a more robust analysis strategy, shifting the focus from the pursuit of absolute accuracy in energy values to capturing the topological differences in the conformational energy landscapes among different variants. We interpret the “low energy” regions as corresponding to thermodynamically stable and highly sampled conformations. Therefore, FEL is regarded as a powerful tool for analysing the motion patterns of RBD in MD trajectories, rather than a quantitative indicator for summarising the conformational changes of RBD. Such an analytical strategy has also been demonstrated in previous studies on the dynamics of RNA conformation.48 Based on this, the minimum energy path connecting the closed state and the open state is defined as the main pathway of conformational opening (opening path, see Fig. S3). We believe that even if the free energy values do not fully converge, significant changes in the FEL topological structure (such as the position, shape, and relative depth of the energy wells) can reliably reveal the differences in conformational intermediates and dynamic behaviors among different variants. To achieve standardised comparisons among variants, we further project FEL onto a two-dimensional space defined by the conformations and the RMSD of the reference “open” and “closed” states (WT shown in Fig. 1C). This projected space constitutes the basis for our cross-variant comparative analysis (Fig. 2).
Based on RMSD FELs, the focus of our analysis is on the occupancy ratio of the “open” and “closed” states. This metric is defined as the proportion of low-energy conformations within each energy basin (the state boundaries are shown in Fig. S1) on the RMSD free energy surface. It directly quantifies the thermodynamic preference of the system for each macroscopic state and the diversity of the conformational ensemble. The quantitative occupancy ratio results of each variant have been labelled on the respective free energy surfaces in Fig. 2. By focusing on the relative changes in state occupancy rather than the absolute energy barriers, the optimisation mechanism of the dynamic behavior of the S protein under evolutionary pressure can be reliably elucidated.
3.2. Divergence of the RBD opening mechanism among the SARS-CoV-2 variants
The comparison of these FELs indicates that the open mechanism of RBD has undergone significant changes as the virus evolves to the Omicron variant. The transition from WT to Delta reflects a conservative optimization strategy. This is well demonstrated in the RMSD FELs of the wild type (Fig. 1C) and the Delta variant (Fig. 2, upper left), showing similar topological features: two distinct closed and open energy basins separated by clear energy barriers. However, there are also significant differences between the two: the low energy occupancy ratio of the open state (occupancy ratio) in the Delta variant (over 80%) has significantly increased from only 7.4% in the WT. This indicates that the evolutionary process of the protein has greatly expanded the conformational set accessible within the open region, suggesting that the Delta variant has a higher thermodynamic advantage in accessing diverse open states. This represents a very effective evolutionary strategy, consistent with previous experimental observations regarding variants containing the D614G mutation.13,15Although variants like Delta, which are pre-Omicron variants, evolved through an optimized conservative kinetic pathway, the emergence of Omicron introduced a major innovation – a new path on the open pathway that features the appearance of an intermediate state. As shown in the geometric FEL of BA.2 (Fig. S2), a stable intermediate state that is far away from both closed and open states was formed, and it can be inferred that this intermediate state significantly reduced the transition barrier from closed toward open. However, this kinetic advantage is deceptive and is accompanied by a thermodynamic cost. Although this intermediate state is easy to enter, the system is difficult to escape from this “trap” to reach the fully open conformation. This is quantitatively reflected in the low energy occupancy rate of the open state of BA.2, being approximately 9.8% (Fig. 2), which is comparable to that of the original WT strain. This mechanism difference provides a reasonable explanation for the puzzling finding observed in the HDX-MS17 experiment, where the early Omicron variants have a lower tendency to be open compared to their predecessors. The conformational dynamics of BA.4&5 represent an integrated feature, resolving the dilemma of BA.2. The key difference from BA.2 lies in the topological structure of its energy landscape: the geometric FEL of BA.4&5 exhibits a smoother energy slope (Fig. S3), which facilitates a gradual transition to the open state. The occupancy rate of its open state has climbed to over 60% (Fig. 2), also strongly supporting this. By combining the kinetic accessibility provided by the intermediate state driving path with a thermodynamically favourable landscape conducive to extensive exploration, the BA.4&5 variants achieve truly efficient RBD opening. This suggests that the overall topological features of the landscape during the optimization of dynamic properties, including the smoothness of the pathway and the presence of intermediate-state traps, perhaps play a more critical role in determining the efficiency of opening. Next, we will attempt to explain the changes in the opening mechanism of different variants through protein structure analysis.
3.3. Glycan dynamics governing the RBD opening mechanism swift
The interaction patterns between key mutations and the surrounding glycans are then investigated to uncover the molecular underpinnings of these distinct energy landscapes. As shown in Fig. 1D–F, during the opening process of the RBD, various mutation sites (e.g., Q493R, E484A, S477N, and L452R) form new and stable contacts with adjacent surface glycans (colored in blue and orange) at different transitional stages. This finding suggests that, given the significant changes in polarity and charge introduced by these amino acid substitutions, they interfere with previously identified glycan–RBD interaction patterns and replace the established RBD opening mechanism in the WT and Delta variant, with a new, mutation-induced mechanism.The representative opening pathways, illustrated in the right panel of Fig. 2, provide primary structural insights into the dynamic differences, the core of which lies in the dynamic behavior of key glycans surrounding the RBD. In both Delta and BA.4&5, as the RBD opens upward, the N165-glycan forms dynamic contacts with the N343-glycan on the lower side of RBD, establishing a “bridge” structure, a key feature of the “with N165–N343 glycan contact” mechanism illustrated in Fig. 4. This observation of N165 and N343 glycans cooperatively stabilizes the open state, building upon and refining previous findings that highlighted their individual and collective roles in modulating the S protein's conformational energy landscape.21,26 In contrast, during the opening process in BA.2, the N165-glycan remains adhered to the RBD surface and fails to establish effective interactions with the N343-glycan, defining the “without N165–N343 glycan contact” mechanism (Fig. 4). The absence of this bridging structure likely contributes significantly to the thermodynamic instability observed in the BA.2 open state. This suggests that viral amino acid mutations can reshape RBD opening dynamics through allosteric modulation of the protein–glycan interaction network, indicating that the well-known “glycan gating” phenomenon plays a central role in this evolutionary trajectory.21,25,26
To investigate the molecular mechanisms driving the differential glycan dynamics among the variants, we focused on key amino acid mutations on the RBD surface. Detailed analyses of the glycan–RBD interactions for the Delta, BA.2, and BA.4&5 variants are organized in Fig. S3, Fig. 3, and Fig. S4, respectively. These complex dynamic data can be distilled into two distinct, variant-specific RBD opening mechanisms, which are illustrated as intuitive models in Fig. 4. These models, classified as “with N165–N343 glycan contact” and “without N165–N343 glycan contact”, demonstrate how mutations directly regulate the motional pathway of the N165-glycan by remodeling the local amino acid–glycan interaction network. The quantitative evidence supporting these mechanistic models comes from our detailed analysis of interaction frequencies, presented as histograms in Fig. S6. This figure quantifies the evolving dynamic contacts between key mutated residues and the N165-glycan (top, blue) or N343-glycan (bottom, orange) along the RBD opening pathway, providing the underlying data for the mechanisms shown in Fig. 4.
In the BA.2 variant, the Q493R mutation at the edge of the receptor-binding motif (RBM) imparts a stage-dependent regulatory effect on the N165-glycan, showcasing a mechanism of interaction during the RBD opening process. This finding reveals that a specific mutation alters spike dynamics through a glycan interaction, thereby reinforcing the view that glycans play important functional roles beyond simply providing surface shielding. In the early stages of RBD opening, the newly introduced arginine residue (R493) acts as a potent electrostatic anchor. Its positive charge forms a specific and strong salt bridge with the negatively charged terminal sialic acid of the N165-glycan. This initial interaction functions as a transient but strong tether, significantly restricting the glycan's conformational freedom and guiding its initial movement along with the RBD surface (see Fig. S5).
However, as the RBD transitions towards a more open state, this specific, localized tethering effect diminishes. The increasing distance and altered orientation between R493 and the glycan weaken the initial salt bridge. The interaction then shifts to a weaker, more diffuse electrostatic attraction. This secondary interaction is governed by the overall electrostatic complementarity between the positive surface potential of the RBD, enhanced by the R493 mutation, and the glycan surface (see Fig. S5A–C). Our additional simulation on BA.2 spike with oligomannose-N165 glycan (see Fig. S5E) demonstrates that this principle holds for different glycoforms, including both sialylated complex and oligomannose types, which provides a structural basis for understanding why the N165 glycan can tolerate significant structural diversity across variants while still performing its crucial role in modulating RBD dynamics. This transition from a strong, localized interaction to a weaker, delocalized one is supported by our electrostatic potential surface calculations and analysis (see Fig. S5).
Consequently, the Q493R mutation does not permanently anchor the glycan but rather modulates its dynamics in a two-fold manner: an initial strong restriction followed by a gentler, more global guidance. This effect collectively ensures the N165-glycan remains associated with the RBD surface, preventing the large-scale swinging motions required for bridge formation. This overall coupled motion is further corroborated by a principal component analysis of the N165-glycan and the RBD in the BA.2 variant (Fig. 3, lower right corner), which reveals a high degree of correlation between their respective centers of mass. To test the reliability of this proposed tethering mechanism, we performed two additional, independent metadynamics simulations of the BA.2 variant: one with re-initialized glycan conformations and another where the N165 glycan was remodelled to the oligomannose-type (Man5) also prevalent in Omicron. Crucially, the key features of this N165-glycan restriction and the resulting thermodynamic instability of the open state remained consistent across all simulations. This consistency suggests that the observed mechanism is a feature of the system and not an artefact of a specific initial model.
The Delta and BA.4&5 variants tried to employ a similar strategy: by introducing a positively charged arginine residue via the L452R mutation to increase the surface's positive charge and attract the N165-glycan. This successfully anchors the root of the glycan, thereby balancing the competing needs of stabilizing the RBD intermediate state and maintaining tip flexibility for N343 bridge formation (see Fig. S3 and S4's lower right corner). However, they diverge in the stabilization effect on the RBD opening intermediate state, revealing how BA.4&5 selectively inherited and refined traits from its predecessor. In the Delta variant, while the L452R mutation anchors the root of the N165-glycan, the presence of the negatively charged glutamic acid at position 484 (E484) contributes to a more diffuse negative electrostatic potential on the surrounding RBD surface. This creates a subtle, yet persistent, unfavorable interaction with the overall partial negative charge of the N165-glycan, thereby weakening the global affinity between the glycan and the RBD. This attenuated attraction makes it difficult for the Delta variant to stabilize a distinct intermediate state during the RBD opening process, unlike what is observed in its predecessors.
BA.4&5 meticulously refines this mechanism. While retaining L452R, it eliminates the unfavorable electrostatic interference through the E484A mutation, leading to a markedly enhanced contact between the N165-glycan and the RBD surface. This strengthened and uniformly distributed interaction network strikes an exceptional balance: it is strong enough to efficiently stabilize the critical intermediate state along the opening pathway, yet it avoids the excessive tether-like restriction of BA.2's Q493R. By retaining the native Q493, whose neutral side chain forms weaker hydrogen bonds compared to the potent electrostatic attraction from arginine's positively charged group, it preserves the conformational freedom necessary for the glycan tip to form the bridge. By resolving the electrostatic hindrance encountered by Delta, BA.4&5 resolves the dilemma between kinetic accessibility and thermodynamic stability, demonstrating a fitness advantage.
3.4. Mutational fine-tuning of the dynamic competition network at the site 477/478
The consequences of these two distinct opening mechanisms, as modeled in Fig. 4, are directly manifested in the formation or dissolution of a dynamic competition network at site 477/478. When the N165-glycan possesses bridging capability (as in Delta and BA.4&5, see in Fig. S3 and S4), its free-swing tip frequently contacts site 477/478, where it encounters the N343-glycan. This establishes a dynamic “N165-glycan – site 477/478 – N343-glycan” triangular competitive relationship. The contact frequency data in Fig. S6 (with site 477/478 highlighted in red) imply the formation of this complex network, which constitutes the core structural and dynamic feature enabling the “glycan bridge” mechanism. Conversely, when the motion of the N165-glycan is restricted (as in BA.2, no 477/478 interaction, see in Fig. 3), it is tethered by R493 and cannot reach the 477/478 region for effective contact. The direct consequence is the dismantling of the triangular relationship. Data in Fig. S6 shows that the contact frequency between N165 and site 477/478 is almost negligible in BA.2, which in turn allows the interaction between the N343-glycan and site 477/478 to become markedly more stable and frequent, as it is no longer subject to competition from N165.Furthermore, the persistent mutational trend observed at site 477/478 throughout the evolution of SARS-CoV-2 implies a critical function for this region in modulating glycan dynamics. We note that the T478K mutation, which became dominant in the Delta variant, and the S477N mutation, prevalent in the Omicron family, could both further modulate the triangular competition network by altering the local chemical environment. Chemically, the T478K mutation substitutes a neutral, polar threonine with a positively charged, long-chain lysine. We hypothesize that this newly introduced positive charge center could more effectively “capture” the polar glycan chains via long-range electrostatic attraction, thereby enhancing the stickiness of this site as a network node and potentially affecting the frequency and duration of N165 or N343 glycan contacts. On the other hand, the S477N mutation replaces a serine with an asparagine; although both are neutral polar residues, asparagine possesses a different hydrogen bond donor–acceptor pattern (due to its amide group) and a longer side chain.
Such a seemingly subtle adjustment could potentially optimize the conformational landscape of the hydrogen-bond network with the glycan chain, achieving a delicate “fine-tuning” of the interaction. This leads to an intriguing hypothesis: these subsequent mutations may not serve to create entirely new mechanisms. Instead, their role could be to seek a more advantageous dynamic equilibrium for RBD opening and closing by optimizing interaction strengths at critical nodes within the pre-existing dynamic framework. This hypothesis provides a new perspective for understanding variant evolution strategies, and its specific mechanisms await further experimental and computational validation. This “fine-tuning” hypothesis is also supported by another of our findings. Our simulations show that in BA.2, the R346 residue is also involved in electrostatic anchoring of the N165 glycan. Notably, the R346T mutation frequently occurs in subsequent, more widely spread Omicron subvariants (such as BA.2.75), strongly suggesting that this site is also a key fine-tuning node for the virus's adaptive evolution to optimize glycan dynamics.
4. Conclusions
In conclusion, this study, through extensive molecular dynamics simulations, elucidates two distinct evolutionary pathways for the RBD opening mechanism of the SARS-CoV-2 spike protein. Pre-Omicron variants, such as Delta, followed a conservative refinement strategy, enhancing the propensity for opening without altering the core mechanism. The emergence of Omicron BA.2, however, marked a radical innovation that achieved kinetic accessibility by introducing a functional intermediate state, albeit at the cost of the “open” state's stability. Ultimately, the BA.4&5 variant represents a culmination of these strategies by integrating the kinetic advantages of BA.2 with the thermodynamic stability of Delta to achieve a strong RBD opening capability. We further propose an atomic basis for this evolutionary logic: key amino acid mutations (L452R/Q493R) gate the RBD opening dynamics by modulating the motional patterns of the N165-glycan, which in turn controls the formation of a critical “glycan bridge”.We also recognize that constructing a precise and comprehensive description for a system as large and complex as the spike glycoprotein is a significant computational challenge. On one hand, although we cross-validated the simulations using multiple standard methods, fully satisfying the ideal standard of multiple reversible transitions for slow, large-scale conformational changes remains a Frontier challenge. On the other hand, our current analysis focuses on direct glycan–protein interactions. We fully acknowledge that water-mediated hydrogen bonds, or “water bridges”, play a crucial role in the interaction landscape and conformational dynamics. Indeed, we observed traces of structural water molecules within our trajectories (as shown in Fig. S5), yet fully characterizing their role requires additional extensive work. A comprehensive analysis of these water bridges, which involves dissecting the intricate coupling between solvent, flexible protein loops, and glycans, represents a substantial undertaking that we are pursuing as an independent, in-depth follow-up study. Therefore, we position the FELs in this study as semi-quantitative physical pictures that reveal key mechanistic divergences. We believe that, building upon this foundation, future work incorporating both longer simulation times and explicit analysis of solvent effects will be able to refine the absolute values of the energy barriers, thereby deepening our understanding of the dynamic mechanisms of viral evolution.
This work provides insights into the physicochemical principles underlying viral adaptive evolution, highlighting how distant residues can allosterically reshape conformational energy landscapes to alter function. Crucially, the detailed mechanisms suggested here offer a robust framework for interpreting perplexing virological data. For instance, the context-dependent RBD dynamics we describe, which are intrinsically tied to the full trimer conformation, provide a plausible mechanistic framework for recently reported discrepancies between the binding affinities of isolated RBDs and intact spike trimers for emerging variants like JN.1.49 By illuminating these sophisticated evolutionary strategies, our findings offer new targets and refined principles for the rational design of future antiviral drugs and vaccines.
Conflicts of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.Data availability
The MD outputs are available from Zenodo at https://doi.org/10.5281/zenodo.16876667. Scripts and glycan sequence information mentioned in Methods section (including 3_Additional_Information.xlsx, 3_find_hbond_between.py) has been uploaded in repo github.com/Venthaul/Spike_Open-Close. Any other input files or trajectory files are available on request.Minimum energy paths in free energy landscapes, and interactions between key residues and adjacent glycans can be found in supplementary information (SI). See DOI: https://doi.org/10.1039/d5cp04391c.
Acknowledgements
We acknowledge the data contributors and the CHARMM-GUI Archive for sharing the structural information. We are grateful to undergraduate students Leung Cheuk Fung Alvin and Lin Sijia from the Department of Chemistry of the Hong Kong University of Science and Technology for their inspiring discussions during the project initiation period. We are grateful to Professor Prabal Kumar Maiti and Professor Z. P. Dong for thought-provoking discussions in the course of this work, which is supported in part by the Society of Interdisciplinary Research (SOIRÉE) and the Research Grants Council of Hong Kong (16305623).References
- W. T. Harvey, A. M. Carabelli, B. Jackson, R. K. Gupta, E. C. Thomson, E. M. Harrison, C. Ludden, R. Reeve and A. Rambaut, COVID-19 Genomics UK (COG-UK) Consortium, S. J. Peacock and D. L. Robertson, Nat. Rev. Microbiol., 2021, 19, 409–424 CrossRef CAS PubMed.
- A. C. Walls, Y.-J. Park, M. A. Tortorici, A. Wall, A. T. McGuire and D. Veesler, Cell, 2020, 181, 281–292.e6 CrossRef CAS PubMed.
- Y. Cai, J. Zhang, T. Xiao, H. Peng, S. M. Sterling, R. M. Walsh Jr., S. Rawson, S. Rits-Volloch and B. Chen, Science, 2020, 369, 1586–1592 CrossRef CAS PubMed.
- D. Wrapp, N. Wang, K. S. Corbett, J. A. Goldsmith, C.-L. Hsieh, O. Abiona, B. S. Graham and J. S. McLellan, Science, 2020, 367, 1260–1263 CrossRef CAS PubMed.
- J. Lan, J. Ge, J. Yu, S. Shan, H. Zhou, S. Fan, Q. Zhang, X. Shi, Q. Wang, L. Zhang and X. Wang, Nature, 2020, 581, 215–220 CrossRef CAS PubMed.
- Y. Liu, J. Liu, K. S. Plante, J. A. Plante, X. Xie, X. Zhang, Z. Ku, Z. An, D. Scharton, C. Schindewolf, S. G. Widen, V. D. Menachery, P.-Y. Shi and S. C. Weaver, Nature, 2022, 602, 294–299 CrossRef CAS PubMed.
- M. McCallum, N. Czudnochowski, L. E. Rosen, S. K. Zepeda, J. E. Bowen, A. C. Walls, K. Hauser, A. Joshi, C. Stewart, J. R. Dillen, A. E. Powell, T. I. Croll, J. Nix, H. W. Virgin, D. Corti, G. Snell and D. Veesler, Science, 2022, 375, 864–868 CrossRef CAS PubMed.
- A. Aggarwal, S. Naskar, N. Maroli, B. Gorai, N. M. Dixit and P. K. Maiti, Phys. Chem. Chem. Phys., 2021, 23, 26451–26458 RSC.
- S. H. Kim, F. L. Kearns, M. A. Rosenfeld, L. Votapka, L. Casalino, M. Papanikolas, R. E. Amaro and R. Freeman, Cell Rep. Phys. Sci., 2023, 4, 101346 Search PubMed.
- M. Lu, P. D. Uchil, W. Li, D. Zheng, D. S. Terry, J. Gorman, W. Shi, B. Zhang, T. Zhou, S. Ding, R. Gasser, J. Prévost, G. Beaudoin-Bussières, S. P. Anand, A. Laumaea, J. R. Grover, L. Liu, D. D. Ho, J. R. Mascola, A. Finzi, P. D. Kwong, S. C. Blanchard and W. Mothes, Cell Host Microbe, 2020, 28, 880–891.e8 Search PubMed.
- Z. Yang, Y. Han, S. Ding, W. Shi, T. Zhou, A. Finzi, P. D. Kwong, W. Mothes and M. Lu, mBio, 2022, 13, e03227–21 CrossRef CAS PubMed.
- T.-J. Yang, P.-Y. Yu, Y.-C. Chang, K.-H. Liang, H.-C. Tso, M.-R. Ho, W.-Y. Chen, H.-T. Lin, H.-C. Wu and S.-T. D. Hsu, Nat. Struct. Mol. Biol., 2021, 28, 731–739 CrossRef CAS PubMed.
- L. Yurkovetskiy, X. Wang, K. E. Pascal, C. Tomkins-Tinch, T. P. Nyalile, Y. Wang, A. Baum, W. E. Diehl, A. Dauphin, C. Carbone, K. Veinotte, S. B. Egri, S. F. Schaffner, J. E. Lemieux, J. B. Munro, A. Rafique, A. Barve, P. C. Sabeti, C. A. Kyratsous, N. V. Dudkina, K. Shen and J. Luban, Cell, 2020, 183, 739–751.e8 CrossRef CAS PubMed.
- D. Weissman, M.-G. Alameh, T. De Silva, P. Collini, H. Hornsby, R. Brown, C. C. LaBranche, R. J. Edwards, L. Sutherland, S. Santra, K. Mansouri, S. Gobeil, C. McDanal, N. Pardi, N. Hengartner, P. J. Lin, Y. Tam, P. A. Shaw, M. G. Lewis, C. Boesler, U. Sahin, P. Acharya, B. F. Haynes, B. Korber and D. C. Montefiori, Cell Host Microbe, 2021, 29, 23–31.e4 Search PubMed.
- S. M.-C. Gobeil, K. Janowska, S. McDowell, K. Mansouri, R. Parks, V. Stalls, M. F. Kopp, K. Manne, D. Li, K. Wiehe, K. O. Saunders, R. J. Edwards, B. Korber, B. F. Haynes, R. Henderson and P. Acharya, Science, 2021, 373, eabi6226 CrossRef CAS PubMed.
- Q. Geng, Y. Wan, F.-C. Hsueh, J. Shang, G. Ye, F. Bu, M. Herbst, R. Wilkens, B. Liu and F. Li, eLife, 2023, 12, e74060 Search PubMed.
- V. Calvaresi, A. G. Wrobel, J. Toporowska, D. Hammerschmid, K. J. Doores, R. T. Bradshaw, R. B. Parsons, D. J. Benton, C. Roustan, E. Reading, M. H. Malim, S. J. Gamblin and A. Politis, Nat. Commun., 2023, 14, 1421 CrossRef CAS PubMed.
- D. Ray, L. Le and I. Andricioaei, Proc. Natl. Acad. Sci. U. S. A., 2021, 118, e2100943118 CrossRef CAS PubMed.
- L. Fallon, K. A. A. Belfon, L. Raguette, Y. Wang, D. Stepanenko, A. Cuomo, J. Guerra, S. Budhan, S. Varghese, C. P. Corbo, R. C. Rizzo and C. Simmerling, J. Am. Chem. Soc., 2021, 143, 11349–11360 Search PubMed.
- V. Ovchinnikov and M. Karplus, J. Phys. Chem. B, 2023, 127, 8565–8575 Search PubMed.
- Y. T. Pang, A. Acharya, D. L. Lynch, A. Pavlova and J. C. Gumbart, Commun. Biol., 2022, 5, 1170 Search PubMed.
- M. I. Zimmerman, J. R. Porter, M. D. Ward, S. Singh, N. Vithani, A. Meller, U. L. Mallimadugula, C. E. Kuhn, J. H. Borowsky, R. P. Wiewiora, M. F. D. Hurley, A. M. Harbison, C. A. Fogarty, J. E. Coffland, E. Fadda, V. A. Voelz, J. D. Chodera and G. R. Bowman, Nat. Chem., 2021, 13, 651–659 CrossRef CAS PubMed.
- R. A. Mansbach, S. Chakraborty, K. Nguyen, D. C. Montefiori, B. Korber and S. Gnanakaran, Sci. Adv., 2021, 7, eabf3671 Search PubMed.
- M. Gur, E. Taka, S. Z. Yilmaz, C. Kilinc, U. Aktas and M. Golcuk, J. Chem. Phys., 2020, 153, 075101 CrossRef CAS PubMed.
- T. Sztain, S.-H. Ahn, A. T. Bogetti, L. Casalino, J. A. Goldsmith, E. Seitz, R. S. McCool, F. L. Kearns, F. Acosta-Reyes, S. Maji, G. Mashayekhi, J. A. McCammon, A. Ourmazd, J. Frank, J. S. McLellan, L. T. Chong and R. E. Amaro, Nat. Chem., 2021, 13, 963–968 Search PubMed.
- L. Casalino, Z. Gaieb, J. A. Goldsmith, C. K. Hjorth, A. C. Dommer, A. M. Harbison, C. A. Fogarty, E. P. Barros, B. C. Taylor, J. S. McLellan, E. Fadda and R. E. Amaro, ACS Cent. Sci., 2020, 6, 1722–1734 CrossRef CAS PubMed.
- D. M. Lyons and A. S. Lauring, Viruses, 2018, 10, 407 CrossRef PubMed.
- R. Sanjuán, A. Moya and S. F. Elena, Proc. Natl. Acad. Sci. U. S. A., 2004, 101, 15376–15379 CrossRef PubMed.
- H. Woo, S.-J. Park, Y. K. Choi, T. Park, M. Tanveer, Y. Cao, N. R. Kern, J. Lee, M. S. Yeom, T. I. Croll, C. Seok and W. Im, J. Phys. Chem. B, 2020, 124, 7128–7137 CrossRef CAS PubMed.
- J. Lee, X. Cheng, J. M. Swails, M. S. Yeom, P. K. Eastman, J. A. Lemkul, S. Wei, J. Buckner, J. C. Jeong, Y. Qi, S. Jo, V. S. Pande, D. A. Case, C. L. Brooks, A. D. MacKerell, J. B. Klauda and W. Im, J. Chem. Theory Comput., 2016, 12, 405–413 CrossRef CAS PubMed.
- B. Webb and A. Sali, Curr. Protoc. Bioinformatics, 2016, 54, 5.6.1–5.6.37 Search PubMed.
- A. Shajahan, L. E. Pepi, B. Kumar, N. B. Murray and P. Azadi, Sci. Rep., 2023, 13, 10053 Search PubMed.
- M. J. Abraham, T. Murtola, R. Schulz, S. Páll, J. C. Smith, B. Hess and E. Lindahl, SoftwareX, 2015, 1–2, 19–25 Search PubMed.
- M. Bonomi, D. Branduardi, G. Bussi, C. Camilloni, D. Provasi, P. Raiteri, D. Donadio, F. Marinelli, F. Pietrucci, R. A. Broglia and M. Parrinello, Comput. Phys. Commun., 2009, 180, 1961–1972 Search PubMed.
- D. J. Evans and B. L. Holian, J. Chem. Phys., 1985, 83, 4069–4074 Search PubMed.
- M. Parrinello and A. Rahman, J. Appl. Phys., 1981, 52, 7182–7190 CrossRef CAS.
- T. Darden, D. York and L. Pedersen, et al. , J. Chem. Phys., 1993, 98, 10089 Search PubMed.
- U. Essmann, L. Perera, M. L. Berkowitz, T. Darden, H. Lee and L. G. Pedersen, J. Chem. Phys., 1995, 103, 8577–8593 CrossRef CAS.
- A. Toukmaji, C. Sagui, J. Board and T. Darden, J. Chem. Phys., 2000, 113, 10913–10927 Search PubMed.
- B. Hess, H. Bekker, H. J. C. Berendsen and J. G. E. M. Fraaije, J. Comput. Chem., 1997, 18, 1463–1472 Search PubMed.
- A. Barducci, M. Bonomi and M. Parrinello, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2011, 1, 826–843 CAS.
- T. M. Schäfer and G. Settanni, J. Chem. Theory Comput., 2020, 16, 2042–2052 CrossRef PubMed.
- E. N. Baker and R. E. Hubbard, Prog. Biophys. Mol. Biol., 1984, 44, 97–179 CrossRef CAS PubMed.
- A. D. Becke, J. Chem. Phys., 1993, 98, 5648–5652 CrossRef CAS.
- P. J. Stephens, F. J. Devlin, C. F. Chabalowski and M. J. Frisch, J. Phys. Chem., 1994, 98, 11623–11627 CrossRef CAS.
- W. J. Hehre, R. Ditchfield and J. A. Pople, J. Chem. Phys., 1972, 56, 2257–2261 CrossRef CAS.
- E. Scrocco and J. Tomasi, New Concepts II, Berlin, Heidelberg, 1973, pp. 95–170 Search PubMed.
- J. D. Prajapati, J. N. Onuchic and K. Y. Sanbonmatsu, J. Mol. Biol., 2022, 434, 167788 CrossRef CAS PubMed.
- Y. Kaku, K. Okumura, M. Padilla-Blanco, Y. Kosugi, K. Uriu, A. A. Hinay, L. Chen, A. Plianchaisuk, K. Kobiyama, K. J. Ishii, J. Zahradnik, J. Ito and K. Sato, Lancet Infect. Dis., 2024, 24, e82 Search PubMed.
| This journal is © the Owner Societies 2026 |




