 Open Access Article
Open Access ArticleFluoroquinolones and their complexes with metal ions, studied with density functional theory
Ran
Friedman

Department of Chemistry and Biomedical Sciences, Linnaeus University, Kalmar, Sweden. E-mail: ran.friedman@lnu.se
First published on 9th March 2026
Abstract
Fluoroquinolone antibiotics might lead to severe side effects, collectively known as fluoroquinolone-associated disability (FQAD). The origin of this phenomenon is unknown, but has been suggested to involve chelation of biologically important ions such as Fe3+. In this study, DFT calculations were used to estimate the Gibbs energies of binding of biologically-relevant ions (Mg2+, Ca2+, Mn2+, Fe3+ and Zn2+) to ciprofloxacin, a prototype of fluoroquinolone antibiotics. The results show preferable binding of Fe3+ to ciprofloxacin, with binding affinity that is over 50 kcal mol−1. The binding of the ions to ciptofloxacin is compared to their binding to tetracycline, a metal binding antibiotic that is not a fluoroquinolone. The affinity of Fe3+ and most other ions to tetracycline was found to be even higher, which leads to the conclusion that ion binding is not the cause for FQAD. Overall, this study demonstrates the usability of a computational-chemistry based approach to a problem within biomedicine. Methodological and structural aspects of the binding are also discussed.
1. Introduction
Quinolone antibiotics have been used to treat bacterial infections since the 1960s and work by inhibition of bacterial topoisomerases.1 The incorporation of a flour atom into the pharmacophore has led to more active compounds (advanced generation quinolones) and the quinolones that are in use today are hence fluoroquinolones (FQ).2 These antibiotics have broad range and many are bioavailable. Hence, they are used to treat many different infections.3 Structural studies reveal binding to the target enzymes through multitude of interactions, including chelation of an Mg2+ ion.After decades of use, reports of serious and persistent adverse effects following use of FQ have gathered the attention of clinicians. Such severe side effects including weakness, reduced balanced and even psychiatric disorders were verified in an animal model and the term “FQ-associated disability”, FQAD, was coined to describe them.4 The use of FQ has since been reduced, but they are still necessary when other antibiotics are insufficient (e.g. due to resistance). Thus, there is an urgent need to understand the causes of FQAD and identify useful treatment.
While the molecular mechanism that underlies FQAD is not understood, metal-ion chelation by the drugs has been suggested to play an important, perhaps crucial factor.5 FQ can chelate many different multivalent ions.6,7 The chelation of Fe3+ appears to be fundamental in this respect. Physiologically relevant concentrations of FQ were enough to reduce the activity of several iron-dependent enzymes.8 In addition, some of the observed side effects can be related to chelation of other ions, such Mg2+ and Zn2+.5
Supplementation with essential metal cofactors is one suggested route to treat FQAD, but to support this further research should show that the chelation of ions is indeed important.5 For this reason, it is important to quantify the affinity of the ions to the drugs. While there are several experimental methods to measure the stability constants between FQ and metals, experimental setups vary which affect the results. A robust theoretical approach is therefore needed which will infer on the binding energies and atomistic interactions involved in ion chelation by FQ. This is the aim of this study. In many previous studies, DFT calculations were used to study ion chelation and for discrimination between ions (e.g. ref. 9–11).
Since most FQ have a common skeleton and two titratable groups, one basic and one acidic, it can be expected that differences with respect to ion chelations will be minimal. Here, ciprofloxacin is used as a prototype of FQ since it is a widely used antibiotic with a relatively simple structure. The complexion of ciprofloxacin with the biologically relevant metal ions Mg2+, Ca2+, Mn2+, Fe3+ and Zn2+ was studied employing a computational approach. The results shed light on the structural and energetic features associated with ion chelation by FQ. Finally, ion complexes formed with tetracycline are also studied since tetracyclines are known for their ability to chelate metal ions but do not lead to similar disabilities as FQ.
2. Theory
2.1. Calculations in implicit solvent
The binding energy ΔGb between an ion and n ligand molecules is calculated as:| ΔGb = G°(complex) − [nG°(ligand) + G°(ion)] | (1) |
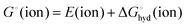 | (2) |
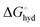 is the hydration energy of the ion, i.e. the energy that is required to transfer an ion from the gas phase (standard state at p = 1 bar) to the solvent (standard state at C = 1 M). Experimental energies, as given in ref. 14 are used here, to which a correction,
is the hydration energy of the ion, i.e. the energy that is required to transfer an ion from the gas phase (standard state at p = 1 bar) to the solvent (standard state at C = 1 M). Experimental energies, as given in ref. 14 are used here, to which a correction, 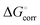 was added:
was added:
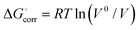 | (3) |
The value of 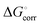 is 1.89 kcal mol−1 and its addition was needed since the values in ref. 14 refer to the solvent standard state (1 M concentration) rather than the standard state in the gas phase (1 bar pressure).
is 1.89 kcal mol−1 and its addition was needed since the values in ref. 14 refer to the solvent standard state (1 M concentration) rather than the standard state in the gas phase (1 bar pressure).
2.2. Calculations with explicit solvent
Implicit solvent models are highly useful in chemistry, but are not always able to capture all the contributions from the solvent correctly. This calls for the use of explicit solvent, i.e. modelling the water molecules explicitly. It is not possible to calculate the binding energy in the same way in explicit solvent, since this would require considering the room temperature dynamics of many water molecules. Realising that the solvent molecules are more important for correctly modelling the hydration of the metal ion (where the charge is concentrated on a single atom) not the ligand (where the charge is delocalised), a mixed model was used here. Consequently, six water molecules were studied in addition to the ion, and implicit solvent is still used. Using even more water molecules makes the problem at hand almost intractable, since there can be multiple minima on the potential energy surface and the additional water molecules do not necessarily making the approximation of the structures and energy more realistic. With this set-up, eqn (1) is replaced by:| ΔGb = G°(complex·(H2O)6) − [nG°(ligand) + G°(ion·(H2O)6)] | (4) |
Since the hydrated ion complexes are much larger in any case, the exact radius of the ion as used to build the cavity in implicit solvent is of lesser important, and the energies in the right-hand side of eqn (4) can be calculated directly from DFT.
Although two water molecules are enough to complete the hydration shell of the ions in their bound state, six were used in the calculations to allow for fully hydrated ions as reference.
2.3. Ion speciation
Fe and Mn exist in the body in both divalent and trivalent states, however, in both cases one form is dominant and was studied here. For Mn, this was the Mn2+ ion. For Fe, Fe3+ was used, because it is the form that exist in physiological pH.15 Fe3+ was modelled as is, i.e. Fe3+ or [Fe·(H2O)6]3+ rather than [FeOH]2+ or [Fe(OH)(H2O)4(H3O)]3+. This is because the hexa-hydrated species was found to be more stable (using the same level of theory as in the calculations of ΔGb). In an aqueous environment, the excess proton would quickly diffuse thereafter leaving [Fe(OH)(H2O)5]2+. However, modelling the speciation of the ion in full complexity would require a much larger number of solvent molecules and would not enable the use of accurate quantum mechanical (QM) methods.3. Computational methods
DFT calculations were performed with ORCA,16,17 version 6.1.0.18 Geometry optimisations were performed employing the ωB97-3c composite method19 in implicit solvent (modelled by SMD20). Convergence to minimum energy was verified by calculating the vibration frequencies and ensuring that none of them was negative. Unless otherwise stated, the energy was thereafter calculated with the ωB97M-V functional21 and the def2-TZVPP basis set.22I.e. the calculations employed the ωB97M-V(SMD)/def2-TZVPP//ωB97X-3C(SMD) setting. Zero point energy (ZPE) corrections to the energy were calculated at 298 K using the ωB97X-3C(SMD) method.Ciprofloxacin was modelled in its zwitterionic state which is dominant in physiological pH. Tetracycline was modelled in its negatively charged, base state (with two negative and one positive charge, yielding formal charge of −1). The molecule's basic group pKa = 9.6 and most acidic group (pKa = 3.3) are charged over a wide range of pH values. Its second acidic group has pKa of 7.8, meaning that some of the molecules will be charged at physiological pH. Because the neutral state is favoured, the binding affinity for tetracycline was shifted by 1.06 kcal mol−1 which corresponds to the distribution of neutral and charged species:
ΔGpHcorr = RT![[thin space (1/6-em)]](https://www.rsc.org/images/entities/i_char_2009.gif) ln(10pKa − pH). ln(10pKa − pH). |
Transition metal ions (Mn2+ and Fe3+) where modelled in their high spin state that was found to be more stable in complexes with the water and drugs using the same level of theory as reported above, except for Mn2+ with tetracycline where the energy of the complex was slightly lower when Mn2+ adopted a lower spin (quartet) state.
4. Results
4.1. Models with implicit solvent
Ciprofloxacin and its coordination with the various metal ions were first studied with a fully implicit solvent model. While such models are simpler and might be less accurate, they alleviate the need to consider the explicit placement of water molecules that can quantitatively modify the results. The coordination of the ion is considered first, followed by binding energies.![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) metal 1
metal 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) 1 complexes.
Three atoms in FQ can coordinate with the metal ions, namely two carboxylate and one keto oxygen. Given the zwitterionic form of these molecules in physiological conditions, there will be a negative charge distributed between those oxygens and the ions are expect to bind two of them, as shown in Fig. 1A and B. All complexes are more stable when the oxidised ring oxygen participates in the coordination, as in Fig. 1A (Table S1).
1 complexes.
Three atoms in FQ can coordinate with the metal ions, namely two carboxylate and one keto oxygen. Given the zwitterionic form of these molecules in physiological conditions, there will be a negative charge distributed between those oxygens and the ions are expect to bind two of them, as shown in Fig. 1A and B. All complexes are more stable when the oxidised ring oxygen participates in the coordination, as in Fig. 1A (Table S1).
 | ||
| Fig. 1 Possible placements of a metal ion that interacts with a single ciprofloxacin molecule. Optimised structures are shown with Fe3+. | ||
Binding energies, ion–oxygen distances and oxygen–ion–oxygen angles for the complexes are given in Table 1. These calculations show favourable binding of Mg2+, Mn2+ and Fe3+ but only by few kcal mol−1. The interaction between Ca2+ and Zn2+ and the drug is repulsive. Mg2+, Ca2+ and Zn2+ are hard ions with filled electronic shells; the small size of Mg2+ however makes it fit well between the oxygens. Zn2+ can also be close to the oxygen atoms but the filled d-shell makes repulsive interactions with the oxygens, making the O–ion–O angle larger and opposes binding. The transitions metals Mn2+ and Fe3+ bind the drug but not strongly as mentioned. Binding energy calculations in implicit solvent are Gibbs (free) energies, since the solvent contributions are approximated as solvation free energies. However, ZPE corrections are normally considered to be useful and were included here. It is worth mentioning that if ZPE are not considered, binding energies are almost the same except that the binding energies for Mn2+ and Fe3+ become slightly positive (Table S2).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ciprofloxacin
1 ciprofloxacin![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) metal complexes. d(ion–OC) is the distance to the coordinating carboxylate oxygen and d(ion–OK) is the distance to the keto oxygen. In this and the following tables energies are in kcal mol−1, distances are in Å and angles are in degrees
metal complexes. d(ion–OC) is the distance to the coordinating carboxylate oxygen and d(ion–OK) is the distance to the keto oxygen. In this and the following tables energies are in kcal mol−1, distances are in Å and angles are in degrees
| Ion | ΔGb | d(ion–OC) | d(ion–OK) | ∠O–ion–O |
|---|---|---|---|---|
| Mg2+ | −5.6 | 1.95 | 1.93 | 93.6 |
| Ca2+ | 6.5 | 2.22 | 2.19 | 78.7 |
| Mn2+ | −1.5 | 1.97 | 1.95 | 93.4 |
| Fe3+ | −0.7 | 1.91 | 1.88 | 95.9 |
| Zn2+ | 17.1 | 1.86 | 1.84 | 108.0 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) metal 2
metal 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) 1 complexes.
Models of ciprofloxacin complexation with metals suggest that the metal binds with multiple drug molecules (e.g. ref. 23). Indeed, using two molecules of the drug instead of one resulted in favourable binding for all ions (Table 2). The binding (Fig. 2) was most favourable for Fe3+, with ΔGb = −73.4 kcal mol−1 and least for Ca2+. The formation of a larger complex decreases the degrees of freedom within each ciprofloxacin molecule, which led to a significant contribution of ZPE opposing binding (Table S3, average ZPE contribution 16.6 kcal mol−1). The structures of the complexes had C2 symmetry. The ions bound in a distorted tetrahedral coordination, leaving room for two extra waters. Structural details are given in Table 2. Ion–oxygen distances are ∼2.0 Å, except in complex with Ca2+ where they are much larger and with Fe3+ where they are <1.9 Å. O–ion–O angles are similar except with Ca2+. Most notably, the OC–ion–OC angle is close to 180° with Ca2+ compared to ∼120° with the other ions. This is because Ca2+ binds more weakly to the drugs, and as a consequence the charged carboxylate oxygens that bind it avoid one another. The OC–OC and OK–OK distances are similar to each other in every complex, except for the complex with Ca2+ where the OC–OC distance is larger.
1 complexes.
Models of ciprofloxacin complexation with metals suggest that the metal binds with multiple drug molecules (e.g. ref. 23). Indeed, using two molecules of the drug instead of one resulted in favourable binding for all ions (Table 2). The binding (Fig. 2) was most favourable for Fe3+, with ΔGb = −73.4 kcal mol−1 and least for Ca2+. The formation of a larger complex decreases the degrees of freedom within each ciprofloxacin molecule, which led to a significant contribution of ZPE opposing binding (Table S3, average ZPE contribution 16.6 kcal mol−1). The structures of the complexes had C2 symmetry. The ions bound in a distorted tetrahedral coordination, leaving room for two extra waters. Structural details are given in Table 2. Ion–oxygen distances are ∼2.0 Å, except in complex with Ca2+ where they are much larger and with Fe3+ where they are <1.9 Å. O–ion–O angles are similar except with Ca2+. Most notably, the OC–ion–OC angle is close to 180° with Ca2+ compared to ∼120° with the other ions. This is because Ca2+ binds more weakly to the drugs, and as a consequence the charged carboxylate oxygens that bind it avoid one another. The OC–OC and OK–OK distances are similar to each other in every complex, except for the complex with Ca2+ where the OC–OC distance is larger.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ciprofloxacin
1 ciprofloxacin![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) metal complexes. OK = keto oxygen, OC = carboxylate oxygen. The notations 'same' and 'other' refer to the oxygens being on the same molecule or not
metal complexes. OK = keto oxygen, OC = carboxylate oxygen. The notations 'same' and 'other' refer to the oxygens being on the same molecule or not
| Ion | ΔGb | Angles | |||||||
|---|---|---|---|---|---|---|---|---|---|
| d(ion–OK) | d(ion–OC) | OK–ion–OC, same | OK–ion–OC, other | OK–ion–OK | OC–ion–OC | d(OK–OK) | d(OC–OC) | ||
| Mg2+ | −42.4 | 1.98 | 1.96 | 90.9 | 118.1 | 122.1 | 119.7 | 3.46 | 3.38 |
| Ca2+ | −24.9 | 2.28 | 2.26 | 76.7 | 106.8 | 115.3 | 173.8 | 3.85 | 4.53 |
| Mn2+ | −45.2 | 2.05 | 2.02 | 88.6 | 118.7 | 122.0 | 124.0 | 3.58 | 3.57 |
| Fe3+ | −73.4 | 1.88 | 1.87 | 95.1 | 115.5 | 119.5 | 117.8 | 3.25 | 3.19 |
| Zn2+ | −45.0 | 1.95 | 1.93 | 95.6 | 115.7 | 115.5 | 120.2 | 3.30 | 3.35 |
4.2. Models with implicit and explicit solvent
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) metal
metal![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) water 1
water 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) 6 complexes.
The ions that were studied here all adopt six-coordinated octahedral structures in water. When optimised with water molecules in the vicinity of the ion, complexes were formed were the ion was ligated by two oxygens of ciprofloxacin and 4–5 water molecules, with the additional waters forming a second shell, Fig. 3. The complexes with Mg2+ and Zn2+ had coordination number (CN) of 6 and an octahedral arrangement. Ca2+ and Fe3+ complexes, with CN = 7 adopted pentagonal bi-pyramidal geometries, that were closer to an ideal arrangement with Ca2+. Finally, the complex with Mn2+ was in a capped octahedral arrangement.
6 complexes.
The ions that were studied here all adopt six-coordinated octahedral structures in water. When optimised with water molecules in the vicinity of the ion, complexes were formed were the ion was ligated by two oxygens of ciprofloxacin and 4–5 water molecules, with the additional waters forming a second shell, Fig. 3. The complexes with Mg2+ and Zn2+ had coordination number (CN) of 6 and an octahedral arrangement. Ca2+ and Fe3+ complexes, with CN = 7 adopted pentagonal bi-pyramidal geometries, that were closer to an ideal arrangement with Ca2+. Finally, the complex with Mn2+ was in a capped octahedral arrangement.
 | ||
| Fig. 3 Optimised structures of complexes between ciprofloxacin and the various ions, optimised with six water molecules at the ion binding site. | ||
Examination of the binding energies (Table 3) reveals favourable binding for all ions, with a strong preference for Fe3+, ΔGb = −20.9. Mn2+ bound with the least favourable Gibbs energy. The distances to the nearest oxygens of ciprofloxacin were ∼2 Å for Mg2+, Fe3+ and Zn2+ and larger for Mn2+ and Ca2+. Distances were larger in comparison to the implicit-ion structures (Table 1).
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6 ciprofloxacin
6 ciprofloxacin![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) metal
metal![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water complexes. d(ion–OC) is the distance to the coordinating carboxylate oxygen and d(ion–OK) is the distance to the keto oxygen. In this and the following tables energies are in kcal mol−1, distances are in Å and angles are in degrees. Oh = octagedral, B-Py = bipyramidal, C-Oh = capped octahedral
water complexes. d(ion–OC) is the distance to the coordinating carboxylate oxygen and d(ion–OK) is the distance to the keto oxygen. In this and the following tables energies are in kcal mol−1, distances are in Å and angles are in degrees. Oh = octagedral, B-Py = bipyramidal, C-Oh = capped octahedral
| Ion | ΔGb | d(ion–OK) | d(ion–OC) | ∠O–ion–O | CN | Geometry |
|---|---|---|---|---|---|---|
| Mg2+ | −9.7 | 2.06 | 2.01 | 86.8 | 6 | Oh |
| Ca2+ | −9.3 | 2.40 | 2.34 | 72.2 | 7 | B-Py |
| Mn2+ | −6.4 | 2.21 | 2.10 | 81.7 | 7 | C-Oh |
| Fe3+ | −20.9 | 2.02 | 1.91 | 87.8 | 7 | B-Py |
| Zn2+ | −13.6 | 2.04 | 2.00 | 89.8 | 6 | Oh |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) metal
metal![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) water 2
water 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/b_char_2009.gif) 6 complexes.
With two ciprofloxacin molecules, all ions bound in an octahedral conformation (Fig. 4). As in all previous calculations, binding was most favourable with Fe3+. Surprisingly, Zn2+ showed repulsive interactions with the two drug molecules and the water, indicating that the ion will only bind one residue. The deviation in distances between pairs of similar oxygen atoms and Zn2+ was the largest among all ions and the octahedral coordination with Zn2+ was much distorted. The increased binding affinity upon incorporation of the second ciprofloxacin molecule was even larger than with the first one for Mn2+ and Fe3+ but not for Mg2+ and Ca2+.
6 complexes.
With two ciprofloxacin molecules, all ions bound in an octahedral conformation (Fig. 4). As in all previous calculations, binding was most favourable with Fe3+. Surprisingly, Zn2+ showed repulsive interactions with the two drug molecules and the water, indicating that the ion will only bind one residue. The deviation in distances between pairs of similar oxygen atoms and Zn2+ was the largest among all ions and the octahedral coordination with Zn2+ was much distorted. The increased binding affinity upon incorporation of the second ciprofloxacin molecule was even larger than with the first one for Mn2+ and Fe3+ but not for Mg2+ and Ca2+.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6 ciprofloxacin
6 ciprofloxacin![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ion
ion![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water complexes (Fig. S1). However, calculations of the binding energies for these complexes showed highly unfavourable interactions.
water complexes (Fig. S1). However, calculations of the binding energies for these complexes showed highly unfavourable interactions.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6 drug
6 drug![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) ion
ion![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water complexes were studied. These complexes were octahedral though slightly deformed (Fig. 5) with all ions, and involved four oxygens from the two drug molecules and two water molecules. Binding energies (Table 5) were favourable for all ions except Mn2+. For all other ions, ΔGb indicated a higher affinity for tetracycline.
water complexes were studied. These complexes were octahedral though slightly deformed (Fig. 5) with all ions, and involved four oxygens from the two drug molecules and two water molecules. Binding energies (Table 5) were favourable for all ions except Mn2+. For all other ions, ΔGb indicated a higher affinity for tetracycline.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6 ciprofloxacin
6 ciprofloxacin![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) metal
metal![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water complexes. d(ion–OC) are the distances to the coordinating carboxylate oxygen, d(ion–OK) to the keto oxygens and d(ion–OW) to the water oxygens
water complexes. d(ion–OC) are the distances to the coordinating carboxylate oxygen, d(ion–OK) to the keto oxygens and d(ion–OW) to the water oxygens
| Ion | ΔGb | d(ion–OK) | d(ion–OC) | d(ion–OW) |
|---|---|---|---|---|
| Mg2+ | −14.6 | 2.00, 2.00 | 1.93, 1.95 | 2.09, 2.09 |
| Ca2+ | −14.1 | 2.36, 2.38 | 2.31, 2.30 | 2.42, 2.42 |
| Mn2+ | −22.3 | 2.17, 2.19 | 2.12, 2.13 | 2.24, 2.25 |
| Fe3+ | −54.7 | 1.98, 1.98 | 1.94, 1.95 | 2.09, 2.12 |
| Zn2+ | 9.3 | 1.96, 2.10 | 2.03, 2.08 | 2.29, 2.27 |
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 6 tetracycline
6 tetracycline![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) metal
metal![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) water complexes
water complexes
| Ion | ΔGb |
|---|---|
| Mg2+ | −24.3 |
| Ca2+ | −24.8 |
| Mn2+ | 32.3 |
| Fe3+ | −60.2 |
| Zn2+ | −28.4 |
5. Discussion
5.1. Explicit versus implicit solvent in calculations of the metal binding affinities
Calculations of absolute (rather than relative) binding energies are known to be challenging and consideration of the modelled reaction is of high importance. Here, these calculations were carried out in implicit solvent, while considering the ion's solvation shell in explicit or implicit terms. As will be outlined below, the calculations suggest that the use of combined explicit/implicit solvation leads to better results.In terms of the structure, calculations with a single drug always show that the ion and its O ligands from the drug are on the same plane with the aromatic rings (Fig. 1 and 3). However, by considering the solvation shell, differences in the coordination between the ions become obvious (Fig. 1 and Table 3). It is clear that different ions bind the single drug with different geometries.
In terms of energies, single ion calculations in implicit solvent suggest that ciprofloxacin does not bind Zn2+ and Ca2+ and barely binds Mn2+ and Fe3+, which stands against multiple lines of experimental evidence.23,28 In contrast, considering the water shell leads to the conclusion that all ions favourably bind to ciprofloxacin at a 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 stoichiometry. At 2
1 stoichiometry. At 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 stoichiometry, implicit water calculations lead to the conclusions that all of the ions bind to ciprofloxacin (Table 2), whereas the explicit/implicit water reveal that Zn2+ would not bind in such case.
1 stoichiometry, implicit water calculations lead to the conclusions that all of the ions bind to ciprofloxacin (Table 2), whereas the explicit/implicit water reveal that Zn2+ would not bind in such case.
Differences in the binding energies in binding to ions (Table 1 compared to Table 3, Table 2 compared to Table 4) are substantial and stem from the contribution of the water molecules to the binding of the ions and the solvation of the drug's oxygens. These are apparently not well captured by the implicit solvent representation, although it works well in many other cases.
5.2. Limitations of the model
Neither experimental nor theoretical studies where binding affinities are estimated are devoid of limitations. Here, the aim was to approximate the binding of drugs to ions that take place under physiological condition in solution while relying on static calculations in implicit solvent. The limitations of the approach are clear. The alternative would be to perform explicit water, out-of-equilibrium simulations such as free energy perturbation or umbrella sampling. Such simulations take into account the multiple degrees of freedom of the system and multiple configurations. Free energy perturbation (FEP) calculations necessitate the use of a molecular mechanics (MM) forcefield to represent the element that is perturbed (here, the ion), because they involve a transformation between the bound and unbound states. Unfortunately, even if forcefield parameters for the ions included in this study could be obtained, considering change-of-coordination is not modelled correctly with such approach.29,30 While machine-learning interatomic potentials can be used for FEP31 this would require the development of potentials that are good enough to model the interactions and geometries as presented which in itself is far from trivial and was hitherto not achieved for such systems. Umbrella sampling calculations can in principle be carried out using quantum mechanical potentials, but the computational cost for systems as modelled here is prohibitive unless very approximate potentials (and small solvation boxes) are used, which would present other limitations.5.3. Binding to tetracyclines
Similar to ciprofloxacin, the calculations show that all but one ion (Mn2+) form complexes with favourable interaction energies with tetracycline, as a structural representative of tetracycline antibiotics. Experimental studies suggest that tetracycline binds Mn2+ under specific experimental conditions (e.g. at the present of oxygen32). It is likely that Mn2+ binds preferably to one molecule of tetracycline rather than two, but since this was not the focus of this study further calculations were not carried out.5.4. Ion chelation does not seem to be a major mechanism for FQAD
Although the results show that FQ bind very strongly to Fe3+, this study does not support the mechanism by which the chelation of Fe3+ leads to FQAD. Tetracycline appear to bind Fe3+ with higher affinity than ciprofloxacin, yet this does not lead to any known disabilities. It might be hypothesised that FQ, either complexed by metals or not, bind off-target in some patients for reasons that are currently unknown.5.5. Ion supplementation might not be a good therapy for FQAD
There is little support for using ion supplements (in the form of Mg- or Ca-salts) as therapy for FQAD following the results presented here. Fe3+ and Mn2+ should not be used for chelation of FQ due to risk for toxicities. Ca2+ and Mg2+ have more modest binding affinities, of less than 15 kcal mol−1. Consequently, none of the ions is expected to be an effective chelator of residual FQ in physiological conditions.6. Conclusions
Accurate calculations of the binding energies of biologically-relevant, multivalent metal ions were performed in order to examine the postulation that FQAD is the result of iron chelation. Consideration of the ions' solvation shell led to more convincing results. While the calculations support strong binding between ciprofloxacin and Fe3+, with ΔGb = −54.7 kcal mol−1, an even stronger binding was shown for tetracycline, which does not lead to similar disabilities in patients. Mg2+ and Ca2+ show more moderate affinity towards ciprofloxacin which limit their usability as drug chelators in vivo.Conflicts of interest
There are no conflicts to declare.Data availability
Supplementary information (SI): Supplementary materials (Tables S1–S3 and Fig. S1) are available in the SI file. Optimised molecular structures are freely available at https://www.doi.org/10.6084/m9.figshare.30515897. See DOI: https://doi.org/10.1039/d5cp04229a.Acknowledgements
The computations were enabled by resources provided by the National Academic Infrastructure for Supercomputing in Sweden (NAISS), partially funded by the Swedish Research Council through grant agreement no. 2022-06725.Notes and references
- T. D. M. Pham, Z. M. Ziora and M. A. T. Blaskovich, MedChemComm, 2019, 10, 1719–1739 Search PubMed.
- P. C. Sharma, A. Jain and S. Jain, Acta Pol. Pharm., 2009, 66, 587–604 CAS.
- B. J. Werth, Fluoroquinolones, 2024, https://www.merckmanuals.com/home/infections/antibiotics/fluoroquinolones Search PubMed.
- K. Kaur, R. Fayad, A. Saxena, N. Frizzell, A. Chanda, S. Das, S. Chatterjee, S. Hegde, M. Baliga, V. Ponemone, M. Rorro, J. Greene, Y. Elraheb, A. Redd, J. Bian, J. Restaino, L. Norris, Z. Qureshi, B. Love, B. Bookstaver, P. Georgantopoulos, O. Sartor, D. Raisch, G. Rao, K. Lu, P. Ray, W. Hrusheshky, R. Schulz, R. Ablin, V. Noxon and C. Bennett, J. Community Supportive Oncol., 2016, 54–65 CrossRef CAS PubMed.
- K. Michalak, A. Sobolewska-Włodarczyk, M. Włdarczyk, J. Sobolewska, P. Wźniak and B. Sobolewski, Oxid. Med. Cell Longev., 2017, 2017, 8023935 CrossRef PubMed.
- A. Serafin and A. Stańczak, Russ. J. Coord. Chem., 2009, 35, 81–95 CrossRef CAS.
- N. Seedher and P. Agarwal, Drug Metab. Drug Interact., 2010, 25, 17–24 CrossRef CAS PubMed.
- S. Badal, Y. F. Her and L. J. Maher, J. Biol. Chem., 2015, 290, 22287–22297 CrossRef CAS PubMed.
- F. Y. Adeowo, B. Honarparvar and A. A. Skelton, RSC Adv., 2016, 6, 79485–79496 RSC.
- S. Kaviani, M. Izadyar and M. R. Housaindokht, J. Mol. Graphics Modell., 2018, 80, 182–189 CrossRef CAS.
- R. Friedman, Phys. Chem. Chem. Phys., 2025, 27, 13705–13713 Search PubMed.
- R. Friedman, J. Phys. Chem. B, 2021, 125, 2251–2257 Search PubMed.
- R. Friedman, Chem. Phys. Chem., 2022, 24, e202200516 Search PubMed.
- Y. Marcus, J. Chem. Soc., Faraday Trans., 1991, 87, 2995–2999 Search PubMed.
- T. Ems, K. St Lucia and M. R. Huecker, StatPearls, StatPearls Publishing, Treasure Island (FL), 2025 Search PubMed.
- F. Neese, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2011, 2, 73–78 Search PubMed.
- F. Neese, F. Wennmohs, U. Becker and C. Riplinger, J. Chem. Phys., 2020, 152, 224108 Search PubMed.
- F. Neese, Wiley Interdiscip. Rev.: Comput. Mol. Sci., 2025, 15, e70019 Search PubMed.
- M. Müller, A. Hansen and S. Grimme, J. Chem. Phys., 2023, 158, 014103 CrossRef PubMed.
- A. V. Marenich, C. J. Cramer and D. G. Truhlar, J. Phys. Chem. B, 2009, 113, 6378–6396 Search PubMed.
- N. Mardirossian and M. Head-Gordon, J. Chem. Phys., 2016, 144, 214110 CrossRef PubMed.
- F. Weigend and R. Ahlrichs, Phys. Chem. Chem. Phys., 2005, 7, 3297–3305 RSC.
- G. Psomas, J. Inorg. Biochem., 2008, 102, 1798–1811 CrossRef CAS PubMed.
- E. K. Efthimiadou, Y. Sanakis, N. Katsaros, A. Karaliota and G. Psomas, Polyhedron, 2007, 26, 1148–1158 CrossRef CAS.
- T. Ohyama and J. A. Cowan, Inorg. Chem., 1995, 34, 3083–3086 CrossRef CAS.
- L. Jin, X. Amaya-Mazo, M. E. Apel, S. S. Sankisa, E. Johnson, M. A. Zbyszynska and A. Han, Biophys. Chem., 2007, 128, 185–196 CrossRef CAS PubMed.
- H. Wang, H. Yao, P. Sun, D. Li and C.-H. Huang, Environ. Sci. Technol., 2015, 50, 145–153 CrossRef.
- E. A. Hartshorn, B. M. Lomaestro and G. R. Bailie, DICP, 1991, 25, 1249–1258 CrossRef PubMed.
- E. Ahlstrand, D. Spångberg, K. Hermansson and R. Friedman, Int. J. Quantum Chem., 2013, 113, 2554–2562 CrossRef CAS.
- E. Ahlstrand, K. Hermansson and R. Friedman, J. Phys. Chem. A, 2017, 121, 2643–2654 CrossRef CAS PubMed.
- D. J. M. van Pinxteren and W. Jespers, J. Chem. Inf. Model., 2025, 65, 9856–9864 CrossRef CAS PubMed.
- W.-R. Chen and C.-H. Huang, Environ. Sci. Technol., 2008, 43, 401–407 CrossRef PubMed.
| This journal is © the Owner Societies 2026 |



