Open Access Article
Open Access ArticleBackbone deprotonation defines a hidden decomposition pathway of diphosphine ligands
Logan J.
Taylor
,
A. Dina
Dilinaer
,
Paul D.
Boyle
and
Marcus W.
Drover
 *
*
Department of Chemistry, Western University, 1151 Richmond Street, London, ON N8K 3G6, Canada. E-mail: marcus.drover@uwo.ca
First published on 12th March 2026
Abstract
Diphosphine ligands are ubiquitous partners in synthetic chemistry, enabling tuneable reactivity and selectivity. Here, we identify an overlooked decomposition pathway: base-induced deprotonation of an ethylene diphosphine carbon backbone to give a strained metallabicycle. The instability of this scaffold, reflected in a short half-life and decomposition even during routine filtration, has implications for diphosphine–metal coordination chemistry and catalysis under basic conditions.
Ligand design is a cornerstone of molecular coordination chemistry, providing a means to stabilize s, p, d, and even f-block metal complexes.1,2 Amongst possible candidates, diphosphine ligands, comprising two phosphine donors linked intramolecularly, have been extensively studied. Both phosphine substituents and ligand backbone are tunable, allowing for variability in length, rigidity, and shape (i.e., aliphatic, aromatic, etc.) (Scheme 1a).3,4 Beyond their primary coordination role, diphosphines can serve as templates for the introduction of secondary coordination sphere (SCS) elements5 or point, axial, or other forms of chirality.6–8
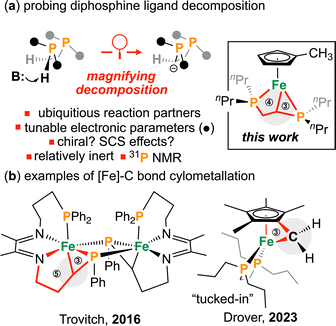 | ||
| Scheme 1 (a) Probing diphosphine decomposition space, SCS = secondary coordination sphere; (b) examples of cycometallated Fe–C complexes. Circles with numbers refer to ring size. | ||
Although often considered spectator ligands, diphosphines can undergo unproductive reactivity, including nucleophilic attack on metal-bound substrates9 or activation of its ligand C–H bonds.10 Larger diphosphine chelates can form cyclic structures upon heating or on exposure to external reagents.11–17 For example, Fogg and co-workers showed that a butylene-linked diphosphine Ru complex reacted with excess silylene ligand to give a [3.5] ruthenacycle (where [n.m] denotes fused n- and m- membered rings).12 Trovitch and co-workers reported that heating a tetradentate Fe-[P,N,N,P] complex led to benzene elimination and formation of a [5.3] cycle (Scheme 1b, left),13 while Lacy and colleagues observed that deprotonation of a Ru-[P,N] complex gave rise to a unique κ3-N,C,P bound Ru complex having a [3.4] cycle, which was reactive toward H2.14 Intricate knowledge of these reactive pathways underpins application of such ligands in catalysis.
As part of our ongoing research program, we have explored diphosphine ligands as anchoring points for the introduction of pendent Lewis acid functionality, reporting scaffolds bearing one,18,19 two,20 and even four21,22 Lewis acid group(s). We have also studied diphosphine ligands in a purely supporting role, reporting the first example of a room-temperature stable iron “tucked-in” complex, formed by treatment of [Cp*Fe(diphosphine)(N2)]+ with base (Scheme 1b, right).23 In this reaction, deprotonation of a peripheral Cp(C![[H with combining low line]](https://www.rsc.org/images/entities/char_0048_0332.gif) 3) group generates a direct {Fe–CH2} bonding interaction.23 This reactive Cp*Fe ‘tucked-in’ complex exhibits high nucleophilicity, undergoing hydroboration, for example, to form a pendent {Cp*CH2BCy2} fragment – an underexploited site for secondary coordination sphere manipulation – which we leveraged for CO2 dihydroboration.24
3) group generates a direct {Fe–CH2} bonding interaction.23 This reactive Cp*Fe ‘tucked-in’ complex exhibits high nucleophilicity, undergoing hydroboration, for example, to form a pendent {Cp*CH2BCy2} fragment – an underexploited site for secondary coordination sphere manipulation – which we leveraged for CO2 dihydroboration.24
To further understand the factors governing tucked-in formation in [{CpR}Fe(diphosphine)]+ (R = functionalized Cp) complexes, we were interested in exploring the chemistry of a related analogue, CpMe, containing a single methylated site. Reactivity of a {CpMe}Fe complex with base, however, did not produce the expected tucked-in iron complex. Instead, in the case of [{CpMe}Fe(diphosphine)(Cl)], deprotonation occurred at the diphosphine backbone, rather than the CpMe group, producing a strained [3.4] ferracycle (Scheme 1a, right). This reactive complex represents, to our knowledge, the first characterized coordination compound featuring a strained κ3-P,C,P ethylene-bridged diphosphine motif. These findings highlight a possible unrecognized decomposition pathway and raise questions about the compatibility of widely used diphosphine ligands with nucleophilic bases.
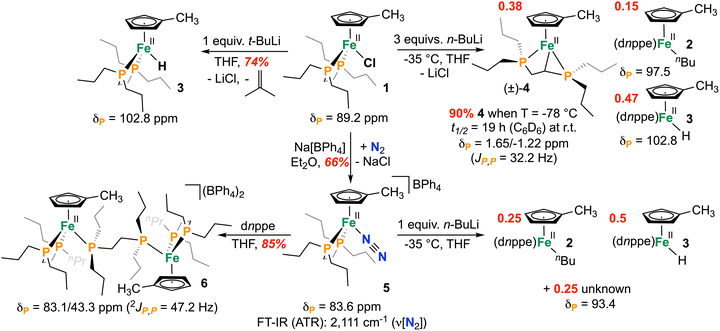
To begin, a THF slurry of sodium methylcyclopentadienide (NaCpMe)25 was reacted with [FeII(dnppe)2(Cl)2] (dnppe = 1,2-bis(di-n-propylphosphino)ethane) to give [{CpMe}Fe(dnppe)(Cl)] (1), with concomitant release of one equivalent of free dnppe ligand. The 31P{1H} NMR spectrum showed a single resonance at δP = 89.2 ppm, approximately 10 ppm downfield of the [Cp*Fe(dnppe)(Cl)] analogue (Scheme 2). 1H NMR spectroscopy revealed Cs symmetry, with two singlets at δH = 4.59 and 2.66 ppm; 2D 1H–13C HMBC and 1H–1H COSY NMR spectroscopic experiments confirmed that these signals correspond to the Cp–H protons. Dissolution of 1 in hexanes and storing the saturated solution at −35 °C overnight afforded purple-brown crystals suitable for X-ray diffraction (Fig. 1, left).
We next investigated 1 as a potential precursor to a tucked-in complex, which was expected to give a single 31P NMR spectral resonance. Treatment of 1 with 3 equiv. of n-BuLi at −35 °C and analysis of the crude reaction mixture by 31P{1H} NMR spectroscopy, however, indicated formation of at least three new diamagnetic compounds: signals at δP = 102.8 and 97.5 ppm along with a pair of ‘roofed’ [AB] doublets at δP = 1.65 and −1.22 ppm (JP,P = 32.2 Hz) in a 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1
1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2.5 ratio (Scheme 2). The signals at δP = 102.8 and 97.5 ppm were assigned as side products arising from n-BuLi attack at iron: [{CpMe}Fe(dnppe)(nBu)] (2) and its β-hydride elimination product, [{CpMe}Fe(dnppe)(H)] (3), respectively, indicating that the targeted tucked-in CpMeFe complex was not prepared. 1H–31P HMBC NMR spectroscopy confirmed the assignment of 2 with the signal at δP = 97.5 ppm correlating to a 1H NMR signal at δH = −0.24 ppm; 1D 1H TOCSY NMR linked this site to three others, consistent with an {Fe-n-Bu} fragment. The most downfield peak at δP = 102.8 ppm corresponds to the iron(II)-hydride, 3, which was independently prepared by reaction of 1 with t-BuLi. Additional support for the structure of 3 comes from a phosphorus-coupled triplet at δH = −17.3 ppm (2JP,H = 72.3 Hz) and a broad Fe—H stretch in the FT-IR ATR spectrum (νFe,H = 1830 cm−1). A molecular ion of the appropriate exact mass and isotope pattern was also observed for complexes 2 (m/z = 454.258) and 3 (m/z = 398.195) by liquid injection field desorption ionization mass spectrometry (LIFDI-HRMS).26
2.5 ratio (Scheme 2). The signals at δP = 102.8 and 97.5 ppm were assigned as side products arising from n-BuLi attack at iron: [{CpMe}Fe(dnppe)(nBu)] (2) and its β-hydride elimination product, [{CpMe}Fe(dnppe)(H)] (3), respectively, indicating that the targeted tucked-in CpMeFe complex was not prepared. 1H–31P HMBC NMR spectroscopy confirmed the assignment of 2 with the signal at δP = 97.5 ppm correlating to a 1H NMR signal at δH = −0.24 ppm; 1D 1H TOCSY NMR linked this site to three others, consistent with an {Fe-n-Bu} fragment. The most downfield peak at δP = 102.8 ppm corresponds to the iron(II)-hydride, 3, which was independently prepared by reaction of 1 with t-BuLi. Additional support for the structure of 3 comes from a phosphorus-coupled triplet at δH = −17.3 ppm (2JP,H = 72.3 Hz) and a broad Fe—H stretch in the FT-IR ATR spectrum (νFe,H = 1830 cm−1). A molecular ion of the appropriate exact mass and isotope pattern was also observed for complexes 2 (m/z = 454.258) and 3 (m/z = 398.195) by liquid injection field desorption ionization mass spectrometry (LIFDI-HRMS).26
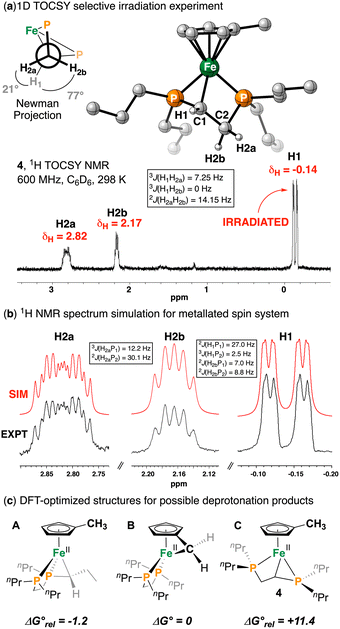
We next sought to identify the species responsible for the 31P NMR signals at δP = 1.65 and −1.22 ppm, which we hypothesized arose from complex 4, formed via direct C–H deprotonation rather than Fe–Cl attack (Scheme 2). Reaction of [{CpMe}Fe(dnppe)(Cl)] (1) with n-BuLi at lower temperature (−78 °C) afforded a deep red solution, predominantly containing the new product 4 in 90% by quantitative 31P NMR spectroscopy. 1H NMR spectroscopy of 4 revealed four Cp—H resonances, consistent with a C1-symmetric molecule, along with a distinctive upfield signal at δH = −0.14 ppm, which was responsive to 31P decoupling. A 1D 1H TOCSY experiment showed coupling to partners at δH = 2.17 and 2.82 ppm, whose spin system was appropriately modelled (Fig. 2a and b). 1H–31P HMBC correlations identified these protons as having a long-range correlation with the diphosphine phosphorus signals at δP = 1.65 and −1.22 ppm. 1H–13C HSQC NMR spectroscopy further indicated that the resonances at δH = 2.17 and 2.82 ppm were geminal (on the same carbon), while the third proton was strongly upfield shifted (δC = −25.1 ppm). This is contrasted with [Cp*Fe(dnppe)(CH3)], for example, which has a 13C NMR signal at δC = −11.3 (2JP,C = 24.1 Hz).21 Taken together, these data are consistent with deprotonation of the diphosphine ligand backbone to form a strained [4.3] ferracycle, [{CpMe}Fe(κ3-P,C,P-dnppe)] (4). Complex 4 was also examined using LIFDI-HRMS, providing a molecular ion at m/z = 396.179, accompanied by an overlapping [M + H]+ pattern at m/z = 397.187 in a 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 ratio.27 To our knowledge, this represents the first observed example of backbone deprotonation at a coordinated ethylene-bridged diphosphine ligand. This outcome also provides an interesting contrast between n-BuLi and t-BuLi reactivity: although attack at iron is observed in both cases, n-BuLi generates Fe–nBu (2), Fe–H (3), and 4, whereas t-BuLi produces only the iron hydride 2. The behaviour of the latter is consistent with t-BuLi being too sterically demanding to access the backbone carbon, precluding deprotonation and thus formation of 4.
3 ratio.27 To our knowledge, this represents the first observed example of backbone deprotonation at a coordinated ethylene-bridged diphosphine ligand. This outcome also provides an interesting contrast between n-BuLi and t-BuLi reactivity: although attack at iron is observed in both cases, n-BuLi generates Fe–nBu (2), Fe–H (3), and 4, whereas t-BuLi produces only the iron hydride 2. The behaviour of the latter is consistent with t-BuLi being too sterically demanding to access the backbone carbon, precluding deprotonation and thus formation of 4.
Attempts to obtain crystalline 4 suitable for single crystal X-ray diffraction were not met with success. Consistent with its strained [{CpMe}Fe(κ3-P,C,P-dnppe)] core, complex 4 decomposes rapidly upon contact with common filter media such as Celite® 545, as well as in solution or in the solid-state at room temperature or even at −35 °C. Using an internal standard and 31P{1H} NMR spectroscopy in C6D6, gradual decay was observed, yielding a half-life of t1/2 ≈ 19 h at 25 °C.
A computational model of 4 revealed a strained three-legged piano stool geometry with κ3-P,C,P diphosphine ligand coordination (Fig. 2a, inset). The Fe–C bond is shortened to 2.070 Å compared to 2.131(1) Å in the related [Cp*Fe(dnppe)(CH3)]21 complex. Similarly, the Fe–P bonds contract to 2.156 and 2.137 Å, relative to 2.162(4) and 2.180(4) Å in [Cp*Fe(dnppe)(CH3)],21 consistent with the effects of κ3-coordination. The reaction of 1 with n-BuLi can, in principle, yield deprotonation at three sites: (a) an n-Pr arm; (b) the diphosphine backbone; or (c) the CpMe ligand, with 8H:4H:3H probability. Compared to tucked-in complex B (Fig. 2c), complex 4 (C) is calculated to be 11.4 kcal mol−1 higher in energy, whereas deprotonation at the n-Pr(CH2) arm (A) is slightly more stable by 1.2 kcal mol−1.23 This indicates that 4 forms as a kinetic product, which is consistent with its clean formation at low temperature and limited stability. NMR chemical shifts were also obtained using DFT calculations. As described previously, cyclometallation of phosphine ligands typically generates small chelate rings, leading to 31P resonances 50–100 ppm upfield of the precursor.28 This is observed here: δP = 89.2 ppm for 1vs. δP = 1.65 and −1.22 ppm for 4 (ΔδP ∼ 90 ppm). Using optimized structures from DFT (see SI), calculated 31P chemical shifts reproduce this trend, giving ΔδP ≈ 99 ppm, once again supporting the assignment of 4 as a diphosphine-backbone-deprotonated species.
To determine whether 4 can arise from precursors other than [{CpMe}Fe(dnppe)(Cl)] (1), we targeted the alternative halide-free complex [{CpMe}Fe(dnppe)(N2)]BPh4 (5), which for the related Cp* complex, was required for clean tucked-in complex preparation.23 Initial attempts combining [FeII(dnppe)2(Cl)2], NaCpMe, and NaBPh4, however, were unsuccessful, as released dnppe was trapped to provide the μ-bridged complex 6. Complex 6 was characterized by single crystal X-ray diffraction (Fig. 1, right), HRESI-MS, and 31P{1H} NMR spectroscopy, which showed a doublet and quartet at δP = 83.1 and 43.3 ppm (2JP,P = 47.2 Hz) (Scheme 2). This outcome suggests reactivity from the desired complex [{CpMe}Fe(dnppe)(N2)]+. Of note, μ-dnppe coordination is not observed for the Cp* analogue, likely due to the more electron-poor iron centre and/or reduced steric bulk of the CpMe system in 5. By carefully omitting exogenous phosphine and reacting 1 with NaBPh4, the desired [{CpMe}Fe(dnppe)(N2)]+ (5) was obtained, exhibiting a single 31P{1H} NMR resonance at δP = 83.6 ppm and an FT-IR N2 stretch of 2111 cm−1. By HRESI-MS, a strong [M–N2]+ peak at m/z = 397.187 was also observed.
Unlike the Fe–Cl complex 1, the [Fe–N2]+ cation 5 does not form the cyclometallated product 4. Reaction of 5 with 3 equiv. n-BuLi at −35 °C gave a 31P{1H} NMR spectrum, exhibiting three signals in a roughly 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2
2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio, assigned as 2, 3 and a previously unobserved singlet at δP = 93.4 ppm, whose identity remains unknown (Scheme 2). Conducting the reaction at −78 °C likewise failed to generate 4 (see SI). Further differences between Cp* and the CpMe systems emerged upon treatment of [Cp*Fe(dnppe)(Cl)] with n-BuLi at −78 °C. This reaction also generated a product consistent with an Fe-κ3-P,C,P-dnppe coordination mode, however, in this case the Fe “tucked-in” complex is formed preferentially (Scheme 1b, right; see SI for details). These observations underscore a fundamental distinction between the CpMe and Cp* platforms. Under comparable conditions, the tucked-in product is more readily accessed in the Cp*Fe system, likely reflecting the greater number of methyl C–H sites available for deprotonation (five-fold higher relative to CpMe). Nevertheless, the κ3-P,C,P coordination motif is common to both systems, highlighting the general accessibility of this binding mode across the two ligand environments.
1 ratio, assigned as 2, 3 and a previously unobserved singlet at δP = 93.4 ppm, whose identity remains unknown (Scheme 2). Conducting the reaction at −78 °C likewise failed to generate 4 (see SI). Further differences between Cp* and the CpMe systems emerged upon treatment of [Cp*Fe(dnppe)(Cl)] with n-BuLi at −78 °C. This reaction also generated a product consistent with an Fe-κ3-P,C,P-dnppe coordination mode, however, in this case the Fe “tucked-in” complex is formed preferentially (Scheme 1b, right; see SI for details). These observations underscore a fundamental distinction between the CpMe and Cp* platforms. Under comparable conditions, the tucked-in product is more readily accessed in the Cp*Fe system, likely reflecting the greater number of methyl C–H sites available for deprotonation (five-fold higher relative to CpMe). Nevertheless, the κ3-P,C,P coordination motif is common to both systems, highlighting the general accessibility of this binding mode across the two ligand environments.
We have identified that treatment of [{CpMe}Fe(dnppe)(Cl)] (1) with base results in deprotonation of the diphosphine ligand backbone, affording a strained κ3-P,C,P Fe iron complex (4). Combined spectroscopic and computational analyses reveal its unusual structure and intrinsic instability and point to a potential phosphine decomposition pathway that may warrant consideration when evaluating the reactivity of metal–diphosphine systems under basic conditions.
The authors are grateful to Western University, the Council of Ontario Universities for a John C. Polanyi award to M. W. D., the Canadian Foundation for Innovation (LOF-212442), the Natural Sciences and Engineering Research Council of Canada (Discovery Grant, RGPIN-2020-04480 (M. W. D.), Discovery Launch Supplement, DGECR-2020-00183), and graduate award (CGRS-M to L. J. T.) for funding. The authors also thank Mr Quentin Lepeintre for initial synthesis of NaCpMe and Dr Mathew Willans for assistance in the simulation of 1H NMR spectral data.
Conflicts of interest
There are no conflicts to declare.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d6cc01011c.CCDC 2527971 (1) and 2527972 (6) contain the supplementary crystallographic data for this paper.29a,b
References
- P. Dierkes and P. W. N. M. van Leeuwen, J. Chem. Soc., Dalton Trans., 1999, 1519–1530 RSC
.
- H. Shet, U. Parmar, S. Bhilare and A. R. Kapdi, Org. Chem. Front., 2021, 8, 1599–1656 RSC
.
- D. Zhao, T. M. Neubauer and B. L. Feringa, Nat. Commun., 2015, 6, 6652 CrossRef CAS PubMed
.
- E. S. Wiendner, A. M. Appel, S. Raugei, W. J. Shaw and R. M. Bullock, Chem. Rev., 2022, 122, 12427–12474 CrossRef PubMed
.
- M. W. Drover, Chem. Soc. Rev., 2022, 51, 1861–1880 RSC
.
- T. Benincori, E. Brenna, F. Sannicolo, L. Trimarco, P. Antongnazza, E. Cesarotti, F. Demartin and T. Pilati, J. Org. Chem., 1996, 61, 6244–6251 CrossRef CAS PubMed
.
- F. Zhang, G.-Q. Chen and X. Zhang, Org. Lett., 2024, 26, 1623–1628 CrossRef CAS PubMed
.
- T. Saito, T. Yokozawa, T. Ishizaki, T. Moroi, N. Sayo, T. Miura and H. Kumobayashi, Adv. Synth. Catal., 2001, 343, 264–267 CrossRef CAS
.
- S. Aghazada, D. Munz, F. W. Heinemann, A. Scheurer and K. Meyer, J. Am. Chem. Soc., 2021, 143, 17219–17225 CrossRef CAS PubMed
.
- T. Ikariya and A. Yamamoto, J. Organomet. Chem., 1976, 118, 65–77 CrossRef CAS
.
- T. Simler, G. Frison, P. Braunstein and A. A. Danopoulos, Dalton Trans., 2016, 45, 2800–2804 RSC
.
- D. Amoroso, M. Haaf, G. P. A. Yap, R. West and D. E. Fogg, Organometallics, 2002, 21, 534–540 CrossRef CAS
.
- C. Ghosh, T. L. Groy, A. C. Bowman and R. J. Trovitch, Chem. Commun., 2016, 52, 4552 Search PubMed
.
- P. M. Fanara, V. Vigneswaran, P. S. Gunasekera, S. N. MacMillan and D. C. Lacy, Organometallics, 2022, 41, 93–98 CrossRef CAS
.
- L. S. Merz, H. Wadepohl, E. Clot and L. H. Gade, Chem. Sci., 2018, 9, 5223–5232 RSC
.
- X. Ye, P. N. Plessow, M. K. Brinks, M. Schelwies, T. Schaub, F. Rominger, R. Paciello, M. Limbach and P. Hofmann, J. Am. Chem. Soc., 2014, 136, 5923–592915 CrossRef CAS PubMed
.
- R. Beck, U. Flörke and H. F. Klein, Organometallics, 2015, 34, 1454–1464 CrossRef CAS
.
- J. A. Zurakowski, B. J. H. Austen, K. R. Brown and M. W. Drover, Chem. Commun., 2022, 58, 2500–2503 RSC
.
- M. L. Clapson, H. Sharma, J. A. Zurakowski and M. W. Drover, Chem. – Eur. J., 2023, 29, e202203763 CrossRef CAS PubMed
.
- B. J. H. Austen, H. Sharma, J. A. Zurakowski and M. W. Drover, Organometallics, 2022, 41, 2709–2715 CrossRef CAS
.
- J. A. Zurakowski, K. R. Brown and M. W. Drover, Inorg. Chem., 2023, 62, 7053–7060 CrossRef CAS PubMed
.
- J. A. Zurakowski, M. Bhattacharyya, D. M. Spasyuk and M. W. Drover, Inorg. Chem., 2020, 60, 37–41 CrossRef PubMed
.
- J. A. Zurakowski and M. W. Drover, Chem. Commun., 2023, 59, 11349–11352 RSC
.
- C. S. Durfy, J. A. Zurakowski and M. W. Drover, Angew. Chem., Int. Ed., 2025, 64, e202421599 CrossRef CAS PubMed
.
- E. J. Palmer, R. J. Strittmatter, K. T. Thornley, J. C. Gallucci and B. E. Bursten, Polyhedron, 2013, 58, 120–128 CrossRef CAS
.
- M. Muhr, P. Heiss, M. Schutz, R. Buhler, C. Gemel, M. H. Linden, H. B. Linden and R. A. Fischer, Dalton Trans., 2021, 50, 9031–9036 RSC
.
- J. H. Gross, Eur. J. Mass Spectrom., 2020, 26, 241–273 CrossRef CAS PubMed
.
- P. E. Garrou, Chem. Rev., 1981, 81, 229–266 CrossRef CAS
.
-
(a)
CCDC 2527971: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2qvkgm
; (b) CCDC 2527972: Experimental Crystal Structure Determination, 2026 DOI:10.5517/ccdc.csd.cc2qvkhn
.
| This journal is © The Royal Society of Chemistry 2026 |