Open Access Article
Open Access ArticleTargeted detection method for locus-specific m6A modifications in low-abundance transcripts based on chemical conversion
Ting
Liu†
a,
Xin
Fang†
a,
Yue
Lu
a,
Xiumin
Liu
a,
Ruiqi
Zhao
*a,
Xiang
Zhou
 ab and
Xiaocheng
Weng
ab and
Xiaocheng
Weng
 *ab
*ab
aCollege of Chemistry and Molecular Sciences, State Key Laboratory of Metabolism and Regulation in Complex Organisms, Wuhan University, Wuhan, Hubei 430072, China. E-mail: rqzhao@whu.edu.cn; xcweng@whu.edu.cn
bWuhan TaiKang Center for Life and Medical Sciences, Wuhan University, Wuhan, Hubei 430071, P. R. China
First published on 18th March 2026
Abstract
In this study, we developed a method for locus-specific m6A detection, which employs sodium nitrite and glyoxal-based chemical conversion with probe hybridization. This approach enables highly sensitive detection of target m6A sites even in low-abundance transcripts.
Since the discovery of pseudouridine in 1957,1 over 170 types of RNA modifications have been identified.2 Among these, N6-methyladenosine (m6A) stands out as one of the most abundant endogenous chemical modifications in eukaryotic mRNAs. Its dynamic regulation is orchestrated by three classes of evolutionarily conserved enzymes: “writers” (e.g., METTL3 and METTL143) that catalyze methylation; “erasers” (e.g., FTO4 and ALKBH55) responsible for demethylation; and “readers” (e.g., the YTH domain family6) which recognize m6A sites. These enzymes regulate m6A dynamics, driving the developement of activity detection, which have enabled applications in single-cell imaging and cancer diagnosis.7,8 The m6A modification directly influences RNA and regulates diverse developmental and cellular processes. Notably, m6A levels are closely linked to the pathogenesis of numerous diseases, positioning m6A as a potential diagnostic biomarker and therapeutic target.9–12 Consequently, the precise detection of m6A sites is critically important for both fundamental research and biomedical applications.
The first transcriptome-wide m6A mapping was achieved in 2012 using m6A-seq13 and MeRIP-seq.14 Subsequent refinements integrated UV crosslinking into antibody-based approaches, such as miCLIP,15 PA-m6A-seq16 and m6ACE-seq17 to achieve higher resolution. However, these methods remain constrained by relatively low resolution and high RNA input requirements. To overcome these limitations, various antibody-independent strategies have emerged. Methods such as m6A-label-seq,18 m6A-SAC-seq,19 MePMe-seq,20 m6A-SEAL-seq,21 MAZTER-seq22 and m6A-REF-seq23 achieved single-base resolution through metabolic labeling or m6A-sensitive enzymes. In parallel, techniques including the DART-seq,24 eTAM-seq,25 GLORI,26,27 and CAM-seq28 employ enzymatic or chemical conversion to induce selective deamination, enabling transcriptome-wide m6A profiling by high-throughput sequencing. While these methods enable transcriptome-wide m6A profiling, clinical diagnostics increasingly require locus-specific detection method. The SCARLET method first validated single-nucleotide m6A detection in 2013.29 Subsequent techniques, such as SELECT,30 LEAD-m6A-seq,31 E-IEXPAR32 and Rol-LAMP33 exploit the ability of m6A to block polymerase or ligase extension, converting modification status into quantifiable amplification signal. Other methods like TARS,34 m6AISH-PLA35 and m6A-PHPEA36 visualize locus-specific m6A sites in cells via fluorescence labeling. Advanced m6A detection methods based on single-molecule counting have also been developed (see reviews for details).37 However, these methods mainly rely on the enzymes selectivity for m6A sites, which generally leads to poor selectivity compared to chemical approaches. Consequently, new detection methods are still needed to accurately identify m6A sites, particularly in low-abundance transcripts.
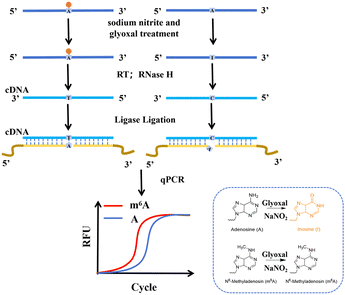
To advance locus-specific m6A detection, we develop a targeted method that integrates chemical conversion with ligation-mediated qPCR amplification, enabling sensitive detection with minimal sample input. The experimental workflow comprises: (1) chemical conversion of unmodified adenosines using sodium nitrite and glyoxal; (2) reverse transcription to generate cDNA; and (3) PCR amplification to enhance signals from low-abundance transcripts. Complementary UP and DOWN probes, flanking the target site, are hybridized to the cDNA. If the adenosine is m6A-modified, it resists chemical conversion, enabling efficient probe ligation via the primer sequences at their ends and subsequent amplification. Conversely, an unmodified adenosine (A) is chemically converted, preventing ligation and reducing product yield. This difference is quantified by a shift in the qPCR cycle threshold (Ct), enabling sensitive and single-base resolution detection of m6A (Fig. 1). If there are varying degrees of modification at this site, it can still be confirmed that there is some modification by comparing the Ct values before and after demethylation.
We first validated the method using synthetic oligonucleotides (45 DNA T and 45 DNA C, Table S1), which mimic cDNA from reverse-transcribed RNA treated with sodium nitrite and glyoxal, containing either m6A (45 DNA T) or unmodified A (45 DNA C). Ligation efficiency was compared under two base-pairing modes (A–T and G–C), corresponding to target sites with m6A (read as A) or converted A (read as G) (Fig. S1A). Corresponding upstream probes (ending in A or G) and a downstream probe were designed to flank the target site. qPCR analysis revealed distinct ligation products between 45 DNA T and 45 DNA C under both pairing modes (Fig. S1B and C), confirming that the ligation-based qPCR strategy effectively discriminates m6A from unmodified A. As the A–T mode showed a more pronounced difference in product yield, it was selected for all subsequent experiments.
To assess the impact of terminal base mismatches on ligation efficiency, a series of probes were designed with variations at terminal or penultimate positions: UP probes with different terminal bases (UP-A, UP-G, UP-T, UP-C) or penultimate-base substitutions (UP-2G, UP-2A, UP-2T, UP-2C), and DOWN probes with varying terminal bases (DOWN-C, DOWN-A, DOWN-T, DOWN-G) (sequences in Table S1). Correctly matched probes (UP-A, UP-G, UP-2G, and DOWN-C) yielded the highest ligation efficiency, whereas all mismatched combinations substantially reduced product formation (Fig. S2). We then optimized the ligation step by comparing Hifi Taq, SplintR, and T4 DNA ligases. Reaction conditions—including enzyme concentration (2, 5, 10, 15, and 25 U), temperature (40, 50, 60, and 70 °C), and time (15, 30, 45, and 60 min) were systematically screened (Fig. S3 and S4). Initial validation showed that T4 DNA Ligase had lower efficiency than the other two enzymes (Fig. S3). Further optimization revealed that HiFi Taq DNA Ligase performs better at lower concentrations. The reduced sequence complexity after chemical treatment allowed higher ligation temperatures, promoting more accurate hybridization. HiFi Taq DNA Ligase achieved maximal product yield at 70 °C, making it the optimal choice (Fig. S4). Based on product yield and discrimination between matched and mismatched probes, 2 U Hifi Taq DNA Ligase at 70 °C for 30 min was selected as optimal. Furthermore, placing the adenosine (A) at the 3′ terminus of the UP probe proved more efficient than at the 5′ end of the DOWN probe (Fig. S3).
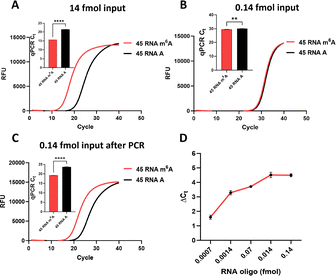
Next, we determined the detection limit using synthetic RNA oligonucleotides (45 RNA A and 45 RNA m6A, Table S1). With 14 fmol of input RNA, the cycle-threshold difference (ΔCt) between m6A and A reached 6.11 cycles, corresponding to 69.07-fold selectivity (Fig. 2A). However, detection became unreliable below 0.14 fmol input RNA (Fig. 2B). Introducing a pre-amplification step (35-cycle PCR of cDNA prior to ligation) restored m6A detection at 0.14 fmol, yielding a ΔCt of 4.1 cycles (Fig. 2C). Further dilution experiments showed the method could distinguish m6A from A at inputs as low as 0.0007 fmol (Fig. 2D)—significantly below the 0.25 fmol required by SELECT.30 These results demonstrate that our approach enables highly sensitive, locus-specific m6A detection, making it suitable for analyzing low-abundance transcripts in limited samples.
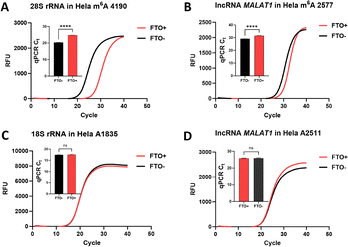
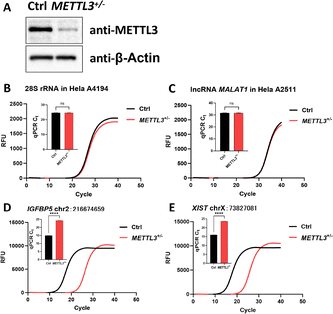
After method validation and optimization, we assessed its performance in biological samples. Using FTO-mediated demethylation to convert m6A to A at specific sites, we directly compared modification states before and after treatment within the same sample (Fig. S7). If this site has certain m6A modifications, after FTO treatment, the m6A/A ratio will show changes. Through chemical treatment, these changes will be amplified, thereby detecting that this site has certain m6A modifications. After validating the enzymatic activity of FTO (Fig. S5 and S6), our method detected specific m6A sites on rRNA, lncRNA, and mRNA by monitoring relative ligation product yields before and after FTO-mediated demethylation. In HeLa cells, the method accurately identified the known m6A site at 28S rRNA 4190 (Fig. 3A), while no significant change was observed at 18S rRNA A1835 after input normalization (Fig. 3C). Similarly, it successfully detected the known m6A site 2577 but not the A site 2511 on lncRNA MALAT1 (Fig. 3B and D). GAPDH served as an internal reference for input normalization throughout.
Subsequently, we selected candidate m6A sites from published sequencing data for experimental validation.30 By comparing the normalized Ct differences between FTO-treated and untreated samples, we confirmed m6A presence at multiple sites on tRNA and mRNA26 (e.g., TRNAK-CUU, IGFBP5, PPRC1; Fig. S8). These results validate both the existence of these sites modification and the broad applicability of our method for locus-specific m6A detection across diverse RNA types.
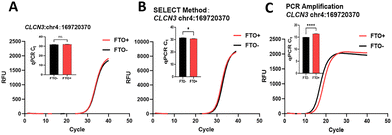
To evaluate our method's performance for low-abundance transcripts, we selected the low-expression mRNA CLCN3 (TPM 13) (TPM stands for Transcripts Per Million, a normalized unit that quantifies gene expression levels38). Using approximately 10 ng of cDNA input, we could not reliably detect the reported m6A site in CLCN325 (Fig. 4A). We attribute this failure to substantial transcript loss during FTO treatment coupled with low transcript abundance. Then we compare our method with SELECT, a widely used approach for detecting m6A modifications due to its simplicity and flexibility. Notably, the SELECT method also failed to detect this site under same amount of RNA input (Fig. 4B). However, after amplifying the target cDNA by 35-cycle PCR, our method successfully confirmed the m6A site on CLCN3 (Fig. 4C). The SELECT method exploits the difference in steric hindrance between m6A and A in RNA to mediate template-driven probe ligation, enabling m6A detection. Although RNA is difficult to amplify and not directly compatible with PCR due to DNA polymerases' template specificity, our approach employs chemical treatment followed by reverse transcription to convert modification information into sequence differences in cDNA. Using cDNA as the substrate for ligation-based detection allows compatibility with PCR amplification, thereby enhancing signal from limited RNA input. For a fair comparison, both methods were assessed under identical conditions, starting with equal amounts of input RNA before processing. The experimental results also indicate a clear advantage over SELECT for low-abundance transcripts and highlights the utility of our approach when sample material is limited.
We next compared METTL3+/− HeLa cells with wild-type controls, as METTL3 knockdown reduces global m6A levels. After knocking out the METTL3, the degree of methylation at some sites changed. Through our method, such changes can be detected. Western blot confirmed markedly lower METTL3 protein in METTL3+/− cells (Fig. 5A, the uncropped images were provided in the Fig. S9).
To control for RNA input variation, samples were normalized by transcript abundance, validating 28S rRNA A4194 sites and lncRNA MALAT1 A2511 in HeLa cells. Following normalization, no significant difference in qPCR Ct values was observed between METTL3+/− cells and wild-type cells for these A sites (Fig. 5B and C). We then validated several previously reported m6A sites (in IGFBP5, xist, RRP12, TPGS2 and USP47) from sequencing datasets.26 Ligation product yields at these sites were consistently lower in METTL3+/− cells than in wild-type controls (Fig. 5D, E and Fig. S10), confirming them as m6A modification sites in HeLa cells.
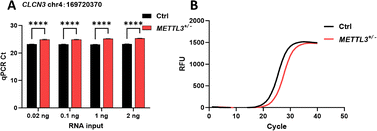
To assess the method's performance for low-abundance transcripts, we selected three low-expression gene loci and successfully detected m6A modifications (Fig. S11). For the low-expression transcript CLCN3, a serial mRNA dilution was performed; after reverse transcription and 35-cycle PCR amplification, m6A was consistently detected with inputs as low as 0.02 ng (Fig. 6). These results confirm the method's applicability for samples with limited transcript abundance. We assessed the detection threshold using model oligonucleotides with varying m6A ratios. As shown in Fig. S12, the method detected m6A levels as low as 20%, with a 2.12 ΔCt relative to the unmodified control, corresponding to a 4.36-fold difference in ligation product abundance.
In summary, we developed a locus-specific m6A detection method and successfully identified m6A sites in various RNA types, including rRNA, lncRNA, and mRNA. Our method converts modification information into sequence differences in cDNA via chemical transformation and reverse transcription. Using amplifiable cDNA as the template enables sensitive detection even with low RNA input. Although the chemical treatment reduces sequence complexity and may limit target selection, the method provides a practical framework for detecting RNA modifications in rare biological samples and, with probe redesign, could be extended to other RNA modifications.
Conflicts of interest
There are no conflicts to declare.Data availability
The data that support the findings of this study are available in the supplementary information (SI) of this article. Supplementary information is available. See DOI: https://doi.org/10.1039/d6cc00494f.Acknowledgements
This work was supported by grants from the Noncommunicable Chronic Diseases-National Science and Technology Major Project (2023ZD0507700), the National Natural Science Foundation of China (22588302).References
- F. F. Davis and F. W. Allen, J. Biol. Chem., 1957, 227, 907–915 CrossRef CAS PubMed.
- P. Boccaletto, F. Stefaniak, A. Ray, A. Cappannini, S. Mukherjee, E. Purta, M. Kurkowska, N. Shirvanizadeh, E. Destefanis, P. Groza, G. Avşar, A. Romitelli, P. Pir, E. Dassi, S. G. Conticello, F. Aguilo and J. M. Bujnicki, Nucleic Acids Res., 2022, 50, D231–D235 CrossRef CAS PubMed.
- J. Liu, Y. Yue, D. Han, X. Wang, Y. Fu, L. Zhang, G. Jia, M. Yu, Z. Lu, X. Deng, Q. Dai, W. Chen and C. He, Nat. Chem. Biol., 2013, 10, 93–95 CrossRef PubMed.
- G. Jia, Y. Fu, X. Zhao, Q. Dai, G. Zheng, Y. Yang, C. Yi, T. Lindahl, T. Pan, Y.-G. Yang and C. He, Nat. Chem. Biol., 2011, 7, 885–887 CrossRef CAS PubMed.
- G. Zheng, J. A. Dahl, Y. Niu, P. Fedorcsak, C.-M. Huang, C. J. Li, C. B. Vågbø, Y. Shi, W.-L. Wang, S.-H. Song, Z. Lu, R. P. G. Bosmans, Q. Dai, Y.-J. Hao, X. Yang, W.-M. Zhao, W.-M. Tong, X.-J. Wang, F. Bogdan, K. Furu, Y. Fu, G. Jia, X. Zhao, J. Liu, H. E. Krokan, A. Klungland, Y.-G. Yang and C. He, Mol. Cell, 2013, 49, 18–29 CrossRef CAS PubMed.
- I. A. Roundtree, G.-Z. Luo, Z. Zhang, X. Wang, T. Zhou, Y. Cui, J. Sha, X. Huang, L. Guerrero, P. Xie, E. He, B. Shen and C. He, eLife, 2017, 6, e31311 CrossRef PubMed.
- Q. Zhang, S. Zhao, C. Su, Q. Han, Y. Han, X. Tian, Y. Li and C.-Y. Zhang, Anal. Chem., 2023, 95, 13201–13210 CrossRef CAS PubMed.
- W.-j Liu, L.-y Wang, Z. Sheng, B. Zhang, X. Zou and C.-y Zhang, Biosens. Bioelectron., 2023, 240, 115645 CrossRef CAS PubMed.
- C. Zhang, Y. Chen, B. Sun, L. Wang, Y. Yang, D. Ma, J. Lv, J. Heng, Y. Ding, Y. Xue, X. Lu, W. Xiao, Y.-G. Yang and F. Liu, Nature, 2017, 549, 273–276 CrossRef CAS PubMed.
- Y. Xiang, B. Laurent, C.-H. Hsu, S. Nachtergaele, Z. Lu, W. Sheng, C. Xu, H. Chen, J. Ouyang, S. Wang, D. Ling, P.-H. Hsu, L. Zou, A. Jambhekar, C. He and Y. Shi, Nature, 2017, 543, 573–576 CrossRef CAS PubMed.
- H. Shi, X. Zhang, Y.-L. Weng, Z. Lu, Y. Liu, Z. Lu, J. Li, P. Hao, Y. Zhang, F. Zhang, Y. Wu, J. Y. Delgado, Y. Su, M. J. Patel, X. Cao, B. Shen, X. Huang, G.-L. Ming, X. Zhuang, H. Song, C. He and T. Zhou, Nature, 2018, 563, 249–253 CrossRef CAS PubMed.
- R. Winkler, E. Gillis, L. Lasman, M. Safra, S. Geula, C. Soyris, A. Nachshon, J. Tai-Schmiedel, N. Friedman, V. T. K. Le-Trilling, M. Trilling, M. Mandelboim, J. H. Hanna, S. Schwartz and N. Stern-Ginossar, Nat. Immunol., 2018, 20, 173–182 CrossRef PubMed.
- D. Dominissini, S. Moshitch-Moshkovitz, S. Schwartz, M. Salmon-Divon, L. Ungar, S. Osenberg, K. Cesarkas, J. Jacob-Hirsch, N. Amariglio, M. Kupiec, R. Sorek and G. Rechavi, Nature, 2012, 485, 201–206 CrossRef CAS PubMed.
- K. D. Meyer, Y. Saletore, P. Zumbo, O. Elemento, C. E. Mason and S. R. Jaffrey, Cell, 2012, 149, 1635–1646 CrossRef CAS PubMed.
- B. Linder, A. V. Grozhik, A. O. Olarerin-George, C. Meydan, C. E. Mason and S. R. Jaffrey, Nat. Methods, 2015, 12, 767–772 CrossRef CAS PubMed.
- K. Chen, Z. Lu, X. Wang, Y. Fu, G. Z. Luo, N. Liu, D. Han, D. Dominissini, Q. Dai, T. Pan and C. He, Angew. Chem., Int. Ed., 2014, 54, 1587–1590 CrossRef PubMed.
- C. W. Q. Koh, Y. T. Goh and W. S. S. Goh, Nat. Commun., 2019, 10, 5636 CrossRef CAS PubMed.
- X. Shu, J. Cao, M. Cheng, S. Xiang, M. Gao, T. Li, X. Ying, F. Wang, Y. Yue, Z. Lu, Q. Dai, X. Cui, L. Ma, Y. Wang, C. He, X. Feng and J. Liu, Nat. Chem. Biol., 2020, 16, 887–895 CrossRef CAS PubMed.
- L. Hu, S. Liu, Y. Peng, R. Ge, R. Su, C. Senevirathne, B. T. Harada, Q. Dai, J. Wei, L. Zhang, Z. Hao, L. Luo, H. Wang, Y. Wang, M. Luo, M. Chen, J. Chen and C. He, Nat. Biotechnol., 2022, 40, 1210–1219 CrossRef CAS PubMed.
- K. Hartstock, N. A. Kueck, P. Spacek, A. Ovcharenko, S. Hüwel, N. V. Cornelissen, A. Bollu, C. Dieterich and A. Rentmeister, Nat. Commun., 2023, 14, 7154 CrossRef CAS PubMed.
- Y. Wang, Y. Xiao, S. Dong, Q. Yu and G. Jia, Nat. Chem. Biol., 2020, 16, 896–903 CrossRef CAS PubMed.
- M. A. Garcia-Campos, S. Edelheit, U. Toth, M. Safra, R. Shachar, S. Viukov, R. Winkler, R. Nir, L. Lasman, A. Brandis, J. H. Hanna, W. Rossmanith and S. Schwartz, Cell, 2019, 178(731–747), e716 Search PubMed.
- H.-X. Chen, Z. Zhang, D.-Z. Ma, L.-Q. Chen and G.-Z. Luo, Methods, 2022, 203, 392–398 CrossRef CAS PubMed.
- K. D. Meyer, Nat. Methods, 2019, 16, 1275–1280 CrossRef CAS PubMed.
- Y.-L. Xiao, S. Liu, R. Ge, Y. Wu, C. He, M. Chen and W. Tang, Nat. Biotechnol., 2023, 41, 993–1003 CrossRef CAS PubMed.
- C. Liu, H. Sun, Y. Yi, W. Shen, K. Li, Y. Xiao, F. Li, Y. Li, Y. Hou, B. Lu, W. Liu, H. Meng, J. Peng, C. Yi and J. Wang, Nat. Biotechnol., 2022, 41, 355–366 CrossRef PubMed.
- H. Sun, B. Lu, Z. Zhang, Y. Xiao, Z. Zhou, L. Xi, Z. Li, Z. Jiang, J. Zhang, M. Wang, C. Liu, Y. Ma, J. Peng, X.-J. Wang and C. Yi, Nat. Methods, 2025, 22, 1226–1236 CrossRef CAS PubMed.
- P. Wang, C. Ye, M. Zhao, B. Jiang and C. He, Nat. Chem., 2025, 17, 1042–1052 CrossRef CAS PubMed.
- N. Liu, M. Parisien, Q. Dai, G. Zheng, C. He and T. Pan, RNA, 2013, 19, 1848–1856 CrossRef CAS PubMed.
- Y. Xiao, Y. Wang, Q. Tang, L. Wei, X. Zhang and G. Jia, Angew. Chem., Int. Ed., 2018, 57, 15995–16000 CrossRef CAS PubMed.
- Y. Wang, Z. Zhang, C. Sepich-Poore, L. Zhang, Y. Xiao and C. He, Angew. Chem., Int. Ed., 2020, 60, 873–880 CrossRef PubMed.
- F. Su, W. Liu, K. Gao, D. Chen, Y. Cheng and Z. Li, Anal. Chim. Acta, 2024, 1302, 342474 CrossRef CAS PubMed.
- J. Li, J. Zhou, Y. Xia, Y. Rui, X. Yang, G. Xie, G. Jiang and H. Wang, Nucleic Acids Res., 2023, 51, e51 CrossRef CAS PubMed.
- Q. Zhang, Y. Dai, X. Teng and J. Li, Angew. Chem., Int. Ed., 2024, 64, e202420977 CrossRef PubMed.
- X. Ren, R. Deng, K. Zhang, Y. Sun, Y. Li and J. Li, Angew. Chem., Int. Ed., 2021, 60, 22646–22651 CrossRef CAS PubMed.
- M. Song, J. Wang, J. Hou, T. Fu, Y. Feng, W. Lv, F. Ge, R. Peng, D. Han and W. Tan, ACS Nano, 2024, 18, 27537–27546 CrossRef CAS PubMed.
- H.-J. Li, J. Hu, W.-J. Liu and C.-Y. Zhang, TrAC, Trends Anal. Chem., 2026, 194, 118537 CrossRef CAS.
- G. P. Wagner, K. Kin and V. J. Lynch, Theory Biosci., 2012, 131, 281–285 CrossRef CAS PubMed.
Footnote |
| † These authors contributed equally. |
| This journal is © The Royal Society of Chemistry 2026 |