Nanobodies targeting ion channels: advancing therapeutics through precision and structural insights
Mukund Sudharsan
M G
 a,
Somanath
Baral
a,
Somanath
Baral
 ab,
Sushree Ankita
Mohapatra
ab,
Sushree Ankita
Mohapatra
 a,
Ithayaraja
Mahendran
a,
Ithayaraja
Mahendran
 a and
Janesh
Kumar
a and
Janesh
Kumar
 *ab
*ab
aMembrane Protein Biology Group, CSIR - Centre for Cellular & Molecular Biology, Uppal Road, Habsiguda, Hyderabad, Telangana-500007, India. E-mail: janesh.ccmb@csir.res.in
bAcademy of Scientific and Innovation Research (AcSIR), Ghaziabad, India
First published on 26th November 2025
Abstract
Ion channels, which are integral to cellular signaling and homeostasis, are implicated in a variety of diseases, including neurological disorders, cardiovascular conditions, and cancers. Despite their importance, targeting ion channels therapeutically is challenging due to issues such as structural conservation and dynamic conformational changes. Nanobodies, derived from camelid heavy-chain antibodies, have emerged as a transformative tool in biotherapeutics due to their small size, remarkable stability, and high specificity for epitope recognition. They represent a significant advancement in targeting ion channels, offering precision and adaptability not achievable with traditional antibodies or small molecules. This review provides an in-depth analysis of the structural and functional basis of ion channel modulation by nanobodies. We explore how they achieve their effects through diverse mechanisms, including direct pore blocking, allosteric modulation of gating, stabilization of specific conformational states, and isoform-selective recognition. We further examine their therapeutic applications, the potential for engineering multivalent constructs to enhance efficacy, and critical advancements in overcoming challenges like blood–brain barrier (BBB) penetration. Finally, we discuss recent clinical developments, the translational gap for ion channel targets, and the transformative potential of artificial intelligence in accelerating the design of next-generation nanobody-based therapeutics.
Introduction
Ion channels are integral membrane proteins that mediate the selective passage of ions across cellular membranes, playing critical roles in maintaining cellular homeostasis, electrical excitability, and signal transduction.1,2 These proteins are essential for various physiological processes, including neuronal firing, muscle contraction, hormone secretion, and fluid regulation. Classified based on their gating mechanisms, ion channels include voltage-gated, ligand-gated, mechanosensitive, and temperature-sensitive channels, each with specialized functions within excitable and non-excitable tissues.1,2The significance of ion channels extends beyond their physiological roles; their dysregulation has been implicated in numerous diseases. Neurological disorders such as epilepsy and chronic pain, cardiovascular abnormalities like arrhythmias and hypertension, and cancers characterized by aberrant ion channel expression and function represent a fraction of the conditions associated with ion channel dysfunction.3 Recent studies have also unveiled the roles of ion channels in immune modulation, metabolic regulation, and stem cell differentiation, underscoring their involvement in systemic pathophysiological processes.4–6
Therapeutically targeting ion channels is a complex endeavour. Structural conservation among channel isoforms, particularly in their pore regions, poses challenges for achieving specificity. For instance, sodium channel isoforms Nav1.7, Nav1.8, and Nav1.9 are highly homologous, yet distinct in their physiological roles in pain signaling, necessitating precise targeting to minimize off-target effects.7 Moreover, the dynamic conformational states of ion channels during gating further complicate drug development, as therapeutics must interact with transient structural configurations that are not readily accessible. While small molecules and peptide toxins have historically served as modulators, their therapeutic application is often limited by insufficient selectivity and systemic toxicity.
While providing enhanced specificity, conventional monoclonal antibodies are hindered by their substantial size (∼150 kDa) and bivalent architecture, which restricts their access to functional epitopes typically concealed within the intricate topography of these transmembrane proteins.8 Nanobodies, the 15 kDa single-variable domains (VHH) of camelid heavy-chain-only antibodies, are smaller than conventional antibodies but have comparable or better specificity, stability, and affinity. They are relatively easier and less expensive to purify compared to conventional antibodies. Nanobodies exhibit low immunogenicity and can be engineered into multivalent or biparatopic formats to enhance avidity and functional specificity, further broadening their therapeutic utility. A particularly transformative aspect of nanobody application lies in the structural elucidation of ion channels.8 Advances in cryo-electron microscopy (cryo-EM) and X-ray crystallography have enabled high-resolution visualization of nanobody-ion channel complexes, revealing mechanisms of action, allosteric modulation, and isoform-specific binding. These structural insights advance our understanding of ion channel biology and facilitate the rational design of next-generation therapeutics. These unique properties make them exceptionally well-suited for targeting proteins and specifically ion channels.
As nanobody-based therapies advance, their ability to cross the blood–brain barrier (BBB) has emerged as a critical advantage for targeting ion channels implicated in central nervous system (CNS) disorders. This review highlights the multifaceted applications of nanobodies in ion channel research and therapy. We organize our discussion around the diverse modulatory mechanisms revealed by structural biology, before examining therapeutic innovation, delivery challenges, and emerging clinical and computational prospects. Through these advancements, nanobody technology is poised to overcome longstanding challenges, offering new avenues for precision medicine in ion channel-associated diseases.
Nanobody – a small, modular and single-domain antibody
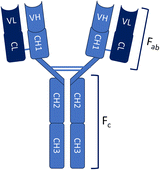
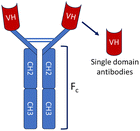
The immune system of the camelid family, which includes camels, llamas, and alpacas, produces a unique class of antibodies known as heavy-chain antibodies (HCAbs).9,10 These Heavy-chain antibodies are composed exclusively of heavy chains and entirely devoid of light chains, distinguished by the lack of the CH1 domain in their constant region, a structural characteristic that typically promotes the pairing of heavy and light chains. The variable region of the HCAb, which possesses the ability to bind to antigens effectively, is referred to as the variable domain of the heavy-chain antibody (VHH) or single-domain antibody (sdAb).11 VHH/sdAb are frequently referred to as nanobodies (Nb), a term that was initially trademarked by the Belgian biopharmaceutical firm Ablynx in 2003.12 In contrast to conventional antibodies (IgG), nanobodies exhibit distinct advantages as detailed in Table 1. Nanobodies are a superior alternative to conventional antibodies(IgG), demonstrating promising potential across a range of applications in structural biology, molecular imaging, disease diagnosis and treatment.| Characteristics | Conventional antibodies | Nanobodies |
|---|---|---|
| a Info compiled from multiple sources.9,11–13,77,100–102 | ||
|
|
|
| Encoded by gene | Two gene fragments (heavy and light chains) | Single-gene fragment (VHH) |
| Molecular weight | ∼150 kDa | ∼15 kDa |
| Structure | Heteromeric (2 heavy + 2 light chains) | Monomeric (single heavy-chain variable domain) |
| Mode of binding | Bivalent | Monovalent |
| Specificity | High | High |
| Affinity | nM–pM range | nM–pM range |
| Preferred expression system | Mammalian (CHO, HEK293) | Bacterial (e.g., E. coli), Yeast (e.g., Pichia pastoris, Saccharomyces cerevisiae) |
| Production yield | Moderate to low (complex folding, glycosylation required) | High (simple folding, no glycosylation required) |
| Solubility | Variable; may aggregate without light chains | Highly soluble (hydrophilic framework) |
| Stability | sensitive to pH and temperature | High thermal and chemical stability |
| Ease of manipulation for enhance affinity | Difficult to manipulate at the genetic level | Easily engineered via CDR3 modification |
| Tissue penetration | Limited due to the Large molecular size | High due to small molecular size |
| Development cost | High | Low to moderate |
| Paratope geometry | Concave/flat (fits protruding epitopes) | Convex (fits concave or buried epitopes) |
Nanobody development requires the generation of a library of nanobody and using it to identify specific high affinity binders against a antigen. The identification of specific nanobodies that effectively target particular antigens can be accomplished through three distinct methodologies, which are as follows: (i) the conventional naïve or immunization approach, (ii) the utilization of synthetic nanobody(Sybodies) libraries, and (iii) the application of computational techniques in conjunction with artificial intelligence-based modeling that focus on the design and characterization of nanobodies.13
(i) Library generation
The immunization of camelids involves the injection of purified antigen (Proteins) or whole cells expressing proteins on surface or virus-like particles into them. Typically, the PBMC's (Peripheral blood mononuclear cells) are isolated from the blood collected from the animal. Nanobody libraries are generated using the mRNA from the PBMCs, that are rich in high-affinity, conformationally selective variable heavy-chain domains (VHHs). Conversely, naïve or synthetic libraries, which are distinguished by their intentionally designed complementarity-determining region (CDR) diversity, facilitate a more rapid identification process in vitro, eliminating the necessity for animal immunization, and proving to be particularly advantageous in scenarios involving human targets or situations where immunization is not feasible or ethical.14(ii) Biopanning to isolate nanobody binders against antigen
The technologies employed for screening and selection play a crucial role in the conversion of VHH repertoires into effective and functional binders that can be utilized for various applications. Among these methodologies, phage display remains the predominant technique for achieving initial enrichment through a series of successive panning processes against antigens that have been immobilized, as seen in enzyme-linked immunosorbent assay (ELISA) formats. Yeast surface display library are panned using magnetic-activated cell sorting (MACS) and fluorescence-activated cell sorting (FACS), both of which allow for the precise differentiation of nanobodies based on key parameters such as binding affinity, specificity, and off-rate kinetics.12 Ribosome and mRNA display methodologies enable researchers to explore extensive synthetic repertoires in a completely controlled in vitro environment, which proves to be instrumental in the process of affinity maturation that enhances the binding capabilities of the nanobodies.15Artificial Intelligence (AI)-Driven technologies have reached a level of sophistication that significantly reduces the expenditure of costly resources and time. AI-Machine Learning (ML) has revolutionized novel protein design field. This topic has been further discussed in the later part of the literature under the section – The Digital Frontier: using AI to design and improve ion channel nanobodies.16
(iii) Validation of nanobody binders
Upon the identification of promising nanobodies, they can be further evaluated using various biophysical techniques and structural biology to ascertain their affinity and binding sites (Fig. 1).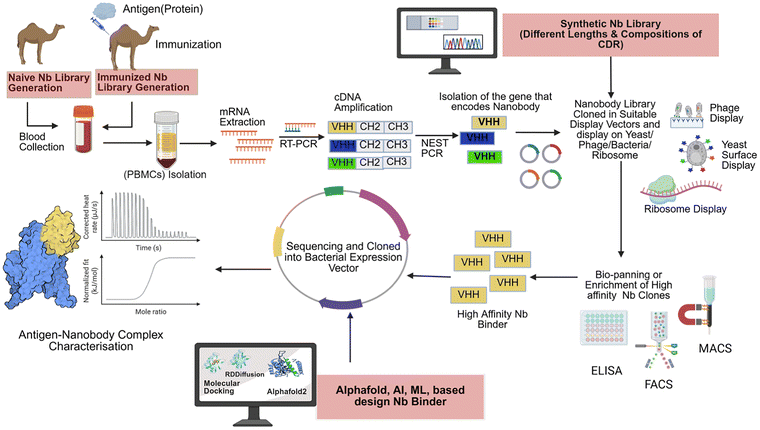 | ||
| Fig. 1 A flowchart for Nanobody generation strategies and methodologies. The image was created using Biorender. | ||
Structural and mechanistic insights into nanobody-mediated ion channel modulation
High-resolution structures of nanobody-channel complexes have revealed an astonishing diversity of modulatory mechanisms. Rather than simply acting as brute-force pore blockers, nanobodies function as sophisticated molecular machines that can stabilize specific conformational states, allosterically regulate gating, or even act as chaperones to enable structural studies of otherwise intractable targets (Table 2). Here, we classify these interactions based on their primary mechanism of action.| Nanobody | Target species and ion channel name | Nanobody library | Display platforms for biopanning | EC50/affinity (Kd) | Property | PDB structure |
|---|---|---|---|---|---|---|
| 13A7 | Mouse P2X7 | Immunized | Phage display | IC50 = 12 nM | Blocking (prevents gating) | No |
| 14D5 | Mouse P2X7 | Immunized | Phage display | EC50 = 6 nM | Potentiating | No |
| Dano1 | Human P2X7 | Immunized | Phage display | IC50 = 0.2 nM | Blocking (high potency) | No |
| A0194009G09 | human Kv1.3 | Immunized | Phage display | IC50 = 8.2 nM | Blocking (prevents gating)/Suppresses Kv1.3-mediated potassium currents/potent inhibitor | 8DFL |
| Nb.C1 | Human ASIC1a | Immunized | Phage display | — | With inhibitor PcTx1 inhibits the gating properties. Nb.C1 stabilizes and prevents aggregation of hASIC1a | 7RNN |
| Nb17, Nb82 | Human NaV1.4/NaV1.5 | Immunized | Phage display | Nb17: 41.1 ± 9. 9 nM (NaV1.4); 60.5 ± 5.80 nM (NaV1.5) Nb82: 50.2 ± 8.87 nM (NaV1.4); 63.2 ± 6.75 nM (NaV1.5) | Detect NaV1.4 & NaV1.5 channels in live cells and tissue homogenates | No, 7R63 (Nb only), Nb82, (no complex) |
| nb.E8 & nb.F3 | Mouse High-voltage activated calcium channels (HVACCs) | Immunized | Phage display | kD(nb.F3) = 13.2 ± 7.2 | Bind to the auxiliary subunit of HVACCS CaVβ1, inhibits recombinant CaV1/CaV2 channels reconstituted with CaVβ1 | 8DAM, 8E0E |
| Nb58 | Human TREK-2 | Immunized | Phage display | Used immunodetection and immunoassays of TREK2 | 8QZ1 | |
| Nb61 | Human TREK-2 | Immunized | Phage display | Kd = 1110 nM, IC50 = 685 ± 18 nM | Inhibiting channel-gating mechanisms reduces K+ conductance | 8QZ2 |
| Nb67 | Human TREK-2 | Immunized | Phage display | EC50 = 101 ± 12 nM | Channel opening/activation enhances K+ efflux | 8QZ3 |
| BiparatropicNb6158 | Human TREK-2 | — | — | Kd = 4.1 nM, 16-fold improvement of IC50 ∼40 nM | More inhibitory responses than Nb61 | EMD-19066 |
| Nb76 | Human TREK-2 | Immunized | Phage display | Not Reported | Channel opening/activation | 8QZ4 |
| Sb1 | Mouse homomeric LRRC8A | Synthetic | Ribosome display | Kd = 23 nM | Channel Inhibition | 7P5V |
| Sb2 | Mouse homomeric LRRC8A | Synthetic | Ribosome display | Kd = 48 nM | Channel Inhibition | 7P5W |
| Sb3 | Mouse homomeric LRRC8A | Synthetic | Ribosome display | Kd = 857 nM | Channel Inhibition | 7P5Y |
| Sb4 | Mouse homomeric LRRC8A | Synthetic | Ribosome display | Kd = 22 nM | Channel Inhibition | 7P60 |
| Sb5 | Mouse homomeric LRRC8A | Synthetic | Ribosome display | Kd = 173 nm | Channel Inhibition | 7P6K |
| Sb1 | Mouse heteromeric LRRC8A/C | Synthetic | Ribosome display | Channel Inhibition, structural stability | 8B41 | |
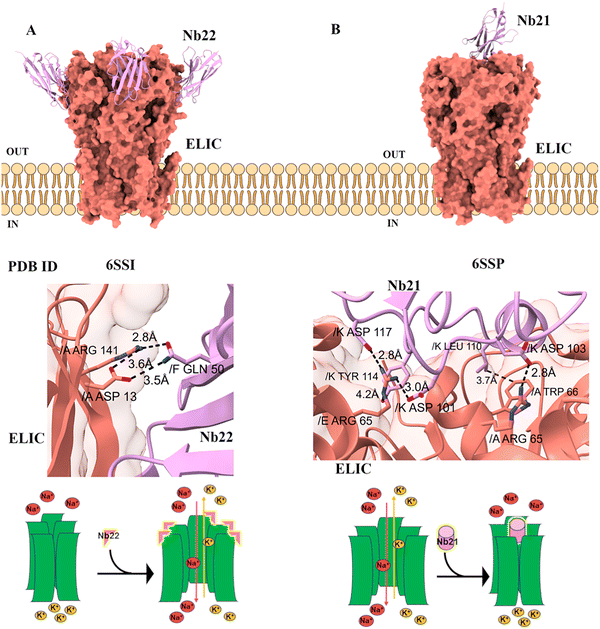
| Nb21 (NAM Nb) | Erwinia ligand-gated ion channel (Cys-loop ligand-gated ion channel) | Immunized | Phage display | IC50 = 0.13 mM | Negative modulation via vestibular binding | 6SSP |
| Nb22 (PAM Nb) | Erwinia ligand-gated ion channel (Cys-loop ligand-gated ion channel) | Immunized | Phage display | EC50 = 4.2 mM | Positive modulation via vestibular binding | 6SSI |
| Nb72 | Erwinia ligand-gated ion channel (Cys-loop ligand-gated ion channel) | Immunized | Phage display | 6HJX | ||
| Nb C4 | Human alpha7 nicotinic receptor/Neuronal acetylcholine receptor subunit alpha-7 | Immunized | Phage display | Kd = 0.2 ± 0.3 nM | Silent allosteric ligand (SAL), showing no effect on ACh-gated currents but inhibiting Nb-E3 elicited potentiation | 8C9X |
| Nb C4 | Human alpha7 nicotinic receptor/Neuronal acetylcholine receptor subunit alpha-7 | Immunized | Phage display | Kd = 0.2 ± 0.3 nM | With nicotine, Silent Allosteric Modulator | 8CAU |
| Nb E3 | Human alpha7 nicotinic receptor/Neuronal acetylcholine receptor subunit alpha-7 | Immunized | Phage display | Kd = 1 ± 0.01 nM | Potentiates ACh-gated currents | 8CE4 |
| Nb E3 | Human alpha7 nicotinic receptor/Neuronal acetylcholine receptor subunit alpha-7 | Immunized | Phage display | Kd = 1 ± 0.01 nM | With nicotine potentiates ACh-gated currents | 8CI1 |
| VHH15 | Mouse 5-HT3 receptor | Immunized | Phage display | Kd = 142 nM | Inhibits channel activity | 4PIR |
| Synthetic Nb SbC2 | Human CALHM2 | Synthetic | Ribosome display | Structural stabilization for cryo-EM | 8RMK | |
| Synthetic Nb SbC4 | Human CALHM2/4 (heteromer) | Synthetic | Ribosome display | Marker for heteromer assembly studies | 8RML | |
| Synthetic Nb SbC4 | Human CALHM4 (decamer dimer) | Synthetic | Ribosome display | Structural capture of large oligomeric states | 8RMN | |
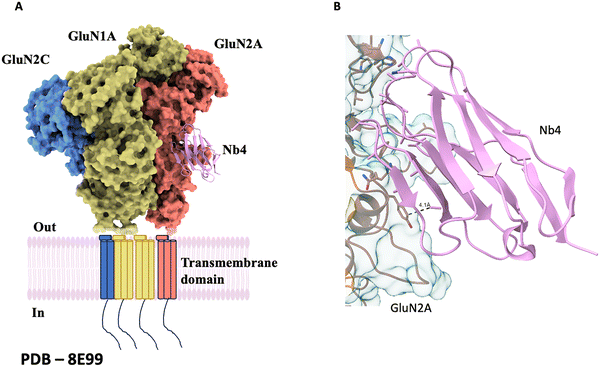
| Nb4 | Human NMDA (GluN1a–GluN2A–GluN2C) | Reduces structural heterogeneity | 8E99 | |||
| DN13 | Human metabotropic glutamate 2 (mGlu2) | Immunized | Phage display | Kd = 3.5 ± 0.6 nM with agonist | suppressed NMDAR gating currents | No |
| DN1 | Human metabotropic glutamate 2 (mGlu2) | Immunized | Phage display | Kd = 2.1 ± 0.4 nM with agonist | increase NMDA receptor (NMDAR) currents | No |
| DN13–DN1 | Human metabotropic glutamate 2 (mGlu2) | Precisely regulate NMDAR function | No | |||
| Nb38 | GABA(A) | Immunized | Phage display | Positive allosteric modulator | 9FFV, 9FFW, 9FFX, 9FFY 9FFZ, 9FG0, 9FG1, 9FG2, 9FG3 | |
| Nb25 | GABA(A) | Immunized | Phage display | Structural stabilization for cryo-EM | 5O8F, 5OJM |
1. Allosteric modulation of channel gating
Many nanobodies function by binding to sites remote from the ion conduction pore, inducing conformational changes that either promote or inhibit channel activity. This allosteric approach offers a powerful route to achieving subtype selectivity.Targeting extracellular domains: the case of TREK-2 and ASIC1a
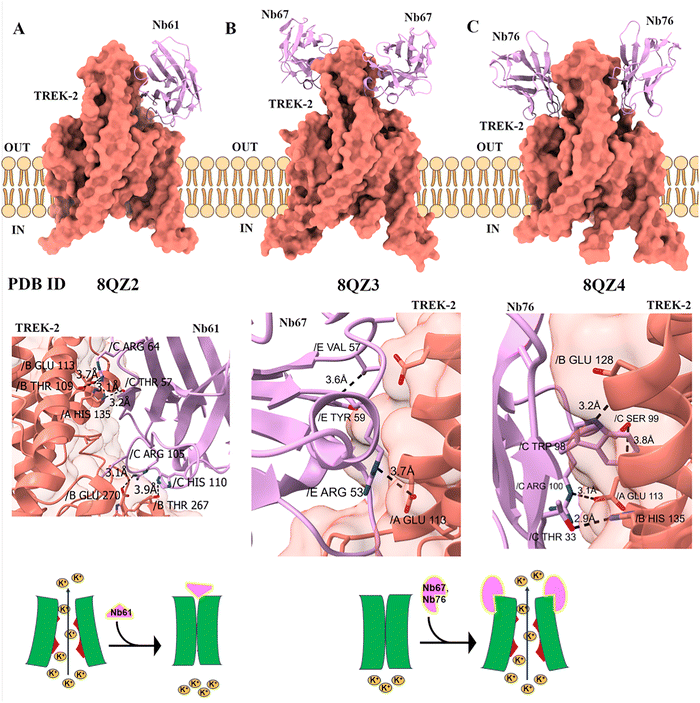
TREK-2 is a two-pore domain potassium (K2P) channel that plays a crucial role in stabilizing the resting membrane potential and regulating neuronal excitability. It responds to mechanical, thermal, and chemical stimuli, contributing to neuroprotection and pain modulation, especially under inflammatory or nerve injury conditions. Approaches using nanobodies have been developed to selectively modulate TREK-2 potassium channel activity by targeting its extracellular domains. Twenty-nine nanobodies were identified to bind TREK2, and three of these nanobodies demonstrated significant modulation of the channel properties of TREK2.Nb61 acts specifically as a negative allosteric modulator for TREK-2, rapidly suppressing TREK-2 currents (IC50 ≈ 685 nM). Notably, this inhibition is highly selective to mouse and human TREK-2, but does not affect related channels like TREK-1 or TASK-3. Structurally, the nanobody operates through a direct, toxin-like mechanism, where its extended CDR3 loop inserts a lysine residue, caps the extracellular ion exit pathway, and decreases potassium currents. However, because the K2P Cap domain of TREK2 restricts direct access to the pore, such a block is unusual, making this finding particularly significant (Fig. 2A). In contrast, Nanobody 67 (Nb67) binds to different extracellular sites and promotes channel opening, enhancing potassium efflux and reducing neuronal hyperexcitability, but also has a weaker affinity to TREK-1, making it partially selective (Fig. 2B). Nb76 binds specifically to the P2-M4 loop of the TREK-2 potassium channel, moves the M4 helix to the open “M4 Up” conformation, thereby allowing the potassium ions to pass. This structural rearrangement closely mirrors the mechanism by which well-established agonists like ML335 activate the channel. By promoting the “M4 Up” conformation, these nanobodies stabilize the selectivity filter in a configuration optimized for maximal potassium ion conductance (Fig. 2C).
Nb61-58 is a bi-paratopic nanobody that enhances inhibition of the TREK-2 potassium channel. It links two nanobody units, Nb-Inhibitor-61 and Nb-Binder-58. They bind distinct, non-overlapping sites on the TREK-2 Cap domain. This design significantly increases binding affinity (over 250-fold) compared to the monomeric inhibitor and improves inhibitory potency about 16-fold (IC50 ∼40 nM). These nanobodies exert their effects allosterically by influencing gating mechanisms linked to the selectivity filter and conformational changes between functional states. Through this precise modulation of TREK-2 activity, these nanobody tools offer valuable therapeutic potential for neuroprotection and pain management by dampening excessive neuronal excitability.17
ASIC1a channel
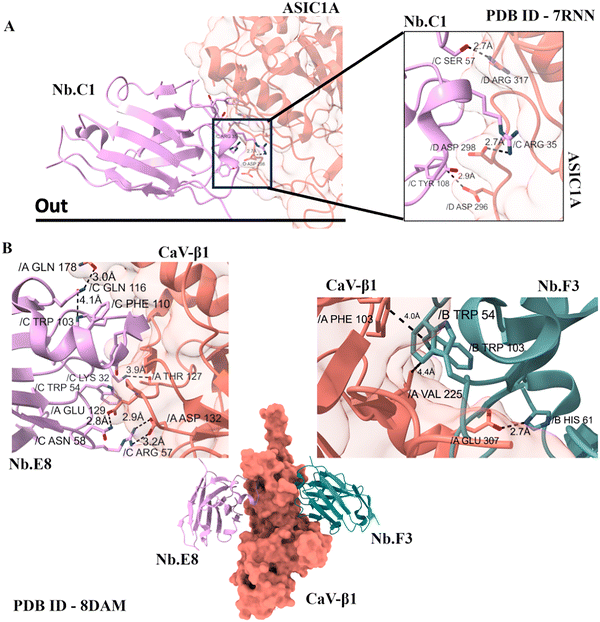
ASIC1a is a proton-gated sodium channel that is primarily involved in pain perception, particularly in conditions like ischemic injury, neuropathic pain, and other forms of nociception. The activation of ASIC1a can lead to the rapid influx of sodium ions into cells, contributing to the generation of pain signals.18 The hASIC1a channel is a trimer with each extracellular domain subunit depicting a palm, thumb, and finger-like architecture.The cryo-EM structure of human ASIC1a with nanobody Nb.C1 revealed that the nanobody binds to the extracellular thumb domain (D298 and L299), which is an important part of the channel that helps detect acidity and control channel opening or closing (Fig. 3A). However, Nb.C1 binding to ASIC1a channels did not affect channel dynamics (Fig. 3A). Interestingly, Nb.C1 attaches to a site on ASIC1a that partly overlaps with the binding sites of some snake toxins, such as MitTx1 and Mambalgin-1, which normally activate or modulate the channel. Thus, Nb.C1 binding to ASIC1a was demonstrated to mitigate the effects of MitTX1 on ASIC1a channels. Because of this, Nb.C1 can block toxins like MitTx1 from activating the channel by taking up their binding space. On its own, Nb.C1 does not strongly change the normal activity of ASIC1a. Interestingly, when Nb.C1 is linked with PcTx1, it acts like a “delivery tool.” It places PcTx1 in the perfect position to block ASIC1a more efficiently, making the inhibition stronger. It's important to note here that this synergy is possible as Nb.C1 does not overlap with the binding site of PcTx1which is another well-known ASIC inhibitor. Functionally, Nb.C1 has two roles: it acts as an antagonist against toxin-driven activation and as a potentiator when fused with inhibitors.19 By selectively binding to ASIC1a, nanobody Nb.C1 can fine-tune channel activity without completely shutting it down. Since ASIC1a is strongly linked to pain perception, inflammation, and neurodegeneration, its modulation using nanobodies offers a powerful therapeutic strategy.
Targeting intracellular auxiliary subunits: a strategy for HVACCs
High-voltage–activated calcium channels (HVACCs) are crucial in neurons and cardiac cells, regulating synaptic transmission and heart rhythm. Dysfunctions in HVACCs contribute to disorders like epilepsy, hypertension, and heart failure.Nb.F3 is a nanobody that does not differentiate between CaVβ isoforms and binds to all the isoforms, while Nb.E8 is specific to CaVβ2a and CaVβ1b. To address these, nanobodies such as nb.F3 have been engineered for isoform-specific inhibition of HVACCs. Nb.F3 interacts with CaVβ through complementing aromatic residues on CDR3 of the nanobody and the SH3 domain of CaVβ. Differences in Nb.E8 CDR3 regions might be essential in differentiating the CaVβ subunits (Fig. 3B). Nb.F3 acts as an allosteric inhibitor by binding the SH3 domain of the CaVβ subunit, reducing channel surface expression, lowering open probability, and accelerating inactivation, thereby suppressing CaVβ-associated currents. A novel approach fuses nb.F3 with ubiquitin ligases (NEDD4L) to promote targeted degradation of CaVβ-bound HVACCs. Nb.F3-NEDD4L inhibited endogenous CaV1/CaV2 channels in mammalian cardiomyocytes, dorsal root ganglion neurons, and pancreatic β cells. Unlike gene knockouts or shRNA, this post-translational method prevents compensatory CaVβ isoform reshuffling, allowing precise study of β-subunit roles. This approach is similar to the concept of PROTACs (proteolysis-targeting chimeras), which use bifunctional molecules to promote the degradation of specific proteins. By selectively targeting HVACCs for degradation, this nanobody-based strategy offers a novel way to treat conditions associated with these channels, such as chronic pain, epilepsy, and cardiovascular diseases.20,21
2. Direct gating modification and state stabilization
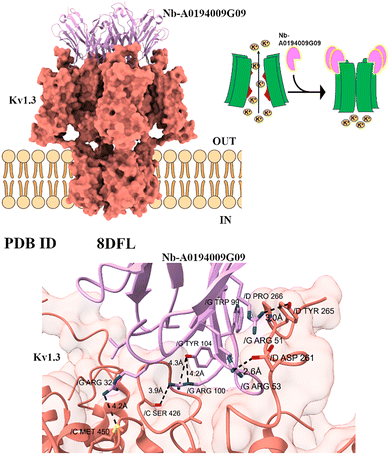
Some nanobodies act more directly on the channel's gating machinery, functioning as “gating modifiers” that lock the channel into a specific, often non-conducting, state.The Kv1.3 channel plays a critical role in the activation of T-cells, which are key components of the immune system. This channel is often overexpressed in various autoimmune diseases, contributing to inappropriate T-cell activation and the progression of conditions such as multiple sclerosis and rheumatoid arthritis.22
The A0194009G09 nanobody is a highly selective and potent modulator of the human Kv1.3 potassium channel. The cryo-EM structure revealed that four nanobody molecules bind to the tetramer's voltage-sensing domains (VSDs) and the adjacent pore domain (Fig. 3A). Unlike conventional channel blockers that physically obstruct the pore-forming EL3 loop, A0194009G09 binds to an allosteric site and inhibits the Kv1.3 channel activity.23 A0194009G09 specifically interacts with Kv1.3 residues Y417 and A421 – G427. This interaction triggers structural rearrangements in a critical part of the channel known as the selectivity filter, focusing on amino acid residues between Tyr436 and Asp449 (Fig. 4). These changes stabilize the channel in a C-type inactivated state, a conformation in which the channel remains closed and unable to conduct potassium ions despite not being physically occluding the channel pore.24 This specificity and prolonged inhibition make Kv1.3-targeting nanobodies a promising therapeutic tool for modulating the immune response and treating autoimmune diseases.
P2X7 ion channel
Similarly, nanobodies targeting the P2X7 receptor, an ATP-gated ion channel critical for inflammation, cell death, and immune responses, demonstrate both inhibitory and potentiating effects. This channel is known to mediate the release of pro-inflammatory cytokines, including IL-1β, which contribute to neuroinflammation, pain, and a variety of other inflammatory diseases.25 When P2X7 is activated, it forms a large pore in the membrane, allowing the passage of ions and other molecules that trigger further inflammatory responses.P2X7-blocking nanobodies offer precise and rapid modulation of the ATP-gated P2X7 ion channel. Nanobody 13A7 (IC50 – 12 nM) efficiently inhibits P2X7 channel activity by preventing the conformational changes triggered by ATP binding in mouse macrophages and T cells. In contrast, nanobody 14D5 (EC50 – 6 nM) enhances P2X7 channel function. These nanobodies act specifically on the P2X7-mediated inflammation pathway. An albumin-binding nanobody fused with 13A7 or 14D5 dimeric nanobodies increased their half-life and prolonged the effects of these nanobodies. Mouse models further verified the binding and modulation of 13A7 and 14D5. Dano1 (IC50 – 0.2 nM), a human-specific nanobody that surpasses traditional small-molecule antagonists by 1000-fold in inhibiting the release of IL-1β by monocytes in LPS and ATP-treated human blood. This superior potency arises from the ability of the nanobody to bind functional epitopes on P2X7 that conventional antibodies and drugs cannot reach.26 While detailed structural data for these nanobodies is still pending, their ability to modulate P2X7 function makes them invaluable tools in studying inflammation and immune responses. As a result, P2X7-targeting nanobodies represent a promising advancement in immunomodulation, with the potential to improve therapies for inflammatory and autoimmune diseases through their speed, selectivity, and enhanced efficacy.
3. Isoform, subunit, and heteromer-specific recognition
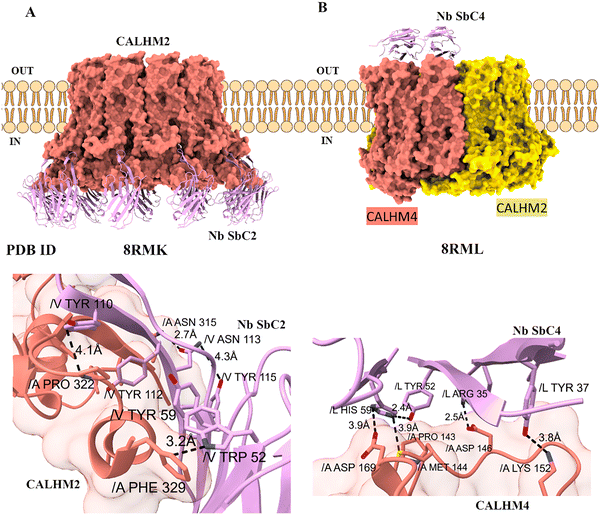
A key advantage of nanobodies is their ability to recognize unique epitopes with surgical precision, enabling the differentiation between closely related channel isoforms or specific subunit compositions within a larger complex.Targeting specific oligomeric assemblies: CALHM and NMDA receptors
While many nanobodies are functional modulators, others serve as high-fidelity detection tools. Nanobodies Nb17 and Nb82 were developed to bind the C-terminal domains of the skeletal muscle (NaV1.4) and cardiac (NaV1.5) voltage-gated sodium channels, respectively. They bind with nanomolar affinities (∼40–60 nM) and minimal cross-reactivity to other isoforms. Although they do not modulate channel gating or ion conductance, they serve as powerful molecular tools for isoform-specific detection, imaging, and biochemical isolation. The specificity and high affinity of Nb17 and Nb82 provide valuable reagents for dissecting the biology of NaV channels and hold promise for developing targeted diagnostics or therapeutic tools, particularly for skeletal and cardiac muscle disorders linked to NaV dysfunction.31
4. Nanobodies as structural chaperones and probes of Cys-loop receptors
The Cys-loop receptor superfamily of pentameric ligand-gated ion channels (pLGICs) has been a particularly fertile ground for nanobody development, where they have served as both functional modulators and indispensable tools for structural biology.Probing a Conserved Allosteric Vestibule: From ELIC to 5-HT3.
Cys-loop receptors
Cys-loop receptors constitute a family of pentameric ligand-gated ion channels, encompassing both cation-selective members such as serotonin 5-HT3 and nicotinic acetylcholine (nACh) receptors, the zinc-activated (ZAC) receptor, as well as anion-selective types including GABA and glycine receptors. The characteristic “Cys-loop” is a conserved disulfide-bonded loop in the extracellular domain essential for structural integrity and function. These receptors mediate fast synaptic transmission by opening their ion channels in response to neurotransmitter binding, allowing selective flow of ions across the membrane for excitatory or inhibitory signalling. Dysfunction in these receptors has been linked to a variety of neurological and psychiatric disorders, including epilepsy, schizophrenia, and neurodegenerative diseases. Therapeutic nanobodies against Cys-loop receptors may be developed to modulate neurological diseases associated with dysfunctional inhibitory or excitatory signaling.32 The prokaryotic channel ELIC serves as an excellent model for its human counterparts. Nanobodies targeting its large extracellular vestibule have revealed a conserved modulatory hotspot.These nanobodies with improved stability, blood–brain barrier penetration, and pharmacokinetics could be used to treat neurological disorders, including epilepsy, schizophrenia, chronic pain, and addiction, offering receptor-specific therapies with high precision and minimal off-target effects.
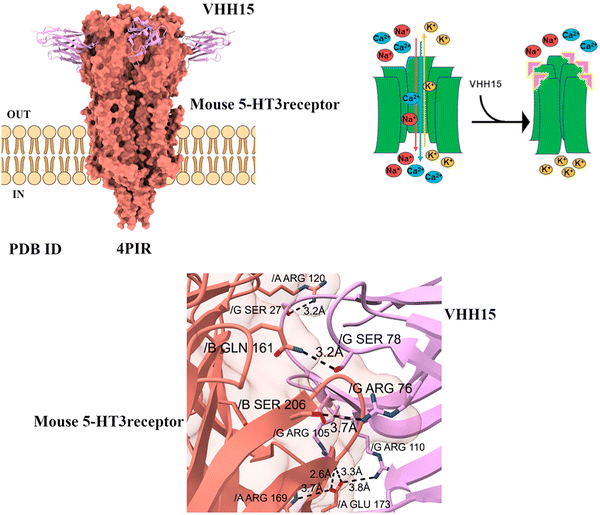
As in ELIC, the vestibule site where nanobody binds is conserved. In the mouse serotonin 5-HT3 receptor, VHH15 binds to the equivalent site. Functional analysis demonstrates that this binding inhibits the channel activation by serotonin. This indicates that the receptor adopts a conformation corresponding to an inhibited, non-conducting state, preventing ion flow through the channel despite the presence of the neurotransmitter. Nanobody binding to the 5-HT3 receptor can influence neurotransmitter recognition by stabilizing the extracellular ligand-binding domain, particularly the flexible loops that form the orthosteric binding site. By capping or interacting with regions such as the C-loop and adjacent structural elements, nanobodies restrict the conformational dynamics necessary for efficient serotonin binding and receptor activation. This stabilization may either lock the receptor in a resting, inactive state or modulate its transition states, thereby altering the affinity and efficacy of neurotransmitter binding (Fig. 8).36 Numerous functional modulator nanobodies (activating and inhibiting) have been generated against 5-HT3AR and m5-HT3AB receptors. These nanobodies were found to have a strong affinity for the receptors, with Kd values ranging from low nanomolar to submicromolar concentrations.32
Diverse mechanisms in human pLGICs: nAChR and GABAARs
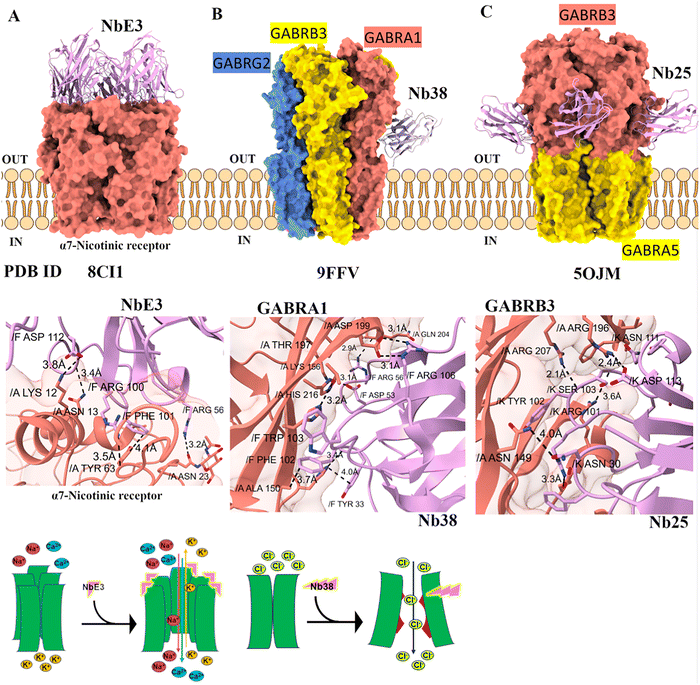
Nanobody38 (Nb38) that specifically binds to the α1 subunit of the GABA_A receptor. Structural and functional studies show that Nb38 binds at the extracellular domain interfaces involving the α1 principal face and the complementary faces of neighbouring β3 and γ2 subunits. This binding stabilizes the receptor in an activated conformation, as evidenced by an approximately 6.5-fold increase in Nb38 binding affinity in the presence of GABA. Functionally, Nb38 strongly potentiates GABA-induced currents, increasing receptor activity by up to 480% in α1β3γ2 engineered receptors and about 290% in wild-type receptors. This potentiation is selective for receptors containing the α1 subunit and is more effective than the benzodiazepine diazepam in amplifying GABA responses. Additionally, Nb38 increases the amplitude and prolongs the decay time of spontaneous inhibitory postsynaptic currents (sIPSCs) in neurons, thereby enhancing inhibitory neurotransmission. The gating mechanism involves Nb38 stabilizing the receptor's activated state without acting as a direct agonist itself. By binding near the ligand-binding site interfaces, Nb38 likely promotes conformational changes that favor channel opening and prolonged activity, enhancing the inhibitory function of GABA_A receptors containing the α1 subunit. Nb38 interacts with the GABA_A receptor via aromatic residues on opposite faces of the binding interface, leading to π–π stacking interactions. Hydrogen bonds between a few polar residues further stabilize the interaction (Fig. 9B).41 Nb25 is a nanobody that specifically binds to the β3 extracellular domain (ECD) of the GABA_A receptor. Structural studies using cryo-EM revealed that three Nb25 molecules bind between adjacent ECD interfaces of the α5β3 GABA_A receptor, particularly at interfaces involving β3 subunits. This binding helps to identify and distinguish β3 subunits in receptor assemblies, which is especially useful because α5 and β3 subunits have similar sequences and structures that make them hard to differentiate at lower resolution (Fig. 9C). Functionally, Nb25 binds at the interfaces between β subunits, and although it binds tightly, it does not show significant potentiation effects like Nb38. Nb25 has been used as a tool in structural studies to facilitate receptor crystallization and improve classification of receptor particles in cryo-EM analysis.42
Contrary to the interactions observed in sybody Sb1, the interaction interface of sybodies Sb2 and Sb3 contains fewer aromatic residues and more hydrophilic residues. All three nanobodies inhibit the LRRC8A by restricting the LRR domain (Sb1) or inducing conformational changes in the LRR domain and pore region (Sb2 and Sb3). Sybodies Sb4 and Sb5 potentiate channel activity by binding to alternate subunits of the LRR domain and are buried in the dimer interfaces of the domains, specifically engaging repeats 2–14 (Sb4) and 2–7 (Sb5). These sites are accessible only in specific subunit positions due to the arrangement of subunits within the channel. Functionally, Sb4 and Sb5 serve as allosteric activators, increasing VRAC channel activity by enhancing LRR domain mobility and triggering conformational changes that likely promote channel opening.44 Beyond mechanistic importance, these findings link LRRC8A modulation to physiological roles such as insulin secretion, glucose control, tumor progression, and ischemic brain injury, highlighting sybody–channel complexes as a promising therapeutic target.
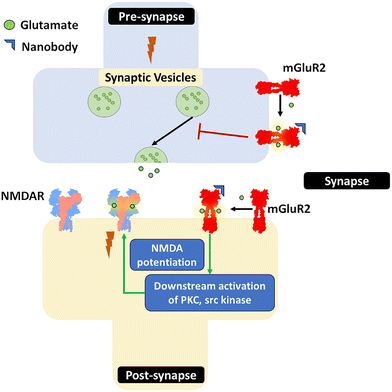
5. Indirect modulation of ion channels via GPCRs: the mGluR2/NMDAR axis
Nanobodies can also modulate ion channels indirectly by targeting associated signaling proteins. Metabotropic glutamate receptors (mGluRs), which are G protein-coupled receptors (GPCRs), form a well-known functional complex with NMDARs. They are known to play crucial roles in modulating excitatory neurotransmission in the brain. Nanobodies DN10 and DN13 were developed as selective positive allosteric modulators of the mGluR2 receptor.These nanobodies bind to the intracellular domain of the receptor, close to the G protein–coupling interface, where they stabilize specific conformations of the receptor. DN13 binds specifically to agonist-bound mGlu2 homomers and enhances their activity. DN1 nanobody demonstrated binding to both homomer and heteromer conformations of mGlu2, without affecting receptor function.45 A DN1–DN13 biparatopic nanobody enhanced the affinity to mGlu2. The DN1–DN13 nanobody improved working memory in GluN1 knockdown mice models.46 By fine-tuning the conformational states of mGluR2, these nanobodies provide a powerful means to probe receptor function, dissect mGluR2–NMDA receptor cross-talk, and inspire novel therapeutic strategies for neurological disorders such as schizophrenia, anxiety, and excitotoxic neurodegeneration. This highlights a sophisticated strategy where a nanobody targeting a GPCR can precisely regulate the function of a downstream ion channel (Fig. 10).
Breaking down the fortress: delivering nanobody across the blood–brain barrier
The blood–brain barrier (BBB) is a selective permeability barrier that protects the brain from harmful substances while regulating the passage of essential nutrients and molecules. However, this selective barrier also presents a significant challenge in delivering therapeutics to the central nervous system (CNS), particularly for large molecules such as antibodies. Nanobodies, with their small size and high stability, offer a promising solution to this challenge, as they can more easily cross the BBB compared to larger antibody constructs.47Delivery of therapeutic or diagnostic nanobodies can be achieved via several mechanisms (Fig. 11). BBB can be breached by several methods, including passive diffusion, Carrier-mediated Transport (CMT), Receptor-mediated transcytosis (RMT), and Adsorptive-mediated transcytosis (AMT).48 It is important to measure how well BBB transport works in order to figure out how useful it might be for treatment. Although data is still limited, a few important studies offer useful benchmarks.
Passive diffusion of any material prerequisites a small size (<400 Da), fat solubility, a less polar surface, and fewer hydrogen bonds (<6) forming residues. 89Zr-nanobody shows a very low level of non-specific, passive diffusion into the brain in rats, approximately ∼0.02% injected dose per gram (% ID per g) after 1 hour. PET imaging studies confirm that without BBB disruption, brain uptake remains minimal, supporting the use of this baseline for assessing targeted delivery strategies in brain imaging and therapy.49
Nbs can traverse the BBB through RMT, a process in which molecules bind to specific receptors on the endothelial cells of the BBB, such as the transferrin receptor or insulin-like growth factor receptors. These receptors facilitate the transport of nanobodies across the BBB, allowing them to reach their target in the CNS. This ability to target the brain with high specificity opens new avenues for the treatment of neurological diseases, including Alzheimer's disease, Parkinson's disease, and glioblastoma. RMT is currently the most widely used method for the delivery of nanobodies through the BBB. FC5 and FC44 are two sdAbs that enter the BBB through RMT.50,51 FC5 specifically binds to the luminal α(2,3)-sialoglycoprotein receptor TMEM30A on brain endothelial cells, an interaction that initiates clathrin-mediated endocytosis of FC5 and its subsequent RMT across the BBB. In vitro studies using Brain Microvascular Endothelial Cells (BMEC) monolayers show that FC5 achieves approximately 4.0 ± 0.56% translocation at 6 hours and 42.0 ± 3.35% at 24 hours, significantly outperforming unrelated single-domain or control antibodies in both specificity and translocation rate. In vivo, FC5 brain uptake ranges from 0.1% to 1% of the injected dose per gram of tissue (% ID per g), with enhanced versions—such as dimerized or Fc-fused constructs—achieving up to ∼0.8% ID per g to 2% ID per g, depending on the experimental model and configuration, reflecting up to a 20-fold increase in delivery efficiency.52 Nbs developed against the extracellular domain of the human insulin-like growth factor 1 receptor (IGF1R), when fused to either mouse or human Fc regions, are capable of crossing the BBB via receptor-mediated transcytosis. These constructs not only achieve brain uptake levels ranging from 5 to 11% ID per g in rats, but have also been shown to deliver non-BBB-penetrant therapeutic small molecules into the CNS.53,54
The transferrin receptor is also a target for RMT.55 Nanobodies targeting the transferrin receptor (TfR) have been employed to deliver neurotensin (NT) into the CNS. One notable construct is the engineered M1 nanobody, particularly its triple histidine mutant M1(His). When combined with other nanobody domains—such as 13A7–13A7 (targeting P2X7 receptor) and Nb80 (binding to albumin)—this multi-domain fusion efficiently crosses the BBB after intravenous administration. Quantitatively, brain uptake reached 13.4% ID per g at 1 nmol kg−1 and 8.8% at 3 nmol kg−1, while a higher dose of 30 nmol kg−1 resulted in 3.5% ID per g, indicating a dose-dependent saturation effect. Incorporating an albumin-binding nanobody extended systemic half-life from 21 minutes to 2.6 hours, significantly enhancing delivery.56–58 Nb targeting a cytosolic prion protein PrPc was capable of crossing the BBB in vitro and in vivo, and it was hypothesized to transmigrate via RMT.59 Bi-specific Nanobodies conjugated with nanobodies targeting RMT fused with human FC region could provide a multipronged approach to overcome the limitations in pharmacokinetics (faster renal clearance of sdAbs) and crossing the BBB. VHH-E9-SCFv8D3 is a bispecific nanobody that uses RMT (scFv8D3 targets transferrin) to deliver a nanobody (VHH-E9) against glial fibrillary acidic protein (GFAP) overexpressed by astroglial cells in regions affected by Alzheimer's disease. VHH-E9 nanobody reached brain concentrations of 0.15% ID per g, while VHH-E9-SCFv8D3 attained 0.44% ID per g 2 hours post-injection.60 Glycosylated nanobodies (Glucoside-Nb) have shown much better ability to cross the BBB and accumulate in glioblastoma tumors in mice. Using RMT, they delivered about 2.9 times more to the brain compared to unmodified nanobodies. In a mouse model of glioma, brain delivery increased by 5.7 times, as confirmed by fluorescence imaging 24 hours after injection. This enhancement is attributed to glycosylation-mediated engagement of the GLUT1 transporter, which is highly expressed on the BBB and glioblastoma cells and is crucial for glucose uptake and tumor metabolism. Quantitatively, glycosylated nanobodies achieve up to about 6% ID per g of brain tissue (ID per g-brain), outperforming unmodified analogues, which typically show less than 2% ID per g-brain.61
The transmigration of Nb's through the BBB by AMT has been demonstrated in a few cases where the pI of the Nb's is high. For instance, Nb R3QV (pI > 8.3) targets Aβ42, Nb A2 (pI > 9.5) targets phosphorylated Tau protein,62 and an Nb that targets GFAP (pI = 9.4).63–65 Alternatively, Nbs have been delivered into the CNS using agents that can transmigrate through the BBB, such as glutathione-targeted PEGylated liposomes, multi-walled carbon nanotubes (MWCNTs), or Fe3O4 nanoparticles.66–69
The study of the bivalent mGlu2-targeting nanobody, DN13-DN1, provides the most complete data. After a single intraperitoneal injection in mice, radiolabeling studies showed that brain uptake peaked at 4 hours, with 0.08% of the injected dose (%ID) was found in total brain extracts. Although this level of penetration appears low, it was sufficient to achieve a therapeutically relevant brain concentration of approximately 9.2 nM, which is significantly higher than the nanobody's binding affinity for its target. Importantly, the nanobody was detectable in the brain for a minimum of 7 days (at approximately 1.5 nM), a duration that directly corresponded with enduring behavioral enhancements in mouse models of schizophrenia. The brain uptake of 0.08% ID is similar in quantity to the 0.01–0.4% reported for the passive diffusion of conventional IgG antibodies.46 However, the pharmacological implications are very different. IgGs have a long serum half-life (days to weeks), while unmodified nanobodies are cleared from circulation within 1 hour. The fact that DN13-DN1 can get IgG-like brain exposure even though it has a short peripheral half-life means that a much more efficient transport mechanism is at work. For context, in the field of PET imaging, a brain uptake of more than 0.1% of the injected dose per cubic centimeter (% ID per cc) is often seen as a benchmark for a successful CNS tracer, which means that the levels achieved by DN13-DN1 are within a functionally significant range.60
The development of these innovative strategies for crossing the BBB holds great promise for advancing nanobody-based treatments for a wide range of CNS disorders.
From bench to bedside: the clinical progression of ion channel-targeting nanobodies
Preclinical studies have shown that nanobodies have a lot of potential, but a close look at their clinical development shows that there is a big gap between proving the technology platform and using it on a specific type of ion channel.The nanobody platform has been very successful in both clinical and business settings. So far, four nanobody-based drugs have been approved by major agencies like the FDA and EMA. These include Caplacizumab (Cablivi®), a bivalent nanobody targeting von Willebrand factor for acquired thrombotic thrombocytopenic purpura;70 Ozoralizumab (Nanozora®), a trivalent, bispecific nanobody targeting TNFalpha for rheumatoid arthritis (approved in Japan);71–73 Envafolimab, a PD-L1-targeting nanobody fused to an Fc domain for solid tumors (approved in China);74,75 and Ciltacabtagene autoleucel (Carvykti®), a CAR-T cell therapy utilizing two B-cell maturation antigen-targeting nanobodies for multiple myeloma.76 A thorough examination of clinical trial databases uncovers a substantial array of nanobody candidates, featuring numerous active trials, predominantly in oncology and inflammatory diseases.77 Nonetheless, a notable finding arises from this analysis: None of the approved nanobody drugs or candidates in late-stage clinical development targets an ion channel. The antigens that have been successfully targeted are primarily soluble proteins (vWF, TNFα) or highly expressed cell surface markers on immune or cancer cells (PD-L1, BCMA). This disparity highlights the unique difficulties associated with ion channel targets. Monoclonal antibodies targeting ion channels such as P2X7 and Orai-1 have commenced clinical trials; however, this advancement is yet to be mirrored in the nanobody domain.
Nanobodies are versatile, small and modular for targeting proteins nevertheless, there are a few limitations to the use of nanobodies to study ion channels. Nanobodies are generated using proteins expressed in a heterologous system. The lack of pivotal regulatory and interacting partners of the receptor in the heterologous system may hinder the efficiency of the nanobody in the native or physiological conditions of the ion channel, since the receptors are known to exist in different conformational states in the presence of auxiliary proteins and other interacting proteins. The ability of a nanobody to trap receptors in specific conformational states might be useful in certain aspects, while it may not be useful to study aspects pertaining to receptor modulation.
Problems that make ion channel targets hard to translate to clinical use
The notable lack of ion channel-targeting nanobodies in the clinical pipeline indicates a substantial translational gap, stemming from the intrinsic biochemical difficulties associated with this category of membrane proteins.First and foremost is the “antigen challenge”. The generation of high-affinity nanobodies typically begins with the immunization of a camelid, which requires substantial quantities of pure, stable, and conformationally correct antigen. Ion channels are notoriously difficult to produce in this manner. They are frequently expressed at low levels in native tissues, and recombinant overexpression may result in misfolding and aggregation.78 Various detergents have been used traditionally to purify membrane proteins. The choice of detergents depends on the membrane protein and might have effect the stability and functionality of membrane proteins. In order to provide a more conducive environment, detergent-purified membrane proteins can be further stabilized by incorporating amphipols, nanodiscs or reconstituting the receptors in liposomes. A recent report comparing nanodiscs and liposome-based purification of ELIC receptor identified that liposomes provided a more native environment to the protein, thereby reducing artefactual conformations as observed in nanodiscs. Such strategies may be adopted for the purification and/or stabilization of challenging membrane protein targets.79
Second, the process of screening is much more complicated. High-throughput ELISA-based tests can help find binders for soluble targets like TNFα. To find functional modulators of an ion channel, on the other hand, you need specialized, low-throughput, and technically challenging tests, like patch-clamp electrophysiology, to see how the channel current changes. This makes the process much more complicated, meaning fewer candidates can be effectively screened, making discovery campaigns take longer and cost more. Finally, the high structural similarity among ion channel subtypes means that very specific nanobodies need to be made to avoid possibly harmful off-target effects. This is a high standard for safety and regulatory approval. The main challenge that must be overcome to move from preclinical promise to clinical reality for ion channel-targeting nanobodies is solving these fundamental problems in antigen production and functional screening. The development of computational tools to identify specific nanobodies against proteins is gaining traction in the field, and the following are some of the AI and ML methods used for this purpose (Table 3).
| Contribution area | Exemplary tools/models | Model type/approach | Key applications in nanobody development |
|---|---|---|---|
| Structure prediction | AlphaFold2, AlphaFold3, IgFold, NanoNet, ImmuneBuilder, RoseTTAFold, NanoBodyBuilder2 | Transformer, ResNet, GNN, AntiBERTy | Accurate modeling of nanobody (VHH) 3D structures, with focus on variable CDR loops (esp. CDR3/H3) |
| Sequence & epitope design | ProteinMPNN, RFdiffusion, JAM, ParaAntiProt, AlphaProteo, BindCraft, Evolvex | Generative AI, transformer, CNN | De novo design of nanobody sequences for specific antigens, identification of strong binding sites. |
| Affinity & developability | DeepAb, ML classifiers, NanoBERTa-ASP & NbX | RNN, ResNet, decision Tree | Predict binding affinity and filter high-affinity candidates; assess developability, solubility, and stability |
| Humanization & immunogenicity | AbNatiV (deep learning) + ML pipelines | Deep learning, ML classifiers | Predict and minimize immunogenicity, improve resemblance to human antibodies |
| High-throughput screening | ML-based interaction models | ML classifiers, CNN | Screening vast protein–antibody interaction datasets, sifting for leads rapidly |
| Generative design | Generative AI platforms, RFdiffusion | Generative AI, transformer | Create diverse in silico nanobody variants targeting multiple or “undruggable” sites |
| Paratope/epitope prediction | Parapred, ParaAntiProt | RNN (LSTM), CNN, transformer | Paratope and epitope prediction to identify binding interfaces |
The digital Frontier: using AI to design and improve ion channel nanobodies
Integrating artificial intelligence (AI) and machine learning (ML) is transforming the traditional, hands-on, and time-consuming process of discovering nanobodies. These computational methods allow for fast and accurate identification, design, and optimization of nanobodies by utilizing advanced deep learning architectures, structural modeling, and large-scale data analysis. This is especially promising for addressing the challenges associated with hard targets, such as ion channels, because of the structural complexity, low expression yields, and the subtle differences among channel subtypes.The emergence of highly precise protein structure prediction tools, particularly AlphaFold, has revolutionized structural biology.80 Advanced structure prediction models like AlphaFold3 have significantly improved nanobody-antigen complex modelling, achieving high-accuracy docking success rates of 8.9% for antibodies and 13.4% for nanobodies.81 While exact binding epitope prediction remains challenging with accuracy below 50%, confidence scores such as ipTM (interface predicted TM-score) reliably distinguish near-native from incorrect models. The prediction quality heavily depends on the nanobody CDR3 loop characteristics, with simple, stretched conformations and shorter loops being modeled more accurately than kinked and longer loops due to their inherent flexibility. Despite these limits, such tools provide a valuable starting point for structure-guided design and candidate selection.82 Complementary deep learning models like IgFold and NanoNet enable precise 3D structure prediction of nanobodies directly from sequences, with IgFold achieving a median CDR3 RMSD accuracy of 2.2 Å for nanobodies and operating at remarkable speed (under 25 seconds). These tools have made large-scale structural analysis feasible, with researchers successfully predicting structures for over 1.4 million paired antibody sequences.83,84
Affinity maturation of nanobodies is an important aspect after discovery implies improving their binding strength to specific target antigens through repeated cycles of design and refinement of their sequences or structures. While this was traditionally done using random mutagenesis followed by experimental screening, recent advances in computational approaches have streamlined and expedited the affinity maturation process.85 Computational affinity maturation platforms like ADAPT and EvolveX combine physics-based models with machine learning to screen millions of virtual mutants, achieving dramatic improvements and selecting top candidates. These approaches have achieved dramatic changes—for example, an anti-CD47 nanobody was computationally optimized using homology modeling combined with ADAPT, resulting in mutants with up to an 87-fold increase in binding affinity and improved thermostability.86 Even more dramatically, another pipeline improved the affinity of a VHH against the immune checkpoint protein Vsig4 by over 1000-fold, transforming a weak binder into a potent nanomolar-affinity molecule.87
Machine learning models for binding prediction have achieved remarkable performance in nanobody-antigen interaction forecasting, with advanced algorithms reaching prediction accuracies exceeding 80%.88 NABP-BERT, a specialized language model developed by Google, represents a breakthrough in sequence-based binding prediction, achieving very high scores in the benchmarks while focusing on surrounding amino acid contexts.89,90 These models leverage extensive datasets of nanobody-protein binding interactions and utilize sophisticated architectures to predict binding purely from sequence data.91,92 Machine learning analyzes large datasets of nanobody–antigen interactions and identifies patterns in sequence, structure, and noncovalent interactions (e.g., hydrogen bonds, aromatic stacking) that govern binding affinity. Feature analysis reveals that hydrogen bonding and aromatic-associated interactions are key mechanistic determinants of binding affinity, with hydrogen bond descriptors accounting for 51.9% of important predictive features and aromatic interactions contributing 29.6%. This enables prediction of new epitopes and the nanobody features critical for strong, specific binding.88
Generative models, including AlphaProteo, BindCraft, and RFdiffusion, further expand capabilities by designing novel sequences targeting challenging epitopes, with BindCraft leveraging AlphaFold2's trained weights to generate nanomolar binders without requiring high-throughput screening.16,93 These comprehensive AI-driven pipelines integrate reinforcement learning, ensemble meta-predictors, and in silico mutagenesis to refine nanobody candidates by predicting how sequence changes affect binding affinity, stability, and developability, thereby accelerating discovery cycles while enhancing success rates against previously difficult targets.94
AI and machine learning continue to reshape the future of nanobody discovery for ion channels by providing powerful capabilities such as structure-guided virtual design, rapid and accurate epitope mapping, and data-driven mutational optimization, where the generation of large amounts of antigen is challenging. These technologies integrate seamlessly with experimental workflows to establish a robust, cutting-edge discovery pipeline tailored for the unique challenges posed by ion channel targets. By combining precise structural modeling with intelligent library selection and functional impact prediction, AI and ML streamline the identification and optimization of high-affinity, specific nanobodies.95–97 This convergence not only accelerates development timelines but also enhances the precision and efficacy of nanobody candidates, paving the way for innovative therapeutics and diagnostics targeting ion channels and other complex membrane proteins.
Future outlook
The field of ion channel-targeted nanobodies stands at a fascinating inflection point. The preclinical evidence is strong, showing that these biologics can change how channels work with unmatched precision and through new allosteric mechanisms. However, the way to use this in the clinic is still mostly unexplored. The future of this field will likely be defined by progress in three key areas: tackling new targets, innovating delivery strategies, and integrating intelligent design. In addition, the need for developing strategies for the administration of nanobodies, increasing the half-life of nanobodies, and generating a large repertoire of nanobodies against multiple ion channels to target very specific diseased states are probably the challenges that lie ahead in this field.A significant gap in the current research landscape is the lack of nanobodies that target ionotropic glutamate receptors (iGluRs), including AMPA, Kainate, NMDA, and Delta receptors.98 This deficiency is particularly notable given their critical role in central nervous system (CNS) physiology and pathology, alongside the recent achievement in creating a brain-penetrating nanobody for their G protein-coupled counterparts, the metabotropic glutamate receptors (mGluRs). The difficulties in targeting iGluRs stem from their considerable structural complexity. Unlike the dimeric mGluRs, iGluRs are large, heavily glycosylated heterotetramers with a complex “dimer-of-dimers” architecture in their extracellular domains.98 This complexity makes them exceptionally difficult to produce as stable antigens for immunization and complicates the identification of functional epitopes. Interestingly, the lack of nanobodies against the structurally simpler Glycine receptor has been attributed to the fact that its structure could be readily solved by other means, suggesting the field has prioritized targets where nanobodies provide a unique enabling advantage. As cryo-EM continues to unravel iGluR structures, the next frontier will be to move beyond structural chaperones and develop functional nanobody modulators, which remain a major untapped therapeutic opportunity.
Author contributions
The manuscript was conceptualized, designed, and written by MS, SB, SAM, IM and supervised by JK.Conflicts of interest
No conflict of interestData availability
No primary research results, software or code have been included and no new data were generated or analysed as part of this review.Acknowledgements
The research program in our laboratory is supported by the DBT-Wellcome Trust India Alliance (IA/S/21/2/505937). Dr Kumar is a Senior Fellow of the DBT-Wellcome Trust India Alliance. This work was supported by the CSIR-FIRST grant (FIR070303) awarded to J. Kumar. MS thanks ANRF, India, for the National Post-Doctoral Fellowship (PDF/2023/000784). SB thanks the intramural funding from CSIR-CCMB. SAM thanks CSIR-FIRST for the fellowship. Molecular graphics and analyses performed with UCSF ChimeraX, developed by the Resource for Biocomputing, Visualization, and Informatics at the University of California, San Francisco, with support from National Institutes of Health R01-GM129325 and the Office of Cyber Infrastructure and Computational Biology, National Institute of Allergy and Infectious Diseases.99References
- M. J. Ackerman and D. E. Clapham, N. Engl. J. Med., 1997, 336, 1575–1586 CrossRef CAS.
- F. Bezanilla, Neuron, 2008, 60, 456–468 CrossRef CAS PubMed.
- C. A. Hübner and T. J. Jentsch, Hum. Mol. Genet., 2002, 11, 2435–2445 CrossRef.
- K. Bohmwald, N. M. S. Gálvez, C. A. Andrade, V. P. Mora, J. T. Muñoz, P. A. González, C. A. Riedel and A. M. Kalergis, Front. Physiol., 2021, 12, 736681 CrossRef PubMed.
- W. Wu, J. Zheng, R. Wang and Y. Wang, Biochem. Pharmacol., 2023, 218, 115863 CrossRef CAS PubMed.
- L. He, M. Ahmad and N. Perrimon, Exp. Cell Res., 2019, 374, 259–265 CrossRef CAS.
- T. Kamei, T. Kudo, H. Yamane, F. Ishibashi, Y. Takada, S. Honda, Y. Maezawa, K. Ikeda and Y. Oyamada, Biochem. Biophys. Res. Commun., 2024, 721, 150126 CrossRef CAS PubMed.
- E. Y. Yang and K. Shah, Front. Oncol., 2020, 10, 1182 CrossRef.
- C. Hamers-Casterman, T. Atarhouch, S. Muyldermans, G. Robinson, C. Hamers, E. B. Songa, N. Bendahman and R. Hamers, Nature, 1993, 363, 446–448 CrossRef CAS PubMed.
- S. S. Rizk, D. M. Moustafa, S. A. ElBanna, H. T. Nour El-Din and A. S. Attia, World J. Microbiol. Biotechnol., 2024, 40, 209 CrossRef PubMed.
- S. Muyldermans, Annu. Rev. Biochem., 2013, 82, 775–797 CrossRef CAS PubMed.
- J. Yong Joon Kim, Z. Sang, Y. Xiang, Z. Shen and Y. Shi, Adv. Drug Delivery Rev., 2023, 195, 114726 CrossRef CAS.
- S. Muyldermans, FEBS J., 2021, 288, 2084–2102 CrossRef CAS.
- C. McMahon, A. S. Baier, R. Pascolutti, M. Wegrecki, S. Zheng, J. X. Ong, S. C. Erlandson, D. Hilger, S. G. F. Rasmussen, A. M. Ring, A. Manglik and A. C. Kruse, Nat. Struct. Mol. Biol., 2018, 25, 289–296 CrossRef CAS.
- Q. Su, W. Shi, X. Huang, S. Yin, X. Yang and X. Lu, MedComm: Biomater. Appl., 2023, 2, e54 Search PubMed.
- J. L. Watson, D. Juergens, N. R. Bennett, B. L. Trippe, J. Yim, H. E. Eisenach, W. Ahern, A. J. Borst, R. J. Ragotte, L. F. Milles, B. I. M. Wicky, N. Hanikel, S. J. Pellock, A. Courbet, W. Sheffler, J. Wang, P. Venkatesh, I. Sappington, S. V. Torres, A. Lauko, V. De Bortoli, E. Mathieu, S. Ovchinnikov, R. Barzilay, T. S. Jaakkola, F. DiMaio, M. Baek and D. Baker, Nature, 2023, 620, 1089–1100 CrossRef CAS PubMed.
- K. E. J. Rödström, A. Cloake, J. Sörmann, A. Baronina, K. H. M. Smith, A. C. W. Pike, J. Ang, P. Proks, M. Schewe, I. Holland-Kaye, S. R. Bushell, J. Elliott, E. Pardon, T. Baukrowitz, R. J. Owens, S. Newstead, J. Steyaert, E. P. Carpenter and S. J. Tucker, Nat. Commun., 2024, 15, 4173 CrossRef PubMed.
- S. Gründer and X. Chen, Int. J. Physiol., Pathophysiol. Pharmacol., 2010, 2, 73–94 Search PubMed.
- Y. Wu, Z. Chen, F. J. Sigworth and C. M. Canessa, eLife, 2021, 10, e67115 CrossRef CAS PubMed.
- T. J. Morgenstern, J. Park, Q. R. Fan and H. M. Colecraft, eLife, 2019, 8, e49253 CrossRef PubMed.
- T. J. Morgenstern, N. Nirwan, E. O. Hernández-Ochoa, H. Bibollet, P. Choudhury, Y. D. Laloudakis, M. Ben Johny, R. A. Bannister, M. F. Schneider, D. L. Minor and H. M. Colecraft, Nat. Commun., 2022, 13, 7556 CrossRef CAS.
- S. Cheng, D. Jiang, X. Lan, K. Liu and C. Fan, Biomed. Pharmacother., 2024, 175, 116651 CrossRef CAS.
- US11149086B2 - KV1.3 binding immunoglobulins - Google Patents, https://patents.google.com/patent/US11149086B2/en? q = (ion + channels + AND + nanobodies + AND + current)&oq = +ion + channels + AND + nanobodies + AND + current, (accessed December 9, 2024).
- P. Selvakumar, A. I. Fernández-Mariño, N. Khanra, C. He, A. J. Paquette, B. Wang, R. Huang, V. V. Smider, W. J. Rice, K. J. Swartz and J. R. Meyerson, Nat. Commun., 2022, 13, 3854 CrossRef CAS PubMed.
- M. L. Cotrina and M. Nedergaard, Purinergic Signalling, 2009, 5, 223–232 CrossRef CAS.
- W. Danquah, C. Meyer-Schwesinger, B. Rissiek, C. Pinto, A. Serracant-Prat, M. Amadi, D. Iacenda, J.-H. Knop, A. Hammel, P. Bergmann, N. Schwarz, J. Assunção, W. Rotthier, F. Haag, E. Tolosa, P. Bannas, E. Boué-Grabot, T. Magnus, T. Laeremans, C. Stortelers and F. Koch-Nolte, Sci. Transl. Med., 2016, 8, 366ra162 Search PubMed.
- H. K. Ganguly, M. B. Elbaum and N. J. Zondlo, Biochemistry, 2025, 64, 2848–2866 CrossRef CAS.
- K. Drożdżyk, M. Peter and R. Dutzler, eLife, 2024, 13, RP96138 CrossRef.
- T.-H. Chou, H. Kang, N. Simorowski, S. F. Traynelis and H. Furukawa, Mol. Cell, 2022, 82, 4548–4563.e4 CrossRef CAS.
- G. Loussouarn, D. Sternberg, S. Nicole, C. Marionneau, F. Le Bouffant, G. Toumaniantz, J. Barc, O. A. Malak, V. Fressart, Y. Péréon, I. Baró and F. Charpentier, Front. Pharmacol., 2016, 6, 314 Search PubMed.
- L. Srinivasan, V. Alzogaray, D. Selvakumar, S. Nathan, J. B. Yoder, K. M. Wright, S. Klinke, J. N. Nwafor, M. S. Labanda, F. A. Goldbaum, A. Schön, E. Freire, G. F. Tomaselli, L. M. Amzel, M. Ben-Johny and S. B. Gabelli, J. Biol. Chem., 2022, 298, 101763 CrossRef CAS PubMed.
- World Intellectual Property Organization, WO2015189162A1, 2015.
- M. Brams, C. Govaerts, K. Kambara, K. L. Price, R. Spurny, A. Gharpure, E. Pardon, G. L. Evans, D. Bertrand, S. C. Lummis, R. E. Hibbs, J. Steyaert and C. Ulens, eLife, 2020, 9, e51511 CrossRef CAS.
- C. M. Hénault, C. Govaerts, R. Spurny, M. Brams, A. Estrada-Mondragon, J. Lynch, D. Bertrand, E. Pardon, G. L. Evans, K. Woods, B. W. Elberson, L. G. Cuello, G. Brannigan, H. Nury, J. Steyaert, J. E. Baenziger and C. Ulens, Nat. Chem. Biol., 2019, 15, 1156–1164 CrossRef PubMed.
- N. M. Barnes, T. G. Hales, S. C. R. Lummis and J. A. Peters, Neuropharmacology, 2009, 56, 273–284 CrossRef CAS PubMed.
- G. Hassaine, C. Deluz, L. Grasso, R. Wyss, M. B. Tol, R. Hovius, A. Graff, H. Stahlberg, T. Tomizaki, A. Desmyter, C. Moreau, X.-D. Li, F. Poitevin, H. Vogel and H. Nury, Nature, 2014, 512, 276–281 CrossRef CAS PubMed.
- M. S. Prevost, N. Barilone, G. Dejean de la Bâtie, S. Pons, G. Ayme, P. England, M. Gielen, F. Bontems, G. Pehau-Arnaudet, U. Maskos, P. Lafaye and P.-J. Corringer, Nat. Commun., 2023, 14, 5964 CrossRef CAS.
- Q. Li, Á. Nemecz, G. Aymé, G. D. de la Bâtie, M. S. Prevost, S. Pons, N. Barilone, R. Baachaoui, U. Maskos, P. Lafaye and P.-J. Corringer, Cell. Mol. Life Sci., 2023, 80, 164 CrossRef CAS PubMed.
- A. Ghit, D. Assal, A. S. Al-Shami and D. E. E. Hussein, J. Genet. Eng. Biotechnol., 2021, 19, 123 CrossRef PubMed.
- S. Masiulis, R. Desai, T. Uchański, I. Serna Martin, D. Laverty, D. Karia, T. Malinauskas, J. Zivanov, E. Pardon, A. Kotecha, J. Steyaert, K. W. Miller and A. R. Aricescu, Nature, 2019, 565, 454–459 CrossRef CAS PubMed.
- P. S. Miller, S. Masiulis, T. Malinauskas, A. Kotecha, S. Rao, S. Chavali, L. D. Colibus, E. Pardon, S. Hannan, S. Scott, Z. Sun, B. Frenz, G. Klesse, S. Li, J. M. Diprose, C. A. Siebert, R. M. Esnouf, F. DiMaio, S. J. Tucker, T. G. Smart, J. Steyaert, M. M. Babu, M. S. P. Sansom, J. T. Huiskonen and A. R. Aricescu, bioRxiv, 2018, preprint DOI:10.1101/338343.
- P. S. Miller, S. Scott, S. Masiulis, L. D. Colibus, E. Pardon, J. Steyaert and A. R. Aricescu, Nat. Struct. Mol. Biol., 2017, 24, 986 CrossRef CAS.
- S. Rutz, D. Deneka, A. Dittmann, M. Sawicka and R. Dutzler, Nat. Struct. Mol. Biol., 2023, 30, 52–61 CrossRef CAS PubMed.
- D. Deneka, S. Rutz, C. A. J. Hutter, M. A. Seeger, M. Sawicka and R. Dutzler, Nat. Commun., 2021, 12, 5435 CrossRef CAS PubMed.
- P. Scholler, D. Nevoltris, D. De Bundel, S. Bossi, D. Moreno-Delgado, X. Rovira, T. C. Møller, D. El Moustaine, M. Mathieu, E. Blanc, H. McLean, E. Dupuis, G. Mathis, E. Trinquet, H. Daniel, E. Valjent, D. Baty, P. Chames, P. Rondard and J.-P. Pin, Nat. Commun., 2017, 8, 1967 CrossRef PubMed.
- M. Oosterlaken, A. Rogliardo, T. Lipina, P.-A. Lafon, M. E. Tsitokana, M. Keck, H. Cahuzac, P. Prieu-Sérandon, S. Diem, C. Derieux, C. Camberlin, C. Lafont, D. Meyer, P. Chames, F. Vandermoere, P. Marin, L. Prézeau, D. Servent, A. Salahpour, A. J. Ramsey, C. Bécamel, J.-P. Pin, J. Kniazeff and P. Rondard, Nature, 2025, 1–9 Search PubMed.
- E. Ruiz-López and A. J. Schuhmacher, Biomolecules, 2021, 11, 1131 CrossRef.
- F. Zheng, Y. Pang, L. Li, Y. Pang, J. Zhang, X. Wang and G. Raes, Front. Immunol., 2022, 13, 978513 CrossRef CAS PubMed.
- W. G. Lesniak, C. Chu, A. Jablonska, B. Behnam Azad, O. Zwaenepoel, M. Zawadzki, A. Lisok, M. G. Pomper, P. Walczak, J. Gettemans and M. Janowski, Eur. J. Nucl. Med. Mol. Imaging, 2019, 46, 1940–1951 CrossRef CAS PubMed.
- A. Muruganandam, J. Tanha, S. Narang and D. Stanimirovic, FASEB J., 2002, 16, 240–242 CrossRef CAS.
- G. K. Farrington, N. Caram-Salas, A. S. Haqqani, E. Brunette, J. Eldredge, B. Pepinsky, G. Antognetti, E. Baumann, W. Ding, E. Garber, S. Jiang, C. Delaney, E. Boileau, W. P. Sisk and D. B. Stanimirovic, FASEB J., 2014, 28, 4764–4778 CrossRef CAS PubMed.
- A. Abulrob, H. Sprong, P. V. B. En Henegouwen and D. Stanimirovic, J. Neurochem., 2005, 95, 1201–1214 CrossRef CAS PubMed.
- W. Alata, A. Yogi, E. Brunette, C. E. Delaney, H. van Faassen, G. Hussack, U. Iqbal, K. Kemmerich, A. S. Haqqani, M. J. Moreno and D. B. Stanimirovic, FASEB J., 2022, 36, e22208 CrossRef CAS PubMed.
- A. Yogi, G. Hussack, H. Van Faassen, A. S. Haqqani, C. E. Delaney, E. Brunette, J. K. Sandhu, M. Hewitt, T. Sulea, K. Kemmerich and D. B. Stanimirovic, Pharmaceutics, 2022, 14, 1452 CrossRef CAS PubMed.
- Y. Wouters, T. Jaspers, B. De Strooper and M. Dewilde, Fluids Barriers CNS, 2020, 17, 62 CrossRef CAS PubMed.
- T. J. Esparza, S. Su, C. M. Francescutti, E. Rodionova, J. H. Kim and D. L. Brody, Fluids Barriers CNS, 2023, 20(1), 64 CrossRef CAS PubMed.
- S. Su, T. J. Esparza and D. L. Brody, PLoS One, 2022, 17, e0276107 CrossRef CAS PubMed.
- Y. Wouters, T. Jaspers, L. Rué, L. Serneels, B. De Strooper and M. Dewilde, Fluids Barriers CNS, 2022, 19, 79 CrossRef CAS PubMed.
- D. R. Jones, W. A. Taylor, C. Bate, M. David and M. Tayebi, PLoS One, 2010, 5, e9804 CrossRef.
- Y. Li and J. Wang, J. Labelled Compd. Radiopharm., 2023, 66, 444–451 CrossRef CAS PubMed.
- S. Zhou, X. Fang, Y. Luo, Y. Yang, W. Wei, G. Huang, X. Zhang and C. Wu, Bioconjugate Chem., 2025, 36, 291–301 CrossRef CAS PubMed.
- R. Abskharon, H. Pan, M. R. Sawaya, P. M. Seidler, E. J. Olivares, Y. Chen, K. A. Murray, J. Zhang, C. Lantz, M. Bentzel, D. R. Boyer, D. Cascio, B. A. Nguyen, K. Hou, X. Cheng, E. Pardon, C. K. Williams, A. L. Nana, H. V. Vinters, S. Spina, L. T. Grinberg, W. W. Seeley, J. Steyaert, C. G. Glabe, R. R. Ogorzalek Loo, J. A. Loo and D. S. Eisenberg, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2300258120 CrossRef CAS PubMed.
- T. Li, J.-P. Bourgeois, S. Celli, F. Glacial, A.-M. Le Sourd, S. Mecheri, B. Weksler, I. Romero, P.-O. Couraud, F. Rougeon and P. Lafaye, FASEB J., 2012, 26, 3969–3979 CrossRef CAS PubMed.
- S. R. Meier, D. Sehlin and S. Syvänen, Nucl. Med. Biol., 2022, 114–115, 128–134 CrossRef CAS PubMed.
- Canada, CA2962275C, 2019.
- M. Rotman, M. M. Welling, A. Bunschoten, M. E. Backer, J. Rip, R. J. A. Nabuurs, P. J. Gaillard, M. A. van Buchem, S. M. van der Maarel and L. van der Weerd, J. Controlled Release, 2015, 203, 40–50 CrossRef CAS PubMed.
- U. Iqbal, H. Albaghdadi, M.-P. Nieh, U. I. Tuor, Z. Mester, D. Stanimirovic, J. Katsaras and A. Abulrob, Nanotechnology, 2011, 22, 195102 CrossRef PubMed.
- U. Iqbal, H. Albaghdadi, Y. Luo, M. Arbabi, C. Desvaux, T. Veres, D. Stanimirovic and A. Abulrob, Br. J. Cancer, 2010, 103, 1606–1616 CrossRef CAS PubMed.
- S. Zhu, A.-G. Huang, F. Luo, J. Li, J. Li, L. Zhu, L. Zhao, B. Zhu, F. Ling and G.-X. Wang, ACS Appl. Mater. Interfaces, 2019, 11, 19006–19016 CrossRef CAS.
- M. Scully, S. R. Cataland, F. Peyvandi, P. Coppo, P. Knöbl, J. A. K. Hovinga, A. Metjian, J. de la Rubia, K. Pavenski, F. Callewaert, D. Biswas, H. D. Winter and R. K. Zeldin, N. Engl. J. Med., 2019, 380, 335–346 CrossRef CAS PubMed.
- S. Oyama, K. Ebina, Y. Etani, M. Hirao, M. Kyuuma, Y. Fujii, K. Iwata, B. Ogawa, T. Hasegawa, S. Kawano, Y. Nakanishi, S. Okada and K. Nakata, Sci. Rep., 2022, 12, 18102 CrossRef CAS PubMed.
- Y. Tanaka, Expert Opin. Biol. Ther., 2023, 23, 579–587 CrossRef CAS PubMed.
- S. J. Keam, Drugs, 2023, 83, 87–92 CrossRef CAS PubMed.
- M. Chen, M. Jiang, X. Wang, L. Shen and J. Li, Expert Opin. Biol. Ther., 2022, 22, 1227–1232 CrossRef CAS.
- A. Markham, Drugs, 2022, 82, 235–240 CrossRef CAS PubMed.
- T. Martin, S. Z. Usmani, J. G. Berdeja, M. Agha, A. D. Cohen, P. Hari, D. Avigan, A. Deol, M. Htut, A. Lesokhin, N. C. Munshi, E. O’Donnell, A. K. Stewart, J. M. Schecter, J. D. Goldberg, C. C. Jackson, T.-M. Yeh, A. Banerjee, A. Allred, E. Zudaire, W. Deraedt, Y. Olyslager, C. Zhou, L. Pacaud, D. Madduri, A. Jakubowiak, Y. Lin and S. Jagannath, J. Clin. Oncol., 2023, 41, 1265–1274 CrossRef CAS PubMed.
- I. Jovčevska and S. Muyldermans, BioDrugs, 2020, 34, 11–26 CrossRef.
- C. J. Hutchings, P. Colussi and T. G. Clark, mAbs, 2018, 11, 265 CrossRef.
- V. Dalal, B. K. Tan, H. Xu and W. W. Cheng, eLife, 2025, 14, RP106728 CrossRef.
- M. Sánchez-Marín, M. Giulini and A. M. J. J. Bonvin, Combining AI structure prediction and integrative modelling for nanobody-antigen complexes, bioRxiv, 2025, preprint DOI:10.1101/2025.07.01.662355.
- F. N. Hitawala and J. J. Gray, What does AlphaFold3 learn about antigen and nanobody docking, and what remains unsolved?, bioRxiv, 2025, preprint DOI:10.1101/2024.09.21.614257.
- F. Eshak and A. Goupil-Lamy, 2024.
- T. Cohen, M. Halfon and D. Schneidman-Duhovny, Front. Immunol., 2022, 13, 958584 CrossRef CAS PubMed.
- J. A. Ruffolo, L. S. Chu, S. P. Mahajan and J. J. Gray, Nat. Commun., 2023, 14(1), 2389 CrossRef CAS.
- L. Wang, X. He, G. Guo, X. Qian and Q. Huang, Computational nanobody design using graph neural networks and Metropolis Monte Carlo sampling, bioRxiv, 2025, preprint DOI:10.1101/2025.06.08.658414.
- X. Cheng, J. Wang, G. Kang, M. Hu, B. Yuan, Y. Zhang and H. Huang, Int. J. Mol. Sci., 2019, 20, 4187 CrossRef PubMed.
- Z. Zhang, R. Van Der Kant, I. Marković, D. Vizarraga, T. Garcia, K. Maragkou, J. Delgado Blanco, D. Cianferoni, G. Orlando, G. Cia, N. Geukens, C. Carolis, A. N. Volkov, S. N. Savvides, M. Dewilde, J. Schymkowitz, L. Serrano Pubul and F. Rousseau, In silico design of stable single-domain antibodies with high affinity, bioRxiv, 2024, preprint DOI:10.1101/2024.04.22.589762.
- H. Feng, X. Sun, N. Li, Q. Xu, Q. Li, S. Zhang, G. Xing, G. Zhang and F. Wang, ACS Omega, 2024, 9, 47893–47902 CrossRef CAS PubMed.
- F. S. Ahmed, S. Aly and X. Liu, Briefings Bioinf., 2024, 26(1), bbae518 CrossRef PubMed.
- J. T. Hadsund, T. Satława, B. Janusz, L. Shan, L. Zhou, R. Röttger and K. Krawczyk, Bioinf. Adv., 2024, 4(1), vbae033 Search PubMed.
- S. Dandibhotla, M. Samudrala, A. Kaneriya and S. Dakshanamurthy, Pharmaceuticals, 2025, 18(3), 329 CrossRef CAS PubMed.
- Y. Zhang, Z. Wang, F. Ge, X. Wang, Y. Zhang, S. Li, Y. Guo, J. Song and D. J. Yu, Briefings Bioinf., 2024, 25(6), bbae489 CrossRef CAS PubMed.
- M. Pacesa, L. Nickel, C. Schellhaas, J. Schmidt, E. Pyatova, L. Kissling, P. Barendse, J. Choudhury, S. Kapoor, A. Alcaraz-Serna, Y. Cho, K. H. Ghamary, L. Vinué, B. J. Yachnin, A. M. Wollacott, S. Buckley, A. H. Westphal, S. Lindhoud, S. Georgeon and B. E. Correia, Nature, 2025, 646(8084), 483–492 CrossRef CAS PubMed.
- V. Dewaker, V. K. Morya, Y. H. Kim, S. T. Park, H. S. Kim and Y. H. Koh, Biomark. Res., 2025, 13(1), 52 CrossRef PubMed.
- N. S. El Salamouni, J. H. Cater, L. M. Spenkelink and H. Yu, FEBS Open Bio, 2025, 15, 236–253 CrossRef CAS PubMed.
- J. Liu, L. Wu, A. Xie, W. Liu, Z. He, Y. Wan and W. Mao, J. Nanobiotechnol., 2025, 23(1), 87 CrossRef PubMed.
- H. Zhu and Y. Ding, Biology, 2025, 14(5), 547 CrossRef CAS.
- E. Karakas, M. C. Regan and H. Furukawa, Trends Biochem. Sci., 2015, 40, 328–337 CrossRef CAS PubMed.
- E. F. Pettersen, T. D. Goddard, C. C. Huang, E. C. Meng, G. S. Couch, T. I. Croll, J. H. Morris and T. E. Ferrin, Protein Sci., 2021, 30, 70–82 CrossRef CAS.
- E. Pardon, T. Laeremans, S. Triest, S. G. F. Rasmussen, A. Wohlkönig, A. Ruf, S. Muyldermans, W. G. J. Hol, B. K. Kobilka and J. Steyaert, Nat. Protoc., 2014, 9, 674–693 CrossRef CAS PubMed.
- M. Arbabi-Ghahroudi, Front Immunol., 2017, 8, 1589 CrossRef PubMed.
- K. A. Henry and C. R. MacKenzie, Front. Immunol., 2018, 9, 41 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |