Open Access Article
Open Access ArticleDiscovery of natural products that modulate signaling in patient-derived cells
Joseph A.
Balsamo†
 a,
Hannah L.
Thirman†
a,
Hannah L.
Thirman†
 bcf,
Kathryn E.
Penton
bcf,
Kathryn E.
Penton
 d,
Jordan T.
Froese
d,
Jordan T.
Froese
 e,
Benjamin J.
Reisman
e,
Benjamin J.
Reisman
 b,
Sierra M.
Lima
b,
Sierra M.
Lima
 c,
Madeline J.
Grider-Hayes
c,
Madeline J.
Grider-Hayes
 cf,
Chad R.
Potts
cf,
Chad R.
Potts
 gh,
Jonathan M.
Irish
gh,
Jonathan M.
Irish
 cfg,
P.
Brent Ferrell
cfg,
P.
Brent Ferrell
 gh and
Brian O.
Bachmann
gh and
Brian O.
Bachmann
 *abc
*abc
aDepartment of Pharmacology, Vanderbilt University, Vanderbilt University School of Medicine, Nashville, Tennessee, USA. E-mail: brian.bachmann@vanderbilt.edu
bProgram in Chemical & Physical Biology, Vanderbilt University School of Medicine, Nashville, Tennessee, USA
cDepartment of Cell & Developmental Biology, Vanderbilt University, Nashville, Tennessee, USA
dDepartment of Chemistry, Vanderbilt University, Nashville, Tennessee, USA
eDepartment of Chemistry, Ball State University, 2000 W. University Ave, Muncie, USA
fDepartment of Pediatrics, University of Colorado, Anschutz Medical Campus, Aurora, CO, USA
gVanderbilt-Ingram Cancer Center, Vanderbilt University Medical Center, Nashville, Tennessee, USA
hDepartment of Medicine, Division of Hematology/Oncology, Vanderbilt University Medical Center, Nashville, Tennessee, USA
First published on 25th February 2026
Abstract
Natural products derived from biosynthetically apt microorganisms represent an important source for chemotherapeutic agents founded first via non-specific, cytotoxicity-guided isolation, then investigated for translational potential. Analyzing biosynthetic gene clusters within microbes predicts significant untapped potential of undiscovered therapeutically relevant natural products; however, stimulating biosynthesis of such compounds in sufficient quantities for isolation, structure determination, and biological assessment remains a gap. We address this with a bioactivity discovery pipeline that first utilizes Multiplexed Activity Profiling to identify stimulus-dependent induction of bioactive secondary metabolites via functional responses in human leukemia cell lines measured with fluorescence flow cytometry. Next, active extracts are analyzed via Multiplexed Activity Metabolomics, which correlates single cell assays with spectrally defined chromatographic arrays to identify bioactive metabolomic features prior to isolation. Two experimental case studies using this workflow with cave-microbe extracts identified new molecular phenotypes of the pyridine-pyrrolidine alkaloid siderochelin and the pyrrolopyrrole-functionalized anthracycline isoquinocycline B. Finally, we investigated the effects of anthracycline functionalization on human primary cells using single cell mass cytometry to understand if changes in anthracycline aglycon structure and glycosylation impacted cell selectivity. Despite sharing an anthracyclinone pharmacophore core, variants differentially impacted responses of leukemia cell populations within and among acute myeloid leukemia (AML) patient samples. This suggests pharmacophore assumptions may not guide assessment of therapeutic potential for anthracyclines and that modest structural differences can elicit marked changes in cellular function in different patients. Taken together, the depth of information afforded by single cell molecular phenotype-based discovery and mass cytometry deep cell profiling provides new patient-level insight into biological mechanisms of new and previously discovered molecules that may find expanded use in the clinic.
Introduction
Microbial natural products, also known as secondary metabolites, form the basis for many human therapeutics for diverse clinical indications.1,2 However, the journey from a microbial source, to newly discovered metabolite, to the clinic, is circuitous and not frequently driven by human biological insights nor clinically relevant endpoints. For example, staurosporines and anthracyclines are well represented in the clinic for treating hematological malignancies3–5 and they are even more highly represented in nature's pharmacopeia. However, these individual natural products were discovered based on cytotoxicity screening in cell lines and their early-stage development in humans was not governed by patient-specific biology. Hence, it is largely possible that many undiscovered scaffolds, known scaffolds, and/or scaffold analogs possess unique pharmacological properties in humans that are not revealed by traditional cytotoxicity and target-based discovery paradigms. Our goal is to accelerate natural product discovery in the context of precision medicine in a way that takes advantage of the structural diversity and variation between and within classes of secondary metabolites to identify compounds with high clinical potential for development. Herein, we identify and provide a single cell chemical biology platform to address three challenges regarding microbial natural product discovery for human medicine: determination of conditions for biosynthetic gene cluster (BGC) activation, identification of the active component in microbial crude extracts, and translational insight into human pharmacology (Fig. 1).There are multiple strategies to explore novel biosynthetic and chemical space including the discovery of novel taxa from nature, genome mining of annotated culture collections,6,7 and metagenomic analysis of complex ecosystems.8–10 Microbial genomic sequencing efforts have uncovered a previously unappreciated richness of cryptic BGCs that encode predicted biosynthesis of known and novel secondary metabolites.11,12 Biochemical inference permits structural prediction of microbial secondary metabolites, enabling genome mining strategies,11–15 in the hope that novel structures engender novel clinical potential. However, one rate-determining step in microbial genome mining approaches is discovering conditions to activate the expression, translation, and production of the products of secondary metabolic gene clusters. This may be accomplished by optimizing production via heterologous expression in chassis organisms,16–18 genetic modification of native producers,19,20 or via discovering conditions for chemical and biological stimulation in native producers.21–26 These methods are currently resource and time-consuming activities that are often prerequisites for the additionally labor-intensive processes of isolation, structure elucidation, and biological assessment of individual metabolites. Another limitation of some genome mining strategies is that prioritization is rarely informed by foreknowledge of potentially interesting biological activity. Resistance genes can provide clues27 and the development methods for sequence-based estimation are an active area of research;28–30 however, it has become uncommon to include screening in the process of discovery, and when it is, it is generally via a single phenotypic assay such as cytotoxicity.31,32 To unambiguously link initial biological phenotypes induced by an extract to a discrete metabolite, it is typically necessary to isolate and structurally identify a metabolite of interest prior to investigating the scope of its bioactivity. Isolation is a second rate-determining step in activity-based workflows.
To address stimulus mining and metabolite isolation bottlenecks, we utilized Multiplexed Activity Profiling (MAP)25,33 to simultaneously correlate multiple stimuli of natural product biosynthesis against multiple single cell phenotypic markers of mammalian cell function to rapidly determine productive activation conditions. Active extracts were then fractioned for Multiplexed Activity Metabolomics (MAM)31,33 to track biological activity to known molecular features within active extracts prior to isolation and rapidly prioritize potentially new molecular features for further characterization. MAP and MAM accelerate the discovery of both previously unreported secondary metabolites and known compounds with novel biological phenotypes with potential for human medicine.
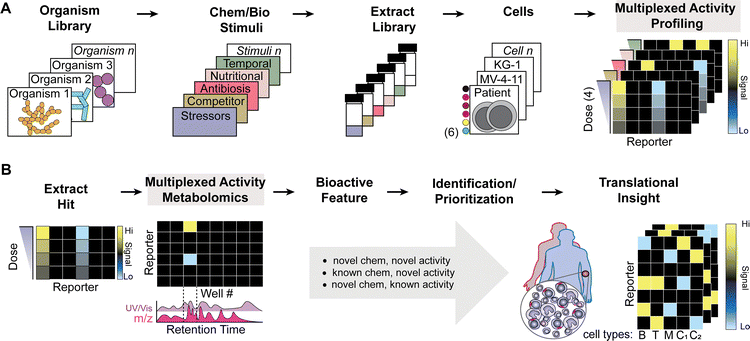
Herein, we demonstrate a bench-to-bedside inspired discovery workflow (Fig. 1) that integrates aforementioned design components for natural product discovery in the context of precision medicine. We provide two compound identification case studies that illustrate MAP and MAM for discovery and isolation of bioactive molecules. Following isolation, compounds are tested in primary samples based on new bioactivity, structure, and clinical precedent. Two genera, isolated from cave systems, were subjected to the above workflow: Amycolatopsis coloradensis ssp. ossimia synthesized the pyridine-pyrrolidine alkaloid siderochelin,34,35 which exhibited new bioactivity that impacted DNA damage and translation response pathways, and Micromonospora phytophila ssp. tarocco manufactured isoquinocycline B,36–39 an anthracycline analog that elicits opposite cell cycle phenotypes of doxorubicin in the MV-4-11 cell line and discretely targets AML patient cell subsets. We propose this workflow identifies clinically relevant human biological insights for microbial compounds and delivers new leads with unprecedented levels of human cell characterization.
Experimental
MV-4-11 cell culture
MV-4-11 is a human AML cell line isolated from the blasts of a 10-year-old male with biphenotypic B-myelomonocytic leukemia.40 MV-4-11 leukemia cells were purchased from ATCC (CRL-9591) and subsequently confirmed as mycoplasma negative. Cells were cultured in IMDM (Gibco 12440-053) that was supplemented to 10% fetal bovine serum at final concentration. Cells were kept in a water-jacketed 5% CO2 incubator at 37 °C and maintained at densities between 100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 and 1
000 and 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000
000![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per mL of culture media. The cells were fed every other day and passaged every 4 days.
000 cells per mL of culture media. The cells were fed every other day and passaged every 4 days.
AML patient samples
All specimens were obtained in accordance with the Declaration of Helsinki following protocols approved by the Vanderbilt University Medical Center Institutional Review Board. Details of patients and sample acquisition were previously published.41 Briefly, consent was obtained via an approved written consent form, and eligibility criteria included ≥18 years of age with suspected AML undergoing clinical evaluation at Vanderbilt. Samples analyzed here were collected from bone marrow prior to any treatment. Once obtained, samples underwent immediate (within <30 min) density gradient separation of mononuclear cells using a BD Vacutainer CPT Cell Preparation Tube with Sodium Heparin (BD Biosciences, Franklin Lakes, NJ). The separated mononuclear cells were then pelleted with low-speed centrifugation (200 × g) and aliquoted into multiple cryotubes in an 88% FBS + 12% DMSO solution. Patient genotypes are documented in Table S1.Method details
Cave strain isolation and identification
Strains were isolated from swab samples from Blue Spring Cave in Sparta, Tennessee, which were vortexed in sterile water (100 mg mL−1). Supernatants were serially diluted (10-fold, 100-fold, and 1000-fold) and plated on minimal medium agar plates (ISP-2 medium/100-fold diluted, 1.5% agar). Plates were then incubated at 30 °C, and colonies were picked over a three-week period. DNA isolations for purified colonies were performed with a commercial kit (Wizard DNA isolation kit; Promega, Inc.). The 16S rRNA genes for these were then amplified with universal primers 27F (AGA GTT TGA TCC TGG CTC AG) and 1525R (AAG GAG GTG ATC CAG CCG CA) and the use of high-fidelity DNA polymerase (Phusion; Thermo, Inc.). The PCR thermocycler conditions involved an initial incubation at 98 °C for 1 min and 30 cycles of (i) denaturation at 98 °C for 0.5 min, (ii) annealing at 59 °C for 1 min, and (iii) extension at 72 °C for 1.5 min. The final extension step was conducted at 72 °C for 10 min. Target 16S rDNA amplicons were purified using a gel extraction kit (QIAquick gel extraction kit). Purified PCR products were directly sequenced from both the 5′ and 3′ ends, and these sequences were combined to give the near-full-length 16S rDNA gene sequences. Preliminary genus identifications were determined by a comparison of 16S rDNA gene sequences to type strains in the EzBioCloud database.42 Phylogenetic analysis was accomplished in MEGA7 by muscle alignment of 16S sequences and phylogenetic tree formation using the maximum likelihood method of 1000 bootstrapped experiments.43Scanning electron microscopy
M. phytophila subsp. tarocco and A. coloradensis ssp. ossimia were plated on ISP-2 agar, overlaid with nylon membrane, and incubated at 30 °C for 3 weeks. Samples were air dried, coated with gold palladium using a Cressington 108 sputter coater and imaged on a FEI Quanta 250 SEM at 5 kV.Preparation of microbial crude extracts
Streptomyces strains were maintained on ISP-2 agar (yeast extract 4 g L−1, malt extract 10 g L−1, glucose 4 g L−1, and agar 20 g L−1, pH 7.2). Loops of mycelia were used to inoculate 5-mL seed cultures in ISP-2 medium (yeast extract 4 g L−1, malt extract 10 g L−1, and glucose 4 g L−1, pH 7.2) for Streptomyces strains, incubating them for 3 days at 30 °C. Seed cultures were then transferred to 250-mL Erlenmeyer flasks containing 25 mL base media (ISP-2) alone or stimulated in ISP-2 with 1.5 mM LaCl3 or 120 nM rifampin or rich media (Table S2) grown for 7 days at 30 °C with shaking. Aqueous fermentation broth was extracted by shaking with Diaion HP20 synthetic absorbent resin (Alfa Aesar) (125 mL of HP20 bead/H2O slurry per 500 mL of aqueous broth) for 2 h. Fermentation broth was then centrifuged (3700 × g, 30 min) and the supernatant was decanted. Metabolites were eluted from absorbent resin and cells with methanol (250 mL of methanol/125 mL of HP20 bead/H2O slurry) by shaking for 1.5 h, followed by centrifugation (3700 × g, 30 min) and decanting of the methanol extract. Further extraction was performed with acetone (250 mL of acetone/125 mL of HP20 bead/H2O slurry) by shaking for 1.5 h, followed by centrifugation (3700 × g, 30 min) and decanting of the acetone extract.Generation of metabolomic arrays
Mass spectrometry was performed by using a TSQ Triple Quantum mass spectrometer equipped with an electrospray ionization source and Surveyor PDA Plus detector. For positive ion mode, the following settings were used: capillary temperature was 270 °C; spray voltage 4.2 kV; spray current 30 mA; capillary voltage 35 V; tube lens 119 V; and skimmer offset 15 V. For negative ion mode, capillary temperature 270 °C; spray voltage 30 kV; spray current 20 mA; capillary voltage 35 V; tube lens 119 V; and skimmer offset 15 V. Fraction plates were prepared by injecting 20 µL of 200 µM purified compounds in methanol or concentrated extract (2.5 mg mL−1) via a Thermo PAL auto injector onto a Phenomenex luna 5 µm C182 reverse-phase HPLC column. The sample was fractionated using a gradient of 100% Buffer A (95% H2O, 5% acetonitrile) to 100% Buffer B (5% acetonitrile, 95% H2O) over 48 min at a flow rate of 1 mL min−1 and a fixed splitter with a 3![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 ratio with three parts going to the photodiode array detector and fraction collector and one part going to the MS. Fractions were collected in 1-min intervals in a 96 deep well plate. A volume of 150 µL of eluent from each well was transferred to four replica plates and dried in vacuo using a Genevac HT-6 system with standard HPLC-Lyo protocol.
1 ratio with three parts going to the photodiode array detector and fraction collector and one part going to the MS. Fractions were collected in 1-min intervals in a 96 deep well plate. A volume of 150 µL of eluent from each well was transferred to four replica plates and dried in vacuo using a Genevac HT-6 system with standard HPLC-Lyo protocol.
Fluorescent cell barcoding of cell-seeded metabolomic arrays
Eight serial 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2.14 dilutions of Pacific Blue were prepared, covering a concentration range from 0.038 to 7.67 µg mL−1. Six serial 1
2.14 dilutions of Pacific Blue were prepared, covering a concentration range from 0.038 to 7.67 µg mL−1. Six serial 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2.5 dilutions of Pacific Orange were prepared, covering a concentration range from 0.22 to 21 µg mL−1. Each dilution of Pacific Blue was added to all wells in a single row of a 96-well plate (10 µL per well), so that the dye concentration in each row decreased from the top to the bottom of the plate. Similarly, each dilution of Pacific Orange was added to all wells in a column of the same 96-well plate (10 µL per well), so that the concentration in each column decreased from columns 1 to 6 and from columns 7 to 12. This procedure yielded two sets of 48 barcoded wells per plate. Approximately 200
2.5 dilutions of Pacific Orange were prepared, covering a concentration range from 0.22 to 21 µg mL−1. Each dilution of Pacific Blue was added to all wells in a single row of a 96-well plate (10 µL per well), so that the dye concentration in each row decreased from the top to the bottom of the plate. Similarly, each dilution of Pacific Orange was added to all wells in a column of the same 96-well plate (10 µL per well), so that the concentration in each column decreased from columns 1 to 6 and from columns 7 to 12. This procedure yielded two sets of 48 barcoded wells per plate. Approximately 200![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells (180 µL suspended in phosphate-buffered saline (PBS)) were added to each well and incubated in the dark at room temperature for 30 min. Staining was then quenched by addition of 75 µL of 1% BSA (Sigma) in PBS.
000 cells (180 µL suspended in phosphate-buffered saline (PBS)) were added to each well and incubated in the dark at room temperature for 30 min. Staining was then quenched by addition of 75 µL of 1% BSA (Sigma) in PBS.
Antibody staining
Cells were stained with antibodies in 100 µL staining medium for 30 min in the dark, unless otherwise noted. Individual antibodies were added in accordance with the manufacturer's instructions. Staining was quenched with 1% BSA in PBS, and stained cells were washed with PBS prior to analysis.Isolation of siderochelin
A. coloradensis subsp. ossimia was plated on ISP-2 (agar yeast extract 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g L−1, malt extract 10
g L−1, malt extract 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g L−1, glucose 4
g L−1, glucose 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g L−1, and agar 20
g L−1, and agar 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g L−1, pH 7.2) from glycerol stocks and incubated at 30 °C for 5–7 days. Cells were then harvested from solid culture, homogenized, and used to inoculate seed cultures (25 mL ISP-2 liquid media: yeast extract 4
g L−1, pH 7.2) from glycerol stocks and incubated at 30 °C for 5–7 days. Cells were then harvested from solid culture, homogenized, and used to inoculate seed cultures (25 mL ISP-2 liquid media: yeast extract 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g L−1, malt extract 10
g L−1, malt extract 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g L−1, glucose 4
g L−1, glucose 4![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) g L−1, pH 7.2) in 250 mL Erlenmeyer flasks. Seed cultures were incubated at 30 °C for 48 hours, cells were then harvested from liquid cultures, homogenized, and used to inoculate 500 mL rich media (Table S2). After 7-day incubation, 100 mL of activated Diaion® HP-20 resin (Supelco)/H2O slurry was added to each 500 mL flask and flasks were incubated with shaking for 3 hours. Cells/resin were then centrifuged (3700 × g, 30
g L−1, pH 7.2) in 250 mL Erlenmeyer flasks. Seed cultures were incubated at 30 °C for 48 hours, cells were then harvested from liquid cultures, homogenized, and used to inoculate 500 mL rich media (Table S2). After 7-day incubation, 100 mL of activated Diaion® HP-20 resin (Supelco)/H2O slurry was added to each 500 mL flask and flasks were incubated with shaking for 3 hours. Cells/resin were then centrifuged (3700 × g, 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) min), the supernatant discarded, and the resin/cellular mass extracted with 200 mL methanol/100 mL Diaion® HP-20 resin (Supelco) used (incubation with shaking for 1 h). Cell mass/resin was centrifuged (3700 × g, 30
min), the supernatant discarded, and the resin/cellular mass extracted with 200 mL methanol/100 mL Diaion® HP-20 resin (Supelco) used (incubation with shaking for 1 h). Cell mass/resin was centrifuged (3700 × g, 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) min), methanol extract removed, concentrated and stored, and the resin/cellular mass extracted with 200 mL acetone/100 mL HP-20 resin used (incubation with shaking for 1 h). The cell mass/resin was again centrifuged (3700 × g, 30
min), methanol extract removed, concentrated and stored, and the resin/cellular mass extracted with 200 mL acetone/100 mL HP-20 resin used (incubation with shaking for 1 h). The cell mass/resin was again centrifuged (3700 × g, 30![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) min), and the acetone extract was removed, concentrated and stored. Crude acetone extract was fractionated with Sephadex LH-20 resin (GE Healthcare Bio-Sciences) with methanol as the eluent. Fractions were analyzed by analytical HPLC/MS, and fractions containing the compound(s) of interest were pooled and further purified by preparative HPLC (Waters, XBridge C18 Prep, 5
min), and the acetone extract was removed, concentrated and stored. Crude acetone extract was fractionated with Sephadex LH-20 resin (GE Healthcare Bio-Sciences) with methanol as the eluent. Fractions were analyzed by analytical HPLC/MS, and fractions containing the compound(s) of interest were pooled and further purified by preparative HPLC (Waters, XBridge C18 Prep, 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) µM) (10
µM) (10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) mL min−1: linear gradient of 100% Solution A to 100% Solution B. Solution A = 95
mL min−1: linear gradient of 100% Solution A to 100% Solution B. Solution A = 95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5, H2O
5, H2O![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeCN, 10
MeCN, 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) mM NH4OAc; Solution B: 5
mM NH4OAc; Solution B: 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 95 H2O
95 H2O![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeCN, 10
MeCN, 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) mM NH4OAc). Siderochelin was dereplicated using a combination of mass spectrometry and nuclear magnetic resonance spectroscopy data (Fig. S1).
mM NH4OAc). Siderochelin was dereplicated using a combination of mass spectrometry and nuclear magnetic resonance spectroscopy data (Fig. S1).
Isolation of isoquinocycline B
Isoquinocycline B was isolated from M. phytophila subsp. tarocco as described in the isolation of siderochelin except seed cultures were scaled in 500 mL base media (Table S2) and isolated as described above except for preparative HPLC isolation (Waters, XBridge C18 Prep, 5 µM) (10 mL min−1, 0–1 min: 100% solution A, 45 min: 100% solution B, 55 min: 100% solution B, 56 min: 100% solution A, 60 min: 100% solution A) (solution A = 95![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5, H2O
5, H2O![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeCN, 10 mM NH4OAc; solution B: 5
MeCN, 10 mM NH4OAc; solution B: 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 95 H2O
95 H2O![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) MeCN, 10 mM NH4OAc). No flash column chromatography was utilized. Isoquinocycline B was dereplicated using a combination of mass spectrometry and nuclear magnetic resonance spectroscopy data (Fig. S2).
MeCN, 10 mM NH4OAc). No flash column chromatography was utilized. Isoquinocycline B was dereplicated using a combination of mass spectrometry and nuclear magnetic resonance spectroscopy data (Fig. S2).
Mass cytometry
Mass cytometry experiments were performed as previously described.44 Briefly, after compound challenge samples were pelleted by centrifugation at 200 × g, resuspended and washed with PBS (HyClone, HyClone Laboratories, Logan, UT), then pelleted, and resuspended in PBS. They were then stained with Cell-ID Cisplatin (Fluidigm, South San Francisco, CA) as per the manufacturer's recommended protocol. The cells were washed and resuspended in staining medium [CSM: PBS + 1% BSA (Fisher Scientific, Fair Lawn, NJ)]. Cells were then stained with a mass cytometry antibody panel of 22 extracellular antibodies designed to characterize AML blasts and most non-AML peripheral blood mononuclear cells (Table S3). A master mix of these antibodies was added to each sample to give a final staining volume of 50 µL and incubated at room temperature for 30 min.CyTOF FCB data preprocessing and computational deconvolution
After acquisition, data was uploaded and stored in Cytobank for scaling, quality control gating, compensation, and analysis of unmixed cytometry data (FCS file format). Raw median intensity values were transformed into a hyperbolic arcsine scale with a cofactor of 5 for all readouts. Quality control gating (QC) was performed as shown in Fig. S3. Scaled and gated samples were compensated and then computationally deconvoluted using the DebarcodeR algorithm. The resulting demultiplexed FCS files for each well were uploaded to Cytobank for storage and further analysis.Dimensionality reduction and fold change on UMAP analyses
Fold change on UMAP calculations were inspired by Sconify, a toolkit published in 2018 by Burns et al.,45 and were performed as follows: after debarcoding, the FCS files corresponding to cellular events from all patients and treatment conditions were uploaded to one Cytobank experiment and fed into a UMAP analysis (settings: population: viable, single, CD45+ cells, channels = all surface markers listed in Table S3 except HLA-DR, event sampling = equal sampling from each patient/treatment condition, num neighbors = 15, minimum distance = 0.01, collapse outliers = yes). FCS files containing intensity measurements for all readouts with the UMAP axes appended were exported into R for subsequent analysis. To compute fold change on the UMAP in R, for each patient, data from one anthracycline and the vehicle was selected, and arcsinh transformed with a cofactor of 5. K nearest neighbors (KNN) was then performed using a K equal to the square root of the total number of cells. Then, for each cell neighborhood, and each of the intracellular markers listed in Table S3, the median of the vehicle data was subtracted from the median of the drug data. For more information on arcsinh scaling in cytometry analysis, see first use in the Supplement of ref. 46 and more recent implementations.31,47–49Quantification and statistical analysis
All methods and details regarding quantifications and analyses are described in the relevant part of the main text, figure legends, and/or Methods section. Experiments were not randomized and the sample sizes are listed within the figures, main text, figure legends and/or Methods section.Results
Single cell methods enable multiplexing of human cell phenotypic markers with dose–response data to prioritize bioeffector isolation from complex metabolic extracts
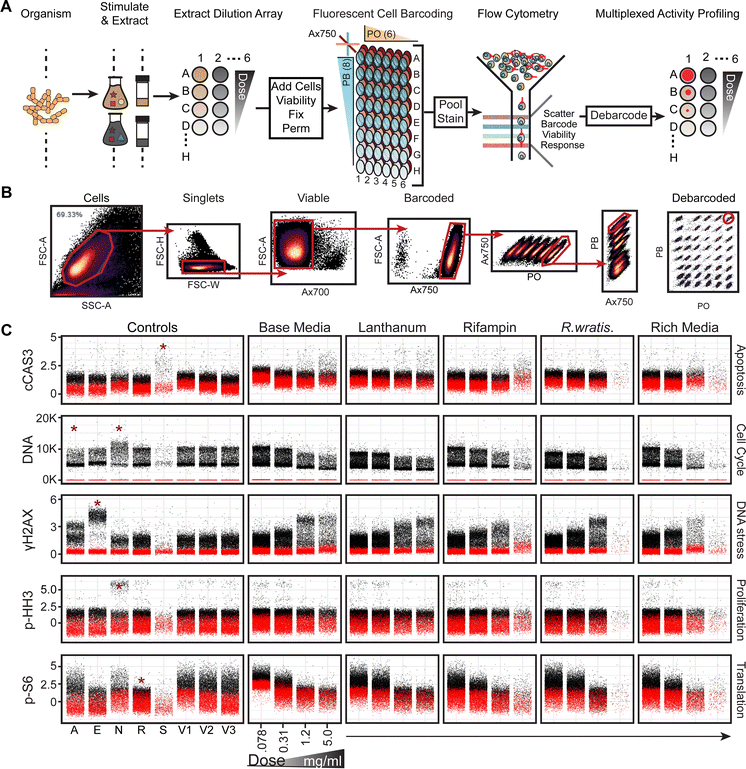
To drive discovery of bioactive natural products, we developed a cell function-centric50 screen that integrates multiple single cell intracellular marker quantification31,51 to coordinate stimulus conditions, dose–response data, and fluorescent reporter assays for interrogation of microbial extracts from diverse culture collections (Fig. 1A). The Vanderbilt Laboratory for Biosynthetic Studies Actinobacteria collection houses >500 actinomycetes purified by restrictive culturing methods from cave ecosystems, an under-utilized source of diverse actinomycete genera with predicted biosynthetic potential.24,52–57 Following phylogenetic assessment by 16S rDNA sequences, microbes were cultivated using chemical and biological stimuli (Fig. 1A and Table S2) in liquid cultures to activate stimulus sensitive secondary metabolite biosynthesis.23,24,56,58–62 Broth and biomass were extracted with hydrophobic solid phase extraction and cataloged in extract libraries. Extracts were serially diluted 64-fold on 96-well plates, dried in vacuo, and dissolved in cancer cell line or primary patient cell containing culture media (Fig. 1A) for overnight incubation.Incubated cells were stained with Alexa Fluor 700 (Ax700) (Table S4) for use in downstream identification and gating of non-viable cells with depolarized cell membranes that would otherwise complicate data interpretation, fixed in 1.6% paraformaldehyde, permeabilized in ice-cold methanol, and stained with NHS-amine reactive fluorescent cell barcoding (FCB) reagents (Table S4) to coordinate cell populations to origin plate wells (Fig. 2A and 3A).31,63,64 We arrayed eight row-oriented levels of Pacific Blue with six column-oriented levels of Pacific Orange and a single level of Alexa Fluor 750 (Ax750) for discrete labeling of forty-eight unique experimental conditions (Fig. S4). Ax750 serves as an uptake control to account for variability of FCB staining across viable cell morphologies in a sample (Fig. S3 and S4). Barcoded cells were pooled and stained with an antibody cocktail designed to measure cleaved Caspase-3 protein (c-CAS3, apoptosis), γH2AX (DNA damage), phosphorylated-Histone H3 (p-HH3, proliferation), and p-S6 (nutrient sensing, cell metabolism, and protein translation) alongside the fluorescent DNA intercalator YoPro (DNA content, cell cycle stage) (Table S4). Use of FCB prior to antibody staining helps ensure uniform staining across samples and dramatically reduces fluorescent reagent cost/assay and time to data acquisition.
Fluorescently labeled and light scattering single cell marker response information was acquired simultaneously by flow cytometry65 and stored in Cytobank. Intact, single cells were gated from cellular debris (Fig. 2B and Fig. S3). Viable, Ax700 negative, cells were selected (Fig. 2B) and gated for Ax750 uptake control positive cells (Fig. S3). Within Ax750 positive cells, six fluorescent levels of Pacific Orange-stained cells were ascertainable (Fig. 2B and Fig. S3). Gating on any Pacific Orange population reveals eight fluorescent levels of Pacific Blue-stained cells (Fig. 2B). Taken together, this gating strategy enables multiplexing of forty-eight unique experimental conditions with six fluorescent biometric readouts to perform 288 single cell assays in approximately ten minutes on a capable flow cytometer (Fig. 1A). Following acquisition, cell events were reassigned, or “debarcoded”, to original wells by coordinated fluorescent dye intensity (Fig. 2A, Fig. 3A). This process is semi-automated by an in-lab developed R code script available on GitHub termed DebarcodeR (Fig. 2B).66 Visualization of debarcoded single cell data is presented using “Rothko” plots (inspired by the painter Mark Rothko). In this representation, each point corresponds to an individual gated, intact, viable cell. Black points denote barcoded cells stained with the antibody cocktail and thus report marker-specific fluorescence. Red points represent FCB control cells that are treated with compound and barcoded but are not stained with the antibody cocktail and therefore capture background and compound-associated fluorescence. A detailed protocol describing the use of flow cytometry in combination with FCB for multiplexing and DebarcodeR for subsequent computational deconvolution is also available.66,67
Since natural product extracts may themselves contain fluorescent molecules and even quenchers, we generate a control pool of treated samples with no antibody stain (shown as red dots in Rothko plots), to detect any sample-specific fluorescence that may interfere with barcode dyes or fluorescent antibodies. In the event of fluorescent interference in samples, the red dots, which correspond to the shift in fluorescence at each channel, will shift, revealing interference. This has not occurred in the data shown herein. However, if endogenous fluorescence is observed in unstained controls, any marker shifts must be validated by either verifying in non-barcoded experiments if the fluorescence overlaps with barcoding dyes, using an alternative fluorophore for the marker of interest if the fluorescence overlaps with markers, or by pre-fractionating the extracts to remove fluorescent compounds.
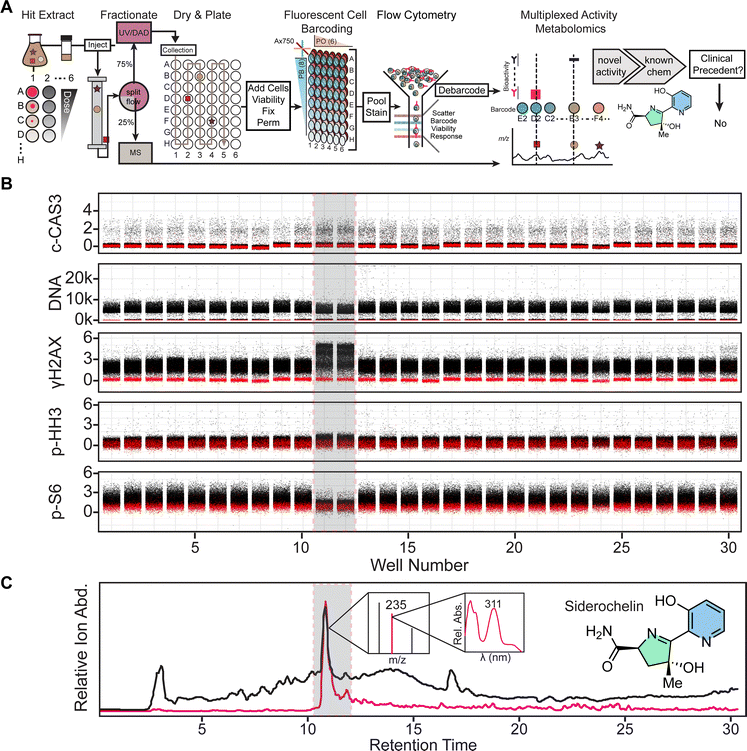
Debarcoded cell populations were evaluated for extract-specific, dose-dependent phenotypic shifts in a five-marker fluorescent panel after exclusion of non-viable cells (Fig. 1A). This allowed for determination if a given extract was active before metabolomic fractionation. Extracts exhibiting activity were fractioned by reverse phase split-flow high performance liquid coupled to polarity switching electrospray mass spectrometric and diode array UV/vis analyzer detection (HPLC/MS/UV) (Fig. 1B and 3A) and collected in 96-well plates. Chromatography is used to separate metabolites into fractions which can be correlated back to their unique mass. Fractions and associated chromatographic features were matched to well plates (Fig. 1B and 3A). Fractions and cells were processed for stimulation and flow cytometry as described above and debarcoded to examine well-specific bioactivity of each fluorescent reporter (Fig. 1B). Bioactive wells were tracked to MS/UV data to infer well-isolated ions and absorbance spectra (Fig. 1B). Bioactive lead secondary metabolites were isolated by varying chromatographic techniques (size exclusion, preparative HPLC, silica plates, etc.) and compared to known compound databases with nuclear magnetic resonance (NMR) and high-resolution mass spectrometry (HRMS), as described in the methods section. Thus, MAM affords an activity-driven approach for discovery of new and known bioeffector scaffolds while simultaneously discovering new uses of previously discovered metabolites, albeit ones potentially not prioritized for future study, ostensibly due to insufficiently interesting biological activity as measured.
MAP and MAM identify siderochelin induced injury via perturbations to phospho-signaling in DNA damage and translation response markers
Fig. 2C illustrates a single cell dose response Rothko plot of extracts from stimulated A. coloradensis subsp. ossmiensis (Fig. S5 and S6). The phenotypic marker for DNA damage responses, γH2AX,68 exhibited elevated single cell fluorescent shifts in a dose-dependent manner across stimuli. Additionally, dose-dependent p-S6 suppression was demonstrated across stimuli. In all, this single cell data indicated the producer synthesized metabolites across chemical and biological stimulus that affects DNA damage response68 and nutrient signaling and response69–71 in the MV-4-11 cell line. We confirmed these interpretations by comparing experimental samples to known bioeffector natural product positive controls31 (Fig. S7), included as internal controls on barcoded plates (Fig. 2C).Fig. 3B and C illustrate MAM data, which correlate single cell marker response data to associated mass spectra and UV/Vis absorbance. MAM of a hit extract from A. coloradensis subsp. ossmiensis (Fig. 2) revealed an isolated increase of γH2AX and decrease of p-S6 fluorescent intensities in wells 11 and 12 (Fig. 3B) consistent with preliminary screening data (Fig. 2). Extract mass chromatograms highlight an abundant total ion current peak residing across retention times 10–12 minutes (Fig. 3C). A number of compounds map to a single well; therefore, candidate m/z ion currents were extracted to ascertain secondary metabolites in wells consistent with bioactivity data and then inspected for UV/Vis absorbance. The candidate bioactive compounds corresponded to an extracted ion current of m/z ∼235, which aligned to wells 11 and 12 with an UV/Vis absorbance maximum of ∼311 nm (Fig. 3C).
The bioactive compound responsible for phenotypic shifts in Fig. 3B was purified by combination of fractionation via Sephadex LH-20 size exclusion chromatography and preparative reversed phase HPLC (See Methods), then dereplicated to the previously described natural product siderochelin34 by UV/Vis, m/z (Fig. 3C) and proton NMR (Fig. S1). The siderochelin family of compounds was first isolated from Nocardia and originally characterized as a ferrous ion chelating agent with weak antibiotic activity.34 Additional analogs with similar metal chelating properties have been isolated; however, antibiotic properties are reported to be modest.35,72 To our knowledge, this is the first report of any siderochelin analog eliciting diverse molecular changes in mammalian cancer cell model signaling associated with DNA damage and ribosomal protein S6 phosphorylation. As such, we decided to test siderochelin's nascent biological activity in primary AML patient samples (Fig. S8). A primary bone marrow aspirate and leukapheresis sample was challenged with increasing concentrations of siderochelin to determine if injury pathways observed in the MV-4-11 cell model translated to primary samples. Viable, singlet cells were separated by scatter properties and CD45 expression as described previously.31 The most notable impact was a reduction in Ki67 within blast and healthy (lymphoid and myeloid) subsets (Fig. S8) and decreased p-S6 exclusively in leukapheresis blast cells; a phenotype that was consistent with the MV-4-11 model cell line. No notable changes were observed for bone marrow aspirates. Taken together this demonstrates how using the MV-4-11 model cell line can capture molecular phenotypes that can reflect responses of primary samples; however, it is not a “catch-all” as bone-marrow aspirates, which represent a different leukemic niche, did not respond to siderochelin challenge. The limited range of biological impact on primary samples, alongside a lacking literature precedent for clinical development, enabled rapid and rational decision making to discontinue investigating siderochelin as a lead compound.
M. phytophila ssp. tarocco synthesizes anthracyclines that induce marked DNA content increases not observed in clinical anthracyclines
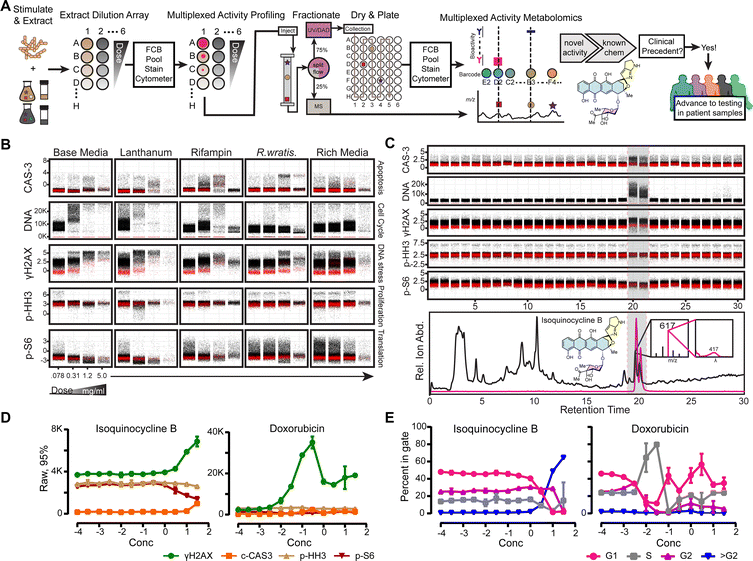
In the second case study, M. phytophila ssp. tarocco microbial extract analyzed by MAP/MAM workflows with MV-4-11 cells (Fig. 4A) exhibit robust, phenotypic, dose-dependent changes across multiple cell status markers (Fig. 4B). Most notably, the DNA binding-dependent fluorescence shift was notable, and increased well beyond typical G2 locked cell fluorescent profiles (Fig. 4B) marked by nocodazole internal positive control (Fig. S7). The metabolite responsible for this phenotype was identified via MAM in wells 20 and 21, consistent with additional γH2AX, c-CAS3, and p-S6 fluorescent shifts (Fig. 4C), as observed in extract screening (Fig. 4B). Extracted positive ion currents indicated candidate metabolite of m/z ∼617 with local UV absorbance of maximum of ∼417 nm (Fig. 4C). The producing bacterium was scaled up for isolation of the bioactive component in base media since this was the optimal condition for biosynthesis of target compound (Fig. S9). The HP20-adsorbed acetone extract from scaled cultures was purified by size-exclusion, preparative, and analytical chromatographic methods. Based upon mass spectrometric and NMR analysis, the structure of the isolated bioactive component was determined to be isoquinocycline B36,37 by NMR (Fig. S2) and HRMS (Fig. 4).Isoquinocycline B is the stereoisomer of kosinostatin and both were previously characterized as weak, Gram positive antibiotics36 with kosinostatin noted to have cytotoxicity against various mammalian cell lines.37 Functional and/or mechanistic characterization of kosinostatin and isoquinocycline B in mammalian systems have not been pursued, perhaps due to the assumption of mechanistic similarity to the daunorubicin/doxorubicin family of natural products. Both molecules maintain an anthracyclinone core, (Fig. 5A) which would suggest a similar mechanism for these molecules involving DNA intercalation and topoisomerase II inhibition,5,73 however it remained unclear if both compounds would engender similar phenotypes in the MV-4-11 cell line. Given the clinical importance of the anthracycline doxorubicin in AML, we assayed isoquinocycline B alongside doxorubicin to compare the phenotypes induced in the MV-4-11 cell line. We incubated MV-4-11 cells for 16 hours with twelve doses of each compound. Isoquinocycline B enhanced γH2AX and c-CAS3 but suppressed p-S6 phosphorylation only at high concentrations (>1 µM) (Fig. 4D). DNA content was measured by fluorescence intensity of YoPro where cells in G1 are YoPro low, Cells in S phase are YoPro medium, and cells in G2 are YoPro high. We also use aphidicolin and nocodazole controls whose mechanisms of action lock cells in G1 or G2 respectively.74,75 MV-4-11 challenged with isoquinocycline B exhibited DNA fluorescence intensity shift consistent with an increase in cellular DNA content beyond G2 as defined by nocodazole controls (Fig. 4E and Fig. S7). Moreover, we confirmed that enhanced YoPro signal occurred with respect to cell doubling times, greater than ten hours, rather than intrinsic fluorescence of the anthracyclines rapidly incorporated, within 30 minutes, into DNA (Fig. S10). In contrast, doxorubicin promoted a robust γH2AX response (Fig. 4D) with minimal impact on translation or apoptosis and depressed DNA fluorescent intensity shifts to sub-G1 (<2N) levels (Fig. 4E). To our knowledge, this is the first observation of such distinctly different single cell molecular phenotype induction by anthracyclinone pharmacophore family compounds.
Mass cytometric assessment of anthracyclines on cellular responses from AML patient samples compared to PBMCs
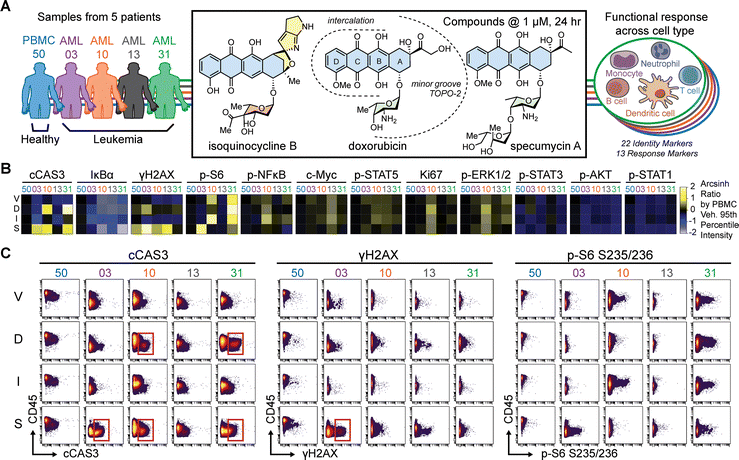
The anthracycline family of molecules is widely prescribed across multiple clinical indications, particularly in acute lymphocytic and myelogenous leukemias.3,76 For their primary mechanism of action, anthracyclines bind DNA and stabilize a topoisomerase II complex with cleaved DNA, preventing replication and inducing cell death.3,77 Anthracycline treatment is also associated with off-target side-effects arising from a variety of other mechanisms,76,78 as well as mutagenicity potentially leading to secondary cancers79 and resistance occurs in refractory or relapsing leukemias.80,81 For their primary mode of action, anthracyclines are hypothesized to possess two pharmacophoric regions.82 The tricyclic anthraquinone core intercalates into DNA82 and the A-ring substitutions, including the glycosyl moiety, face the minor groove and interact with topoisomerase in the ternary complex (Fig. S11).To observe the effects of chemical functionalization of the A-ring in single cell subsets from heterogeneous primary patient samples, we incubated a healthy peripheral blood mononuclear cell sample (PBMC50) and four AML leukapheresis patient samples (AML03, 10, 13, and 31) (Table S1) with 1 µM of isoquinocycline B, doxorubicin, specumycin A1, a glycovariant of daunomycin previously identified using MAM,31 and a vehicle control for 24 hours (Fig. 5A). For analysis by time-of-flight mass cytometry (CyTOF), cells were stained with cisplatin for viability, fixed, permeabilized, and barcoded with a total of 35 heavy-metal conjugated antibodies (See Methods). In all, twenty-two cell surface identity markers were used to identify AML blast subsets and non-AML peripheral blood mononuclear cells, alongside thirteen intracellular cell status markers (Fig. 5A and Table S3).
Intracellular response markers across all cells in treated patient samples were initially quantified and visualized as heatmaps (Fig. 5B). Bulk PBMC exhibited some γH2AX response to specumycin but otherwise appeared largely insensitive to anthracycline challenge (Fig. 5B). The largest variation in effects between patients and treatments was seen for c-CAS3, a marker of apoptosis, IκBα, an immunomodulation marker,83 γH2AX, a readout of the DNA damage response, and p-S6, part of central nutrient sensing pathways affecting translation (Fig. 5B). Differences in responses between patient samples are readily ascertainable. Doxorubicin and specumycin activated c-CAS3 in AML10 and AML31 but not in AML13. In contrast, isoquinocycline B did not activate c-CAS3 changes in any of the patient samples (Fig. 5B). All three anthracyclines demonstrated decreased detection of IκBα, an inhibitor of the stress signaling NF-κB transcription factor, in all leukemia samples except for specumycin in AML03 (Fig. 5B). Whereas specumycin activated γH2AX in all samples except AML31, isoquinocycline B did not demonstrate a pronounced DNA damage response in any of the samples. Lastly, doxorubicin and isoquinocycline B reduced p-S6 in only AML03 and AML13, while specumycin reduced p-S6 in AML10. In summary, while some patient sample differential responses may be due to potency at these single dose studies, despite sharing a similar anthracyclinone core, the three anthracyclines demonstrate AML patient sample differential effects for markers of apoptosis, immunomodulation, DNA damage response, and modulation of nutrient-dependent signaling translation.
Heterogeneous signaling responses to three anthracyclines are revealed across patient samples
Typically, mature lymphocytes from healthy PBMC highly express CD45 protein, whereas CD45 is generally low in leukemic blast cells and stem and progenitor cells.84,85 Consistent with these observations, PBMC50 cells had higher overall CD45 expression than the four AML samples (Fig. 5C and 6A). Analysis of anthracycline-dependent cellular responses, particularly injury associated responses, reveals markedly divergent responses among patients (Fig. 5C). For example, samples AML03, 10, and 31, but not 13, demonstrate apoptotic signaling via enhanced c-CAS3 on specumycin exposure while only samples AML10 and 31 demonstrate similar responses with doxorubicin (Fig. 5C, red boxes). In contrast, only AML03 exhibited a strong DNA stress response via γH2AX (Fig. 5C, red box). The cells responsible for increased marker intensity in Fig. 5C were generally CD45 low, or likely to be blasts; this provided supporting evidence that the anthracyclines were largely activating regulated cell death pathways targeting leukemia cell populations (Fig. 5C).Cell subset responses vary across patient samples
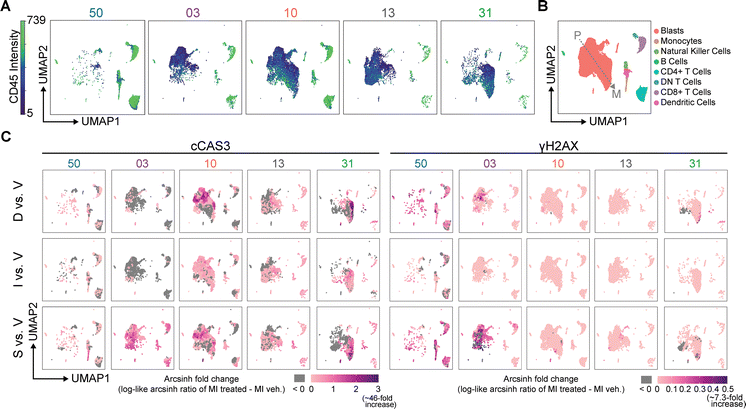
To further identify the distinct cell populations responding to anthracyclines, UMAP was performed to project cellular identities inferred from the 22 identity markers among all five patient samples into two-dimensional space for visualization and immunophenotyping.86 The UMAP plots of the five human samples (1 PBMC, 4 AML) were subdivided based on patient sample of origin to visualize differences in cell populations across patients (Fig. 6A and Fig. S12 and S13). Healthy cell subsets such as B and T cells, despite their low abundance in AML patient samples, clustered in UMAP regions consistent with healthy PBMC and clearly separated from blast cells, demonstrating the ability of this approach to identify blast cells unique to patients while also capturing canonically healthy subsets (Fig. 6B and Fig. S14). Moreover, UMAP blast cell embeddings captured interpatient and intratumor heterogeneity by distinguishing primitive and mature blasts (Fig. 6B and Fig. S13). Based on high CD34 and low to medium CD38 intensity, the left side of the blast populations (Fig. 6B) can be considered more primitive AML; the high CD33 and CD123 on the right side of the blast populations generally correlate to more mature blast cells (Fig. 6B and Fig. S12).87 The four patient samples tested here represented a spectrum of possessing mostly primitive blasts (AML03 and AML13), mostly mature blasts (AML31) and a combination of both (AML10) (Fig. 6A and Fig. S12 and S13). Therefore, mass cytometry allows interrogation of small molecule effects on more primitive blast cells and mature blast cells, which correspond to distinct disease biology.To further illustrate this point, we quantified the patient molecular phenotypes of cell populations undergoing induction of apoptosis and/or a DNA damage response following anthracycline treatment as determined by signal intensity of c-CAS3 and γH2AX from the UMAP clustering (Fig. 6A). Specifically, the fold change in median signal intensity for each anthracycline vs. vehicle was calculated using K-nearest neighbors (KNN). This visualization highlighted the heterogeneity across both patient and drug in response to anthracycline treatment as compared with vehicle (Fig. 6C).45 For example, PBMC50 cells incubated with anthracyclines exhibited limited apoptosis and demonstrated a uniform DNA damage response across cell populations induced by doxorubicin and more prominently specumycin A1 (Fig. 6C), validating the findings from the biaxial plots in Fig. 5C. Among the populations with the highest γH2AX activation by specumycin A in PBMC50 cells were the CD4+ T cells, with a median arcsinh fold change in γH2AX of 0.255 and the natural killer cells, dendritic cells, and monocytes, which had a median arcsinh fold change of 0.323 (Fig. S15). AML03 treated with specumycin exhibited an increased c-CAS3 and γH2AX signature in blast cells and healthy cell subsets whereas doxorubicin and isoquinocycline were relatively inert (Fig. 6C). In addition, marked γH2AX activation was seen within the AML03 blast cell subset by specumycin (95th percentile: 0.773), compared with isoquinocycline B (95th percentile: 0.00), and doxorubicin (95th percentile: 0.328). The blasts of AML10 showed a c-CAS3 response to all three anthracyclines; the highest arcsinh fold change in c-CAS3 within the blast cells was induced by doxorubicin (95th percentile: 1.98), followed by specumycin (95th percentile: 1.60), and finally isoquinocycline (95th percentile: 0.54) (Fig. 6C and Fig. S15). This is consistent with the CD45 low, c-CAS3 high regions observed in AML10 in response to specumycin and doxorubicin treatment seen in Fig. 5C. The replot in Fig. 6C illustrates that only a small subset of primitive blasts had a strong apoptotic response to doxorubicin and specumycin A (darkest purple regions), with this subset having a median arcsinh fold change in c-CAS3 of 1.68 in response to doxorubicin and of 1.17 in response specumycin A1. AML13 exhibited relatively weak c-CAS3 and γH2AX shifts compared to the other AML samples (Fig. 6C). Finally, AML31 demonstrated positive c-CAS3 responses to all three anthracyclines; interestingly, doxorubicin and specumycin A impacted different blast cell regions for c-CAS3 but not γH2AX. The c-CAS3 high region within the blast cells of AML31 (darkest purple regions) had a higher median arcsinh fold change of 2.57 in response to doxorubicin as compared with specumycin A1, which had a median arcsinh fold change of 1.76 (Fig. S15).
DBSCAN was applied to the areas of the “D vs. V” and “S vs. V” UMAPs in the top quartile of c-CAS3 fold change and subsequently subjected to Marker Enrichment Modeling (MEM) (Fig. S16). MEM quantifies contextual protein enrichment on a scale of 0 (no enrichment) to 10 (highest possible enrichment).88 Subtracting the doxorubicin MEM label from the specumycin MEM label (“ΔMEM”),89 illuminated the prominent phenotypic difference between the two blast cell regions was in enrichment of CD123, a surface marker overexpressed in many hematologic malignancies.90
In summary, the mass cytometric data illustrate contrasting responses across and within patient samples to three structurally divergent anthracyclines that may have been previously assumed to be mechanistically similar, based on conservation of a core anthracyclinone aglycon scaffold. The structural changes presented by the variable A-ring substitutions and glycovariants, engender differential cellular subset responses within samples and between patients. Taken together, this suggests that decorations to the anthracyclinone pharmacophore in anthracyclines, which face the DNA-enzyme complex, can have markedly different effects on DNA functions resulting in differential cellular responses.
Discussion
Herein, we applied a framework for secondary metabolite discovery that employs multiplexed molecular phenotyping in single cells to identify bioactivities of interest in crude extracts (MAP), prioritizes compounds for isolation and structural elucidation (MAM), and then uses those compounds in primary cells to integrate the analysis of novel bioactivities for assessment of patient-specific responses (mass cytometry). In this way, we aim to provide lead molecules, comprising known or new compounds, with preclinical insights that exceed the standard of traditional hit-to-lead development pipelines.We provide two case studies that utilize MAP and MAM to rapidly identify bioactive secondary metabolites in two organisms by stimulating them with five chemical and biological stimuli that generated lead extracts based on dose-dependent marker responses for MAM. In the first case study, we identified siderochelin (Fig. 2 and 3) via previously undescribed γH2AX induction, providing new insights into what was formerly assigned as an antimicrobial siderophore.34 In the second case study, we identified isoquinocycline B via enhanced DNA fluorescence in addition to stimulating c-CAS3 (Fig. 4). These activities are previously unreported for these two known, but not commercially available molecules, expanding the interest in understanding their utility. Since isoquinocycline B belonged to the clinically impactful anthracycline family of compounds and demonstrated an unexpected molecular phenotype, we selected it to be the subject of an extended study in the application of single cell deep cellular profiling in primary cells (Fig. 5 and 6). In this assessment, we illustrate the potential of mass cytometric deep cell profiling to distinguish the effects of structural differences in the two pharmacophoric regions in the anthracycline class.
Like many highly successful natural products in the clinic, anthracycline family compounds target a central cellular process – topoisomerase 2-catalyzed DNA unwinding during replication. The therapeutic strategy of targeting a central function stands in contrast with targeted approaches, for example discovery that targets oncogenic kinases, which offer the hope of higher therapeutic index, but can also be compensated by rapid evolution of resistance in cancer. Despite their toxicity, anthracyclines have been widely used in various chemotherapeutic strategies for over 50 years and are an indispensable component of regimens in hematological malignancies.91 For example, ‘7 + 3’ therapy of daunomycin + cytarabine is a staple of AML therapy but many patients are refractory to this regimen or eventually relapse, thus identifying alternative treatment modalities is critical. The anthracycline therapeutic index is thought to arise from the increased sensitivity of rapidly proliferating cancer cells in comparison to normal cells.92 The anthracyclinone ring intercalates into DNA, and the A-ring and daunosamine sugar interact with the minor groove and topoisomerase 2,93 leading to DNA double-strand breaks, resulting in the formation of a drug-DNA-enzyme ternary complex.73 The interaction of anthracyclines with both DNA and topoisomerase has been extensively studied and leveraged to enhance topoisomerase interactions; for example, the glycosyl moiety has been identified as a structural determinant for topoisomerase interaction and as such, modifications to this region of the molecule have been attempted with some success.77,94 In this context, it is not entirely surprising that isoquinocycline B, doxorubicin, and specumycin A1 appear to induce differences in apoptosis, immunomodulation, DNA damage response, and nutrient response signaling (Fig. 4) given their different glycosylation and anthracycline core scaffold substituents (Fig. 5A). However, to our knowledge this is the first study to demonstrate functional differences in naturally occurring anthracycline analogs through investigating primary samples across multiple functional and hematopoietic identity markers. Of the three anthracyclines, specumycin differs by glycosylation on the daunosamine sugar with an additional acetal moiety, and isoquinocycline via a significant modification to the anthracycline core scaffold, a spirocyclic pyrrolopyrrole. The additional acid sensitive acetal functional group on daunosamine in specumycin reflects increased potency across several patient samples and cell subsets.
Intratumor heterogeneity is highly prevalent in AML (Fig. S13) and leukemia cells can be organized into a hierarchy from primitive to mature based on normal cell development.95 In our small cohort, the samples with more mature blast cells (AML10 and AML31) and IDH1 mutations were commonly sensitive to both doxorubicin and specumycin, and perhaps more sensitive to the three anthracyclines compared to AML03 and AML13. Taken together, multiplexed, single cell experimental and analysis approaches revealed that these anthracyclines elicited distinct, patient dependent functional outcomes. This approach may be beneficial in other heterogeneous tumor types to help identify specific cell subsets sensitive to different secondary metabolites.
Future studies will investigate if structural variations in anthracyclines present a potential therapeutic advantage in treatment of leukemia. There are hundreds of anthracycline natural products reported comprising many core anthraquinone modifications and A-ring substitutions with an unknown potential for modification of enzyme client binding to the DNA complex.96 The abundant structure-activity relationships (SAR) present in naturally occurring anthracyclines is typical among natural product classes. Using MAP to perform deep cell profiling in primary samples can prioritize important SAR and help stratify patients for successful therapy for a given molecule.
Conclusion
To realize the untapped potential of microbes for producing metabolites with translational potential in humans, we applied a cytometric platform for accelerating discovery of bioactive microbial metabolites and mapping them to functional roles in human primary cells. Multiplexed activity profiling (MAP) of microbial extracts was used to identify chemical and biological stimuli capable of inducing bacteria to produce metabolites affecting mammalian cell functional marker perturbation. Multiplexed activity metabolomics (MAM) identified the functionally perturbing components of active extracts prior to isolation, revealing new potential mechanisms for new and known metabolites. After isolation, patient-level deep cellular responses of primary cells were quantified via mass cytometric analysis and categorized via machine learning. Together, these methods can accelerate the discovery of conditions for production and identification of metabolite families, and structure-activity relationships within them, with previously unrecognized human translational potential.Author contributions
J. B. Experimental design, screening campaign, data acquisition and analysis, figure generation, manuscript writing and revision; H. T. Data analysis, figure generation, manuscript writing and revision; K. P. Screening campaign, data acquisition and analysis; J. F. Screening campaign, data acquisition and analysis; B. R. Experimental design, screening campaign; S. L. Screening campaign, data analysis; M. H. Screening campaign, data acquisition; C. P. Data acquisition; J. I. Screening campaign, project conception, support, and supervision; B. F. Screening campaign, project conception, support, and supervision, manuscript writing and revision; B. B. Screening campaign, project conception, support, and supervision, manuscript writing and revision.Conflicts of interest
The authors have no conflict of interest to declare.Data availability
The FCS files used for generating all figures in the main manuscript and supplemental information are deposited at Zenodo and publicly available at the following URL: https://doi.org/10.5281/zenodo.16333899. Supporting data for the article is included in the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5cb00203f.Acknowledgements
The authors are grateful to the VUMC flow cytometry core for instrument training and scheduling; the VUMC cell imaging shared resource for SEM imaging; and the small molecule NMR core for instrument training and scheduling. The authors thank Claire Cross for assistance with Sconify. We gratefully acknowledge Mr. Lonnie Carr for permission to collect microorganisms from Blue Spring Cave. Research was supported by the following funding resources: NIH and NCI grants R01 CA226833-07, T32 GM065086, T32 GM007628-43, and U01 TR002625-04.References
- D. J. Newman, Natural products and drug discovery, Natl. Sci. Rev., 2022, 9, 11 Search PubMed.
- W. H. Organization, The selection and use of essential medicines: report of the WHO Expert Committee on Selection and Use of Essential Medicines, 2023 (including the 23rd WHO Model List of Essential Medicines and the 9th WHO Model List of Essential Medicines for Children), World Health Organization, 2024 Search PubMed.
- K. Krohn, Anthracycline chemistry and biology/volume, ed K. Krohn, Springer, Berlin, 2008 Search PubMed.
- S. F. A. Rizvi, S. Tariq and M. Mehdi, Anthracyclines: mechanism of action, classification, pharmacokinetics and future–a mini review, Int. J. Biotech. Bioeng., 2018, 4(4), 81–85 Search PubMed.
- D. Gewirtz, A critical evaluation of the mechanisms of action proposed for the antitumor effects of the anthracycline antibiotics adriamycin and daunorubicin, Biochem. Pharmacol., 1999, 57(7), 727–741 CrossRef CAS PubMed.
- J. Chu, X. Vila-Farres and S. F. Brady, Bioactive Synthetic-Bioinformatic Natural Product Cyclic Peptides Inspired by Nonribosomal Peptide Synthetase Gene Clusters from the Human Microbiome, J. Am. Chem. Soc., 2019, 141(40), 15737–15741 CrossRef CAS PubMed.
- M. H. Medema, T. de Rond and B. S. Moore, Mining genomes to illuminate the specialized chemistry of life, Nat. Rev. Genet., 2021, 22(9), 553–571 Search PubMed.
- J. Chu, X. Vila-Farres, D. Inoyama, M. Ternei, L. J. Cohen and E. A. Gordon, et al., Discovery of MRSA active antibiotics using primary sequence from the human microbiome, Nat. Chem. Biol., 2016, 12(12), 1004–1006 Search PubMed.
- H. A. Iqbal, L. Low-Beinart, J. U. Obiajulu and S. F. Brady, Natural Product Discovery through Improved Functional Metagenomics in Streptomyces, J. Am. Chem. Soc., 2016, 138(30), 9341–9344 CrossRef CAS PubMed.
- J. G. Owen, Z. Charlop-Powers, A. G. Smith, M. A. Ternei, P. Y. Calle and B. V. Reddy, et al., Multiplexed metagenome mining using short DNA sequence tags facilitates targeted discovery of epoxyketone proteasome inhibitors, Proc. Natl. Acad. Sci. U. S. A., 2015, 112(14), 4221–4226 Search PubMed.
- N. Ziemert, M. Alanjary and T. Weber, The evolution of genome mining in microbes - a review, Nat. Prod. Rep., 2016, 33(8), 988–1005 RSC.
- B. O. Bachmann, S. G. Van Lanen and R. H. Baltz, Microbial genome mining for accelerated natural products discovery: is a renaissance in the making?, J. Ind. Microbiol. Biotechnol., 2014, 41(2), 175–184 CrossRef CAS PubMed.
- L. Laureti, L. Song, S. Huang, C. Corre, P. Leblond and G. L. Challis, et al., Identification of a bioactive 51-membered macrolide complex by activation of a silent polyketide synthase in Streptomyces ambofaciens, Proc. Natl. Acad. Sci. U. S. A., 2011, 108(15), 6258–6263 CrossRef CAS PubMed.
- C. S. McCaughey, J. A. van Santen, J. J. J. van der Hooft, M. H. Medema and R. G. Linington, An isotopic labeling approach linking natural products with biosynthetic gene clusters, Nat. Chem. Biol., 2022, 18(3), 295–304 CrossRef CAS PubMed.
- R. D. Kersten, Y. L. Yang, Y. Xu, P. Cimermancic, S. J. Nam and W. Fenical, et al., A mass spectrometry-guided genome mining approach for natural product peptidogenomics, Nat. Chem. Biol., 2011, 7(11), 794–802 CrossRef CAS PubMed.
- T. Beites and M. V. Mendes, Chassis optimization as a cornerstone for the application of synthetic biology based strategies in microbial secondary metabolism, Front. Microbiol., 2015, 6, 906–915 Search PubMed.
- J. Liu, X. Wu, M. Yao, W. Xiao and J. Zha, Chassis engineering for microbial production of chemicals: from natural microbes to synthetic organisms, Curr. Opin. Biotechnol, 2020, 66, 105–112 CrossRef CAS PubMed.
- L. K. Caesar, M. T. Robey, M. Swyers, Md N. Islam, R. Ye and P. P. Vagadia, et al., Heterologous Expression of the Unusual Terreazepine Biosynthetic Gene Cluster Reveals a Promising Approach for Identifying New Chemical Scaffolds, mBio, 2020, 11(4), e01691-20 CrossRef PubMed.
- J. Kormanec, B. Rezuchova, D. Homerova, D. Csolleiova, B. Sevcikova and R. Novakova, et al., Recent achievements in the generation of stable genome alterations/mutations in species of the genus Streptomyces, Appl. Microbiol. Biotechnol., 2019, 103(14), 5463–5482 CrossRef CAS PubMed.
- A. C. Ross, L. E. S. Gulland, P. C. Dorrestein and B. S. Moore, Targeted Capture and Heterologous Expression of the Pseudoalteromonas Alterochromide Gene Cluster in Escherichia coli Represents a Promising Natural Product Exploratory Platform, ACS Synth. Biol., 2015, 4(4), 414–420 CrossRef CAS PubMed.
- K. Scherlach and C. Hertweck, Triggering cryptic natural product biosynthesis in microorganisms, Org. Biomol. Chem., 2009, 7(9), 1753–1760 RSC.
- P. J. Rutledge and G. L. Challis, Discovery of microbial natural products by activation of silent biosynthetic gene clusters, Nat. Rev. Microbiol., 2015, 13(8), 509–523 CrossRef CAS PubMed.
- Y. Imai, S. Sato, Y. Tanaka, K. Ochi and T. Hosaka, Lincomycin at Subinhibitory Concentrations Potentiates Secondary Metabolite Production by Streptomyces spp, Appl. Environ. Microbiol., 2015, 81(11), 3869–3879 CrossRef CAS PubMed.
- B. C. Covington, J. A. McLean and B. O. Bachmann, Comparative mass spectrometry-based metabolomics strategies for the investigation of microbial secondary metabolites, Nat. Prod. Rep., 2017, 34(1), 6–24 RSC.
- J. T. Froese, J. A. Balsamo, B. J. Reisman, S. M. Barone, J. M. Irish and B. O. Bachmann, Multiplexed activity metabolomics for isolation of filipin macrolides from a hypogean actinomycete, J. Antibiot., 2025, 78(2), 78–89 CrossRef CAS PubMed.
- C. Covington Brett, M. Spraggins Jeffrey, E. Ynigez-Gutierrez Audrey, B. Hylton Zachary and O. Bachmann Brian, Response of Secondary Metabolism of Hypogean Actinobacterial Genera to Chemical and Biological Stimuli, Appl. Environ. Microbiol., 2018, 84(19), e01125 Search PubMed.
- J. T. Wibowo, A. Bayu, W. D. Aryati, C. Fernandes, A. Yanuar and A. Kijjoa, et al., Secondary Metabolites from Marine-Derived Bacteria with Antibiotic and Antibiofilm Activities against Drug-Resistant Pathogens, Marine Drugs, 2023, 21(1), 50 CrossRef CAS PubMed.
- A. S. Walker and J. Clardy, A Machine Learning Bioinformatics Method to Predict Biological Activity from Biosynthetic Gene Clusters, J. Chem. Inf. Model., 2021, 61(6), 2560–2571 CrossRef CAS PubMed.
- C. M. F. Ancajas, A. S. Oyedele, C. M. Butt and A. S. Walker, Advances, opportunities, and challenges in methods for interrogating the structure activity relationships of natural products, Nat. Prod. Rep., 2024, 41(10), 1543–1578 RSC.
- M. A. Skinnider, C. W. Johnston, M. Gunabalasingam, N. J. Merwin, A. M. Kieliszek and R. J. MacLellan, et al., Comprehensive prediction of secondary metabolite structure and biological activity from microbial genome sequences, Nat. Commun., 2020, 11(1), 6058 CrossRef CAS PubMed.
- D. C. Earl, P. B. Ferrell, N. Leelatian, J. T. Froese, B. J. Reisman and J. M. Irish, et al., Discovery of human cell selective effector molecules using single cell multiplexed activity metabolomics, Nat. Commun., 2018, 9(1), 39 CrossRef PubMed.
- M. H. Woehrmann, W. M. Bray, J. K. Durbin, S. C. Nisam, A. K. Michael and E. Glassey, et al., Large-scale cytological profiling for functional analysis of bioactive compounds, Mol. BioSyst., 2013, 9(11), 2604–2617 CrossRef CAS PubMed.
- J. A. Balsamo, K. E. Penton, Z. Zhao, M. J. Hayes, S. M. Lima and J. M. Irish, et al., An immunogenic cell injury module for the single-cell multiplexed activity metabolomics platform to identify promising anti-cancer natural products, J. Biol. Chem., 2022, 298(9), 102300 CrossRef CAS PubMed.
- W. C. Liu, S. M. Fisher, J. S. Wells Jr., C. S. Ricca, P. A. Principe and W. H. Trejo, et al., Siderochelin, a new ferrous-ion chelating agent produced by Nocardia, J. Antibiot., 1981, 34(7), 791–799 CrossRef CAS PubMed.
- C.-H. Lu, F.-W. Ye and Y.-M. Shen, Siderochelins with anti-mycobacterial activity from Amycolatopsis sp. LZ149, Chin. J. Nat. Med., 2015, 13(1), 69–72 CrossRef CAS PubMed.
- Y. Igarashi, H. Higuchi, T. Oki and T. Furumai, NMR analysis of quinocycline antibiotics: structure determination of kosinostatin, an antitumor substance from Micromonospora sp. TP-A0468, J. Antibiot., 2002, 55(2), 134–140 CrossRef CAS PubMed.
- T. Furumai, Y. Igarashi, H. Higuchi, N. Saito and O. T. Kosinostatin, A quinocycline antibiotic with antitumor activity from Micromonospora sp. TP-A0468, J. Antibiot., 2002, 55(2), 128–133 CrossRef CAS PubMed.
- H. M. Ma, Q. Zhou, Y. M. Tang, Z. Zhang, Y. S. Chen and H. Y. He, et al., Unconventional origin and hybrid system for construction of pyrrolopyrrole moiety in kosinostatin biosynthesis, Chem. Biol., 2013, 20(6), 796–805 CrossRef CAS PubMed.
- Z. Zhang, Y. K. Gong, Q. Zhou, Y. Hu, H. M. Ma and Y. S. Chen, et al., Hydroxyl regioisomerization of anthracycline catalyzed by a four-enzyme cascade, Proc. Natl. Acad. Sci. U. S. A., 2017, 114(7), 1554–1559 Search PubMed.
- M. J. S. Asmaa, H. A. Al-Jamal, A. R. Hussein, B. H. Yahaya, R. Hassan and F. A. Hussain, et al., Transcriptomic Profiles of MV4-11 and Kasumi 1 Acute Myeloid Leukemia Cell Lines Modulated by Epigenetic Modifiers Trichostatin A and 5-Azacytidine, Int. J. Hematol. Oncol. Stem Cell Res., 2020, 14(1), 72–92 Search PubMed.
- P. B. Ferrell Jr., K. E. Diggins, H. G. Polikowsky, S. R. Mohan, A. C. Seegmiller and J. M. Irish, High-Dimensional Analysis of Acute Myeloid Leukemia Reveals Phenotypic Changes in Persistent Cells during Induction Therapy, PLoS One, 2016, 11(4), e0153207 Search PubMed.
- M. Chalita, Y. O. Kim, S. Park, H. S. Oh, J. H. Cho and J. Moon, et al., EzBioCloud: a genome-driven database and platform for microbiome identification and discovery, Int. J. Syst. Evol. Microbiol., 2024, 74, 6 CrossRef PubMed.
- S. Kumar, G. Stecher and K. Tamura, MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets, Mol. Biol. Evol., 2016, 33(7), 1870–1874 Search PubMed.
- M. Roussel, P. B. Ferrell Jr., A. R. Greenplate, F. Lhomme, S. Le Gallou and K. E. Diggins, et al., Mass cytometry deep phenotyping of human mononuclear phagocytes and myeloid-derived suppressor cells from human blood and bone marrow, J. Leukocyte Biol., 2017, 102(2), 437–447 CrossRef CAS PubMed.
- T. J. Burns, G. P. Nolan and N. Samusik, Continuous visualization of differences between biological conditions in single-cell data, bioRxiv, preprint, 2018, 337485 DOI:10.1101/337485.
- J. M. Irish, J. H. Myklebust, A. A. Alizadeh, R. Houot, J. P. Sharman and D. K. Czerwinski, et al., B-cell signaling networks reveal a negative prognostic human lymphoma cell subset that emerges during tumor progression, Proc. Natl. Acad. Sci. U. S. A., 2010, 107(29), 12747–12754 Search PubMed.
- B. Bodenmiller, E. R. Zunder, R. Finck, T. J. Chen, E. S. Savig and R. V. Bruggner, et al., Multiplexed mass cytometry profiling of cellular states perturbed by small-molecule regulators, Nat. Biotechnol., 2012, 30(9), 858–867 Search PubMed.
- S. M. Barone, A. G. A. Paul, L. M. Muehling, J. A. Lannigan, W. W. Kwok and R. B. Turner, et al., Unsupervised machine learning reveals key immune cell subsets in COVID-19, rhinovirus infection, and cancer therapy, eLife, 2021, 10, e64653 Search PubMed.
- H. L. Thirman, M. J. Grider-Hayes, L. C. Geben, R. A. Ihrie, L. E. Brown and J. A. Porco, et al., Connecting chemical structure to single cell signaling profiles, Commun. Biol., 2025, 8(1), 1137 CrossRef CAS PubMed.
- A. D. Steele, C. N. Teijaro, D. Yang and B. Shen, Leveraging a large microbial strain collection for natural product discovery, J. Biol. Chem., 2019, 294(45), 16567–16576 Search PubMed.
- K. E. Diggins, P. B. Ferrell Jr. and J. M. Irish, Methods for discovery and characterization of cell subsets in high dimensional mass cytometry data, Methods, 2015, 82, 55–63 CrossRef CAS PubMed.
- R. Subramani and W. Aalbersberg, Culturable rare Actinomycetes: diversity, isolation and marine natural product discovery, Appl. Microbiol. Biotechnol., 2013, 97(21), 9291–9321 Search PubMed.
- P. S. Salazar-Hamm, F. E. Homan, S. A. Good, J. J. M. Hathaway, A. E. Clements and E. G. Haugh, et al., Subterranean marvels: microbial communities in caves and underground mines and their promise for natural product discovery, Nat. Prod. Rep., 2025, 42(3), 592–622 RSC.
- P. Rangseekaew and W. Pathom-Aree, Cave Actinobacteria as Producers of Bioactive Metabolites, Front. Microbiol., 2019, 10, 387 Search PubMed.
- H. K. Kumar, H. M. Gan, M. H. Tan, W. W. Eng, H. A. Barton and A. O. Hudson, et al., Genomic characterization of eight Ensifer strains isolated from pristine caves and a whole genome phylogeny of Ensifer (Sinorhizobium), J. Genomics, 2017, 5, 12–15 CrossRef PubMed.
- D. K. Derewacz, B. C. Covington, J. A. McLean and B. O. Bachmann, Mapping Microbial Response Metabolomes for Induced Natural Product Discovery, ACS Chem. Biol., 2015, 10(9), 1998–2006 Search PubMed.
- C. R. Goodwin, B. C. Covington, D. K. Derewacz, C. R. McNees, J. P. Wikswo and J. A. McLean, et al., Structuring Microbial Metabolic Responses to Multiplexed Stimuli via Self-Organizing Metabolomics Maps, Chem. Biol., 2015, 22(5), 661–670 CrossRef CAS PubMed.
- D. K. Derewacz, C. R. Goodwin, C. R. McNees, J. A. McLean and B. O. Bachmann, Antimicrobial drug resistance affects broad changes in metabolomic phenotype in addition to secondary metabolism, Proc. Natl. Acad. Sci. U. S. A., 2013, 110(6), 2336–2341 Search PubMed.
- C. T. Nguyen, D. Dhakal, V. T. T. Pham, H. T. Nguyen and J. K. Sohng, Recent Advances in Strategies for Activation and Discovery/Characterization of Cryptic Biosynthetic Gene Clusters in Streptomyces, Microorganisms., 2020, 8, 4 Search PubMed.
- K. Ochi and T. Hosaka, New strategies for drug discovery: activation of silent or weakly expressed microbial gene clusters, Appl. Microbiol. Biotechnol., 2013, 97(1), 87–98 CrossRef CAS PubMed.
- B. K. Okada and M. R. Seyedsayamdost, Antibiotic dialogues: induction of silent biosynthetic gene clusters by exogenous small molecules, FEMS Microbiol. Rev., 2017, 41(1), 19–33 CrossRef CAS PubMed.
- F. J. Reen, S. Romano, A. D. Dobson and F. O'Gara, The Sound of Silence: Activating Silent Biosynthetic Gene Clusters in Marine Microorganisms, Mar. Drugs, 2015, 13(8), 4754–4783 Search PubMed.
- P. O. Krutzik, M. R. Clutter, A. Trejo and G. P. Nolan, Fluorescent cell barcoding for multiplex flow cytometry, Current protocols in cytometry, 2011, p. 31 Search PubMed.
- K. Sukhdeo, R. I. Paramban, J. G. Vidal, J. Elia, J. Martin and M. Rivera, et al., Multiplex flow cytometry barcoding and antibody arrays identify surface antigen profiles of primary and metastatic colon cancer cell lines, PLoS One, 2013, 8(1), e53015 Search PubMed.
- B. F. Alfonso and M. Al-Rubeai, Flow Cytometry, Comprehensive Biotechnology, 2011, pp. 559–78 Search PubMed.
- B. J. Reisman, S. M. Barone, B. O. Bachmann and J. M. Irish, DebarcodeR increases fluorescent cell barcoding capacity and accuracy, Cytometry, Part A, 2021, 99(9), 946–953 Search PubMed.
- H. A. M. Schares, M. J. Hayes, J. A. Balsamo, H. L. Thirman, B. O. Bachmann and J. M. Irish, Multiplexed cytometry for single cell chemical biology, Methods in Cell Biology, Academic Press, 2023 Search PubMed.
- M. Podhorecka, A. Skladanowski and P. Bozko, H2AX Phosphorylation: Its Role in DNA Damage Response and Cancer Therapy, J. Nucleic Acids, 2010, 2010 Search PubMed.
- Y. Ma, Y. Vassetzky and S. Dokudovskaya, mTORC1 pathway in DNA damage response, Biochim. Biophys. Acta, Mol. Cell Res., 2018, 1865(9), 1293–1311 Search PubMed.
- C. Porta, C. Paglino and A. Mosca, Targeting PI3K/Akt/mTOR Signaling in Cancer, Front. Oncol., 2014, 4, 64 Search PubMed.
- D. Saleiro and L. C. Platanias, Intersection of mTOR and STAT signaling in immunity, Trends Immunol., 2015, 36(1), 21–29 Search PubMed.
- L. A. Mitscher, T. Högberg, S. D. Drake, A. W. Burgstahler, M. Jackson and B. Lee, et al., Isolation and structural determination of siderochelin C, a fermentation product of an unusual Actinomycetes sp, J. Antibiot., 1984, 37(10), 1260–1263 CrossRef CAS PubMed.
- C. Temperini, L. Messori, P. Orioli, C. Di Bugno, F. Animati and G. Ughetto, The crystal structure of the complex between a disaccharide anthracycline and the DNA hexamer d(CGATCG) reveals two different binding sites involving two DNA duplexes, Nucleic Acids Res., 2003, 31(5), 1464–1469 CrossRef CAS PubMed.
- G. Pedrali-Noy, S. Spadari, A. Miller-Faurès, A. O. Miller, J. Kruppa and G. Koch, Synchronization of HeLa cell cultures by inhibition of DNA polymerase alpha with aphidicolin, Nucleic Acids Res., 1980, 8(2), 377–387 Search PubMed.
- R. J. Vasquez, B. Howell, A. M. Yvon, P. Wadsworth and L. Cassimeris, Nanomolar concentrations of nocodazole alter microtubule dynamic instability in vivo and in vitro, Mol. Biol. Cell, 1997, 8(6), 973–985 CrossRef CAS PubMed.
- G. Minotti, P. Menna, E. Salvatorelli, G. Cairo and L. Gianni, Anthracyclines: molecular advances and pharmacologic developments in antitumor activity and cardiotoxicity, Pharmacol. Rev., 2004, 56(2), 185–229 Search PubMed.
- J. Marinello, M. Delcuratolo and G. Capranico, Anthracyclines as Topoisomerase II Poisons: From Early Studies to New Perspectives, Int. J. Mol. Sci., 2018, 19, 11 Search PubMed.
- N. R. Neuendorff, K. P. Loh, A. S. Mims, K. Christofyllakis, W. K. Soo and B. Bolukbasi, et al., Anthracycline-related cardiotoxicity in older patients with acute myeloid leukemia: a Young SIOG review paper, Blood Adv., 2020, 4(4), 762–775 Search PubMed.
- V. Brower, Tracking Chemotherapy's Effects on Secondary Cancers, JNCI: Journal of the National Cancer Institute, 2013; vol. 105(19), pp. 1421–1422 Search PubMed.
- P. Ferguson, R. K. Hills, A. Grech, S. Betteridge, L. Kjeldsen and M. Dennis, et al., An operational definition of primary refractory acute myeloid leukemia allowing early identification of patients who may benefit from allogeneic stem cell transplantation, Haematologica, 2016, 101(11), 1351–1358 CrossRef CAS PubMed.
- F. Caiado, D. Maia-Silva, C. Jardim, N. Schmolka, T. Carvalho and C. Reforco, et al., Lineage tracing of acute myeloid leukemia reveals the impact of hypomethylating agents on chemoresistance selection, Nat. Commun., 2019, 10(1), 4986 CrossRef PubMed.
- Y. Pommier, P. Pourquier, Y. Fan and D. Strumberg, Mechanism of action of eukaryotic DNA topoisomerase I and drugs targeted to the enzyme, Biochim. Biophys. Acta, Gene Struct. Exp., 1998, 1400(1), 83–106 CrossRef CAS PubMed.
- A. Oeckinghaus and S. Ghosh, The NF-kappaB family of transcription factors and its regulation, Cold Spring Harbor Perspect. Biol., 2009, 1(4), a000034 Search PubMed.
- P. S. Kala and M. Zubair, Flow Cytometry Blood Cell Identification, StatPearls, StatPearls Publishing, 2024 Search PubMed.
- B. L. Wood, M. Arroz, D. Barnett, J. DiGiuseppe, B. Greig and S. J. Kussick, et al., 2006 Bethesda International Consensus recommendations on the immunophenotypic analysis of hematolymphoid neoplasia by flow cytometry: Optimal reagents and reporting for the flow cytometric diagnosis of hematopoietic neoplasia, Cytometry, Part B, 2007, 72B(S1), S14–S22 Search PubMed.
- L. McInnes, J. Healy and J. Melville, UMAP: Uniform Manifold Approximation and Projection for Dimension Reduction. 2018 Search PubMed.
- V. Weeda, S. G. C. Mestrum and M. P. G. Leers, Flow Cytometric Identification of Hematopoietic and Leukemic Blast Cells for Tailored Clinical Follow-Up of Acute Myeloid Leukemia, Int. J. Mol. Sci., 2022, 23(18), 10529 CrossRef CAS PubMed.
- K. E. Diggins, A. R. Greenplate, N. Leelatian, C. E. Wogsland and J. M. Irish, Characterizing cell subsets using marker enrichment modeling, Nat. Methods, 2017, 14(3), 275–278 CrossRef CAS PubMed.
- A. R. Greenplate, D. D. McClanahan, B. K. Oberholtzer, D. B. Doxie, C. E. Roe and K. E. Diggins, et al., Computational Immune Monitoring Reveals Abnormal Double-Negative T Cells Present across Human Tumor Types, Cancer Immunol. Res., 2019, 7(1), 86–99 CrossRef CAS PubMed.
- H. El Achi, E. Dupont, S. Paul and J. D. Khoury, CD123 as a Biomarker in Hematolymphoid Malignancies: Principles of Detection and Targeted Therapies, Cancers, 2020, 12(11), 3087 CrossRef CAS PubMed.
- Medicinal Chemistry: A Series of Monographs. 171981. pp. ii–ii Search PubMed.
- T. Murphy and K. W. L. Yee, Cytarabine and daunorubicin for the treatment of acute myeloid leukemia, Expert Opin. Pharmacother., 2017, 18(16), 1765–1780 CrossRef CAS PubMed.
- Y. Pommier, A. Nussenzweig, S. Takeda and C. Austin, Human topoisomerases and their roles in genome stability and organization, Nat. Rev. Mol. Cell Biol., 2022, 23(6), 407–427 CrossRef CAS PubMed.
- C. A. Geisberg and D. B. Sawyer, Mechanisms of anthracycline cardiotoxicity and strategies to decrease cardiac damage, Curr. Hypertens. Rep., 2010, 12(6), 404–410 CrossRef CAS PubMed.
- A. G. X. Zeng, S. Bansal, L. Jin, A. Mitchell, W. C. Chen and H. A. Abbas, et al., A cellular hierarchy framework for understanding heterogeneity and predicting drug response in acute myeloid leukemia, Nat. Med., 2022, 28(6), 1212–1223 CrossRef CAS PubMed.
- W. E. Running, Chapman and Hall Dictionary of Natural Products on CD-ROM, J. Chem. Inf. Comput. Sci., 1993, 33(6), 934–935 Search PubMed.
Footnote |
| † Contributed equally to this manuscript. |
| This journal is © The Royal Society of Chemistry 2026 |