Open Access Article
Open Access ArticleStructure and characterisation of CMP-Kdn synthetase from the haptophyte microalgae Prymnesium parvum
Claire
Morley
 a,
Alexandra J.
Munro-Clark
a,
Ben A.
Wagstaff
a,
Irina
Ivanova
c,
Evgenia
Dubinskaya
a,
Ava
Rostock
a,
Mary
Ortmayer
a,
Colin W.
Levy
a and
Robert A.
Field
a,
Alexandra J.
Munro-Clark
a,
Ben A.
Wagstaff
a,
Irina
Ivanova
c,
Evgenia
Dubinskaya
a,
Ava
Rostock
a,
Mary
Ortmayer
a,
Colin W.
Levy
a and
Robert A.
Field
 *abc
*abc
aSchool of Chemistry and Manchester Institute of Biotechnology, The University of Manchester, 131 Princess Street, Manchester, M1 7DN, UK
bSchool of Chemistry, Pharmacy and Pharmacology, University of East Anglia, Norwich Research Park, Norwich, NR4 7TJ, UK. E-mail: r.a.field@uea.ac.uk
cIceni Glycoscience, Norwich Research Park, Norwich, NR4 7TJ, UK
First published on 25th February 2026
Abstract
Sialic acids – 9-carbon ulosonic acids – are implicated in many cell–cell and host–pathogen interactions due to their prevalent location at the non-reducing end of glycoconjugates. Sialic acids have recently been observed in microalgae, including the toxic bloom-forming Prymnesium parvum, which produces the deaminated sialic acid, ketodeoxynonulosonic acid (Kdn), through de novo biosynthesis. Here we report on the key CMP-sialic acid synthetase enzyme (CMAS), PpNeuA, which activates Kdn to its sugar nucleotide congener, CMP-Kdn. In the present study, the X-ray crystal structure of PpNeuA was determined to 1.8 Å resolution and shows that it adopts a similar overall fold to that of other sialic acid synthetase enzymes, with which it shares ca 30% amino acid sequence identity. PpNeuA specificity for Kdn is dependent upon Arg196, a hydrophilic residue that is only found in Kdn-specific sialic acid synthetases. R196L mutation switches the substrate preference of PpNeuA from Kdn to N-acetylneuraminic acid (Neu5Ac). Kinetic analysis shows that Arg196 plays both a role in substrate binding (impact on KM) and catalysis (impact on kcat). In the context of generating metabolic probes to identify the location and context (glycolipid vs glycoprotein) of Kdn in P. parvum, we also report on the ability of PpNeuA to accept both 5Az-Kdn and 9Az-Kdn as substrates.
Introduction
Sialic acids are a diverse family of acidic nine-carbon sugars that are widespread in nature; they can be highly modified, leading to at least 60 known variations.1 Most often they are found at the non-reducing terminal end of glycoconjugates (glycoproteins and glycolipids), where their exposure at the cell surface presents them to pathogens.2 In humans, the most common sialic acid is Neu5Ac, while variations at C5 lead to other sialic acids, for example, the hydroxylated variant N-glycolylneuraminic acid (Neu5Gc) in mammals other than humans and the deaminated variant Kdn in cold blooded vertebrates (Fig. 1).3 Variations at C5, C7, and C8 lead to pseudaminic acid (Pse), a non-mammalian sialic acid present in bacteria.4 3-deoxy-D-manno-octulosonic acid (Kdo) is a related eight-carbon sugar present in the cell wall polysaccharides of Gram-negative bacteria.5Human and avian flu viruses are well known to exploit sialic acids to attach and gain entry to the cell.6 Studies have also shown that Kdn may be a mediator of viral infection in another haptophyte microalgae, Emiliania huxleyi.7,8 Phylogenetically related to E. huxleyi, Prymnesium parvum is a unicellular mixotrophic haptophyte microalga that is responsible for harmful algal blooms (HABs) across the globe.9,10 These HABs result in large-scale fish kills, with devasting effects on both the local ecology and economy.11 Algal viruses and their role in the regulation of algal blooms are well studied,12 but the role of such viruses in regulation of Prymnesium blooms is less well explored.9 We previously reported that a lytic, double-stranded DNA virus, PpDNAV, contributes to the demise of P. parvum HABs in the Norfolk Broads, UK.13,14 We also showed that PpDNAV only infects those strains of Prymnesium that produce Kdn.13 Although the molecular mechanism behind viral infection remains to be understood, the possibility of PpDNAV-Kdn interactions emulating those of the well-studied influenza virus-NeuAc interactions is an interesting prospect.
Kdn has been widely found in lower vertebrates and pathogenic bacteria, as well as in mammalian tissues, cancers and microalgae.2,15 Kdn incorporation into glycoconjugates is a result of either a specific Kdn biosynthetic pathway,16,17 or through promiscuity in the Neu5Ac biosynthetic pathway.18 The first step of incorporation of sialic acids into glycoconjugates involves the activation of sialic acid to the cytidine monophosphate diester by CMP-sialic acid synthetase (CMAS) in the presence of Mg2+ (Fig. 1).19 The activated sialic acid is then incorporated into glycoconjugates by sialyltransferases.19 In a similar way, Kdo is activated by CMP-Kdo synthetase (CKS), and Pse is activated by CMP-Pse synthetase (PseF) where it is then incorporated into lipooligosaccharides (LOS) and lipopolysaccharide (LPS), or glycoproteins of pathogenic bacteria (Fig. 1).5,20 This activation strategy differs from the more common nucleoside diphosphate diester activation of other monosaccharides.21 The structures of several CMAS and CKS enzymes have been solved from both eukaryotic mammalian organisms22 and bacterial sources,23–27 recently the first structure of PseF was published.20 The X-ray crystallographic structures of CMAS, CKS and PseF show that they share a common 3D structure, and catalytic mechanism.24,26 The enzyme transitions between an open conformation, when CTP is bound, to the active closed conformation, after sialic acid binding. In the closed conformation, the catalytic arginine, and two divalent cations are positioned for catalysis and the formation of CMP-sialic acid.24
While the majority of characterised CMAS enzymes have a preference for Neu5Ac, they often show some level of substrate promiscuity.28 However, there are also examples of CMAS with a Kdn preference – e.g. from Oncorhynchus mykiss,29Danio rerio,30Bacteriodetes thetaiotamicron16 and P. parvum.17 There is limited understanding of what governs the difference in substrate specificity between CMAS enzymes, and there is currently no published X-ray crystal structure for a Kdn-specific CMAS enzyme.
The production of sialic acid glycosides is very important for fundamental research and pharmaceutical chemistry, such as for determining structure–function relationships31 and in the development of anti-influenza drugs.32 However, the chemical synthesis is demanding; therefore enzymatic syntheses are often employed. The CMAS enzymes utilised in such syntheses typically show specificity for Neu5Ac.24 Recently, Kdn has been utilised as a temporary protecting group in the efficient production of α2,6-Neu5Ac-glycoconjugates,33 highlighting the importance of and the requirement for Kdn-specific CMAS enzymes.
Herein, we present the X-ray crystal structure of the Kdn-specific CMAS enzyme, PpNeuA, to underpin greater understanding of the strong preference of this enzyme for Kdn. We analyse the molecular basis for Kdn specificity, and through single site substitution mutants demonstrate the importance of Arg196 in CMAS substrate specificity. We also assess the potential of the PpNeuA enzyme to activate azide-substituted Kdn, which is central to metabolic Kdn engineering studies.
Materials and methods
Expression of CMP-Kdn synthetase and mutants
CMP-Kdn synthetase (CAMPEP_0191219004, with transmembrane domain truncation), PpNeuA-WT, was expressed as previously described.17 PpNeuA mutants were cloned using IDT's gBlock gene fragments. The PpNeuA-WT sequence with the desired mutation and overhangs for In-Fusion™ cloning into the pOPINF vector34 were used for cloning. A full list of sequences used in this study can be found in the supporting information. gBlock fragments were cloned into pOPINF vectors using In-Fusion™ Snap Assembly kit (Takara) per the manufacturer's instructions.The wild-type and mutant enzymes were expressed in BL21 (DE3)™ cells (NEB). 1 L cultures were grown at 37 °C in Luria-Bertani (LB) medium containing 100 µg mL−1 ampicillin until mid-exponential phase. Protein expression was induced with 0.5 mM isopropyl β-D-thiogalactopyranoside, and the cells were incubated at 18 °C overnight. Cells were harvested by centrifugation and then lysed, using a cell disruptor (25 kpsi, Constant Systems) in buffer containing 50 mM Tris-HCl, pH 7.5, 0.5 M NaCl, 20 mM imidazole, protease inhibitor (SigmaFast™ Protease Inhibitor Cocktail Tables, Sigma) and 10 µg mL−1 DNase (Roche). All proteins were purified using nickel affinity chromatography with a linear gradient of imidazole from 20 mM to 500 mM. Protein sample fractions judged to be >95% pure by SDS-PAGE were pooled and dialysed into TBS and concentrated using an Amicon Ultra 10 kDa MWCO filter.
For protein crystallisation, the His-tag was cleaved using HRV 3C protease (GenScript). The protein was then repurified using nickel affinity chromatography, as before, but the flow through was collected. The protein was subjected to a final purification polishing step using gel filtration with a HiLoad 16/600 Superdex 200 pg column (GE Healthcare), eluted with TBS, at 0.5 mL min−1. Fractions containing protein were pooled and concentrated to 10 mg mL−1 using an Amicon Ultra 15 10 kDa MWCO concentrator.
Circular dichroism
Data was collected on a ChiraScan V100. The CD signal for measurement in the far-UV region (190–260 nm) was recorded in a 1 mm path length cell with a protein concentration of 1 mg mL−1. CD spectra were recorded with a bandwidth of 0.5 nm and a response time of 15 seconds, averaging three scans.Crystallisation, X-ray data collection and structure solution
Recombinant PpNeuA with His-tag removed was crystallized at 10 mg mL−1 in TBS. Crystallization conditions were identified using the PACT Premier screen (Molecular Dimensions). Crystals suitable for diffraction experiments were obtained by sitting drop vapour diffusion in 400 nL drops containing equal volumes of protein and reservoir solution, after two weeks at 4 °C. The reservoir solution comprised of 0.2 M CaCl2, 0.1 M Tris, pH 8.0, 20% w/v PEG 6000. Prior to data collection crystals were cryoprotected with PEG 200 and flash-cooled in liquid nitrogen. All data were collected at Diamond Light Source (Harwell, UK). Data reduction was performed with Dials and the structure solved using a model derived from murine CMAS (PDB ID: 1QWJ). Iterative rounds of model building, and refinement were performed in COOT35 and Phenix.refine36 respectively. Validation with MOLPROBITY37 and PDBREDO38 were incorporated into the iterative rebuild and refinement process. The resolution cut off was validated using a paired refinement in PDBREDO. Data collection and refinement statistics are shown in Table S1. Pairwise structural alignments were made using the TM-align algorithm via RSCB protein Data Bank.39Molecular docking
CTP and CMP-Kdn were docked into a receptor derived from the apo PpNeuA X-ray crystal structure using Molsoft ICM-Pro. The crystal structure of the apo form of PpNeuA was in an open conformation, a model of the closed conformation was generated prior to docking studies. The structure of Neisseria meningitidis CMP-sialic acid synthetase (NmCMAS, PDB ID: 6CKL) was chosen as the basis for the closed model of PpNeuA. The two structures were superimposed before imposing the conformation of Chain B from 6CKL onto the structure of PpNeuA. This PpNeuA model of the closed conformation was used as the starting point for all subsequent docking studies. Additional molecular dynamics for R196 and subsequent energy minimisation steps were performed during the various docking procedures. A more detailed explanation of the process is included in the supplementary information.31P NMR biochemical assay
5Az-Kdn40 and 9Az-Kdn41 were prepared using variations on published methods. 10 mM Kdn, Neu5Ac, 5Az-Kdn or 9Az-Kdn was mixed with 8 mM CTP in 50 mM Tris-HCl (pH 9.0). 31P NMR spectra of each reaction was recorded before the addition of 20 or 1 µg CMP-sialic acid synthetase enzyme (t = 0 min), and after incubation of the enzymes in 30-minute intervals over 6 h at 298 K and 400 MHz. The reaction followed via the reduction of CTP signals (−5.79 ppm doublet; −10.71 doublet; and −19.38 singlet), and the appearance of CMP-sialic acid signals (−5.51 ppm singlet; −4.65 ppm singlet).Kinetic assay
The CMAS enzyme activity was followed from the initial rates of pyrophosphate formation as detected using the EnzChek Phosphate Assay Kit (Invitrogen), following the manufacturer's instructions. The reaction was carried out in 50 mM Tris-HCl, pH 9.0 at 200 µL volume in triplicate. The reaction was initiated by the addition of 200–800 ng PpNeuA or mutants and followed by absorbance at 360 nm. The amount of pyrophosphate produced was calculated from comparison with a pyrophosphate standard curve. The initial rates of reaction were measured over a range of sialic acid (Neu5Ac, Kdn) concentrations with a fixed amount of CTP (0.5 mM). Vmax, KM and kcat were calculated according to the Michaelis Menten equation using GraphPad Prism.Results and discussion
Overall structure of PpNeuA
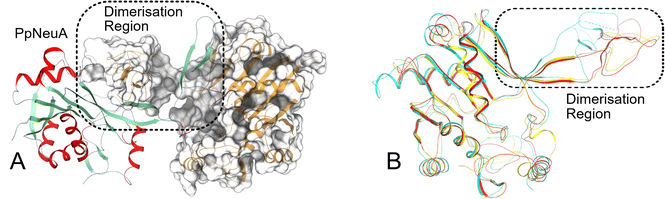
The X-ray crystal structure of ligand-free PpNeuA, without the C-terminal transmembrane domain, was solved to 1.8 Å resolution. PpNeuA belongs to space group P1211. The asymmetric unit (ASU) has dimensions 56.43, 47.78, 87.85 Å. Much like bacterial CKS and CMAS,22–24,27,42 the model consists of a homodimer, which is consistent with gel filtration analysis (Fig. S1). The two monomers superimpose with a low RMSD of 0.34 Å over 219 Cα atoms. The PpNeuA model (Fig. 2) consist of two domains, an N-terminal globular α/β domain (residues 9 to 143, 181 to 233), and a smaller dimerization domain (residues 144 to 180). The dimerization domain of each monomer forms tight interactions with the globular domain of the opposite monomer, through hydrogen bonds, salt bridges and hydrophobic contacts. The dimer interfacial area (3730 Å2) is large and helps generate a tightly entwined complex. The globular domain is an α-β-α sandwich, with the central β-sheet composed of six parallel β-strands (β1–5, 10) and one anti-parallel β-strand (β9) sandwiched between α-helices on both sides. The dimerization domain is composed of two β-strands (β6–7). The loop between β3 and αB (residues 81–88) is unmodelled due to low electron density.The CMAS crystals structures with the lowest RMSD are from M. musculus (PDB ID: 1QWJ) and Vibrio cholera (PDB ID: 6IFD)24,25 with RMSD of 1.45 Å and 1.41 Å, respectively. The PpNeuA active site is formed at the dimer interface, and includes the CTP binding pocket, and the sialic acid binding pocket. The apo-PpNeuA adopts the open conformation with no divalent metals bound, in the same way as other apo-CMAS crystal structures.
Substrate binding in PpNeuA
PpNeuA shares considerable overall structural similarity with previously published CMAS and CKS structures, however sequence identity ranges between only 25 to 40% (CMAS), 12 to 17% (CKS), and 26% (PseF). Structural and sequence alignments show that residues involved in metal ion binding and CTP binding are wholly conserved, whilst there is more variation in the amino acids that bind sialic acid (Fig. S2 and S3).The sialic acid binding site of PpNeuA is most like Neu5Ac specific CMAS, as the only difference between Neu5Ac and Kdn is the substitution at C5, Neu5Ac contains the larger and more hydrophobic N-acetyl group, whilst Kdn has a smaller and more hydrophilic hydroxyl group. The C5 group of Neu5Ac is accommodated within CMAS in a hydrophobic pocket composed of at least three hydrophobic and bulky residues in an otherwise polar active site (Fig. S2).43 However, in PpNeuA the hydrophobic pocket is formed of Leu109, Tyr183 and Arg196, the third hydrophobic residue is replaced by hydrophilic arginine. CMAS enzymes from organisms that produce only Kdn – Klebsiella ozaenae,44Brevibacterium sp.45 and Streptomyces albus46 – all contain arginine in the same position (Fig. S2), while the Kdn-specific CMAS from Danio rerio has the hydrophilic threonine in this position.30 Interestingly, the CMAS from Oncorhynchus mykiss (omCMAS), has leucine in this position despite Kdn being the preferred substrate.28 The Kdn-preference of omCMAS is likely due to decreased hydrophobicity of the C5 binding pocket, which is composed of fewer hydrophobic residues than bacterial CMAS (NmCMAS and Vibrio cholerae CMAS (VcCMAS)).23,24 Compared to the eukaryotic murine CMAS (MmCMAS),22 omCMAS has a shorter loop in the sialic acid binding site, this is proposed to alter the positioning of hydrophobic residues that are required to bind the C5 substituent, and thus weaken interactions with Neu5Ac compared to MmCMAS.22,28
Despite the differences in stereochemistry and substituents between Kdn and Pse (Fig. 1), there are similarities in the sialic acid binding site. In PseF, the hydrophobic pocket contains three additional hydrophobic residues, Phe161, Leu184 add Phe215, compared to PpNeuA, this is to bind the additional N-acetyl group at C7.20 Arg196 (PpNeuA) is equivalent to Ile214 of PseF, the hydrophobic Ile can accommodate the C5 N-acetyl substituent of Pse.
To further understand the structural basis for Kdn-specificity of PpNeuA, co-crystallisation in the presence of CTP and Kdn was attempted. However, these attempts did not result in co-crystals. Therefore CTP, CMP-Kdn and CMP-Neu5Ac were docked into a receptor derived from the apo PpNeuA X-ray crystal structure using Molsoft ICM-Pro.
CTP binding residues
Previously characterised CMAS enzymes, NmCMAS24 and MmCMAS,22 showed that CTP bound in the open configuration, whilst the VcCMAS23 was found in a partially closed conformation when CDP was bound. The partially closed conformation seen for VcCMAS is facilitated by Gln177, this residue is not conserved in NmCMAS, MmCMAS and PpNeuA, therefore, CTP was docked into the open conformation of PpNeuA. Most CTP binding residues are invariant (Fig. S2) and the interactions are conserved within PpNeuA with the docked pose matching previously co-crystalised CMAS24 and CKS27,47 structures (Fig. 3). Arg78 hydrogen bonds with O2 and N3 of cytosine base, giving specificity for cytosine. An additional interaction between Thr82 and N4 is noted in PpNeuA that is also seen in eukaryotic MmCMAS, but not in bacterial CMAS (NmCMAS and VcCMAS). Asn29 donates a hydrogen bond to the 2′OH and 3′OH of the ribose moiety. CTP triphosphate interacts with PpNeuA through Arg19, Lys28, Ser22 to the α-, β- and γ-phosphates much in the same way as other CMAS enzymes. Interestingly, PpNeuA and other Kdn specific CMAS enzymes do not have a Lys residue that is invariant in Neu5Ac-specifc enzymes (Fig. S2), in PpNeuA the residue is Val23.24 In characterised CMAS enzymes this Lys (Lys16 NmCMAS numbering), can interact through hydrogen bonding interactions with the backbone amino group to the γ-phosphate,23,24 and also through the amino group to the β-phosphate.23 It has been demonstrated that this Lys is not essential for CMAS activity, although mutation to alanine does reduce activity.48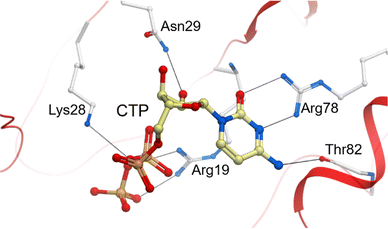 | ||
| Fig. 3 Modelled binding interaction between CTP and PpNeuA. PpNeuA is shown in grey, CTP carbon atoms are shown in cream. Hydrogen bonds are shown as grey lines. | ||
Sialic acid binding residues
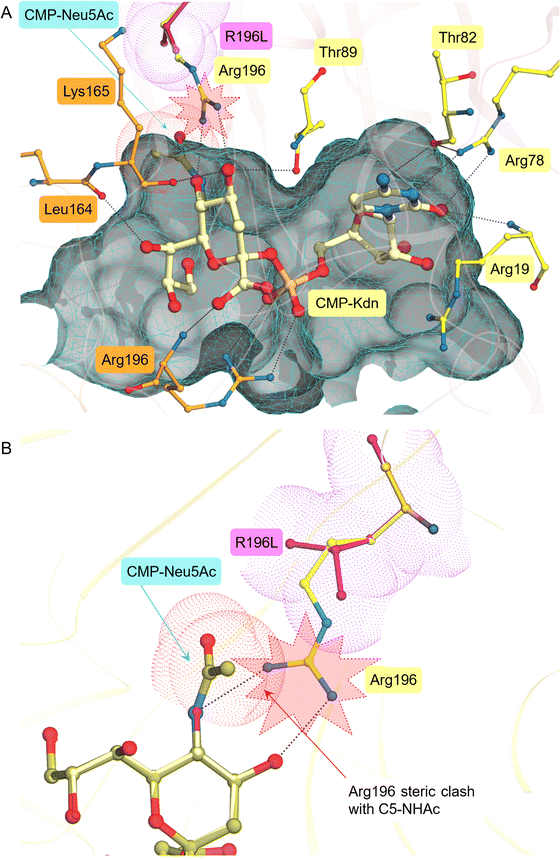
In previously characterised CMAS enzymes, Neu5Ac makes various contacts with the active site, many of which are highly conserved (Fig. S2).23,24,43In silico docking simulations of PpNeuA and its product, CMP-Kdn, showed that many of the interactions are conserved in PpNeuA. In structurally characterised CMAS, there are three key interactions between enzyme and CMP-Neu5Ac: the interaction between O4 and a serine; the interaction between Gln and O8, and the nitrogen of the N-acetyl group; and the interaction between the hydrophobic pocket and the C5 N-acetyl group. The docked pose of CMP-Kdn is similar to CMP-Neu5Ac in co-crystalised CMAS structures,22,24 and makes two of the conserved interactions. In PpNeuA Thr89 hydrogen bonds with the O4 atom, in other CMAS enzymes this threonine is substituted by serine, which maintains the same hydrogen bonding abilities. In the modelled closed structure of PpNeuA, the strictly conserved Gln110 does not make a hydrogen bonding interaction with O8 of CMP-Kdn (Fig. 4), however, the backbone of Leu164 hydrogen bonds with O7 on CMP-Kdn. The hydrophobic pocket in PpNeuA, contains Arg196 which is stacked against Lys165 in an extended position, in this position, Arg196 can hydrogen bond with the O5 of CMP-Kdn (Fig. 4). Attempts to dock CMP-Neu5Ac were unsuccessful in generating a plausible pose, due to a large steric clash between the C5 N-acetyl group of CMP-Neu5Ac and Arg196. This clash occluded CMP-Neu5Ac from binding and infers that Neu5Ac is not a substrate for PpNeuA, this is in agreement with experimental results from Wagstaff et al.17 As Neu5Ac active CMAS enzymes commonly have Leu instead of Arg in this position, this mutation was made in silico and CMP-Neu5Ac was docked successfully, thus, suggesting the role of Arg196 as gate-keeping residue for CMP-Kdn specificity.Importance of sialic acid binding residues
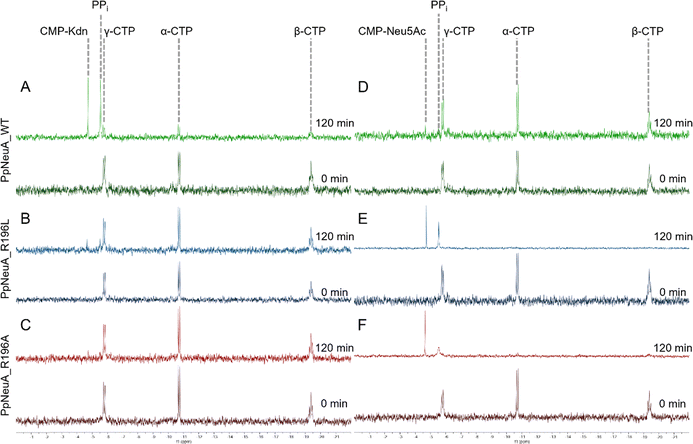
To test the importance of Arg196 in Kdn specificity and binding, two mutants were made, with Arg196 mutated to alanine or leucine (Fig. S4). Circular dichroism was conducted and showed that the mutants did not destabilise the fold of PpNeuA (Fig. S5). 31P NMR was used to follow the reaction of PpNeuA and mutants (Fig. 5). The loss of the signals for CTP, corresponding to the α-, β-, and γ-phosphates, and the formation of signals for CMP-sialic acid and pyrophosphate followed the reaction's progress. As previously shown, the wild-type enzyme does not convert Neu5Ac to CMP-Neu5Ac, whilst conversion of Kdn occurs rapidly.17 Both mutants were able to utilise Neu5Ac as a substrate. PpNeuA_R196L showed greater conversion of Neu5Ac than PpNeuA_R196A. Neither mutant was able to convert Kdn at the concentrations tested.As other CMAS enzymes show substrate promiscuity,28,30,49 reactions were carried out with twenty times more enzyme to assess substrate promiscuity. After 60 minutes of reaction, PpNeuA_WT converted less than 1% Neu5Ac to product, and both mutants converted less than 1% Kdn to product (Fig. S6).
To understand the role of Arg196, the steady state kinetic parameters for PpNeuA_WT and mutants were calculated from monitoring the release of pyrophosphate in a continuous spectrophotometric assay (Table 1 and Fig. S6). The KM values for PpNeuA_WT and mutants are in a similar range to earlier literature values for CMAS enzymes with their preferred substrate. However, compared to Kdn-specific CMAS, the KM values for PpNeuA are 10-fold lower than both the enzymes from O. mykiss28 and B. thetaiotamicron.16 For both mutants, the KM for Kdn is at least 50 times higher than the wild type, showing the importance of Arg196 in substrate binding. For Neu5Ac, PpNeuA_R196A has weaker binding compared to the larger and more hydrophobic PpNeuA_R196L mutant and demonstrates the importance of the larger hydrophobic residue in binding the N-acetyl group in the otherwise polar active site. Interestingly, the single residue mutation can change substrate specificity in PpNeuA. omCMAS despite having leucine in this position is still more active towards Kdn than Neu5Ac,28 this highlights the complexity and intricacy of substrate specificity in CMAS. omCMAS is the only sialic acid synthetase within O. mykiss, and as O. mykiss produces both Kdn- and Neu5Ac-containing glycans it is understandable that omCMAS has a relaxed substrate specificity.28
| K M/mM | k cat/s−1 | |||
|---|---|---|---|---|
| Kdn | Neu5Ac | Kdn | Neu5Ac | |
| WT | 0.12 | 4.3 | 4.7 | 0.0070 |
| R196L | 6.7 | 0.27 | 0.028 | 0.93 |
| R196A | 30 | 0.60 | 0.037 | 0.46 |
In addition to the role of Arg196 in binding, as evidenced by the increased KM for the mutants, the kcat values of the mutants are also significantly reduced, indicating that Arg196 may also play a role in transition state stabilisation and hence catalysis. The kcat values of the mutants with Kdn are more than 100 times reduced. As well as being involved in substrate binding, Arg196 is proposed to hold the substrate in a productive position for catalysis through the hydrogen bonding network. The loss of Arg196 disrupts the hydrogen bonding network, Horsfall et al., proposed that Tyr179 (NmCMAS numbering) plays a similar role to Arg196, in substrate binding and positioning.43
Substrate specificity with azide-substituted sialic acids
We next assessed the substrate scope of PpNeuA with Kdn with azide modification at C5 or C9. Previous studies with C9-azide-modified Neu5Ac, Neu5Gc and Kdn have shown significantly lower incorporation of 9Az-Kdn compared to 9Az-Neu5Ac and 9Az-Neu5Gc,50 low incorporation is partly due to the lack of Kdn specificity of CMAS, but also the identity of the sialyltransferase, where some sialyltransferases show low activity toward CMP-Kdn compared to other CMP-sialic acids.51 Activity was measured by 31P NMR after a two-hour reaction. Compared to the unsubstituted Kdn, these modified substrates showed lower conversions, 9Az-Kdn showed 31% conversion, whilst 5Az-Kdn only showed 10% conversion (Fig. S8). Unsurprisingly, the larger azido group in the C5 position, much like the N-acetyl group of Neu5Ac, may sterically clash with Arg196. In the docked model of PpNeuA with CMP-Kdn, there appears to be a larger pocket surrounding O9, which better allows the 9Az-Kdn to bind in a productive orientation for catalysis compared to 5Az-Kdn. The activity of PpNeuA towards both 9Az-Kdn and 5Az-Kdn augurs well for the efficient in vitro production of azide-labelled CMP-Kdn and related glycoconjugates; it also encourages metabolic glycoengineering studies in P. parvum to better understand the location and role of this acidic sugar in microalgal biology and viral infection processes.Conclusions
In the present study, we conducted a structural characterisation of CMP-Kdn synthetase from P. parvum, to understand its substrate specificity. Whilst all previously structurally characterised CMAS enzymes show Neu5Ac substrate specificity, PpNeuA has specificity for Kdn. Arg196 is shown to be a gate-keeper residue for Kdn specificity and play a role in both substrate binding and catalysis. The ability of PpNeuA to utilise azide-modified Kdn was demonstrated, which opens the way for PpNeuA use in the production of Kdn-containing glycoconjugates suitable for bio-orthogonal modification for both environmental, biotechnological and biomedical applications.Author contributions
C.M.: experimental data, writing – original draft preparation. A.J.M. and B.A.W.: experimental data, review-editing. I.I, E.D., A.R., M.E.O. and C.W.L.: experimental data. R.A.F: writing – review, editing, funding and conceptualization. All authors contributed to the article and approved the submitted version.Conflicts of interest
The authors declare no conflict of interest.Data availability
The atomic coordinates and structure factors for PpNeuA have been deposited in the Protein Data Bank (PDB) under accession number 9T4Z. The data are publicly available at the RCSB PDB website via the following DOI: https://doi.org/10.2210/pdb9t4z/pdb.Additional data, supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5cb00285k.
Acknowledgements
CM, AM-C and BW were supported by the University of Manchester. We gratefully acknowledge engagement and financial support from John Currie and colleagues of the Norfolk Pike Angling Club. The authors would like to thank Diamond Light Source for beamtime on i03 under proposal MX24447-63.References
- A. Varki, Trends Mol. Med., 2008, 14, 351–360 CrossRef CAS PubMed.
- S. Dedola, S. Ahmadipour, P. de Andrade, A. N. Baker, A. N. Boshra, S. Chessa, M. I. Gibson, P. J. Hernando, I. M. Ivanova, J. E. Lloyd, M. J. Marin, A. J. Munro-Clark, G. Pergolizzi, S. J. Richards, I. Ttofi, B. A. Wagstaff and R. A. Field, RSC Chem. Biol., 2024, 5, 167–188 RSC.
- A. L. Lewis, X. Chen, R. L. Schnaar and A. Varki, in Essentials of Glycobiology, ed. A. Varki, R. D. Cummings, J. D. Esko, P. Stanley, G. W. Hart, M. Aebi, D. Mohnen, T. Kinoshita, N. H. Packer, J. H. Prestegard, R. L. Schnaar and P. H. Seeberger, Cold Spring Harbor, NY, 4th edn, 2022, pp. 185–204 DOI:10.1101/glycobiology.4e.15.
- H. S. Chidwick and M. A. Fascione, Org. Biomol. Chem., 2020, 18, 799–809 RSC.
- S. Jelakovic, K. Jann and G. E. Schulz, FEBS Lett., 1996, 391, 157–161 CrossRef CAS PubMed.
- J. E. Stencel-Baerenwald, K. Reiss, D. M. Reiter, T. Stehle and T. S. Dermody, Nat. Rev. Microbiol., 2014, 12, 739–749 CrossRef CAS PubMed.
- J. M. Fulton, H. F. Fredricks, K. D. Bidle, A. Vardi, B. J. Kendrick, G. R. DiTullio and B. A. Van Mooy, Environ. Microbiol., 2014, 16, 1137–1149 CrossRef CAS PubMed.
- S. L. Rose, J. M. Fulton, C. M. Brown, F. Natale, B. A. Van Mooy and K. D. Bidle, Environ. Microbiol., 2014, 16, 1150–1166 CrossRef CAS PubMed.
- B. A. Wagstaff, E. S. Hems, M. Rejzek, J. Pratscher, E. Brooks, S. Kuhaudomlarp, E. C. O'Neill, M. I. Donaldson, S. Lane, J. Currie, A. M. Hindes, G. Malin, J. C. Murrell and R. A. Field, Biochem. Soc. Trans., 2018, 46, 413–421 CrossRef CAS PubMed.
- S. R. Manning and J. W. La Claire, Mar. Drugs, 2010, 8, 678–704 CrossRef CAS PubMed.
- J. Sobieraj and D. Metelski, Toxins, 2023, 15, 403 CrossRef CAS PubMed.
- C. P. Brussaard, J. Eukaryot Microbiol., 2004, 51, 125–138 CrossRef PubMed.
- B. A. Wagstaff, I. C. Vladu, J. E. Barclay, D. C. Schroeder, G. Malin and R. A. Field, Viruses, 2017, 9, 40 CrossRef PubMed.
- B. A. Wagstaff, J. Pratscher, P. P. L. Rivera, E. S. Hems, E. Brooks, M. Rejzek, J. D. Todd, J. C. Murrell and R. A. Field, Environ. Sci. Technol., 2021, 55, 16538–16551 CrossRef CAS PubMed.
- S. Inoue and K. Kitajima, Glycoconj. J., 2006, 23, 277–290 CrossRef CAS PubMed.
- L. Wang, Z. Lu, K. N. Allen, P. S. Mariano and D. Dunaway-Mariano, Chem. Biol., 2008, 15, 893–897 CrossRef CAS PubMed.
- B. A. Wagstaff, M. Rejzek and R. A. Field, J. Biol. Chem., 2018, 293, 16277–16290 CrossRef CAS PubMed.
- J. Hao, W. F. Vann, S. Hinderlich and M. Sundaramoorthy, Biochem. J., 2006, 397, 195–201 CrossRef CAS PubMed.
- M. Sellmeier, B. Weinhold and A. Munster-Kuhnel, Top. Curr. Chem., 2015, 366, 139–167 CrossRef CAS PubMed.
- T. Keenan, A. R. Cowan, E. K. P. Flack, N. E. Hatton, A. J. Walklett, G. H. Thomas, G. R. Hemsworth and M. A. Fascione, Structure, 2024, 32, 2399–2409 CrossRef CAS PubMed.
- D. Kapitonov and R. K. Yu, Glycobiology, 1999, 9, 961–978 CrossRef CAS PubMed.
- S. Krapp, A. K. Munster-Kuhnel, J. T. Kaiser, R. Huber, J. Tiralongo, R. Gerardy-Schahn and U. Jacob, J. Mol. Biol., 2003, 334, 625–637 CrossRef CAS PubMed.
- S. Bose, D. Purkait, D. Joseph, V. Nayak and R. Subramanian, Acta Crystallogr., Sect. D:Struct. Biol., 2019, 75, 564–577 CrossRef CAS PubMed.
- M. M. Matthews, J. B. McArthur, Y. Li, H. Yu, X. Chen and A. J. Fisher, Biochemistry, 2020, 59, 3157–3168 CrossRef CAS PubMed.
- S. C. Mosimann, M. Gilbert, D. Dombroswki, R. To, W. Wakarchuk and N. C. Strynadka, J. Biol. Chem., 2001, 276, 8190–8196 CrossRef CAS PubMed.
- D. J. Heyes, C. Levy, P. Lafite, I. S. Roberts, M. Goldrick, A. V. Stachulski, S. B. Rossington, D. Stanford, S. E. Rigby, N. S. Scrutton and D. Leys, J. Biol. Chem., 2009, 284, 35514–35523 CrossRef CAS PubMed.
- S. Jelakovic and G. E. Schulz, J. Mol. Biol., 2001, 312, 143–155 CrossRef CAS PubMed.
- D. Nakata, A. K. Munster, R. Gerardy-Schahn, N. Aoki, T. Matsuda and K. Kitajima, Glycobiology, 2001, 11, 685–692 CrossRef CAS PubMed.
- T. Haselhorst, A. K. Munster-Kuhnel, A. Stolz, M. Oschlies, J. Tiralongo, K. Kitajima, R. Gerardy-Schahn and M. von Itzstein, Biochem. Biophys. Res. Commun., 2005, 327, 565–570 CrossRef CAS PubMed.
- W. Schaper, J. Bentrop, J. Ustinova, L. Blume, E. Kats, J. Tiralongo, B. Weinhold, M. Bastmeyer and A. K. Munster-Kuhnel, J. Biol. Chem., 2012, 287, 13239–13248 CrossRef CAS PubMed.
- S. Wang, C. Chen, M. R. Gadi, V. Saikam, D. Liu, H. Zhu, R. Bollag, K. Liu, X. Chen, F. Wang, P. G. Wang, P. Ling, W. Guan and L. Li, Nat. Commun., 2021, 12, 3573 CrossRef CAS PubMed.
- P. Laborda, S.-Y. Wang, A.-M. Lu, M. He, X.-C. Duan, Y.-J. Qian, Y.-S. Jung, L. Liu and J. Voglmeir, Adv. Synth. Catal., 2017, 359, 3120–3125 CrossRef CAS.
- Y. Zhou, Y. Li, J. Wen, Y. Zhang, Z. Hu, K. Zhong, H. Cao and J. Cheng, Chin. J. Chem., 2025, 43, 1479–1486 CrossRef CAS.
- N. S. Berrow, D. Alderton, S. Sainsbury, J. Nettleship, R. Assenberg, N. Rahman, D. I. Stuart and R. J. Owens, Nucleic Acids Res., 2007, 35, e45 CrossRef PubMed.
- P. Emsley and K. Cowtan, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2004, 60, 2126–2132 CrossRef PubMed.
- D. Liebschner, P. V. Afonine, M. L. Baker, G. Bunkoczi, V. B. Chen, T. I. Croll, B. Hintze, L. W. Hung, S. Jain, A. J. McCoy, N. W. Moriarty, R. D. Oeffner, B. K. Poon, M. G. Prisant, R. J. Read, J. S. Richardson, D. C. Richardson, M. D. Sammito, O. V. Sobolev, D. H. Stockwell, T. C. Terwilliger, A. G. Urzhumtsev, L. L. Videau, C. J. Williams and P. D. Adams, Acta Crystallogr., Sect. D:Struct. Biol., 2019, 75, 861–877 CrossRef CAS PubMed.
- C. J. Williams, J. J. Headd, N. W. Moriarty, M. G. Prisant, L. L. Videau, L. N. Deis, V. Verma, D. A. Keedy, B. J. Hintze, V. B. Chen, S. Jain, S. M. Lewis, W. B. Arendall, 3rd, J. Snoeyink, P. D. Adams, S. C. Lovell, J. S. Richardson and D. C. Richardson, Protein Sci., 2018, 27, 293–315 CrossRef CAS PubMed.
- R. P. Joosten, F. Long, G. N. Murshudov and A. Perrakis, IUCrJ, 2014, 1, 213–220 CrossRef CAS PubMed.
- S. Bittrich, J. Segura, J. M. Duarte, S. K. Burley and Y. Rose, Bioinformatics, 2024, 40(6), btae370 CrossRef CAS PubMed.
- C. H. Lin, T. Sugai, R. L. Halcomb, Y. Ichikawa and C. H. Wong, J. Am. Chem. Soc., 1992, 114, 10138–10145 CrossRef CAS.
- D. C. M. Kong and M. von Itzstein, Carbohydr. Res., 1997, 305, 323–329 CrossRef CAS.
- H. J. Yoon, M. J. Ku, B. Mikami and S. W. Suh, Acta Crystallogr., Sect. D:Biol. Crystallogr., 2008, 64, 1292–1294 CrossRef CAS PubMed.
- L. E. Horsfall, A. Nelson and A. Berry, FEBS J., 2010, 277, 2779–2790 CrossRef CAS PubMed.
- Y. A. Knirel, N. A. Kocharova, A. S. Shashkov, N. K. Kochetkov, V. A. Mamontova and T. F. Soloveva, Carbohydr. Res., 1989, 188, 145–155 CrossRef CAS PubMed.
- N. V. Potekhina, L. I. Evtushenko, S. N. Senchenkova and A. S. Shashkov, Russ. J. Bioorg. Chem., 2007, 33, 66–72 CrossRef CAS PubMed.
- A. S. Shashkov, G. M. Streshinskaya, E. M. Tul'skaya, S. N. Senchenkova, L. M. Baryshnikova, A. S. Dmitrenok, B. E. Ostash and V. A. Fedorenko, Antonie van Leeuwenhoek, 2016, 109, 923–936 CrossRef CAS PubMed.
- H. Schmidt, J. R. Mesters, J. Wu, R. W. Woodard, R. Hilgenfeld and U. Mamat, PLoS One, 2011, 6, e23231 CrossRef CAS PubMed.
- D. M. Stoughton, G. Zapata, R. Picone and W. F. Vann, Biochem. J., 1999, 343(Pt 2), 397–402 CrossRef CAS PubMed.
- I. B. Mertsalov, B. N. Novikov, H. Scott, L. Dangott and V. M. Panin, Biochem. J., 2016, 473, 1905–1916 CrossRef CAS PubMed.
- B. Cheng, L. Dong, Y. Zhu, R. Huang, Y. Sun, Q. You, Q. Song, J. C. Paton, A. W. Paton and X. Chen, ACS Chem. Biol., 2019, 14, 2141–2147 CAS.
- O. Blixt and J. C. Paulson, Adv. Synth. Catal., 2003, 345, 687–690 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |