 Open Access Article
Open Access ArticleDynamic granular hydrogels to assess pancreatic cancer cell fate
Ellen
Frahm
a and
Chien-Chi
Lin
 *ab
*ab
aWeldon School of Biomedical Engineering, Purdue University, 206 Martin Jischke Dr, West Lafayette, IN 47907, USA
bIndiana University Simon Comprehensive Cancer Center, Indianapolis, IN, USA. E-mail: lin711@purdue.edu; Tel: +(765) 495-7791
First published on 15th January 2026
Abstract
Granular hydrogels are an emerging biomaterial platform increasingly used in biomedical applications, including therapeutic delivery and tissue regeneration. Assembled from micron-scale hydrogel particles through physical assembly or chemical cross-linking, granular hydrogels possess micro- and macroscopic pores that facilitate molecular transport and cell migration. However, current granular hydrogels are typically fabricated with defined stiffness, porosity, and compositions that do not recapitulate the dynamic nature of native tissues, including the tumor microenvironment. To address this challenge, we have developed dynamic granular hydrogels formed by gelatin-norbornene-carbohydrazide (GelNB-CH) microgels. GelNB-CH microgels were first prepared from a microfluidic droplet generator coupled with the rapid thiol-norbornene photo-click gelation. The collected microgels were annealed via inverse electron-demand Diels–Alder (iEDDA) click reaction to form granular hydrogels, which were dynamically stiffened via hydrazone bonding. Notably, adjusting the concentration of the stiffening reagent (i.e., oxidized dextran, oDex) enabled dynamic stiffening of the granular hydrogels without affecting the void fraction. Pancreatic cancer-associated fibroblasts (CAFs) seeded in the granular hydrogels spread rapidly throughout the scaffold and induced cancer cell migration. This work enhances the design of granular hydrogels, offering a highly adaptable biomaterial platform for in vitro cancer modeling.
1. Introduction
Pancreatic ductal adenocarcinoma (PDAC) comprises about 90% of all pancreatic cancer cases, with a five-year survival rate of 13% in 2025.1 The tumor microenvironment (TME) of PDAC is marked by excess deposition of extracellular matrices (ECM),2 resulting in high matrix stiffness that limits the permeability of therapeutics.3 The excess matrices are deposited by cancer-associated fibroblasts (CAFs), which also interact closely with pancreatic cancer cells (PCCs) to guide their cell-fate processes.4,5 Studies have shown that healthy pancreatic tissues are substantially softer (Young's modulus E ∼ 3 kPa, equivalent to shear modulus G′ ∼ 1 kPa) than their cancerous counterparts.6,7 The variation in stiffness manifests spatiotemporally in the TME. To this end, many hydrogel-based PDAC models have been developed to present the cells with a microenvironment of varying stiffness. For example, we have reported biomimetic hydrogel models for PDAC research, in which the gels were crosslinked via a modular thiol-norbornene photo-click reaction between multi-arm poly(ethylene glycol)-norbornene (PEGNB) and bis-cysteine-bearing peptides.8–11 Zhang et al. created gelatin-methacrylate (GelMA) hydrogels with stiffness ranging from 1 kPa to 20 kPa to demonstrate enhanced cancer cell colony formation in stiffer gels.12 Curvello et al. utilized collagen-nanocellulose matrices to demonstrate that PCC/CAF co-cultures resulted in a threefold increase in stiffness compared to a PCC monoculture control.13 To mimic the dynamic tumor tissue properties, we have reported several hydrogel systems whose crosslinking density could be adjusted on-demand through modular click chemistry, hydrazone bonding,14,15 enzymatic reaction,16,17 or viscoelastic interactions.18,19Most chemically crosslinked hydrogels are considered mesoporous, with an average ‘mesh size’ of a few tens of nanometers.20,21 Unless the gels were designed to degrade, these mesoporous hydrogels would restrict macromolecular transport and cell migration.22 To this end, hydrogels with micron-scale pores have been engineered to overcome the constraints of mesoporous bulk hydrogels.23–25 One such example is granular hydrogels assembled from individual microgels, which can be fabricated from inverse suspension/bulk emulsion,14,26 in-air27 or droplet chip microfluidics,28 extrusion or volumetric bioprinting,29 or mechanical crushing of bulk hydrogels.24 These microgels can be physically packed into a tight scaffold30,31 with limited porosity and diffusivity.32 Microgels can also be assembled into granular hydrogels through secondary chemical reactions between the microgels, forming “microporous annealed particles” or MAP scaffolds.33 MAP scaffolds typically exhibit large voids between the microgels, enabling cells to migrate more freely without significantly degrading the biomaterial matrix.34 The void fraction in MAP scaffolds or granular hydrogels could be controlled by adjusting microgel packing density or by incorporating sacrificial microgels that degrade post-assembly.35 For example, Seymour et al. create scaffolds with 60% of void space at the expense of a compromised scaffold integrity.35 An alternate example by Qazi et al. created continuously longer void spaces with rod-shaped microgels.36
The microgel size and annealing process can affect the scaffold's void space and adhesion properties.37–40 For example, Widener et al. showed that homogeneous microspheres were more suitable for culturing Jurkat T cells as they maintained even cell distribution and higher viability.40 Caldwell et al. demonstrated that microgels with diameters in the hundred-micron range promote a spindling morphology of mesenchymal stem cells compared to those with ten-micron diameters.38 Truong et al. used norbornene-modified hyaluronic acid to prepare microgels for further conjugating cell-adhesive peptide RGD to promote fibroblastic growth for wound healing.37 Most granular hydrogel scaffolds are annealed with secondary reactions such as photocrosslinking,41,42 Michael-type addition,43 guest–host interaction,40 or inverse electron demand Diels–Alder (iEDDA) click reaction.14 Additionally, cell-generated traction force or cell-secreted ECM may serve as a mechanism to assemble the microgels and further stabilize the scaffold alongside the chemical annealing reaction.44
Granular hydrogels have been exploited to model cancer in vitro. For example, Kieda et al. generated MCF-7 breast cancer spheroids in dextran-alginate microgels for drug testing.28 Zhang et al. used GelMA/Matrigel microgels to model cellular clustering for lung cancer.45 Clara-Trujillo et al. demonstrated the proliferation of multiple myeloma cells in fibronectin/hyaluronic acid microgels in a suspension culture.46 Nguyen et al. demonstrated the conjugation of collagen I to polyacrylamide microgels to improve glioblastoma spreading.34 Molley et al. modeled melanoma–endothelium interactions using a packed granular hydrogel.26 Nonetheless, few studies have exploited granular hydrogels to model the PDAC microenvironment. Atkins et al. used granular hydrogels to probe the mechanical effects of PDAC cells when the cells were transferred between soft and stiff granular hydrogels.47 Siciliano et al. utilized alginate microgels containing pH-sensitive fluorescent probes to measure optically the extracellular pH values of PDAC cells treated with chemotherapeutics.48 Nonetheless, these microgels/granular hydrogels did not exhibit dynamically tunable properties.
In this study, we developed a dynamic granular hydrogel platform to investigate the impact of matrix stiffening on pancreatic CAFs and PCCs. We synthesized microgels from gelatin-norbornene-carbohydrazide (GelNB-CH) due to its capacity to support three orthogonal click reactions: thiol-norbornene photocrosslinking, tetrazine-norbornene iEDDA click reaction, and hydrazide-aldehyde hydrazone click reaction.15 Our prior work has established dynamic GelNB-CH hydrogels and microgels that promote cell spreading and proliferation as stiffness increases.14,15 In this work, we designed a new microgel processing workflow to prepare annealed GelNB-CH granular hydrogels, followed by dynamic tuning of granular hydrogel stiffness to assess pancreatic CAF spreading and cancer cell growth patterns.
2. Materials & methods
2.1. Materials
Type A gelatin (230–280 bloom) was purchased from VWR Scientific. 4-arm PEG-thiol (PEG4SH, 10 kDa) and 4-arm PEG-amido-succinic-acid (PEG4ASA) was purchased from Laysan Bio. Lithium phenyl-2,4,6-trimethylbenzoylphosphinate (LAP), dextran (15–25 kDa), and FITC-dextran (2 MDa) were acquired from Sigma-Aldrich. Sodium metaperiodate (NaIO4) was purchased from Fisher Scientific. Tetrazine-amine was purchased from Click Chemistry Tools. Carbic anhydride, carbohydrazide, and N-(3-dimethylaminopropyl)-N′-ethyl carbodiimide (EDC) hydrochloride were purchased from Acros Organics. HOBt hydrate was acquired from Oakwood Chemical. O-(7-Azabenzotriazole-1-yl)-N,N,N,N′-tetramethyluronium hexafluorophosphate (HATU) was purchased from Chem-Implex International. 5% Pico-Surf in Novec 7500 and Novec 7500 solutions were acquired from Sphere Fluidics and 3M, respectively. PDMS for droplet chip replicas (SYLGARD™ 184) and for annealing chambers used in material characterization and cell study (DOWSIL™ SE 1700) was purchased from Dow Chemical. CF-488A-carbohydrazide was purchased from Biotium. Fluoraldehyde and 2,4,6-trinitrobenzene sulfonic acid (TNBSA) were obtained from Fisher Scientific. All other chemicals were obtained from Fisher Scientific unless otherwise noted.2.2. Macromer synthesis and characterization
GelNB-CH was synthesized according to our published protocol.15 First, the primary amines on pristine gelatin (1 g) were reacted with carbic anhydride (0.6 g) to form GelNB in 18 mL of PBS at pH 8 (adjusted by adding 1 N NaOH) and room temperature for 18 hours.49 The product was dialyzed against ddH2O for 48 hours and freeze-dried. The degree of NB substitution was characterized by a fluoraldehyde assay and determined to be 5.1–5.4 mM per wt% gelatin, equivalent to 85–90% norbornene substitution. Next, GelNB was reacted with carbohydrazide (CH) with a ratio of 1 g GelNB to 1.2 g CH. The reaction occurred through EDC/HOBt-assisted carbodiimide chemistry, using 2.25 mmol of EDC and HOBt each, in 13 mL 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 DMF/ddH2O at pH 5 and room temperature for 24 hours. The product was dialyzed against ddH2O for 48 hours and freeze-dried, to which the degree of CH substitution was determined to be 3.5–4 mM per wt% (by TNBSA assay). Oxidized dextran (oDex) was synthesized by reacting 1 gram of 15–25 kDa dextran with 1.58 grams of NaIO4, yielding two aldehyde groups on C2 and C4 positions on dextran backbone.50 Fluorescent CF488A-oDex was prepared by quickly vortexing 5 μL of 5 mM CF488A-carbohydrazide within 100 μL of a 5% oDex solution at 4 °C.
1 DMF/ddH2O at pH 5 and room temperature for 24 hours. The product was dialyzed against ddH2O for 48 hours and freeze-dried, to which the degree of CH substitution was determined to be 3.5–4 mM per wt% (by TNBSA assay). Oxidized dextran (oDex) was synthesized by reacting 1 gram of 15–25 kDa dextran with 1.58 grams of NaIO4, yielding two aldehyde groups on C2 and C4 positions on dextran backbone.50 Fluorescent CF488A-oDex was prepared by quickly vortexing 5 μL of 5 mM CF488A-carbohydrazide within 100 μL of a 5% oDex solution at 4 °C.
2.3. Fabrication of GelNB-CH microgels via microfluidic droplet chip generation
Microfluidic droplet chips were formed via soft lithography from a silicon wafer master device with SYLGARD™ 184 PDMS to produce replicas. The replicas were cleaned with acetone and treated with air plasma treatment and mounted on a glass slide. A precursor solution containing GelNB-CH (5 or 10 wt%), PEG4SH (1 or 2 wt%), and LAP (2 mM), was mixed in a 1-milliliter syringe. Two syringe pumps (Braintree Scientific, BS-300) were used concurrently to pump the aqueous precursor phase (10–40 microliters per minute) and the oil phase (Novec 7500 with 2% Pico-Surf, 30–200 microliters per minute) through the droplet chip and collected in a microcentrifuge tube. The droplets were photocrosslinked through exposure to 365 nm light at 5 mW cm−2 for 2 minutes. The resultant microgels were collected by passing through a 40-micron cell strainer, with the oil drained into a 50 mL conical tube. The strainer and conical tube were centrifuged for five seconds to remove the remaining oil from the microgels. The collected microgels were imaged with BioTek Lionheart FX microscope (Agilent Technologies) and microgel diameters were obtained using ImageJ software. The polydispersity index (PDI) of microgel populations was determined using the equation shown below.51 | (1) |
2.4. Hydrogel characterization
Precursor solutions with 5 or 10 wt% of GelNB-CH and 1–3 wt% PEG4SH were prepared for determining their viscosity. Viscosity testing occurred with a parallel plate rheometer (Anton Paar MCR102) under ramp logarithmic shear stress reported as a function of viscosity at a given shear rate. Thus, the primary interest for the viscosity of each precursor solution occurs at the specific shear rate as experienced in the microfluidic system. The shear rate at the optimized syringe pumping rate was determined by the Hagen/Poiseuille equation as shown below.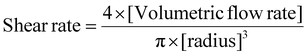 | (2) |
The radius of the microchannel is 125 microns and the volumetric flow rate of the aqueous precursor is 18 microliters per minute, resulting in a final shear rate of 195.58 1 s−1. Per the definition of Poiseuille's law, this shear rate is applicable to fluids of all viscosities that flow through the microchannel of the microfluidic system. Bulk hydrogels of all precursors were formulated for strain-sweep shear modulus measurement with oscillating shear strain applied in a ramp logarithmic increase from 0.1% to 5% strain from a parallel plate rheometer.
2.5. Scaffold assembly and characterization
The microgels were washed with PBS and incubated in PBS containing oDex (0.0125 to 0.1 wt%) at room temperature for 24 hours. Phase contrast images were taken of microgels to observe their changes in diameter due to the formation of additional hydrazone bonding. Bulk hydrogels were incubated in oDex solutions of the same concentrations and measured with strain-sweep shear rheometry. A combination of the results of bulk hydrogel and microgel behavior determined the stiffening parameters for microgels to form scaffolds. The microgels were prepared into a scaffold with initial vortexing within a 0.5 wt% PEG-tetra-tetrazine (PEG4Tz) solution and centrifuged at 6000 RPM (Thermo Scientific Sorvall ST 8R) for 90 seconds. The aggregate of microgels was incubated at room temperature for one hour. The PEG4Tz supernatant was aspirated and reserved for molding the scaffold. The microgels were transferred to 7.5 mm diameter circular molds (DOWSIL™ SE 1700 PDMS) to create scaffolds that were roughly 1 mm in thickness. The PEG4Tz supernatant was refreshed over the microgels and incubated at room temperature for another hour. The resultant scaffold was removed from the mold and placed within an oDex solution between 0.0125 wt% and 0.1 wt% for 24 hours at room temperature. To determine the relative oDex distribution through the scaffold, solutions of CF488A-oDex were used to observe the distribution of oDex in the granular hydrogel. Fluorescence images throughout the granular hydrogels were captured with Oxford BC43 benchtop confocal microscope (Andor Instruments). Scaffolds were characterized for their shear modulus with parallel-plate rheometry (Anton Paar MCR102) and their Young's modulus with micro-indentation (Cell Scale Micro Tester G2) with a 0.3048-millimeter diameter beam and a 1-millimeter spherical indenter.2.6. Cancer cell culture and spheroid assembly
1303-GFP-49-hT-CAFs and Td-tomato Pa03Cs (gifts from Dr Melissa Fishel, Indiana University School of Medicine) were maintained in high glucose DMEM supplemented with 10% FBS and 1% antibiotics/antimycotics and maintained at 37 °C with 5% CO2. Cell spheroids were formed via the protocol for the AggreWell™400 culture plate from STEMCELL Technologies. Briefly, the microwells within the well plate are rinsed with an anti-adherence solution and culture medium. Cells were seeded within the well and allowed to settle in the microwells for 24 hours before harvesting.2.7. CAF spheroids embedding in dynamic granular hydrogels
CAF spheroids were assembled with roughly 100 cells per spheroid, and roughly 6000 spheroids were harvested. Separately, microgels were formed under sterile conditions as previously described and annealed for one hour via incubation with 0.5 wt% PEG4Tz at room temperature. Next, CAF spheroids were mixed amongst the microgels and pipetted into a 10 mm glass-bottom tissue culture dish, forming a disc-shaped spheroid-laden granular hydrogel. Each granular hydrogel contained up to 1000 spheroids. A layer of 0.5 wt% PEG4Tz solution was added on top of the scaffolds for one hour. Culture medium was added over the granular hydrogels after the one-hour period. The granular hydrogels either remained in the culture medium or were supplemented with 0.1 wt% oDex one day post-embedding and were cultured for 4 days with daily imaging via confocal microscope.2.8. Individual CAF seeding in Pa03C spheroids-laden dynamic granular hydrogels
Pa03c spheroids were assembled with roughly 150 cells per spheroid, and roughly 6000 spheroids were harvested. Pa03C spheroid-laden granular hydrogels were prepared as described in the section above. Individual CAFs were suspended in culture medium at 106 cells per mL and seeded atop the Pa03C spheroid-laden granular hydrogel. Dynamic stiffening and cell culture processes were performed as described above.2.9. Immunostaining CAF from 2D culture
For 2D culture of CAF on hydrogels, flat bulk hydrogels were formed with the same precursor as the microgels. CAF spheroids were formed and seeded on top of the hydrogels, which either remained in the culture medium or were stiffened via supplementing the media with 0.1 wt% oDex one day post-seeding. The spheroid-seeded flat hydrogels were cultured for 4 days, with daily confocal microscopy imaging. For immunostaining, CAFs were fixed with 4% paraformaldehyde 24 hours at 4 °C after the culture day 4 imaging and then rinsed three times with PBS at ten-minute intervals. Fixed cells were permeabilized with 0.1% Triton-X in PBS for one hour and then rinsed three times with PBS at ten-minute intervals. Phalloidin-rhodamine was diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100 in PBS and incubated over the hydrogels on a rocker plate for 1 hour. The gels were rinsed three times with PBS at ten-minute intervals. DAPI was diluted 1
100 in PBS and incubated over the hydrogels on a rocker plate for 1 hour. The gels were rinsed three times with PBS at ten-minute intervals. DAPI was diluted 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 200 in PBS and incubated over the hydrogels on a rocker plate for 30 minutes. The gels were rinsed three times with PBS at ten-minute intervals and imaged via confocal microscope.
200 in PBS and incubated over the hydrogels on a rocker plate for 30 minutes. The gels were rinsed three times with PBS at ten-minute intervals and imaged via confocal microscope.
2.10. Statistical analysis
All data were reported as mean with a standard error of the mean. All statistical analyses were performed with GraphPad Prism software. One-way ANOVA was used to analyze microgel diameter (Fig. 3B, C and 4D), polydispersity index (Fig. 2D), and void fraction (Fig. 5D and S2). Two-way ANOVA was used to analyze microgel diameter (Fig. 2C), precursor solution viscosity (Fig. 2E). Tukey's HSD test was used to determine significant groups for all one- and two-way ANOVA. Unpaired t-tests with the Holm-Sidak method were used to analyze differences in maximum-intensity projection areas (Fig. 6C, F and S3B). Values of p < 0.05 were statistically significant. Levels of significance were designated as * (p < 0.05), ** (p < 0.01), *** (p < 0.001) and **** (p < 0.0001).3. Results & discussion
3.1. GelNB-CH hydrogel crosslinking and dynamic stiffening
In this study, primary hydrogel crosslinking was achieved through thiol-norbornene photo-click reaction between GelNB-CH and PEG4SH, using LAP as the photoinitiator (Fig. 1A). Irradiation of LAP under 365 nm light generates radical species to extract protons from sulfhydryl groups on PEG4SH, yielding thiyl radicals that react with the unsaturated norbornene groups on GelNB-CH. The norbonyl radical then propagates through another thiol group to produce a thioether bond, regenerating a thiyl radical to continue participating in the thiol-norbornene photo-click reaction until all radicals are consumed/terminated (Fig. 1A). The benefits of using GelNB-CH as the macromer for hydrogel fabrication include: (1) inherent bioactivity: GelNB-CH is synthesized from gelatin, denatured collagen with inherent bioactive sites for protease cleavage and cell adhesion. (2) Modular crosslinking: GelNB-CH can be crosslinked into hydrogels with either inert (e.g., PEG4SH) or bioactive crosslinker (e.g., thiolated hyaluronic acid or THA), allowing orthogonal control of hydrogel crosslinking density and bioactive molecules. (3) Dynamic modification: both NB and CH moieties can be leveraged for post-gelation on-demand modification of hydrogel physicochemical properties. For example, the excess NB group can be used for post-gelation conjugation of thiol or tetrazine-bearing molecules, whereas the CH moiety is reactive toward aldehyde-containing macromers (e.g., oxidized dextran or oDex) via a hydrazone reaction.In situ photo-rheometry was conducted to assess the effect of photoinitiator LAP concentration on crosslinking of GelNB-CH (5 wt%) and PEG4SH (1 wt%). Increasing LAP concentration from 0.5 mM to 2 mM led to a faster gel point (the crossover point where storage modulus exceeds loss modulus), decreasing from ∼30 seconds to within 10 seconds upon light exposure (Fig. 1B). This is comparable to the crosslinking of GelNB, where initial gelation occurs between 5 and 30 seconds of light exposure.49,52–54 Like other thiol-norbornene hydrogel systems, the crosslinking density and stiffness of GelNB-CH hydrogels could be independently tuned by adjusting either the thiol (from PEG4SH) or norbornene (from GelNB-CH) content (Fig. 1C and D). These GelNB-CH hydrogels were thin (thickness ∼ 500 μm) and crosslinked rapidly (under 2 min) using low-intensity (5 mW cm−2) longwave UV light (365 nm) and a very low concentration of photoinitiator LAP (2 mM or 0.059 wt%). These photocrosslinking parameters are considered cytocompatible for in situ cell encapsulation.55 Furthermore, modular control of gel moduli was readily achieved independent of the bioactive component, enabling the decoupling of the effect of matrix biophysical and biochemical properties on cell fate processes.49
3.2. Effect of polymer compositions on microgel fabrication
Microfluidic droplet-generating chips have been widely used to create microgels with uniform sizes.56,57 In a typical microfluidic chip, the aqueous prepolymer solution (i.e., GelNB-CH, PEG4SH, LAP) and the oil phase (i.e., Pico-Surf in Novec 7500) flow through the respective channels and join at the junction where droplets of the aqueous solution are ‘pinched off’ by the oil stream. The droplets are then crosslinked into microgels through appropriate chemistry (e.g., thiol-norbornene photo-crosslinking, Fig. 2A). We have previously reported the fabrication of GelNB-CH microgels in a microfluidic chip with defined channel dimensions.14 However, the microfluidic parameters governing the size and polydispersity index (PDI) of the resulting microgels were not evaluated systematically. In the current contribution, we first studied the effect of precursor solution composition (i.e., [GelNB-CH] and [PEG4SH]) on microgel sizes (Fig. 2B). Here, LAP concentration was fixed at 2 mM as complete gelation was achieved within 60 seconds (Fig. 1B). Increasing GelNB-CH concentration from 5 wt% to 10 wt% with 1 wt% PEG4SH increased average microgel sizes, from ∼140 μm to ∼206 μm, respectively (Fig. 2B and C). Increasing crosslinker content to 2 wt% PEG4SH increased the microgel diameter to ∼162 μm and ∼352 μm for 5 wt% and 10 wt% GelNB-CH, respectively. Monodisperse microgel populations have an average PDI of 0.1 or less (eqn (1)).51 Increasing the concentration of GelNB-CH significantly increased the PDIs of the microgels (Fig. 2D). Furthermore, microgels fabricated from 5 wt% GelNB-CH and 1 wt% PEG4SH were especially monodisperse, with a PDI of ∼0.01. We hypothesized that increasing the polymer content drastically increases solution viscosity, creating difficulty for the oil phase to ‘pinch off’ the viscous aqueous phase. To test this hypothesis, we measured the viscosities of all precursor solutions at the aqueous phase flow rate in the microchannel of the droplet generation chip. It was determined that the shear rate of any fluid through the channel would be 195.58 s−1 (eqn (2)). Hence, precursor solutions were subjected to increases in shear stress through a corresponding shear rate of 200 s−1. A 7.5-fold increase in viscosity (i.e., ∼25 mPa s to ∼189 mPa s) was observed between the lowest and highest macromer-containing precursors (5 wt%/1 wt% and 10 wt%/2 wt% of GelNB-CH/PEG4SH, respectively, Fig. 2E). This observation suggested that the viscosity of the aqueous phase might play a role in determining the size of the resulting microgels. Due to their low PDI of 0.01, microgels fabricated from 5 wt% GelNB-CH and 1 wt% PEG4SH were used for subsequent studies.3.3. Effect of microfluidic flow rates on microgel fabrication
Beyond precursor formulation, microgel properties can be further customized by manipulating the flow rates of aqueous precursor and surfactant oil during droplet generation. Microfluidic droplet generator chips feature precisely defined microchannels that spatially constrict the aqueous and oil phases. External syringe pumping controls the flow rates of aqueous and oil phases. This flow behavior within the microchannels further dictates the size and uniformity of the microgels.58 We first tested the effect of oil phase flow rate at a fixed aqueous phase flow rate of 10 µL min−1. At a lower oil phase flow rate (e.g., 30 µL min−1), the microgels formed were larger and with more polydispersity (Fig. 3A, B and Table S1). Increasing the oil phase flow rate to 40 µL min−1 resulted in a significant reduction in both the average microgel diameter and the PDI. No further difference was found when the oil-phase flow rate exceeded 50 µL min−1. We then fixed the oil-to-aqueous phase ratio at 5 while increasing the aqueous phase flow rate from 14 to 40 µL min−1. A higher aqueous phase flow rate would increase the droplet generation throughput. We found that the aqueous phase flow rate did not drastically alter the size of the microgels, but higher PDIs were obtained at above 18 µL min−1 (Fig. 3C and Table S2). After the above optimization process, we selected an aqueous and oil flow rate of 18 and 90 µL min−1, respectively, for the remaining experiments in this study.3.4. Dynamic stiffening of microgels via hydrazone bonding
GelNB-CH is a multifunctional macromer amenable to multiple orthogonal reactions, including thiol-norbornene photo-click reaction (for hydrogel crosslinking), tetrazine-norbornene iEDDA click reaction (for microgel annealing), and hydrazone bonding (for dynamic stiffening). After identifying suitable parameters for fabricating uniform GelNB-CH microgels with thiol-norbornene photo-click gelation, we evaluated hydrogel stiffening using initiator-free hydrazone bonding with an aldehyde-containing reagent (e.g., oDex, Fig. 4A).15 The effect of oDex-mediated hydrazone bonding on hydrogel stiffness was first evaluated using GelNB-CH hydrogels with an elastic shear modulus of ∼2000 Pa.15 Without oDex, the stiffness of GelNB-CH hydrogels decreased over 48 hours of incubation due to hydrogel swelling. In the presence of a sufficiently high amount of oDex (e.g., 0.1 wt%), however, hydrogel stiffness increased gradually and reached ∼8000 Pa, a 4-fold increase from before stiffening (Fig. 4B). As oDex reacted with GelNB-CH, the overall polymer density in the hydrogels increased, resulting in smaller hydrogel mesh sizes, lower swelling, and hence higher stiffness.20 The formation of hydrazone bonding in oDex-stiffened hydrogels was verified via strain-sweep rheometry (Fig. 4C). The stiffness of GelNB-CH hydrogels without additional hydrazone bonding was independent of applied strain. However, hydrogels infused with higher oDex content (e.g., 0.05 and 0.1 wt%) exhibited strain-induced softening, indicating that some of the chemical bonds in these hydrogels were ‘ruptured’ at higher strains. It is worth noting that, while hydrazone bonding is not covalent, we have shown that oDex-stiffened GelNB-CH hydrogels were stable for at least 72 hours without any drop in shear moduli.15 Another factor to consider is that, while hydrazone bonds are not covalent, these bonds were formed within a pre-crosslinked hydrogel via thiol-norbornene photo-click reaction. In other words, the bonding between carbohydrazide on GelNB-CH and aldehyde on oDex was restricted within the hydrogels, thereby increasing their local concentration and bond stability. In another study, hydrogels crosslinked by pure carbohydrazide-aldehyde bonding were stable for at least 7 days for supporting microvascular formation.59 Nonetheless, the degree of dynamic stiffening (from 2 to 10 kPa) was relevant towards modeling PDAC progression.60Next, the effect of hydrazone-induced dynamic stiffening was evaluated using microgels. Microgels were fabricated from 5 wt% GelNB-CH and 1 wt% PEG4SH and washed with PBS, with an average diameter of ∼163 μm before stiffening (Fig. 4D). After 24 hours of oDex incubation, the average diameter of microgels decreased to ∼100 μm for 0.1 wt% oDex (Fig. 4D and Fig. S1). The difference between the largest and smallest microgels within a population is similar between non-stiffened (e.g., ∼72 μm difference) and highly stiffened (e.g., ∼77 μm difference) microgels due to the initial uniformity of the microgels. This uniformity is maintained after stiffening, regardless of oDex concentration (e.g., PDI values between ∼0.008 and ∼0.01, Table S3).
3.5. Annealing and dynamic stiffening of granular hydrogels
In this study, GelNB-CH microgels were crosslinked by thiol-norbornene photoclick reaction using only a small portion of the norbornene group (up to 15%). The remaining norbornene moiety was leveraged for annealing microgels via tetrazine-norbornene iEDDA click reaction. An advantage to using this additional click chemistry is that it is independent of light penetration, allowing the creation of thicker gel structures.61 Microgels were annealed into a scaffold using cycloaddition between electron-poor tetrazine and electron-rich norbornene (Fig. 5A). To achieve annealing, microgels were vortexed in 0.5 wt% PEG4Tz solution, followed by centrifugation at 6000 rpm for 90 seconds and incubation for a minimum of two hours. It is worth noting that our prior work on GelNB-CH granular hydrogels utilized hydrazone-stiffened microgels before their annealing into granular hydrogels by iEDDA click reaction.14 The orthogonality of hydrazone bonding and iEDDA click reaction would allow for an annealing-first strategy, where the entire annealed granular hydrogels were subjected to hydrazone-induced stiffening. To this end, the annealed scaffolds were incubated in CF488A-oDex for 24 hours to induce dynamic stiffening. Compared with non-stiffened control, CF488A-oDex uniformly distributed in the annealed granular hydrogels, suggesting that the diffusion of oDex into the granular scaffold was not hindered (Fig. 5B). The void fraction of the stiffened granular scaffold was maintained at around 35% regardless of the process of scaffold formation or degree of stiffening (Fig. 5C, D and S2). This is due to the initial uniformity of the microgels before any stiffening. Additionally, particle packing theory expects that monodisperse microgels produce similar void fractions when packed into a scaffold.62 Conversely, microgels formed by inverse suspension created microgels with a much wider diameter range distribution (i.e., ∼250 μm).14 The void fraction of these scaffolds with pre-stiffened microgels created void fractions ranging from ∼28% up to ∼40%.143.6. Effect of dynamic stiffening on CAF and PCC spheroids
CAFs are known to facilitate cancer cell spreading in PDAC and are activated in response to increased matrix stiffness.5 Here, we evaluated the effect of dynamic granular hydrogel stiffening on pancreatic CAFs and cancer cells. CAF spheroids were assembled in Aggrewell plate, recovered, and embedded in the void space of granular scaffolds, followed by annealing using PEG4Tz via iEDDA click reaction. After annealing, CAF-laden granular hydrogels were incubated in 0.1% oDex for 24 hours to induce dynamic stiffening, followed by tracking of cell morphology for four days (Fig. 6A and B). CAFs dissipated from the central spheroid region within two days after embedding. In stiffened bulk hydrogels, the spreading of CAFs was limited due to spatial restriction from the reduced hydrogel mesh size.15 On the other hand, in granular hydrogels annealed from stiffened and smaller microgels, CAF outgrowth was limited due to the larger void fraction.14 In contrast, dynamic stiffening of granular hydrogel did not significantly alter their void fractions (Fig. 5D), allowing the observed differences to be attributed to the variation in granular gel stiffness. Over four days, CAFs cultured within the soft granular scaffolds grew to over 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 μm2 in maximum-intensity projected area per cellular cluster (Fig. 6C). Conversely, CAFs within the stiffened scaffolds grew outward more slowly, with an average maximum-intensity projected area of ∼20
000 μm2 in maximum-intensity projected area per cellular cluster (Fig. 6C). Conversely, CAFs within the stiffened scaffolds grew outward more slowly, with an average maximum-intensity projected area of ∼20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 μm2. Cell migration is affected by matrix stiffness and the cell's ability to generate traction force.63 For example, cells form long, spindling filopodia to probe the stiffness of their surroundings and form lower stability attachments if the stiffness is lower than ‘desired’.63,64 Meanwhile, cells form more secure lamellipodia if the matrix stiffness is ideal for migration.63,64 As these cellular processes were not easily discerned in 3D granular hydrogels, we prepared 2D flat hydrogels (with the same formulations as those in microgels) on which CAF spheroids were seeded. CAF-seeded hydrogels were stiffened with 0.1% oDex supplemented in the culture medium. Over four days, the maximum-intensity projection areas were evaluated, which revealed similar measurements between cell clusters (i.e., spheroids and the outgrown cells on the hydrogel surface (Fig. S3A and S3B)). However, CAFs on the soft hydrogel surface appeared longer and spindled, while CAFs on the stiff hydrogel surface appeared rounder. Immunostaining of F-actin revealed a brighter appearance of actin bundles in the spindling cells on the soft hydrogel. Meanwhile, F-actin appeared duller in the rounder cells on the stiff hydrogel. These morphological observations indicate that CAFs might be making temporary attachments to the hydrogel matrix as they ‘explored’ their local microenvironment.64 On the other hand, CAFs on the stiffened hydrogel formed more stable adhesions to the matrix, potentially generating stronger traction forces.64
000 μm2. Cell migration is affected by matrix stiffness and the cell's ability to generate traction force.63 For example, cells form long, spindling filopodia to probe the stiffness of their surroundings and form lower stability attachments if the stiffness is lower than ‘desired’.63,64 Meanwhile, cells form more secure lamellipodia if the matrix stiffness is ideal for migration.63,64 As these cellular processes were not easily discerned in 3D granular hydrogels, we prepared 2D flat hydrogels (with the same formulations as those in microgels) on which CAF spheroids were seeded. CAF-seeded hydrogels were stiffened with 0.1% oDex supplemented in the culture medium. Over four days, the maximum-intensity projection areas were evaluated, which revealed similar measurements between cell clusters (i.e., spheroids and the outgrown cells on the hydrogel surface (Fig. S3A and S3B)). However, CAFs on the soft hydrogel surface appeared longer and spindled, while CAFs on the stiff hydrogel surface appeared rounder. Immunostaining of F-actin revealed a brighter appearance of actin bundles in the spindling cells on the soft hydrogel. Meanwhile, F-actin appeared duller in the rounder cells on the stiff hydrogel. These morphological observations indicate that CAFs might be making temporary attachments to the hydrogel matrix as they ‘explored’ their local microenvironment.64 On the other hand, CAFs on the stiffened hydrogel formed more stable adhesions to the matrix, potentially generating stronger traction forces.64
To investigate the spreading of PCCs within the granular hydrogel environment, we formed spheroids of pancreatic cancer patient-derived cell line Pa03C in Aggrewell plate and embedded them within microgels before scaffold annealing. Singular CAFs were seeded onto the scaffold surface. Scaffolds were stiffened with 0.1% oDex for 24 hours, followed by tracking of cell morphology for 4 days. Dispersed CAFs spread out quickly and formed an extensive interconnected network around microgels and the PCC spheroid (Fig. 6D). Interestingly, PCCs began to migrate away from their original spheroids within 4 days of culture, a behavior not observed in the absence of CAF in bulk hydrogels.15 CAF-induced PCC spreading was also observed in a previous study, in which CAFs and PCCs were mixed within the same spheroids.14 Regardless of how CAFs were presented (i.e., dispersed in the void space outside of PCC spheroids or in mixed spheroids), their presence increased PCC migration (Fig. 6E). However, no statistical significance was found in PCC spreading area between soft and stiffened granular hydrogels (Fig. 6F).
In the TME, cancer cells proliferate and secrete ECM that stiffens the matrix, leading to compression of stromal tissue65 that may restrict cancer cell migration.66 However, higher matrix stiffness can activate pancreatic stellate cells to become CAFs,5 which not only remodel the TME but also facilitate cancer cell spreading.67 Furthermore, CAFs secrete pro-migratory cytokines such as transforming growth factor-beta (TGFβ) and fibroblastic growth factor (FGF), which also promote the migratory behavior of PCCs.68 Conversely, the ECM deposited by CAFs may also restrict cancer cell metastasis by localizing deposition around the PCCs and within the tumor.69,70 The increasing stiffness of the tumor leads to a poorer prognosis of PDAC,71 with increasing stiffness of the PCCs demonstrating a more invasive phenotype.72 In the current study, we show that dynamically stiffened granular hydrogels restricted CAF spreading (Fig. 6C), likely due to increased cell adhesion strength. However, the difference in CAF adhesion did not alter PCC migration (Fig. 6F). This was different from what we observed in a previous study, in which a dynamically stiffened 2D hydrogel surface seeded with CAFs induced PCC migration away from spheroids.73 The discrepancy in PCC migration results may be attributed to the different dimensionality of the culture format (i.e., flat 2D surface vs. 3D granular hydrogel) that led to different CAF responses in cytokine secretion or additional ECM deposition. Future studies will focus on integrating additional pancreatic TME components into the dynamic granular hydrogel system to tease out the molecular mechanisms.
4. Conclusion
In conclusion, this work demonstrates a biofabrication workflow for creating dynamic granular hydrogels. Microgels were first fabricated using GelNB-CH, a dually functionalized gelatin with modular reactivities, via thiol-norbornene photo-click reaction. Microfluidic parameters for forming microgels with uniform sizes were identified, including macromer concentrations and relative flow rates of the oil and aqueous phases. Next, the GelNB-CH microgels were annealed using PEG4Tz via iEDDA click reaction. Finally, the annealed granular hydrogels were dynamically stiffened by oDex-mediated hydrazone bonding. The annealing and dynamic stiffening processes were cytocompatible for entrapping pancreatic CAFs and PCCs. Finally, dynamic stiffening of GelNB-CH granular hydrogels reduced CAF spreading without hindering CAF-induced PCC migration. Future work will focus on adapting this dynamic granular hydrogel platform to induce physical stress/confinement and to evaluate the effect of cytokine-mediated PCC migration.Author contributions
E. F.: conceptualization, formal analysis, data curation, methodology, software, validation, visualization, writing – original draft, writing – review & editing. C.-C. Lin: conceptualization, formal analysis, funding acquisition, methodology, project administration, resources, supervision, visualization, writing – review & editing.Conflicts of interest
The authors declare no conflict of interest.Data availability
The data supporting this article have been included as part of the supplementary information (SI). Supplementary information is available. See DOI: https://doi.org/10.1039/d5bm00997a.Acknowledgements
The authors thank Dr Melissa Fishel for providing the CAFs and Pa03Cs. This project was supported in part by the US Department of Defense (W81XWH2210864) and National Institutes of Health (R01CA227737).References
- American Cancer Society, ACS Facts and Figures, 2025 Search PubMed.
- K. Yamashita and Y. Kumamoto, Int. J. Mol. Sci., 2024, 25, 6003 CrossRef CAS PubMed.
- D. Wang, Y. Li, H. Ge, T. Ghadban, M. Reeh and C. Güngör, Cancers, 2022, 14, 3998 Search PubMed.
- P. Gamradt, K. Thierry, M. Masmoudi, Z. X. Wu, H. Hernandez-Vargas, S. Bachy, T. Antonio, B. Savas, Z. Hussain, R. Tomasini, P. Milani, P. Bertolino and A. Hennino, PNAS Nexus, 2023, 2, 405 CrossRef PubMed.
- D. Öhlund, A. Handly-Santana, G. Biffi, E. Elyada, A. Almeida, M. Ponz-Sarvise, V. Corbo, T. Oni, S. Hearn, E. J. Lee, I. I. Chio, C. Hwang, H. Tiriac, L. Baker, D. Engle, C. Feig, A. Kultti, M. Egeblad, D. Fearon, J. M. Crawford, H. Clevers, Y. Park and D. Tuveson, J. Exp. Med., 2017, 214, 579–596 CrossRef PubMed.
- A. Nabavizadeh, T. Payen, A. C. Iuga, I. R. Sagalovskiy, D. Desrouilleres, N. Saharkhiz, C. F. Palermo, S. A. Sastra, P. E. Oberstein, V. Rosario, M. D. Kluger, B. A. Schrope, J. A. Chabot, K. P. Olive and E. E. Konofagou, Theranostics, 2020, 10, 4614–4626 CrossRef CAS PubMed.
- A. Rubiano, D. Delitto, S. Han, M. Gerber, C. Galitz, J. Trevino, R. M. Thomas, S. J. Hughes and C. S. Simmons, Acta Biomater., 2018, 67, 331–340 CrossRef PubMed.
- A. Raza, C. S. Ki and C.-C. Lin, Biomaterials, 2013, 34, 5117–5127 CrossRef CAS PubMed.
- C. S. Ki, H. Shih and C.-C. Lin, Biomacromolecules, 2013, 14, 3017–3026 CrossRef CAS PubMed.
- T.-Y. Lin, C. S. Ki and C.-C. Lin, Biomaterials, 2014, 35, 6898–6906 CrossRef CAS PubMed.
- C. S. Ki, T.-Y. Lin, M. Korc and C.-C. Lin, Biomaterials, 2014, 35, 9668–9677 Search PubMed.
- H. Zhang, J. Chen, X. Hu, J. Bai and T. Yin, Bioeng. Transl. Med., 2023, 8, e10518 Search PubMed.
- R. Curvello, V. Kast, M. H. Abuwarwar, A. L. Fletcher, G. Garnier and D. Loessner, Front. Digit. Health, 2021, 26, 704584 CrossRef PubMed.
- C.-Y. Chang, H. Nguyen, E. Frahm, K. Kolaczyk and C.-C. Lin, RSC Appl. Polym., 2024, 2, 656–669 RSC.
- C. Y. Chang, H. C. Johnson, O. Babb, M. L. Fishel and C. C. Lin, Acta Biomater., 2021, 130, 161–171 Search PubMed.
- H.-Y. Liu, M. Korc and C.-C. Lin, Biomaterials, 2018, 160, 24–36 Search PubMed.
- H. D. Nguyen, H.-Y. Liu, B. N. Hudson and C.-C. Lin, ACS Biomater. Sci. Eng., 2019, 5, 1247–1256 Search PubMed.
- F. Y. Lin, C. Y. Chang, H. Nguyen, H. Li, M. L. Fishel and C. C. Lin, Mater. Today Bio, 2023, 19, 100576 Search PubMed.
- H. D. Nguyen and C.-C. Lin, Acta Biomater., 2024, 177, 203–215 Search PubMed.
- C.-C. Lin and A. T. Metters, Adv. Drug Delivery Rev., 2006, 58, 1379–1408 Search PubMed.
- C.-C. Lin and K. S. Anseth, Pharm. Res., 2009, 26, 631–643 CrossRef CAS PubMed.
- M. W. Tibbitt and K. S. Anseth, Biotechnol. Bioeng., 2009, 103, 655–663 Search PubMed.
- Z. Mahdieh, M. D. Cherne, J. P. Fredrikson, B. Sidar, H. S. Sanchez, C. B. Chang, D. Bimczok and J. N. Wilking, Biomed. Mater., 2022, 17, 045020 CrossRef CAS PubMed.
- V. G. Muir, S. Weintraub, A. P. Dhand, H. Fallahi, L. Han and J. A. Burdick, Adv. Sci., 2023, 10, e2206117 Search PubMed.
- N. Annabi, X. Zhong, C. Ji, S. Koshy, A. Khademhosseini and F. Kdehghani, Tissue Eng., Part B, 2010, 16, 1937–3376 Search PubMed . (Electronic).
- T. Molley, G. K. Jalandhra, S. R. Nemec, A. S. Tiffany, A. Patkunarajah, K. Poole, B. Harley, T. T. Hung and K. Kilian, Biomater. Sci., 2021, 9, 4496–4509 Search PubMed.
- M. Schot, M. Becker, C. A. Paggi, F. Gomes, T. Koch, T. Gensheimer, C. Johnbosco, L. P. Nogueira, A. van der Meer, A. Carlson, H. Haugen and J. Leijten, Adv. Mater., 2023, 36, 2308949 Search PubMed.
- J. Kieda, S. Appak-Baskoy, M. Jeyhani, M. Navi, K. W. Y. Chan and S. S. H. Tsai, ACS Biomater. Sci. Eng., 2023, 9, 1043–1052 Search PubMed.
- D. Ribezzi, M. Gueye, S. Florczak, F. Dusi, D. de Vos, F. Manente, A. Hierholzer, M. Fussenegger, M. Caiazzo, T. Blunk, J. Malda and R. Levato, Adv. Mater., 2023, 35, e2301673 Search PubMed.
- L. Riley, L. Schirmer and T. Segura, Curr. Opin. Biotechnol, 2019, 60, 1–8 Search PubMed.
- A. Charlet, F. Bono and E. Amstad, Chem. Sci., 2022, 13, 3082–3093 Search PubMed.
- A. C. Daly, L. Riley, T. Segura and J. A. Burdick, Nat. Rev. Mater., 2020, 5, 20–43 Search PubMed.
- D. R. Griffin, W. M. Weaver, P. O. Scumpia, D. Di Carlo and T. Segura, Nat. Mater., 2015, 14, 737–744 Search PubMed.
- D. T. Nguyen, D. I. Pedro, A. Pepe, J. G. Rosa, J. I. Bowman, L. Trachsel, G. R. Golde, I. Suzuki, J. M. Lavrador, N. T. Y. Nguyen, M. A. Kis, R. A. Smolchek, N. Diodati, R. Liu, S. R. Phillpot, A. R. Webber, P. Castillo, E. J. Sayour, B. S. Sumerlin and W. G. Sawyer, Biointerphases, 2023, 18, 021001 Search PubMed.
- A. J. Seymour, S. Shin and S. C. Heilshorn, Adv. Healthcare Mater., 2021, 10, 2100644 Search PubMed.
- T. H. Qazi, J. Wu, V. G. Muir, S. Weintraub, S. E. Gullbrand, D. Lee, D. Issadore and J. A. Burdick, Adv. Mater., 2022, 34, 2109194 Search PubMed.
- N. F. Truong, E. Kurt, N. Tahmizyan, S. C. Lesher-Pérez, M. Chen, N. J. Darling, W. Xi and T. Segura, Acta Biomater., 2019, 94, 160–172 CrossRef CAS PubMed.
- A. S. Caldwell, G. T. Campbell, K. M. T. Shekiro and K. S. Anseth, Adv. Healthc. Mater., 2017, 6, 1700254 Search PubMed.
- C. Piras, A. G. Kay, P. G. Genever and D. Smith, Chem. Sci., 2021, 12, 3958–3965 Search PubMed.
- A. E. Widener, S. Duraivel, T. E. Angelini and E. A. Phelps, Adv. NanoBiomed Res., 2022, 2, 2200030 CrossRef CAS PubMed.
- N. Di Caprio, M. D. Davidson, A. C. Daly and J. A. Burdick, Adv. Mater., 2024, 36, 2312226 CrossRef PubMed.
- B. G. Carvalho, A. Nakayama, H. Miwa, S. W. Han, L. G. de la Torre, D. Di Carlo, J. Lee, H.-J. Kim, A. Khademhosseini and N. R. de Barros, Aggregate, 2024, 5, e464 Search PubMed.
- B. N. Pfaff, L. J. Pruett, N. J. Cornell, J. de Rutte, D. Di Carlo, C. B. Highley and D. R. Griffin, ACS Biomater. Sci. Eng., 2021, 7, 422–427 Search PubMed.
- S. Bulut, D. Günther, M. Bund, C. Haats, T. Bissing, C. Bastard, M. Wessling, L. De Laporte and A. Pich, Adv. Healthcare Mater., 2023, 13, 2302957 CrossRef.
- Y. Zhang, Q. Hu, Y. Pei, H. Luo, Z. Wang, X. Xu, Q. Zhang, J. Dai, Q. Wang, Z. Fan, Y. Fang, M. Ye, B. Li, M. Chen, Q. Xue, Q. Zheng, S. Zhang, M. Huang, T. Zhang, J. Gu and Z. Xiong, Nat. Commun., 2024, 15, 3382 Search PubMed.
- S. Clara-Trujillo, L. Tolosa, L. Cordón, A. Sempere, G. Gallego Ferrer and J. L. Gómez Ribelles, Biomater. Adv., 2022, 135, 212749 CrossRef CAS PubMed.
- D. J. Atkins, J. M. Rosas, L. K. Månsson, N. Shahverdi, S. S. Dey and A. A. Pitenis, ACS Biomater. Sci. Eng., 2024, 10, 2177–2187 Search PubMed.
- A. C. Siciliano, S. Forciniti, V. Onesto, H. Iuele, D. D. Cave, F. Carnevali, G. Gigli, E. Lonardo and L. L. del Mercato, Adv. Healthcare Mater., 2024, 13, 2401138 CrossRef CAS.
- Z. Mũnoz, H. Shih and C.-C. Lin, Biomater. Sci., 2014, 2, 1063–1072 RSC.
- A. Jeanes and C. A. Wilham, J. Am. Chem. Soc., 1950, 72, 2655–2657 CrossRef CAS.
- J. M. Hughes, P. M. Budd, A. Grieve, P. Dutta, K. Tiede and J. Lewis, J. Appl. Polym. Sci., 2015, 132, 42061 CrossRef.
- J. Van Hoorick, A. Dobos, M. Markovic, T. Gheysens, L. Van Damme, P. Gruber, L. Tytgat, J. Van Erps, H. Thienpont, P. Dubruel, A. Ovsianikov and S. Van Vlierberghe, Biofabrication, 2020, 13, 015017 Search PubMed.
- V. T. Duong and C.-C. Lin, Macromol. Biosci., 2023, 23, 2300213 CrossRef CAS PubMed.
- M. H. Kim, H. Nguyen, C. Y. Chang and C. C. Lin, ACS Biomater. Sci. Eng., 2021, 7, 4196–4208 CrossRef CAS PubMed.
- M. R. Arkenberg, H. D. Nguyen and C.-C. Lin, J. Mater. Chem. B, 2020, 8, 7835–7855 RSC.
- W. Li, L. Zhang, X. Ge, B. Xu, W. Zhang, L. Qu, C.-H. Choi, J. Xu, A. Zhang, H. Lee and D. A. Weitz, Chem. Soc. Rev., 2018, 47, 5646–5683 RSC.
- H. Shieh, M. Saadatmand, M. Eskandari and D. Bastani, Sci. Rep., 2021, 11, 1565 Search PubMed.
- W. H. Tan and S. Takeuchi, Adv. Mater., 2007, 19, 2696–2701 CrossRef CAS.
- T. Hozumi, T. Kageyama, S. Ohta, J. Fukuda and T. Ito, Biomacromolecules, 2018, 19, 288–297 Search PubMed.
- J.-H. Zheng, Y.-H. Zhu, J. Yang, P.-X. Ji, R.-K. Zhao, Z.-H. Duan, H.-F. Yao, Q.-Y. Jia, Y.-F. Yin, L.-P. Hu, Q. Li, S.-H. Jiang, Y.-M. Huo, W. Liu, Y.-W. Sun and D.-J. Liu, Cell Rep., 2024, 43, 114633 Search PubMed.
- N. Contessi Negrini, A. Angelova Volponi, P. T. Sharpe and A. D. Celiz, ACS Biomater. Sci. Eng., 2021, 7, 4330–4346 CrossRef CAS PubMed.
- L. Riley, G. Wei, Y. Bao, P. Cheng, K. L. Wilson, Y. Liu, Y. Gong and T. Segura, Small, 2023, 19, e2303446 CrossRef PubMed.
- C. Ji and Y. Huang, Commun. Biol., 2023, 6, 1169 CrossRef PubMed.
- M. Innocenti, Cell Adhes. Migr., 2018, 12, 401–416 Search PubMed.
- T. P. Padera, B. R. Stoll, J. B. Tooredman, D. Capen, E. d. Tomaso and R. K. Jain, Nature, 2004, 427, 695–695 CrossRef CAS PubMed.
- R. F. Hwang, T. Arumugam, V. Ramachandran, K. D. Amos, A. Rivera, B. Ji, D. B. Evans, C. D. Logsdon and T. Moore, Cancer Res., 2008, 68, 918–926 CrossRef CAS PubMed.
- C. Gaggioli, S. Hooper, C. Hidalgo-Carcedo, R. Grosse, J. F. Marshall, K. Harrington and E. Sahai, Nat. Cell Biol., 2007, 9, 1392–1400 CrossRef CAS PubMed.
- B. Erdogan and D. J. Webb, Biochem. Soc. Trans., 2017, 45, 229–236 CrossRef CAS PubMed.
- M. Nurmik, P. Ullman, F. Rodriguez, S. Haan and E. Letellier, Int. J. Cancer, 2019, 146, 895–905 CrossRef PubMed.
- D. Pankova, Y. Chen, M. Terajima, M. J. Schliekelman, B. N. Baird, M. Fahrenholtz, L. Sun, B. J. Gill, T. J. Vadakkan, M. P. Kim, Y. H. Ahn, J. D. Roybal, X. Liu, E. R. Parra Cuentas, J. Rodriguez, I. I. Wistuba, C. J. Creighton, D. L. Gibbons, J. M. Hicks, M. E. Dickinson, J. L. West, K. J. Grande-Allen, S. M. Hanash, M. Yamauchi and J. M. Kurie, Mol. Cancer Res., 2016, 14, 287–295 CrossRef CAS PubMed.
- R. Alvarez, E. Garcia-Garcia, P. P. Lopez-Casas, D. Megias, C. Guerra, M. Muñoz, Y. Quijano, A. Cubillo, J. Rodriguez-Pascual, C. Plaza, E. de Vicente, S. Prados, S. Tabernero, M. Barbacid, F. Lopez-Rios, M. Hidalgo and M. Musteanu, Br. J. Cancer, 2013, 109, 926–933 CrossRef CAS PubMed.
- A. V. Nguyen, K. D. Nyberg, M. B. Scott, A. M. Welsh, A. H. Nguyen, N. Wu, S. V. Hohlbauch, N. A. Geisse, E. A. Gibb, A. G. Robertson, T. R. Donahue and A. C. Rowat, Integr. Biol., 2018, 8, 1232–1245 CrossRef PubMed.
- C. Y. Chang and C. C. Lin, ChemBioChem, 2025, 26, e202400955 CrossRef CAS PubMed.
| This journal is © The Royal Society of Chemistry 2026 |






