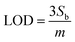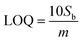Development of a cellulose acetate-based polymeric membrane for the extraction and quantification of fluoxetine and norepinephrine in biological samples
Seyedeh
Susan Sayyedi
a,
Tahere
Khezeli
 *a and
Ali
Daneshfar
b
*a and
Ali
Daneshfar
b
aDepartment of Chemistry, Faculty of Science, Ilam University, Ilam, 69315-516, Iran. E-mail: t.khezeli@ilam.ac.ir
bDepartment of Chemistry, Lorestan University, Khoramabad, Iran
First published on 15th December 2025
Abstract
In this study, a cellulose acetate based polymer membrane was synthesized via wet phase inversion and applied as an adsorptive medium for the extraction and determination of fluoxetine (FL) and norepinephrine (NE) from human blood serum and urine. The membrane was fabricated by incorporating polyvinylpyrrolidone (PVP) as a pore-forming agent, ethylene glycol dimethacrylate (EGDMA) as a cross-linker monomer, azobisisobutyronitrile (AIBN) as an initiator, and polyacrylonitrile (PAN) as a structural support. Structural and morphological characterization was performed using Fourier Transform Infrared (FT-IR), X-ray diffraction (XRD), and Field Emission Scanning Electron Microscopy (FE-SEM), confirming the successful formation of an amorphous, porous membrane with suitable functional groups for adsorption. Extraction parameters were systematically optimized using a Central Composite Design (CCD). The method exhibited excellent linearity for NE (0.1–200 µg L−1, R2 = 0.9984) and FL (0.5–200 µg L−1, R2 = 0.9992). Limits of detection were 0.05–0.2 µg L−1, while limits of quantification ranged from 0.16 to 0.66 µg L−1. Precision was demonstrated with intra- and inter-day RSD values below 5.6%. Recoveries from spiked urine and serum samples exceeded 90.30% with RSD < 5.7%.
Introduction
Fluoxetine (FL) is a selective serotonin reuptake inhibitor that primarily functions by inhibiting serotonin reuptake at the presynaptic membrane, leading to elevated synaptic serotonin concentrations.1 It has been primarily utilized for the treatment of depression and has found extensive application in diverse therapeutic areas, including obsessive-compulsive disorder, appetite modulation and weight management in adults, enhancing post-stroke recovery, ameliorating neurocognitive functions in children with Attention-Deficit/Hyperactivity Disorder (ADHD), and mood regulation.2–5 Furthermore, studies indicate that FL influences bone and its mechanical properties in stressed rodent models, highlighting potential therapeutic benefits extending beyond its established psychiatric indications.6 Collectively, the multifaceted role of FL in addressing a spectrum of conditions underscores its significance in clinical practice. FL exhibits extensive binding to plasma proteins, predominantly albumin and alpha-1-acid glycoprotein. Consequently, its plasma concentration remains elevated for several weeks following the initiation of therapy. The protracted half-life and slow renal elimination kinetics of both the parent drug and its metabolite suggest drug accumulation, which can lead to potential drug–drug interactions with other co-administered medications.7Norepinephrine (NE), also identified as noradrenaline (NA), functions as a crucial monoamine neurotransmitter, first characterized in the 1940s by Ulf von Euler. This catecholamine is critically involved in the modulation of arousal, attention, cognitive function, and stress responses. Furthermore, NE operates hormonally within the peripheral nervous system, particularly during the acute stress response, often termed the “fight or flight” mechanism. Consequently, its pivotal role positions it as a significant pharmacological target in the therapeutic management of various psychiatric, neurological, and cardiovascular disorders.7 NE, a significant monoamine neurotransmitter in both the central nervous system (CNS) and the sympathetic nervous system, plays a crucial role in diverse physiological and pathophysiological processes. It is involved in regulating cortical and hippocampal neural circuits and modulating the immune system. Furthermore, reduced cerebral NE levels are currently recognized as an underlying factor in major depressive disorder. Despite its multifaceted importance, the development of methods enabling high spatio-temporal resolution detection of NE in living systems remains a significant challenge.8 Nevertheless, NE has been shown to enhance cardiac function, thereby underscoring its therapeutic potential.
Given that both FL and NE occur at trace levels in biological matrices, reliable quantification requires an efficient sample preparation step prior to instrumental analysis. Traditionally, classical liquid–liquid extraction (LLE) and solid-phase extraction (SPE) have been employed for this purpose; however, these approaches are often labor-intensive, time-consuming, and solvent-consuming, and they may suffer from limited selectivity and matrix interference.9 Consequently, growing attention has been directed toward the development of new techniques such as membrane-based extraction, which provide higher efficiency, improved selectivity, and greater environmental compatibility compared with conventional methods.
The urgent demand for greener, faster, and simpler analytical workflows has catalyzed the emergence of innovative sample preparation technologies based on sustainable materials, particularly cellulose membranes and discs. Methods such as the Rotating Paper Disc (RPD), Cellulose Pad Solid-Phase Extraction (CPSE), and Dispersive ICP (DICP) represent a paradigm shift towards miniaturized and solvent-minimized solid-phase microextraction (SPME) techniques. RPD utilizes the mechanical rotation of a cellulose disc to dramatically enhance mass transfer kinetics, facilitating rapid extraction of analytes from complex liquid samples before direct analysis. Similarly, CPSE provides a highly effective, portable platform for solid-phase preconcentration, where the analyte is efficiently retained on a small cellulose pad, minimizing solvent use and simplifying subsequent elution or direct introduction. DICP, often employed for trace element analysis, leverages the cellulosic material's extensive surface area and functional groups for dispersive solid-phase extraction (D-SPE), allowing for highly efficient analyte enrichment prior to instrumental detection, collectively streamlining sample handling and significantly accelerating the overall analytical process.10,11
Membrane extraction (ME) is one such technique, employed for the separation and enrichment of compounds.12,13 The operational principle involves the diffusion of analyte molecules across solvent-impregnated hydrophobic porous membranes. This diffusion is driven by gradients, such as concentration differences or electrical potential, between the donor and acceptor phases.14 Common membrane materials include polyamide and polysulfone, which are selected based on the specific compounds targeted for extraction.13
The characteristics of membranes, including polymer type and physicochemical properties, play a key role in extraction efficiency, selectivity, and process stability. Employing suitable polymers in membrane fabrication enables precise design and performance optimization. Many conventional polymers are non-degradable, leading to undesirable environmental consequences such as air and groundwater pollution. As a result, naturally derived biodegradable polymers, such as cellulose acetate (CA), are supplanting non-degradable polymers. CA is a semi-synthetic biodegradable polymer produced via the chemical reaction of cellulose with acetic anhydride and acetic acid in the presence of sulfuric acid. CA possesses several advantageous characteristics, including thermal stability, heat resistance, water insolubility, non-toxicity, favorable mechanical stability, biocompatibility, biodegradability, relatively low cost, ease of processing, excellent chemical resistance, and good film-forming qualities. Its properties are highly dependent on the degree of esterification, quantified by the number of acetate groups replacing hydroxyl groups, which determines whether the final compound is an acetate, diacetate, or triacetate.14,15 CA-based membranes are utilized across various fields, including gas separation and removal, pharmaceutical industries, adsorption, water desalination, and wastewater treatment, employing diverse membrane separation processes such as nanofiltration, ultrafiltration, microfiltration, and reverse osmosis.16–18
In this research, a novel CA-based polymer membrane was synthesized for the first time using the wet phase inversion method. This polymer membrane was subsequently evaluated as a adsorptive phase in the determination and extraction of FL and NE from human blood serum and urine samples using high-performance liquid chromatography with ultraviolet detection (HPLC-UV). During the membrane preparation, PVP was incorporated as a pore-forming agent, EGDMA as a cross-linking monomer agent, AIBN as an initiator, and PAN, which provides structural support through its polar cyano groups, was used to formulate the polymer solution. Dimethylformamide (DMF) was employed as the casting solvent for membrane preparation. To the best of our knowledge, a suitable new polymer membrane for the targeted extraction and separation of FL and NE in biological matrices has not been previously reported. Several variables influencing optimal extraction performance, including the number of extraction cycles, elution solvent characteristics, salt concentration, pH, and elution solvent volume, were systematically evaluated and optimized using a Central Composite Design (CCD) and response surface methodology (RSM).
Experimental procedure
Materials and reagents
HPLC-grade solvents, including acetonitrile, ethanol, methanol, water, and dimethylformamide (DMF), were supplied by Merck (Darmstadt, Germany). EGDMA (97.5%), AIBN (99.0%), PAN, PVP with an average molecular weight of 25![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 g mol−1, CA, FL, and NE were all purchased from Sigma-Aldrich (St. Louis, MO, USA). Primary stock solutions of FL and NE were prepared at a concentration of 100 mg L−1 in deionized water and stored in the refrigerator. These stock solutions were subsequently diluted with double-distilled water to obtain working solutions at the desired concentrations for analysis.
000 g mol−1, CA, FL, and NE were all purchased from Sigma-Aldrich (St. Louis, MO, USA). Primary stock solutions of FL and NE were prepared at a concentration of 100 mg L−1 in deionized water and stored in the refrigerator. These stock solutions were subsequently diluted with double-distilled water to obtain working solutions at the desired concentrations for analysis.
Instrumentation
FL and NE analysis were performed using an HPLC system (model SCL-10Avp, Shimadzu, Japan). This system comprised a UV detector (Model SPD-10Avp) set at a wavelength of 254 nm, a dual solvent pump (Model LC-10Avp), an injection valve (Model EIG 001), and a Eurospher 100-5 C8 column (150 mm length, 4.6 mm internal diameter, 5 µm particle size). Optimal chromatographic separation was achieved with a mobile phase consisting of acetonitrile and water (50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50, v/v) at a flow rate of 0.5 mL min−1. pH measurements were conducted using a Metrhom Model 780 digital pH meter (Switzerland) equipped with an Ag/AgCl glass electrode. The synthesized membrane was characterized for functional groups using a Bruker-vertex 70 Fourier Transform Infrared (FT-IR) spectrometer (Germany). X-ray diffraction (XRD) patterns of the new membrane were obtained using a PHILIPS PW1730 diffractometer (Netherlands) equipped with a Cu Kα radiation source (λ = 1.541838 Å) operating at 30 kV, with 2θ ranging from 5° to 80°. Field Emission Scanning Electron Microscopy (FE-SEM) images of the fabricated membrane were acquired using a TESCAN MIRA II FE-SEM instrument (Czech Republic).
50, v/v) at a flow rate of 0.5 mL min−1. pH measurements were conducted using a Metrhom Model 780 digital pH meter (Switzerland) equipped with an Ag/AgCl glass electrode. The synthesized membrane was characterized for functional groups using a Bruker-vertex 70 Fourier Transform Infrared (FT-IR) spectrometer (Germany). X-ray diffraction (XRD) patterns of the new membrane were obtained using a PHILIPS PW1730 diffractometer (Netherlands) equipped with a Cu Kα radiation source (λ = 1.541838 Å) operating at 30 kV, with 2θ ranging from 5° to 80°. Field Emission Scanning Electron Microscopy (FE-SEM) images of the fabricated membrane were acquired using a TESCAN MIRA II FE-SEM instrument (Czech Republic).
Polymer membrane preparation
For the preparation of the membrane, 0.3 g of CA was dissolved in 6 mL of DMF, serving as a suitable solvent. To this solution, 0.1 g of PAN, which provides structural support through its polar cyano groups, 280 µL of EGDMA, 0.4 g of PVP, and 0.025 g of AIBN were added. To facilitate the synthesis of the polymeric membranes, the resulting mixture was stirred in a water bath at 50 °C at 600 rpm for 30 min. After a one-hour resting period, the mixture was cast as thin circular films onto a glass plate. These films were then placed in a water vapor bath at 60 °C for 2 h to achieve stabilization and coagulation. Following detachment, the membrane was immersed in deionized water for 24 h. Given PVP's water-solubility, the membrane is immersed in deionized water, a process engineered to leach the PVP from the material matrix to the formation of a well-defined porous structure within the membrane. The detailed steps for membrane preparation are schematically presented in Fig. 1.Membrane extraction method for FL and NE
The comprehensive steps for setting up the filtration system and the subsequent extraction of FL and NE are illustrated in Fig. 2. Initially, the prepared membrane was securely positioned within the filtration unit using a clamp. Subsequently, 50 mL of an aqueous solution containing FL and NE, each at a concentration of 100 µg L−1, was passed through the membrane. This filtration process was repeated four times. During this procedure, the analytes were effectively extracted and adsorbed onto the polymer membrane. For the elution of the adsorbed analytes, the membrane was rinsed with 4 mL of methanol. The collected elution phase was then evaporated in an oven at 60 °C. The remaining residue was redissolved in 100 µL of methanol and injected into the HPLC-UV system for quantitative analysis.Preparation of urine and serum samples
Adult volunteers provided urine and serum samples with their informed consent, in line with the ethical standards of the declaration of Helsinki. The study included no invasive techniques, and no personally identifiable information was gathered. According to institutional guidelines, ethical approval was not necessary for this form of low-risk sample collection. No special pre-treatment was performed on human urine sample prior to the membrane procedure. Instead, 5 mL aliquots of urine were filtered using filter paper and subsequently stored at 4 °C until analysis. Blood serum sample was also initially filtered. For protein precipitation, a 5 mL serum aliquot was mixed with 20 mL of acetonitrile. Following this, the supernatant was separated by centrifugation and stored at 4 °C. After these preparatory steps, a specific volume of FL and NE standard solutions was spiked into both the prepared urine and serum samples. The spiked samples were then diluted to a final volume of 50 mL with deionized water, ensuring the target analyte concentrations were within the established calibration curve range.Results and discussion
Polymerization mechanism
The polymerization involved in the membrane preparation follows a free radical polymerization mechanism, initiated by AIBN and employing EGDMA as the cross-linking monomer.19,20 Upon heating at 50 °C, AIBN generate free radicals, which attack the vinyl double bonds of EGDMA, initiating the polymerization process. The propagation stage proceeds through successive addition of EGDMA molecules, leading to the formation of a three-dimensional cross-linked polymeric network. This cross-linked structure is formed within the CA and PAN matrix, providing mechanical stability to the resulting composite membrane. During the final immersion step in deionized water, most of the PVP serving as a pore-forming agent is leached out, leaving behind a well-defined porous architecture.Characterization of membrane
FT-IR spectroscopy was utilized to identify the functional groups present in the structure of the synthesized CA-based membrane, developed for the separation of FL and NE. The spectroscopic results, as presented in Fig. 3, clearly indicated the presence of various functional groups attributed to the membrane's constituent components. A relatively broad band was observed at 3463 cm−1, assigned to the stretching vibration of the hydroxyl group (–OH), which could originate from free hydroxyl groups within the CA structure.21 The C–H stretching vibration was also detected at 2927 cm−1, corresponding to the methylene and methyl groups in the polymeric chains of the various membrane components.21 A distinct peak appeared at 2245 cm−1, attributed to the stretching vibration of the nitrile triple bond (C![[triple bond, length as m-dash]](https://www.rsc.org/images/entities/char_e002.gif) N), thereby confirming the presence of the nitrile group in the PAN structure. Furthermore, a strong band at 1750 cm−1 was observed, indicative of ester carbonyl (C
N), thereby confirming the presence of the nitrile group in the PAN structure. Furthermore, a strong band at 1750 cm−1 was observed, indicative of ester carbonyl (C![[double bond, length as m-dash]](https://www.rsc.org/images/entities/char_e001.gif) O) stretching, which can be attributed to the ester groups present in the EGDMA monomer and CA. Bands observed at 1440 cm−1 and 1377 cm−1 are, respectively, attributed to the bending vibrations of CH2 and CH3 groups.21 In the mid-range of the spectrum, specifically at 1162 cm−1 and 1050 cm−1, bands of medium to high intensity were observed, corresponding to the stretching vibrations of C–O–C (ether) and C–O (ester) bonds, primarily originating from the ether and ester groups within the CA and EGDMA structures. In the low-frequency region (900–500 cm−1), bands appeared at 903, 604, and 475 cm−1, can be related to out-of-plane C–H bending vibrations. Overall, the presence of these characteristic bands confirms the expected polymeric structure and indicates the successful polymerization of the monomers employed in the membrane's synthesis.
O) stretching, which can be attributed to the ester groups present in the EGDMA monomer and CA. Bands observed at 1440 cm−1 and 1377 cm−1 are, respectively, attributed to the bending vibrations of CH2 and CH3 groups.21 In the mid-range of the spectrum, specifically at 1162 cm−1 and 1050 cm−1, bands of medium to high intensity were observed, corresponding to the stretching vibrations of C–O–C (ether) and C–O (ester) bonds, primarily originating from the ether and ester groups within the CA and EGDMA structures. In the low-frequency region (900–500 cm−1), bands appeared at 903, 604, and 475 cm−1, can be related to out-of-plane C–H bending vibrations. Overall, the presence of these characteristic bands confirms the expected polymeric structure and indicates the successful polymerization of the monomers employed in the membrane's synthesis.
As shown in the FE-SEM micrographs (Fig. 4), the polymer membrane exhibits a porous morphology characterized by irregularly shaped and unevenly distributed pores. The variation in pore size and their heterogeneous distribution across the surface suggest that non-uniform phase separation occurred during the membrane formation process. Such morphological features confirm the development of a porous network structure, which is expected to play a crucial role in governing the membrane's permeability and separation efficiency.
XRD analysis was used to investigate the crystalline structure of the synthesized membrane. The resulting diffraction pattern, presented in Fig. 5, exhibited a broad peak in the 2θ range of 8 to 11°, indicative of the membrane's amorphous nature. This pattern is attributed to the inherent lack of long-range order in the constituent polymers: specifically, CA, PAN, and EGDMA, as well as the formation of a cross-linked network structure through polymerization.
Optimization of factor
To optimize the extraction process, specifically membrane performance, a comprehensive approach was adopted to overcome the limitations inherent in simple single-variable optimization. Such methods are often time-consuming and laborious, requiring extensive experimentation without adequately accounting for interactive effects among variables. Consequently, RSM, employing a CCD for experimental design, was utilized to systematically investigate critical quantitative parameters, including volume of eluent, ionic strength, and pH. A crucial aspect of this optimization involved the careful selection of an appropriate eluent to facilitate the complete desorption of analytes from the membrane. Factors such as eluent polarity, analyte solubility within the eluent, and compatibility with HPLC-UV detection were thoroughly considered in this selection. Methanol, ethanol, 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 v/v methanol/water, and 50
50 v/v methanol/water, and 50![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 50 v/v ethanol/water were thus chosen as candidate eluents, and their elution efficiencies were rigorously assessed under identical conditions. As depicted in Fig. 6, the elution efficiency decreased in the following order: methanol > ethanol > methanol/water > ethanol/water, a trend consistent with the dielectric constants of the respective solvents. Among the investigated eluents, methanol demonstrated superior effectiveness in desorbing analytes from the membrane, as clearly illustrated by the results presented in Fig. 6. Therefore, methanol was selected as the optimal eluent for all subsequent experimental stages.
50 v/v ethanol/water were thus chosen as candidate eluents, and their elution efficiencies were rigorously assessed under identical conditions. As depicted in Fig. 6, the elution efficiency decreased in the following order: methanol > ethanol > methanol/water > ethanol/water, a trend consistent with the dielectric constants of the respective solvents. Among the investigated eluents, methanol demonstrated superior effectiveness in desorbing analytes from the membrane, as clearly illustrated by the results presented in Fig. 6. Therefore, methanol was selected as the optimal eluent for all subsequent experimental stages.
Experimental design by CCD
This study employed a CCD for experimental design, modeling, and optimization. Specifically, a three-factor CCD, incorporating two central points and consisting of 16 experimental runs, was utilized. Table 1 details the experimental factors and their corresponding responses, with all experiments meticulously conducted in triplicate. The obtained responses were subsequently analyzed using a Pareto chart (Fig. 7). On this chart, the vertical dashed line represents the 95% confidence limit, signifying that variables with linear effects extending beyond this threshold exert a statistically significant influence on the experimental response. As observed, the linear effect of the pH variable clearly surpassed this confidence limit, indicating its significant impact on the experimental outcome.| Factors | High (+1) | Levels central (0) | Low (−1) |
|---|---|---|---|
| X 1 = pH | 11 | 7 | 3 |
| X 2 = NaCl | 10 | 5 | 0 |
| X 3 = volume of methanol | 5 | 4 | 3 |
| Run | X 1 | X 2 | X 3 | Average peak area |
|---|---|---|---|---|
| 8 | 11 | 10 | 5 | 686![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 791 791 |
| 6 | 11 | 0 | 5 | 606![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 823 823 |
| 1 | 3 | 0 | 3 | 345![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 678 678 |
| 13 | 7 | 5 | 3 | 443![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 210 210 |
| 15 (C) | 7 | 5 | 4 | 480![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 983 983 |
| 12 | 7 | 10 | 4 | 432![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 901 901 |
| 9 | 3 | 5 | 4 | 304![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 571 571 |
| 2 | 3 | 0 | 5 | 320![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 400 400 |
| 16 (C) | 7 | 5 | 4 | 489![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 921 921 |
| 7 | 11 | 10 | 3 | 634![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 709 709 |
| 11 | 7 | 0 | 4 | 423![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 709 709 |
| 4 | 3 | 10 | 5 | 297![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 605 605 |
| 3 | 3 | 10 | 3 | 287![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 604 604 |
| 14 | 7 | 5 | 5 | 357![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 120 120 |
| 10 | 11 | 5 | 4 | 648![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 532 532 |
| 5 | 11 | 0 | 3 | 650![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 271 271 |
Analysis of variance (ANOVA) was applied to estimate the goodness of fit and evaluate the effect of factors on the analytical response. The sum of squares (SS), degree of freedom (DF), mean square (MS), F-values, p-values, and the determination of coefficient (R2) of obtained data were analyzed by ANOVA (Table 2). R2 quantifies the proportion of variance in the dependent variable (average peak area) explained by the independent variable(s), thereby indicating the goodness of fit; a value approaching unity signifies superior model fidelity. The obtained R2 value of 0.95336 (Table 2) suggests a strong correlation between the model and the experimental data. Furthermore, an adjusted R2 value of 0.92226 further corroborates the robustness of this correlation. To precisely assess the statistical significance of the variables incorporated into the polynomial equation, an F-test was employed with a 95% confidence interval. Within the corresponding ANOVA table, a p-value below the 0.05 significance threshold indicates that the variable exerts a statistically significant effect on the analytical response under scrutiny. Based on these results, only the pH of the medium exhibited the most significant statistical impact.
| Factors | Sum of square (SS) | Degree of freedom (df) | Mean square (MS) | F-Value | p-Value |
|---|---|---|---|---|---|
| a X 1 = pH; X2 = NaCl; X3 = volume of methanol. | |||||
| (X1) | 2.793137 × 1011 | 1 | 2.793137 × 1011 | 6992.645 | 0.007613 |
| (X1)2 | 6.841399 × 109 | 1 | 6.841399 × 109 | 171.275 | 0.048550 |
| (X2) | 5.286744 × 106 | 1 | 5.286744 × 106 | 0.132 | 0.777871 |
| (X2)2 | 1.914479 × 107 | 1 | 1.914479 × 107 | 0.479 | 0.614498 |
| (X3) | 8.599409 × 108 | 1 | 8.599409 × 108 | 21.529 | 0.135138 |
| (X3)2 | 1.706939 × 109 | 1 | 1.706939 × 109 | 42.733 | 0.096637 |
| Lack of fit | 1.405272 × 1010 | 8 | 1.756590 × 109 | 43.976 | 0.116130 |
| Pure error | 3.994392 × 107 | 1 | 3.994392 × 107 | ||
| Total SS | 3.021618 × 1011 | 15 | |||
| R 2 squared | 0.95336 | ||||
| R 2 adjusted | 0.92227 | ||||
Fig. S1 presents three-dimensional diagrams, illustrating the interactions among the variables (X1 × X2, X1 × X3, X2 × X3) investigated in the experimental design. Accordingly, the pH of the aqueous solution of FL and NE was adjusted using 0.1 M HCl/NaOH, with the extraction medium's pH varied from 3 to 11. As depicted in Fig. S1, an increase in the extraction efficiency was observed as the pH rose from 3 to 11. This enhanced extraction in basic conditions is attributed to the synergistic effect of functional groups present on both the synthesized membrane and the analytes, leading to greater adsorption of FL and NE. It is well-established that the acidity or basicity of the aqueous phase significantly impacts the structure and performance of CA membranes, influencing their hydrophilicity, permeability, and reactivity.22,23 Alkaline conditions disrupt both intermolecular and intramolecular hydrogen bonds within cellulose, thereby destabilizing its crystalline structure and increasing the accessibility of its hydroxyl groups.24,25 Furthermore, the swelling of cellulose fibers under alkaline conditions induces subsequent morphological alterations and an expansion of the specific surface area, which collectively facilitate greater access to the cellulose surface and augment extraction efficiency. The utility of CA in alkaline media is also well-documented in applications such as membranes and drug delivery systems.26 Consequently, based on these critical observations, a pH of 11 was determined to be the optimal parameter for the extraction process.
The influence of salt percentage and ionic strength was investigated across a range of 0–10% w/v NaCl. As shown in Fig. S1, changing the ionic strength of the aqueous solution did not have much effect on the extraction efficiency, but since the best fit between the experimental data and the proposed model was achieved at 2.5% w/v salt, this value was reported as the optimal value. Subsequently, the optimal eluent volume was examined within a range of 3 to 5 mL. Fig. S1 indicated that the highest extraction efficiency was achieved at a volume of 4 mL. Volumes of less than 4 mL of methanol were insufficient to wash the analytes from the membrane surface and resulted in a decrease in the analytical signal.
The congruence between predicted and experimental values was quantitatively assessed using a desirability function, as detailed in Fig. 8. This methodology systematically profiles the desirability of responses by assigning predicted values of the dependent variable (average peak area) to a continuous scale ranging from 0.0 (undesirable) to 1.0 (highly desirable). As illustrated in Fig. 8, the excellent agreement observed between the experimental data (average peak area: 686![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 791) and the model-predicted data (average peak area: 664
791) and the model-predicted data (average peak area: 664![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 662) strongly affirms the robustness and practical utility of the developed predictive model. Based on this comprehensive desirability analysis, the optimal conditions for each factor were consequently determined as follows: sample solution pH (X1): 11; [NaCl] (X2): 2.5% w/v; and methanol volume (X3): 4 mL.
662) strongly affirms the robustness and practical utility of the developed predictive model. Based on this comprehensive desirability analysis, the optimal conditions for each factor were consequently determined as follows: sample solution pH (X1): 11; [NaCl] (X2): 2.5% w/v; and methanol volume (X3): 4 mL.
Validation of method
The proposed polymer membrane extraction method for FL and NE was thoroughly evaluated by assessing several key analytical figures of merit. These included the linear range, limit of detection (LOD), limit of quantification (LOQ), relative standard deviation (RSD), and extraction recovery, all of which were used to characterize the method, as detailed in Table 3. To establish the linear range, ten standard solutions of FL and NE were prepared at specified concentrations spanning 0.1 to 200 µg L−1. Following extraction using a CA membrane, these samples were injected into an HPLC-UV system, and the corresponding peak areas at each concentration were recorded. Calibration curves for NE demonstrated linearity across the 0.1–200 µg L−1 and for FL was 0.5–200 µg L−1 concentration range, achieving R2 value more than 0.9984.| Analyte | Linear range (µg L−1) | Regression equation | R 2 | LOD (µg L−1) | LOQ (µg L−1) | Repeatability and reproducibility (RSD%) (n = 3) | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Intra-day | Inter-day | ||||||||||
| 1 µg L−1 | 50 µg L−1 | 150 µg L−1 | 1 µg L−1 | 50 µg L−1 | 150 µg L−1 | ||||||
| NE | 0.1–200 |
y = 8826.3x + 18![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 745.0 745.0 |
0.9984 | 0.05 | 0.16 | 4.7 | 3.8 | 5.1 | 5.1 | 4.5 | 5.3 |
| FL | 0.5–200 |
y = 4711.1x + 10![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 803.0 803.0 |
0.9992 | 0.20 | 0.66 | 4.9 | 4.0 | 5.0 | 5.1 | 5.0 | 5.6 |
The LODs and LOQs for the target compounds were determined from 10 replicate analysis performed at the lowest calibration curve concentration (0.1 µg L−1 for NE and 0.5 µg L−1 for FL). These values were calculated using established LOD and LOQ equations, where ‘m’ represents the slope of the calibration curve and ‘Sb’ is the standard deviation of the 10 replicate measurements at the lowest calibration concentration.
Specifically, the LODs and LOQs for FL and NE ranged from 0.05–0.2 µg L−1 and 0.16–0.66 µg L−1, respectively. Both intra-day and inter-day repeatability were assessed through three consecutive experiments involving the extraction of FL and NE from three pure standard solution at a concentrations of 1, 50 and 150 µg L−1. The results of these experiments, summarized in Table 3, demonstrated calculated RSDs consistently below 5.6%.
To evaluate the practical applicability of the proposed method, it was utilized for the quantification of FL and NE in two authentic biological specimens: urine and blood serum after standard addition method. For this propose standard solutions of FL and NE were spiked into each prepared real sample, targeting concentrations of 1 and 50 µg L−1 within the range of the established calibration curves. Subsequently, the analytes were recovered using a synthesized membrane. All experiments were performed in triplicate at each concentration level to enable the calculation of RSD. As detailed in Table 4, the recovery rates for the target compounds from the real samples consistently exceeded 90.30%, with corresponding RSD values remaining below 5.7%.
| Sample | Spiking level (µg L−1) | NE | FL | ||
|---|---|---|---|---|---|
| Found (µg L−1) | Recovery% (RSD%) | Found (µg L−1) | Recovery% (RSD%) | ||
| Urine | 0 | ND | — | ND | — |
| 1 | 0.942 | 94.20 (5.3) | 0.953 | 95.30 (5.7) | |
| 50 | 49.87 | 99.75 (4.7) | 49.09 | 98.18 (3.9) | |
| Serum | 0 | ND | — | ND | — |
| 1 | 0.918 | 91.80 (5.1) | 0.902 | 90.30 (4.9) | |
| 50 | 48.12 | 96.24 (5.5) | 50.01 | 100.03 (5.3) | |
Fig. S2 presents the chromatograms obtained for standard solutions of FL and NE, as well as for urine and blood serum blank samples, following their processing with the synthesized membrane. Within the standard chromatogram, distinct peaks corresponding to FL and NE were observed at retention times of 8.3 and 15.2 min, respectively.
Comparison with other extraction methods
Table 5 compares the analytical performance of the present HPLC-UV method coupled with a membrane extraction procedure with several previously reported methods for the determination of norepinephrine (NE) and fluoxetine (FL).10,11,26–29 Prior investigations utilizing UPLC-MS/MS in conjunction with SPE or LC-MS/MS with cation-exchange SPE (CX-SPE) generally provided adequate sensitivity; however, they necessitated expensive equipment, extensive sample preparation, and costly cartridges. Although the CX-SPE/MS approach achieved a relatively low LOQ of 5 ng mL−1, its linear range (10–500 ng mL−1) was narrower compared to what is accomplished in this study.| Parameter | Ref. 10 | Ref. 11 | Ref. 26 | Ref. 27 | Ref. 28 | Ref. 29 | This work |
|---|---|---|---|---|---|---|---|
| Instrument | GC-MS | GC-MS | UPLC-MS/MS | LC-MS/MS | HPLC–PDA | GC-MS | HPLC-UV |
| Method | OS-RPD | CPSE | SPE | CX-SPE | FPSE membranes | BFS | Membrane |
| Linear range | 0.10–10 µg mL−1 | 0.12–2 µg mL−1 | 2–5000 pg mL−1 | 10–500 ng mL−1 | 0.100–20 µg mL−1 | 0.100–10 µg mL−1 | 0.1–200 µg L−1 |
| LOD | 0.022–0.025 µg mL−1 | 0.029 µg mL−1 | — | — | 0.04 µg mL−1 | 0.017–0.022 µg mL−1 | 0.05–0.20 µg L−1 |
| LOQ | 0.073–0.083 µg mL−1 | 0.095 µg mL−1 | 20 pg mL−1 | 5 ng mL−1 | 0.1 µg mL−1 | 0.056–0.072 µg mL−1 | 0.16–0.66 µg L−1 |
| R 2 | 0.9990–0.9995 | 0.9990 | >0.9908 | 0.9990 | 0.9858–0.9980 | 0.9984–0.9998 | >0.9984 |
| RSD (%) | 1.2–8.3 | <10.0 | <15.0 | <15.0 | 15.0 | 2.4–6.4 | <5.7 |
| Recovery (%) | 50.4–98.2 | 90.1 to 98.2 | 86.0–107.7 | 111.4 | 86.4–114% | 49.0–62.3% | 90.30–100.03 |
| Matrix | Blood and urine | Whole blood | Urine and serum | Urine | Whole blood, urine, and saliva | Blood and urine | Urine, serum |
| Analyte | FL | FL | NE | NE | FL | FL | NE, FL |
Meanwhile, methods involving GC-MS paired with Octanol-Supported Rotating Sorptive Paper Discs (OS-RPD) and those incorporating Cellulose Paper Sorptive Extraction (CPSE) demonstrated acceptable detection limits (0.022–0.029 µg mL−1); however, they were hindered by limited linearity, and lower recoveries for FL (around ∼50%). In contrast, the membrane extraction system developed in this work achieved dramatically higher recovery values (90.30–100.05%), showcasing its enhanced extraction efficacy. Likewise, HPLC-PDA used with Fabric Phase Sorptive Extraction (FPSE) membranes presented a wider linear range (0.1–200 µg mL−1) and commendable analytical performance, yet still showed a higher LOD of 0.04 µg mL−1 compared to the LOD attained in the current method (0.05–0.2 µg L−1), underscoring the considerably improved sensitivity of the proposed membrane-based HPLC-UV technique. Furthermore, GC-MS utilizing Biofluid Sampler (BFS) extraction exhibited acceptable outcomes but faced higher RSD values (up to 6.4%) and moderate recovery (49.0–62.0%). In summary, the developed membrane-based microextraction coupled with HPLC-UV displays numerous significant advantages over established methods, including extremely low detection limits, wide linear dynamic range (0.1–200 µg L−1), high extraction recovery, excellent precision, and simplicity, lower cost, and complete compatibility with routine laboratory procedures, in contrast to methods reliant on MS and GC. These results affirm that the proposed method serves as a highly effective, economical, and analytically robust alternative for the simultaneous quantification of NE and FL in biological samples.
Analytical greenness metric for membrane based extraction of FL and NE
The increasing imperative of Green Analytical Chemistry (GAC) has led to the development and widespread adoption of quantitative analytical greenness metric tools, which provide objective, standardized assessments of the environmental impact and sustainability profile of chemical methodologies. These metrics, such as the analytical greenness calculator (AGREE) or National Environmental Methods Index (NEMI) indices, operate by scoring various method parameters including reagent toxicity, energy consumption, and waste volume to generate a singular or visual representation of the method's overall ecological footprint. A significant advancement in this domain is the Blue Analytical Greenness Index (BAGI), which often expands the evaluation criteria beyond strict ecological parameters to incorporate crucial economic and efficiency factors, thus offering a more holistic “blue” assessment that aligns with the principles of sustainable development.30,31The number inside the BAGI pictogram represents the overall score assigned to the analytical method, which ranges from 25 (for the worst performance of the method) to 100 (for the best performance of the method). In this research, a polymeric membrane was employed for the quantification of FL and NE utilizing HPLC-UV instrumentation. The membrane's synthesis was conducted in-lab with standard laboratory equipment. The method exhibited a sample throughput of 4 analyses per hour. Preconcentration, involving both extraction and evaporation, was a requisite step to achieve the desired sensitivity. The analytical procedure necessitated a 50 mL sample aliquot and multi-step sample preparation, with all operations performed manually. Reflecting these parameters, a BAGI score of 50 was assigned to the method (Fig. S3).
Conclusion
A cellulose acetate-based polymer membrane was successfully synthesized and applied for the selective extraction and determination of FL and NE in biological samples. Structural and morphological analyses confirmed the formation of a stable, porous, and functional material suitable for adsorption. The developed method exhibited wide linear ranges, low detection limits, high precision, and excellent recovery rates in both serum and urine matrices. Compared with conventional LC-MS/MS and solid-phase extraction techniques, the proposed membrane approach provides comparable analytical performance while being simpler, cost-effective, and environmentally friendly. These results highlight the potential of biodegradable polymeric membranes as sustainable and efficient alternatives for pharmaceutical and biomedical analysis.Author contributions
SS. S., T. KH., and A. D. conceptualized the study. SS. S. prepared the membrane. Data collection was performed by SS. S., and T. KH. Data analysis was performed by T. KH. Original draft was prepared by SS. S., and T. KH. Further review and editing of the manuscript was done by SS. S., T. KH., and A. D. Resources and supervision was provided by T. KH., and A. D.Conflicts of interest
The authors affirm that they have no identifiable financial conflicts of interest or personal relationships that may have influenced the findings presented in this manuscript.Data availability
The data that support the findings of this study are available on request from the corresponding author.Supplementary information (SI) is available. See DOI: https://doi.org/10.1039/d5ay01724f.
Acknowledgements
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors. The authors thank Ilam University for supporting this research.References
- K. T. Fitzgerald and A. C. Bronstein, Top. Companion Anim. Med., 2013, 28, 13–17 CrossRef PubMed.
- B. P. Leeuwenburgh, W. A. Helbing, P. Steendijk, P. H. Schoof and J. Baan, Am. J. Physiol.: Heart Circ. Physiol., 2001, 281, H2697–H2704 CrossRef CAS PubMed.
- A. E. Serralde-Zuñiga, A. G. González-Garay, Y. Rodríguez-Carmona and G. Meléndez-Mier, Obes. Facts, 2022, 15, 473–486 CrossRef PubMed.
- P. Mushivi, A. Tsarkov, P. Petlovanyi and R. Paul, World J. Adv. Res. Rev., 2022, 15, 405–416 CrossRef CAS.
- R. W. Sommi, M. L. Crismon and C. L. Bowden, Pharmacotherapy, 1987, 7, 1–14 CrossRef CAS PubMed.
- R. W. Lam, H.-K. Wong, R. A. Kumarsing, A. N. Chua, R. C. Ho, R. S. McIntyre and C. S. Ho, Transl. Psychiatry, 2022, 12, 339 CrossRef CAS PubMed.
- L. S. Hussain, V. Reddy and C. V. Maani, Physiology, Noradrenergic Synapse, 2025 Search PubMed.
- Y. Li, J. Guo, H. Li, Y. Wang, H. Yang and D. Zhang, Research Square, 2023, 1–17 Search PubMed.
- M. S. Jagirani, O. Ozalp and M. Soylak, Crit. Rev. Anal. Chem., 2022, 52, 1343–1369 CrossRef CAS PubMed.
- B. Jain, R. Jain, S. Kaur, S. K. M. Haque, S. Sharma, M. M. Ghoneim and L. A. Al-Khateeb, Sustainable Chem. Pharm., 2024, 41, 101724 CrossRef CAS.
- B. Jain, R. Jain, A. Kabir, A. Ghosh, T. Zughaibi, V. Chauhan, S. Koundal and S. Sharma, Separations, 2023, 10, 281 CrossRef CAS.
- V. Raks, H. Al-Suod and B. Buszewski, Chromatographia, 2018, 81, 189–202 CrossRef CAS PubMed.
- T. Msagati, L. Chimuka and E. Cukrowska, Water, 2018, 34, 421 Search PubMed.
- R. Abu-Zurayk, N. Alnairat, A. Khalaf, A. A. Ibrahim and G. Halaweh, Processes, 2023, 11, 489 CrossRef CAS.
- A. Torre-Celeizabal, F. Russo, F. Galiano, A. Figoli, C. Casado-Coterillo and A. Garea, ACS Sustain. Chem. Eng., 2025, 13, 1253–1270 CrossRef CAS PubMed.
- A. Khamwichit, K. Wongsuwan and W. Dechapanya, Polymers, 2025, 17, 224 CrossRef CAS PubMed.
- V. Vatanpour, M. E. Pasaoglu, H. Barzegar, O. O. Teber, R. Kaya, M. Bastug, A. Khataee and I. Koyuncu, Chemosphere, 2022, 295, 133914 CrossRef CAS PubMed.
- I. Ounifi, F. Khaled, M. Kahloul, H. Ajari and A. Hafiane, in Cellulose – Fundamentals and Conversion into Biofuel and Useful Chemicals, IntechOpen, 2023 Search PubMed.
- D. Rapacz, K. Smolinska-Kempisty and J. Wolska, J. Environ. Chem. Eng., 2024, 12, 112159 CrossRef CAS.
- S. Haoue, H. Derdar, M. Belbachir and A. Harrane, Bull. Chem. React. Eng. Catal., 2020, 15, 221–230 CrossRef CAS.
- R. M. Silverstein and G. C. Bassler, J. Chem. Educ., 1962, 39, 546 CrossRef CAS.
- Y. Zhang, J. Xi, Y. Lou, Y. Chu, Z. Xu, H. Xiao and W. Wu, Colloids Surf., A, 2023, 676, 132116 CrossRef CAS.
- F. S. Tekin and P. Z. Çulfaz-Emecen, ACS Appl. Polym. Mater., 2023, 5, 2185–2194 CrossRef CAS.
- H. Yokota, J. Appl. Polym. Sci., 1985, 30, 263–277 CrossRef CAS.
- W. R. Kunusa, I. Isa, L. A. Laliyo and H. Iyabu, J. Phys.:Conf. Ser., 2018, 1028, 012199 CrossRef.
- M. V. T. Nguyen, N. Sherck, S. Köhler, E. Schreiner, R. Gupta, G. H. Fredrickson and M. S. Shell, Biomacromolecules, 2024, 25, 5809–5818 CrossRef CAS PubMed.
- W. Wang, T. Liu, Q. Li, E. Ji, W. Xu, S. Qiao, Y. Cui, B. Li and H. Xu, J. Chromatogr. Sci., 2025, 63, bmaf021 CAS.
- A. Tartaglia, S. Covone, E. Rosato, M. Bonelli, F. Savini, K. Furton, I. Gazioglu, C. D'Ovidio, A. Kabir and M. Locatelli, Adv. Sample Prep., 2022, 3, 100022 CrossRef.
- B. Jain, R. Jain, A. Kabir, N. Ali, M. R. Khan and S. Sharma, J. Chromatogr. B, 2024, 1247, 124327 CrossRef CAS PubMed.
- N. Manousi, W. Wojnowski, J. Płotka-Wasylka and V. Samanidou, Green Chem., 2023, 25, 7598–7604 RSC.
- Z. H. A. Aljboory, M. Ghani and J. B. Raoof, J. Chromatogr. A, 2025, 25, 465615 CrossRef PubMed.
| This journal is © The Royal Society of Chemistry 2026 |