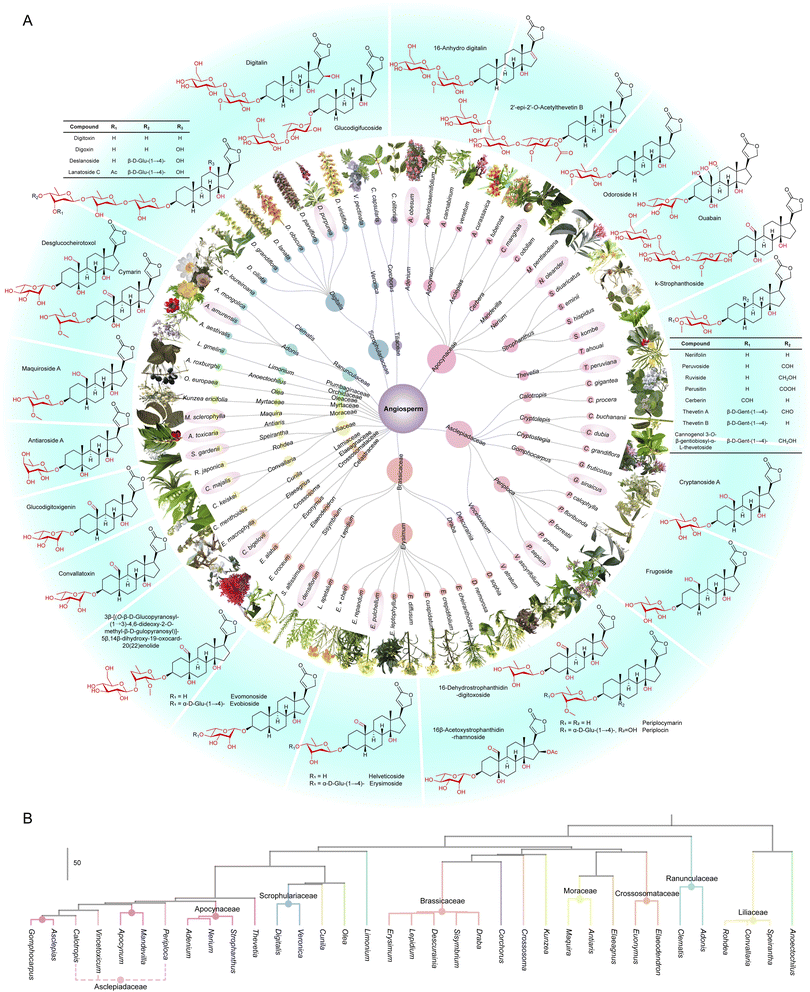
Cardiac glycosides: structural diversity, chemical ecology, bioactivity, and artificial synthesis
Dian
Jiao
 a,
Yibo
Zhang
a,
Yibo
Zhang
 a,
Wending
Guo
a,
Wending
Guo
 a,
Shuang
Liu
a,
Shuang
Liu
 *b,
Ping
Su
*b,
Ping
Su
 *a and
Luqi
Huang
*a and
Luqi
Huang
 *a
*a
aState Key Laboratory for Quality Ensurance and Sustainable Use of Dao-di Herbs, National Resource Center for Chinese Materia Medica, China Academy of Chinese Medical Sciences, Beijing 100700, China. E-mail: suping120@nrc.ac.cn; huangluqi01@126.com
bExperimental Research Center, China Academy of Chinese Medical Sciences, Beijing 100700, China. E-mail: scarlett0706@163.com
First published on 21st October 2025
Abstract
Covering: up to 2025
Cardiac glycosides (CGs), a class of metabolites found in nature, comprise sugar residues, unsaturated lactone rings, and steroidal cores. As renowned phytotoxins, they play vital roles in maintaining ecological balance. CGs have been widely used in the treatment of cardiovascular diseases such as heart failure and tachyarrhythmia for more than 200 years. Recent studies have revealed that CGs have numerous applications in various disease therapeutic areas, including anticancer, immunomodulatory, anti-inflammatory, antiviral, and neuroprotective effects. However, the medicinal resources of CGs are mainly reliant on natural plant and animal extracts, which not only limits their sustainable supply but also increases development costs. With the growing understanding of the pharmacological value of CGs and their increasing demand in the pharmaceutical industry, the sustainable supply of medicinal resources will become a bottleneck limiting their further development. Therefore, the artificial synthesis of target active ingredients, including chemical (semi)synthesis and biosynthesis, is becoming a hot topic among scholars worldwide. This paper presents the first systematic review of recent research advances in the structure, distribution, chemical ecology, biological activities, and artificial synthesis of CGs. Finally, we discuss the current challenges and urgent issues in this field, aiming to promote the widespread application of CGs in medicine through comprehensive pharmacological studies and exploration of synthesis techniques.
1. Introduction
Cardiac glycosides (CGs) have been used in the clinical treatment of cardiovascular diseases such as heart failure and atrial fibrillation for more than 200 years. The use of foxglove extract for the treatment of patients with congestive heart failure was first described in 1785 by the English physician William Withering.1 Over the following decade, its use in 163 patients with swelling led to the publication of the medical classic An Account of the Foxglove and Some of Its Medical Uses. Since then, foxglove has become one of the most important herbs in modern medicine. In recent times, this botanical has been replaced by its purified active ingredients—CGs, digitoxin,2 and digoxin.3 With the development of chemical analytical techniques (including chromatography, mass spectrometry, and nuclear magnetic resonance), more CGs have been isolated, characterized, and synthesized with greater precision.4,5 Some CGs are Food and Drug Administration (FDA)-approved Na+/K+-ATPase (NKA) inhibitors. NKA is a ubiquitous membrane protein found throughout the living world and abundantly present in cell membranes. It maintains the ionic concentration gradient across the cell membrane by actively transporting Na+ out of the cell and replacing it with K+. Since its discovery sixty years ago, NKA has been extensively studied, and its key physiological roles in regulating cardiac function and maintaining neural signaling have been established.6,7 To date, hundreds of CGs have been identified from a wide range of angiosperms. Plants produce CGs that target the NKA of natural enemies, thereby serving as a self-defense mechanism.8 Notably, more than 100 insect species are known to have evolved mechanisms to tolerate CG toxicity.9,10 This plant–insect interaction is an example of co-evolution, whereby plants produce CGs as a defense mechanism, while certain insects have adapted to use these plants as a food source. In addition, some animals consume CG-containing plants not to overcome plant defenses but to acquire toxins that protect them from predators.11–13Herbs containing CGs have been used in Traditional Chinese medicine (TCM) for thousands of years. For example, Periploca forrestii Schltr. is a commonly used folk medicine in TCM for the treatment of rheumatoid arthritis and wounds, suggesting that its main ingredient, CG, has a promoting effect on inflammation and wound healing. In recent times, numerous studies have found that, in addition to their well-known cardiac-strengthening effects, CGs possess considerable therapeutic potential as anticancer, anti-inflammatory, immunomodulatory, antiviral, and neuroprotective agents—providing a rich source of therapeutic leads for the pharmaceutical industry and ongoing inspiration for medicinal design. However, plants of the genus Digitalis are still the only viable source of CGs.14 The amount of CGs can vary among different organs of the same plant, and the yield is greatly affected by the harvesting time and environmental conditions.15 Some CGs are obtained through the biotransformation of precursor compounds extracted from toads. This method of extraction seriously disrupts the ecological balance, and the use of pesticides has also led to a decline in amphibian populations worldwide.16,17 Therefore, relying solely on existing natural toad resources will inevitably lead to the decline or even extinction of wild species of toads. Breeding toads on an artificial scale not only requires specific growth cycles but also faces several challenges, including difficulties in product purification, species degradation, and heavy metal contamination.18–20 Low natural availability of CGs in plants and animals, complicated extraction procedures, and environmental pollution by extraction solvents limit the broader application of CGs, and alternative production methods are needed. Currently, artificial synthesis using techniques such as biosynthesis and chemical synthesis is regarded as a promising alternative production method, attracting long-term interest and remaining one of the most exciting and dynamic topics. In particular, advancements in biosynthesis technology in the past two decades have provided theoretical and technological support for shifting the production of active ingredients from the traditional resource-based methods to a new industrial production mode that is highly efficient, green, stable, and controllable. Synthetic biology research on tanshinone,21 paclitaxel,22 artemisinin,23 vinblastine,24 and others has made great progress. The total synthesis of some CGs has been achieved.25,26 In the case of CG biosynthesis, many key genes have also been identified, offering promising prospects for the heterologous synthesis of these natural compounds.
Research on the pharmacological effects, chemical composition, synthetic biology, and chemical synthesis of CGs has progressed rapidly in recent years, and these new findings offer fresh directions for their clinical application and drug development. Although there have been many reviews on CGs in the past, most of them have focused only on isolated aspect of pharmacological activity (e.g., anticancer effects). More than two hundred pharmacological studies on CGs are published annually. While previous reviews focused on pre-2022 findings, our analysis will integrate recent studies to emphasize the latest breakthroughs. Moreover, comprehensive reviews on the artificial synthesis of CGs remain scarce. Herein, we present the first systematic review of the recent research advancements on the structure, distribution, chemical ecology, biological activity, and artificial synthesis of CGs. We expect that this review will facilitate the translation of CGs from basic research to clinical applications and further support the exploration of their potential in treating a wide range of diseases.
2. Natural sources, ecological functions, and isolation
2.1. Structure and distribution
CGs comprise a steroid nucleus and sugars, with a basic skeleton containing 17 carbon atoms (Fig. 1). At the C-17 position, CGs have a five- or six-membered unsaturated lactone ring, a structural feature that confers unique pharmacological activity, based on which they can be categorized as cardenolides and bufadienolides. Steroid nuclei found in nature have four rings: A, B, C, and D. The A/B ring typically exhibits a cis-fused junction (e.g., digitoxigenin), and some CGs, like uzarigenin, also contain a trans-fused junction; the B/C ring displays a trans-fused junction, while the C/D ring exhibits a cis-fused junction. CGs have a β-hydroxyl group at the C-14 position, which is an important feature that distinguishes CGs from other steroidal compounds. Sugars are typically attached to the C-3 position of the steroid parent nucleus, predominantly in the β-configuration, and can be present either as monosaccharides or oligosaccharides. In terms of sugar composition, cardenolides exhibit considerable diversity in their sugar moieties, encompassing a wide range of monosaccharides and unique deoxy sugars, whereas bufadienolides are typically conjugated with more common sugars such as glucose or rhamnose. Currently, more than 20 types of sugar moieties in CGs have been reported. Based on the presence or absence of a hydroxyl group at the C-2 position, these sugars can be classified into α-hydroxy sugars and α-deoxy sugars.2.2. Ecological functions
The ecological roles of CGs primarily manifest in their function as plant-derived secondary metabolites, particularly in plant defense and interspecific interactions. These roles significantly shape the relationships between plants and other organisms (such as herbivores, insects, and microorganisms), as well as their dynamics within the broader ecosystem.2.3. Extraction and isolation from natural sources
The extraction and isolation of CGs remain challenging in natural product research, primarily due to their occurrence as complex mixtures of structurally analogous compounds with similar polarities, coupled with their susceptibility to degradation under high temperature, acidic or alkaline conditions, and enzymatic hydrolysis.77 The solubility of CGs varies significantly with solvent choice; they are generally water-soluble but insoluble in non-polar solvents, except for chloroform and ethyl acetate. The degree of glycosylation and the number of hydroxyl groups both enhance hydrophilicity and influence pharmacokinetic properties. However, solubility cannot be accurately predicted based solely on molecular structure. For example, digitoxin—a compound containing five hydroxyl groups and three sugar units—is poorly soluble in water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 100
100![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000) yet highly soluble in chloroform (1
000) yet highly soluble in chloroform (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 40). In contrast, ouabain, which possesses eight hydroxyl groups and one sugar unit, is readily soluble in water (1
40). In contrast, ouabain, which possesses eight hydroxyl groups and one sugar unit, is readily soluble in water (1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 75) but exhibits low solubility in chloroform.
75) but exhibits low solubility in chloroform.
Conventional extraction employs polar organic solvents such as ethanol,30 methanol, or aqueous methanol/ethanol mixtures.69 Preliminary enrichment is often achieved through liquid–liquid partitioning using petroleum ether–ethyl acetate–water or n-butanol systems.27,78 Alternatively, dichloromethane extraction followed by washing with 1% NaCl solution can remove phenolic contaminants.33 Bufadienolides are typically extracted with methanol.40 Further purification is commonly performed using silica gel column chromatography (CC), preparative thin-layer chromatography (TLC), or high-performance liquid chromatography (HPLC). Normal-phase CC utilizing binary gradients of medium- to low-polarity solvents (e.g., chloroform, ethyl acetate, dichloromethane) mixed with methanol as a polar modifier effectively separates CGs of varying polarities.33 HPLC, particularly reversed-phase C18 columns with acetonitrile–water78 or methanol–water gradients,68 is preferred over gas chromatography (GC) due to the non-volatility of CGs and no need for derivatization. Method selection depends heavily on the specific matrix and target compounds, often requiring optimization of solvent systems and gradient conditions to achieve high recovery and resolution. TLC remains useful for microgram-scale screening, with reagents like SbCl3 employed for bufadienolides detection.40 Although limited by low volatility, GC-MS has been applied in some cases, while high-performance capillary electrophoresis (HPCE) offers an alternative with high resolution and efficiency.77
Structural elucidation of CGs is complicated by their chromatographic similarities and sensitivity, often requiring a combination of chromatographic and spectroscopic techniques to determine glycosylation patterns, stereochemistry, and substituent locations.77,79 Acid hydrolysis followed by sugar derivatization further aids structural characterization.27,34 Up to now, more than five hundred CG structures have been successfully elucidated through these integrated approaches.31 These elucidated structures provide a critical foundation for understanding structure–activity relationships, guiding the discovery of novel analogues with improved pharmacological profiles. Future efforts may focus on optimizing extraction and separation workflows to better preserve labile structures and enable the identification of even lower-abundance CGs. Moreover, integration with computational and bioactivity-guided approaches could accelerate the targeted discovery of compounds with desired therapeutic properties.
3. Biological activities
CGs exhibit a wide range of medicinal properties (Fig. 5 and Table 1). Digoxin is FDA-approved for intravenous use in acute heart failure or acute exacerbations of chronic heart failure. In ophthalmic diseases, compounded esculin and digitalis glycoside eye drops are used to treat visual fatigue and macular degeneration, and some experiments are underway to explore their use in the treatment of dry eye disease.80 Clinical trials are underway globally to explore the therapeutic potential of CGs in anticancer, anti-inflammatory, antiviral activities, among others. | ||
| Fig. 5 Bioactivities, clinical and potential applications of CGs. The superscript * indicates findings based on clinical evidence, while others are based on in vivo and in vitro results. | ||
| CGs | Structure | Indications/bioactivity | Market information |
|---|---|---|---|
| a This table lists CGs, including both current and historical agents, with selected compounds under investigation for novel indications (Phase II and beyond). Abbreviations: HF, heart failure; NASH, nonalcoholic steatohepatitis. N/A, not applicable. US: United States; CN: China; EG: Egypt; GB: United Kingdom; NL: Netherlands. | |||
| Digoxin |
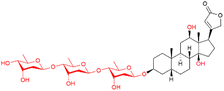
|
Atrial fibrillation, HF, arrhythmias, tachycardia | In current use (Lanoxin tablet, digoxin injection) |
| Anti-NASH activity | Phase II clinical trial (US: NCT06588699; CN: NCT04216693) | ||
| Anti-rheumatoid arthritis activity | Phase II trial completion (EG: NCT04834557) | ||
| Phase I trial completion (US: NCT03131973; GB: NCT01355354) | |||
| Anti-cutaneous warts activity | Phase II trial completion (NL: NCT02333643) | ||
| Digitoxin |
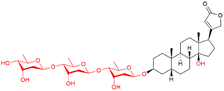
|
Arrhythmias, HF | Discontinued (FDA); available elsewhere (CRYSTODIGIN injection) |
| Anti-tumor activity in pancreatic cancer | Phase II clinical trial (CN: CTIS2024-512128-12-01) | ||
| Anti-cystic fibrosis activity | Phase II clinical trial (US: NCT00782288) | ||
| Deslanoside |
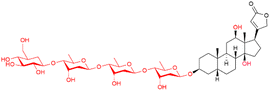
|
Arrhythmias, HF, tachycardia | Discontinued (Cedilanid-D injection) |
| Ouabain (Strophanthin G) |
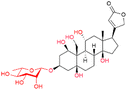
|
HF | Discontinued (FDA); available elsewhere (Uabasin injection) |
| Strophanthin K |
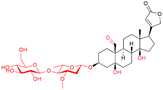
|
HF | Not approved (FDA); available elsewhere (Strophanthin K injection) |
| Convallatoxin |
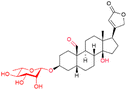
|
HF | Not approved (FDA); available elsewhere (Convallatoxin injection) |
| Lanatoside C |
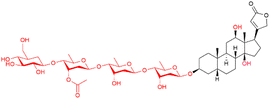
|
Arrhythmias, HF | Not approved (FDA); discontinued elsewhere (Lanatoside C injection) |
| Proscillaridin A |
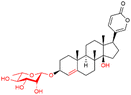
|
HF | Not approved (FDA); discontinued elsewhere (Prost tablet) |
| Digitalis glycosides (Digital purpurea total CGs) | N/A | Visual fatigue and macular degeneration | Not approved (FDA); available elsewhere (Esculin and Digitalis glycosides eye drops) |
| Rodealin (Rohdea japonica total CGs) | N/A | Atrial fibrillation, HF, tachycardia | Not approved (FDA); discontinued elsewhere (Rodealin injection) |
| Thevetosid (Thevetia peruviana total CGs) | N/A | Atrial fibrillation, HF | Not approved (FDA); discontinued elsewhere (Thevetosidum) |
In addition, CGs were found to promote wound healing. Digitoxigenin considerably improves skin condition by increasing the hydroxyproline content in the wound tissues.81 Periplocin notably promotes cell proliferation and migration and stimulates collagen production in fibroblast L929 cells through activation of the NKA-mediated Src/ERK and PI3K/AKT pathways.82 CG from Streptocaulon juventas (Lour.) Merr. induces collagen synthesis in predominantly human skin fibroblasts by protecting LLCPK1 cells from hypoxic injury.32
CGs can markedly improve the level of homeostasis in the body. Digoxin can improve cerebral metabolic disorders caused by chronic cerebral hypoperfusion and attenuate cognitive deficits.83 Ouabain reduces infrared-induced skeletal muscle dysfunction by preventing the reduction of α2 NKA function in rats.84 Nanomolar concentrations of ouabain induce selective cytotoxicity in tissue-resident macrophages and enhance white adipose tissue homeostasis, highlighting its potential application in the treatment of metabolic syndrome, a condition characterized by pathogenic macrophage infiltration and activation.85 In addition, ouabain is selectively cytotoxic to senescent cells, possibly inducing apoptosis by blocking ion pumps and depleting intracellular K+.86
Recently, CGs have also been found to have therapeutic potential in reducing intraocular pressure87 and improving reproduction.88,89 Further pharmacological activities and mechanisms of action of CGs are being explored.
3.1. Structure–activity relationship
NKA represents the most well-established target of CGs and also functions as a signal transducer that regulates cell metabolism, survival, and death.90 As the largest protein complex within the P-type cation pump family, it hydrolyzes ATP to drive the transport of K+ into and Na+ out of the cell at a 2![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 3 stoichiometric ratio, thereby establishing and maintaining the transmembrane electrochemical gradient.
3 stoichiometric ratio, thereby establishing and maintaining the transmembrane electrochemical gradient.
The aglycone moiety of CG is considered the essential pharmacophore responsible for the pharmacological efficacy of CGs.91 X-ray crystallography has revealed that CGs embed into the CG-binding pocket of the NKA α-subunit. The conserved 14β-hydroxyl group mediates a common interaction mode with NKA across multiple CGs, primarily through hydrogen bonding with the carbonyl oxygen atom of Thr797.92,93 Additional substituents interact with the protein via specific polar contacts.93 The β-oriented surface of CGs facilitates extensive hydrogen-bonding networks with residues in the αM1, αM2, and αM6 helices of NKA.30,33,94 A/B cis-junction is important for high activity.95 Due to the hydrophobic nature of the B ring, the introduction of oxygen-containing groups at C-7 or C-8 can significantly enhance cytotoxic activity. Most substituents at C-10, C-13, and C-17 positions of naturally occurring CGs are in the β-configuration. The C-10 (e.g., 10-epi-uzarigenin) or C-13 (e.g., 13-epidigitoxigenin) α-configurations are typically achieved through chemical modification or semisynthesis, whereas the C-17 α-configuration (e.g., (+)-17β-hydroxystrebloside) is associated with a loss of cardiotonic activity. This is attributed to its ability to adopt at least three distinct conformations upon binding to the cation-binding site, leading to disrupted interaction with NKA.92 Substituents such as the C-10 angular methyl, aldehyde, or hydroxymethyl groups, the C-12 β-hydroxyl group, and the C-17β side chain are critical for mediating cardiotonic activity, whereas hydroxylation at C-16 is detrimental.27 C1, C5, C11 and 19 hydroxyls may enhance the activity, but their presence is not essential.95
A defining feature of CGs is the unsaturated lactone ring at the C-17β position, which distinguishes them from steroid hormones. This lactone contains two oxygen atoms that generate a highly negative electrostatic potential, enhancing both orientation and strength of hydrogen bonding. The interaction energy contributed by the lactone ring is comparable to that of the steroid core (−22.7 kJ mol−1). Notably, α-pyrone-type lactones (as in bufadienolides, −27.6 kJ mol−1) exhibit stronger interaction than butenolide-type rings (as in cardenolides, −20.5 kJ mol−1), correlating with their higher cardiotonic potency.96
The types and attachment sites of sugars considerably influence the solubility and biological activity of CGs. Aglycones are generally absorbed more rapidly and metabolized more easily than their glycosylated counterparts.91 Notably, glycosylation at the C-3 position often enhances cytotoxic activity several-fold compared to the aglycone form, indicating that the hydrophilic group at this position plays a critical role in potency.27 Monoglycosides typically exhibit the highest cytotoxicity, while increased sugar chain length and molecular size tend to reduce toxic effects.97
3.2. Therapeutic effect of CGs on cardiovascular diseases
CGs were initially employed in the treatment of cardiovascular diseases, particularly heart failure and arrhythmias. Their therapeutic effects are primarily mediated through specific binding to and inhibition of NKA, which subsequently alters Na+–Ca2+ exchanger (NCX) activity and promotes Ca2+ release from the sarcoplasmic reticulum. The resulting increase in intracellular Ca2+ concentration enhances myocardial contractility and improves cardiac output, thereby exerting positive inotropic effects and alleviating symptoms of heart failure. At nanomolar concentrations, CGs can activate protective signaling pathways (e.g., calcium oscillations, SRC kinase, MAPK) without significantly inhibiting ion pumping, suggesting potential for safer cardiovascular drugs.91Pharmacological studies have shown that periplocin attenuates cardiac remodeling, improves cardiomyocyte contractility, and enhances cardiac diastolic function in heart failure rats, making it a potential treatment for heart failure with preserved ejection fraction.98 C-reactive protein (CRP) is associated with inflammation in cardiovascular disease, and digoxin inhibits CRP synthesis. This is the first report on the inhibition of CRP by CGs.99 Through live cell screening, Magadum et al. found that CGs can increase myocardial contractility and induce cardiomyocyte proliferation, thereby promoting cardiac repair and enhancing cardiomyocyte cycle activity.100 Clinical studies have found that in an elderly population whose digoxin therapy was discontinued prior to hospital admission, prognosis was considerably affected even after receiving other medications, suggesting that digoxin continues to play an irreplaceable role in the treatment and prognosis of chronic heart failure.101 Digoxin may partially diminish the expected decrease in RV systolic function and increase in RV systolic size through its positive inotropic effect, suggesting that it may improve cardiac function and potentially reduce mortality.102
3.3. Anticancer effects of CGs
Several studies have shown that CGs can be used as potential antitumor drugs and exhibit potent cytotoxicity against a wide range of cancer cells.33,40,103 CGs exert anticancer activity across various body systems, mainly by targeting key signaling axes, affecting the transcription and expression of cancer cell metabolic pathway proteins, inhibiting the malignant behaviors of cancer cells (proliferation, migration, differentiation, and invasion),104 inducing apoptosis and autophagy,105 and ameliorating the clinical symptoms of cancer,106 among other mechanisms.Breast cancer (BC) is the most common malignant tumor in women and the leading cause of female cancer incidence worldwide.107 Lanatoside C, peruvoside, and strophanthidin ameliorate the dysregulation of EGR1 and downstream proteins of the MAPK/ERK signaling pathway and reduce the proliferation and invasion of MCF-7 BC cells.108 Ouabain induces estrogen receptor α (ERα) degradation, thereby killing ERα-positive BC cells.109 Oleandrin induces immunogenic cell death by stimulating the ER.110 In triple-negative BC cells, 3′-epi-12β-hydroxyfroside (hyfs) induces complete autophagic flux, and combination treatment with an autophagy inhibitor may enhance activity.111 Preclinical 3D model screening has demonstrated that digoxin has therapeutic potential for treating dedifferentiated endometrial carcinoma.112 Proscillaridin A and lanatoside C target UCP2 to increase ROS, inhibit uterine leiomyosarcoma cell growth, and induce cell death.113 Neriifolin exerts anticancer activity against prostate cancer cells through endoplasmic reticulum stress (ERS)–mediated DNA damage, G2/M blockade, and apoptosis induction.114,115
CGs and their derivatives exhibit substantial anticancer activities in different cancer models and play a role in inhibiting tumor growth, overcoming drug resistance, and altering tumor microenvironmental aspects. Intrahepatic cholangiocarcinoma is a type of hepatocellular carcinoma with poor prognosis and high mortality. Digitoxin can inhibit the proliferation and migration of intrahepatic cholangiocarcinoma cells by targeting the NF-κB/ST6GAL1 signaling axis.116 CGs from Thevetia peruviana (Pers.) K. Schum. considerably induced apoptosis in human hepatocellular carcinoma HepG2 cells in a dose-dependent manner.117 Binding of ouabain, oleandrin, and digoxin to NKA downregulates the THADA–LAT1 pathway to inhibit the proliferation of HepG2 cells and human epidermoid carcinoma KB cells.118 CGs have agonistic and potentiating effects on RORγ/RORγT nuclear receptors, modulating Th17 or Tc17 lymphocyte differentiation.119 Periplocymarin synchronously activates apoptosis and initiates the AMPK/ULK1 and mTOR signaling pathways, leading to protective autophagy.120 Lanatoside C downregulates the expression of signal transducer and activator of transcription 3 (STAT3), increases ROS levels, and decreases MMP, thereby slowing proliferation and inducing apoptosis in HuCCT-1 and TFK-1 cholangiocarcinoma cells.121 Ouabain at a nanomolar concentration interferes with spheroid growth and is cytotoxic to bile duct cancer cells.122 Digitoxin can promote apoptosis in colon cancer cells with drug-resistant KRAS mutations by decreasing HIF-1α and STAT3 levels, thereby inhibiting cellular proliferation and migration.123 Proscillaridin A enhances the sensitivity of drug-resistant cells.124 Moreover, digitoxin activates the ROS-induced RhoA/ROCK pathway, inhibits cancer cell proliferation and migration, and reverses the pro-angiogenic effects mediated by the tumor microenvironment.125 Different phenotypes affect the rate of ROS production by digitoxin, which differentially induces apoptosis in four pancreatic cancer cell lines.126 Oleandrin is a novel and potent inhibitor of ATM and ATR kinases that mediates DNA damage response to enhance the sensitivity of lung cancer to radiotherapy.127 In human multiple myeloma AMO1 cells, periplocin considerably inhibited the oncogene c-MYC via α1-NKA.128,129 Convallatoxin downregulates the PTHR1 and Wnt/β-catenin pathways, inhibiting the malignant development of osteosarcoma cells.130 Ouabain induces apoptosis in leukemic stem cells by promoting the loss of Mcl-1 and c-Myc expression.131 17β-neriifolin induces apoptosis by inhibiting the overexpression of HOXA9.132 A synthetic digitoxin derivative downregulates the phosphorylation of MEK1/2 and induces a cellular G2/M cell cycle arrest in leukemia cells, resulting in cell death.133 Indoleamine-pyrrole 2′,3′-dioxygenase 1 (IDO1) plays an important role in cancer cell metabolism. Ouabain and digoxin downregulate IDO1 mRNA and protein levels at concentrations that do not affect cell viability.134 CGs can downregulate the expression of the glucose transporter GLUT1 and affect glucose metabolism in various cancer cells by inhibiting NKA α3-isoform activity.135
3.4. Immunomodulatory and anti-inflammatory effects of CGs
Neuroinflammation is present in virtually every neurological disorder and contributes to disease pathogenesis by promoting neuronal loss and blood–brain barrier dysfunction.136,137 Ouabain promotes antioxidant effects in retinal cells and has been demonstrated for the first time to protect retinal ganglion cells through the activation of autophagic pathways.138 Jansson et al. found that digoxin and lanatoside C attenuated meningeal and choroid plexus inflammatory responses and blocked inflammatory propagation, which was also confirmed in a 3D in vitro model.139 Moreover, digoxin can reverse vascular dilation and pulp expansion, suggesting that it can attenuate pulp inflammation.140Respiratory diseases are common and prevalent conditions that constitute a major public health issue.141 COVID-19 is a pandemic of acute respiratory disease caused by the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).142 Digitoxin and ouabain counteract the CFTR protein loss induced by SARS-CoV-2 infection, thereby reducing inflammation in the airway caused by COVID-19.143 In addition to its role in acute models of inflammation, ouabain also acts on chronic inflammatory phenotypes. It inhibits the activation of the p38 MAPK signaling pathway, thus modulating allergic asthma.144
Digoxin exhibits a protective effect against intervertebral disk degeneration by decreasing the transcriptional levels of proinflammatory cytokines (COX-2, iNOS, IL-1β, and IL-6). It inhibits destructive metalloproteinases and rescues extracellular matrix (ECM) loss, activating the AKT and ERK1/2 signaling pathways to promote ECM anabolism.145 Osteoarthritis (OA), a common chronic joint disease with no known drugs to alter its progression, has become a leading cause of disability worldwide.146 Jia et al. found that digoxin improves the OA inflammatory microenvironment and promotes chondrogenesis by downregulating the M1-like macrophage-derived exosome miR-146b-5p/Usp3&Sox5 axis.147 Wang et al. demonstrated that ouabain and digoxin limited OA development in vitro. Low-density lipoprotein receptor-related protein 4 (LRP4), a novel target of digoxin, was targeted for regulating chondrocyte metabolism.148
FDA-approved CGs have demonstrated potent anti-inflammatory activity in the development of a wide range of other inflammatory conditions. Ouabain downregulates the expression of the amino acid transporter SLC7A11 to inhibit glutathione synthesis. Excessive oxidative stress notably induces G2/M cycle arrest in human HaCaT keratinocytes, demonstrating anti-psoriatic activity.149 Ouabain blocks electrophysiological disturbances and claudin-2 overexpression, offering potential to ameliorate ionizing radiation-induced intestinal dysfunction.150 Minami et al. designed a specific perilipin 1-derived LD-targeting domain that binds to the optimized LIR-interacting domain, which recognizes digoxin as an activator of lipophagy to inhibit the transition to NASH in vivo.151 Multiple models of inflammation have been used to study CGs. Digoxin exhibits excellent anti-inflammatory activity by inhibiting soluble epoxide hydrolase activity and exerts antipyretic effects in rectal temperature measurements.152 Convallatoxin153 and the synthesized CG BD-8 (ref. 154) also showed anti-inflammatory effects both in vivo and in vitro. The immunomodulatory and anti-inflammatory effects of CGs are shown in Table 2.
| CGs | Anti-inflammatory effects | Experimental subject | Administration mode and dosage |
|---|---|---|---|
| a Abbreviations: i.p., intraperitoneal; p.o., oral; i.a., intra-articular. N/A, not applicable (the administration mode and dosage were not addressed). | |||
| Ouabain138 | Reduce TNFR1/2, TLR4 and CD14 expression; increase TNFR2 levels; decrease the density of Iba-1 and the production of ROS | RGC | 3 nM |
| Ouabain149 | Inhibit GSH synthesis, inducing oxidative stress in psoriatic keratinocytes | HaCaT | 0–200 nM |
| Ouabain144 | Reduce migration of eosinophils, lymphocytes and macrophages; decrease IL-33, TSLP, IL-4, IL-1β, TNF-α and TGF-β; attenuate collagen deposition and mucus production | Female BALB/c mice | 0.56 mg per kg per day, i.p. |
| Ouabain150 | Prevent occludin overexpression and pore-forming claudin-2 expression | Male Wistar rats | 1 μg per kg per day, i.p. |
| Digoxin140 | Reverse ZOL-induced vasodilation/dilatation; reduce IL-17, TNF-α and TGF-β and increase of IL-6 expression | Male Wistar rats | 1 μg per kg per day, i.p. |
| Digoxin152 | Elevate EETs levels, inhibiting inflammatory mediators; reduce lung edema, neutrophilic infiltration and alveolar septal thickening | Swiss albino mice, Wistar albino rats | 0.1–0.2 mg kg−1, p.o. |
| Digoxin145 | Suppress TNF-α-induced inflammation; attenuate ECM destruction; promote ECM anabolism | Male rats | 50 nM, i.p. |
| Human nucleus pulposus cells | 10–100 nM | ||
| Digoxin147 | Reduce iNOS and IL-6 secretion; improve joint inflammatory microenvironment; promote chondrocyte production | RAW 264.7, OA patient synovial macrophages | 20–80 μM |
| Male C57BL/6 mice | 0.02, 0.2 mg per kg per day, i.a. | ||
| Digoxin151 | Activate lipophagy; attenuate inflammatory and profibrotic gene expression | C57BL/6J mice | 0.8 mg per kg per day, p.o. |
| Human retrospective cohort study | N/A | ||
| Digitoxin, Ouabain143 | Prevent CFTR loss, reducing TNFα/NFκB- and ENaC-dependent inflammation in COVID-19 airways | Vero E6 | 50 nM |
| Digoxin, Ouabain148 | Stimulate chondrocyte differentiation and metabolism; relieve OA-associated pain; target LRP4 to promote chondrocyte anabolism | C3H10T1/2, C28I2, mouse primary chondrocytes | 50–100 nM |
| Male C57BL/6 mice | 50–100 nM, i.a. | ||
| Human retrospective cohort study | N/A | ||
| Digoxin, Lanatoside C139 | Reduce IL-1β-induced gCCL2, sICAM-1, sVCAM-1 and CX3CL1; block endothelial inflammatory secretion | Primary human brain mixed glial, pericyte, endothelial | 0.1, 1 μM |
| BD-8 (ref. 154) | Inhibit NO production; decrease IL-1β; reduce phosphorylation of inflammation-associated signaling pathways | Primary mouse macrophages | 0.01–100 μM |
| Female Swiss albino mice | 0.28, 0.56, 1.12 mg kg−1, i.p. | ||
| Convallatoxin154 | Inhibit ZFP91-mediated pro-IL-1β ubiquitination and caspase-8 inflammasome activity, reducing IL-1β production | THP-1, BMM | 3–100 nM |
| C57BL/6 male mice | 150 μg per kg per day, p.o. | ||
3.5. Antiviral effects of CGs
The binding of CGs to NKA is involved in antiviral activity against a wide range of viruses, either by activating signaling cascades or altering the concentration of intracellular ions.155 CGs are involved in almost every step of the viral life cycle, including RNA synthesis, protein translation, and post-transcriptional processing, except viral attachment to the host cell.156 CGs have been described as promising compounds in addition to ongoing clinical viral vaccines with strong immune responses.As FDA-approved inhibitors targeting NKA, ouabain and digoxin inhibit Japanese encephalitis virus (JEV) infection during the viral replication phase.157 Amarelle et al. found that ouabain inhibits influenza A virus (IAV) replication in alveolar epithelial cells by decreasing intracellular potassium levels.158 Zika virus (ZIKV) is a novel mosquito-borne flavivirus associated with neurological disorders.159 Ouabain and digoxin have been shown to reduce the viral burden of ZIKV in adult mice and prevent ZIKV infection-induced microcephaly, which is fatal in pregnant mice.160 Ouabain reduces ZIKV replication by decreasing RNA copy number, while NS5-RdRp and NS3-helicase proteins are suggested to be the targets of its direct action on ZIKV proteins.161 Moreover, Du et al. demonstrated that ouabain, digoxin, lanatoside C, and digitoxin have anti-Ebola virus (EBOV) replicative and transcriptional activities, which may specifically inhibit the entry of EBOV into host HeLa cells by mediating intracellular calcium disorder.162 Compared to drugs such as cyclosporin A, sunitinib, and chloroquine, digoxin has the lowest inhibitory concentration (IC50) to block Bunyamwera virus (BUNV) infection by inhibiting NKA-induced high electron-density and swollen cristae in mitochondria.163 Ouabain and the digitoxigenin derivative PST2238 (rostafuroxin) specifically bind to the α1 subunit of the NKA (ATP1A1) and blocks EGFR Tyr845 phosphorylation, thereby inhibiting infection caused by efficient uptake of human respiratory syncytial virus (RSV).164 Prion diseases are rapidly progressing and ultimately fatal neurodegenerative disorders in mammals.165 CGs are targeted NKA inhibitors that reduce cellular prion protein (PrPC) levels.166 C4′-Dehydro-oleandrin reaches higher concentrations in the brain and exhibits lower toxicity than other CGs.167 In 2024, Wu et al. found that lanatoside C inhibited Herpes simplex virus type 1 (HSV-1) replication both in vitro and in vivo by inducing the localization of intracellular nuclear factor erythroid 2-related factor 2 (NRF2) at the periphery of the cell nucleus.168 In addition, ten unreported CGs from S. asper exhibited potent antiviral effects against HSV-1 in vitro.78 Lytic reactivation of Epstein–Barr virus (EBV) plays an important role in virus-driven malignancies. Cai et al. reported for the first time that two CGs from Sonchus asper (L.) Hill could effectively inhibit early antigen expression in vitro to limit EBV lytic replication.37 Notably, Yang et al. revealed that the notable antiviral effects of CGs on porcine transmissible gastroenteritis coronavirus (TGEV) and human HCoV-OC43 were dependent on an NKA-independent signaling axis. Ouabain inhibits the viral activities of coronaviruses by activating Ndfip1/2 and NEDD4, leading to Janus kinase 1 (JAK1) proteolysis and downregulation.169 Cutaneous warts are caused by the human papillomavirus (HPV).170 Topical ionic contra-viral therapy with digoxin considerably reduced the diameter of cutaneous warts and HPV load compared with placebo.171 In addition, CGs inhibited the expression of both early and late vaccinia virus (VACV) proteins at different concentrations.172
The SARS-CoV-2 epidemic has led to growing interest in the potential therapeutic applications of CGs for COVID-19. Souza et al. found that CGs reduced viral replication and inhibited NF-κB from directly interfering with SARS-CoV-2 yield and inflammatory cytokine production.173 Digoxin and ouabain treatments considerably inhibited SARS-CoV-2 replication by more than 99% in vitro, demonstrating greater efficacy than approved drugs such as chloroquine and remdesivir.174 Pollard et al. found that digitoxin notably reduced the levels of cytokines TNFα, GRO/KC, MIP2, MCP1, and IFNγ in cotton rats infected with influenza viruses.175 As the initiation of SARS-CoV-2 infection requires the receptor binding domain (RBD) on the viral spike protein to bind to the host receptor ACE2 protein, CGs (ouabain, digitoxin, and digoxin) may prevent viral penetration and reduce infectivity by first inhibiting ACE2:RBD binding.176 Oleandrin also demonstrates potent prophylactic and therapeutic activity in inhibiting SARS-CoV-2. A defined extract of Nerium oleander L. applied to the relevant hamster model of COVID-19 showed a substantial reduction of nasal viral load in the nasal turbinates.177 In addition, oleandrin inhibits the expression of 78 kilodalton glucose-regulated protein (GRP78), which can reduce SARS-CoV-2, Beta, and Delta, as well as highly infectious Omicron variants. When combined with oleandrin at low nanomolar dosages, the treatment of COVID-19 is enhanced by remdesivir or nirmatrelvir.178 The antiviral effects of CGs are shown in Table 3.
| CGs | Virus | Mechanism | Experimental subject | Dosage |
|---|---|---|---|---|
| a Abbreviations: i.p.,intraperitoneal; p.o., oral; t.c., topical cutaneous; s.l., sublingual; s.c., subcutaneous; N/A, not applicable (the mechanism was not addressed). | ||||
| Ouabain | JEV157 | Attenuate virus replication via NKA blockade | Vero, Huh-7, U251 | 49–784 nM |
| BALB/c mice | 3 mg per kg per day i.p. | |||
| IAV158 | Attenuate virus replication via NKA blockade | Alveolar epithelial cells | 1–20 nM | |
| RSV164 | Inhibit viral uptake by targeting ATP1A1 | A549, HSAEC | 25 nM | |
| Ouabain | EBOV162 | Inhibit virus entry and replication | Hela, HEK293T, THP-1 | 50–500 nM |
| ZIKV (ref. 160 and 161) | Attenuate virus replication via NKA blockade | Vero | 12.25–784 nM | |
| Adult mice, pregnant mice | 2 mg per kg per day, i.p.; 3 mg per kg per day, p.o. | |||
| Attenuate virus replication via NS5-RdRp/NS3-helica | Vero | 2.5–20 nM | ||
| TGEV/HCoV-OC43 (ref. 169) | Attenuate virus replication through JAK1_STAT1/3 | ST, HCT-8 | 30–300 nM | |
| SARS-CoV-2 (ref. 174 and 176) | Attenuate virus replication | Vero | 100 nM | |
| Reduce the ACE2/RBD binding | A549 | 80–120 nM | ||
| Digoxin | JEV157 | Inhibit viral RNA synthesis via NKA blockade | Huh-7 | 49–784 nM |
| ZIKV160 | Attenuate virus replication via NKA blockade | Vero | 12.25–784 nM | |
| EBOV162 | Inhibit virus entry and replication | Hela, HEK293T, THP-1 | 50–500 nM | |
| BUNV163 | Attenuate virus replication via NKA blockade | Vero, HEK293T, BHK-21 | 0–500 nM | |
| HPV171 | Attenuate virus replication | Human study | 5–30 mg per day, t.c. | |
| SARS-CoV-2 (ref. 174 and 176) | Attenuate virus replication | Vero | 150 nM | |
| Reduce the ACE2/RBD binding | A549 | 80–120 nM | ||
| Lanatoside C | HSV-1 (ref. 168) | Attenuate virus replication via NRF2 | ARPE-19, Vero | 50–800 nM |
| EBOV162 | Inhibit virus entry and replication | Hela, HEK293T, THP-1 | 50–500 nM | |
| Digitoxin | EBOV162 | Inhibit virus entry and replication | Hela, HEK293T, THP-1 | 50–500 nM |
| SARS-CoV-2 (ref. 175 and 176) | Decrease THFα, GRO/KC, MIP2, MCPl, and IFNy | Cotton rat | 30–300 μg per kg per day, i.p. | |
| Reduce the ACE2/RBD binding | A549 | 80–120 nM | ||
| Oleandrin | SARS-CoV-2 (ref. 177 and 178) | Inhibit NKA | Vero | 0.005–1.0 μg mL−1 |
| Golden Syrian hamster | 0.0325, 0.325, 3.25 mg per day, s.l. | |||
| Induce viral infection through GRP78 blockade | Vero E6-ACE2, Caco-2-ACE2, A549-ACE2 | 4.38–100 nM | ||
| Rostafuroxin (PST2238) | RSV164 | Inhibit viral uptake by targeting ATP1A1 | A549, HSAEC | 20 nM |
| C4′-Dehydro-oleandrin (KDC203) | PrPC167 | Inhibit NKA | T98G, LN-229, 104C1, GPC-16, Guinea pig neurons/astrocytes/cardiomyocytes | 12–800 nM |
| Guinea pig | 0.15, 0.225, 0.3 mg per kg per day, s.c. | |||
| CGs from Streblus asper | HSV-1 (ref. 78) | N/A | Vero | 0.19–1.03 μM |
| CGs from Sonchus asper | EBV37 | Inhibit NKA | Raji, A549, THP-1, C666-1 | 0.35–5.8 nM |
| CGs | VACV172 | Attenuate virus replication | Hela | 5–100 nM |
3.6. Pharmacological activity of CGs in neuroprotection
As the α3 subunit of NKA serves as a neuronal receptor for agrin, its activation is implicated in various physiological activities in the nervous system.179,180 As FDA-approved NKA inhibitors, CGs trigger signaling pathways or regulate Ca2+ levels in a concentration-dependent manner to maintain neuronal homeostasis,181 and are therefore frequently used in studies of neurotransmitter-related diseases. Multiple sclerosis, a central nervous system autoimmune disease, causes loss of nerve myelin, and digoxin stimulates myelin regeneration.182 By combining both anti-inflammatory and neuroprotective activities, digoxin exerts a healing effect on peripheral nerve injuries.183 Neuroinflammation drives the development of epilepsy, and digoxin exhibits a clear anticonvulsant potential.184 Low doses of digoxin stimulate dendritic spine formation or recirculation and promote motor learning ability.185 Moreover, digoxin reduces hippocampal neuronal death, neuroinflammation, and cholinergic deficits, which can help in the treatment of cognitive impairment and Alzheimer's disease.186 At subnanomolar concentrations, ouabain counteracts calcium overload and neuronal apoptosis during excitotoxic stress.187 Dysregulation of microRNAs (miRNAs) is involved in the development of a wide range of diseases; however, no clinical approvals have been granted for the treatment of neurological disorders by modulating miRNAs. Nguyen et al. screened CGs as novel miR-132 inducers capable of protecting rodent and human neurons, suggesting a promising therapeutic approach for neurodegenerative diseases.188 In addition, CGs have been shown to restore WDR45 autophagy abnormalities and exhibit neuroprotective effects during hypoxia and glucose starvation, with therapeutic potential for beta-propeller protein-associated neurodegeneration.1893.7. Toxicology and safety profile
Despite their broad therapeutic potential, CGs face a major challenge in clinical translation due to their narrow therapeutic index—the fine margin between effective and toxic doses. Their mechanism, mediated through NKA inhibition, induces intracellular calcium overload and predisposes to arrhythmias. This cardiotoxicity severely limits systemic use and necessitates meticulous dosing and therapeutic drug monitoring.Digoxin is currently the only FDA-approved CG still in clinical use for humans, indicated for conditions such as atrial fibrillation, heart failure, arrhythmias, and tachycardia,190 as well as in veterinary medicine.191 Other CGs like digitoxin and deslanoside have been discontinued by the FDA due to high toxicity and have been replaced by alternative agents. Digoxin is commercially available in capsules, tablets, and ampules for oral and intravenous administration.192 Thus, using digoxin as a representative example, the therapeutic window in humans is well established at 0.5–2.0 ng mL−1 (equivalent to 0.6–2.6 nM).193 Published data on acute toxicity report toxic doses ranging from 50 to 200 μg kg−1 in humans, varying by sex, age, and route of administration. The frequency and severity of side effects—50% cardiac and 25% gastrointestinal—depend on dosage, patient condition, and concomitant medications.194,195 In recent human trials, therapeutic drug monitoring has detected digoxin levels below the quantitation limit (300 pg mL−1).171 Given that clinical trials for their emerging pharmacological effects (e.g., anticancer, antiviral) remain scarce and largely in early stages, the majority of reported in vivo data are still derived from preclinical animal models. In mice, acute toxic doses vary widely: 8.15 μg kg−1 (subcutaneous), 124–7670 μg kg−1 (other injections), and up to 17![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 780 μg kg−1 (oral). Rats generally exhibit higher toxicity thresholds than mice.194 Many in vivo studies demonstrate efficacy at non-toxic doses, though high or prolonged exposure may worsen health outcomes. These findings support the therapeutic potential of CGs at low doses, over short durations, or with altered administration routes—strategies that may reduce toxicity and side effects.158 Combination therapy is also under consideration.193 However, much of the evidence remains limited to cellular models, underscoring the need for further validation. Additionally, toxicological data for other CGs remain limited. Subtle structural variations can significantly alter their toxicological profiles, clinical manifestations, and histopathological outcomes,192 underscoring the need for more comprehensive studies to support clinical translation.
780 μg kg−1 (oral). Rats generally exhibit higher toxicity thresholds than mice.194 Many in vivo studies demonstrate efficacy at non-toxic doses, though high or prolonged exposure may worsen health outcomes. These findings support the therapeutic potential of CGs at low doses, over short durations, or with altered administration routes—strategies that may reduce toxicity and side effects.158 Combination therapy is also under consideration.193 However, much of the evidence remains limited to cellular models, underscoring the need for further validation. Additionally, toxicological data for other CGs remain limited. Subtle structural variations can significantly alter their toxicological profiles, clinical manifestations, and histopathological outcomes,192 underscoring the need for more comprehensive studies to support clinical translation.
4. Synthesis and biosynthesis
4.1. (Semi)synthesis of CGs
The structural diversity of CGs arises from variations in C-3, C-5, and C-17 stereoconfigurations, as well as the presence or absence of oxidizing groups and double bonds, ultimately influencing their biological efficacy.77 Extracts from natural plants have struggled to meet the market demand, and owing to the incomplete and immature understanding of their biosynthetic pathways, structural modification of these compounds remains limited and challenging. Therefore, chemical synthesis remains the predominant synthetic approach for CGs at present.196 Baran and co-workers accomplished the seminal total synthesis of ouabagenin from adrenosterone. This work pioneered a solution to the long-standing challenge of selective C–H hydroxylation at saturated carbon centers, achieving controllable oxidation processes and enabling access to diverse ouabagenin analogues.197,198 By 2019, the total chemical synthesis of the cardiac aglycons (+)-digitoxigenin199 and ouabagenin,25,26,200 alongside the CGs ouabain25,26 and rhodexin A,201,202 had been accomplished. In the past five years, total syntheses of other cardenolide aglycones, including 19-hydroxysarmentogenin,203 ouabagenin,203 and cinobufagin,204 have also been reported. Detailed chemical (semi)synthetic strategies for accessing cardiac aglycones and glycosides can be found in Table S1.The optimization of steps and synthetic conditions in recent years has led to the total chemical synthesis of the CGs cannogenol 3-O-α-L-rhamnoside (89),95 acovenoside A (90) and its congeners,97 rhodexin B (91),205 oleandrin (92), and beaumontoside206 (Fig. 6). The raw materials for chemical synthesis are primarily commercially available steroids or those obtained through simple, rapid syntheses of common steroids. Then, the process focuses on introducing a C-14 β-hydroxyl group and an unsaturated endolactone ring at the C-17 position, followed by a glycosylation reaction. The steps in the reaction flow to modify the steroidal parent nucleus are selected in the appropriate order to introduce specific functional groups—primarily hydroxyl or acetyl groups—onto the target CGs. The current reactive inertia of the C(sp3)–H bond at the C-14 position of CGs as well as spatial site-blocking effects makes direct C–H hydroxylation to install a hydroxyl group at C-14 extremely challenging. Most programs choose to introduce hydroxyl groups by first installing a double bond in ring D, followed by a hydration reaction. In 2019, Khatri et al. employed a Cu(II)-catalyzed diastereoselective Michael/aldol cascade approach to achieve rapid assembly of functionalized C-14 hydroxyl group-containing steroidal skeletons.95 The modification of C-17 is primarily achieved by subjecting the precursor to Barton vinyl iodide synthesis, followed by Stille or Suzuki coupling with a commercially available tin reagent. The modification of the steroidal scaffold primarily involves stereoselective protection and deprotection, epoxidation, and ring-opening reactions. The glycosylation step emphasizes the selection and optimization of catalysts and reaction conditions, considering the inherent differences in the reactivity of alcohol hydroxyl groups at various positions on the steroid scaffold. Most synthetic schemes select phosphine salts as catalysts, and different phosphine salts of various strong acids lead to different levels of α:β selectivity. Liu et al. used gold(I)-catalyzed glycosylation with superarmed glycosyl ortho-alkynylbenzoates as donors.97 The final step of the chemical synthesis is to perform global deprotection to obtain the target CG.
 | ||
| Fig. 6 Total chemical synthesis of CGs in recent years. TBS: tert-butyldimethylsilyl; TIPS: triisopropylsilyl; TMS: trimethylsilyl. | ||
In addition to conventional purely chemical synthesis, a chemoenzymatic approach has emerged. By combining the advantages of biocatalysis and chemical synthesis, this strategy demonstrates high regioselectivity and stereoselectivity, providing low-cost and eco-friendly synthesis routes to natural products that were previously difficult or inaccessible to produce using only contemporary chemical methods. This efficient approach to synthesizing natural products and their derivatives has become a growing trend.207–209 In recent years, researchers have begun exploring chemoenzymatic approaches to tackle the key steps in the synthesis of complex CGs, achieving notable progress (Fig. 7). The underlying synthetic principles of chemoenzymatic and chemical methods are similar. Currently, the main biocatalytic step is the formation of C-14 hydroxyl group, which offers considerable advantages over the cumbersome steps and low yields associated with chemical catalysis. Two P450 enzymes, Calotropis gigantea (L.) W. T. Aiton CYP11411 and B. gargarizans CYP44476, directly convert androstenedione (AD) to 14α-OH-AD. Subsequent chemical reactions produce key 14α-hydroxyl group steroid intermediates, ultimately enabling CG synthesis through the modular installation of a five- or six-membered lactone ring at the C-17 position. When the desired 17β-substituted orientation cannot be achieved through the above coupling reaction, an SN2-type-free radical reaction can be employed to control the configuration at C-17, guided by NOE correlations between the C-18 methyl group and the C-20 hydrogen atom.210 Moreover, a novel steroidal C14α-hydroxylase (CYP14A) with high catalytic activity and broad substrate specificity was mined and identified from Cochliobolus lunatus R.R. Nelson & F.A. Haasis (strain CGMCC 3.3589). Based on RoseTTaFold de novo prediction and molecular docking, a binding model for the CYP14A steroidal substrate was developed. By applying fixed-point saturating mutagenesis and a combined mutation strategy, mutants with notably improved catalytic efficiency and regioselectivity were obtained. The synthesis of cardenolide periplogenin, (+)-digitoxigenin, and its three diastereomers was further achieved through chemical methods.211
 | ||
| Fig. 7 The chemoenzymatic synthesis pathways of CGs. TBS: tert-butyldimethylsilyl; TMS: trimethylsilyl. | ||
4.2. Biosynthesis of CGs
The elucidation of the biosynthetic pathways of CGs is one of the most promising research areas for the future. It not only helps to reveal the molecular mechanisms underlying their natural synthesis but also provides a theoretical foundation and technical support for their efficient production. Biotransformation employs biological systems—such as bacteria, fungi, plant tissues, or isolated enzymes—to structurally modify exogenous compounds under mild conditions. This approach enables diverse chemical transformations that are often challenging in conventional organic synthesis, facilitating access to novel structural analogues.212 Although historically applied in the modification of CGs,213 biotransformation has been increasingly supplanted by advances in biosynthetic methods. Currently, based on advancements in genomics, transcriptomics, metabolomics, and enzyme functional studies, the hypothesized and partially elucidated biosynthetic pathways of CGs involve complex metabolic processes. These primarily include the synthesis of the steroidal backbone, formation of sugar moieties, and glycosylation of the aglycone with sugar units. The NCBI accession numbers of the relevant enzymes and their species information are provided in Table S2.β-Sitosterol, campesterol, and cholesterol biosynthesis. Cycloartenol serves as the common precursor for both phytosterols (e.g., campesterol and β-sitosterol) and cholesterol.216 The pathways diverge through early enzymatic steps: the phytosterol pathway is initiated by sterol C-24 methyltransferase 1 (SMT1)217,218 and phytosterol-specific C4 sterol methyl oxidase 1 (SMO1),219 while the cholesterol pathway begins with C24 sterol side-chain reductase 2 (SSR2)220,221 and pathway-specific C4 sterol methyl oxidase 3 (SMO3).222 A set of intermediate enzymes—cyclopropylsterol isomerase (CPI),223 sterol C-14 demethylase (CYP51),224 sterol C-14 reductase (C14-R),225 and sterol 8,7 isomerase (8,7-SI)226—are common to both pathways. Following these shared steps, the pathways diverge again. In the phytosterol branch, sterol C-24 methyltransferase 2 (SMT2) catalyzes a second methylation to form the C24-ethyl group characteristic of β-sitosterol.227 This pathway subsequently utilizes sterol methyl oxidase 2 (SMO2),228 sterol C-5(6) desaturase 1 (C5-SD1),229 7-dehydrocholesterol reductase 1 (7-DR1),230 and 24-sterol reductase (SSR1)231 to complete demethylation, dehydrogenation, hydrogenation, and side-chain reduction, yielding either β-sitosterol or campesterol. Conversely, cholesterol biosynthesis is completed through the actions of sterol methyl oxidase 4 (SMO4), sterol C-5(6) desaturase 2 (C5-SD2), and 7-dehydrocholesterol reductase 2 (7-DR2).216
Pregnenolone biosynthesis. The conversion of three sterols to pregnenolone involves the mitochondrial cytochrome P450 side-chain cleavage enzyme (P450scc) catalyzing the hydroxylation of C-22 and C-20 on the side chain and the break between the two carbons,232 which is the first and rate-limiting step in the control of steroid biosynthesis.233 P450scc has been characterized in both animals and plants. In animals, the Homo sapiens (Linnaeus, 1758) CYP11A1 gene234 and the B. gargarizans CYP11A1 gene235 encode cholesterol P450scc, which catalyzes the conversion of cholesterol to pregnenolone. Although the activity of plant P450scc on different sterol substrates was demonstrated decades ago,232 it was not definitively identified and characterized until 2023. As CGs are present only in leaves and not in roots, CYP87A4 from D. lanata was identified through differential gene expression analysis and confirmed to act on cholesterol or campesterol via yeast heterologous expression experiments.236 In the same year, on the basis of isotope labeling that confirmed pregnenolone as a precursor in CG biosynthesis, two P450scc enzymes—CYP87A106 from D. purpurea and CYP87A103 from Calotropis procera (Aiton) W. T. Aiton—were identified through differential expression analysis between leaf and root transcriptomes, along with BLAST searches. This finding was demonstrated in further experiments showing that both enzymes can produce pregnenolone using cholesterol, campesterol, and β-sitosterol as substrates. Additional CYP87A homologs identified across plant species via BLAST similarly exhibit sterol side-chain cleavage activity.237,238 The evolutionary mechanism by which CYP87A family proteins in plants acquire the catalytic activity required for pregnenolone biosynthesis remains unclear. These proteins differ in sequence from their mammalian counterpart, CYP11A1. Notably, amino acid substitutions such as A355V or L357A, based on CYP11A1, result in the loss of side-chain cleavage activity in DlCYP87A4 and EcCYP87A126. This suggests that mammalian P450scc is specialized for steroid hormone biosynthesis, whereas plant P450scc has evolved to synthesize specific metabolites.236,239,240
Progesterone biosynthesis. The current study suggests that the conversion of pregnenolone to progesterone occurs in two steps: the first step involves the oxidation of the C3-hydroxyl group to produce isoprogesterone, catalyzed by NAD+-dependent 3β-hydroxysteroid dehydrogenase (3βHSD), and the second step involves the migration of the double bond from position 5 to position 4. The currently characterized plant 3βHSDs241,242 belong to the short-chain dehydrogenase/reductase gene family,243 and serve to oxidize and isomerize the precursor compound sterol. The second step of isomerization to progesterone is irreversible.
Pregnanolone biosynthesis. The types of CGs reported in plants are mostly in the C-3 β-configuration and have a cis-fused A/B ring junction. In these plants, progesterone is converted by NADPH-dependent co-reactive progesterone-5β-reductases (St5βR)244,245 and 3βHSD to pregnanolone (3β-hydroxy-5β-pregane-20-one). Enzymes exhibiting St5βR activity are present in numerous plants that both produce and do not produce CGs. Recent studies have revealed that these enzymes also function as iridoid synthases, and are now collectively referred to as PRISEs (progesterone 5β-reductase/iridoid synthase-like enzymes).246,247 In plants producing CGs, the presence of progesterone-5α-reductases (St5αR),248 3βHSD, and 3α-hydroxysteroid 5β-oxidoreductases (3αHSD)249 enables the generation of the corresponding products with a C-3 α-configuration or trans-fused A/B ring junction. Notably, enzymes with confirmed 3αHSD activity remain limited and generally demonstrate low catalytic efficiency—an issue that still requires resolution.
Downstream biosynthesis processes for the steroid nucleus. In addition to the two CYP450 s used in the chemoenzymatic synthesis described above, which catalyze the 14α-hydroxylation of precursors, enzymes involved in the 14β-hydroxylation of pregnanolone in plants have not been identified. In 2022, Zhao et al. identified the key P450 enzyme (CYP11411) from C. gigantea that catalyzed the direct C–H activation at the C-14 position of AD, resulting in the retention of the 14α-OH-AD product configuration via the canonical “oxygen rebound” mechanism.156 To date, all reported enzymatic hydroxylation reactions of tertiary carbon centers in natural products have maintained their stereoconfigurations, as shown in the previous example.250 Therefore, 14β-hydroxylation of tertiary carbon centers in progesterone may occur through hydrogen isomerization (α-H to β-H) followed by hydroxylation, or via direct hydroxylation through a stereoinversion, which involves hydrogen atom abstraction, electron transfer, the formation of a stable cation intermediate via isomerization, and the subsequent attack of the cation by water. The 21-hydroxylase CYP21A1 has been reported only in mammals (e.g., Mus musculus (Linnaeus, 1758)).251 21-Hydroxypregnane 21-O-malonyltransferases (21MAT) transfer the malonyl group from malonyl-CoA to the C-21 hydroxyl group of the pregnane precursor 5β-pregnane-3β,14β,21-triol-20-one generated in the above two steps. Among the tested substrates, the free ketol exhibited the lowest acceptance in this acyltransferase reaction, whereas the 3-O-glycosylated ketol demonstrated significantly higher enzyme affinity, supporting the hypothesis that glycosylation may occur at an early stage in CG biosynthesis.252 The enzymes involved in the subsequent cyclization to generate unsaturated five-membered or six-membered lactone rings at C-17 remain to be elucidated. Notably, under physiological conditions, analogues of 21-O-malonyl-5-pregnane-3β,14β-diol-20-one undergo only very slow lactonization, whereas those lacking the 14β-hydroxyl group show negligible reactivity, indicating that 14β-hydroxylation precedes lactone formation.253
Studies on the biosynthetic reaction mechanisms, reaction sequences, and related enzyme genes of bufadienolides have been rarely reported. The putative biosynthetic pathway of bufadienolides in plants initiates from the precursor 5β-pregnane-3β,14β,21β-triol-20-one. This intermediate undergoes acylation with oxaloacetyl-CoA, followed by aldol condensation, decarboxylation, reduction, and elimination reactions, ultimately producing a six-membered unsaturated lactone ring.51 Isotope tracing revealed that cholesterol is the biosynthetic precursor of marinobufotoxin and marinobufagenin.254 Unlike in plants, the biosynthesis of bufadienolides in toads does not involve the cleavage of cholesterol side chains.255 Thus, pregnenolone does not act as a precursor.256,257 These findings align with the aforementioned evolutionary implications regarding P450scc and further suggests the existence of an alternative biosynthetic pathway in animals. Based on transcriptome analysis, the cholesterol-bile acid–bufadienolide pathway is currently proposed. In toads, the process begins with the MVA pathway, then, lanosterol undergoes a series of reactions to produce the key intermediate, cholesterol. CYP27A1 initiates the conversion of cholesterol to lithocholic acid, a bile acid salt intermediate,258 which is subsequently converted into bufadienolides through a series of unknown steps.259 The enzymes 3βHSD260 and steroid 5β-reductase (SRD5β),261 which modify the steroid skeleton, have also been reported in toads. In addition, Lei et al. identified five CYP450 enzymes from toads,262 which are similar to the previously reported fungal CYP enzyme Sth10 from Thanatephorus cucumeris (A.B. Frank) Donk (starin NBRC 6298).263 This enzyme catalyzes the hydroxylation of bufalin and resibufogenin at different sites to yield compounds 93–105.
S3GTs capable of glycosylation modification on intact CGs have been identified, including microbial-derived OleD from Streptomyces antibioticus (Waksman & Woodruff, 1941) Waksman & Henrici, 1948 (ref. 271) and YjiC1 from Bacillus subtilis (Ehrenberg, 1835) Cohn, 1872,272 but suffer from low catalytic efficiency and poor regioselectivity. Three plant GTs catalyze the glycosylation of substrates 106–117 to form their corresponding CGs (Fig. 9). In 2017, Wen et al. reported for the first time that UDP-glycosyltransferase (UGT) 74AN1 from Asclepias curassavica Lour., a plant UDP-GT with substrate promiscuity, catalyzes the formation of 3-O-β-D-glucosides from various intermediates in the CG biosynthesis pathway and is the first to catalyze cardiac steroid 3α-hydroxyglycosylation of GT. UGT74AN1 accepts UDP-Glc as the primary sugar donor and only minimally utilizes UDP-GlcNAc.267 In addition, based on the use of suspension cultures of plant cells from several non-cardiac steroid-producing species for CG biotransformation,52 the team identified the first UGT74AN3 from the non-CG-producing plant C. roseus that accepts only UDP-Glc as its major sugar donor and is catalytically active toward eight structurally distinct CTSs and phenolic compounds.273 In 2022, Huang et al. identified and characterized the plant S3GT UGT74AN2 from C. gigantea, capable of generating various cardiotonic steroid 3-O-glycosides using UDP-Glc, UDP-GlcNAc, and UDP-Gal as sugar donors, and through structure- and sequence-based engineering, yielded the triple mutant UGT74AN2 I284R/W390H/V391G, which exhibits broader sugar donor specificity and enhanced catalytic activity.274
| Products | Microbial strains | Fermentation vessels | Carbon source | Yield | References |
|---|---|---|---|---|---|
| Campesterol | Y. lipolytica | 5 L bioreactor | Sunflower seed oil | 942 mg L−1 | 277 |
| S. cerevisiae | 5 L fermenter | Glucose | 916.9 mg L−1 | 281 | |
| Y. lipolytica | 5 L fermenter | Glucose | 837 mg L−1 | 280 | |
| C. jadinii | Shake flask | Glucose | 92.1 mg L−1 | 282 | |
| 5 L bioreactor | 807 mg L−1 | ||||
| S. cerevisiae | 96-Well plates | Glucose | 18.4 mg L−1 | 279 | |
| β-Sitosterol | S. cerevisiae | 96-Well plates | Glucose | 2 mg L−1 | 278 |
| Cholesterol | S. cerevisiae | 5 L fermenter | Glucose | 5.5 g L−1 | 283 |
| C. jadinii | Shake flask | Glucose | 81.8 mg L−1 | 282 | |
| 5 L bioreactor | 1.52 g L−1 | ||||
| S. cerevisiae | 5 L bioreactor | Glucose | 1.78 g L−1 | 288 | |
| Pregnenolone | Y. lipolytica | 5 L bioreactor | Glucose or sunflower seed oil | 78 mg L−1 | 287 |
| S. cerevisiae | 5 L bioreactor | Glucose | 0.83 g L−1 | 288 | |
| Progesterone | M. neoaurum | Shake flask | Glucose | 235 mg L−1 | 290 |
| S. cerevisiae | 5 L fermenter | Glucose and ethanol | 1.06 g L−1 | 238 |
In 2016, Du et al. introduced an exogenous 7-dehydrocholesterol reductase (DHCR7) into Yarrowia lipolytica (Wick., Kurtzman & Herman) Van der Walt & Arx, using sunflower seed oil as a carbon source. This enabled campesterol production with a yield of 453 ± 24.7 mg L−1via high-cell-density fermentation in a 5 L bioreactor.276 In the following year, they screened DHCR7 from Danio rerio (Hamilton, 1822) and enhanced the expression of the key gene POX2, which increased the yield of campesterol to 942 ± 50.1 mg L−1.277 By 2020, Xu et al. reconstituted the multi-enzyme pathway for synthesizing various phytosterols in Saccharomyces cerevisiae (Desm.) Meyen through metabolic engineering, strain evolution, and fermentation engineering, achieving a campesterol yield of 7 mg L−1 and β-sitosterol yield of 2 mg L−1 through fermentation in 96-well plates,278 and the campesterol yield was increased to 18.4 mg L−1 in 2023 by partially restoring the activity of the sterol acyltransferase ARE2 in yeast and enhancing the upstream FPP supply.279 In 2020, Qian et al. similarly achieved the de novo synthesis of campesterol in Y. lipolytica by knocking down multifunctional β-oxidation protein (Mfe1) and peroxisomal biogenesis factor 10 (PEX10) and heterologously introducing DHCR7, to obtain a yield of 837 mg L−1 through incubation in a 5 L fermenter for 144 h.280 In 2021, Zhou et al. achieved a campesterol yield of 916.9 mg L−1 through fermentation by overexpressing a screened heterologous DHCR7 gene combined with an endogenous promoter engineering strategy, using a 5 L fermenter for fed-batch fermentation process.281 By 2023, Gu et al. developed a highly efficient CRISPR/Cas9 technique for metabolic pathway modification in the polyploid industrial yeast Cyberlindnera jadinii (Sartory, R. Sartory, J. Weill & J. Mey.) Minter 2009, achieving yields of 92.1 and 81.8 mg L−1 of campesterol and cholesterol, respectively, via shake flask fermentation. High-density replenishment of batch fermentation further increased the yields to 807 mg L−1 and 1.52 g L−1, marking the first gram-scale production of a steroidal compound in C. jadinii.282 In addition, Xu et al. achieved a cholesterol yield of 5.5 g L−1 during high-density replenishment batch fermentation, enabling the production of 2.03 g L−1 of diosgenin through synthetic-pathway optimization, fine-tuning of gene expression, and elimination of competing pathways.283
Except for sterols, the de novo synthesis of pregnenolone in S. cerevisiae was achieved as early as 1988, yielding 60 mg L−1 using glucose as the carbon source;284 however, no substantial breakthrough has been reported to date. The P450scc system comprises the P450 monooxygenase CYP11A1 and its natural redox partners adrenodoxin (Adx) and adrenodoxin reductase.285 CYP11A1 catalyzes the conversion of sterols to pregnenolones, which is the first and limiting step in steroidogenesis. By 2013, Makeeva et al. constructed a stable transgenic Escherichia coli (Migula, 1895) Castellani & Chalmers, 1919 strain with a functionally reconstructed bovine cholesterol pathway, yielding 0.42 ± 0.015 mg L−1 of pregnenolone from 500 μM (193 mg L−1) of cholesterol in 24 h.286 In 2019, Zhang et al. introduced the mammalian P450scc CYP11A1 and its electron transport chain into campesterol-producing engineered Y. lipolytica strains developed in their earlier study,277 achieving a pregnenolone yield of 78.0 mg L−1.287 In 2025, Chen et al. achieved cholesterol and pregnenolone titers of 1.78 g L−1 and 0.83 g L−1, respectively, in engineered yeast via rational engineering of DHCR7 and RgCYP87A3 guided by computational simulations and QM/MM analysis, coupled with mitochondrial co-localization of electron transfer components and pathway optimization.288
In 2014, Strizhov et al. achieved a progesterone yield of 25 mg L−1 from 7.5 mM cholesterol using Mycolicibacterium smegmatis (Trevisan 1889) Gupta et al. 2018 (strain mc2155).289 By 2022, Liu et al. reported that CYP11A1 can also catalyze the side-chain cleavage of 4-HBC to produce progesterone. They introduced CYP11A1 into Mycobacterium neoaurum Tsukamura, 1972, a strain with high 4-HBC yield, and further accelerated the electron transfer to achieve the first green and sustainable fermentation production of phytosterols to progesterone, achieving a yield of 45 mg L−1 under light-emitting diode light-driven conditions. By combining engineered M. neoaurum and InP nanoparticles to form a novel inorganic–biological hybrid system, a yield up to 235 ± 50 mg L−1 was achieved.290 Moreover, an engineered M. smegmatis mc2155 strain produced 85.2 ± 4.7 mol% 3-methoxymethyl-pregnenolone within 48 hours. Subsequent acid hydrolysis removed the protecting group, yielding high-purity pregnenolone via crystallization.291 Over the past two decades, yeast-engineered strains have remained suboptimal for progesterone production compared to microbial conversion. In 2025, Li et al. screened P450scc, cytochrome P450 reductase (CPR), and HSD from Marsdenia tenacissima (Roxb.) Wight & Arn. By minimizing intracellular shuttling of substrates and enzymes in engineered S. cerevisiae yeast, they achieved progesterone production of 1.06 g L−1via fermentation using simple carbon sources.238
Optimization of secondary metabolite production of CGs in plant factories is also a key research direction. In D. purpurea stem cultures, the accumulation of digitoxin and digoxin was increased up to 9.1-fold and 11.9-fold, respectively, through induction and feeding with the precursor progesterone (200–300 mg L−1).292 By expressing P5βR from Arabidopsis thaliana (L.) Heynh. in D. purpurea cultured in vitro, digitoxin and digoxin contents were increased up to 3.8-fold and 2.2-fold, respectively.14
5. Conclusions and prospects
CGs have been a much publicized and controversial compound since their discovery. Achieving a balance between the therapeutic efficacy and side effects of CGs remains a critical concern. The pharmacological activity of CGs in the treatment of cardiovascular diseases was among the earliest areas to be investigated and applied in the clinic. Digoxin is not per se cheap, while digoxin-based medications are often administered at very small doses, making it accordingly low-cost medications available for the treatment of heart failure; however, its frequency of use is currently declining dramatically.293 Contrastingly, for patients with atrial fibrillation who cannot control their heart rate with other medications, CGs remain one of the most commonly used medications worldwide.294 With the deepening research on the pharmacological activities of CGs in multidisciplinary medical fields, their wide range of pharmacological effects has been gradually discovered, which is expected to lead to the development of new therapeutic drugs for clinical diseases across various fields, including oncology, neurology, respiratory diseases, and immune system disorders. Besides the widely-known NKA, several studies have found that CGs act on various new drug targets and are expected to be the most promising broad-spectrum drugs for application. Notably, given the pleiotropic effects of CGs, their optimal concentration as drugs warrants further investigation to ensure the stability and survival of all animal cells for maximum cost-effectiveness; it is essential to avoid cytotoxicity to functionally important differentiated cells while targeting harmful tumor or senescent cells.Although hundreds of CGs are found in plants, only 20 to 30 of them are clinically applicable, with merely six or seven being commonly used. An examination of pharmacologic studies of CGs revealed that there is adequate data only for digoxin and digitoxin. The complex chemical structure and diversity of sugar residues of CGs enhance their variety and biological activity as drugs. Low solubility and severe cardiotoxicity pose serious limitations to the therapeutic use of CGs.295 Nonetheless, C-3 glycosylation modification enhances the solubility and reduces the toxicity of this class of drugs, which is considered a practical approach to improve their pharmacodynamic and pharmacokinetic properties and enhance their biological activity.296–298 Studies on different structural CGs for the treatment of clinical diseases can help us better understand the intrinsic link between structure and pharmacodynamics. Large-scale, high-quality prospective efficacy studies or clinical trials with low bias and minimal randomization error are needed in the future. These studies should utilize evidence from ongoing pharmacokinetic and pharmacodynamic studies to comprehensively evaluate the safety, efficacy, and therapeutic effects of CGs on pharmacological actions, as well as assess the efficacy of combination therapeutic strategies.
Furthermore, since the 1980s, discoveries and pharmacological studies on endogenous CGs have enhanced the understanding of human physiological and pathophysiological processes, opening new avenues for therapeutic targeting. Numerous studies have confirmed that endogenous ouabain and marinobufagenin, two of the most prominent steroid hormones, play an important role in the development of cardiovascular diseases such as hypertension and heart failure.64 However, their existence, structure, biosynthetic pathways, and mechanisms of action remain unclear or controversial.1,63 Urgent research priorities for endogenous CGs include elucidating their biosynthetic pathways, defining their physiological and pathophysiological roles in cardiovascular and metabolic regulation, deciphering NKA-mediated signaling mechanisms, standardizing detection methods, and exploring their therapeutic targeting potential. The resolution of these questions will not only advance drug discovery but also improve our understanding of evolutionary biology and adaptive physiology.
The traditional methods of obtaining active ingredients from TCM, such as raw plant extraction, do not fully meet the clinical needs and have a negative impact on the natural environment. The scarcity of medicinal plant resources and the incomplete analysis of biosynthetic pathways continue to constrain the study of CGs. Although the total chemical synthesis of some CGs has been achieved, challenges remain, such as cumbersome steps, low yields, and harsh reaction conditions. Especially, the synthesis of the C-14 β-hydroxyl group and C-17 unsaturated lactone rings—the key structures of CG compounds—remains challenging due to spatial site resistance. Most chemically synthesized raw materials, including unsaturated lactone rings and different types of sugar donors, are still obtained commercially and cannot be produced cost-effectively. Currently, biocatalysts are applied in chemoenzymatic synthesis approach for the synthesis of the C-14 hydroxyl group, which considerably simplifies the synthesis steps and demonstrates great potential for future applications. In nature, many biologically active CGs are decorated with specific functional groups such as hydroxyl, carbonyl, and glycosyl moieties at various positions. Representative examples including digoxin, ouabain, and convallatoxin possess characteristic hydroxylations at C-12 or C-19, which significantly contribute to their potent bioactivities. Chemically introducing these functional groups—especially the C-19 hydroxyl—remains challenging due to issues such as poor regio- and stereoselectivity, lengthy synthetic routes, and low yields. As a promising alternative, chemoenzymatic synthesis offers a more efficient and selective approach. For instance, a CYP450 capable of catalyzing C-19 hydroxylation of steroidal substrates has been reported, providing a potential biocatalytic tool for modifying CGs at this position.263 This enzyme and similar biocatalysts could be harnessed in the future for the synthesis of complex natural CGs and their analogues. Furthermore, there is a continuing need to discover and characterize novel enzymes that can functionalize distinct sites (e.g., C-5, C-14, and C-19) of the cardenolide scaffold. Expanding this biocatalytic toolbox will facilitate precise structural diversification and efficient synthesis of CGs.
Access to rare sugar donors through biosynthesis has also been explored to some extent,299 suggesting that biosynthesis could be the most promising and widely applied alternative method to produce CGs. In recent years, new technologies and methods, such as high-throughput screening, molecular probes, and genome-wide association studies, have emerged, accelerating the elucidation of the complete biosynthetic pathway of CGs. Low-cost genome sequencing, advances in computational tools, and the application of artificial intelligence and machine learning technologies are revolutionizing the ability to identify, understand, and manipulate biosynthetic pathways. Currently reported P450 enzymes have low activity, and heterologous biosynthesis is still limited; therefore, the modification of enzyme function must be addressed. Methods and applications based on structure and machine learning-guided enzyme design make enzymes programmable and predictable, enabling them to execute chemical reactions that do not occur naturally. This advancement will greatly contribute to the popularization of biocatalysis.300 Protein engineering utilizing tools such as AlphaFold for protein structure prediction301 and protein crystal structure resolution technology,302 enables the precise modification of the catalytic properties of enzymes with low catalytic activity. This approach will lay the foundation for the heterologous production of CGs in microorganisms by using synthetic biology methods. In summary, synergistic breakthroughs in underlying technology clusters and intelligent design of enzyme functions will lead to a digital revolution in biosynthesis, facilitating the discovery of new CG drugs, more sustainable production methods, and a deeper understanding of complex biological processes.
6. Author contributions
Conceptualization, L. H., P. S., and S. L.; formal analysis and investigation, D. J.; writing – original draft preparation, D. J., Y. Z., and W. G.; writing – review and editing, D. J., Y. Z., P. S., and S. L.; visualization, D. J., and W. G.; project administration, P. S., and S.L. All authors have read and agreed to the published version of the manuscript.7. Conflicts of interest
There are no conflicts to declare.8. Data availability
This review synthesizes existing data from publicly available sources. No primary research results, software or code have been included and no new data were generated or analysed as part of this review. Drug approval and clinical trial data were extracted from: WHO International Clinical Trials Registry Platform (https://www.who.int/ictrp), U.S. FDA Drugs Database (https://www.accessdata.fda.gov/scripts/cder/daf), China Center for Drug Evaluation (https://www.cde.org.cn), U.S. National Library of Medicine ClinicalTrials.gov (https://clinicaltrials.gov). All data supporting the conclusions are derived from the cited references and repositories, with access details provided in the reference list.Supplementary information: detailed chemical synthetic approaches to cardiac aglycons and glycosides and the NCBI accession numbers for relevant enzymes. See DOI: https://doi.org/10.1039/d5np00050e.
9. Acknowledgements
This work was supported by National Natural Science Foundation of China (82204568, 82304672), the Fundamental Research Funds for the Central Public Welfare Research Institutes (ZZ16-YQ-037, ZZ16-YQ-042, ZZXT202201), key project at central government level: the ability establishment of sustainable use for valuable Chinese medicine resources (2060302), and Scientific and Technological Innovation Project of the China Academy of Chinese Medical Sciences (CI2023E002, CI2024C004YN).10. Notes and references
- D. Pavlovic, Cell Calcium, 2020, 86, 102156 CrossRef CAS.
- R. C. Batterman and A. C. De Graff, Am. Heart J., 1947, 34, 663–673 CrossRef CAS.
- M. B. Kreidberg, H. L. Chernoff and W. L. Lopez, N. Engl. J. Med., 1963, 268, 23–30 CrossRef CAS.
- P. G. Louw, Nature, 1949, 163, 30 CrossRef CAS.
- M. Friedman and R. Bine Jr, J. Clin. Invest., 1949, 28, 32–34 CrossRef CAS.
- N. U. Fedosova, M. Habeck and P. Nissen, Compr. Physiol., 2021, 12, 2659–2679 CrossRef.
- P. L. Jorgensen, K. O. Hakansson and S. J. Karlish, Annu. Rev. Physiol., 2003, 65, 817–849 CrossRef CAS.
- X. López-Goldar and A. A. Agrawal, Evolution, 2023, 77, 2431–2441 CrossRef PubMed.
- M. Herbertz, S. Dalla, V. Wagschal, R. Turjalei, M. Heiser and S. Dobler, Mol. Ecol., 2024, 33, e17041 CrossRef CAS PubMed.
- A. A. Agrawal and A. P. Hastings, Proc. Natl. Acad. Sci. U. S. A., 2023, 120, e2302251120 CrossRef CAS.
- A. Betz, R. Bischoff and G. Petschenka, Proc. Biol. Sci., 2024, 291, 20232721 CAS.
- F. Beran and D. G. Heckel, Mol. Ecol., 2024, 33, e17443 CrossRef CAS PubMed.
- L. Espinosa Del Alba and G. Petschenka, J. Insect Physiol., 2023, 147, 104508 CrossRef CAS PubMed.
- E. Kairuz, N. Pérez-Alonso, A. Capote-Pérez, A. Pérez-Pérez, A. A. Espinosa-Antón, G. Angenon, E. Jiménez and B. Chong-Pérez, Ind. Crops Prod., 2020, 146, 11 CrossRef.
- S. K. Verma, S. Gantait, B. R. Jeong and S. J. Hwang, Sci. Rep., 2018, 8, 18009 CrossRef CAS PubMed.
- C. Leeb, C. Brühl and K. Theissinger, Sci. Total Environ., 2020, 706, 134430 CrossRef CAS PubMed.
- P. Knozowski, J. J. Nowakowski, A. M. Stawicka, A. Górski and B. Dulisz, Sci. Total Environ., 2023, 898, 165280 CrossRef CAS.
- R. Zheng, X. Chen, C. Ren, Y. Teng, Y. Shen, M. Wu, H. Wang and M. Huang, Chemosphere, 2020, 247, 125925 CrossRef CAS.
- M. Y. Huang, R. Y. Duan, J. W. Yin, Q. Zhao, Y. Y. Wan and Y. Liu, Aquat. Toxicol., 2020, 229, 105671 CrossRef CAS PubMed.
- Z. Y. Gao, P. Jiang and C. S. Zhan, Zhongguo Zhongyao Zazhi, 2022, 47, 6034–6041 CAS.
- J. Guo, Y. J. Zhou, M. L. Hillwig, Y. Shen, L. Yang, Y. Wang, X. Zhang, W. Liu, R. J. Peters, X. Chen, Z. K. Zhao and L. Huang, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 12108–12113 CrossRef CAS.
- D. K. Ro, E. M. Paradise, M. Ouellet, K. J. Fisher, K. L. Newman, J. M. Ndungu, K. A. Ho, R. A. Eachus, T. S. Ham, J. Kirby, M. C. Chang, S. T. Withers, Y. Shiba, R. Sarpong and J. D. Keasling, Nature, 2006, 440, 940–943 CrossRef CAS PubMed.
- B. Jiang, L. Gao, H. Wang, Y. Sun, X. Zhang, H. Ke, S. Liu, P. Ma, Q. Liao, Y. Wang, H. Wang, Y. Liu, R. Du, T. Rogge, W. Li, Y. Shang, K. N. Houk, X. Xiong, D. Xie, S. Huang, X. Lei and J. Yan, Science, 2024, 383, 622–629 CrossRef CAS PubMed.
- J. Zhang, L. G. Hansen, O. Gudich, K. Viehrig, L. M. M. Lassen, L. Schrübbers, K. B. Adhikari, P. Rubaszka, E. Carrasquer-Alvarez, L. Chen, V. D'Ambrosio, B. Lehka, A. K. Haidar, S. Nallapareddy, K. Giannakou, M. Laloux, D. Arsovska, M. A. K. Jørgensen, L. J. G. Chan, M. Kristensen, H. B. Christensen, S. Sudarsan, E. A. Stander, E. Baidoo, C. J. Petzold, T. Wulff, S. E. O'Connor, V. Courdavault, M. K. Jensen and J. D. Keasling, Nature, 2022, 609, 341–347 CrossRef CAS.
- H. Zhang, M. Sridhar Reddy, S. Phoenix and P. Deslongchamps, Angew Chem. Int. Ed. Engl., 2008, 47, 1272–1275 CrossRef CAS PubMed.
- M. S. Reddy, H. Zhang, S. Phoenix and P. Deslongchamps, Chem.–Asian J., 2009, 4, 725–741 CrossRef CAS.
- H. Y. Yang, Y. X. Chen, S. Luo, Y. L. He, W. J. Feng, Y. Sun, J. J. Chen and K. Gao, RSC Adv., 2022, 12, 23240–23251 RSC.
- Y. Takada, K. Kaneko and Y. Kawakami, J. Membr. Biol., 2023, 256, 229–241 CrossRef CAS PubMed.
- J. Long, J. C. Ouyang, Y. H. Luo, Q. J. Wu, X. T. Liao, Z. L. Chen, Q. L. Wang, X. Y. Liang, L. Liu, X. M. Yang and X. S. Li, Nat. Prod. Res., 2024, 38, 211–219 CrossRef CAS PubMed.
- H. L. Ran, S. Z. Huang, H. Wang, L. Yang, C. J. Gai, R. J. Duan, H. F. Dai, Y. L. Guan and W. L. Mei, Phytochemistry, 2023, 210, 113668 CrossRef CAS PubMed.
- Y. L. He, H. Y. Yang, L. Zhang, Z. Gong, G. L. Li and K. Gao, Chem. Biodiversity, 2024, 21, e202401460 CrossRef CAS.
- Y. Xu, J. Xu, W. Zhu, Y. Yan, X. Jiang, Z. Xie, F. Feng and J. Zhang, Planta Med., 2023, 89, 1444–1456 CrossRef CAS.
- Y. Ren, E. N. Kaweesa, L. Tian, S. Wu, K. Sydara, M. Xayvue, C. E. Moore, D. D. Soejarto, X. Cheng, J. Yu, J. E. Burdette and A. D. Kinghorn, J. Nat. Prod., 2023, 86, 1411–1419 CrossRef CAS.
- A. J. Ji, Q. Ma, M. Y. Kong, L. Y. Li, X. L. Chen, Z. Q. Liu, J. J. Wu and R. R. Zhang, Chin. J. Nat. Med., 2022, 20, 202–209 CAS.
- M. Huang, S. Shen, C. Luo and Y. Ren, Molecules, 2019, 24, 2749 CrossRef.
- X. Shang, X. Miao, F. Yang, C. Wang, B. Li, W. Wang, H. Pan, X. Guo, Y. Zhang and J. Zhang, Front. Pharmacol., 2019, 10, 25 CrossRef CAS.
- J. Cai, B. D. Zhang, Y. Q. Li, W. F. Zhu, T. Akihisa, T. Kikuchi, J. Xu, W. Y. Liu, F. Feng and J. Zhang, Bioorg. Chem., 2022, 127, 106004 CrossRef CAS PubMed.
- A. A. Agrawal, G. Petschenka, R. A. Bingham, M. G. Weber and S. Rasmann, New Phytol., 2012, 194, 28–45 CrossRef CAS PubMed.
- E. Kakouri, C. Kanakis, P. Trigas and P. A. Tarantilis, Anal. Bioanal. Chem., 2019, 411, 3135–3150 CrossRef CAS.
- O. Potterat, M. Kaufmann, C. Tschopp, M. Caj, J. K. Reinhardt, A. Prescimone, D. Shah, S. Baumgartner, M. A. Sciotti and L. Suter-Dick, Phytochemistry, 2025, 230, 114329 CrossRef CAS PubMed.
- J. Guo, F. Chen, W. Zhang, H. Bai, L. Li, Y. Ma and Z. Yang, J. Sep. Sci., 2024, 47, e2400145 CrossRef.
- J. Kolodziejczyk-Czepas and A. Stochmal, Phytochem. Rev., 2017, 16, 1155–1171 CrossRef CAS PubMed.
- L. Krenn, B. Kopp, A. Deim, W. Robien and W. Kubelka, Planta Med., 1994, 60, 63–69 CrossRef CAS.
- B. Kopp, L. Krenn, M. Draxler, A. Hoyer, R. Terkola, P. Vallaster and W. Robien, Phytochemistry, 1996, 42, 513–522 CrossRef CAS PubMed.
- M. Iizuka, T. Warashina and T. Noro, Chem. Pharm. Bull., 2001, 49, 282–286 CrossRef CAS.
- W. Cheng, Y. F. Tan, H. Y. Tian, X. W. Gong, K. L. Chen and R. W. Jiang, Nat. Prod. Res., 2014, 28, 901–908 CrossRef CAS PubMed.
- T. Brillatz, M. Jacmin, K. Vougogiannopoulou, E. A. Petrakis, E. Kalpoutzakis, J. Houriet, L. Pellissier, A. Rutz, L. Marcourt, E. F. Queiroz, A. D. Crawford, A. L. Skaltsounis and J. L. Wolfender, J. Ethnopharmacol., 2020, 259, 112954 CrossRef CAS.
- Y. Li, H. Zhang, X. Liang, B. Song, X. Zheng, R. Wang, L. Liu, X. Song and J. Liu, Nat. Prod. Res., 2020, 34, 950–957 CrossRef CAS PubMed.
- O. M. Khattab, D. M. El-Kersh, S. A. M. Khalifa, N. Yosri, H. R. El-Seedi and M. A. Farag, Plants, 2023, 12, 2078 CrossRef CAS.
- J. Yang, Y. H. Zhang, F. Miao, L. Zhou and W. Sun, Fitoterapia, 2010, 81, 636–639 CrossRef CAS.
- P. S. Steyn and F. R. van Heerden, Nat. Prod. Rep., 1998, 15, 397–413 RSC.
- M. Ye, J. G. Dal, H. Z. Guo, Y. J. Cui and D. Guo, Tetrahedron Lett., 2002, 43, 8535–8538 CrossRef CAS.
- C. Zhu, X. Lu, T. Cai, K. Zhu, L. Shi, Y. Chen, T. Wang, Y. Yang, D. Tu, Q. Fu, J. Huang and Y. Zhen, PNAS Nexus, 2024, 3, pgae215 CrossRef CAS.
- W. L. Wei, J. J. Hou, X. Wang, Y. Yu, H. J. Li, Z. W. Li, Z. J. Feng, H. Qu, W. Y. Wu and D. A. Guo, J. Ethnopharmacol., 2019, 237, 215–235 CrossRef CAS PubMed.
- J. Slagboom, C. Kaal, A. Arrahman, F. J. Vonk, G. W. Somsen, J. J. Calvete, W. Wüster and J. Koo, Microchem. J., 2022, 175, 107187 CrossRef CAS.
- J. M. Hamlyn, M. P. Blaustein, S. Bova, D. W. DuCharme, D. W. Harris, F. Mandel, W. R. Mathews and J. H. Ludens, Proc. Natl. Acad. Sci. U. S. A., 1991, 88, 6259–6263 CrossRef CAS PubMed.
- S. N. Orlov, A. M. Tverskoi, S. V. Sidorenko, L. V. Smolyaninova, O. D. Lopina, N. O. Dulin and E. A. Klimanova, Genes Dis., 2021, 8, 259–271 CrossRef CAS PubMed.
- J. G. Wang, J. A. Staessen, E. Messaggio, T. Nawrot, R. Fagard, J. M. Hamlyn, G. Bianchi and P. Manunta, J. Hypertens., 2003, 21, 1475–1481 CrossRef CAS.
- S. Harwood, A. M. Mullen, A. C. McMahon and A. Dawnay, Am. J. Hypertens., 2001, 14, 1112–1115 CrossRef CAS.
- E. Berendes, P. Cullen, H. Van Aken, W. Zidek, M. Erren, M. Hübschen, T. Weber, S. Wirtz, M. Tepel and M. Walter, Crit. Care Med., 2003, 31, 1331–1337 CrossRef CAS PubMed.
- L. K. Lewis, T. G. Yandle, J. G. Lewis, A. M. Richards, G. B. Pidgeon, R. J. Kaaja and M. G. Nicholls, Hypertension, 1994, 24, 549–555 CrossRef CAS.
- S. Baecher, M. Kroiss, M. Fassnacht and M. Vogeser, Clin. Chim. Acta, 2014, 431, 87–92 CrossRef CAS.
- L. K. Lewis, T. G. Yandle, P. J. Hilton, B. P. Jensen, E. J. Begg and M. G. Nicholls, Hypertension, 2014, 64, 680–683 CrossRef CAS PubMed.
- C. Staehr, C. Aalkjaer and V. V. Matchkov, Clin. Sci., 2023, 137, 1595–1618 CrossRef CAS.
- A. Y. Bagrov, J. I. Shapiro and O. V. Fedorova, Pharmacol. Rev., 2009, 61, 9–38 CrossRef CAS PubMed.
- R. Xu, J. Du, L. Deng, F. Yang, J. Zhang, D. Wang and Y. Zhang, Zhongguo Zhongyao Zazhi, 2012, 37, 2286–2288 CAS.
- Y. Li, Y. B. Liu, S. S. Yu, X. G. Chen, X. F. Wu, S. G. Ma, J. Qu, Y. C. Hu, J. Liu and H. N. Lv, Steroids, 2012, 77, 375–381 CrossRef CAS PubMed.
- S. Martucciello, G. Paolella, A. M. Romanelli, S. Sposito, L. Meola, A. Cerulli, M. Masullo, S. Piacente and I. Caputo, Molecules, 2022, 27, 4874 CrossRef CAS.
- R. M. Osman Mohammed, Y. Huang, X. Guan, X. Huang, S. Deng, R. Yang, J. Li and J. Li, Phytochemistry, 2022, 200, 113239 CrossRef CAS PubMed.
- M. Mirzaei, G. C. Younkin, A. F. Powell, M. L. Alani, S. R. Strickler and G. Jander, Plants, 2024, 13, 466 CrossRef CAS.
- M. Mirzaei, T. Züst, G. C. Younkin, A. P. Hastings, M. L. Alani, A. A. Agrawal and G. Jander, J. Chem. Ecol., 2020, 46, 1131–1143 CrossRef CAS.
- H. J. McCoy, C. Zeng, E. McCoy, P. MacKinley, J. Vickruck, L. A. Calhoun and H. H. Tai, J. Agric. Food Chem., 2022, 70, 14613–14621 CrossRef CAS PubMed.
- H. Burger, S. Buttala, H. Koch, M. Ayasse, S. D. Johnson and P. C. Stevenson, J. Exp. Biol., 2024, 227, jeb246156 CrossRef PubMed.
- M. Hoogshagen, A. P. Hastings, J. Chavez, M. Duckett, R. Pettit, A. P. Pahnke, A. A. Agrawal and J. C. de Roode, J. Chem. Ecol., 2024, 50, 52–62 CrossRef CAS.
- A. A. Agrawal, L. Espinosa Del Alba, X. López-Goldar, A. P. Hastings, R. A. White, R. Halitschke, S. Dobler, G. Petschenka and C. Duplais, Proc. Natl. Acad. Sci. U. S. A., 2022, 119, e2205073119 CrossRef CAS PubMed.
- M. Baum and S. Dobler, J. Chem. Ecol., 2024, 50, 63–70 CrossRef CAS PubMed.
- H. R. El-Seedi, S. A. M. Khalifa, E. A. Taher, M. A. Farag, A. Saeed, M. Gamal, M. F. Hegazy, D. Youssef, S. G. Musharraf, M. M. Alajlani, J. Xiao and T. Efferth, Pharmacol. Res., 2019, 141, 123–175 CrossRef CAS.
- Q. Ouyang, Y. X. He, Y. L. Zhang, J. Q. You, M. H. Yu, C. Lei and A. J. Hou, Phytochemistry, 2024, 219, 113990 Search PubMed.
- H. J. Kwon, H. J. Sim, S. I. Lee, Y. M. Lee, Y. D. Park and S. P. Hong, J. Pharm. Biomed. Anal., 2011, 54, 217–221 CrossRef CAS.
- Y. Wang, J. Yuan, Y. Wen, S. Li, L. Tian and Y. Jie, Sci. Rep., 2025, 15, 5747 CrossRef.
- Y. Li, X. Feng and F. Qiu, Chem. Biodiversity, 2024, 21, e202301585 CrossRef CAS.
- L. Chen, P. Jiang, J. Li, Z. Xie, Y. Xu, W. Qu, F. Feng and W. Liu, Phytomedicine, 2019, 57, 72–83 CrossRef CAS.
- J. Cao, D. Yao, R. Li, X. Guo, J. Hao, M. Xie, J. Li, D. Pan, X. Luo, Z. Yu, M. Wang and W. Wang, Neurosci. Bull., 2022, 38, 181–199 CrossRef CAS.
- V. V. Kravtsova, A. A. Fedorova, M. V. Tishkova, A. A. Livanova, O. V. Vetrovoy, A. G. Markov, V. V. Matchkov and I. I. Krivoi, Int. J. Mol. Sci., 2022, 23, 10921 CrossRef CAS.
- A. Olona, C. Hateley, A. Guerrero, J. H. Ko, M. R. Johnson, P. K. Anand, D. Thomas, J. Gil and J. Behmoaras, Br. J. Pharmacol., 2022, 179, 1874–1886 CrossRef CAS PubMed.
- A. Shatrova, E. Burova, N. Pugovkina, A. Domnina, N. Nikolsky and I. Marakhova, Sci. Rep., 2022, 12, 11194 CrossRef CAS PubMed.
- V. A. Barathi, A. Katz, S. Chaudhary, H. L. Li, D. M. Tal, A. Marcovich, C. W. Do and S. J. D. Karlish, Am. J. Physiol.: Cell Physiol., 2024, 326, C1505–C1519 CrossRef.
- R. Zhang, M. Chu, Y. Chen and P. Yan, Reprod. Domest. Anim., 2022, 57, 587–597 CrossRef CAS.
- J. Yue, H. Wu, Y. Yang, H. Song, S. Zhou, X. Li, N. Lin, M. Xie, Y. Xing and L. Chen, Reprod. Sci., 2023, 30, 1594–1607 CrossRef CAS PubMed.
- N. S. Dhalla, V. Elimban and A. D. Adameova, Int. J. Mol. Sci., 2024, 25, 10807 CrossRef CAS PubMed.
- I. Prassas and E. P. Diamandis, Nat. Rev. Drug Discovery, 2008, 7, 926–935 CrossRef CAS.
- Y. Ren, Q. Tan, K. Heath, S. Wu, J. R. Wilson, J. Ren, P. Shriwas, C. Yuan, T. Ngoc Ninh, H. B. Chai, X. Chen, D. D. Soejarto, M. E. Johnson, X. Cheng, J. E. Burdette and A. D. Kinghorn, Bioorg. Med. Chem., 2020, 28, 115301 CrossRef CAS.
- M. Laursen, J. L. Gregersen, L. Yatime, P. Nissen and N. U. Fedosova, Proc. Natl. Acad. Sci. U. S. A., 2015, 112, 1755–1760 CrossRef CAS.
- M. Laursen, L. Yatime, P. Nissen and N. U. Fedosova, Proc. Natl. Acad. Sci. U. S. A., 2013, 110, 10958–10963 CrossRef CAS.
- H. R. Khatri, B. Bhattarai, W. Kaplan, Z. Li, M. J. Curtis Long, Y. Aye and P. Nagorny, J. Am. Chem. Soc., 2019, 141, 4849–4860 CrossRef CAS.
- C. P. Melero, M. Medarde and A. S. Feliciano, Molecules, 2000, 5, 51–81 CrossRef CAS.
- B. Liu, S. Bi, J. Wang, P. Xu and B. Yu, Org. Lett., 2024, 26, 8725–8729 CrossRef CAS PubMed.
- J. Hao, L. Chang, D. Wang, C. Ji, S. Zhang, Y. Hou and Y. Wu, Journal of Cardiovascular Translational Research, 2023, 16, 127–140 CrossRef PubMed.
- M. Zaczkiewicz, K. Kostenzer, M. Graf, B. Mayer, O. Zimmermann and J. Torzewski, J. Clin. Med., 2022, 11, 1762 CrossRef CAS PubMed.
- A. Magadum, H. V. Renikunta, N. Singh, C. Estaras, R. Kishore and F. B. Engel, Front. Cardiovasc. Med., 2022, 9, 901396 CrossRef CAS.
- A. Malik, R. Masson, S. Singh, W. C. Wu, M. Packer, B. Pitt, F. Waagstein, C. J. Morgan, R. M. Allman, G. C. Fonarow and A. Ahmed, J. Am. Coll. Cardiol., 2019, 74, 617–627 CrossRef CAS PubMed.
- K. R. Kumar, A. Flair, E. J. Thompson, K. O. Zimmerman, N. D. Andersen, K. D. Hill and C. P. Hornik, Pediatric Critical Care Medicine, 2022, 23, 453–463 CrossRef.
- B. D. Zhang, W. F. Zhu, T. Akihisa, T. Kikuchi, M. Ukiya, F. Maya, J. Xu, W. Y. Liu, F. Feng and J. Zhang, Phytochemistry, 2021, 181, 112544 CrossRef CAS PubMed.
- M. P. Teixeira, N. F. Haddad, E. F. Passos, M. N. Andrade, M. L. A. Campos, J. M. C. da Silva, C. S. de Figueiredo, E. Giestal-de-Araujo, D. P. de Carvalho, L. Miranda-Alves and L. S. de Paiva, Cancers, 2022, 14, 6168 CrossRef CAS.
- J. Škubník, V. Svobodová Pavlíčková, J. Psotová and S. Rimpelová, Cells, 2021, 10, 3341 CrossRef PubMed.
- S. Zhu, M. Abudupataer, S. Yan, C. Wang, L. Wang and K. Zhu, Biosens. Bioelectron., 2022, 218, 114747 CrossRef CAS PubMed.
- N. Harbeck and M. Gnant, Lancet, 2017, 389, 1134–1150 CrossRef.
- H. Pavithran, R. Kumavath and P. Ghosh, Int. J. Mol. Sci., 2023, 24, 15922 CrossRef CAS.
- F. Acconcia, Int. J. Mol. Sci., 2022, 23, 11102 CrossRef CAS.
- X. Li, J. Zheng, S. Chen, F. D. Meng, J. Ning and S. L. Sun, Cell Death Dis., 2021, 12, 314 CrossRef CAS.
- F. Y. Huang, S. Z. Dai, W. T. Xu, W. Xiong, Y. Sun, Y. H. Huang, J. Y. Wang, Y. Y. Lin, H. Chen, G. H. Tan and W. P. Zheng, Pharmacol. Res., 2023, 187, 106613 CrossRef CAS PubMed.
- P. P. Kumar, D. Smith, J. Key, H. Dong, A. Ganapathysamy, V. Maranda, N. K. Y. Wong, M. L. Fernandez, H. Kim, G. Zhang, C. Ewanowich, L. Hopkins, A. Freywald, L. M. Postovit, M. Köbel, Y. Fu, F. S. Vizeacoumar, F. J. Vizeacoumar, M. S. Carey and C. H. Lee, Gynecol. Oncol., 2024, 188, 162–168 CrossRef CAS PubMed.
- Y. Nagao, A. Yokoi, K. Yoshida, M. Sugiyama, E. Watanabe, K. Nakamura, M. Kitagawa, E. Asano-Inami, Y. Koya, M. Yoshihara, S. Tamauchi, Y. Shimizu, Y. Ikeda, N. Yoshikawa, T. Kato, Y. Yamamoto and H. Kajiyama, Pharmacol. Res., 2023, 189, 106693 CrossRef CAS.
- W. Zhao, G. Li, Q. Zhang, M. Chen, L. He, Z. Wu, Y. Zhang, M. Fan, Y. Liang, W. Zhang, F. Zeng and F. Deng, Biochem. Pharmacol., 2023, 209, 115453 CrossRef CAS PubMed.
- M. Liu, Q. Huang, J. A, L. Li, X. Li, Z. Zhang and J. T. Dong, Cancers, 2021, 13, 5809 CrossRef CAS.
- Y. Zhan, R. Wang, C. Huang, X. Xu, X. Xiao, L. Wu, J. Wei, T. Long and C. Gao, Oncol. Rep., 2024, 52, 103 CrossRef CAS PubMed.
- M. S. Liu, H. Y. Deng, X. L. Mei, W. B. Yuan, W. B. Feng, A. Z. Huang, L. Liu, H. Luo, X. M. Yang and X. S. Li, Fitoterapia, 2024, 177, 8 Search PubMed.
- M. Katoh, T. Fujii, Y. Tabuchi, T. Shimizu and H. Sakai, J. Physiol. Sci., 2024, 74, 23 CrossRef CAS.
- I. Karwaciak, J. Pastwińska, A. Sałkowska, R. A. Bachorz and M. Ratajewski, Arch. Biochem. Biophys., 2024, 759, 110085 CrossRef CAS PubMed.
- Y. Hao, T. Song, M. Wang, T. Li, C. Zhao, T. Li, Y. Hou and H. He, Int. J. Oncol., 2023, 62, 44 CrossRef CAS.
- C. Zhang, H. Y. Yang, L. Gao, M. Z. Bai, W. K. Fu, C. F. Huang, N. N. Mi, H. D. Ma, Y. W. Lu, N. Z. Jiang, L. Tian, T. Cai, Y. Y. Lin, X. X. Zheng, K. Gao, J. J. Chen and W. B. Meng, Front. Pharmacol., 2023, 14, 1098915 CrossRef CAS.
- C. Mayr, T. Kiesslich, D. Bekric, M. Beyreis, M. Kittl, C. Ablinger, E. Neureiter, M. Pichler, F. Prinz, M. Ritter, D. Neureiter, M. Jakab and H. Dobias, PLoS One, 2023, 18, e0287769 CrossRef CAS.
- C. Mi, X. Cao, K. Ma, M. Wei, W. Xu, Y. Lin, J. Zhang and T. Y. Wang, Chem.-Biol. Interact., 2022, 351, 109729 CrossRef CAS PubMed.
- M. Semba, S. Takamatsu, S. Komazawa-Sakon, E. Miyoshi, C. Nishiyama, H. Nakano and K. Moriwaki, Int. J. Mol. Sci., 2022, 23, 6973 CrossRef CAS PubMed.
- C. Boscaro, G. Schimdt, A. Cignarella, L. Dal Maso, C. Bolego and L. Trevisi, Biochem. Pharmacol., 2024, 222, 116049 CrossRef CAS.
- H. Lindholm, K. Ejeskär and F. Szekeres, Int. J. Mol. Sci., 2022, 23, 8237 CrossRef CAS.
- Q. Wu, X. Liu, L. M. Wang, Y. H. Yang, L. F. Pan, J. J. Zhang, Y. Q. Wang, Q. H. Yao, S. L. Ma and S. R. Zhang, Phytother. Res., 2024, 38, 4151–4167 CrossRef CAS.
- M. Tokugawa, Y. Inoue, H. Aoki, C. Miyajima, K. Ishiuchi, K. Tsurumi, C. Kujirai, D. Morishita, M. Matsuno, H. Mizukami, M. Ri, S. Iida, T. Makino, M. Aoyama and H. Hayashi, J. Biochem., 2024, 175, 253–263 CrossRef CAS PubMed.
- M. Tokugawa, Y. Inoue, K. Ishiuchi, C. Kujirai, M. Matsuno, M. Ri, Y. Itoh, C. Miyajima, D. Morishita, N. Ohoka, S. Iida, H. Mizukami, T. Makino and H. Hayashi, Sci. Rep., 2021, 11, 9528 CrossRef CAS PubMed.
- X. Liu, Z. Geng, X. Ding, Y. Lou and X. Zhang, Bioengineered, 2022, 13, 13280–13292 CrossRef CAS PubMed.
- J. Poohadsuan, G. A. O'Doherty, W. Owattanapanich, S. Kungwankiattichai, Y. Rojanasakul, S. Issaragrisil and S. Luanpitpong, Cell Commun. Signaling, 2023, 21, 283 CrossRef CAS.
- S. Takase, Y. S. Yun, F. Moriya, S. Sekine, S. Yotsumoto, K. Miyake, T. Yahagi, A. Ito and H. Inoue, J. Nat. Med., 2023, 77, 180–187 CrossRef CAS PubMed.
- M. Zhou, J. C. Boulos, S. M. Klauck and T. Efferth, Cell Biol. Toxicol., 2023, 39, 2971–2997 CrossRef CAS.
- M. A. Shandell, A. L. Capatina, S. M. Lawrence, W. J. Brackenbury and D. Lagos, J. Biol. Chem., 2022, 298, 101707 CrossRef CAS.
- T. Fujii, M. Katoh, M. Ootsubo, O. T. T. Nguyen, M. Iguchi, T. Shimizu, Y. Tabuchi, Y. Shimizu, H. Takeshima and H. Sakai, J. Cell. Physiol., 2022, 237, 2980–2991 CrossRef CAS PubMed.
- F. Leng and P. Edison, Nat. Rev. Neurol., 2021, 17, 157–172 CrossRef PubMed.
- R. M. Ransohoff, Science, 2016, 353, 777–783 CrossRef CAS PubMed.
- T. Mázala-de-Oliveira, C. S. de Figueiredo, G. de Rezende Corrêa, M. S. da Silva, R. L. Miranda, M. A. de Azevedo, M. Cossenza, A. A. Dos Santos and E. Giestal-de-Araujo, Neurochem. Res., 2022, 47, 723–738 CrossRef PubMed.
- D. Jansson, V. B. Dieriks, J. Rustenhoven, L. C. D. Smyth, E. Scotter, M. Aalderink, S. Feng, R. Johnson, P. Schweder, E. Mee, P. Heppner, C. Turner, M. Curtis, R. Faull and M. Dragunow, Commun. Biol., 2021, 4, 260 CrossRef CAS.
- A. Carlos, J. V. M. Lemos, M. M. F. Borges, M. C. P. Albuquerque, F. B. Sousa, A. Alves, T. S. Dantas and P. G. B. Silva, J. Appl. Oral Sci., 2023, 31, e20230230 CrossRef CAS.
- G. C. R. D. Collaborators, Lancet Respir. Med., 2020, 8, 585–596 CrossRef.
- C. Huang, Y. Wang, X. Li, L. Ren, J. Zhao, Y. Hu, L. Zhang, G. Fan, J. Xu, X. Gu, Z. Cheng, T. Yu, J. Xia, Y. Wei, W. Wu, X. Xie, W. Yin, H. Li, M. Liu, Y. Xiao, H. Gao, L. Guo, J. Xie, G. Wang, R. Jiang, Z. Gao, Q. Jin, J. Wang and B. Cao, Lancet, 2020, 395, 497–506 CrossRef CAS.
- H. Caohuy, O. Eidelman, T. Chen, O. Mungunsukh, Q. Yang, N. I. Walton, B. S. Pollard, S. Khanal, S. Hentschel, C. Florez, A. S. Herbert and H. B. Pollard, Sci. Rep., 2024, 14, 16895 CrossRef.
- J. Galvão, L. H. A. Cavalcante-Silva, É. de Almeida Lima, D. C. M. Carvalho, A. F. Alves and S. R. Mascarenhas, Int. Immunopharmacol., 2022, 109, 108808 CrossRef PubMed.
- Q. Meng, K. Liu, Z. Liu, J. Liu, Z. Tian, S. Qin, J. Wei and L. Cheng, Front. Immunol., 2023, 14, 1251517 CrossRef CAS PubMed.
- G. Misseri, C. Gregoretti and G. Lo Bianco, Jama, 2024, 331, 706–707 CrossRef.
- H. Jia, L. Duan, P. Yu, Y. Zhou, R. Liu and H. Wang, Int. Immunopharmacol., 2022, 111, 109135 CrossRef CAS PubMed.
- K. D. Wang, X. Ding, N. Jiang, C. Zeng, J. Wu, X. Y. Cai, A. Hettinghouse, A. Khleborodova, Z. N. Lei, Z. S. Chen, G. H. Lei and C. J. Liu, Ann. Rheum. Dis., 2022, 81, 544–555 CrossRef CAS.
- X. Zhou, F. Fei, W. Song, H. Ma, Z. Xu, J. Yue, B. Cao, R. Sun, Y. Zhao, Y. Yang, J. Jiang, Y. Geng, Z. Weng and J. Li, Mol. Genet. Genomics, 2023, 298, 567–577 CrossRef CAS PubMed.
- A. G. Markov, A. A. Livanova, A. A. Fedorova, V. V. Kravtsova and I. I. Krivoi, Int. J. Mol. Sci., 2023, 25, 278 CrossRef PubMed.
- Y. Minami, A. Hoshino, Y. Higuchi, M. Hamaguchi, Y. Kaneko, Y. Kirita, S. Taminishi, T. Nishiji, A. Taruno, M. Fukui, Z. Arany and S. Matoba, Nat. Commun., 2023, 14, 4084 CrossRef CAS.
- S. Patel, R. Gururani, S. Jain, N. Tripathi, S. Paliwal, S. Paliwal, S. Paliwal and S. Sharma, Drug Dev. Res., 2022, 83, 1097–1110 CrossRef CAS.
- Y. Xing, J. Y. Wang, M. Y. Li, Z. H. Zhang, H. L. Jin, H. X. Zuo, J. Ma and X. Jin, Br. J. Pharmacol., 2022, 179, 1887–1907 CrossRef CAS.
- D. A. Ferreira, A. B. A. Medeiros, M. M. Soares, É. A. Lima, G. Oliveira, M. Leite, M. V. Machado, J. Villar, L. A. Barbosa, C. Scavone, M. T. Moura and S. Rodrigues-Mascarenhas, Cells, 2024, 13, 1568 CrossRef CAS.
- J. M. de Sales-Neto, D. C. Madruga Carvalho, D. W. Arruda Magalhães, A. B. Araujo Medeiros, M. M. Soares and S. Rodrigues-Mascarenhas, Int. Immunopharmacol., 2024, 127, 111368 CrossRef CAS PubMed.
- J. Škubník, J. Bejček, V. S. Pavlíčková and S. Rimpelová, Molecules, 2021, 26, 5627 CrossRef.
- J. Guo, X. Jia, Y. Liu, S. Wang, J. Cao, B. Zhang, G. Xiao and W. Wang, Antimicrob. Agents Chemother., 2020, 64, e02373–19 CAS.
- L. Amarelle, J. Katzen, M. Shigemura, L. C. Welch, H. Cajigas, C. Peteranderl, D. Celli, S. Herold, E. Lecuona and J. I. Sznajder, Am. J. Physiol.: Lung Cell. Mol. Physiol., 2019, 316, L1094–L1106 CrossRef CAS.
- Z. Y. Liu, W. F. Shi and C. F. Qin, Nat. Rev. Microbiol., 2019, 17, 131–139 CrossRef CAS PubMed.
- J. Guo, X. Jia, Y. Liu, S. Wang, J. Cao, B. Zhang, G. Xiao and W. Wang, Commun. Biol., 2020, 3, 380 CrossRef CAS.
- D. C. M. Carvalho, P. G. da Silva, W. M. Dantas, S. J. R. da Silva, C. T. A. da Silva, E. J. F. Chaves, D. A. M. de Araújo, R. N. de Oliveira, S. Rodrigues-Mascarenhas and L. J. Pena, Sci. Rep., 2022, 12, 12598 CrossRef CAS.
- X. Du, X. Zuo, F. Meng, F. Wu, X. Zhao, C. Li, G. Cheng and F. X. Qin, Biochem. Biophys. Res. Commun., 2020, 522, 862–868 CrossRef CAS PubMed.
- B. Pacheco, A. Fernández-Oliva, M. García-Serradilla and C. Risco, J. Gen. Virol., 2023, 104, 001838 CAS.
- M. Lingemann, T. McCarty, X. Liu, U. J. Buchholz, S. Surman, S. E. Martin, P. L. Collins and S. Munir, PLoS Pathog., 2019, 15, e1007963 CrossRef CAS.
- I. Zerr, A. Ladogana, S. Mead, P. Hermann, G. Forloni and B. S. Appleby, Nature Reviews Disease Primers, 2024, 10, 14 CrossRef PubMed.
- M. Mehrabian, X. Wang, S. Eid, B. Q. Yan, M. Grinberg, M. Siegner, C. Sackmann, M. Sulman, W. Zhao, D. Williams and G. Schmitt-Ulms, PLoS One, 2022, 17, e0270915 CrossRef CAS.
- S. Eid, W. Zhao, D. Williams, Z. Nasser, J. Griffin, P. Nagorny and G. Schmitt-Ulms, PLoS One, 2024, 19, e0308821 CrossRef CAS.
- S. Wu, S. Wang, X. Lin, S. Yang, X. Ba, D. Xiong, L. Xiao and R. Li, Phytomedicine, 2024, 124, 155308 CrossRef CAS PubMed.
- C. W. Yang, H. Y. Hsu, H. Y. Chang, Y. Z. Lee and S. J. Lee, Biochem. Pharmacol., 2020, 180, 114122 CrossRef CAS PubMed.
- S. Biglari, A. S. Moghaddam, M. A. Tabatabaiefar, R. Sherkat, L. Youssefian, A. H. Saeidian, F. Vahidnezhad, L. C. Tsoi, J. E. Gudjonsson, H. Hakonarson, J. L. Casanova, V. Béziat, E. Jouanguy and H. Vahidnezhad, Genet. Med., 2024, 26, 101028 CrossRef CAS PubMed.
- M. Rijsbergen, T. Niemeyer-van der Kolk, G. Hogendoorn, S. Kouwenhoven, C. Lemoine, E. S. Klaassen, M. de Koning, S. Beck, J. N. Bouwes Bavinck, G. Feiss, J. Burggraaf and R. Rissmann, Br. J. Dermatol., 2019, 180, 1058–1068 CrossRef CAS PubMed.
- J. Samolej, I. J. White, B. L. Strang and J. Mercer, J. Gen. Virol., 2024, 105, 001971 CAS.
- K. F. C. Souza E Souza, B. P. T. Moraes, I. C. N. P. Paixão, P. Burth, A. R. Silva and C. F. Gonçalves-de-Albuquerque, Front. Pharmacol., 2021, 12, 624704 CrossRef PubMed.
- J. Cho, Y. J. Lee, J. H. Kim, S. I. Kim, S. S. Kim, B. S. Choi and J. H. Choi, Sci. Rep., 2020, 10, 16200 CrossRef CAS.
- B. S. Pollard, B. L. JC and J. R. Pollard, In Vivo, 2020, 34, 3723–3730 CrossRef CAS PubMed.
- H. Caohuy, O. Eidelman, T. Chen, S. Liu, Q. Yang, A. Bera, N. I. Walton, T. T. Wang and H. B. Pollard, Sci. Rep., 2021, 11, 22195 CrossRef CAS.
- K. S. Plante, V. Dwivedi, J. A. Plante, D. Fernandez, D. Mirchandani, N. Bopp, P. V. Aguilar, J. G. Park, P. P. Tamayo, J. Delgado, V. Shivanna, J. B. Torrelles, L. Martinez-Sobrido, R. Matos, S. C. Weaver, K. J. Sastry and R. A. Newman, Biomed. Pharmacother., 2021, 138, 111457 CrossRef CAS.
- D. P. Ha, W. J. Shin, Z. Liu, M. E. Doche, R. Lau, N. M. Leli, C. S. Conn, M. Russo, A. Lorenzato, C. Koumenis, M. Yu, S. M. Mumenthaler and A. S. Lee, Cell Biosci., 2024, 14, 115 CrossRef CAS.
- L. G. Hilgenberg, H. Su, H. Gu, D. K. O'Dowd and M. A. Smith, Cell, 2006, 125, 359–369 CrossRef CAS.
- L. Desfrere, M. Karlsson, H. Hiyoshi, S. Malmersjö, E. Nanou, M. Estrada, A. Miyakawa, H. Lagercrantz, A. El Manira, M. Lal and P. Uhlén, Proc. Natl. Acad. Sci. U. S. A., 2009, 106, 2212–2217 CrossRef CAS.
- P. F. Kinoshita, A. M. M. Orellana, V. W. Nakao, N. M. de Souza Port's, L. E. M. Quintas, E. M. Kawamoto and C. Scavone, Br. J. Pharmacol., 2022, 179, 1512–1524 CrossRef CAS.
- H. E. Titus, H. Xu, A. P. Robinson, P. A. Patel, Y. Chen, D. Fantini, V. Eaton, M. Karl, E. D. Garrison, I. V. L. Rose, M. Y. Chiang, J. R. Podojil, R. Balabanov, S. A. Liddelow, R. H. Miller, B. Popko and S. D. Miller, Glia, 2022, 70, 1950–1970 CrossRef CAS.
- G. Gurkan, M. A. Erdogan, G. Karahan, H. Kaya, G. Yigitturk, C. Kizmazoglu and O. Erbas, Turkish Neurosurgery, 2023, 33, 258–264 Search PubMed.
- V. Tsyvunin, S. Shtrygol, M. Mishchenko, D. Lytkin, A. Taran, D. Shtrygol and T. Gorbach, Ceska Slov. Farm., 2023, 72, 172–183 CrossRef CAS.
- J. Hashimoto, E. Fujita, K. Tanimoto, S. Kondo and K. Matsumoto-Miyai, Neuroscience, 2024, 541, 77–90 CrossRef CAS.
- M. A. Erdogan, M. Kirazlar, G. Yigitturk and O. Erbas, Neurochem. Res., 2022, 47, 1290–1298 CrossRef CAS PubMed.
- D. A. Sibarov, Z. D. Zhuravleva, M. A. Ilina, S. I. Boikov, Y. D. Stepanenko, T. V. Karelina and S. M. Antonov, Cells, 2023, 12, 2011 CrossRef CAS PubMed.
- L. D. Nguyen, Z. Wei, M. C. Silva, S. Barberán-Soler, J. Zhang, R. Rabinovsky, C. R. Muratore, J. M. S. Stricker, C. Hortman, T. L. Young-Pearse, S. J. Haggarty and A. M. Krichevsky, Nat. Commun., 2023, 14, 7575 CrossRef CAS.
- A. Papandreou, N. Singh, L. Gianfrancesco, D. Budinger, K. Barwick, A. Agrotis, C. Luft, Y. Shao, A. S. Lenaerts, A. Gregory, S. Y. Jeong, P. Hogarth, S. Hayflick, S. Barral, J. Kriston-Vizi, P. Gissen, M. A. Kurian and R. Ketteler, bioRxiv, 2024, preprint, DOI:10.1101/2023.09.13.556416.
- R. D. Lopes, R. Rordorf, G. M. De Ferrari, S. Leonardi, L. Thomas, D. M. Wojdyla, P. Ridefelt, J. H. Lawrence, R. De Caterina, D. Vinereanu, M. Hanna, G. Flaker, S. M. Al-Khatib, S. H. Hohnloser, J. H. Alexander, C. B. Granger and L. Wallentin, J. Am. Coll. Cardiol., 2018, 71, 1063–1074 CrossRef CAS.
- B. W. Keene, C. E. Atkins, J. D. Bonagura, P. R. Fox, J. Häggström, V. L. Fuentes, M. A. Oyama, J. E. Rush, R. Stepien and M. Uechi, J. Vet. Intern. Med., 2019, 33, 1127–1140 CrossRef PubMed.
- A. F. M. Botelho, F. Pierezan, B. Soto-Blanco and M. M. Melo, Toxicon, 2019, 158, 63–68 CrossRef CAS PubMed.
- W. Zhuang, T. He, B. B. Jia, Z. Z. Wang, L. Zhang, X. Z. Dong and S. Y. Xi, Front. Pharmacol., 2023, 14, 1040778 CrossRef CAS PubMed.
- J. Patocka, E. Nepovimova, W. Wu and K. Kuca, Environ. Toxicol. Pharmacol., 2020, 79, 103400 CrossRef CAS PubMed.
- O. J. Ziff and D. Kotecha, Trends Cardiovasc. Med., 2016, 26, 585–595 CrossRef CAS.
- M. Michalak, K. Michalak and J. Wicha, Nat. Prod. Rep., 2017, 34, 361–410 RSC.
- H. Renata, Q. Zhou and P. S. Baran, Science, 2013, 339, 59–63 CrossRef CAS PubMed.
- H. Renata, Q. Zhou, G. Dünstl, J. Felding, R. R. Merchant, C. H. Yeh and P. S. Baran, J. Am. Chem. Soc., 2015, 137, 1330–1340 CrossRef CAS PubMed.
- M. Honma and M. Nakada, Tetrahedron Lett., 2007, 48, 1541–1544 CrossRef CAS.
- K. Mukai, S. Kasuya, Y. Nakagawa, D. Urabe and M. Inoue, Chem. Sci., 2015, 6, 3383–3387 RSC.
- M. E. Jung and D. Yoo, Org. Lett., 2011, 13, 2698–2701 CrossRef CAS.
- M. E. Jung and H. V. Chu, Org. Lett., 2008, 10, 3647–3649 CrossRef CAS.
- J. Sun, Y. Chen, S. S. Ragab, W. Gu, Z. Tang, Y. Tang and W. Tang, Angew Chem. Int. Ed. Engl., 2023, 62, e202303639 CrossRef CAS PubMed.
- C. Tichvon, E. Zviagin, Z. Surma and P. Nagorny, Org. Lett., 2024, 26, 2445–2450 CrossRef CAS.
- Z. Fejedelem, N. Carney and P. Nagorny, J. Org. Chem., 2021, 86, 10249–10262 CrossRef CAS.
- N. Carney, N. Perry, J. Garabedian and P. Nagorny, Org. Lett., 2023, 25, 966–971 CrossRef CAS PubMed.
- X. Zhang, E. King-Smith, L. B. Dong, L. C. Yang, J. D. Rudolf, B. Shen and H. Renata, Science, 2020, 369, 799–806 CrossRef CAS PubMed.
- J. D. Wang, Q. Xie, X. M. Wang, M. F. Long, Y. Y. Chen, Z. Liu, M. Xia, J. Guo, Z. P. Wang, R. F. Wang, S. Y. Shen, Y. Lu, Y. Yin, Y. T. Hu, W. Gao, X. Zhang, P. Su and L. Q. Huang, ACS Catal., 2025, 15, 2690–2702 CrossRef CAS.
- S. Liu, S. Yang and P. Su, Science of Traditional Chinese Medicine, 2024, 2, 95–103 CrossRef.
- Y. Zhao, B. Zhang, Z. Q. Sun, H. Zhang, W. Wang, Z. R. Wang, Z. K. Guo, S. Yu, R. X. Tan and H. M. Ge, ACS Catal., 2022, 12, 9839–9845 CrossRef CAS.
- F. Song, M. Zheng, J. Wang, H. Liu, Z. Lin, B. Liu, Z. Deng, H. Cong, Q. Zhou and X. Qu, Nat. Synth., 2023, 2, 729–739 CrossRef CAS.
- J. Feng, Q. Wu, D. Zhu and Y. Ma, Chemsuschem, 2022, 15, e202102399 CrossRef CAS.
- J. Zhao, S. H. Guan, X. B. Chen, W. Wang, M. Ye and D. A. Guo, Chin. Chem. Lett., 2007, 18, 1316–1318 CrossRef CAS.
- E. Leete, H. Gregory and E. G. Gros, J. Am. Chem. Soc., 1965, 87, 3475–3479 CrossRef CAS.
- H. Zhao, Q. Tang, C. Mo, L. Bai, D. Tu and X. Ma, Acta Pharm. Sin. B, 2017, 7, 215–222 CrossRef.
- P. D. Sonawane, J. Pollier, S. Panda, J. Szymanski, H. Massalha, M. Yona, T. Unger, S. Malitsky, P. Arendt, L. Pauwels, E. Almekias-Siegl, I. Rogachev, S. Meir, P. D. Cárdenas, A. Masri, M. Petrikov, H. Schaller, A. A. Schaffer, A. Kamble, A. P. Giri, A. Goossens and A. Aharoni, Nat. Plants, 2016, 3, 16205 CrossRef.
- A. C. Diener, H. Li, W. Zhou, W. J. Whoriskey, W. D. Nes and G. R. Fink, Plant Cell, 2000, 12, 853–870 CrossRef CAS.
- N. Holmberg, M. Harker, C. L. Gibbard, A. D. Wallace, J. C. Clayton, S. Rawlins, A. Hellyer and R. Safford, Plant Physiol., 2002, 130, 303–311 CrossRef CAS PubMed.
- A. Rahier, Steroids, 2011, 76, 340–352 CrossRef CAS PubMed.
- S. Sawai, K. Ohyama, S. Yasumoto, H. Seki, T. Sakuma, T. Yamamoto, Y. Takebayashi, M. Kojima, H. Sakakibara, T. Aoki, T. Muranaka, K. Saito and N. Umemoto, Plant Cell, 2014, 26, 3763–3774 CrossRef CAS PubMed.
- P. D. Cárdenas, P. D. Sonawane, J. Pollier, R. Vanden Bossche, V. Dewangan, E. Weithorn, L. Tal, S. Meir, I. Rogachev, S. Malitsky, A. P. Giri, A. Goossens, S. Burdman and A. Aharoni, Nat. Commun., 2016, 7, 10654 CrossRef PubMed.
- I. Raghavan, B. Ravi Gopal, E. Carroll and Z. Q. Wang, Plant Cell Physiol., 2023, 64, 107–116 CrossRef CAS.
- A. Rahier and F. Karst, Biochem. J., 2014, 459, 289–299 CrossRef CAS PubMed.
- M. Kushiro, T. Nakano, K. Sato, K. Yamagishi, T. Asami, A. Nakano, S. Takatsuto, S. Fujioka, Y. Ebizuka and S. Yoshida, Biochem. Biophys. Res. Commun., 2001, 285, 98–104 CrossRef CAS.
- K. Schrick, U. Mayer, A. Horrichs, C. Kuhnt, C. Bellini, J. Dangl, J. Schmidt and G. Jürgens, Genes Dev., 2000, 14, 1471–1484 CrossRef CAS.
- M. Souter, J. Topping, M. Pullen, J. Friml, K. Palme, R. Hackett, D. Grierson and K. Lindsey, Plant Cell, 2002, 14, 1017–1031 CrossRef CAS.
- J. Wang, J. Liu, Z. Song and W. D. Nes, Bioorg. Med. Chem. Lett., 2008, 18, 232–235 CrossRef CAS PubMed.
- S. Darnet and A. Rahier, Biochem. J., 2004, 378, 889–898 CrossRef CAS.
- S. Choe, T. Noguchi, S. Fujioka, S. Takatsuto, C. P. Tissier, B. D. Gregory, A. S. Ross, A. Tanaka, S. Yoshida, F. E. Tax and K. A. Feldmann, Plant Cell, 1999, 11, 207–221 CAS.
- D. Silvestro, T. G. Andersen, H. Schaller and P. E. Jensen, PLoS One, 2013, 8, e56429 CrossRef CAS.
- M. Zhang, C. Wang, Q. Lin, A. Liu, T. Wang, X. Feng, J. Liu, H. Han, Y. Ma, D. Bonea, R. Zhao and X. Hua, Plant J., 2015, 83, 582–599 CrossRef CAS.
- P. Lindemann and M. Luckner, Phytochemistry, 1997, 46, 507–513 CrossRef CAS.
- I. C. Guo, M. C. Hu and B. C. Chung, J. Biomed. Sci., 2003, 10, 593–598 CAS.
- N. Strushkevich, F. MacKenzie, T. Cherkesova, I. Grabovec, S. Usanov and H. W. Park, Proc. Natl. Acad. Sci. U. S. A., 2011, 108, 10139–10143 CrossRef CAS PubMed.
- G. Li, T. An, Y. Li, J. Yue, R. Huang, J. Huang, J. Liang, W. Yao, L. Huang, Y. Chen, R. Zhang, A. Ji and L. Duan, Front. Genet., 2022, 13, 828877 CrossRef CAS PubMed.
- E. Carroll, B. Ravi Gopal, I. Raghavan, M. Mukherjee and Z. Q. Wang, Nat. Commun., 2023, 14, 4042 CrossRef CAS PubMed.
- M. Kunert, C. Langley, R. Lucier, K. Ploss, C. E. Rodríguez López, D. A. Serna Guerrero, E. Rothe, S. E. O'Connor and P. D. Sonawane, Nat. Plants, 2023, 9, 1607–1617 CrossRef CAS PubMed.
- R. Li, S. Guo, D. Wang, T. Yang, W. Li, J. Wang, L. Huang, X. Zhang and Z. Dai, Metab. Eng., 2025, 90, 197–208 CrossRef CAS PubMed.
- G. C. Younkin, M. L. Alani, A. Páez-Capador, H. D. Fischer, M. Mirzaei, A. P. Hastings, A. A. Agrawal and G. Jander, New Phytol., 2024, 242, 2719–2733 CrossRef CAS PubMed.
- P. Hartz, S. J. Strohmaier, B. M. El-Gayar, A. Abdulmughni, M. C. Hutter, F. Hannemann, E. M. J. Gillam and R. Bernhardt, FEBS J., 2021, 288, 6510–6527 CrossRef CAS PubMed.
- Y. Han, M. Xia, Y. F. Zhang, P. Su and L. Q. Huang, Zhongguo Zhongyao Zazhi, 2024, 49, 370–378 CAS.
- J. Munkert, M. Ernst, F. Müller-Uri and W. Kreis, Phytochemistry, 2014, 100, 26–33 CrossRef CAS PubMed.
- M. Sameeullah, M. Yildirim, N. Aslam, M. C. Baloğlu, B. Yucesan, A. G. Lössl, K. Saba, M. T. Wah eed and E. Gurel, Int. J. Mol. Sci., 2021, 22, 11736 CrossRef CAS PubMed.
- P. Pérez-Bermúdez, A. A. García, I. Tuñón and I. Gavidia, New Phytol., 2010, 185, 687–700 CrossRef.
- J. Klein, E. Horn, M. Ernst, T. Leykauf, T. Leupold, M. Dorfner, L. Wolf, A. Ignatova, W. Kreis and J. Munkert, Plant Cell Rep., 2021, 40, 1631–1646 CrossRef CAS PubMed.
- L. Martinelli, C. Bihanic, A. Bony, F. Gros, C. Conart, S. Fiorucci, H. Casabianca, F. Schiets, G. Chietera, B. Boachon, B. Blerot, S. Baudino, F. Jullien and D. Saint-Marcoux, Plant Physiol., 2024, 194, 1006–1023 CrossRef CAS PubMed.
- K. Schmidt, J. Petersen, J. Munkert, C. Egerer-Sieber, M. Hornig, Y. A. Muller and W. Kreis, Phytochemistry, 2018, 156, 9–19 CrossRef CAS.
- G. Shiko, M. J. Paulmann, F. Feistel, M. Ntefidou, V. Hermann-Ene, W. Vetter, B. Kost, G. Kunert, J. A. Z. Zedler, M. Reichelt, R. Oelmüller and J. Klein, New Phytol., 2023, 240, 318–337 CrossRef CAS.
- U. Stuhlemmer, W. Haussmann, F. Milek, W. Kreis and E. Reinhard, Z. Naturforsch., C:J. Biosci., 1993, 48, 713–721 CrossRef CAS PubMed.
- Z. Z. Xu, Z. Zhuang, R. Cai, G. Q. Lin, Z. She, Q. Zhao and Q. L. He, Angew Chem. Int. Ed. Engl., 2023, 62, e202216989 CrossRef CAS PubMed.
- C. Rieck, D. Geiger, J. Munkert, K. Messerschmidt, J. Petersen, J. Strasser, N. Meitinger and W. Kreis, Microbiologyopen, 2019, 8, e925 CrossRef.
- M. Tropper, S. Höhn, L. S. Wolf, J. Fritsch, N. Kastner-Detter, C. Rieck, J. Munkert, N. Meitinger, H. Lanig and W. Kreis, Phytochemistry, 2021, 187, 112710 CrossRef CAS.
- R. M. Pádua, N. Meitinger, M. Hennemann, P. Schebitz, R. Waibel, S. Löber, P. Gmeiner, T. Clark and W. Kreis, Tetrahedron, 2016, 72, 4556–4563 CrossRef.
- M. D. Siperstein, A. W. Murray and E. Titus, Arch. Biochem. Biophys., 1957, 67, 154–160 CrossRef CAS PubMed.
- R. I. Dmitrieva, A. Y. Bagrov, E. Lalli, P. Sassone-Corsi, D. M. Stocco and P. A. Doris, Hypertension, 2000, 36, 442–448 CrossRef CAS.
- C. Chen and M. V. Osuch, Biochem. Pharmacol., 1969, 18, 1797–1802 CrossRef CAS.
- A. M. Porto and E. G. Gros, Experientia, 1970, 26, 11 CrossRef CAS PubMed.
- S. Liu, S. Yang, B. Blazekovic, L. Li, J. Zhang and Y. Wang, Engineering, 2024, 38, 13–26 CrossRef.
- L. Zheng, Y. Grigorova, K. Ramirez, V. Zernetikina, W. Wei, N. Petrashevskaya, O. Juhasz, R. Fenner, E. Lakatta, A. Bagrov and O. Federova, FASEB J., 2019, 33, lb331 Search PubMed.
- D. Xu, M. Wu, X. Li, M. Xia, D. Liu, Y. Dai, Q. Yu, B. Wu and D. Wang, Int. J. Biol. Macromol., 2018, 120, 673–682 CrossRef CAS.
- Y. Zhang, X. Li, D. Xu, M. Wu, Y. Dai, M. Xia and D. Wang, Int. J. Biol. Macromol., 2021, 175, 67–78 CrossRef CAS.
- X. Lei, X. Wang, W. Xiong, H. Xiao, Y. Wu, T. Huang, R. Liang, Y. Li and S. Lin, ACS Chem. Biol., 2024, 19, 1169–1179 CrossRef CAS PubMed.
- J. Wang, Y. Zhang, H. Liu, Y. Shang, L. Zhou, P. Wei, W. B. Yin, Z. Deng, X. Qu and Q. Zhou, Nat. Commun., 2019, 10, 3378 CrossRef PubMed.
- T. M. Beale and M. S. Taylor, Org. Lett., 2013, 15, 1358–1361 CrossRef CAS PubMed.
- R. W. Gantt, P. Peltier-Pain and J. S. Thorson, Nat. Prod. Rep., 2011, 28, 1811–1853 RSC.
- P. Ullmann, A. Ury, D. Rimmele, P. Benveniste and P. Bouvier-Navé, Biochimie, 1993, 75, 713–723 CrossRef CAS PubMed.
- C. Wen, W. Huang, X. L. Zhu, X. S. Li, F. Zhang and R. W. Jiang, Org. Lett., 2018, 20, 534–537 CrossRef CAS.
- N. H. Thuan, T. Yamaguchi, J. H. Lee and J. K. Sohng, Enzyme Microb. Technol., 2013, 52, 234–240 CrossRef CAS.
- N. H. Hoang, S. Y. Hong, N. L. Huong and J. W. Park, J. Microbiol. Biotechnol., 2016, 26, 477–482 CrossRef.
- L. K. Sharma, B. R. Madina, P. Chaturvedi, R. S. Sangwan and R. Tuli, Arch. Biochem. Biophys., 2007, 460, 48–55 CrossRef CAS PubMed.
- M. Zhou, Y. Hou, A. Hamza, C. Pain, C. G. Zhan, T. S. Bugni and J. S. Thorson, Org. Lett., 2012, 14, 5424–5427 CrossRef CAS PubMed.
- K. Li, J. Feng, Y. Kuang, W. Song, M. Zhang, S. Ji, X. Qiao and M. Ye, Adv. Synth. Catal., 2017, 359, 3765–3772 CrossRef CAS.
- C. Wen, W. Huang, M. M. He, W. L. Deng and H. H. Yu, Biotechnol. Lett., 2020, 42, 135–142 CrossRef CAS PubMed.
- W. Huang, Y. He, R. W. Jiang, Z. X. Deng and F. Long, ACS Catal., 2022, 12, 2927–2937 CrossRef CAS.
- X. Xu, Y. Liu, G. Du, R. Ledesma-Amaro and L. Liu, Trends Biotechnol., 2020, 38, 779–796 CrossRef CAS PubMed.
- H. X. Du, W. H. Xiao, Y. Wang, X. Zhou, Y. Zhang, D. Liu and Y. J. Yuan, PLoS One, 2016, 11, e0146773 CrossRef.
- Y. Zhang, Y. Wang, M. Yao, H. Liu, X. Zhou, W. Xiao and Y. Yuan, Biotechnol. Lett., 2017, 39, 1033–1039 CrossRef CAS.
- S. Xu, C. Chen and Y. Li, ACS Synth. Biol., 2020, 9, 3157–3170 CrossRef CAS.
- S. Xu, X. Teng and Y. Li, ACS Synth. Biol., 2023, 12, 1109–1118 CrossRef CAS.
- Y. D. Qian, S. Y. Tan, G. R. Dong, Y. J. Niu, C. Y. Hu and Y. H. Meng, Appl. Microbiol. Biotechnol., 2020, 104, 7165–7175 CrossRef CAS.
- W. Zhou, H. Gao, Y. Wu, X. Zhang, M. Xu, T. Yang, M. Shao and Z. Rao, J. Ind. Eng. Chem., 2021, 72, 4314–4324 CAS.
- L. Gu, R. Zhang, X. Fan, Y. Wang, K. Ma, J. Jiang, G. Li, H. Wang, F. Fan and X. Zhang, ACS Synth. Biol., 2023, 12, 2947–2960 CrossRef CAS PubMed.
- L. Xu, D. Wang, J. Chen, B. Li, Q. Li, P. Liu, Y. Qin, Z. Dai, F. Fan and X. Zhang, Metab. Eng., 2022, 70, 115–128 CrossRef CAS PubMed.
- C. Duport, R. Spagnoli, E. Degryse and D. Pompon, Nat. Biotechnol., 1998, 16, 186–189 CrossRef CAS PubMed.
- W. L. Miller and R. J. Auchus, Endocr. Rev., 2011, 32, 81–151 CrossRef CAS.
- D. S. Makeeva, D. V. Dovbnya, M. V. Donova and L. A. Novikova, Am. J. Mol. Biol., 2013, 3, 173–182 CrossRef.
- R. Zhang, Y. Zhang, Y. Wang, M. Yao, J. Zhang, H. Liu, X. Zhou, W. Xiao and Y. Yuan, ACS Synth. Biol., 2019, 8, 2666–2678 CrossRef CAS PubMed.
- Q. Chen, W. Wei, Z. Chao, R. Qi, J. He, H. Chen, K. Wang, X. Wang, Y. Rao and J. Zhou, Nat. Commun., 2025, 16, 3740 CrossRef CAS PubMed.
- N. Strizhov, V. Fokina, G. Sukhodolskaya, D. Dovbnya, M. Karpov, A. Shutov, L. Novikova and M. Donova, New Biotechnol., 2014, 31, S67 CrossRef.
- K. Liu, F. Q. Wang, K. Liu, Y. Zhao, B. Gao, X. Tao and D. Wei, Bioresources and Bioprocessing, 2022, 9, 93 CrossRef PubMed.
- M. Karpov, N. Strizhov, L. Novikova, T. Lobastova, S. Khomutov, A. Shutov, A. Kazantsev and M. Donova, Microb. Cell Fact., 2024, 23, 105 CrossRef CAS PubMed.
- J. G. Patil, M. L. Ahire, K. M. Nitnaware, S. Panda, V. P. Bhatt, P. B. Kishor and T. D. Nikam, Appl. Microbiol. Biotechnol., 2013, 97, 2379–2393 CrossRef CAS PubMed.
- H. Zheng, Am. J. Med., 2022, 135, 670–672 CrossRef CAS.
- F. Ferrari, I. Santander and R. Stein, Curr. Cardiol. Rev., 2020, 16, 141–146 CrossRef PubMed.
- A. F. M. Botelho, A. L. S. Miranda, T. G. Freitas, P. F. Milani, T. Barreto, J. S. Cruz and M. M. Melo, Cardiovasc. Toxicol., 2020, 20, 539–547 CrossRef CAS PubMed.
- J. Zhang, L. V. Ponomareva, N. S. Nandurkar, Y. Yuan, L. Fang, C. G. Zhan and J. S. Thorson, ACS Med. Chem. Lett., 2015, 6, 1053–1058 CrossRef CAS PubMed.
- X. S. Li, Y. C. Ren, Y. Z. Bao, J. Liu, X. K. Zhang, Y. W. Zhang, X. L. Sun, X. S. Yao and J. S. Tang, Eur. J. Med. Chem., 2018, 145, 252–262 CrossRef CAS PubMed.
- J. M. Langenhan, N. R. Peters, I. A. Guzei, F. M. Hoffmann and J. S. Thorson, Proc. Natl. Acad. Sci. U. S. A., 2005, 102, 12305–12310 CrossRef CAS.
- Y. J. Yan, J. Yang, L. Wang, D. D. Xu, Z. Y. Yu, X. W. Guo, G. P. Horsman, S. J. Lin, M. F. Tao and S. X. Huang, Chem. Sci., 2020, 11, 3959–3964 RSC.
- R. Buller, S. Lutz, R. J. Kazlauskas, R. Snajdrova, J. C. Moore and U. T. Bornscheuer, Science, 2023, 382, eadh8615 CrossRef CAS PubMed.
- J. Abramson, J. Adler, J. Dunger, R. Evans, T. Green, A. Pritzel, O. Ronneberger, L. Willmore, A. J. Ballard, J. Bambrick, S. W. Bodenstein, D. A. Evans, C. C. Hung, M. O'Neill, D. Reiman, K. Tunyasuvunakool, Z. Wu, A. Žemgulytė, E. Arvaniti, C. Beattie, O. Bertolli, A. Bridgland, A. Cherepanov, M. Congreve, A. I. Cowen-Rivers, A. Cowie, M. Figurnov, F. B. Fuchs, H. Gladman, R. Jain, Y. A. Khan, C. M. R. Low, K. Perlin, A. Potapenko, P. Savy, S. Singh, A. Stecula, A. Thillaisundaram, C. Tong, S. Yakneen, E. D. Zhong, M. Zielinski, A. Žídek, V. Bapst, P. Kohli, M. Jaderberg, D. Hassabis and J. M. Jumper, Nature, 2024, 630, 493–500 CrossRef CAS PubMed.
- N. E. Chayen and E. Saridakis, Nat. Methods, 2008, 5, 147–153 CrossRef CAS.
| This journal is © The Royal Society of Chemistry 2026 |