Open Access Article
Open Access ArticleDiscovery of semisynthetic derivatives of (R)- and (S)-usnic acids as potential antifungal agents against C. tropicalis and T. rubrum†
Anna
Fontana‡
a,
Alessio
Colleoni§‡
 a,
Roberta
Listro
a,
Giacomo
Rossino
a,
Roberta
Listro
a,
Giacomo
Rossino
 a,
Pasquale
Linciano
a,
Barbara
Vigani
a,
Caterina
Valentino
a,
Valeria
Cavalloro
a,
Pasquale
Linciano
a,
Barbara
Vigani
a,
Caterina
Valentino
a,
Valeria
Cavalloro
 bc,
Marta Elisabetta Eleonora
Temporiti
bc,
Solveig
Tosi
bc,
Emanuela
Martino
bc,
Marta Elisabetta Eleonora
Temporiti
bc,
Solveig
Tosi
bc,
Emanuela
Martino
 bc and
Simona
Collina
bc and
Simona
Collina
 *a
*a
aDepartment of Drug Sciences, University of Pavia, viale Taramelli 12, 27100 Pavia, Italy. E-mail: simona.collina@unipv.it
bDepartment of Earth and Environmental Sciences, University of Pavia, via Sant'Epifanio 14, 27100 Pavia, Italy
cNational Biodiversity Future Center, Piazza Marina 61, Palermo, 90133, Italy
First published on 10th July 2025
Abstract
The prevalence of human fungal infections (FIs) is rapidly increasing worldwide, posing substantial challenges to public health. The underestimation of FIs risk led to a limited knowledge of the fungal pathogenicity and a concomitant paucity of antimycotic drugs that are increasingly unable to effectively address resistance liabilities. The identification of innovative antifungal drugs is therefore an urgent need. Natural products have always been under scrutiny in the drug discovery process. Of these, usnic acid (UA) represents a compelling starting point for antifungal drug development due to its natural occurrence as a secondary metabolite in various lichen species, where it serves as a natural defence mechanism against fungal invasion. This dibenzofuran derivative possesses an intrinsically rigid three-dimensional architecture with stereogenic center, providing a pre-organized chiral scaffold with potential for selective interaction with fungal targets. Despite its high therapeutic potential as antimicrobial agent, UA suffers from poor solubility and hepatotoxicity issues. The proposed research explores the modification of UA scaffold to generate the series of semisynthetic compounds 1–9 by derivatizing the (R)- and (S)-UA as enamines. Considering the inherent chirality of UA, this work aims to identify structure–activity relationships that optimize antifungal efficacy while improving the pharmacokinetic properties of UA. The resulting compounds were evaluated for their antifungal activity against three strains, showing significant differences in potency concerning their absolute configuration. This research addresses the urgent need for novel antifungal agents in an era of increasing resistance to conventional treatments, identifying (9bS,15S)-1, 3, 4, and 8 compounds as promising compounds for developing antifungal therapeutics.
1. Introduction
Fungal infections (FIs) or mycoses have historically received less attention than bacterial and viral infections both from academia and pharmaceutical companies, due to their relatively low incidence and mortality rates in developed countries. Nonetheless, fungal infections account for 300 million cases per year, resulting in 1.5 million deaths, most of which occur in neglected populations.1–5In the last years, FIs, particularly among hospitalized and immunocompromised patients, have raised several public health concerns, prompting the World Health Organization to release a priority list of the main fungal threats in 2022.6,7 This formal report emphasised the need for an unprecedented research effort to thoroughly understand the mechanisms underpinning fungal virulence and, consequently, to expand the limited arsenal of available antifungal drugs.7 Notably, recommended therapy for the treatment of FIs has remained largely unchanged over the last decade, accounting for the use of broad-spectrum antimycotics such as amphotericin B (AmB) and azoles (e.g. fluconazole, FCZ) as mono- or combination therapy.8,9 Multi-drug approaches are regularly employed to counteract the increasing drug resistance acquired by several fungal strains.10,11
Among pathogenic fungi, Candida spp. (e.g. Candida albicans, Candida tropicalis and Candida auris) represent the most common etiological agents of mucocutaneous and invasive FIs, while Trichophyton spp. (e.g. Trichophyton rubrum and Trichophyton mentagrophytes) is responsible for up to 50% of dermatophytosis, superficial FIs occurring on keratinous substrates.12–14 The increasing use of antifungal drugs for cutaneous mycosis treatment, and the long-term therapeutic regimens required to eradicate the vast majority of recurrent FIs, have led to the emergence of drug-resistant fungal strains especially against AmB and azoles.11,15
Given the magnitude of this global health issue, launching new drug discovery programs is essential for identifying new chemical entities capable of tackling, or at least limiting, the spread of FIs.16
In medicinal chemistry campaigns, natural products are extensively investigated due to their intrinsic properties and distinctive structural complexity, often leading to the discovery of biologically active metabolites and inspiring the design of new compounds.17–19 Although still poorly studied, lichens have increasingly emerged as rich sources of secondary metabolites with multiple pharmacological activities, as evidenced by their use in traditional medicine.20–22 Among these, usnic acid (UA) is one of the best-characterized, being abundantly biosynthesised by numerous lichen species.23 Due to the presence of a stereogenic centre, UA can occur in nature in both enantiopure and racemic forms.24,25 Specifically, (+)-(R)-UA is the most abundant in several genera including Ramalina and Usnea. Conversely, (−)-(S)-UA is found in small amounts in a few species of Cladonia and Alectoria. This explains the low commercial availability and the high retail cost of the (S)-enantiomer which has limited its investigation in a therapeutic perspective.24,26
UA exhibits a broad spectrum of biological assets, including antineoplastic, anti-inflammatory and antimicrobial activities.23,27–29 Extensive literature has examined the anti-infective properties of UA, but only a limited number of studies have investigated and demonstrated its significant antimycotic activity, primarily focusing on the (R)-enantiomer.25,30–34
Despite its pharmacological potentiality, UA has faced significant challenge in therapeutic development. Documented cases of hepatotoxicity prompted regulatory intervention, with FDA ordering the withdrawal of UA-containing products from the market due to safety concerns.35,36 Further, UA suffers from poor water solubility, strong binding affinity for serum proteins, and low bioavailability.37–39 These pharmaceutical limitations restrict its use for both systemic and topical treatment.40–42 To improve the pharmacokinetic properties of UA, different chemical modifications on the scaffold have been explored, leading to different classes of biologically active compounds (e.g. enamine, benzofuran-2-one, heteroaryl, and pyrazole derivatives).26,32–34,43,44
Building on these findings, herein we prepared a series of semisynthetic UA-based enamines (compounds 1–9, Table 1) as potential agents against FIs. We also investigated whether the configuration of the stereogenic center in UA affects antifungal potency and how systematic modifications of each UA enantiomer can further modulate the antifungal activity.26 Indeed, the importance of chirality in the drug discovery process is well known, and the field of antifungal medicines is no exception.45 Many antifungals are chiral, ranging from simple synthetic azoles to more intricate natural and semi-synthetic scaffolds (e.g. amphotericin B, echinocandins, anidulafungin).46–48 To evaluate the impact of the absolute configuration of UA scaffold on its antifungal activity, both the commercially available (R)-UA, and the (S)-UA isolated from Cladonia foliacea were properly functionalized, generating compounds 1–9. These latter were subsequently tested against three fungal strains, Candida albicans, Candida tropicalis, and Trichophyton rubrum to evaluate their antimycotic and fungicidal activity. Moreover, to assess the safety, the cytotoxicity on human dermal fibroblasts was assessed.
| Cpds | MIC99 (μM) | MIC99 (μM) |
|---|---|---|
| C. tropicalis | T. rubrum | |
| a Experiments were performed in triplicate. | ||
| AmB | >400 | >400 |
| FCZ | >200 | >200 |
| (R)-UA | 17.4 | 580 |
| (S)-UA | 4.54 | 580 |
| (9bR,15S)-1 | 6.70 | 450 |
| (9bS,15S)-1 | 0.22 | 28 |
| (9bR,15S)-2 | 12.0 | 400 |
| (9bS,15S)-2 | 24.0 | 100 |
| (9bR,15S)-3 | 1.59 | 405 |
| (9bS,15S)-3 | 0.40 | 405 |
| (9bR,15S)-4 | 11.8 | 394 |
| (9bS,15S)-4 | 1.54 | 394 |
| (9bR,15R)-5 | 223 | 446 |
| (9bS,15R)-5 | >446 | 446 |
| (R)-6 | 214 | 427 |
| (S)-6 | 26.7 | 427 |
| (9bR,15S)-7 | >246 | 246 |
| (9bS,15S)-7 | 15.4 | 7.40 |
| (9bR,15S)-8 | 7.80 | >260 |
| (9bS,15S)-8 | 1.00 | >260 |
| (9bR,15S)-9 | >253 | >253 |
| (9bS,15S)-9 | >253 | >253 |
2. Results and discussion
2.1 Design and synthesis of UA-based enamines
To improve both safety and drug-like properties of UA while retaining its antifungal properties, a series of semisynthetic derivatives of (R)-UA and (S)-UA were designed (1–9). A comprehensive review of the literature has revealed that the approved antifungal drugs are medium-to-large sized molecules, characterised by intrinsic flexibility.8,49,50 Accordingly, we decided to derivatize the constrained and disk-shape UA enantiomers as enamines, since this chemical modification, unlike the others applied on the scaffold, allowed the insertion of fragments endowed with different flexibility, length, and polarity (Fig. 1). Specifically, the ketone in position 13 was derivatized with amino acids (i.e. serine, arginine, phenylalanine and tyrosine), and hydrophobic benzyl moieties (i.e. 1-methyl-benzylamine and 3-chloro-benzylamine). We also explored the conjugation of compounds 4 with flexible PEG chains or linear carbon chains. While the introduction of PEG moiety may offer many benefits, including the reduced renal excretion and proteolysis as well as increased water solubility, the saturated carbon chain may facilitate molecule permeability through membranes or disrupt their integrity.51The semisynthetic derivatives 1–9 were prepared starting from homochiral UA. (S)-UA was isolated from Cladonia foliacea following our in-house well-established protocol, whereas the opposite enantiomer was purchased.26
The synthesis of compounds 1–6 is described in the Scheme 1.
UA enantiomers were condensed with the appropriate amines in presence of triethylamine as base, in ethanol under heating affording the corresponding enamine derivatives 1–6, as described in the Scheme 1. To attain a green and efficient synthesis, microwaves irradiation was investigated. With the sole exception of compound 2, mw irradiation resulted an efficient procedure for the preparation of the UA-based enamines, since it allowed obtain the product in 45 minutes with yield ranging from 37 to 87%, depending on the substrate. For the synthesis of compounds 7–9, long chain amines 10–13 were prepared first, according to the Scheme 2.
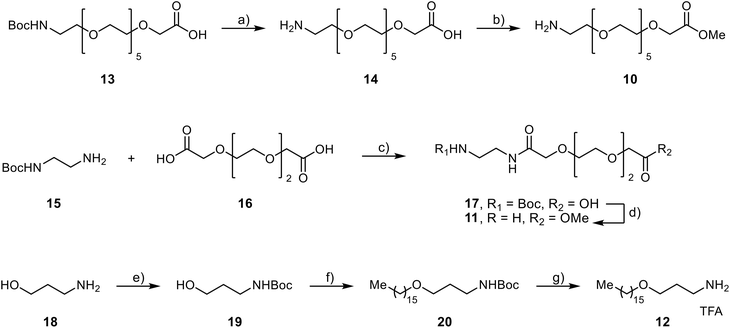
Compound 10 was obtained starting from the commercially available synthon 13 which was Boc-deprotected using trifluoroacetic acid in dichloromethane at 0 °C for 2 hours, to give amino acid 14 which underwent a mw-assisted Fisher esterification in methanol, using sulfuric acid as catalyst to give the corresponding methyl ester 10. For the preparation of compound 11, the N-Boc-ethylenediamine 15 was condensed with the dicarboxylic acid 16 using 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide hydrochloride (EDC-HCl) and a catalytic amount of hydroxy-benzotriazole (HOBt) as coupling agents, and diisopropylethylamine (DIPEA) as base in anhydrous DMF under nitrogen atmosphere for 16 hours, to provide the corresponding amide 17. Treating 17 with trimethylsilyl chloride (TMSCl) in methanol allowed both the simultaneous methyl esterification and the removal of the Boc-group, thus obtaining the amine 11. For the synthesis of amine 12, 3-amino-1-propanol 18 was alkylated with hexadecyl bromide affording the intermediate 19 followed by Boc-deprotection. Lastly, compound 4 was coupled with amines 10–12 in presence of EDC-HCl, HOBt and DIPEA, affording the corresponding enamines 7–6 (Scheme 3).
 | ||
| Scheme 3 Reagents and conditions. a) Amine 10–12 (1–1.5 equiv.), TEA (2 equiv.), EDC-HCl (1.5 equiv.), HOBt (0.1 equiv.), DIPEA (4.0 equiv.), anh. DMF, N2 atm, from 0 °C to r.t., 3–14 h. | ||
All newly synthesized compounds were fully characterized by 1H and 13C NMR spectroscopy. The observed chemical shifts were in accordance with previously reported values for structurally analogous compounds, confirming the regioselective derivatization of usnic acid at the C-13 position.52–54
2.2 Antifungal activity
The antifungal activity of the newly synthesized compounds 1–9, alongside the parent enantiomers (R)-UA and (S)-UA, was evaluated against the following clinical strains: Candida albicans ATCC 1023146, isolated from man with bronchomycosis Candida tropicalis ATCC 750 isolated from bronchitis patient, and a strain of Trichophyton rubrum LM 237 isolated from patient with foot onychomycosis and maintained in the mycological collection of the University of Pavia (LM). The MIC99 values (defined as the minimum concentration required to inhibit 99% of fungal growth) was determined by means of multiwell microplates.To assess the therapeutic potential of our compounds, we selected clinical isolates with known resistance to FCZ and AmB. Consistent with their resistant profiles, both reference drugs displayed limited effectiveness (MIC99 > 200 μM or >400 μM, Table 1). This strategic choice is in line with medical need and clinical urgency. Whilst FCZ is widely used to treat candidemia, especially against C. albicans infections, its efficacy has started to be affected by the emergence tolerance. Persistent candidemia with C. tropicalis is showing a worrying increase in FCZ resistance, reaching rates up to 25%.55T. rubrum, a predominant cause of chronic dermatophytosis, is increasingly resistant to azoles, especially FCZ, often exhibiting MIC50 values ≥160 μM following prolonged therapeutic exposures.
The situation with amphotericin B is somewhat different. Both C. albicans and C. tropicalis exhibit low resistance to AmB, but some cases of resistant strains have started to being reported, particularly among immunocompromised patients, likely because of prior exposure to antifungal agents, as the ones used in the present study.56,57 Although no evidence of resistance against T. rubrum has been observed, its use against dermatophytes is not common due to its nephrotoxicity and the superficial nature of dermatophyte infections.58
Regarding the newly developed series, a first screening at a concentration of 200 μM was performed. Since no effect of the UA derivatives was observed with C. albicans, this strain was not further investigated. A different behaviour was observed for C. tropicalis and T. rubrum, as summarized in Table 1.
Specifically, UA enantiomers as well as most semisynthetic derivatives (except for compounds 5, (R)-6, (9bS,15S)-7 and 9) displayed outstanding antifungal activity against C. tropicalis with MIC99 values in the low to sub-micromolar range. Among these, compounds (9bR,15S)-1, (9bS,15S)-1, (9bR,15S)-3, (9bS,15S)-3, (9bS,15S)-4, (9bR,15S)-8, and (9bS,15S)-8 stood out for their single-digit/sub-micromolar activity with MIC99 values of 6.70, 0.22, 1.59, 0.40, 1.54, 7.80 and 1.00 μM, respectively. To determine whether the observed antifungal activity was fungicidal or fungistatic, the minimum fungicidal concentrations (MFC) were determined for the most potent compounds. Compounds (9bS,15S)-3 and (9bS,15S)-4 exhibited MFC values of 25.4 μM and 49.3 μM, with an MFC/MIC ratio of 63.5 and 32, respectively, thus highlighting a prominent fungistatic rather than fungicidal mode of action.
T. rubrum exhibited modest susceptibility toward to both UA enantiomers and most semisynthetic derivatives 1–9 with MIC99 values ranging from 100 to 580 μM. Notably, compounds (9bS,15S)-1 and (9bS,15S)-7 represented significative exceptions, exhibiting MIC99 values of 28 and 7.4 μM, respectively, resulting in a 20-fold and 78-fold potency gains over the parent (S)-UA.
2.3 Effect of stereochemistry at position 9b on antifungal activity
Analysis of MIC99 values of compounds bearing the (R)- and (S)-UA as scaffold reveals significant insight into the role of chirality in the antifungal efficacy. Results reported in Table 1 suggest that the absolute configuration impacts antifungal activity, with structural modifications of the parent scaffold amplifying potency differences between the UA enantiomers. This relationship can be quantified by calculating the ratio between the MIC99 values of (R)-UA (or its derivative) and the MIC99 of (S)-UA (or its derivative). It should be noted that many compounds in our series contain an additional chiral center on the side chain at C-15, resulting in diastereomeric rather than enantiomeric pairs. However, since the absolute configuration of this additional stereocenter remains consistent across compared pairs, we can reasonably attribute the differences in the activity to the absolute configuration of configuration at position 9b of UA scaffold.Focusing on the parent enantiomers (R)-UA and (S)-UA, they exhibited modest difference in antimycotic activity against C. tropicalis, with (S)-UA resulting 4-fold more potent (MIC99 of 17.42 and 4.54 μM, for (R)-UA and (S)-UA, respectively). However, upon derivatization, several derivatives demonstrated significantly enhanced potency and increased eudysmic ratio (ER, defined as the ratio of activity between stereoisomers at position 9b), underscoring the critical role in activity (Fig. 2). Notably, compounds with (S)-configuration at position 9b demonstrated superior activity. For example, (9bS,15S)-1 exhibited ERs of 15 and 31 in the MIC99 against T. rubrum and C. tropicalis, respectively. Similarly, compounds (9bS,15S)-3, (9bS,15S)-4, (9bS,15S)-6 and (9bS,15S)-8 showed ERs between 4 and 8 toward C. tropicalis. The most pronounced stereochemical dependence was observed with compound (9bS,15S)-7 which showed the highest eudysmic ratio in the whole series (ER = 33) in the activity against T. rubrum.
These data collectively suggest that the (S)-configuration at position 9b of UA scaffold generally confers optimal antifungal activity. This stereochemical arrangement likely facilitates more favourable interaction with fungal targets or more efficient disruption of fungal cellular processes. Our findings highlight the potential of (S)-UA and its semisynthetic derivatives as promising candidates for addressing resistance mechanisms in clinically challenging fungal pathogens.26
2.4 Cytotoxicity studies
A primary requirement for in vivo application of novel antifungal agents is their biocompatibility, which includes both non-toxicity and good tissue tolerance. To determine the safety margin—the concentration range that is effective against fungi but safe for human cells—of our newly developed derivatives, we conducted cytotoxicity studies on normal human dermal fibroblasts at two different concentrations (50 μM, and 125 μM). This cell line was selected because both C. albicans and T. rubrum are responsible for mucocutaneous and cutaneous infections, making dermal fibroblast physiologically relevant for evaluation.Results demonstrated that the enamine derivatization decreases the cytotoxicity profile of the parent UA (Fig. S1 and S2†). Specifically, most of the new derivatives display good cytocompatibility profile with cell viability percentages above 90% at both tested concentrations (Fig. 3). Exceptions include compound (9bR,15S)-7, which resulted completely cytotoxic, and the pair of diastereomers (9bR,15S)-8 and (9bS,15S)-8 which exerted cytotoxicity only at the highest concentration tested.
2.5 Water thermodynamic solubility.
Given the poor water solubility associated with UA core, we envisaged a possible topical formulation for the enamine derivatives. In this context, we experimentally evaluated water solubility for the most promising compounds as this parameter is critical for developing effective topical formulations ensuring the drug delivery across skin barriers. Aqueous solubility was determined by HPLC for the best-performing compounds (9bR,15S)-1, 3, 4, 7 and 8 as well as their corresponding diastereoisomer, applying a method already developed by us.59 Results reported in Table 2 evidence that enamines (9bR,15S)-1, 3 and 4 incorporating amino acids motifs, exhibit a markedly enhanced water solubility ranging from 0.15 to 4.7 mM. However, no solubility improvements were observed for (9bR,15S)-7 and 8 despite the insertion of PEGylated lateral chains.| Cmpd | Aqueous solubility (mM) at 25 °C |
|---|---|
| LOD: limit of detection (0.54 μM); LOQ: limit of quantification (61.13 μM). | |
| UA | <0.3 (lit.)60 |
| (9bS,15S)-1 | 0.15 |
| (9bR,15S)-3 | 4.36 |
| (9bS,15S)-3 | 4.70 |
| (9bR,15S)-4 | 3.81 |
| (9bS,15S)-4 | 1.92 |
| (9bS,15S)-7 | <LOD |
| (9bS,15S)-8 | <LOQ |
3. Conclusion
This study explored the potential of UA as a chiral scaffold for the development of semisynthetic antimycotic agents. A series of semisynthetic enamine derivatives of both (R)- and (S)-UA were rationally designed, synthesized, and evaluated for their antifungal properties against clinically relevant, drug-resistant strains of C. tropicalis and T. rubrum. Among the series, compounds (9bS,15S)-1, 3, 4, and 8 demonstrated remarkable low to sub-micromolar activity against C. tropicalis, outperforming both the parent UA and the clinical antifungals amphotericin B and fluconazole. Notably, compound (9bS,15S)-1 also exhibited activity against T. rubrum indicating a broader spectrum efficacy. Biological evaluation revealed that configuration at the 9b stereocenter of UA affects the antifungal activity. In almost all the diastereomeric pairs, derivatives bearing the 9bS configuration outperformed their 9bR counterparts, establishing structure activity relationship that can guide future optimization. Furthermore, the systematic derivatization of UA effectively addressed some liabilities of the parent compound, i.e. enhanced safety and improved solubility. Cytocompatibility assays on human dermal fibroblasts confirmed that enamine derivatization reduced the UA intrinsic toxicity, with most compounds maintaining >90% cell viability at therapeutically relevant concentrations. The enhanced water solubility observed for several amino acid derivatives significantly improves their drug-likeness profile, supporting further development for both topical and potentially systemic applications.Overall, this study establishes the (S)-enantiomer of UA as a privileged chiral scaffold for the development of antifungal agents and highlights the derivatisation with amino acids as a valuable strategy for optimizing physicochemical and biological properties. Based on their antifungal activity, safety, and solubility profiles, (9bS,15S)-1, (9bS,15S)-3, and (9bS,15S)-4 have been identified as the most promising compounds and will be prioritized for further medicinal chemistry and pharmacological investigation.
4. Material and methods
4.1 General
Reagents and solvents for synthesis were purchased from Sigma-Aldrich (Italy) or VWR and were used as received, unless otherwise specified. Thin-layer-chromatography (TLC) was carried out on silica gel precoated aluminium-plates (Fluka Kieselgel 60 F254, Merck) and visualized by ultra-violet (UV) lamp and ninhydrin stain.The monomodal oven Discover® SP instrument (CEM Corporation, Matthews, NC, USA) was used to perform reactions with microwave irradiations.
NMR spectra were recorded on i) a Bruker Avance 400 spectrometer with 1H at 400.134 MHz and 13C at 100.62 MHz, and ii) a Bruker NMR Avance Neo 700 MHz with 13C at 176 MHz. Proton chemical shifts (δ) were reported in ppm and referenced to the solvent residual peak (CDCl3, δ = 7.26 ppm; CD3OD, δ = 3.31 ppm; DMSO-d6, δ = 2.50 ppm). Signals were abbreviated as s (singlet), d (doublet), t (triplet), q (quartet), m (multiplet) and br (broad). The coupling constant values (J) are reported in Hertz (Hz). 13C NMR spectra were recorded with complete proton decoupling. Carbon chemical shifts (δ) were reported in ppm and referenced to the solvent residual peak (CDCl3, δ = 77.23 ppm; CD3OD, δ = 49.00 ppm; DMSO-d6, δ = 39.52 ppm). Compound purity was evaluated by HPLC-UV/vis on a Jasco (Tokyo, Japan) system consisting of a PU-1580 pump and a MD-1510 photodiode array (PDA) detector. Chromatogram acquisitions and elaborations were performed using the ChromNAV software (Tokyo, Japan). Analyses were run on a XBridge™ Phenyl, (4.6 × 150 mm, 5 μm) column, at room temperature. The mobile phases were A: water containing 0.1% of formic acid, and B: acetonitrile containing 0.1% of formic acid. Elution was performed on a linear gradient from 50% to 100% B over 10 min, followed by an isocratic hold at 100% B for 3 min. The flow rate was 1.0 mL min−1 and the injection volume 10 μL. The chromatograms were recorded at 308 nm wavelength. All the final synthesized compounds showed a purity ≥95%. Optical rotation values were recorded using a Jasco photoelectric polarimeter DIP 1000 with a 0.5 dm quartz cell at the sodium D line (λ = 589 nm). The IUPAC names of each compound were generated using ChemDraw Professional 16.0.
4.2 (−)-(S)-Usnic acid extraction from Cladonia foliacea
30 g of C. foliacea matrix was freshly prepared by grinding the lichen thalli. Three aliquots of 10 g were suspended in EtOH and then heated three-times under mw irradiation (100 W, 120 psi) at 80 °C for 5 minutes. The extract was filtered, and the solvent was evaporated under reduced pressure. The resulting green syrup was resuspended in MeOH to remove polar undesirable metabolites, filtered, and concentrated. The residue was partitioned between water and DCM. The organic phase was collected, dried over anhydrous Na2SO4, filtered, and concentrated. (−)-(S)-UA was purified from the crude by crystallisation with CHCl3/EtOH 1![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 2. 250 mg (0.08% yield) of (−)-(S)-UA were obtained as a yellow solid. 1H NMR (400 MHz, CDCl3) δ 13.34 (s, 1H), 11.05 (s, 1H), 6.00 (s, 1H), 2.70 (s, 3H), 2.69 (s, 3H), 2.13 (s, 3H), 1.79 (s, 3H). 13C NMR (101 MHz, CDCl3) δ 201.93, 200.48, 198.20, 191.86, 179.52, 164.03, 157.65, 155.35, 109.47, 105.37, 104.09, 101.67, 98.48, 59.22, 32.26, 31.43, 28.05, 7.69. ESI-MS (m/z): [M–H]− calcd for C18H15O7−, 343.1; found 343.0. HPLC k = 3.83, mp 204 °C, [α]20D −477° (c = 0.2%, CHCl3).
2. 250 mg (0.08% yield) of (−)-(S)-UA were obtained as a yellow solid. 1H NMR (400 MHz, CDCl3) δ 13.34 (s, 1H), 11.05 (s, 1H), 6.00 (s, 1H), 2.70 (s, 3H), 2.69 (s, 3H), 2.13 (s, 3H), 1.79 (s, 3H). 13C NMR (101 MHz, CDCl3) δ 201.93, 200.48, 198.20, 191.86, 179.52, 164.03, 157.65, 155.35, 109.47, 105.37, 104.09, 101.67, 98.48, 59.22, 32.26, 31.43, 28.05, 7.69. ESI-MS (m/z): [M–H]− calcd for C18H15O7−, 343.1; found 343.0. HPLC k = 3.83, mp 204 °C, [α]20D −477° (c = 0.2%, CHCl3).
4.3 Chemistry
Methyl (1-((R)-6-acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2(1H)-ylidene)ethyl)-L-serinate ((9bR,15S)-1). The title compound was purified over silica gel, mobile phase DCM/MeOH, 95
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5. White solid (50% yield). 1H NMR (400 MHz, MeOD) δ 5.85 (s, 1H), 4.61 (s, 1H), 4.15 (dd, J = 11.5, 3.7 Hz, 2H), 3.96 (td, J = 10.9, 3.4 Hz, 2H), 3.86 (s, 3H), 2.69 (s, 3H), 2.65 (s, 3H), 2.06 (s, 5H), 1.74 (s, 3H), proton on heteroatoms exchange with the solvent. 13C NMR (176 MHz, CDCl3) δ 200.77, 200.61, 198.93, 175.45, 174.90, 174.58, 168.86, 168.76, 163.94, 163.75, 158.20, 157.89, 155.90, 155.62, 108.51, 108.28, 104.50, 101.47, 101.31, 63.28, 62.95, 58.79, 58.31, 53.49, 31.33, 19.25, 7.55. ESI-MS (m/z): [M–H]− calcd for C22H22NO9−, 444.13; found 444.08 and [M + Cl]− calcd for C22H2335[Cl]NO9−, 480.11; found 480.78. HPLC-UV/vis: k = 2.23, [α]20D +264.00° (c 0.25, CHCl3).
5. White solid (50% yield). 1H NMR (400 MHz, MeOD) δ 5.85 (s, 1H), 4.61 (s, 1H), 4.15 (dd, J = 11.5, 3.7 Hz, 2H), 3.96 (td, J = 10.9, 3.4 Hz, 2H), 3.86 (s, 3H), 2.69 (s, 3H), 2.65 (s, 3H), 2.06 (s, 5H), 1.74 (s, 3H), proton on heteroatoms exchange with the solvent. 13C NMR (176 MHz, CDCl3) δ 200.77, 200.61, 198.93, 175.45, 174.90, 174.58, 168.86, 168.76, 163.94, 163.75, 158.20, 157.89, 155.90, 155.62, 108.51, 108.28, 104.50, 101.47, 101.31, 63.28, 62.95, 58.79, 58.31, 53.49, 31.33, 19.25, 7.55. ESI-MS (m/z): [M–H]− calcd for C22H22NO9−, 444.13; found 444.08 and [M + Cl]− calcd for C22H2335[Cl]NO9−, 480.11; found 480.78. HPLC-UV/vis: k = 2.23, [α]20D +264.00° (c 0.25, CHCl3).
Methyl (1-((S)-6-acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]-furan-2(1H)-ylidene)ethyl)-L-serinate ((9bS,15S)-1). The title compound was purified over silica gel, mobile phase DCM/MeOH, 95
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5. White solid (37% yield). 1H NMR (400 MHz, CDCl3) δ 13.29 (s, 1H), 11.63 (s, 1H), 5.76 (s, 1H), 4.55 (dt, J = 8.1, 4.0 Hz, 2H), 4.20–4.08 (m, 1H), 4.03–3.99 (m, 1H), 3.79 (s, 3H), 2.61 (s, 3H), 2.57 (s, 3H), 2.03 (s, 3H), 1.65 (s, 3H), proton on enaminic NH and on serin hydroxyl exchange with the solvent. 13C NMR (176 MHz, CDCl3) δ 200.81, 200.70, 199.02, 198.90, 175.41, 174.53, 168.86, 168.74, 163.85, 163.73, 158.26, 158.07, 155.94, 155.77, 108.29, 105.01, 104.75, 101.52, 63.14, 62.96, 58.53, 58.21, 53.58, 31.44, 19.17, 7.63. ESI-MS (m/z): [M–H]− calcd for C22H22NO9−, 444.13; found 444.10 and [M + Cl]− calcd for C22H2335[Cl]NO9−, 480.11; found 480.51. HPLC-UV/vis: k = 1.70, [α]20D −420.40° (c 0.25, CHCl3).
5. White solid (37% yield). 1H NMR (400 MHz, CDCl3) δ 13.29 (s, 1H), 11.63 (s, 1H), 5.76 (s, 1H), 4.55 (dt, J = 8.1, 4.0 Hz, 2H), 4.20–4.08 (m, 1H), 4.03–3.99 (m, 1H), 3.79 (s, 3H), 2.61 (s, 3H), 2.57 (s, 3H), 2.03 (s, 3H), 1.65 (s, 3H), proton on enaminic NH and on serin hydroxyl exchange with the solvent. 13C NMR (176 MHz, CDCl3) δ 200.81, 200.70, 199.02, 198.90, 175.41, 174.53, 168.86, 168.74, 163.85, 163.73, 158.26, 158.07, 155.94, 155.77, 108.29, 105.01, 104.75, 101.52, 63.14, 62.96, 58.53, 58.21, 53.58, 31.44, 19.17, 7.63. ESI-MS (m/z): [M–H]− calcd for C22H22NO9−, 444.13; found 444.10 and [M + Cl]− calcd for C22H2335[Cl]NO9−, 480.11; found 480.51. HPLC-UV/vis: k = 1.70, [α]20D −420.40° (c 0.25, CHCl3).
(1-((R)-6-Acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2-(1H)-ylidene)ethyl)-L-phenylalanine ((9bR,15S)-3). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. Brownish oil (87% yield). 1H NMR (400 MHz, CDCl3) δ, 13.57 (s, 1H), 13.36 (s, 1H), 12.16 (s, 1H), 7.47–6.84 (m, 5H), 5.76 (s, 1H), 4.53 (s, 1H), 3.42–3.40 (m, 1H), 3.12–3.05 (m, 1H), 2.66 (s, 3H), 2.16 (s, 3H), 2.07 (s, 3H), 1.66 (s, 3H), proton on enaminic NH exchanges with the solvent. 13C NMR (101 MHz, CDCl3) δ 200.82, 197.94, 173.81, 171.26, 163.45, 158.52, 156.08, 136.80, 129.60, 128.71, 127.18, 107.78, 105.45, 102.90, 101.38, 60.49, 45.36, 31.36, 21.14, 18.70, 14.28, 8.61. ESI-MS (m/z): [M–H]− calcd for C27H24NO8−, 490.15; found 490.03 and [M + Cl]− calcd for C27H2535[Cl]NO8−, 526.13; found 526.46. HPLC-UV/vis: k = 2.84, [α]20D +103.80° (c 0.25, CHCl3).
1. Brownish oil (87% yield). 1H NMR (400 MHz, CDCl3) δ, 13.57 (s, 1H), 13.36 (s, 1H), 12.16 (s, 1H), 7.47–6.84 (m, 5H), 5.76 (s, 1H), 4.53 (s, 1H), 3.42–3.40 (m, 1H), 3.12–3.05 (m, 1H), 2.66 (s, 3H), 2.16 (s, 3H), 2.07 (s, 3H), 1.66 (s, 3H), proton on enaminic NH exchanges with the solvent. 13C NMR (101 MHz, CDCl3) δ 200.82, 197.94, 173.81, 171.26, 163.45, 158.52, 156.08, 136.80, 129.60, 128.71, 127.18, 107.78, 105.45, 102.90, 101.38, 60.49, 45.36, 31.36, 21.14, 18.70, 14.28, 8.61. ESI-MS (m/z): [M–H]− calcd for C27H24NO8−, 490.15; found 490.03 and [M + Cl]− calcd for C27H2535[Cl]NO8−, 526.13; found 526.46. HPLC-UV/vis: k = 2.84, [α]20D +103.80° (c 0.25, CHCl3).
(1-((S)-6-Acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2-(1H)-ylidene)ethyl)-L-phenylalanine ((9bS,15S)-3). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. Brownish oil (51% yield). 1H NMR (400 MHz, CDCl3) δ 13.52 (s, 1H), 13.36 (s, 1H), 12.11 (s, 1H), 7.40–6.99 (m, 5H), 5.75 (s, 1H), 4.52 (td, J = 9.0, 3.8 Hz, 1H), 3.42 (dd, J = 13.8, 4.0 Hz, 1H), 3.04 (d, J = 7.6 Hz, 1H), 2.67 (s, 3H), 2.18 (s, 3H), 2.08 (s, 3H), 1.67 (s, 3H), proton on enaminic NH exchanges with the solvent. 13C NMR (176 MHz, CDCl3) δ 200.73, 198.35, 174.03, 163.63, 158.44, 156.01, 136.61, 129.75, 128.79, 127.36, 108.12, 105.22, 102.50, 101.46, 45.46, 40.49, 32.07, 31.38, 29.84, 29.50, 28.52, 22.83, 18.54, 14.26, 8.66, 7.62, 1.16. ESI-MS (m/z): [M–H]− calcd for C27H24NO8−, 490.15; found 490.14 and [M + Cl]− calcd for C27H2535[Cl]NO8−, 526.13; found 526.91. HPLC-UV/vis: k = 2.90, [α]20D −267.15° (c 0.25, CHCl3).
1. Brownish oil (51% yield). 1H NMR (400 MHz, CDCl3) δ 13.52 (s, 1H), 13.36 (s, 1H), 12.11 (s, 1H), 7.40–6.99 (m, 5H), 5.75 (s, 1H), 4.52 (td, J = 9.0, 3.8 Hz, 1H), 3.42 (dd, J = 13.8, 4.0 Hz, 1H), 3.04 (d, J = 7.6 Hz, 1H), 2.67 (s, 3H), 2.18 (s, 3H), 2.08 (s, 3H), 1.67 (s, 3H), proton on enaminic NH exchanges with the solvent. 13C NMR (176 MHz, CDCl3) δ 200.73, 198.35, 174.03, 163.63, 158.44, 156.01, 136.61, 129.75, 128.79, 127.36, 108.12, 105.22, 102.50, 101.46, 45.46, 40.49, 32.07, 31.38, 29.84, 29.50, 28.52, 22.83, 18.54, 14.26, 8.66, 7.62, 1.16. ESI-MS (m/z): [M–H]− calcd for C27H24NO8−, 490.15; found 490.14 and [M + Cl]− calcd for C27H2535[Cl]NO8−, 526.13; found 526.91. HPLC-UV/vis: k = 2.90, [α]20D −267.15° (c 0.25, CHCl3).
(1-((R)-6-Acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2-(1H)-ylidene)ethyl)-L-tyrosine ((9bR,15S)-4). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1. White solid (48% yield). 1H NMR (400 MHz, CDCl3) δ 13.36 (s, 1H), 12.19 (s, 1H), 7.01 (d, J = 8.4 Hz, 2H), 6.69 (d, J = 8.4 Hz, 2H), 5.77 (s, 1H), 4.47 (td, J = 8.2, 4.5 Hz, 1H), 3.25 (dd, J = 13.8, 4.4 Hz, 1H), 3.04 (dd, J = 13.8, 8.2 Hz, 1H), 2.66 (s, 3H), 2.28 (s, 3H), 2.08 (s, 3H), 1.67 (s, 3H), protons on enaminic NH, carboxylic acid and phenol of tyrosine exchange with the solvent. 13C NMR (101 MHz, DMSO) δ 200.98, 196.84, 188.18, 172.55, 162.48, 157.82, 156.05, 155.84, 130.38, 129.17, 128.23, 127.30, 115.09, 106.20, 105.27, 102.59, 101.40, 100.84, 56.03, 45.31, 31.76, 31.07, 18.61, 8.84, 7.52. ESI-MS (m/z): [M–H]− calcd for C27H24NO9−, 506.15; found 506.64, [M + Cl]− calcd for C27H2535[Cl]NO9−, 542.12; found 542.91 and C27H2537[Cl]NO9−, 544.12; found 544.05. HPLC-UV/vis: k = 1.81, [α]20D +80.35° (c 0.2, CHCl3).
1. White solid (48% yield). 1H NMR (400 MHz, CDCl3) δ 13.36 (s, 1H), 12.19 (s, 1H), 7.01 (d, J = 8.4 Hz, 2H), 6.69 (d, J = 8.4 Hz, 2H), 5.77 (s, 1H), 4.47 (td, J = 8.2, 4.5 Hz, 1H), 3.25 (dd, J = 13.8, 4.4 Hz, 1H), 3.04 (dd, J = 13.8, 8.2 Hz, 1H), 2.66 (s, 3H), 2.28 (s, 3H), 2.08 (s, 3H), 1.67 (s, 3H), protons on enaminic NH, carboxylic acid and phenol of tyrosine exchange with the solvent. 13C NMR (101 MHz, DMSO) δ 200.98, 196.84, 188.18, 172.55, 162.48, 157.82, 156.05, 155.84, 130.38, 129.17, 128.23, 127.30, 115.09, 106.20, 105.27, 102.59, 101.40, 100.84, 56.03, 45.31, 31.76, 31.07, 18.61, 8.84, 7.52. ESI-MS (m/z): [M–H]− calcd for C27H24NO9−, 506.15; found 506.64, [M + Cl]− calcd for C27H2535[Cl]NO9−, 542.12; found 542.91 and C27H2537[Cl]NO9−, 544.12; found 544.05. HPLC-UV/vis: k = 1.81, [α]20D +80.35° (c 0.2, CHCl3).
(1-((S)-6-Acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2-(1H)-ylidene)ethyl)-L-tyrosine ((9bS,15S)-4). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 + 0.1% NH3 (in MeOH). Pale yellow solid (54% yield). 1H NMR (400 MHz, MeOD) δ 7.08 (d, J = 8.4 Hz, 2H), 6.70 (d, J = 8.4 Hz, 2H), 5.76 (s, 1H), 4.57–4.54 (m, 1H), 3.33–3.31 (m, 1H, under MeOD), 2.96 (dd, J = 13.9, 9.3 Hz, 1H), 2.65 (s, 3H), 2.23 (s, 3H), 2.01 (s, 3H), 1.66 (s, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (101 MHz, DMSO) δ 201.00, 197.43, 188.82, 173.92, 172.80, 170.81, 162.68, 156.30, 155.81, 133.15, 130.55, 126.28, 115.17, 106.29, 105.17, 102.43, 101.60, 100.89, 56.30, 45.35, 31.64, 31.08, 18.40, 8.50, 7.53. ESI-MS (m/z): [M–H]− calcd for C27H24NO9−, 506.15; found 506.29 and [M + Cl]− calcd for C27H2535[Cl]NO9−, 542.12; found 542.36. HPLC-UV/vis: k = 1.56, [α]20D −176.5° (c 0.25, CHCl3).
1 + 0.1% NH3 (in MeOH). Pale yellow solid (54% yield). 1H NMR (400 MHz, MeOD) δ 7.08 (d, J = 8.4 Hz, 2H), 6.70 (d, J = 8.4 Hz, 2H), 5.76 (s, 1H), 4.57–4.54 (m, 1H), 3.33–3.31 (m, 1H, under MeOD), 2.96 (dd, J = 13.9, 9.3 Hz, 1H), 2.65 (s, 3H), 2.23 (s, 3H), 2.01 (s, 3H), 1.66 (s, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (101 MHz, DMSO) δ 201.00, 197.43, 188.82, 173.92, 172.80, 170.81, 162.68, 156.30, 155.81, 133.15, 130.55, 126.28, 115.17, 106.29, 105.17, 102.43, 101.60, 100.89, 56.30, 45.35, 31.64, 31.08, 18.40, 8.50, 7.53. ESI-MS (m/z): [M–H]− calcd for C27H24NO9−, 506.15; found 506.29 and [M + Cl]− calcd for C27H2535[Cl]NO9−, 542.12; found 542.36. HPLC-UV/vis: k = 1.56, [α]20D −176.5° (c 0.25, CHCl3).
(R)-6-Acetyl-7,9-dihydroxy-8,9b-dimethyl-2-(1-(((R)-1-phenylethyl)amino)ethylidene) dibenzo[b,d]furan-1,3(2H,9bH)-dione ((9bR,15R)-5). The title compound was purified over silica gel, mobile phase DCM/MeOH, 95
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5. Yellow solid (61% yield). 1H NMR (400 MHz, MeOD) δ 7.49–7.36 (m, 5H), 5.86 (s, 1H), 5.19 (q, J = 6.7 Hz, 1H), 2.68 (s, 3H), 2.59 (s, 3H), 2.05 (s, 3H), 1.71 (s, 3H), 1.68 (d, J = 6.7 Hz, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (176 MHz, CDCl3) δ 200.78, 198.64, 191.02, 174.41, 163.60, 158.37, 155.98, 141.74, 129.44, 128.25, 125.77, 108.10, 105.17, 102.43, 102.24, 101.47, 57.49, 54.46, 31.98, 31.39, 24.26, 18.96, 7.59. ESI-MS (m/z): [M–H]− calcd for C26H24NO6−, 446.16; found 446.34, [M + Cl]− calcd for C26H2535[Cl]NO6−, 482.14; found 482.28 and calcd for C26H2537[Cl]NO6−, 484.14; found 484.51. HPLC-UV/vis: k = 3.94, [α]20D +186.3° (c 0.5, CHCl3).
5. Yellow solid (61% yield). 1H NMR (400 MHz, MeOD) δ 7.49–7.36 (m, 5H), 5.86 (s, 1H), 5.19 (q, J = 6.7 Hz, 1H), 2.68 (s, 3H), 2.59 (s, 3H), 2.05 (s, 3H), 1.71 (s, 3H), 1.68 (d, J = 6.7 Hz, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (176 MHz, CDCl3) δ 200.78, 198.64, 191.02, 174.41, 163.60, 158.37, 155.98, 141.74, 129.44, 128.25, 125.77, 108.10, 105.17, 102.43, 102.24, 101.47, 57.49, 54.46, 31.98, 31.39, 24.26, 18.96, 7.59. ESI-MS (m/z): [M–H]− calcd for C26H24NO6−, 446.16; found 446.34, [M + Cl]− calcd for C26H2535[Cl]NO6−, 482.14; found 482.28 and calcd for C26H2537[Cl]NO6−, 484.14; found 484.51. HPLC-UV/vis: k = 3.94, [α]20D +186.3° (c 0.5, CHCl3).
(S)-6-Acetyl-7,9-dihydroxy-8,9b-dimethyl-2-(1-(((R)-1-phenylethyl)amino)ethylidene) dibenzo[b,d]furan-1,3(2H,9bH)-dione ((9bS,15R)-5). The title compound was purified over silica gel, mobile phase DCM/MeOH, 95
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 5. Yellowish oil (58% yield). 1H NMR (400 MHz, MeOD) δ 7.38–7.22 (m, 5H), 5.74 (s, 1H), 5.08 (q, J = 6.7 Hz, 1H), 2.57 (s, 3H), 2.47 (s, 3H), 1.93 (s, 3H), 1.59 (s, 3H), 1.56 (d, J = 6.8 Hz, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (101 MHz, CDCl3) δ 200.85, 198.88, 192.39, 188.68, 174.92, 163.66, 159.34, 156.18, 142.42, 130.33, 128.30, 125.81, 108.19, 105.21, 102.48, 101.54, 54.50, 32.01, 31.44, 24.31, 18.99, 7.62, 1.16. ESI-MS (m/z): [M–H]− calcd for C26H24NO6−, 446.16; found 446.21, [M + Cl]− calcd for C26H2535[Cl]NO6−, 482.14; found 482.47 and calcd for C26H2537[Cl]NO6−, 484.14; found 484.48. HPLC-UV/vis: k = 4.05, [α]20D −96.5° (c 0.25, CHCl3).
5. Yellowish oil (58% yield). 1H NMR (400 MHz, MeOD) δ 7.38–7.22 (m, 5H), 5.74 (s, 1H), 5.08 (q, J = 6.7 Hz, 1H), 2.57 (s, 3H), 2.47 (s, 3H), 1.93 (s, 3H), 1.59 (s, 3H), 1.56 (d, J = 6.8 Hz, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (101 MHz, CDCl3) δ 200.85, 198.88, 192.39, 188.68, 174.92, 163.66, 159.34, 156.18, 142.42, 130.33, 128.30, 125.81, 108.19, 105.21, 102.48, 101.54, 54.50, 32.01, 31.44, 24.31, 18.99, 7.62, 1.16. ESI-MS (m/z): [M–H]− calcd for C26H24NO6−, 446.16; found 446.21, [M + Cl]− calcd for C26H2535[Cl]NO6−, 482.14; found 482.47 and calcd for C26H2537[Cl]NO6−, 484.14; found 484.48. HPLC-UV/vis: k = 4.05, [α]20D −96.5° (c 0.25, CHCl3).
(R)-6-Acetyl-2-(1-((3-chlorobenzyl)amino)ethylidene)-7,9-dihydroxy-8,9b-dimethyldibenzo [b,d]furan-1,3(2H,9bH)-dione ((R)-6). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 + 0.1% NH3 (in MeOH). Yellow solid (47% yield). 1H NMR (400 MHz, CDCl3) δ 13.88 (s, 1H), 13.36 (s, 1H), 11.88 (s, 1H), 7.37–7.32 (m, 2H), 7.29 (s, 1H), 7.23–7.14 (m, 1H), 5.81 (s, 1H), 4.66 (d, J = 5.7 Hz, 2H), 2.68 (s, 3H), 2.65 (s, 3H), 2.10 (s, 3H), 1.72 (s, 3H). 13C NMR (176 MHz, CDCl3) δ 200.82, 198.80, 191.10, 175.43, 174.69, 163.69, 158.38, 155.98, 137.14, 135.39, 130.78, 128.87, 127.60, 125.42, 108.29, 105.12, 102.61, 102.43, 101.55, 57.63, 47.31, 32.05, 29.85, 18.72, 7.64. ESI-MS (m/z): [M–H]− calcd for C25H2135[Cl]NO6−, 466.11; found 466.03 and calcd for C25H2137[Cl]NO6−, 468.11; found 468.19. HPLC-UV/vis: k = 4.07, [α]20D +41.30° (c 0.5, CHCl3).
1 + 0.1% NH3 (in MeOH). Yellow solid (47% yield). 1H NMR (400 MHz, CDCl3) δ 13.88 (s, 1H), 13.36 (s, 1H), 11.88 (s, 1H), 7.37–7.32 (m, 2H), 7.29 (s, 1H), 7.23–7.14 (m, 1H), 5.81 (s, 1H), 4.66 (d, J = 5.7 Hz, 2H), 2.68 (s, 3H), 2.65 (s, 3H), 2.10 (s, 3H), 1.72 (s, 3H). 13C NMR (176 MHz, CDCl3) δ 200.82, 198.80, 191.10, 175.43, 174.69, 163.69, 158.38, 155.98, 137.14, 135.39, 130.78, 128.87, 127.60, 125.42, 108.29, 105.12, 102.61, 102.43, 101.55, 57.63, 47.31, 32.05, 29.85, 18.72, 7.64. ESI-MS (m/z): [M–H]− calcd for C25H2135[Cl]NO6−, 466.11; found 466.03 and calcd for C25H2137[Cl]NO6−, 468.11; found 468.19. HPLC-UV/vis: k = 4.07, [α]20D +41.30° (c 0.5, CHCl3).
(S)-6-Acetyl-2-(1-((3-chlorobenzyl)amino)ethylidene)-7,9-dihydroxy-8,9b-dimethyldibenzo [b,d]furan-1,3(2H,9bH)-dione ((S)-6). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 + 0.1% NH3 (in MeOH). Yellow solid (51% yield). 1H NMR (400 MHz, CDCl3) δ 13.88 (s, 1H), 13.36 (s, 1H), 11.88 (s, 1H), 7.37–7.32 (m, 2H), 7.29 (s, 1H), 7.23–7.14 (m, 1H), 5.81 (s, 1H), 4.66 (d, J = 5.7 Hz, 2H), 2.68 (s, 3H), 2.65 (s, 3H), 2.10 (s, 3H), 1.72 (s, 3H). 13C NMR (176 MHz, CDCl3) δ 200.82, 198.80, 191.10, 175.43, 174.69, 163.69, 158.38, 155.98, 137.14, 135.39, 130.78, 128.87, 127.60, 125.42, 108.29, 105.12, 102.61, 102.43, 101.55, 57.63, 47.31, 32.05, 29.85, 18.72, 7.64. ESI-MS (m/z): [M–H]− calcd for C25H2135[Cl]NO6−, 466.11; found 466.08 and calcd for C25H2137[Cl]NO6−, 468.11; found 468.22. HPLC-UV/vis: k = 4.07, [α]20D −41.10° (c 0.5, CHCl3).
1 + 0.1% NH3 (in MeOH). Yellow solid (51% yield). 1H NMR (400 MHz, CDCl3) δ 13.88 (s, 1H), 13.36 (s, 1H), 11.88 (s, 1H), 7.37–7.32 (m, 2H), 7.29 (s, 1H), 7.23–7.14 (m, 1H), 5.81 (s, 1H), 4.66 (d, J = 5.7 Hz, 2H), 2.68 (s, 3H), 2.65 (s, 3H), 2.10 (s, 3H), 1.72 (s, 3H). 13C NMR (176 MHz, CDCl3) δ 200.82, 198.80, 191.10, 175.43, 174.69, 163.69, 158.38, 155.98, 137.14, 135.39, 130.78, 128.87, 127.60, 125.42, 108.29, 105.12, 102.61, 102.43, 101.55, 57.63, 47.31, 32.05, 29.85, 18.72, 7.64. ESI-MS (m/z): [M–H]− calcd for C25H2135[Cl]NO6−, 466.11; found 466.08 and calcd for C25H2137[Cl]NO6−, 468.11; found 468.22. HPLC-UV/vis: k = 4.07, [α]20D −41.10° (c 0.5, CHCl3).
(1-((R)-6-Acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2(1H)-ylidene)ethyl)-L-arginine ((9bR,15S)-2). A suspension of (+)-(R)-UA (50 mg, 0.15 mmol), L-arginine (25 mg, 0.15 mmol), and TEA (41 μl, 0.30 mmol) in EtOH (1.5 mL, 0.1 M) was refluxed at 80 °C for 14 h under N2. The crude was purified by crystallisation with diisopropyl ether at 0 °C, affording the title compound as pale yellow solid (35 mg, 47%). 1H NMR (400 MHz, DMSO-d6) δ 13.39 (s, 1H), 13.27 (s, 1H), 12.48 (s, 1H), 8.90 (s, 1H), 7.43 (s, 4H), 5.71 (s, 1H), 4.25 (s, 1H), 3.15–3.08 (m, 2H), 2.59 (s, 3H), 2.57 (s, 3H), 1.94 (s, 3H), 1.87–1.76 (m, 2H), 1.59 (s, 3H), 1.55–1.47 (m, 2H). 13C NMR (176 MHz, DMSO-d6) δ 200.67, 197.02, 188.47, 172.29, 172.12, 172.05, 162.62, 157.73, 157.10, 155.67, 106.18, 105.15, 102.27, 101.59, 100.69, 58.25, 56.22, 40.33, 31.96, 30.94, 30.08, 24.85, 18.95, 7.47. ESI-MS (m/z): [M + H]+ calcd for C24H29N4O8+, 501.20; found 501.27 and [M–H]− calcd for C24H27N4O8−, 499.18; found 499.31. HPLC-UV/vis: k = 0.39, [α]20D +255.0° (c 0.2%, DMSO).
Synthesis of (1-((S)-6-Acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2(1H)-ylidene)ethyl)-L-arginine ((9bS,15S)-2). A suspension of (+)-(S)-UA (22 mg, 0.065 mmol), L-arginine (12 mg, 0.065 mmol), and TEA (18 μl, 0.13 mmol) in EtOH (1.5 mL) was refluxed at 80 °C for 14 h under nitrogen atmosphere. The solvent was removed under reduced pressure and the crude was purified by crystallisation with diisopropyl ether at 0 °C, affording the title compound as pale yellow solid (15 mg, 46% yield). 1H NMR (400 MHz, DMSO-d6) δ 13.06 (bs, 1H), 9.10 (bs, 1H), 7.59 (bs, 6H), 5.82 (s, 1H), 4.22 (t, J = 5.8 Hz, 1H), 3.15–3.07 (m, 2H), 2.63 (s, 3H), 2.54 (s, 3H), 1.96 (s, 3H), 1.87–1.76 (m, 2H), 1.65 (s, 3H), 1.58–1.50 (m, 2H). 13C NMR (176 MHz, MeOD) δ 200.90, 198.13, 189.61, 174.31, 173.98, 163.07, 158.05, 157.26, 156.12, 107.09, 105.12, 102.10, 102.01, 100.92, 58.84, 56.73, 40.66, 31.01, 30.02, 29.78, 24.62, 17.90, 6.21. ESI-MS (m/z): [M + H]+ calcd for C24H29N4O8+, 501.20; found 501.48 and [M–H]− calcd for C24H27N4O8−, 499.18; found 499.03. HPLC-UV/vis: k = 0.28, [α]20D −182.4° (c 0.2%, DMSO).
20-Amino-3,6,9,12,15,18-hexaoxaicosanoic acid (14). To a well-stirred solution of Boc-NH-PEG6-acid (13, 100 mg, 0.22 mmol, 1 equiv.) in DCM (1 mL), TFA was added dropwise (422 μl, 5.1 mmol, 5 equiv.) at 0 °C and the reaction was allowed to stir at r.t. for 14 h. The solvent was evaporated, and the residue was diluted with MeOH followed by the treatment with Amberlyst-15 ion-exchange acid resin for 5 h. Upon filtration, the resin was collected and swelled by alternate washing with DCM and MeOH. The resin was suspended in 50 mL of methanol in presence of 0.1% NH3 in MeOH and the resulting solution was shaken for 14 h. The suspension was filtrated and the concentrated to give the titled compound as a brownish oil (80 mg, quantitative yield). 1H NMR (400 MHz, CDCl3) δ 3.62 (t, J = 10.2 Hz, 20H), 2.55 (t, J = 6.5 Hz, 4H), 1.06 (t, J = 6.5 Hz, 2H).
Methyl 20-amino-3,6,9,12,15,18-hexaoxaicosanoate (10). To a solution of 14 (80 mg, 0.22 mmol, 1 equiv.) in MeOH (2.2 mL), 1 N H2SO4 (4 drops) was added dropwise under stirring. The mixture was heated three times under mw irradiation (200 W, 250 psi) at 120 °C for 3 minutes. Upon completion of the reaction, the mixture was treated with Amberlyst-15 ion-exchange acid resin and shaken for 5 h. After filtration, the resin was collected and swelled by alternating washing with DCM and MeOH. The resin was suspended in 50 mL of methanol in presence of 0.1% NH3 in MeOH and was shaken for 14 h. The suspension was filtrated and the concentrated to give the titled compound as a brownish oil (35 mg, 42% yield). 1H NMR (400 MHz, CDCl3) δ 3.75 (t, J = 6.4 Hz, 2H), 3.70–3.55 (m, 19H), 2.98 (t, J = 6.5 Hz, 2H), 2.60 (t, J = 6.5 Hz, 2H), 2.12–1.95 (br, 4H), 1.31–1.16 (m, 2H). 2.07–1.90 (br, 2H), 0.94–0.78 (m, 4H).
2,2-Dimethyl-4,9-dioxo-3,11,14,17-tetraoxa-5,8-diazanonadecan-19-oic acid (17). To a solution of 3,6,9-trioxaundecanedioic acid (16, 139 mg, 0.624 mmol, 2 equiv.) in anhydrous DMF (3.12 mL), EDC-HCl (72 mg, 0.374 mmol, 1.2 equiv.), HOBt (21 mg, 0.156 mmol, 0.5 equiv.) and DIPEA (273 μL, 1.56 mmol, 5 equiv.) were added, and the resulting solution was stirred at r.t. under nitrogen atmosphere for 30 minutes. N-Boc-ethylenediamine (15, 49 μL, 0.312 mmol, 1.0 equiv.) was added, and the reaction mixture was stirred for additional 6 h. n-Heptane was added to form the azeotrope with DMF, and the solvent was removed under reduced pressure. The crude was purified on silica gel (mobile phase, DCM/MeOH + 0.1% HCOOH, from 40
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 to 20
1 to 20![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1), to give the title compound as viscous clear oil (84 mg, 70% yield). 1H NMR (400 MHz, CDCl3) δ 7.61 (t, J = 6.1 Hz, 1H), 5.38 (s, 1H), 4.08 (s, 4H), 3.96 (d, J = 5.2 Hz, 2H), 3.72–3.58 (m, 6H), 3.38–3.29 (m, 2H), 3.24–3.15 (m, 2H), 1.35 (s, 9H).
1), to give the title compound as viscous clear oil (84 mg, 70% yield). 1H NMR (400 MHz, CDCl3) δ 7.61 (t, J = 6.1 Hz, 1H), 5.38 (s, 1H), 4.08 (s, 4H), 3.96 (d, J = 5.2 Hz, 2H), 3.72–3.58 (m, 6H), 3.38–3.29 (m, 2H), 3.24–3.15 (m, 2H), 1.35 (s, 9H).
Methyl 1-amino-4-oxo-6,9,12-trioxa-3-azatetradecan-14-oate hydrochloride (11). To a solution of 17 (84 mg, 0.215 mmol, 1 equiv.) in MeOH (2 mL), TMSCl (72 mg, 0.374 mmol, 1.2 equiv.) was added dropwise at 0 °C. The reaction was allowed to stir at r.t. for 6 h. The solvent was removed in vacuo and the residue was triturated with n-hexane and Et2O, to give the title compound as light-yellow oil (quantitative yield). 1H NMR (400 MHz, MeOD) δ 4.23–4.20 (m, 4H), 3.77–3.77 (m, 5H), 3.76–3.71 (m, 6H), 3.71–3.67 (m, 4H).
tert-Butyl (3-hydroxypropyl)carbamate (19). To a solution of 18 in anhydrous DCM, Boc2O was added at r.t., and the resulting mixture was stirred in the same conditions for 3 h. Upon completion of the reaction, the solvent was removed in vacuo and the crude was filtered through a small plug of silica gel eluting with a DCM/MeOH, 20
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 to give the desired product as transparent oil (quantitative yield). 1H NMR (400 MHz, CDCl3) δ 4.87 (s, 1H), 3.59 (q, J = 5.4 Hz, 2H), 3.21 (q, J = 6.3 Hz, 2H), 1.60 (p, J = 6.0 Hz, 2H), 1.37 (s, 9H).
1 to give the desired product as transparent oil (quantitative yield). 1H NMR (400 MHz, CDCl3) δ 4.87 (s, 1H), 3.59 (q, J = 5.4 Hz, 2H), 3.21 (q, J = 6.3 Hz, 2H), 1.60 (p, J = 6.0 Hz, 2H), 1.37 (s, 9H).
tert-Butyl (3-(hexadecyloxy)propyl)carbamate (20). To a solution of 19 (500 mg, 1.84 mmol, 1 equiv.) in anhydrous DMF (10 mL) at 0 °C, 60% NaH in mineral oil was added (66 mg, 2.75 mmol, 1.5 equiv.). After 1 h, 1-bromohexadecane (263 μL, 2.21 mmol, 1.2 equiv.) was added, and the resulting mixture was allowed to stir at r.t. for 16 h. The reaction was quenched with a saturated NH4Cl solution (20 mL) and then extracted with EtOAc. The combined organic layers were dried over anhydrous Na2SO4, filtered and concentrated. The crude was purified on silica gel (petroleum ether/EtOAc, from 6
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 to 5
1 to 5![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1) to afford the title compound as orange oil (497 mg, 75% yield). 1H NMR (400 MHz, CDCl3) δ 4.93–4.89 (bs, 1H), 3.40 (t, J = 5.9 Hz, 2H), 3.32 (t, J = 6.6 Hz, 2H), 3.15 (q, J = 6.3 Hz, 2H), 1.67 (p, J = 6.2 Hz, 2H), 1.49 (dt, J = 14.7, 7.1 Hz, 2H), 1.37 (s, 9H), 1.22–1.14 (m, 26H), 0.81 (t, J = 6.8 Hz, 3H).
1) to afford the title compound as orange oil (497 mg, 75% yield). 1H NMR (400 MHz, CDCl3) δ 4.93–4.89 (bs, 1H), 3.40 (t, J = 5.9 Hz, 2H), 3.32 (t, J = 6.6 Hz, 2H), 3.15 (q, J = 6.3 Hz, 2H), 1.67 (p, J = 6.2 Hz, 2H), 1.49 (dt, J = 14.7, 7.1 Hz, 2H), 1.37 (s, 9H), 1.22–1.14 (m, 26H), 0.81 (t, J = 6.8 Hz, 3H).
3-(Hexadecyloxy)propan-1-aminium 2,2,2-trifluoroacetate (12). A solution of 20 (99 mg, 0.248 mmol, 1 equiv.) in anhydrous DCM (1.24 mL) was cooled to 0 °C, and TFA was added dropwise under nitrogen atmosphere. The reaction mixture was allowed to stir 0 °C for 2 h, until the consumption of the starting material. The solvent was removed in vacuo, affording the title compound as yellow solid (quantitative yield). 1H NMR (400 MHz, CDCl3) δ 10.79–10.39 (bs, 1H), 7.66–7.31 (bs, 2H), 3.56 (t, J = 5.2 Hz, 2H), 3.36 (t, J = 6.9 Hz, 2H), 3.13 (s, 2H), 1.87 (p, J = 5.2 Hz, 2H), 1.48 (p, J = 6.6 Hz, 2H), 1.28–1.15 (m, 26H), 0.80 (t, J = 6.8 Hz, 3H).
Methyl (S,E)-2-((R)-6-acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2(1H)-ylidene)-4-(4-hydroxybenzyl)-5-oxo-9,12,15,18,21,24-hexaoxa-3,6-diazahexacosan-26-oate ((9bR,15S)-7). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 + 0.1% NH3 (in MeOH). Yellow solid (56% yield). 1H NMR (400 MHz, CDCl3) δ 13.83 (s, 1H), 13.30 (s, 1H), 11.89 (s, 1H), 7.03 (d, J = 8.1 Hz, 2H), 6.76 (d, J = 8.1 Hz, 2H), 6.55 (s, 1H), 5.75 (s, 1H), 4.38 (d, J = 7.1 Hz, 1H), 3.67 (t, J = 6.4 Hz, 2H), 3.64–3.51 (m, 25H), 3.50–3.35 (m, 2H), 3.04 (d, J = 7.0 Hz, 2H), 2.61 (s, 3H), 2.42 (s, 3H), 2.03 (s, 3H), 1.63 (s, 3H), proton on enaminic NH exchanges with the solvent. 13C NMR (101 MHz, MeOD) δ 205.51, 201.63, 198.54, 189.78, 172.44, 167.61, 163.90, 156.59, 155.93, 131.22, 126.16, 112.08, 109.31, 103.54, 101.85, 100.93, 99.71, 70.20, 70.14, 70.05, 69.99, 69.85, 68.89, 66.23, 59.24, 56.98, 50.73, 39.21, 34.31, 31.35, 30.87, 30.03, 29.26, 22.30, 17.61, 13.02, 6.20. ESI-MS (m/z): [M + Na]+ calcd for C42H54N2NaO16+, 865.34; found 865.39 and [M–H]− calcd for C42H53N2O16−, 841.34; found 841.44. HPLC-UV/vis: k = 2.11, [α]20D +42.30° (c 0.2, CHCl3).
1 + 0.1% NH3 (in MeOH). Yellow solid (56% yield). 1H NMR (400 MHz, CDCl3) δ 13.83 (s, 1H), 13.30 (s, 1H), 11.89 (s, 1H), 7.03 (d, J = 8.1 Hz, 2H), 6.76 (d, J = 8.1 Hz, 2H), 6.55 (s, 1H), 5.75 (s, 1H), 4.38 (d, J = 7.1 Hz, 1H), 3.67 (t, J = 6.4 Hz, 2H), 3.64–3.51 (m, 25H), 3.50–3.35 (m, 2H), 3.04 (d, J = 7.0 Hz, 2H), 2.61 (s, 3H), 2.42 (s, 3H), 2.03 (s, 3H), 1.63 (s, 3H), proton on enaminic NH exchanges with the solvent. 13C NMR (101 MHz, MeOD) δ 205.51, 201.63, 198.54, 189.78, 172.44, 167.61, 163.90, 156.59, 155.93, 131.22, 126.16, 112.08, 109.31, 103.54, 101.85, 100.93, 99.71, 70.20, 70.14, 70.05, 69.99, 69.85, 68.89, 66.23, 59.24, 56.98, 50.73, 39.21, 34.31, 31.35, 30.87, 30.03, 29.26, 22.30, 17.61, 13.02, 6.20. ESI-MS (m/z): [M + Na]+ calcd for C42H54N2NaO16+, 865.34; found 865.39 and [M–H]− calcd for C42H53N2O16−, 841.34; found 841.44. HPLC-UV/vis: k = 2.11, [α]20D +42.30° (c 0.2, CHCl3).
Methyl (S,E)-2-((S)-6-acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2(1H)-ylidene)-4-(4-hydroxybenzyl)-5-oxo-9,12,15,18,21,24-hexaoxa-3,6-diazahexacosan-26-oate ((9bS,15S)-7). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 + 0.1% NH3 (in MeOH). Yellow solid (49% yield). 1H NMR (400 MHz, MeOD) δ 7.09 (d, J = 8.5 Hz, 2H), 6.73 (d, J = 8.5 Hz, 2H), 5.81 (s, 1H), 4.72 (dd, J = 8.5, 5.6 Hz, 1H), 4.01 (s, 2H), 3.71 (t, J = 6.2 Hz, 2H), 3.66 (s, 3H), 3.65–3.54 (m, 20H), 3.45–3.36 (m, 2H), 3.18 (dd, J = 13.8, 5.6 Hz, 1H), 3.00 (dd, J = 13.7, 8.6 Hz, 1H), 2.65 (s, 3H), 2.34 (s, 3H), 2.02 (s, 3H), 1.67 (s, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (176 MHz, MeOD) δ 200.90, 198.66, 174.59, 174.12, 173.83, 172.42, 169.94, 163.10, 157.99, 156.60, 156.09, 130.47, 129.16, 129.06, 128.09, 126.07, 115.14, 107.22, 104.98, 101.88, 100.91, 70.13, 69.84, 68.88, 66.28, 59.29, 50.73, 39.20, 38.98, 38.82, 34.31, 33.35, 30.74, 30.03, 29.36, 28.69, 24.15, 23.52, 22.62, 17.44, 10.02, 6.19. ESI-MS (m/z): [M + Na]+ calcd for C42H54N2NaO16+, 865.34; found 865.13 and [M–H]− calcd for C42H53N2O16−, 841.34; found 841.25. HPLC-UV/vis: k = 1.81, [α]20D −48.30° (c 0.2, CHCl3).
1 + 0.1% NH3 (in MeOH). Yellow solid (49% yield). 1H NMR (400 MHz, MeOD) δ 7.09 (d, J = 8.5 Hz, 2H), 6.73 (d, J = 8.5 Hz, 2H), 5.81 (s, 1H), 4.72 (dd, J = 8.5, 5.6 Hz, 1H), 4.01 (s, 2H), 3.71 (t, J = 6.2 Hz, 2H), 3.66 (s, 3H), 3.65–3.54 (m, 20H), 3.45–3.36 (m, 2H), 3.18 (dd, J = 13.8, 5.6 Hz, 1H), 3.00 (dd, J = 13.7, 8.6 Hz, 1H), 2.65 (s, 3H), 2.34 (s, 3H), 2.02 (s, 3H), 1.67 (s, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (176 MHz, MeOD) δ 200.90, 198.66, 174.59, 174.12, 173.83, 172.42, 169.94, 163.10, 157.99, 156.60, 156.09, 130.47, 129.16, 129.06, 128.09, 126.07, 115.14, 107.22, 104.98, 101.88, 100.91, 70.13, 69.84, 68.88, 66.28, 59.29, 50.73, 39.20, 38.98, 38.82, 34.31, 33.35, 30.74, 30.03, 29.36, 28.69, 24.15, 23.52, 22.62, 17.44, 10.02, 6.19. ESI-MS (m/z): [M + Na]+ calcd for C42H54N2NaO16+, 865.34; found 865.13 and [M–H]− calcd for C42H53N2O16−, 841.34; found 841.25. HPLC-UV/vis: k = 1.81, [α]20D −48.30° (c 0.2, CHCl3).
Methyl (S,E)-2-((R)-6-acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2(1H)-ylidene)-4-(4-hydroxybenzyl)-5,10-dioxo-12,15,18-trioxa-3,6,9-triazaicosan-20-oate ((9bR,15S)-8). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 + 0.1% NH3 (in MeOH). Yellow solid (10% yield). 1H NMR (400 MHz, CDCl3) δ 13.82 (s, 1H), 13.37 (s, 1H), 11.86 (s, 1H), 7.06 (d, J = 7.6 Hz, 2H), 6.81 (d, J = 7.6 Hz, 2H), 5.82 (s, 1H), 4.45 (s, 1H), 4.17 (s, 2H), 3.85–3.65 (m, 13H), 3.46–3.30 (m, 4H), 3.03–2.98 (m, 2H), 2.68 (s, 3H), 2.49 (s, 3H), 2.10 (s, 3H), 1.71 (s, 3H), proton on enaminic NH exchanges with the solvent. 13C NMR (101 MHz, CDCl3) δ 200.86, 198.62, 172.24, 171.04, 171.00, 169.48169.38, 167.98, 167.54, 163.61, 158.34, 156.05, 131.02, 130.78, 128.94, 126.53, 115.93, 108.09, 105.23, 101.51, 60.23, 52.16, 52.00, 41.80, 39.04, 38.87, 38.41, 32.01, 30.50, 29.06, 23.88, 23.11, 18.86, 14.18, 11.09, 7.62. ESI-MS (m/z): [M + Na]+ calcd for C38H45N3NaO14+, 790.28; found 790.07 and [M–H]− calcd for C38H44N3O14−, 766.28; found 766.36. HPLC-UV/vis: k = 1.65, [α]20D +172.69° (c 0.3, CHCl3).
1 + 0.1% NH3 (in MeOH). Yellow solid (10% yield). 1H NMR (400 MHz, CDCl3) δ 13.82 (s, 1H), 13.37 (s, 1H), 11.86 (s, 1H), 7.06 (d, J = 7.6 Hz, 2H), 6.81 (d, J = 7.6 Hz, 2H), 5.82 (s, 1H), 4.45 (s, 1H), 4.17 (s, 2H), 3.85–3.65 (m, 13H), 3.46–3.30 (m, 4H), 3.03–2.98 (m, 2H), 2.68 (s, 3H), 2.49 (s, 3H), 2.10 (s, 3H), 1.71 (s, 3H), proton on enaminic NH exchanges with the solvent. 13C NMR (101 MHz, CDCl3) δ 200.86, 198.62, 172.24, 171.04, 171.00, 169.48169.38, 167.98, 167.54, 163.61, 158.34, 156.05, 131.02, 130.78, 128.94, 126.53, 115.93, 108.09, 105.23, 101.51, 60.23, 52.16, 52.00, 41.80, 39.04, 38.87, 38.41, 32.01, 30.50, 29.06, 23.88, 23.11, 18.86, 14.18, 11.09, 7.62. ESI-MS (m/z): [M + Na]+ calcd for C38H45N3NaO14+, 790.28; found 790.07 and [M–H]− calcd for C38H44N3O14−, 766.28; found 766.36. HPLC-UV/vis: k = 1.65, [α]20D +172.69° (c 0.3, CHCl3).
Methyl (S,E)-2-((S)-6-acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2(1H)-ylidene)-4-(4-hydroxybenzyl)-5,10-dioxo-12,15,18-trioxa-3,6,9-triazaicosan-20-oate ((9bS,15S)-8). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 + 0.1% NH3 (in MeOH). Yellow solid (10% yield). 1H NMR (400 MHz, MeOD) δ 7.09 (d, J = 8.5 Hz, 2H), 6.74 (d, J = 8.5 Hz, 2H), 5.79 (s, 1H), 4.70 (dd, J = 8.8, 5.3 Hz, 1H), 4.15 (s, 2H), 3.98 (s, 2H), 3.73 (s, 3H), 3.71–3.63 (m, 8H), 3.41–3.37 (m, 4H), 3.21 (dd, J = 13.8, 5.2 Hz, 1H), 3.00 (dd, J = 13.8, 8.8 Hz, 1H), 2.65 (s, 3H), 2.31 (s, 3H), 2.02 (s, 3H), 1.67 (s, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (101 MHz, MeOD) δ 200.84, 198.68, 189.94, 174.54, 174.02, 171.94, 171.34, 170.27, 163.11, 157.92, 156.60, 156.01, 130.45, 126.13, 115.13, 107.22, 104.91, 102.19, 101.85, 100.87, 70.57, 70.46, 70.11, 69.97, 69.75, 67.69, 59.42, 57.10, 50.86, 39.01, 38.75, 38.22, 30.78, 30.03, 17.39, 6.21. ESI-MS (m/z): [M + Na]+ calcd for C38H45N3NaO14+, 790.28; found 790.41 and [M–H]− calcd for C38H44N3O14−, 766.28; found 766.62. HPLC-UV/vis: k = 1.39, [α]20D −180.85° (c 0.3, CHCl).
1 + 0.1% NH3 (in MeOH). Yellow solid (10% yield). 1H NMR (400 MHz, MeOD) δ 7.09 (d, J = 8.5 Hz, 2H), 6.74 (d, J = 8.5 Hz, 2H), 5.79 (s, 1H), 4.70 (dd, J = 8.8, 5.3 Hz, 1H), 4.15 (s, 2H), 3.98 (s, 2H), 3.73 (s, 3H), 3.71–3.63 (m, 8H), 3.41–3.37 (m, 4H), 3.21 (dd, J = 13.8, 5.2 Hz, 1H), 3.00 (dd, J = 13.8, 8.8 Hz, 1H), 2.65 (s, 3H), 2.31 (s, 3H), 2.02 (s, 3H), 1.67 (s, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (101 MHz, MeOD) δ 200.84, 198.68, 189.94, 174.54, 174.02, 171.94, 171.34, 170.27, 163.11, 157.92, 156.60, 156.01, 130.45, 126.13, 115.13, 107.22, 104.91, 102.19, 101.85, 100.87, 70.57, 70.46, 70.11, 69.97, 69.75, 67.69, 59.42, 57.10, 50.86, 39.01, 38.75, 38.22, 30.78, 30.03, 17.39, 6.21. ESI-MS (m/z): [M + Na]+ calcd for C38H45N3NaO14+, 790.28; found 790.41 and [M–H]− calcd for C38H44N3O14−, 766.28; found 766.62. HPLC-UV/vis: k = 1.39, [α]20D −180.85° (c 0.3, CHCl).
(S)-2-(((E)-1-((R)-6-Acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2(1H)-ylidene)ethyl)amino)-N-(3-(hexadecyloxy)propyl)-3-(4-hydroxyphenyl)propanamide ((9bR,15S)-9). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 + 0.1% NH3 (in MeOH). Colourless oil (25% yield). 1H NMR (400 MHz, CDCl3) δ 13.87 (s, 1H), 13.37 (s, 1H), 11.81 (s, 1H), 7.10 (d, J = 7.8 Hz, 2H), 6.80 (d, J = 7.6 Hz, 2H), 6.50 (s, 1H), 5.84 (s, 1H), 4.38 (s, 1H), 3.48 (t, J = 5.3 Hz, 2H), 3.40–3.32 (m, 4H), 3.27–3.20 (m, 1H), 3.12–3.06 (m, 1H), 2.69 (s, 3H), 2.43 (s, 3H), 2.11 (s, 3H), 1.82–1.72 (m, 5H), 1.37–1.20 (m, 28H), 0.90 (t, J = 5.3 Hz, 3H), proton on enaminic NH exchanges with the solvent. 13C NMR (101 MHz, CDCl3) δ 200.83, 198.72, 168.51, 163.70, 158.29, 155.96, 155.55, 132.84, 131.04, 130.74, 128.95, 127.01, 116.01, 108.29, 104.57, 101.54, 71.58, 70.30, 68.33, 60.76, 39.40, 39.05, 38.88, 32.07, 31.43, 30.51, 29.85, 29.80, 29.64, 29.50, 29.07, 28.75, 26.28, 23.89, 23.13, 22.83, 18.88, 14.26, 14.19, 11.10, 7.63, 1.16. ESI-MS (m/z): [M + Na]+ calcd for C46H64N2NaO9, 811.45; found 811.94 and [M–H]− calcd for C46H63N2O9, 787.45; found 787.12. HPLC-UV/vis: k = 5.40, [α]20D +116.93° (c 0.3, CHCl3).
1 + 0.1% NH3 (in MeOH). Colourless oil (25% yield). 1H NMR (400 MHz, CDCl3) δ 13.87 (s, 1H), 13.37 (s, 1H), 11.81 (s, 1H), 7.10 (d, J = 7.8 Hz, 2H), 6.80 (d, J = 7.6 Hz, 2H), 6.50 (s, 1H), 5.84 (s, 1H), 4.38 (s, 1H), 3.48 (t, J = 5.3 Hz, 2H), 3.40–3.32 (m, 4H), 3.27–3.20 (m, 1H), 3.12–3.06 (m, 1H), 2.69 (s, 3H), 2.43 (s, 3H), 2.11 (s, 3H), 1.82–1.72 (m, 5H), 1.37–1.20 (m, 28H), 0.90 (t, J = 5.3 Hz, 3H), proton on enaminic NH exchanges with the solvent. 13C NMR (101 MHz, CDCl3) δ 200.83, 198.72, 168.51, 163.70, 158.29, 155.96, 155.55, 132.84, 131.04, 130.74, 128.95, 127.01, 116.01, 108.29, 104.57, 101.54, 71.58, 70.30, 68.33, 60.76, 39.40, 39.05, 38.88, 32.07, 31.43, 30.51, 29.85, 29.80, 29.64, 29.50, 29.07, 28.75, 26.28, 23.89, 23.13, 22.83, 18.88, 14.26, 14.19, 11.10, 7.63, 1.16. ESI-MS (m/z): [M + Na]+ calcd for C46H64N2NaO9, 811.45; found 811.94 and [M–H]− calcd for C46H63N2O9, 787.45; found 787.12. HPLC-UV/vis: k = 5.40, [α]20D +116.93° (c 0.3, CHCl3).
(S)-2-(((E)-1-((S)-6-Acetyl-7,9-dihydroxy-8,9b-dimethyl-1,3-dioxo-3,9b-dihydrodibenzo[b,d]furan-2(1H)-ylidene)ethyl)amino)-N-(3-(hexadecyloxy)propyl)-3-(4-hydroxyphenyl)propanamide ((9bS,15S)-9). The title compound was purified over silica gel, mobile phase DCM/MeOH, 9
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) :
:![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 1 + 0.1% NH3 (7 M in MeOH). White solid (11% yield). 1H NMR (400 MHz, MeOD) δ 7.10 (d, J = 8.4 Hz, 2H), 6.75 (d, J = 8.3 Hz, 2H), 5.84 (s, 1H), 4.69 (t, J = 7.2 Hz, 1H), 3.41–3.38 (m, 6H, under MeOD), 3.22–3.13 (m, 1H), 3.01 (dd, J = 13.8, 8.3 Hz, 1H), 2.68 (s, 3H), 2.36 (s, 3H), 2.05 (s, 3H), 1.76–1.70 (m, 5H), 1.33–1.24 (m, 28H), 0.97 (t, J = 7.5 Hz, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (100 MHz, CDCl3) δ 200.75, 190.65, 179.76, 175.24, 168.74, 167.77, 163.75, 158.29, 156.20, 155.43, 132.61, 131.03, 130.87, 128.95, 115.98, 107.83, 104.97, 101.53, 71.64, 70.44, 68.32, 60.93, 39.54, 38.88, 32.07, 31.43, 30.51, 29.84, 29.81, 29.78, 29.66, 29.51, 29.08, 28.75, 26.24, 23.90, 23.13, 22.83, 18.99, 14.26, 14.19, 11.11, 7.63. ESI-MS (m/z): [M + Na]+ calcd for C46H64N2NaO9, 811.45; found 812.03 and [M–H]− calcd for C46H63N2O9, 787.45; found 787.21. HPLC-UV/vis: k = 4.69, [α]20D −163.32° (c 0.25, CHCl3).
1 + 0.1% NH3 (7 M in MeOH). White solid (11% yield). 1H NMR (400 MHz, MeOD) δ 7.10 (d, J = 8.4 Hz, 2H), 6.75 (d, J = 8.3 Hz, 2H), 5.84 (s, 1H), 4.69 (t, J = 7.2 Hz, 1H), 3.41–3.38 (m, 6H, under MeOD), 3.22–3.13 (m, 1H), 3.01 (dd, J = 13.8, 8.3 Hz, 1H), 2.68 (s, 3H), 2.36 (s, 3H), 2.05 (s, 3H), 1.76–1.70 (m, 5H), 1.33–1.24 (m, 28H), 0.97 (t, J = 7.5 Hz, 3H), all the protons on heteroatoms exchange with the solvent. 13C NMR (100 MHz, CDCl3) δ 200.75, 190.65, 179.76, 175.24, 168.74, 167.77, 163.75, 158.29, 156.20, 155.43, 132.61, 131.03, 130.87, 128.95, 115.98, 107.83, 104.97, 101.53, 71.64, 70.44, 68.32, 60.93, 39.54, 38.88, 32.07, 31.43, 30.51, 29.84, 29.81, 29.78, 29.66, 29.51, 29.08, 28.75, 26.24, 23.90, 23.13, 22.83, 18.99, 14.26, 14.19, 11.11, 7.63. ESI-MS (m/z): [M + Na]+ calcd for C46H64N2NaO9, 811.45; found 812.03 and [M–H]− calcd for C46H63N2O9, 787.45; found 787.21. HPLC-UV/vis: k = 4.69, [α]20D −163.32° (c 0.25, CHCl3).
4.4 Water solubility determination
Thermodynamic aqueous solubility was determined following the previously reported procedure.59 Excess solid (about 10–20 mg) of each compound was suspended in 5 mL of ultrapure water for HPLC in a volumetric bottle and magnetically stirred (1500 rpm) for 24 hours at 25 ± 1 °C.The resulting suspension was filtered on a Nylon Syringe Filters (13 mm, 0.45 μm pore size) and the concentration of the compound in solution was quantified by HPLC-UV/PDA on a Jasco (Tokyo, Japan) system, consisting of a PU-1580 pump and a MD-1510 photodiode array (PDA) detector. Chromatogram acquisitions and elaborations were performed using the ChromNAV software (Tokyo, Japan). Analyses were carried out on a XBridge™ Phenyl, (4.6 × 150 mm, 5 μm) column, at room temperature. The mobile phases were A: water containing 0.1% of formic acid, and B: acetonitrile containing 0.1% of formic acid. Elution was performed on a linear gradient from 50% to 100% B over 10 min, followed by an isocratic hold at 100% B for 3 min. The flow rate was 1.0 mL min−1 and the injection volume 10 μL. Standard calibration curves of (+)-(R)-UA (from 0.073 mM to 2.904 mM, R2 = 0.998) (Fig. S3†) was determined. Since all the derivatives share the same chromophore, the same response factor (at λ = 308 nm, the relative maximum absorption peak) was applied. Limits of detections (LOD) and quantification (LOQ) were 0.54 μM and LOQ 61.13 μM, respectively. Solubility was determined from the mean peak areas of duplicate injections.
4.5 In vitro assay
The antifungal activity was evaluated by microdilution method using 96 microwell plates (Microtiter®), according to Clinical and Laboratory Standards Institute (CLSI; formerly NCCLS) procedures, which considers this method the best for antifungal susceptibility testing of yeasts and filamentous fungi.61,62
Both C. albicans, C. tropicalis and T. rubrum were cultured on Sabouraud dextrose agar (SDA) (Merck KGaA, Darmstadt, Germany) at 27 °C for one week before microdilution tests. Dermatophytes suspension was performed by collecting actively growing mycelium in test tubes containing broken coverslips and 10 mL of sterile distillate water, while yeasts suspension was made resuspended the plate-grown cells in sterile distillate water to obtain the starting fungal inoculum of 1.0 × 107 CFU mL−1.
All tested substances were dissolved in 5% dimethyl sulfoxide (DMSO) aqueous solution and tested ranging from 2.0 × 10−1 to 2.4 × 10−5 mg mL−1. Twofold serial broth dilution method in SDA was performed. All microwell plates were incubated at 30 °C and visually evaluated after 24 and 48 hours for Candida strains or 7 days T. rubrum.
All experiments were conducted in triplicate, and solvent blanks were included. Amphotericin B and fluconazole reference antimycotic compound were used.
![[thin space (1/6-em)]](https://www.rsc.org/images/entities/char_2009.gif) 000 cells per cm2) and after 24 h, medium was removed, cells were washed with phosphate buffer solution (PBS, Sigma Aldrich, Italy) and samples were added (200 μl). Test compounds were prepared as 10 mM stock solutions in DMSO and diluted with complete culture medium (CM). Aliquots of the working solutions were transferred to 96-well plates to afford final concentrations of 125 μM and 50 μM. Cells were exposed to the compounds for 24 h at 37 °C. Vehicle (0.5% DMSO) was included on every plate.
000 cells per cm2) and after 24 h, medium was removed, cells were washed with phosphate buffer solution (PBS, Sigma Aldrich, Italy) and samples were added (200 μl). Test compounds were prepared as 10 mM stock solutions in DMSO and diluted with complete culture medium (CM). Aliquots of the working solutions were transferred to 96-well plates to afford final concentrations of 125 μM and 50 μM. Cells were exposed to the compounds for 24 h at 37 °C. Vehicle (0.5% DMSO) was included on every plate.
To assess cell viability, Alamar blue (ThermoFisher Scientific, Italy) assay was performed. The medium was removed and 100 μL of a 10% v/v solution of Alamar blue in DMEM was added to each well and left in contact for 3 h. After 3 h, fluorescence was detected by means of a multi-mode microplate reader (FLUOstar Omega Microplate Reader, BMG LabTech, Ortenberg, G) at two different wavelengths: at 570 nm to detect the reduced form (red) of the Alamar blue, and 655 nm, to detect the oxidized one (blue). Results were expressed as cell viability% by normalizing the fluorescence measured after contact with each sample with that measured for CM, used as reference. Six replicates were performed for each sample/control.
Abbreviations
| AmB | Amphotericin B |
| DCM | Dichloromethane |
| DIPEA | Diisopropylethylamine |
| DMF | N,N-Dimethylformamide |
| DMSO | Dimethyl sulfoxide |
| EDC | 1-Ethyl-3-(3-dimethylaminopropyl)carbodiimide |
| ESI-MS | Electrospray ionization-mass spectrometry |
| EtOAc | Ethyl acetate |
| EtOH | Ethanol |
| FCZ | Fluconazole |
| FIs | Fungal infections |
| HOBt | Hydroxy-benzotriazole |
| ACN | Acetonitrile |
| MeOH | Methanol |
| r.t. | Room temperature |
| SAR | Structure–activity relationship |
| TEA | Triethylamine |
| TFA | Trifluoroacetic acid |
| TMSCl | Trimethylsilyl chloride |
| UA | Usnic acid |
Data availability
All relevant data are within the manuscript and its ESI† files.Author contributions
A. F. and A. C.: synthesis of compounds; G. R.: assistance in synthesis; R. L.: solubility studies; C. V.: cell viability studies; V. C.: extraction of UA; B. V.: supervising cell viability studies; M. E. E. T.: antifungal studies; S. T.: supervising antifungal studies; all the authors: writing – original draft; G. R. and P. L.: validation, writing and editing; S. C. and E. M.: validation, writing, review and editing, supervision, funding acquisition.Conflicts of interest
There is no conflict of interest to declare.Acknowledgements
This research was funded by the Italian University Ministry Project “ONE HEALTH BASIC AND TRANSLATIONAL RESEARCH ACTIONS (INF-ACT)” grant number PE13_INFACT_PNRR-U.A. 14.01 to S. C., and by the National Recovery and Resilience Plan (NRRP), Mission 4 Component 2 Investment 1.4 – Call for tender No. 3138 of 16 December 2021, rectified by Decree No. 3175 of 18 December 2021 of Italian Ministry of University and Research funded by the European Union – NextGenerationEU; Award Number: Project code CN_00000033, Concession Decree No. 1034 of 17 June 2022 adopted by the Italian Ministry of University and Research, Project title “National Biodiversity Future Center – NBFC” to E. M. and S. T. The authors thank INFACT for the fellowship to R. L. and NBFC for the fellowship to V. C. and to M. E. E. T. The authors gratefully acknowledge Centro Grandi Strumenti of University of Pavia for the NMR Bruker Avance 400 and 700 MHz spectrometer facilities and for the support and assistance in this work.Notes and references
- S. Sedik, S. Wolfgruber, M. Hoenigl and L. Kriegl, Diagnosing Fungal Infections in Clinical Practice: A Narrative Review, Expert Rev. Anti-infect. Ther., 2024, 22(11), 935–949, DOI:10.1080/14787210.2024.2403017.
- S. Dellière, A. Jabet and A. Abdolrasouli, Current and Emerging Issues in Dermatophyte Infections, PLoS Pathog., 2024, 20(6), e1012258, DOI:10.1371/journal.ppat.1012258.
- W. Fang, J. Wu, M. Cheng, X. Zhu, M. Du, C. Chen, W. Liao, K. Zhi and W. Pan, Diagnosis of Invasive Fungal Infections: Challenges and Recent Developments, J. Biomed. Sci., 2023, 30(1), 42, DOI:10.1186/s12929-023-00926-2.
- S. Gnat, D. Łagowski, A. Nowakiewicz and M. Dyląg, A Global View on Fungal Infections in Humans and Animals: Opportunistic Infections and Microsporidioses, J. Appl. Microbiol., 2021, 131(5), 2095–2113, DOI:10.1111/jam.15032.
- E. E. Seagle, S. L. Williams and T. M. Chiller, Recent Trends in the Epidemiology of Fungal Infections, Infect. Dis. Clin. North Am., 2021, 35(2), 237–260, DOI:10.1016/j.idc.2021.03.001.
- M. C. Fisher and D. W. Denning, The WHO Fungal Priority Pathogens List as a Game-Changer, Nat. Rev. Microbiol., 2023, 21(4), 211–212, DOI:10.1038/s41579-023-00861-x.
- J. D. Jenks, O. A. Cornely, S. C.-A. Chen, G. R. Thompson and M. Hoenigl, Breakthrough Invasive Fungal Infections: Who Is at Risk?, Mycoses, 2020, 63(10), 1021–1032, DOI:10.1111/myc.13148.
- T. Vanzolini and M. Magnani, Old and New Strategies in Therapy and Diagnosis against Fungal Infections, Appl. Microbiol. Biotechnol., 2024, 108(1), 147, DOI:10.1007/s00253-023-12884-8.
- F. B. Cavassin, J. L. Baú-Carneiro, R. R. Vilas-Boas and F. Queiroz-Telles, Sixty Years of Amphotericin B: An Overview of the Main Antifungal Agent Used to Treat Invasive Fungal Infections, Infect. Dis. Ther., 2021, 10(1), 115–147, DOI:10.1007/s40121-020-00382-7.
- L. Brescini, S. Fioriti, G. Morroni and F. Barchiesi, Antifungal Combinations in Dermatophytes, J. Fungi, 2021, 7(9), 727, DOI:10.3390/jof7090727.
- J. Berman and D. J. Krysan, Drug Resistance and Tolerance in Fungi, Nat. Rev. Microbiol., 2020, 18(6), 319–331, DOI:10.1038/s41579-019-0322-2.
- C. Lass-Flörl, S. S. Kanj, N. P. Govender, G. R. Thompson, L. Ostrosky-Zeichner and M. A. Govrins, Invasive Candidiasis, Nat. Rev. Dis. Primers, 2024, 10(1), 20, DOI:10.1038/s41572-024-00503-3.
- N. Barantsevich and E. Barantsevich, Diagnosis and Treatment of Invasive Candidiasis, Antibiotics, 2022, 11(6), 718, DOI:10.3390/antibiotics11060718.
- F. O. AL-Khikani, Dermatophytosis a Worldwide Contiguous Fungal Infection: Growing Challenge and Few Solutions, Biomed. Biotechnol. Res. J., 2020, 4(2), 117, DOI:10.4103/bbrj.bbrj_1_20.
- A. Logan, A. Wolfe and J. C. Williamson, Antifungal Resistance and the Role of New Therapeutic Agents, Curr. Infect. Dis. Rep., 2022, 24(9), 105–116, DOI:10.1007/s11908-022-00782-5.
- X. Gong, M. Y. Wani, A. S. Al-Bogami, A. Ahmad, K. Robinson and A. Khan, The Road Ahead: Advancing Antifungal Vaccines and Addressing Fungal Infections in the Post-COVID World, ACS Infect. Dis., 2024, 10(10), 3475–3495, DOI:10.1021/acsinfecdis.4c00245.
- R. J. Young, S. L. Flitsch, M. Grigalunas, P. D. Leeson, R. J. Quinn, N. J. Turner and H. Waldmann, The Time and Place for Nature in Drug Discovery, JACS Au, 2022, 2(11), 2400–2416, DOI:10.1021/jacsau.2c00415.
- A. G. Atanasov, S. B. Zotchev, V. M. Dirsch, I. E. Orhan, M. Banach, J. M. Rollinger, D. Barreca, W. Weckwerth, R. Bauer, E. A. Bayer, M. Majeed, A. Bishayee, V. Bochkov, G. K. Bonn, N. Braidy, F. Bucar, A. Cifuentes, G. D'Onofrio, M. Bodkin, M. Diederich, A. T. Dinkova-Kostova, T. Efferth, K. El Bairi, N. Arkells, T.-P. Fan, B. L. Fiebich, M. Freissmuth, M. I. Georgiev, S. Gibbons, K. M. Godfrey, C. W. Gruber, J. Heer, L. A. Huber, E. Ibanez, A. Kijjoa, A. K. Kiss, A. Lu, F. A. Macias, M. J. S. Miller, A. Mocan, R. Müller, F. Nicoletti, G. Perry, V. Pittalà, L. Rastrelli, M. Ristow, G. L. Russo, A. S. Silva, D. Schuster, H. Sheridan, K. Skalicka-Woźniak, L. Skaltsounis, E. Sobarzo-Sánchez, D. S. Bredt, H. Stuppner, A. Sureda, N. T. Tzvetkov, R. A. Vacca, B. B. Aggarwal, M. Battino, F. Giampieri, M. Wink, J.-L. Wolfender, J. Xiao, A. W. K. Yeung, G. Lizard, M. A. Popp, M. Heinrich, I. Berindan-Neagoe, M. Stadler, M. Daglia, R. Verpoorte and C. T. Supuran, Natural Products in Drug Discovery: Advances and Opportunities, Nat. Rev. Drug Discovery, 2021, 20(3), 200–216, DOI:10.1038/s41573-020-00114-z.
- D. J. Newman and G. M. Cragg, Natural Products as Sources of New Drugs over the Nearly Four Decades from 01/1981 to 09/2019, J. Nat. Prod., 2020, 83(3), 770–803, DOI:10.1021/acs.jnatprod.9b01285.
- A. Pasinato and G. Singh, Lichens Are a Treasure Chest of Bioactive Compounds: Fact or Fake?, New Phytol., 2025, 246(2), 389–395, DOI:10.1111/nph.70034.
- O. T. Adenubi, I. M. Famuyide, L. J. McGaw and J. N. Eloff, Lichens: An Update on Their Ethnopharmacological Uses and Potential as Sources of Drug Leads, J. Ethnopharmacol., 2022, 298, 115657, DOI:10.1016/j.jep.2022.115657.
- P. White, R. Oliveira, A. Oliveira, M. Serafini, A. Araújo, D. Gelain, J. Moreira, J. Almeida, J. Quintans, L. Quintans-Junior and M. Santos, Antioxidant Activity and Mechanisms of Action of Natural Compounds Isolated from Lichens: A Systematic Review, Molecules, 2014, 19(9), 14496–14527, DOI:10.3390/molecules190914496.
- H. Wang, M. Xuan, C. Huang and C. Wang, Advances in Research on Bioactivity, Toxicity, Metabolism, and Pharmacokinetics of Usnic Acid In Vitro and In Vivo, Molecules, 2022, 27(21), 7469, DOI:10.3390/molecules27217469.
- M. Xu, E. Oppong-Danquah, X. Wang, S. Oddsson, A. Abdelrahman, S. V. Pedersen, M. Szomek, A. E. Gylfason, B. S. Snorradottir, E. A. Christensen, D. Tasdemir, C. J. Jameson, S. Murad, O. S. Andresson, K. P. Magnusson, H. J. de Boer, M. Thorsteinsdottir, S. Omarsdottir, S. Heidmarsson and E. S. Olafsdottir, Novel Methods to Characterise Spatial Distribution and Enantiomeric Composition of Usnic Acids in Four Icelandic Lichens, Phytochemistry, 2022, 200, 113210, DOI:10.1016/j.phytochem.2022.113210.
- A. Galanty, P. Paśko and I. Podolak, Enantioselective Activity of Usnic Acid: A Comprehensive Review and Future Perspectives, Phytochem. Rev., 2019, 18(2), 527–548, DOI:10.1007/s11101-019-09605-3.
- V. Cavalloro, G. Marrubini, R. Stabile, D. Rossi, P. Linciano, G. Gheza, S. Assini, E. Martino and S. Collina, Microwave-Assisted Extraction and HPLC-UV-CD Determination of (S)-Usnic Acid in Cladonia Foliacea, Molecules, 2021, 26(2), 455, DOI:10.3390/molecules26020455.
- M. Varlı, S. R. Bhosle, E. Kim, Y. Yang, İ. Taş, R. Zhou, S. Pulat, C. D. B. Gamage, S.-Y. Park, H.-H. Ha and H. Kim, Usnic Acid Targets 14-3-3 Proteins and Suppresses Cancer Progression by Blocking Substrate Interaction, JACS Au, 2024, 4(4), 1521–1537, DOI:10.1021/jacsau.3c00774.
- Y. Zhang, Y. Han, Z. Huang, Y. Huang, J. Kong, Y. Sun, J. Cao and T. Zhou, Restoring Colistin Sensitivity and Combating Biofilm Formation: Synergistic Effects of Colistin and Usnic Acid against Colistin-Resistant Enterobacteriaceae, ACS Infect. Dis., 2023, 9(12), 2457–2470, DOI:10.1021/acsinfecdis.3c00315.
- R. Galla, S. Ferrari, S. Ruga, B. Mantuano, G. Rosso, S. Tonello, L. Rosa, P. Valenti and F. Uberti, Effects of Usnic Acid to Prevent Infections by Creating a Protective Barrier in an In Vitro Study, Int. J. Mol. Sci., 2023, 24(4), 3695, DOI:10.3390/ijms24043695.
- A. S. Filimonov, O. I. Yarovaya, A. V. Zaykovskaya, N. B. Rudometova, D. N. Shcherbakov, V. Yu. Chirkova, D. S. Baev, S. S. Borisevich, O. A. Luzina, O. V. Pyankov, R. A. Maksyutov and N. F. Salakhutdinov, (+)-Usnic Acid and Its Derivatives as Inhibitors of a Wide Spectrum of SARS-CoV-2 Viruses, Viruses, 2022, 14(10), 2154, DOI:10.3390/v14102154.
- A. Priya, C. B. M. Kumar, A. Valliammai, A. Selvaraj and S. K. Pandian, Usnic Acid Deteriorates Acidogenicity, Acidurance and Glucose Metabolism of Streptococcus Mutans through Downregulation of Two-Component Signal Transduction Systems, Sci. Rep., 2021, 11(1), 1374, DOI:10.1038/s41598-020-80338-6.
- P. Nithyanand, R. M. Beema Shafreen, S. Muthamil and S. Karutha Pandian, Usnic Acid Inhibits Biofilm Formation and Virulent Morphological Traits of Candida Albicans, Microbiol. Res., 2015, 179, 20–28, DOI:10.1016/j.micres.2015.06.009.
- R. H. Pires, R. Lucarini and M. J. S. Mendes-Giannini, Effect of Usnic Acid on Candida Orthopsilosis and C. Parapsilosis, Antimicrob. Agents Chemother., 2012, 56(1), 595–597, DOI:10.1128/AAC.05348-11.
- O. A. Luzina and N. F. Salakhutdinov, Usnic Acid and Its Derivatives for Pharmaceutical Use: A Patent Review (2000–2017), Expert Opin. Ther. Pat., 2018, 28(6), 477–491, DOI:10.1080/13543776.2018.1472239.
- S. Chen, Z. Ren and L. Guo, Hepatotoxicity of Usnic Acid and Underlying Mechanisms, J. Environ. Sci. Health, Part C: Toxicol. Carcinog., 2025, 43(1), 1–22, DOI:10.1080/26896583.2024.2366737.
- N. Croce, M. Pitaro, V. Gallo and G. Antonini, Toxicity of Usnic Acid: A Narrative Review, J. Toxicol., 2022, 2022, 1–12, DOI:10.1155/2022/8244340.
- H. D. A. de Araújo, H. A. M. F. Silva, J. G. da Silva Júnior, M. C. P. de A. Albuquerque, L. C. B. B. Coelho and A. de L. Aires, The Natural Compound Hydrophobic Usnic Acid and Hydrophilic Potassium Usnate Derivative: Applications and Comparisons, Molecules, 2021, 26(19), 5995, DOI:10.3390/molecules26195995.
- Y. Yang, W. K. Bae, J.-Y. Lee, Y. J. Choi, K. H. Lee, M.-S. Park, Y. H. Yu, S.-Y. Park, R. Zhou, İ. Taş, C. Gamage, M.-J. Paik, J. H. Lee, I. J. Chung, K. K. Kim, J.-S. Hur, S. K. Kim, H.-H. Ha and H. Kim, Potassium Usnate, a Water-Soluble Usnic Acid Salt, Shows Enhanced Bioavailability and Inhibits Invasion and Metastasis in Colorectal Cancer, Sci. Rep., 2018, 8(1), 16234, DOI:10.1038/s41598-018-34709-9.
- T. Kristmundsdóttir, E. Jónsdóttir, H. M. Ögmundsdóttir and K. Ingólfsdóttir, Solubilization of Poorly Soluble Lichen Metabolites for Biological Testing on Cell Lines, Eur. J. Pharm. Sci., 2005, 24(5), 539–543, DOI:10.1016/j.ejps.2005.01.011.
- D. C. S. Macedo, F. J. F. Almeida, M. S. O. Wanderley, M. S. Ferraz, N. P. S. Santos, A. M. Q. López, N. S. Santos-Magalhães and M. C. B. Lira-Nogueira, Usnic Acid: From an Ancient Lichen Derivative to Promising Biological and Nanotechnology Applications, Phytochem. Rev., 2021, 20(3), 609–630, DOI:10.1007/s11101-020-09717-1.
- A. Galanty, J. Popiół, M. Paczkowska-Walendowska, E. Studzińska-Sroka, P. Paśko, J. Cielecka-Piontek, E. Pękala and I. Podolak, (+)-Usnic Acid as a Promising Candidate for a Safe and Stable Topical Photoprotective Agent, Molecules, 2021, 26(17), 5224, DOI:10.3390/molecules26175224.
- A. Zugic, V. Tadic and S. Savic, Nano- and Microcarriers as Drug Delivery Systems for Usnic Acid: Review of Literature, Pharmaceutics, 2020, 12(2), 156, DOI:10.3390/pharmaceutics12020156.
- V. Kartsev, B. Lichitsky, A. Geronikaki, A. Petrou, M. Smiljkovic, M. Kostic, O. Radanovic and M. Soković, Design, Synthesis and Antimicrobial Activity of Usnic Acid Derivatives, MedChemComm, 2018, 9(5), 870–882, 10.1039/C8MD00076J.
- S. P. Kwong, H. Wang, L. Shi, Z. Huang, B. Lu, X. Cheng, G. Chou, L. Ji and C. Wang, Identification of Photodegraded Derivatives of Usnic Acid with Improved Toxicity Profile and UVA/UVB Protection in Normal Human L02 Hepatocytes and Epidermal Melanocytes, J. Photochem. Photobiol., B, 2020, 205, 111814, DOI:10.1016/j.jphotobiol.2020.111814.
- R. U. McVicker and N. M. O'Boyle, Chirality of New Drug Approvals (2013–2022): Trends and Perspectives, J. Med. Chem., 2024, 67(4), 2305–2320, DOI:10.1021/acs.jmedchem.3c02239.
- H. C. de Oliveira, B. T. Bezerra and M. L. Rodrigues, Antifungal Development and the Urgency of Minimizing the Impact of Fungal Diseases on Public Health, ACS Bio Med Chem Au, 2023, 3(2), 137–146, DOI:10.1021/acsbiomedchemau.2c00055.
- R. He, D. Guo, Z. Huang, Y. Kong, C. Ji, J. Gu, Z. Zhang, J. Diao, Z. Zhou, M. Zhao, J. Fan and W. Zhang, Systematic Investigation of Stereochemistry, Stereoselective Bioactivity, and Antifungal Mechanism of Chiral Triazole Fungicide Metconazole, Sci. Total Environ., 2021, 784, 147194, DOI:10.1016/j.scitotenv.2021.147194.
- W. Shi, B. A. Nacev, S. Bhat and J. O. Liu, Impact of Absolute Stereochemistry on the Antiangiogenic and Antifungal Activities of Itraconazole, ACS Med. Chem. Lett., 2010, 1(4), 155–159, DOI:10.1021/ml1000068.
- A. Ahmadi, E. Mohammadnejadi, P. Karami and N. Razzaghi-Asl, Current Status and Structure Activity Relationship of Privileged Azoles as Antifungal Agents (2016–2020), Int. J. Antimicrob. Agents, 2022, 59(3), 106518, DOI:10.1016/j.ijantimicag.2022.106518.
- K. C. Howard, E. K. Dennis, D. S. Watt and S. Garneau-Tsodikova, A Comprehensive Overview of the Medicinal Chemistry of Antifungal Drugs: Perspectives and Promise, Chem. Soc. Rev., 2020, 49(8), 2426–2480, 10.1039/C9CS00556K.
- S. Nishimura and N. Matsumori, Chemical Diversity and Mode of Action of Natural Products Targeting Lipids in the Eukaryotic Cell Membrane, Nat. Prod. Rep., 2020, 37(5), 677–702, 10.1039/C9NP00059C.
- O. A. Luzina and N. F. Salakhutdinov, Biological Activity of Usnic Acid and Its Derivatives: Part 1. Activity against Unicellular Organisms, Russ. J. Bioorg. Chem., 2016, 42(2), 115–132, DOI:10.1134/S1068162016020084.
- T. E. Kornienko, A. A. Chepanova, A. L. Zakharenko, A. S. Filimonov, O. A. Luzina, N. S. Dyrkheeva, V. P. Nikolin, N. A. Popova, N. F. Salakhutdinov and O. I. Lavrik, Enhancement of the Antitumor and Antimetastatic Effect of Topotecan and Normalization of Blood Counts in Mice with Lewis Carcinoma by Tdp1 Inhibitors—New Usnic Acid Derivatives, Int. J. Mol. Sci., 2024, 25(2), 1210, DOI:10.3390/ijms25021210.
- X. Yu, Q. Guo, G. Su, A. Yang, Z. Hu, C. Qu, Z. Wan, R. Li, P. Tu and X. Chai, Usnic Acid Derivatives with Cytotoxic and Antifungal Activities from the Lichen Usnea Longissima, J. Nat. Prod., 2016, 79(5), 1373–1380, DOI:10.1021/acs.jnatprod.6b00109.
- E. Berkow and S. Lockhart, Fluconazole Resistance in Candida Species: A Current Perspective, Infect. Drug Resist., 2017, 10, 237–245, DOI:10.2147/IDR.S118892.
- Z. Rezaei and H. Moghimi, Fungal-Bacterial Consortia: A Promising Strategy for the Removal of Petroleum Hydrocarbons, Ecotoxicol. Environ. Saf., 2024, 280, 116543, DOI:10.1016/j.ecoenv.2024.116543.
- M. K. Calgin and Y. Cetinkol, Distribution and Antifungal Susceptibility Patterns of Candida Species at a University Hospital in Northern Turkey, J. Infect. Dev. Countries, 2018, 12(02), 97–101, DOI:10.3855/jidc.9698.
- P. Keshwania, N. Kaur, J. Chauhan, G. Sharma, O. Afzal, A. S. Alfawaz Altamimi and W. H. Almalki, Superficial Dermatophytosis across the World's Populations: Potential Benefits from Nanocarrier-Based Therapies and Rising Challenges, ACS Omega, 2023, 8(35), 31575–31599, DOI:10.1021/acsomega.3c01988.
- D. Rossi, A. Marra, P. Picconi, M. Serra, L. Catenacci, M. Sorrenti, E. Laurini, M. Fermeglia, S. Pricl, S. Brambilla, N. Almirante, M. Peviani, D. Curti and S. Collina, Identification of RC-33 as a Potent and Selective Σ1 Receptor Agonist Potentiating NGF-Induced Neurite Outgrowth in PC12 Cells. Part 2: G-Scale Synthesis, Physicochemical Characterization and in Vitro Metabolic Stability, Bioorg. Med. Chem., 2013, 21(9), 2577–2586, DOI:10.1016/j.bmc.2013.02.029.
- M. J. O'Neil, The Merck Index - An Encyclopedia of Chemicals, Drugs, and Biologicals, 13th edn, 2006 Search PubMed.
- E. L. Berkow, S. R. Lockhart and L. Ostrosky-Zeichner, Antifungal Susceptibility Testing: Current Approaches, Clin. Microbiol. Rev., 2020, 33(3), e00069-19, DOI:10.1128/CMR.00069-19.
- CLSI Standard M27, Reference Methods for Broth Dilution Antifungal Susceptibility Testing of Yeast, Clinical and Laboratory Standards Institute, Wayne, PA, 4th edn, 2017 Search PubMed.
Footnotes |
| † Electronic supplementary information (ESI) available. See DOI: https://doi.org/10.1039/d5md00457h |
| ‡ Both authors contributed equally. |
| § Present address: Department of Pharmaceutical Sciences, University of Milan, via Mangiagalli 25, 20133, Milan, Italy. |
| This journal is © The Royal Society of Chemistry 2025 |